- 1Department of Pharmacy, The Second Affiliated Hospital of Shantou University Medical College, Shantou, China
- 2Department of Critical Care Medicine, The Second Affiliated Hospital of Shantou University Medical College, Shantou, China
Background: Multidrug-resistant Acinetobacter baumannii (MDR-AB) poses a critical threat in neonatal intensive care units (NICUs), however few studies have reported the molecular characterization of A. baumannii isolates from the NICU in China. This study aims to describe the molecular, biofilm and motility characteristics related to carbapenem resistance genes in A. baumannii isolates from the NICU.
Methods: Following an outbreak in March–April 2024, 21 A. baumannii isolates from neonatal and environmental sources were collected. These isolates underwent antimicrobial susceptibility testing, whole-genome sequencing (WGS) for resistance gene profiling and multi-locus sequence typing (MLST). Biofilm formation and twitching motility were measured in representative strains exposed to subinhibitory concentrations of gentamicin, levofloxacin, and meropenem.
Results: All 21 A. baumannii isolates were carbapenem-resistant Acinetobacter baumannii (CRAB), showing resistance to cephalosporins, aminoglycosides, and quinolones. WGS identified 25 resistance genes, including the universal blaOXA-23, blaOXA-66, blaADC-25and blaTEM-1D, aminoglycoside resistance genes (armA, aph genes), macrolide resistance genes (mphE, msrE), and tetracycline (tetB) resistance genes. Efflux pumps (adeABC, adeFGH, adeIJK) were ubiquitous. Two sequence types emerged: ST195 (20 isolates) and ST1959 (1 isolate), with high genetic similarity suggesting nosocomial transmission. ST1959 exhibited stronger biofilm formation than ST195. Biofilm formation of ST1959 was significantly enhanced by gentamicin but inhibited by levofloxacin. Meropenem suppressed only the ST195 biofilms. Twitching motility was markedly reduced in ST1959 and also decreased after levofloxacin exposure.
Conclusion: This NICU outbreak was driven by CRAB ST195 harboring a conserved resistome. Strain-specific differences in biofilm formation and motility under antibiotic stress highlight the interplay between genetic lineage and phenotypic adaptability. Levofloxacin demonstrates dual anti-biofilm and anti-motility effects, which could guide targeted infection control strategies against high-risk A. baumannii clones.
1 Introduction
Acinetobacter baumannii (AB) is a Gram-negative, non-fermenting bacterium that has been frequently identified in intensive care units(ICUs) in recent years. It predominantly infects critically ill, immunocompromised patients, those undergoing invasive procedures, and individuals receiving broad-spectrum antibiotics. AB is associated with a range of diseases, including meningitis, pneumonia, peritonitis, and bacteremia (Muller et al., 2023). The mortality rate of invasive AB infections remains alarmingly high. Recent data from systematic reviews and meta-analyses indicate that crude mortality rates for carbapenem-resistant A. baumannii (CRAB) infections can range from 30% to 44%, particularly among vulnerable populations such as neonates and immunocompromised individuals (Lee et al., 2017; Tacconelli et al., 2018; Luo et al., 2025). CRAB has shown an alarming upward trend in global epidemiology over the past decade, with significant regional variations. High endemicity is reported in Asia, Europe, and the Americas (Mohd Sazlly Lim et al., 2019; Luo et al., 2024). Notably, China faces a critical challenge with CRAB, where reported resistance rates to carbapenems frequently exceed 60-70% in some regions, establishing it as a priority pathogen for infection control (Wang et al., 2024). A similar concerning trend is observed in neighboring countries, including India, Thailand where the high prevalence of CRAB contributes to substantial public health burdens (Kamolvit et al., 2015; Kumar et al., 2018).
The success of A. baumannii as a hospital-acquired pathogen is largely attributable to its remarkable ability to develop multidrug resistance and persist in the environment. Key virulence factors include biofilm formation, which protects bacterial communities from antibiotics and host immune responses, and surface-associated motility, which facilitates colonization and spread (Jain et al., 2019; Armalyte et al., 2023). These mechanisms confer resistance to desiccation, nutritional stress, and disinfection protocols, thereby contributing to the persistence of endemic clones and sustained hospital outbreaks.
Although frequently reported as a major pathogen in adults, A. baumannii is equally menacing in neonatal intensive care unit (NICU) (Vogiantzi et al., 2024). Risk factors for NICU-acquired AB infections include low birth weight, prolonged NICU stay, umbilical catheterization, central venous catheterization, assisted ventilation, and prior antibiotic exposure (Wei et al., 2015; Phatigomet et al., 2022). Several outbreaks of CRAB in the NICU have been documented in the last decade (Zarrilli et al., 2012; Tsiatsiou et al., 2015; Wei et al., 2015; Ulu-Kilic et al., 2018).
Therapeutic options for CRAB infections remain severely limited, complicating clinical management. The emergence of extensively drug-resistant (XDR) strains has led to the reliance on last-resort agents, such as colistin and tigecycline, often used in combination therapy (Isler et al., 2019). Although such combinations have demonstrated some clinical success, their efficacy is often offset by concerns regarding toxicity (e.g., nephrotoxicity from colistin) and the emergence of resistance. Recent advances with novel beta-lactam/beta-lactamase inhibitor combinations (e.g., cefiderocol, sulbactam-durlobactam) offer promising alternatives, demonstrating enhanced in vitro activity and improved clinical outcomes in certain settings. However, their accessibility and high cost limit widespread use, especially in resource-limited settings (Iovleva et al., 2025). This underscores that infection control and prevention remain the cornerstone of managing CRAB.
Enhancing awareness and establishing robust epidemiological surveillance, including molecular characterization of circulating strains, are fundamental to successful infection control and are particularly imperative in high-risk environments like the NICU. However, despite the significant burden of CRAB, detailed studies on the molecular epidemiology, biofilm formation, and motility characteristics of CRAB isolates specifically from NICUs in South China remain scarce. The present study aims to address this gap by providing a comprehensive molecular, biofilm, and motility characterization of CRAB clinical isolates obtained from a children’s hospital NICU in South China. We report the distribution of carbapenem resistance genes and examine the correlation between resistance profiles and virulence traits, thereby offering valuable insights to inform local infection control strategies and optimize antimicrobial therapy.
2 Materials and methods
2.1 Bacterial strains and culture conditions
A total of twenty-one clinical isolates of A. baumannii investigated in this study were collected during the period of 2024 by medical laboratories of tertiary care hospitals in Shantou city, Guangdong province, China. All isolates were cultured in Luria-Bertani (LB) broth with shaking at 37°C. Antimicrobial susceptibility testing was performed using Mueller-Hinton (MH) broth or agar plates.
2.2 Antimicrobial susceptibility testing
Antimicrobial susceptibility testing was performed using the BD Phoenix-100 Automated Microbiology System (BD Diagnostics, Sparks, MD, USA) with the NMIC/ID-402 panel for Gram-negative bacteria, following the manufacturer’s instructions. Results were interpreted according to the Clinical and Laboratory Standards Institute guidelines (CLSI M100, 2024). The tested antibiotics included those pre-configured in the panel, such as imipenem (10 µg), meropenem (10 µg), amikacin (30 µg), ampicillin-sulbactam (10/10 µg); piperacillin-tazobactam (36 µg); ceftazidime (30 µg); cefotaxime (30 µg); cefepime (30 µg); Aztreonam; CIP: ciprofloxacin (5µg); AMK: amikacin (30 µg); gentamicin (10 µg); Levofloxacin (5 µg); SXT: trimethoprim-sulfamethoxazole (1.25/23.75 µg); colistin (30 µg).
2.3 Whole genome sequencing, genome annotation analysis and visualization
Whole-genome sequencing was performed using the Illumina HiSeq 2500 second-generation high-throughput sequencing platform with a 2×100 bp paired-end configuration. The Illumina-based workflow comprised three primary steps: library preparation from extracted nucleic acids, amplification sequencing, and quality control. Raw sequencing reads in FASTQ format were assembled into contigs (FASTA files) using SPAdes v2.0 software on a Linux/CentOS system. Command-line execution was performed according to the SPAdes (v3.16.1) documentation (https://github.com/ablab/spades).
