- 1Searle Chemistry Laboratory, Computation Institute, University of Chicago, Chicago, IL, USA
- 2Computing, Environment and Life Sciences, Argonne National Laboratory, Chicago, IL, USA
- 3School of Earth, Atmospheric, and Environmental Sciences, University of Manchester, Manchester, UK
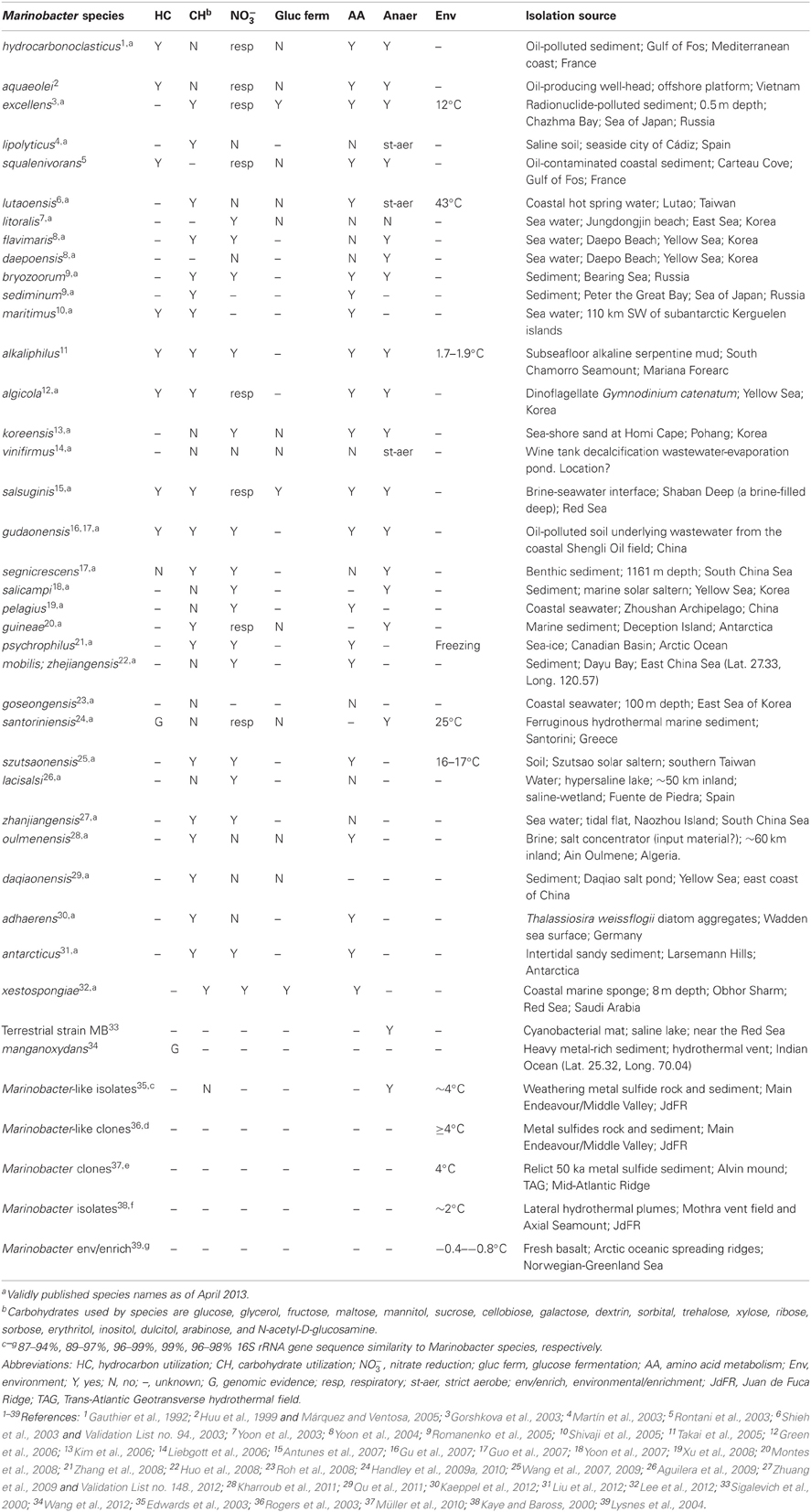
The Marinobacter genus comprises widespread marine bacteria, found in localities as diverse as the deep ocean, coastal seawater and sediment, hydrothermal settings, oceanic basalt, sea-ice, sand, solar salterns, and oil fields. Terrestrial sources include saline soil and wine-barrel-decalcification wastewater. The genus was designated in 1992 for the Gram-negative, hydrocarbon-degrading bacterium Marinobacter hydrocarbonoclasticus. Since then, a further 31 type strains have been designated. Nonetheless, the metabolic range of many Marinobacter species remains largely unexplored. Most species have been classified as aerobic heterotrophs, and assessed for limited anaerobic pathways (fermentation or nitrate reduction), whereas studies of low-temperature hydrothermal sediments, basalt at oceanic spreading centers, and phytoplankton have identified species that possess a respiratory repertoire with significant biogeochemical implications. Notable physiological traits include nitrate-dependent Fe(II)-oxidation, arsenic and fumarate redox cycling, and Mn(II) oxidation. There is also evidence for Fe(III) reduction, and metal(loid) detoxification. Considering the ubiquity and metabolic capabilities of the genus, Marinobacter species may perform an important and underestimated role in the biogeochemical cycling of organics and metals in varied marine habitats, and spanning aerobic-to-anoxic redox gradients.
Introduction
Marine habitats are host to a diverse range of substrates and physicochemical regimes. Among these, hydrothermal features attract particular interest owing to ore-grade concentrations of metals, physicochemical extremes, and the presence of chemolithoautotrophic macrofauna and microbiota. Bacteria and Archaea occupying marine habitats have a substantial physical presence. There are an estimated 3.9 × 1030 prokaryotic cells in the open ocean and unconsolidated marine sediments, comprising ~3.1 × 1011 tonnes of carbon (Whitman et al., 1998). This is slightly more than the estimated carbon content of terrestrial prokaryotes, and just under half of that in all plant life. Prokaryotes can contribute significantly to marine ecosystem functioning and biogeochemical cycles (Jørgensen, 2006), owing to their prevalence and enormous capacity for transforming their environments through metabolism of organic and inorganic matter (Gadd, 2010). Yet much of the marine microbial biomass remains unexplored, and there is still much to learn about heterotrophic and autotrophic bacterial functioning in the ocean (e.g., Moran et al., 2004; Emerson et al., 2010; Holden et al., 2012).
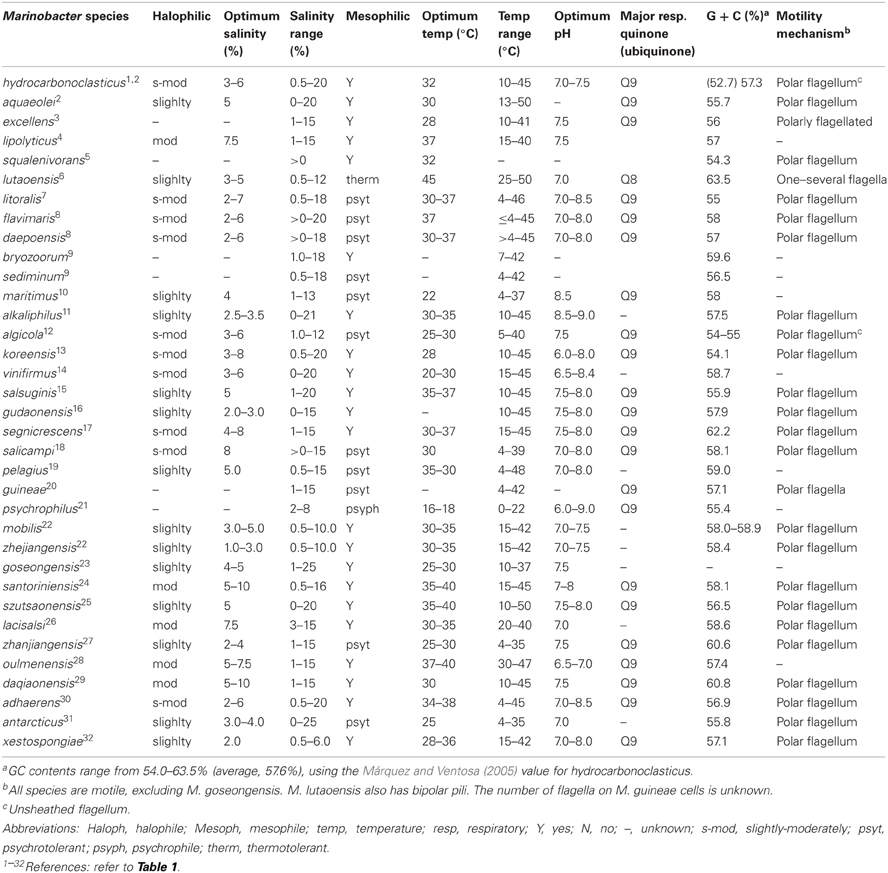
Marinobacter is a heterotrophic, and in some instances mixotrophic (Dhillon et al., 2005; Handley et al., 2009a,b), metabolically flexible genus found in an exceptionally wide range of marine and saline terrestrial settings, including various low-temperature hydrothermal environments (Table 1). The genus comprises Gram-negative Gammaproteobacteria within the Alteromonadales order. All known species are motile with polar flagella (excluding M. goseongensis, Roh et al., 2008), slightly to moderately halophilic (cf. DasSarma and Arora, 2012), aerobic heterotrophs (Table 2). However, few are confirmed strict aerobes, and several are facultative anaerobes (Table 1). All are rod-shaped, with the exception of the ellipsoidal M. Segnicrescens (Guo et al., 2007), and most are neutrophilic, except the slightly alkaliphilic M. alkaliphilus, which grows optimally at pH 8.5–9.0 (Takai et al., 2005; also see Al-Awadhi et al., 2007; Table 2). Although most species are mesophilic, many are psychrotolerant (also known as psychrotrophic) and capable of growth down to ~4°C (Table 2; Moyer and Morita, 2007). A couple of other species are either psychrophilic (with a growth optimum near 15°C) or thermotolerant (with growth up to 50°C and an optimum of 45°C). This phenotypic versatility contributes to the ubiquity of this genus, and its ability to occupy diverse physicochemical regimes.
Marinobacter Hydrocarbonoclasticus, Denitrification and Hydrocarbons
The genus was created for M. hydrocarbonoclasticus, which was isolated from hydrocarbon-polluted sediment, collected from the mouth of an oil refinery outlet along the French Mediterranean coast (Gauthier et al., 1992), and includes the later heterotypic synonym, M. aquaeolei (Huu et al., 1999; Márquez and Ventosa, 2005). The species has an obligate requirement for sodium. It grows readily on complex organic media containing yeast extract and peptone, and aerobically on a range of organic acids (acetate, butyrate, caproate, fumarate, adipate, lactate, citrate), and the amino acids L-glutamate, L-glutamine, and L-proline. Under anaerobic conditions M. hydrocarbonoclasticus can perform denitrification using a membrane-bound respiratory NarGHI complex to reduce nitrate (Correia et al., 2008). The nitrite formed is reduced to N2 via nitrite reductase cytochrome cd1 (Besson et al., 1995), nitric oxide reductase NorBC (EMBL-EBI ABM20188.1, ABM20189.1), and nitrous oxide reductase (Prudêncio et al., 2000). M. hydrocarbonoclasticus is most notable, however, for its ability to aerobically degrade liquid and solid, aliphatic (pristane, heneicosane, eicosane, hexadecane, tetradecane) and aromatic (phenanthrene, phenyldecane) hydrocarbons. It uses each hydrocarbon as a sole energy source, and produces large quantities of bioemulsifier. Bioemulsifiers (biosurfactants), are thought to aid in bacterial adhesion to hydrophobic surfaces, water-immiscible material breakdown, and competitor inhibition, and are attracting increasing interest for various industrial applications (Banat et al., 2000; Nerurkar et al., 2009; Williams, 2009; Soberón-Chávez and Maier, 2011).
Of the 34 species named since the genus was created, several exhibit hydrocarbonoclastic activity, while others remain untested (Table 1). Additional hydrocarbons utilized by Marinobacter species include squalene, which is metabolized under denitrifying conditions (Rontani et al., 2003), polycyclic aromatic hydrocarbons (PAHs) (Cui et al., 2008), hexane, heptane, petroleum ether (Shivaji et al., 2005; Antunes et al., 2007), n-pentadecane, n-tridecane, n-undecane, n-decane, n-nonane, butane, and kerosene (Takai et al., 2005).
This hydrocarbonoclastic capacity in Marinobacter has attracted attention owing to the potential for these bacteria to remediate crude oil contamination in environments as diverse as the Arabian Gulf (Al-Awadhi et al., 2007) and Artic sea ice (Gerdes et al., 2005). Nitrate reduction by Marinobacter species has also been exploited for potential use in oilfield maintenance. Dunsmore et al. (2006) showed reduction of added nitrate prevented deleterious growth of sulfate-reducing bacteria in produced water from a North Sea oilfield oil reservoir, controlling microbial souring reactions. The beneficial reduction of nitrate was largely attributed to indigenous Marinobacter species.