ResFinder (version 4.6.0) (https://cge.food.dtu.dk/services/ResFinder/) from the Center for Genomic Epidemiology (CGE, DTU) was employed to identify antimicrobial resistance genes in the contigs of 21 CRAB strains. Finally, a heatmap visualizing the distribution of resistance genes across the 21 CRAB strains was generated using MORPHEUS (https://software.broadinstitute.org/morpheus/).
2.4 Multi-locus sequence typing
Multi-locus sequence typing (MLST) was performed in silico on the assembled genomes using the MLST software (v2.25.0, https://github.com/tseemann/mlst) on a Linux system. The software automatically queried the PubMLST database (https://pubmlst.org/organisms/acinetobacter-baumannii) against the Oxford scheme. The seven housekeeping genes (gltA, gyrB, gdhB, recA, cpn60, gpi, and rpoD) were aligned and compared using BLAST algorithms. A unique integer identifier was assigned to each allele of these genes. The combination of these integers (allelic profile) defined the Sequence Type (ST) for each isolate. Whole-genome single nucleotide polymorphism (SNP) analysis was performed using CSI Phylogeny 1.4 from the Center for Genomic Epidemiology (CGE; https://cge.food.dtu.dk/services/CSIPhylogeny/). The complete, high-quality genome of Acinetobacter baumannii ATCC 19606 (GenBank accession: CP045113.1) and ATCC 17978 (GenBank accession: CP053094.1) was used as the reference for the whole-genome alignment. This standard reference strain was selected to ensure consistent and reproducible SNP calling, facilitating reliable phylogenetic inference and cross-study comparisons. A maximum-likelihood phylogenetic tree was constructed from the core genome SNP alignment using FigTree v1.4.4. The genetic relatedness among the 21 A. baumannii strains was assessed based on the resulting tree topology to evaluate their homology and elucidate potential transmission pathways.
2.5 Biofilm formation assays
Biofilm production was evaluated using the crystal violet (CV) staining assay as previously described by O’Toole and Kolter with slight modifications (O'Toole and Kolter, 1998). Briefly, A. baumannii overnight cultures were adjusted to a turbidity equivalent to 0.5 McFarland standard in 0.85% saline solution. For the quantitative assessment of biofilm formation, biofilms were formed in 24-well flat-bottom polystyrene plates. Bacterial suspensions were added to the wells and were statically incubated at 37°C for 48 h. After incubation, the planktonic cells were removed, and the adhered biofilms were gently washed twice with phosphate-buffered saline (PBS), air-dried, and stained with 1 mL of 0.7% (w/v) crystal violet solution per well for 15 min. The excess stain was rinsed off with distilled water, and the bound CV was solubilized with 1 mL of 33% (v/v) glacial acetic acid per well. The optical density (OD) of the solubilized CV was then measured at 600 nm using a Tecan Infinite M200 Pro microplate reader. Additionally, for qualitative visualization and representative imaging of the biofilm structures, the assay was performed in parallel using sterile 15 mL round-bottom polystyrene tubes. The processing, staining, and solubilization steps of the biofilm were identical to those used in the 24-well plates.
The OD values from the 24-well plate assay were corrected for background staining. This was done by subtracting the value for CV bound to uninoculated Müller Hinton Broth (MHB) control wells. E. coli J53 and P. aeruginosa PAO1 were included as the negative and positive controls, respectively, in both setups. Furthermore, all experiments were performed in triplicate and repeated independently on three different days with consistent results.
2.6 Surface-associated motility
The motility assay was conducted using Motility Test Medium, which was inoculated on the surface and incubated overnight at 37°C. This procedure followed the manufacturer’s detailed instructions to ensure accurate and reliable results. Modified LB broth (tryptone-10 g/l; NaCl -5 g/l; yeast extract-5 g/l) with either 0.4 or 0.8% agar was used for all the motility assays. Freshly grown cultures were stabbed to enable spread of bacteria on the surface of the medium (0.4% semisolid) for swarming motility and the interphase between the bottom of the Petri dish and medium (0.8% semisolid) for twitching motility, as described previously (Clemmer et al., 2011). Plates were prepared on the same day as the inoculation. After inoculation, the plates were sealed with parafilm and incubated at 37°C for 48 h.
3 Results
3.1 Clinical information of the neonates
The primary case of neonatal in the NICU was identified on 23rd March, following which up to 15 April 2024, sudden surge of 8 neonates (Total 21 isolates) due to A. baumannii were noticed from NICU (Table 1). A total of 21 strains were identified, comprising of 9 female, 8 male, and 3 of unknown gender. The youngest neonate was 2 hours old and the oldest was 22 days old. The isolates were collected from various sources: nose swabs (6 isolates), bed sheets (4 isolates), ventilator patient ends (2 isolates), endotracheal tube, ETT (4 isolates), hands (3 isolates), condensate water (1 isolate), and sputum (1 isolate).
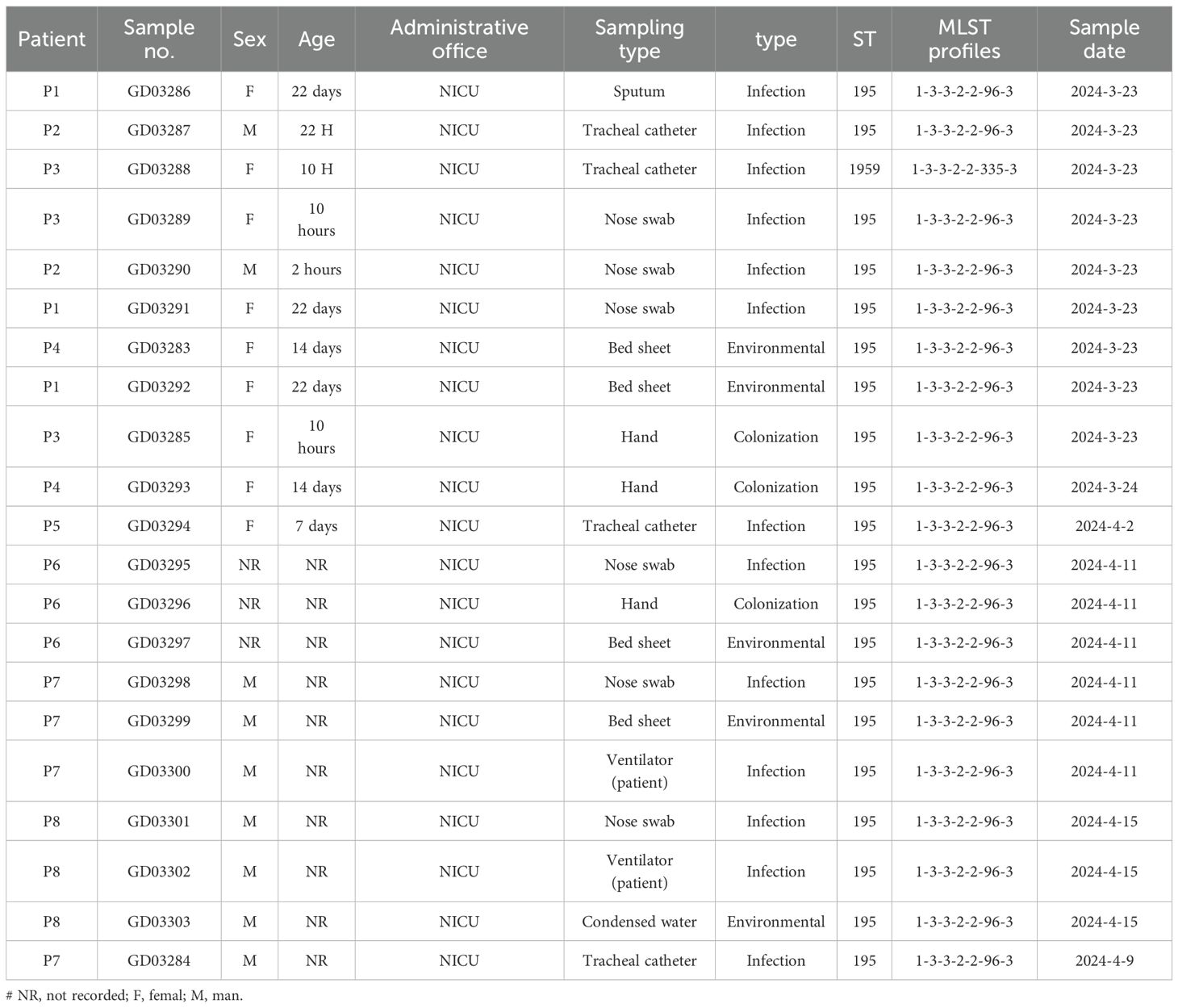
Table 1. Clinical characteristics of the neonates with carbapenem-resistant Acinetobacter baumannii infection.
3.2 Antimicrobial susceptibility testing, MIC determination
All 21 A. baumannii isolates collected in this study were confirmed as CRAB strains based on their resistance to imipenem and meropenem. These isolates exhibited multidrug resistance, as summarized in Table 2, with consistent resistance to piperacillin, piperacillin-tazobactam, ceftazidime, cefepime, and gentamicin. Additionally, they showed varying levels of sensitivity (MIC) to sulfamethoxazole and polymyxin.
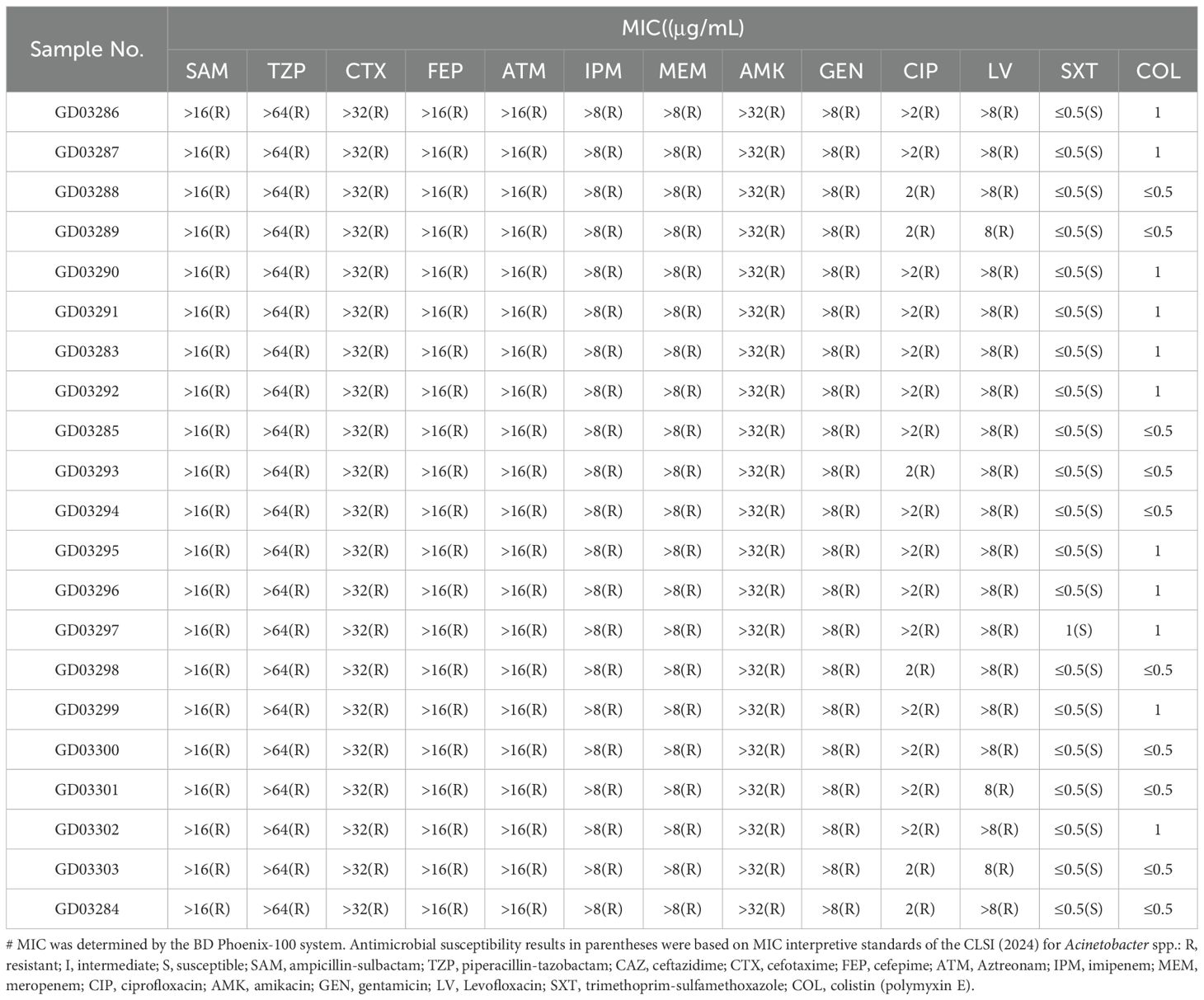
Table 2. Antimicrobial susceptibility profiles of the 21 Acinetobacter baumannii isolates causing infection at the neonatal intensive care unit.
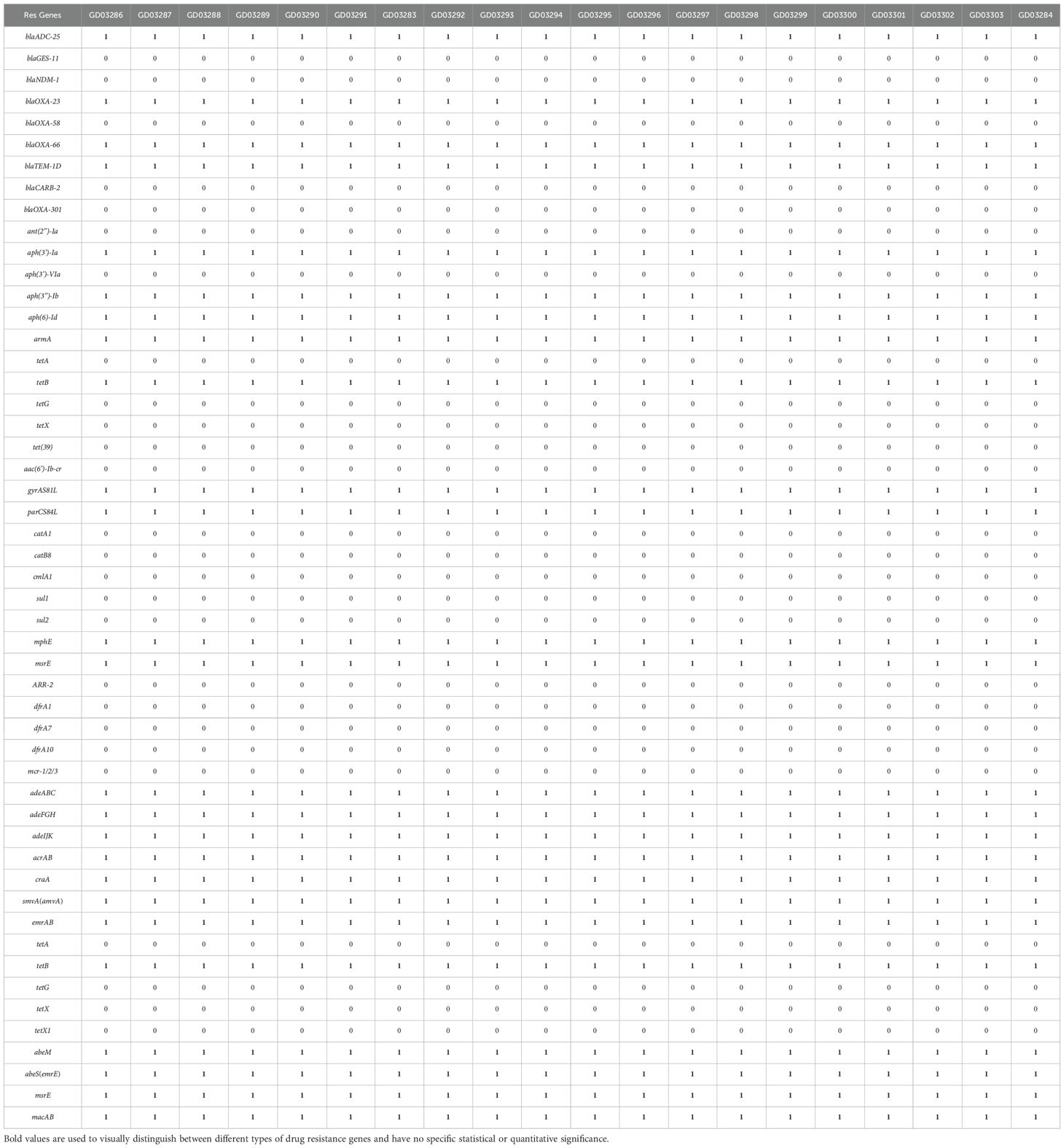
3.3 The distribution of drug-resistant genotypes carried by Acinetobacter baumannii
Analysis of 21 A. baumannii via WGS identified 25 resistance genes, comprising carbapenem resistance gene, β-Lactamase, multiple efflux pump-related, and antibiotic modified enzyme resistance genes (Table 3). All 21 strains (100.0%) had the same four β-lactam resistant genes: blaADC-25, blaOXA-23,blaOXA-66, and blaTEM-1D. Among these, the acquired carbapenemase gene blaOXA-23 and the intrinsic blaOXA-66 are primarily responsible for conferring carbapenem resistance. The blaADC-25 (AmpC) and blaTEM-1D genes contribute to the broader-spectrum resistance to other β-lactams and cephalosporins. Among the six aminoglycoside resistance genes all (100.0%) carried armA, aph (3”) Ib, aph (3 ‘) - Ia, and aph (6) - Id. 100% of the macrolide resistance genes mph (E) and msr (E) were detected, as well as 100% of the gyrAS81L and parCS84L Quinolone Resistance-Determining Region (QRDR) mutations. Additionally, 21 strains carried the tetracycline resistance gene tet (B). Efflux pump-related genes, such as adeABC, adeFGH, adeIJK and acrAB. No sulfonamide resistance gene (sul1), trimethoprim resistance gene (dfrA1, dfrA7, dfrA19), chloramphenicol resistance gene (cat A1, catB8, cmlA1), Rifampicin resistance gene (ARR-2), or Colistin resistance gene (mcr-1/2/3) were found.