Expanded Functional Traits of the Genus
Following the characterization of M. hydrocarbonoclasticus, the functional range of the genus has been further expanded to include (non-exhaustively) fermentation; the ability to respire at least 19 different carbohydrates (Table 1) and several extra amino (e.g., L-alanine, D-glutamate, L-phenylalanine; Antunes et al., 2007 and Green et al., 2006) and organic acids [e.g., malonate, formate, pyruvate, alpha-ketoglutarate; Kim et al. (2006) and Kharroub et al. (2011)]; degradation of the isoprenoid ketone 6,10,14-trimethylpentadecan-2-one (Rontani et al., 1997); growth on ethanol (Gu et al., 2007; Huo et al., 2008), phenol (Liebgott et al., 2006), and various Tweens (e.g., Takai et al., 2005; Green et al., 2006) following enzymatic evidence in M. hydrocarbonoclasticus; utilization of fumarate as an electron acceptor (Takai et al., 2005; Handley et al., 2009a); and oxidation/reduction of arsenic, iron or manganese (Handley et al., 2009a,b; Wang et al., 2012).
As for M. hydrocarbonoclasticus, all subsequently described species are able to grow aerobically on complex organic matter, and oxidize organic acids. Many, but not all are enzymatically able to reduce nitrate (Table 1). Lack of fermentation by M. hydrocarbonoclasticus was initially proposed as a distinctive feature of the genus; however, a number of subsequently isolated type strains exhibit both fermentative and respiratory metabolisms, owing to their ability to ferment glucose (Table 1), lactate (Handley et al., 2009a) and other substrates (Lee et al., 2012). Evaluation of recently available genome sequences also suggests certain Marinobacter species may exhibit enzymatic resistance to arsenic and heavy metals (e.g., Wang et al., 2012).
Biogeography and Phylogeny
Marinobacter colonize diverse saline habitats, e.g., sea ice and hydrothermal sediments, facilitated by psychrophilic to thermotolerant physiologies, and an ability to metabolize an array of (in)organic compounds under aerobic or anaerobic conditions. However, evaluation of phylogenetic trees, constructed using 16S rRNA gene sequences, suggest the genus is monophyletic, forming a single clade distinct from other closely related epsilonproteobacterial lineages (Figure 1).
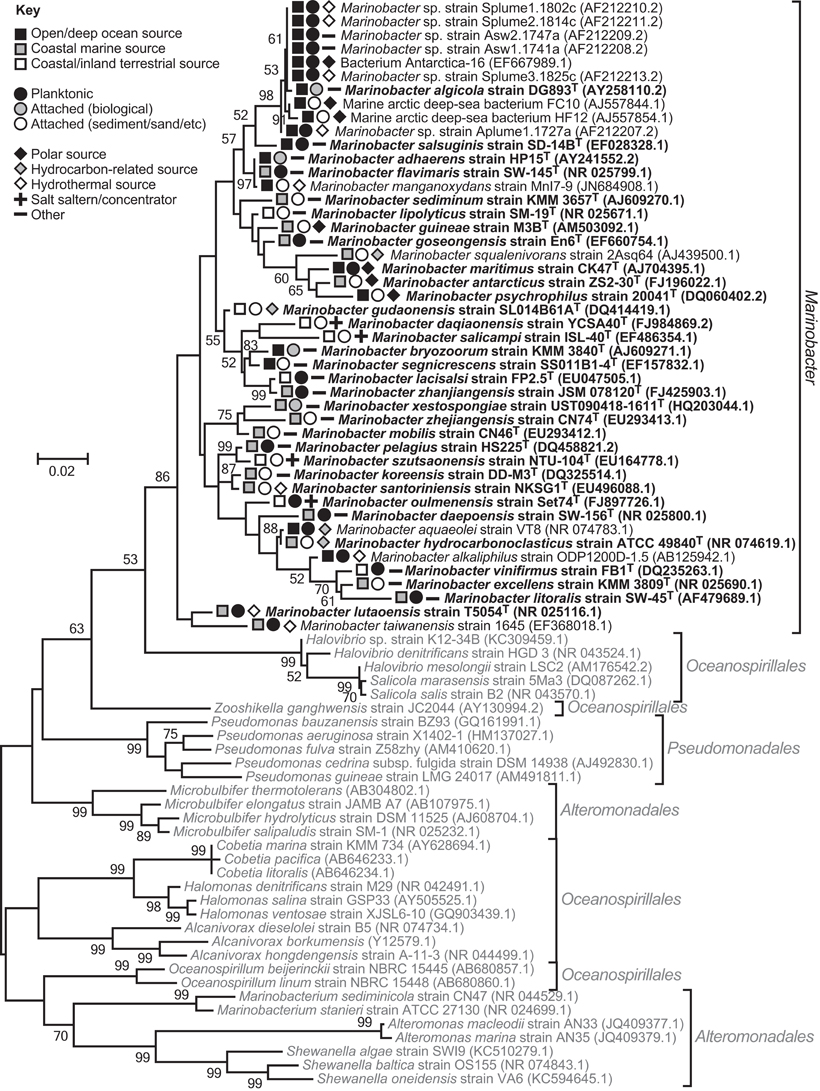
Figure 1. 16S rRNA gene phylogenetic maximum-likelihood tree comparing Marinobacter species and their nearest neighbors within the epsilonproteobacterial orders, Alteromonadales, Pseudomonadales, and Oceanospirillales. The tree indicates the genus is monophyletic, despite the three orders being non-monophyletic (Williams et al., 2010). The same result was obtained using the neighbor-Joining method. Trees were constructed using MEGA v5.0 (Tamura et al., 2011), Clustal W alignments (Thompson et al., 1994), and 1000 bootstrap replicates. Bootstrap values ≥50 are shown. Sequences used were ≥1350 bp long. Marinobacter isolates are in dark font with type species bolded, and closely related Gammaproteobacteria are in pale font. GenBank accession numbers are given in parentheses. The symbols indicate Marinobacter isolate sources.
Despite the physiological versatility of the genus, and the ability of some strains to grow (non-optimally) without salt (e.g., Huu et al., 1999; Sigalevich et al., 2000; Liebgott et al., 2006), Marinobacter appear to be geographically restricted to marine or terrestrial environments rich in sodium salts. This observation is consistent with the hypothesis that microorganisms exhibit non-random biogeographical differentiation and distribution, due in part to environmental selection (Martiny et al., 2006).
In marine environments, dispersal does not appear to be a limiting factor for Marinobacter. Strains have been isolated from, and phylogenetically detected in, oceans (Pacific, Atlantic, Indian, Arctic, and Antarctic) and seas, spanning the globe from pole to equator (Table 1; Kaye et al., 2011). They display both attached and planktonic lifestyles, and the distribution of the genus extends from deep-ocean (hydrothermal) benthic sediment and exposed basalt to surface water, or coastal (hydrothermal) sediment, hot spring water and sand (Table 1; Figure 1).
In many instances, terrestrial isolation sources can be clearly linked to the ocean (e.g., coastal solar salterns and hot springs, and polluted soil from a coastal oil field; Table 1). Isolation of species from terrestrial sources, up to 50–60 km inland, implies a greater degree of terrestrial dispersal (Table 1; Figure 1 and Table S2 in Kaye et al., 2011). However, there is insufficient information regarding the nature of terrestrial isolation sources, and too few isolate and phylogenetic data, to judge how well-dispersed this genus is on land, or whether terrestrial sources are strictly independent from marine influences.