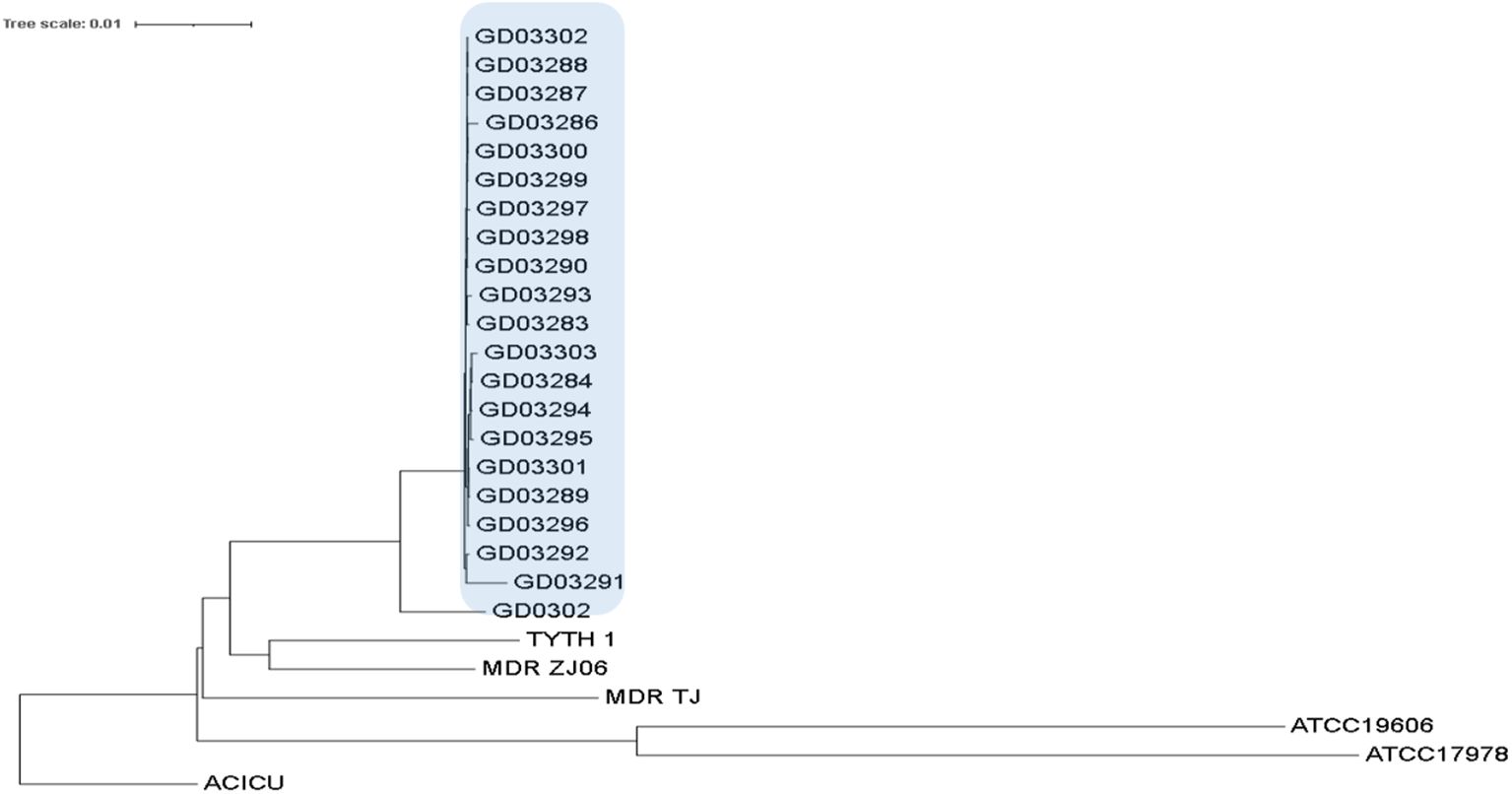
3.4 Phylogenetic tree diagram of 21 strains of Acinetobacter baumannii
The distribution of the isolates has been clearly depicted in Figure 1. Notably, two distinct sequence types (STs) were identified: ST195, which includes GD03286 and several other isolates, and ST1959, which consists only of GD03288. Despite the classification into different STs, the strains exhibited remarkable similarities, with the only significant variation being found in the gpi gene sequence. This close genetic relationship raises concerns regarding the potential for nosocomial transmission, suggesting these strains may have been circulated within healthcare settings, thereby posing a risk for infection among patients.
3.5 Biofilm of the strain GD03286 and GD03288 in the presence or absence of antibiotics
Representative polystyrene tubes displayed a range of biofilm formations, which were subsequently stained with crystal violet following a 24-hour incubation period to visualize the extent of biofilm development. Notably, when compared to the control strain AYE, strain GD03288, which belongs to sequence type ST1959 and was isolated from an endotracheal tube (ETT), exhibited a markedly enhanced capacity for biofilm formation. This was in stark contrast to strain GD03286, classified as ST195 and isolated from sputum samples, as well as other clinical isolates, which demonstrated a comparatively weaker biofilm formation ability, as shown in Figure 2A.
Furthermore, the presence of gentamicin at concentrations of both ½xMIC and MIC significantly augmented the biofilm formation capacity of strain GD03288, as illustrated in Figure 2B. Conversely, levofloxacin, administered at the same concentrations, was shown to have a pronounced inhibitory effect on biofilm formation, as depicted in Figure 2C. Additionally, meropenem, at concentrations of ½xMIC and MIC, effectively inhibited the biofilm formation of strain GD03286; however, it did not exert a significant impact on the biofilm development of strain GD03288 at the ½xMIC concentration, as shown in Figure 2D. These findings underscore the complex interactions between antibiotic treatments and biofilm formation in these bacterial strains, highlighting the potential for strain-specific responses to antibiotic exposure.
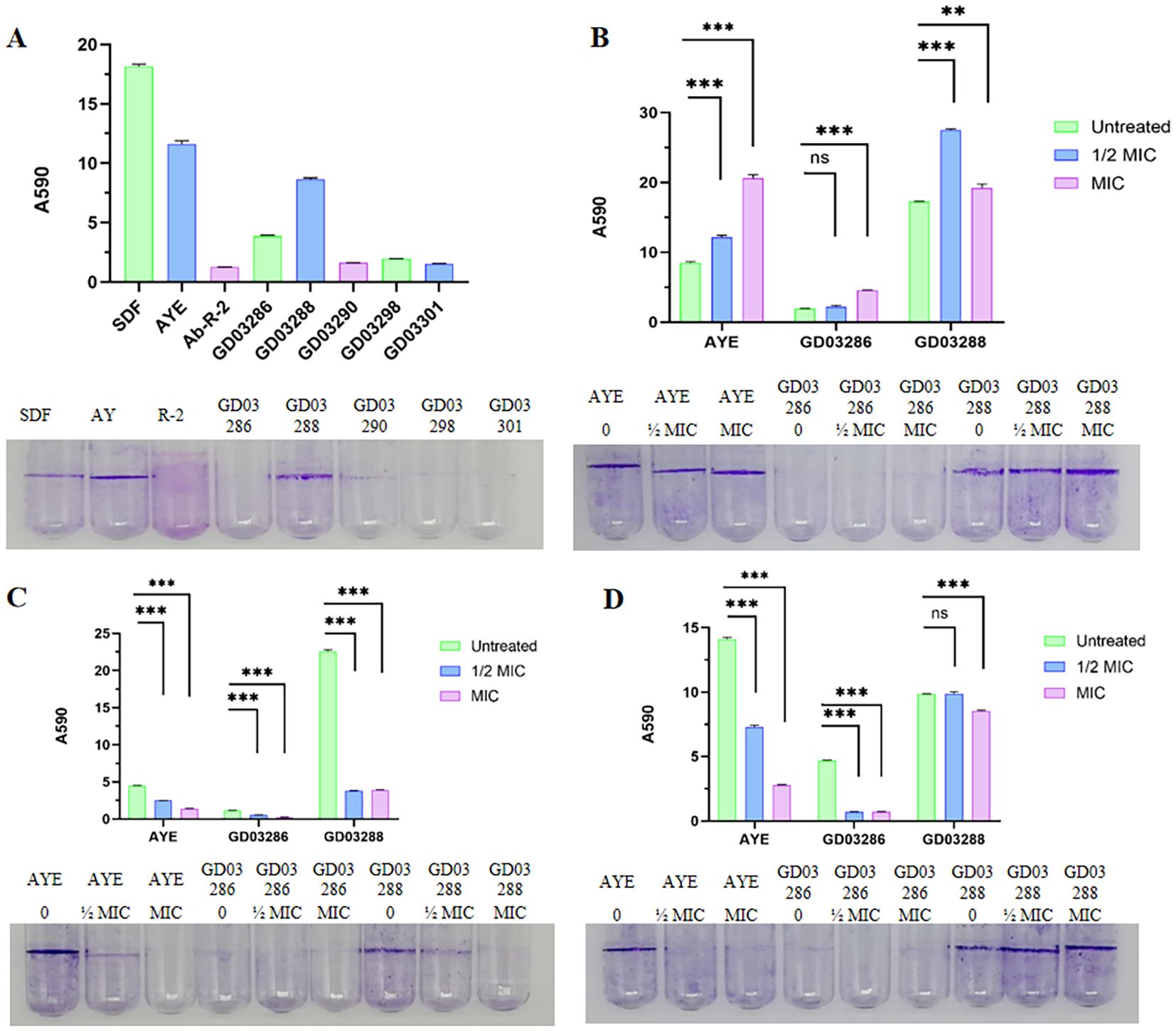
Figure 2. Biofilm of the strain in the presence or absence of antibiotics. (A) OD=1 1% inoculant, 37°C stand still for 48h, no antibiotic; (B) OD=1 1% inoculant, 37°C stand still for 48h, with Gentamicin, ½xMIC=4 μg/mL and MIC = 8 μg/mL; (C) OD = 1 1% inoculant, 37°C stand still for 48h, with Levofloxacin, ½xMIC=4 μg/mL and MIC = 8 μg/mL; (D) OD=1 1% inoculant, 37°C stand still for 48h, with Meropenem, ½xMIC=0.5 μg/mL and MIC = 1 μg/mL. ns: not significant; "**“:P<0.05, "***":P<0.01.
3.6 Motility of the strain GD03286 and GD03288 in the presence or absence of antibiotics
The twitching motility of A. baumannii was assessed qualitatively by visually evaluating the characteristic subsurface hazy growth that developed around the inoculation site, providing a clear indication of the bacterial movement. Notably, the antibiotic levofloxacin(Lv 8 µg/mL) was observed to substantially inhibit this motility pattern (Figure 3), highlighting its potential as a therapeutic agent. Furthermore, the clinical isolate GD03288 (Figure 3C) exhibited a visibly weaker and more constrained motility phenotype compared to reference strains AYE (Figure 3A) and R-2 (Figure 3B), as well as the more motile clinical strain GD03286. This reduced motility in GD03288 may indicate an adaptation or resistance mechanism distinct from other strains.
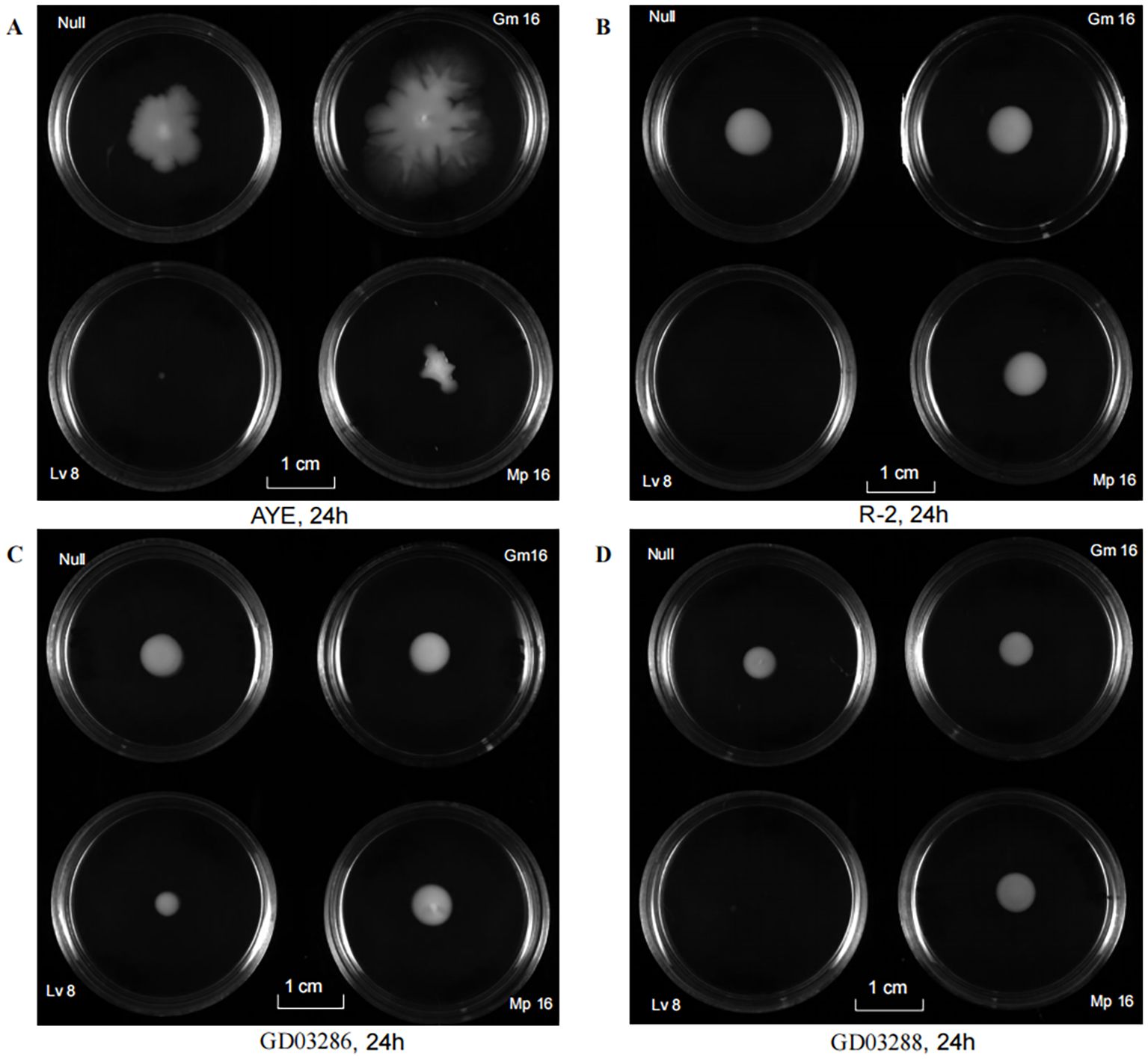
Figure 3. Motility of the strain in the presence or absence of antibiotics. (A) Motility suppression effect against representative strains (AYE) of Gm, Lv and Mp. (B) Motility suppression effect against representative strains (R-2) of Gm, Lv and Mp. (C) Motility suppression effect against representative strains (GD03286) of Gm, Lv and Mp. (D) Motility suppression effect against representative strains (GD03288) of Gm, Lv and Mp. Gm, Gentamicin; Lv, Levofloxacin; Mp, Meropenem.