Lifesytles
In many respects Marinobacter species are generalists like their marine and terrestrial Alteromonadales cousins in the Shewanella genus. Shewanella species are respiratory generalists (e.g., Heidelberg et al., 2002), and at least one species (S. baltica) has been described as “very close to the ultimate [marine] r-strategist,” starkly contrasting with genomically streamlined K-strategist (or oligotrophic) marine bacteria like Prochlorococcus (Caro-Quintero et al., 2011). In the presence of surplus organic carbon, Marinobacter can grow rapidly, out-competing other bacteria in enrichment cultures (e.g., Handley et al., 2010). This r-strategist (or copiotrophic) behavior renders them weed-like and relatively easy to cultivate, even compared with other heterotrophic marine bacteria (Kaye and Baross, 2000). Marinobacter can also excel under aerobic-to-anaerobic conditions with no added substrate, while in the presence of Fe(II) (Edwards et al., 2003; Handley et al., 2013a).
This type of lifestyle exhibited by Marinobacter strains has been described as opportunistic or “opportunitrophic” (Singer et al., 2011), following the definition given by Moran et al. (2004) in describing the ability of the marine bacterium Silicibacter pomeroyi to switch rapid between lithoheterotrophy to heterotrophy in response to nutrient pulses. Use of “-troph” in this context describes the mode of obtaining nourishment (as for the term “psychrotroph”) rather than the source of the nourishment (as in “organotroph”), and as such may be considered a misnomer. The term was applied in order to differentiate between types of fast growing and nominally r-strategist bacteria, specifically between specialists (e.g., Geobacter, Mahadevan and Lovley, 2008), and generalists like Marinobacter, Shewanella, Pseudomonas, Vibrio and Roseobacter (Singer et al., 2011)—with the latter two genera already having been deemed opportunistic based on their versatile lifestyles (“opportunitrophs”; Polz et al., 2006). Singer et al. (2011) also identified other potential commonalities shared among the genomes of opportunistic bacteria and M. aquaeolei VT8, including a large genomic toolkit for responding to environmental stimuli and for defense (cf. Polz et al., 2006).
There are few phylogenetic studies of the environments from which Marinobacter species have been isolated that evaluate their in situ relative abundance. Nevertheless, the studies that have been published show two different scenarios for Marinobacter. Strains may be characterized as r-strategists or opportunistic (Polz et al., 2006; Singer et al., 2011), and dominate communities sporadically when stimulated by high nutrient loads, encountered, for example, in marine aggregates or enrichment cultures (Balzano et al., 2009; Handley et al., 2010). In contrast, relatively high in situ abundances of Marinobacter (Müller et al., 2010) and uncultured organisms closely related to Marinobacter (Rogers et al., 2003; Edwards et al., 2004) have been observed in some hydrothermal systems, implying these organisms may play an important and sustained role in post-depositional mineral alteration.
Hydrothermal Settings
Marine hydrothermal systems are dispersed throughout the world's oceans (Martin et al., 2008), and support abundant psychrophilic-to-mesophilic life even in close proximity to high-temperature venting (Reysenbach and Cady, 2001; Edwards et al., 2003). A number of studies suggest Marinobacter may be significant in ‘low-temperature’ hydrothermal systems, defined by low-temperature hydrothermalism (e.g., ~8 to ~40°C, McCollom and Shock, 1997) or ambient seawater temperatures (e.g., ~2° in the deep ocean). This is due to their documented association with several different hydrothermal features (Table 1), and to their ability to heterotrophically or mixotrophically respire inorganic compounds abundant in hydrothermal systems.
A common hydrothermal feature found at plate boundaries, and with which Marinobacter or near-relatives have been associated (Edwards et al., 2003; Rogers et al., 2003; Müller et al., 2010), are massive sulfides, which comprise an estimated 6 × 108 tonnes of material globally (Hannington et al., 2011), and adjunct metalliferous sediments. While much of the material for massive sulfides originates from high-temperature hydrothermal fluids (>350°C) emanating from black smoker chimneys (Hannington et al., 2011), particulates distributed locally by plumes and talus from mound and chimney collapse can equilibrate with ambient temperatures (Edwards et al., 2003), or entire mounds can be inactive (Müller et al., 2010). Adjacent to massive sulfide deposits are low-temperature iron- and manganese-rich metalliferous sediments, derived from distal plume fallout with contributions from mound mass wasting (Jannasch and Mottl, 1985; Mills et al., 1993; Hannington et al., 1998)—possibly of the type from which M. manganoxydans was isolated (Wang et al., 2012).
Deposits consisting of iron oxyhydroxides, nontronite (a ferric iron-rich clay) and iron-manganese crusts can form independently at plate boundaries or at places of intra-plate volcanism (e.g., Alt, 1988; Karl et al., 1988; Boyd and Scott, 2001; Kennedy et al., 2004; Edwards et al., 2011). They form from diffuse low-temperature venting (Karl et al., 1988; Edwards et al., 2011), and can span areas >100 m2 (Boyd and Scott, 2001). Similar deposits exist in shallow marine settings, such as the ferruginous arsenic-rich sediments found in Papua New Guinea and Santorini (Smith and Cronan, 1983; Pichler and Veizer, 1999). M. santoriniensis was isolated from the Santorini sediment (Handley et al., 2009a).
Further examples of low-temperature hydrothermal habitats, with which Marinobacter or near relatives are associated, include those created by sharp temperature gradients that form across high-temperature chimney walls (Rogers et al., 2003), or buoyant plumes (Kaye and Baross, 2000) that rise 200–300 m up from these vents and spread laterally (German et al., 1991). Exposed, iron-rich basalt, delivered by oceanic spreading centers, provides another environment associated with many deep-sea hydrothermal systems (Lysnes et al., 2004), whereas M. alkaliphilus was isolated from alkaline serpentine mud (Takai et al., 2005) from a setting peculiar to mud volcanoes on the non-accretionary Mariana forearc (Fryer et al., 1999).
Function, Biogeochemistry and Hydrothermal Systems
The various low-temperature hydrothermal settings Marinobacter(-like) species inhabit are rich in metals/metalloids, such as iron, manganese, arsenic, copper and zinc (Smith and Cronan, 1983; Hannington et al., 1998) that certain Marinobacter strains can transform enzymatically. Moreover, oxygen gradients established in these sediments may be exploited by Marinobacter species able to grow heterotrophically under anaerobic/aerobic conditions and mixotrophically under aerobic conditions.