4 Discussion
This study provides a comprehensive analysis of A. baumannii isolates from a 2024 outbreak in a neonatal intensive care unit (NICU) in South China. The extensive multidrug-resistant (MDR) phenotype observed, which included resistance to carbapenems, cephalosporins, aminoglycosides, and quinolones, was largely explained by the complex resistome uncovered through whole-genome sequencing (WGS).
Resistance to CRAB is complex and involves the upregulation of efflux pumps, reduced expression of membrane porin proteins, alteration of drug targets, and production of carbapenemases (Pannek et al., 2006; Poirel and Nordmann, 2006; Lee et al., 2017). All the A. baumannii isolates in this study were found to harbor carbapenemase resistance genes blaOXA-23 and blaOXA-66, with a detection rate of 100%. blaOXA-23 and blaOXA-66 are both Ambler D-class enzymes, and transmission of blaOXA-23 is usually through plasmids or chromosomes (Mugnier et al., 2010; Ayoub Moubareck and Hammoudi Halat, 2020). ISAba1, an insertion sequence located upstream of blaOXA-23, suggested to have a coordinating effect on it (Royer et al., 2018). blaOXA-23 and blaOXA-66 may be key contributors to CRAB’s resistance to carbapenem antibiotics in this hospital, which is in line with reports from both domestic and international sources (Sung et al., 2016). Recent studies have revealed a high prevalence of the blaOXA-23 gene in Acinetobacter spp., and it is hypothesized that the plasmids pAZJ221 and Tn2009 are likely to be involved in its horizontal transmission (Liu et al., 2015). Supporting this, Hu et al. (2011) reported that all 17 isolates from a NICU outbreak carried the blaOXA-23 gene, underscoring the value of detecting this gene for managing nosocomial infections in neonatology (Hu et al., 2011).
This multidrug-resistant phenotype was further compounded by the universal presence of blaADC-25 (an inherent, often overexpressed AmpC cephalosporinase) and blaTEM-1D (an extended-spectrum β-lactamase), which collectively contribute to high-level resistance to β-lactams. Resistance to aminoglycosides was primarily mediated by the 16S rRNA methyltransferase gene armA, which confers high-level resistance to virtually all aminoglycosides, correlating perfectly with our susceptibility testing results. The concomitant presence of aph genes may further augment this phenotype (Rizk and Abou El-Khier, 2019; Ababneh et al., 2023). Furthermore, the detection of macrolide (mphE, msrE) and tetracycline (tetB) resistance genes explains the resistance to these drug classes. Critically, the ubiquitous presence of three efflux pump systems (adeABC, adeFGH, adeIJK) constitutes a fundamental pillar of resistance. AdeABC, in particular, is a key contributor to resistance against multiple drug classes (including aminoglycosides, fluoroquinolones, and tetracyclines), and its overexpression is closely linked to the challenges in developing efflux pump inhibitors for clinical use (Sharma et al., 2023). The superposition of these multifactorial resistance mechanisms endows this clone with formidable adaptability, posing a severe challenge to clinical management in the NICU.
Molecular typing identified the outbreak was dominated by the internationally disseminated high-risk clone ST195 (20/21), with one isolate belonging to ST1959. ST195, a member of the global clonal complex CC92/CC2, is a predominant strain responsible for recent hospital outbreaks in China, particularly in NICUs (Percival et al., 2015). The high genetic similarity strongly indicates clonal nosocomial transmission, underscoring the urgent need for enhanced infection control measures in this vulnerable clinical setting.
Beyond antimicrobial resistance, bacterial persistence and dissemination capacity are critical virulence determinants, therefore, we investigated two phenotypes critically associated with pathogenicity and environmental adaptability: biofilm formation and twitching motility. A particularly intriguing and potentially clinically significant observation was that isolates recovered from the inner surface of neonatal endotracheal tubes exhibited the strongest biofilm formation and the weakest motility. This finding is highly congruent with existing theory. Biofilms are protective communities formed by bacteria on the surface of medical devices (e.g., ETTs, central venous catheters) and are central to the persistence of device-associated infections, making them notoriously difficult to eradicate (Percival et al., 2015; Arciola et al., 2018; Deshmukh-Reeves et al., 2025). In the specific niche of an ETT, robust biofilm formation is advantageous for resisting antibiotics and host immune defenses, enabling long-term colonization and recurrent infection. Conversely, reduced motility may signify a switch from a “dissemination mode” to a “colonization mode” whereby bacteria reallocate energy resources from the synthesis of motility organelles (e.g., pili) to the production of biofilm matrix and adherence. Our results suggest that within the actual clinical environment, strains residing at critical sites of infection may be selected for this high-biofilm, low-motility phenotype.
Notably, despite being part of the same outbreak, the ST1959 isolate demonstrated significantly stronger biofilm formation than the predominant ST195 clone, indicating that even within a highly clonal population, minor genotypic variations can lead to significant phenotypic divergence, potentially influencing niche adaptation (Penesyan et al., 2021).
Exposure to subinhibitory concentrations of antibiotics significantly modulated these virulence-associated phenotypes. Specifically, gentamicin enhanced biofilm formation in ST1959, a finding consistent with reports that aminoglycosides can induce a bacterial stress response promoting biofilm development (Sato et al., 2018). This phenomenon raises the clinical concern that sublethal exposure during therapy might inadvertently foster bacterial persistence. In contrast, levofloxacin exhibited potent inhibitory effects, suppressing both biofilm formation and markedly reducing motility. This aligns with the known capacity of fluoroquinolones to interfere with key bacterial processes such as quorum sensing and pilus assembly, thereby curbing biofilm formation and motility (Aboelenin et al., 2024; Zhang et al., 2024; Baraka et al., 2025). Recent studies have revealed promising strategies to inhibit A. baumannii biofilm formation, a key contributor to multidrug resistance. Significant efficacy has been demonstrated by essential oils and natural compounds; for instance, cinnamaldehyde, onion ethanolic extract, Tween 80, and ginger essential oil achieved 70.8%–77.3% biofilm inhibition at low concentrations (Yadav et al., 2024). Synthetic approaches, such as N-acyl-2-aminopyrimidine derivatives, have also showed potent activity by disrupting quorum sensing and production of extracellular polysaccharide substances (Jia et al., 2022). Notably, conventional antibiotics like minocycline and polymyxin B were effective in preventing biofilm formation in >90% and 54% of isolates, respectively, at clinically achievable doses (Beganovic et al., 2019). Additionally, macrolides like erythromycin targeted the quorum sensing system (downregulating abaI and abaR genes), leading to biofilm disruption (Dong et al., 2024). Novel agents including cannabidiol (CBD), not only inhibit and eradicate biofilms but also synergiz with conventional antibiotics (e.g., colistin and meropenem), reducing effective concentrations by up to 1000-fold through membrane damage mechanisms (Yosboonruang et al., 2025). Our results suggest that, beyond direct bactericidal activity, levofloxacin may exert an “anti-virulence” effect at sub-MIC concentrations by impairing colonization and dissemination capabilities. This providing a novel rationale for its potential application in combination therapies or decolonization strategies.
5 Conclusion
This NICU outbreak was driven by a CRAB ST195 clone harboring a conserved and complex resistome. The isolates exhibiting the strongest biofilm formation coupled with the poorest motility, which were predominantly recovered from ETTs, likely represent a key adaptive strategy for successful persistence on medical devices. Our research reveals genotypic and phenotypic heterogeneity within a high-risk clone and delineate how subinhibitory antibiotic concentrations can critically modulate these virulence-associated phenotypes. These insights underscore the necessity of integrating both genetic background and phenotypic plasticity in infection control paradigms. The potent dual inhibitory effects of levofloxacin on biofilm formation and motility warrant further mechanistic and clinical exploration, potentially paving the way for novel therapeutic strategies against resilient CRAB infections.
Data availability statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Author contributions
ZY: Investigation, Methodology, Software, Writing – original draft, Writing – review & editing. XH: Data curation, Funding acquisition, Resources, Writing – original draft, Writing – review & editing. HF: Investigation, Software, Supervision, Writing – review & editing. YZ: Formal analysis, Project administration, Writing – review & editing. XL: Visualization, Writing – review & editing. YW: Methodology, Writing – review & editing. DC: Formal analysis, Software, Writing – review & editing. ZK: Writing – review & editing. XW: Writing – review & editing.
Funding
The author(s) declare financial support was received for the research and/or publication of this article. This work was supported by the Medical Science and Technology Research Foundation of Guangdong Province, China under Grant (NO. B2024169); Shantou medical and healthy science and technology program (NO. 221122176494952).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2025.1705282/full#supplementary-material
References
Ababneh, Q., Al Sbei, S., Jaradat, Z., Syaj, S., Aldaken, N., Ababneh, H., et al. (2023). Extensively drug-resistant Acinetobacter baumannii: role of conjugative plasmids in transferring resistance. PeerJ 11, e14709. doi: 10.7717/peerj.14709
Aboelenin, A. M., El-Mowafy, M., Saleh, N. M., Shaaban, M. I., and Barwa, R. (2024). Ciprofloxacin- and levofloxacin-loaded nanoparticles efficiently suppressed fluoroquinolone resistance and biofilm formation in Acinetobacter baumannii. Sci. Rep. 14, 3125. doi: 10.1038/s41598-024-53441-1
Arciola, C. R., Campoccia, D., and Montanaro, L. (2018). Implant infections: adhesion, biofilm formation and immune evasion. Nat. Rev. Microbiol. 16, 397–409. doi: 10.1038/s41579-018-0019-y
Armalyte, J., Cepauskas, A., Sakalyte, G., Martinkus, J., Skerniskyte, J., Martens, C., et al. (2023). A polyamine acetyltransferase regulates the motility and biofilm formation of Acinetobacter baumannii. Nat. Commun. 14, 3531. doi: 10.1038/s41467-023-39316-5
Ayoub Moubareck, C. and Hammoudi Halat, D. (2020). Insights into Acinetobacter baumannii: A Review of Microbiological, Virulence, and Resistance Traits in a Threatening Nosocomial Pathogen. Antibiotics (Basel) 9, 119. doi: 10.3390/antibiotics9030119
Baraka, K., Abozahra, R., Khalaf, E., Bennaya, M. E., and Abdelhamid, S. M. (2025). Repurposing of paroxetine and fluoxetine for their antibacterial effects against clinical Pseudomonas aeruginosa isolates in Egypt. AIMS Microbiol. 11, 126–149. doi: 10.3934/microbiol.2025007
Beganovic, M., Luther, M. K., Daffinee, K. E., and LaPlante, K. L. (2019). Biofilm prevention concentrations (BPC) of minocycline compared to polymyxin B, meropenem, and amikacin against Acinetobacter baumannii. Diagn. Microbiol. Infect. Dis. 94, 223–226. doi: 10.1016/j.diagmicrobio.2019.01.016
Clemmer, K. M., Bonomo, R. A., and Rather, P. N. (2011). Genetic analysis of surface motility in Acinetobacter baumannii. Microbiol. (Reading) 157, 2534–2544. doi: 10.1099/mic.0.049791-0
Deshmukh-Reeves, E., Shaw, M., Bilsby, C., and Gourlay, C. W. (2025). Biofilm formation on endotracheal and tracheostomy tubing: A systematic review and meta-analysis of culture data and sampling method. Microbiologyopen 14, e70032. doi: 10.1002/mbo3.70032
Dong, H., Sun, J., Liu, Y., Li, Q., Huang, J., Xu, P., et al. (2024). Erythromycin disrupts Acinetobacter baumannii biofilms through destruction of the quorum sensing system. Med. (Baltimore) 103, e38341. doi: 10.1097/md.0000000000038341
Hu, Z., Wang, Z., Liu, D., Chen, P., Wang, H., Chen, Y., et al. (2011). Clinical and molecular microbiological characteristics of carbapenem-resistant Acinetobacter baumannii strains in an NICU. Pediatr. Int. 53, 867–872. doi: 10.1111/j.1442-200X.2011.03397.x
Iovleva, A., Fowler, V. G., Jr., and Doi, Y. (2025). Treatment approaches for carbapenem-resistant Acinetobacter baumannii infections. Drugs 85, 21–40. doi: 10.1007/s40265-024-02104-6
Isler, B., Doi, Y., Bonomo, R. A., and Paterson, D. L. (2019). New Treatment Options against Carbapenem-Resistant Acinetobacter baumannii Infections. Antimicrob. Agents Chemother. 63, 01110-18. doi: 10.1128/aac.01110-18
Jain, M., Sharma, A., Sen, M. K., Rani, V., Gaind, R., and Suri, J. C. (2019). Phenotypic and molecular characterization of Acinetobacter baumannii isolates causing lower respiratory infections among ICU patients. Microb. Pathog. 128, 75–81. doi: 10.1016/j.micpath.2018.12.023
Jia, X. M., Cheng, C., Liu, T., Zhao, Y. L., Guo, B., Tang, L., et al. (2022). Synthesis and antibiofilm evaluation of N-acyl-2-aminopyrimidine derivatives against Acinetobacter baumannii. Bioorg Med. Chem. 76, 117095. doi: 10.1016/j.bmc.2022.117095
Kamolvit, W., Sidjabat, H. E., and Paterson, D. L. (2015). Molecular epidemiology and mechanisms of carbapenem resistance of Acinetobacter spp. in Asia and Oceania. Microb. Drug Resist. 21, 424–434. doi: 10.1089/mdr.2014.0234
Kumar, S., Jan, R. A., Fomda, B. A., Rasool, R., Koul, P., Shah, S., et al. (2018). Healthcare-associated pneumonia and hospital-acquired pneumonia: bacterial aetiology, antibiotic resistance and treatment outcomes: A study from north India. Lung 196, 469–479. doi: 10.1007/s00408-018-0117-7
Lee, C. R., Lee, J. H., Park, M., Park, K. S., Bae, I. K., Kim, Y. B., et al. (2017). Biology of Acinetobacter baumannii: pathogenesis, antibiotic resistance mechanisms, and prospective treatment options. Front. Cell Infect. Microbiol. 7. doi: 10.3389/fcimb.2017.00055
Liu, L. L., Ji, S. J., Ruan, Z., Fu, Y., Fu, Y. Q., Wang, Y. F., et al. (2015). Dissemination of blaOXA-23 in Acinetobacter spp. in China: main roles of conjugative plasmid pAZJ221 and transposon Tn2009. Antimicrob. Agents Chemother. 59, 1998–2005. doi: 10.1128/AAC.04574-14
Luo, Q., Chang, M., Lu, P., Guo, Q., Jiang, X., Xiao, T., et al. (2025). Genomic epidemiology and phylodynamics of Acinetobacter baumannii bloodstream isolates in China. Nat. Commun. 16, 3536. doi: 10.1038/s41467-025-58772-9