Among the functions Marinobacter may perform in these environments is ferrous iron oxidation. The potential for Marinobacter Fe(II) oxidation was first suggested by Edwards et al. (2003) after isolating iron-oxidizing bacteria, phylogenetically resembling M. aquaeolei, from low-temperature hydrothermal metal sulfides. The isolates were able to grow chemoautotrophically on pyrite, basalt glass and siderite under micro-aerobic conditions. This promoted subsequent study of M. aquaeolei, including genome sequencing, and identification of its ability to anaerobically oxidize Fe(II) under mixotrophic conditions (Dhillon et al., 2005; Edwards et al., 2006; Singer et al., 2011). Subsequently, M. santoriniensis, which was isolated from iron-rich hydrothermal sediment, was also shown to perform nitrate-dependent Fe(II) oxidation when supplemented with a small amount of organic carbon (Handley et al., 2009a).
Interestingly, M. santoriniensis was isolated from sediment rife with stalk-like cells and bacteria phylogenetically resembling iron-oxidizing Zetaproteobacteria (Handley et al., 2010). Other Marinobacter (or near relatives) were also cultivated from environments containing stalks (Edwards et al., 2003; Lysnes et al., 2004) that speculatively belong to this increasingly characteristic phylum of marine iron-oxidizers—the Zetaproteobacteria (Emerson et al., 2007, 2010; Edwards et al., 2011).
As Marinobacter are reputedly more versatile than Mariprofundus ferrooxydans strains (the sole representatives of the Zetaproteobacteria) it is possible they perform other functions in these environments instead of, or in addition to, Fe(II) oxidation. For instance, Marinobacter and Marinobacter-like isolates have been implicated in Fe(III) reduction, but only in complex or simple co-cultures with other bacteria (Lysnes et al., 2004; Balzano et al., 2009; Handley et al., 2010). This metabolic trait remains to be demonstrated in definitively anexic cultures. M. santoriniensis has the genetic potential to reductively detoxify arsenate and mercury using proteins encoded by an Escherichia coli-like arsC and merRTA genes (Handley et al., 2013c), in addition to being able to conserve energy for growth via arsenate [As(V)] respiration using an unidentified mechanism, and mixotrophically oxidize arsenite [As(III)] using the aro gene cluster—making it one of a handful of bacteria currently known to completely redox-cycle arsenic (Handley et al., 2009b). This is particularly relevant given that the bacterium was isolated from sediment containing ~400 ppm of arsenic.
It remains to be explored whether other Marinobacter species share this ability to respire arsenic. However, there is cursory evidence for non-respiratory arsenate reductase (plus/minus putative respiratory arsenite oxidase) genes in several publically available Marinobacter genomes (namely, M. hydrocarbonoclasticus ATCC49840, GenBank FO203363.1, Grimaud et al., 2012; M. aquaeolei VT8, GenBank CP000514.1, Singer et al., 2011; M. adhaerens HP15, GenBank CP001978.1, Gärdes et al., 2010; M. algicola DG893, GenBank ABCP00000000.1; Marinobacter spp. BSs20148 and ELB17, GenBank CP003735.1 and AAXY00000000.1). Likewise, in a recent genome announcement Wang et al. (2012) described a Marinobacter candidate, M. manganoxydans MnI7-9 that has not only a putative arsC gene for arsenic detoxification (GenBank YP_005884959.1), but also a host of other genes that may be used for nickel, mercury, copper, chromate, zinc, cobalt, and cadmium resistance. This bacterium adsorbs and tolerates high levels of metals/metalloids, alongside a demonstrated ability to oxidize manganese, Mn(II), to a mixed-valency Mn(III)/Mn(IV) product via an unidentified genetic mechanism. Bacterial manganese oxidation is not thought to be an energy conserving process, but it is considered significant in environmental Mn(IV) oxide formation (Geszvain et al., 2012).
Conclusions and Future Directions
Although the genus is widespread in marine settings, and dozens of cultivated representatives and several sequenced genomes exist, the functional breath of Marinobacter species remains largely unexplored. The ability to metabolize hydrocarbons and inorganic elements (e.g., iron, arsenic, manganese) has been tested in relatively few species. Information, based on cultures and isolation source characteristics, suggests species within the genus are able to contribute, for example, to the degradation of hydrocarbons in oil-polluted sediment, and the oxidation of Fe(II) in ferruginous sediment or basalt. However, we know little about the nature and magnitude of their actual function in the environment. High-throughput omics (genomics, transcriptomics, proteomics) techniques promise to expand our knowledge into the uncultivated black box that encompasses much of the microbiome, and to facilitate in situ investigations of communities (e.g., Ram et al., 2005; Lo et al., 2007; Baker et al., 2012; Handley et al., 2013b), but are limited in part by the large number of genes of unknown function. Much can still be achieved from cultivation experiments. In moving forward, a combination of omics, functional gene expression studies, isotope tracer and cultivation techniques will provide a powerful complement of tools for characterizing both the real and potential function of microorganisms in marine settings and elsewhere, and elucidating the (opportunistic?) role of Marinobacter species in environmental biogeochemical cycles.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We acknowledge funding support from the European Union through the BIOtransformations of TRace elements in AquatiC Systems (BIOTRACS) EST programme, Marie Curie Actions.
References
Aguilera, M., Jiménez-Pranteda, M. L., Kharroub, K., González-Paredes, A., Durban, J. J., Russell, N. J., et al. (2009). Marinobacter lacisalsi sp. nov., a moderately halophilic bacterium isolated from the saline-wetland wildfowl reserve Fuente de Piedra in southern Spain. Int. J. Syst. Evol. Microbiol. 59, 1691–1695.
Al-Awadhi, H., Sulaiman, R. H., Mahmoud, H. M., and Radwan, S. S. (2007). Alkaliphilic and halophilic hydrocarbon-utilizing bacteria from Kuwaiti coasts of the Arabian Gulf. Appl. Microbiol. Biotechnol. 77, 183–186.
Alt, J. C. (1988). Hydrothermal oxide and nontronite deposits on seamounts in the eastern Pacific. Mar. Geol. 81, 227–239.
Antunes, A., Franca, L., Rainey, F. A., Huber, R., Nobre, M. F., Edwards, K. J., et al. (2007). Marinobacter salsuginis sp. nov., isolated from the brine-seawater interface of the Shaban Deep, Red Sea. Int. J. Syst. Evol. Microbiol. 57, 1035–1040.
Baker, B. J., Lesniewski, R. A., and Dick, G. J. (2012). Genome-enabled transcriptomics reveals archaeal populations that drive nitrification in a deep-sea hydrothermal plume. ISME J. 6, 2269–2279.
Balzano, S., Statham, P. J., Pancost, R. D., and Lloyd, J. R. (2009). Role of microbial populations in the release of reduced iron to the water column from marine aggregates. Aquat. Microb. Ecol. 54, 291–303.
Banat, I. M., Makkar, R. S., and Cameotra, S. S. (2000). Potential commercial applications of microbial surfactants. Appl. Microbiol. Biotechnol. 53, 495–508.
Besson, S., Carneiro, C., Moura, J. J., Moura, I., and Fauque, G. (1995). A cytochrome cd1-type nitrite reductase isolated from the marine denitrifier Pseudomonas nautica 617: purification and characterization. Anaerobe 1, 219–226.