Luo, Q., Lu, P., Chen, Y., Shen, P., Zheng, B., Ji, J., et al. (2024). ESKAPE in China: epidemiology and characteristics of antibiotic resistance. Emerg. Microbes Infect. 13, 2317915. doi: 10.1080/22221751.2024.2317915
Mohd Sazlly Lim, S., Zainal Abidin, A., Liew, S. M., Roberts, J. A., and Sime, F. B. (2019). The global prevalence of multidrug-resistance among Acinetobacter baumannii causing hospital-acquired and ventilator-associated pneumonia and its associated mortality: A systematic review and meta-analysis. J. Infect. 79, 593–600. doi: 10.1016/j.jinf.2019.09.012
Mugnier, P. D., Poirel, L., Naas, T., and Nordmann, P. (2010). Worldwide dissemination of the blaOXA-23 carbapenemase gene of Acinetobacter baumannii. Emerg. Infect. Dis. 16, 35–40. doi: 10.3201/eid1601.090852
Muller, C., Reuter, S., Wille, J., Xanthopoulou, K., Stefanik, D., Grundmann, H., et al. (2023). A global view on carbapenem-resistant Acinetobacter baumannii. mBio 14, e0226023. doi: 10.1128/mbio.02260-23
O'Toole, G. A. and Kolter, R. (1998). Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: a genetic analysis. Mol. Microbiol. 28, 449–461. doi: 10.1046/j.1365-2958.1998.00797.x
Pannek, S., Higgins, P. G., Steinke, P., Jonas, D., Akova, M., Bohnert, J. A., et al. (2006). Multidrug efflux inhibition in Acinetobacter baumannii: comparison between 1-(1-naphthylmethyl)-piperazine and phenyl-arginine-beta-naphthylamide. J. Antimicrob. Chemother. 57, 970–974. doi: 10.1093/jac/dkl081
Penesyan, A., Paulsen, I. T., Kjelleberg, S., and Gillings, M. R. (2021). Three faces of biofilms: a microbial lifestyle, a nascent multicellular organism, and an incubator for diversity. NPJ Biofilms Microbiomes 7, 80. doi: 10.1038/s41522-021-00251-2
Percival, S. L., Suleman, L., Vuotto, C., and Donelli, G. (2015). Healthcare-associated infections, medical devices and biofilms: risk, tolerance and control. J. Med. Microbiol. 64, 323–334. doi: 10.1099/jmm.0.000032
Phatigomet, M., Thatrimontrichai, A., Maneenil, G., Dissaneevate, S., and Janjindamai, W. (2022). Risk factors for 30-day mortality in neonates with carbapenem-resistant A. baumannii sepsis. Pediatr. Infect. Dis. J. 41, 1012–1016. doi: 10.1097/INF.0000000000003721
Poirel, L. and Nordmann, P. (2006). Carbapenem resistance in Acinetobacter baumannii: mechanisms and epidemiology. Clin. Microbiol. Infect. 12, 826–836. doi: 10.1111/j.1469-0691.2006.01456.x
Rizk, M. A. and Abou El-Khier, N. T. (2019). Aminoglycoside resistance genes in Acinetobacter baumannii clinical isolates. Clin. Lab. 65. doi: 10.7754/Clin.Lab.2019.190103
Royer, S., de Campos, P. A., Araujo, B. F., Ferreira, M. L., Goncalves, I. R., Batistao, D., et al. (2018). Molecular characterization and clonal dynamics of nosocomial blaOXA-23 producing XDR Acinetobacter baumannii. PloS One 13, e0198643. doi: 10.1371/journal.pone.0198643
Sato, Y., Unno, Y., Ubagai, T., and Ono, Y. (2018). Sub-minimum inhibitory concentrations of colistin and polymyxin B promote Acinetobacter baumannii biofilm formation. PloS One 13, e0194556. doi: 10.1371/journal.pone.0194556
Sharma, S., Kaushik, V., Kulshrestha, M., and Tiwari, V. (2023). Different efflux pump systems in Acinetobacter baumannii and their role in multidrug resistance. Adv. Exp. Med. Biol. 1370, 155–168. doi: 10.1007/5584_2023_771
Sung, J. Y., Koo, S. H., Kim, S., and Kwon, G. C. (2016). Persistence of Multidrug-Resistant Acinetobacter baumannii Isolates Harboring blaOXA-23 and bap for 5 Years. J. Microbiol. Biotechnol. 26, 1481–1489. doi: 10.4014/jmb.1604.04049
Tacconelli, E., Carrara, E., Savoldi, A., Harbarth, S., Mendelson, M., Monnet, D. L., et al. (2018). Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327. doi: 10.1016/s1473-3099(17)30753-3
Tsiatsiou, O., Iosifidis, E., Katragkou, A., Dimou, V., Sarafidis, K., Karampatakis, T., et al. (2015). Successful management of an outbreak due to carbapenem-resistant Acinetobacter baumannii in a neonatal intensive care unit. Eur. J. Pediatr. 174, 65–74. doi: 10.1007/s00431-014-2365-8
Ulu-Kilic, A., Gundogdu, A., Cevahir, F., Kilic, H., Gunes, T., and Alp, E. (2018). An outbreak of bloodstream infection due to extensively resistant Acinetobacter baumannii among neonates. Am. J. Infect. Control 46, 154–158. doi: 10.1016/j.ajic.2017.08.007
Vogiantzi, G., Metallinou, D., Tigka, M., Deltsidou, A., and Nanou, C. I. (2024). Bloodstream infections in the neonatal intensive care unit: A systematic review of the literature. Cureus 16, e68057. doi: 10.7759/cureus.68057
Wang, M., Ge, L., Chen, L., Komarow, L., Hanson, B., Reyes, J., et al. (2024). Clinical outcomes and bacterial characteristics of Carbapenem-resistant Acinetobacter baumannii among patients from different global regions. Clin. Infect. Dis. 78, 248–258. doi: 10.1093/cid/ciad556
Wei, H. M., Hsu, Y. L., Lin, H. C., Hsieh, T. H., Yen, T. Y., Lin, H. C., et al. (2015). Multidrug-resistant Acinetobacter baumannii infection among neonates in a neonatal intensive care unit at a medical center in central Taiwan. J. Microbiol. Immunol. Infect. 48, 531–539. doi: 10.1016/j.jmii.2014.08.025
Yadav, P., Shrestha, S., Basyal, D., Tiwari, A., Sah, R., Sah, A. K., et al. (2024). Characterization and biofilm inhibition of multidrug-resistant Acinetobacter baumannii isolates. Int. J. Microbiol. 2024, 5749982. doi: 10.1155/ijm/5749982
Yosboonruang, A., Kiddee, A., Siriphap, A., Pook-In, G., Suwancharoen, C., Duangjai, A., et al. (2025). Potential of Cannabidiol (CBD) to overcome extensively drug-resistant Acinetobacter baumannii. BMC Complement Med. Ther. 25, 308. doi: 10.1186/s12906-025-05056-w
Zarrilli, R., Di Popolo, A., Bagattini, M., Giannouli, M., Martino, D., Barchitta, M., et al. (2012). Clonal spread and patient risk factors for acquisition of extensively drug-resistant Acinetobacter baumannii in a neonatal intensive care unit in Italy. J. Hosp Infect. 82, 260–265. doi: 10.1016/j.jhin.2012.08.018
Zhang, Y., Jiao, F., Zeng, D., Yu, X., Zhou, Y., Xue, J., et al. (2024). Synergistic Effects of Pyrrosia lingua Caffeoylquinic Acid Compounds with Levofloxacin Against Uropathogenic Escherichia coli: Insights from Molecular Dynamics Simulations, Antibiofilm, and Antimicrobial Assessments. Molecules. 29, 5679. doi: 10.3390/molecules29235679
Keywords: Acinetobacter baumannii, biofilm, motility, multidrug resistance, children
Citation: Yu Z, Fu H, Zhou Y, Luo X, Wu Y, Kuang Z, Cai D, Wu X and Hong X (2025) Molecular, biofilm and motility characterization of Acinetobacter baumannii isolated from a neonatal intensive care unit at a children’s hospital in South China. Front. Cell. Infect. Microbiol. 15:1705282. doi: 10.3389/fcimb.2025.1705282
Received: 14 September 2025; Accepted: 05 November 2025; Revised: 30 October 2025;
Published: 28 November 2025.
Edited by:
Xihui Shen, Northwest A&F University, ChinaCopyright © 2025 Yu, Fu, Zhou, Luo, Wu, Kuang, Cai, Wu and Hong. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiaobing Hong, eGlhb2Jpbmdob25nQDEyNi5jb20=
†These authors have contributed equally to this work
 Zelin Yu1†
Zelin Yu1† Yanqiong Zhou
Yanqiong Zhou Xiajiong Luo
Xiajiong Luo Xiaobing Hong
Xiaobing Hong