Boyd, T. D., and Scott, S. D. (2001). Microbial and hydrothermal aspects of ferric oxyhydroxides and ferrosic hydroxides: the example of Franklin Seamount, Western Woodlark Basin, Papua New Guinea. Geochem. Trans. 2, 45.
Caro-Quintero, A., Deng, J., Auchtung, J., Brettar, I., Höfle, M. G., Klappenbach, J., et al. (2011). Unprecedented levels of horizontal gene transfer among spatially co-occurring Shewanella bacteria from the Baltic Sea. ISME J. 5, 131–140.
Correia, C., Besson, S., Brondino, C. D., Gonzalez, P. J., Fauque, G., Lampreia, J., et al. (2008). Biochemical and spectroscopic characterization of the membrane-bound nitrate reductase from Marinobacter hydrocarbonoclasticus 617. J. Biol. Inorg. Chem. 13, 1321–1333.
Cui, Z., Lai, Q., Dong, C., and Shao, Z. (2008). Biodiversity of polycyclic aromatic hydrocarbon-degrading bacteria from deep sea sediments of the Middle Atlantic Ridge. Environ. Microbiol. 10, 2138–2149.
DasSarma, S., and Arora, P. (2012). “Halophiles,” in eLS. Chichester: John Wiley and Sons, Ltd. doi: 10.1002/9780470015902.a0000394.pub3
Dhillon, A., Edwards, K. J., Webb, E., Rogers, D., and Sogin, M. L. (2005). “Marinobacter aquaeolei gene expression studies for clues to neutrophilic iron oxidation,” in NASA Astrobiology Institute biennial meeting, Abstract 829 (University of Colorado, Boulder, CO).
Dunsmore, B., Youldon, J., Thrasher, D. R., and Vance, I. (2006). Effects of nitrate treatment on a mixed species, oil field microbial biofilm. J. Ind. Microbiol. Biotechnol. 33, 454–462.
Edwards, K., Sogin, M., and Dhillon, A. (2006). Why Sequence Marinobacter aquaeolei? DOE Joint Genome Institute, Available online at: http://www.jgi.doe.gov/sequencing/why/marinobacter.html
Edwards, K. J., Bach, W., McCollom, T. M., and Rogers, D. R. (2004). Neutrophilic iron-oxidizing bacteria in the ocean: their babitats, diversity, and roles in mineral deposition, rock alteration, and biomass production in the deep-sea. Geomicrobiol. J. 21, 393–404.
Edwards, K. J., Glazer, B. T., Rouxel, O. J., Bach, W., Emerson, D., Davis, R. E., et al. (2011). Ultra-diffuse hydrothermal venting supports Fe-oxidizing bacteria and massive umber deposition at 5000 m off Hawaii. ISME J. 5, 1748–1758.
Edwards, K. J., Rogers, D. R., Wirsen, C. O., and McCollom, T. M. (2003). Isolation and characterization of novel psychrophilic, neutrophilic, Fe-oxidizing, chemolithoautotrophic alpha- and gamma-proteobacteria from the deep sea. Appl. Environ. Microbiol. 69, 2906–2913.
Emerson, D., Fleming, E. J., and McBeth, J. M. (2010). Iron-oxidizing bacteria: an environmental and genomic perspective. Annu. Rev. Microbiol. 64, 561–583.
Emerson, D., Rentz, J. A., Lilburn, T. G., Davis, R. E., Aldrich, H., Chan, C., et al. (2007). A novel lineage of proteobacteria involved in formation of marine Fe-oxidizing microbial mat communities. PLoS ONE 2:e667. doi: 10.1371/journal.pone.0000667
Fryer, P., Wheat, C. G., and Mottl, M. J. (1999). Mariana blueschist mud volcanism: implications for conditions within the subduction zone. Geology 27, 103–106.
Gadd, G. M. (2010). Metals, minerals and microbes: geomicrobiology and bioremediation. Microbiology 156, 609–643.
Gärdes, A., Kaeppel, E., Shehzad, A., Seebah, S., Teeling, H., Yarza, P., et al. (2010). Complete genome sequence of Marinobacter adhaerens type strain (HP15), a diatom-interacting marine microorganism. Stand. Genomic. Sci. 3, 97–107.
Gauthier, M. J., Lafay, B., Christen, R., Fernandez, L., Acquaviva, M., Bonin, P., et al. (1992). Marinobacter hydrocarbonoclasticus gen. nov., sp. nov., a new, extremely halotolerant, hydrocarbon-degrading marine bacterium. Int. J. Syst. Bacteriol. 42, 568–576.
Gerdes, B., Brinkmeyer, R., Dieckmann, G., and Helmke, E. (2005). Influence of crude oil on changes of bacterial communities in Arctic sea-ice. FEMS Microbiol. Ecol. 53, 129–139.
German, C. R., Campbell, A. C., and Edmond, J. M. (1991). Hydrothermal scavenging at the Mid-Atlantic Ridge: modification of trace element dissolved fluxes. Earth Planet. Sc. Lett. 107, 101–114.
Geszvain, K., Butterfield, C., Davis, R. E., Madison, A. S., Lee, S. W., Parker, D. L., et al. (2012). The molecular biogeochemistry of manganese(II) oxidation. Biochem. Soc. Trans. 40, 1244–1248.
Gorshkova, N. M., Ivanova, E. P., Sergeev, A. F., Zhukova, N. V., Alexeeva, Y., Wright, J. P., et al. (2003). Marinobacter excellens sp. nov., isolated from sediments of the Sea of Japan. Int. J. Syst. Evol. Microbiol. 53, 2073–2078.
Green, D. H., Bowman, J. P., Smith, E. A., Gutierrez, T., and Bolch, C. J. (2006). Marinobacter algicola sp. nov., isolated from laboratory cultures of paralytic shellfish toxin-producing dinoflagellates. Int. J. Syst. Evol. Microbiol. 56, 523–527.
Grimaud, R., Ghiglione, J. F., Cagnon, C., Lauga, B., Vaysse, P. J., Rodriguez-Blanco, A., et al. (2012). Genome sequence of the marine bacterium Marinobacter hydrocarbonoclasticus SP17, which forms biofilms on hydrophobic organic compounds. J. Bacteriol. 194, 3539–3540.
Gu, J., Cai, H., Yu, S. L., Qu, R., Yin, B., Guo, Y. F., et al. (2007). Marinobacter gudaonensis sp. nov., isolated from an oil-polluted saline soil in a Chinese oilfield. Int. J. Syst. Evol. Microbiol. 57, 250–254.
Guo, B., Gu, J., Ye, Y. G., Tang, Y. Q., Kida, K., and Wu, X. L. (2007). Marinobacter segnicrescens sp. nov., a moderate halophile isolated from benthic sediment of the South China Sea. Int. J. Syst. Evol. Microbiol. 57, 1970–1974.
Handley, K. M., Boothman, C., McBeth, J., Charnock, J. M., Wincott, P. L., Vaughan, D. J., et al. (2013a). Effect of microbially mediated iron redox transformations on arsenic solid-phase associations in a high-iron, arsenic-rich hydrothermal sediment. Geochim. Cosmochim. Acta 102, 124–142.
Handley, K. M., Verberkmoes, N. C., Steefel, C. I., Williams, K. H., Sharon, I., Miller, C. S., et al. (2013b). Biostimulation induces syntrophic interactions that impact C, S and N cycling in a sediment microbial community. ISME J. 7, 800–816.
Handley, K. M., Upton, M., Beatson, S. A., Héry, M., and Lloyd, J. R. (2013c). Genome sequence of hydrothermal arsenic-respiring bacterium Marinobacter santoriniensis strain NKSG1T. Genome Announc. 1:e00231-13. doi: 10.1128/genomeA.00231-13
Handley, K. M., Boothman, C., Mills, R. A., Pancost, R. D., and Lloyd, J. R. (2010). Functional diversity of bacteria in a ferruginous hydrothermal sediment. ISME J. 4, 1193–1205.
Handley, K. M., Héry, M., and Lloyd, J. R. (2009a). Marinobacter santoriniensis sp. nov., an arsenate-respiring and arsenite-oxidizing bacterium isolated from hydrothermal sediment. Int. J. Syst. Evol. Microbiol. 59, 886–892.
Handley, K. M., Héry, M., and Lloyd, J. R. (2009b). Redox cycling of arsenic by the hydrothermal marine bacterium Marinobacter santoriniensis. Environ. Microbiol. 11, 1601–1611.
Hannington, M., Jamieson, J., Monecke, T., Petersen, S., and Beaulieu, S. (2011). The abundance of seafloor massive sulfide deposits. Geology 39, 1155–1158.
Hannington, M. D., Galley, A. G., Herzig, P. M., and Petersen, S. (1998). “Comparison of the TAG mound and stockwork complex with Cyprus type massive sulfide deposits,” in Proceedings of the Ocean Drilling Program, Scientific Results, Vol. 158 (College Station, TX), 380–415.
Heidelberg, J. F., Paulsen, I. T., Nelson, K. E., Gaidos, E. J., Nelson, W. C., Read, T. D., et al. (2002). Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat. Biotechnol. 20, 1118–1123.
Holden, J. F., Breier, J. A., Rogers, K. L., Schulte, M. D., and Toner, B. M. (2012). Biogeochemical processes at hydrothermal vents: microbes and minerals, bioenergetics, and carbon fluxes. Oceanography 25, 196–208.
Huo, Y. Y., Wang, C. S., Yang, J. Y., Wu, M., and Xu, X. W. (2008). Marinobacter mobilis sp. nov. and Marinobacter zhejiangensis sp. nov., halophilic bacteria isolated from the East China Sea. Int. J. Syst. Evol. Microbiol. 58, 2885–2889.
Huu, N. B., Denner, E. B., Ha, D. T., Wanner, G., and Stan-Lotter, H. (1999). Marinobacter aquaeolei sp. nov., a halophilic bacterium isolated from a Vietnamese oil-producing well. Int. J. Syst. Bacteriol. 49(Pt 2), 367–375.
Jannasch, H. W., and Mottl, M. J. (1985). Geomicrobiology of deep-sea hydrothermal vents. Science 229, 717–725.
Jørgensen, B. B. (2006). “Bacteria and marine biogeochemistry,” in Marine Geochemistry, eds. H. D. Schultzand M. Zabel (Berlin: Springer), 169–206.
Kaeppel, E. C., Gardes, A., Seebah, S., Grossart, H. P., and Ullrich, M. S. (2012). Marinobacter adhaerens sp. nov., isolated from marine aggregates formed with the diatom Thalassiosira weissflogii. Int. J. Syst. Evol. Microbiol. 62, 124–128.
Karl, D. M., McMurtry, G. M., Malahoff, A., and Garcia, M. O. (1988). Loihi Seamount, Hawaii: a mid-plate volcano with a distinctive hydrothermal system. Nature 335, 532–535.
Kaye, J. Z., and Baross, J. A. (2000). High incidence of halotolerant bacteria in Pacific hydrothermal-vent and pelagic environments. FEMS Microbiol. Ecol. 32, 249–260.
Kaye, J. Z., Sylvan, J. B., Edwards, K. J., and Baross, J. A. (2011). Halomonas and Marinobacter ecotypes from hydrothermal vent, subseafloor and deep-sea environments. FEMS Microbiol. Ecol. 75, 123–133.
Kennedy, C. B., Scott, S. D., and Ferris, F. G. (2004). Hydrothermal phase stabilization of 2-line ferrihydrite by bacteria. Chem. Geol. 212, 269–277.
Kharroub, K., Aguilera, M., Luján, M., Jiménez-Pranteda, M. L., González-Paredes, A., Ramos-Cormenzana, A., et al. (2011). Marinobacter oulmenensis sp. nov., a moderately halophilic bacterium isolated from brine of a salt concentrator. Int. J. Syst. Evol. Microbiol. 61, 2210–2214.
Kim, B. Y., Weon, H. Y., Yoo, S. H., Kim, J. S., Kwon, S. W., Stackebrandt, E., et al. (2006). Marinobacter koreensis sp. nov., isolated from sea sand in Korea. Int. J. Syst. Evol. Microbiol. 56, 2653–2656.
Lee, O. O., Lai, P. Y., Wu, H. X., Zhou, X. J., Miao, L., Wang, H., and Qian, P. Y. (2012). Marinobacter xestospongiae sp. nov., isolated from the marine sponge Xestospongia testudinaria collected from the Red Sea. Int. J. Syst. Evol. Microbiol. 62, 1980–1985.
Liebgott, P. P., Casalot, L., Paillard, S., Lorquin, J., and Labat, M. (2006). Marinobacter vinifirmus sp. nov., a moderately halophilic bacterium isolated from a wine-barrel-decalcification wastewater. Int. J. Syst. Evol. Microbiol. 56, 2511–2516.
Liu, C., Chen, C. X., Zhang, X. Y., Yu, Y., Liu, A., Li, G. W., et al. (2012). Marinobacter antarcticus sp. nov., a halotolerant bacterium isolated from Antarctic intertidal sandy sediment. Int. J. Syst. Evol. Microbiol. 62, 1838–1844.
Lo, I., Denef, V. J., Verberkmoes, N. C., Shah, M. B., Goltsman, D., DiBartolo, G., et al. (2007). Strain-resolved community proteomics reveals recombining genomes of acidophilic bacteria. Nature 446, 537–541.
Lysnes, K., Thorseth, I. H., Steinsbu, B. O., Ovreas, L., Torsvik, T., and Pedersen, R. B. (2004). Microbial community diversity in seafloor basalt from the Arctic spreading ridges. FEMS Microbiol. Ecol. 50, 213–230.
Mahadevan, R., and Lovley, D. R. (2008). The degree of redundancy in metabolic genes is linked to mode of metabolism. Biophys. J. 94, 1216–1220.
Márquez, M. C., and Ventosa, A. (2005). Marinobacter hydrocarbonoclasticus Gauthier et al. 1992 and Marinobacter aquaeolei Nguyen et al. 1999 are heterotypic synonyms. Int. J. Syst. Evol. Microbiol. 55, 1349–1351.
Martín, S., Márquez, M. C., Sánchez-Porro, C., Mellado, E., Arahal, D. R., and Ventosa, A. (2003). Marinobacter lipolyticus sp. nov., a novel moderate halophile with lipolytic activity. Int. J. Syst. Evol. Microbiol. 53, 1383–1387.
Martin, W., Baross, J., Kelley, D., and Russell, M. J. (2008). Hydrothermal vents and the origin of life. Nat. Rev. Microbiol. 6, 805–814.
Martiny, J. B., Bohannan, B. J., Brown, J. H., Colwell, R. K., Fuhrman, J. A., Green, J. L., et al. (2006). Microbial biogeography: putting microorganisms on the map. Nat. Rev. Microbiol. 4, 102–112.
McCollom, T. M., and Shock, E. L. (1997). Geochemical constraints on chemolithoautotrophic metabolism by microorganisms in seafloor hydrothermal systems. Geochim. Cosmochim. Acta 61, 4375–4391.
Mills, R., Elderfield, H., and Thomson, J. (1993). A dual origin for the hydrothermal component in a metalliferous sediment core from the Mid-Atlantic Ridge. J. Geophys. Res. 98, 9671–9681.
Montes, M. J., Bozal, N., and Mercade, E. (2008). Marinobacter guineae sp. nov., a novel moderately halophilic bacterium from an Antarctic environment. Int. J. Syst. Evol. Microbiol. 58, 1346–1349.
Moran, M. A., Buchan, A., Gonzalez, J. M., Heidelberg, J. F., Whitman, W. B., Kiene, R. P., et al. (2004). Genome sequence of Silicibacter pomeroyi reveals adaptations to the marine environment. Nature 432, 910–913.
Moyer, C. L., and Morita, R. Y. (2007). “Psychrophiles and psychrotrophs,” In eLS. Chichester: John Wiley and Sons Ltd. doi: 10.1002/9780470015902.a0000402.pub2
Müller, M., Handley, K. M., Lloyd, J., Pancost, R. D., and Mills, R. A. (2010). Biogeochemical controls on microbial diversity in seafloor sulphidic sediments. Geobiology 8, 309–326.
Nerurkar, A. S., Hingurao, K. S., and Suthar, H. G. (2009). Bioemulsifiers from marine microorganisms. J. Sci. Ind. Res. 68, 273–277.
Pichler, T., and Veizer, J. (1999). Precipitation of Fe(III) oxyhydroxide deposits from shallow-water hydrothermal fluids in Tutum Bay, Ambitle Island, Papua New Guinea. Chem. Geol. 162, 15–31.
Polz, M. F., Hunt, D. E., Preheim, S. P., and Weinreich, D. M. (2006). Patterns and mechanisms of genetic and phenotypic differentiation in marine microbes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 361, 2009–2021.
Prudêncio, M., Pereira, A. S., Tavares, P., Besson, S., Cabrito, I., Brown, K., et al. (2000). Purification, characterization, and preliminary crystallographic study of copper-containing nitrous oxide reductase from Pseudomonas nautica 617. Biochemistry 39, 3899–3907.
Qu, L., Zhu, F., Zhang, J., Gao, C., and Sun, X. (2011). Marinobacter daqiaonensis sp. nov., a moderate halophile isolated from a Yellow Sea salt pond. Int. J. Syst. Evol. Microbiol. 61, 3003–3008.
Ram, R. J., Verberkmoes, N. C., Thelen, M. P., Tyson, G. W., Baker, B. J., Blake, R. C. I. I., et al. (2005). Community proteomics of a natural microbial biofilm. Science 308, 1915–1920.
Reysenbach, A. L., and Cady, S. L. (2001). Microbiology of ancient and modern hydrothermal systems. Trends Microbiol. 9, 79–86.
Rogers, D. R., Santelli, C. M., and Edwards, K. J. (2003). Geomicrobiology of deep-sea deposits: estimating community diversity from low-temperature seafloor rocks and minerals. Geobiology 1, 109–117.
Roh, S. W., Quan, Z. X., Nam, Y. D., Chang, H. W., Kim, K. H., Rhee, S. K., et al. (2008). Marinobacter goseongensis sp. nov., from seawater. Int. J. Syst. Evol. Microbiol. 58, 2866–2870.
Romanenko, L. A., Schumann, P., Rohde, M., Zhukova, N. V., Mikhailov, V. V., and Stackebrandt, E. (2005). Marinobacter bryozoorum sp. nov. and Marinobacter sediminum sp. nov., novel bacteria from the marine environment. Int. J. Syst. Evol. Microbiol. 55, 143–148.
Rontani, J. F., Gilewicz, M. J., Michotey, V. D., Zheng, T. L., Bonin, P. C., and Bertrand, J. C. (1997). Aerobic and anaerobic metabolism of 6, 10, 14-trimethylpentadecan-2-one by a denitrifying bacterium isolated from marine sediments. Appl. Environ. Microbiol. 63, 636–643.
Rontani, J. F., Mouzdahir, A., Michotey, V., Caumette, P., and Bonin, P. (2003). Production of a polyunsaturated isoprenoid wax ester during aerobic metabolism of squalene by Marinobacter squalenivorans sp. nov. Appl. Environ. Microbiol. 69, 4167–4176.
Shieh, W. Y., Jean, W. D., Lin, Y. T., and Tseng, M. (2003). Marinobacter lutaoensis sp. nov., a thermotolerant marine bacterium isolated from a coastal hot spring in Lutao, Taiwan. Can. J. Microbiol. 49, 244–252.
Shivaji, S., Gupta, P., Chaturvedi, P., Suresh, K., and Delille, D. (2005). Marinobacter maritimus sp. nov., a psychrotolerant strain isolated from sea water off the subantarctic Kerguelen islands. Int. J. Syst. Evol. Microbiol. 55, 1453–1456.
Sigalevich, P., Baev, M. V., Teske, A., and Cohen, Y. (2000). Sulfate reduction and possible aerobic metabolism of the sulfate-reducing bacterium Desulfovibrio oxyclinae in a chemostat coculture with Marinobacter sp. Strain MB under exposure to increasing oxygen concentrations. Appl. Environ. Microbiol. 66, 5013–5018.
Singer, E., Webb, E. A., Nelson, W. C., Heidelberg, J. F., Ivanova, N., Pati, A., et al. (2011). Genomic potential of Marinobacter aquaeolei, a biogeochemical “opportunitroph”. Appl. Environ. Microbiol. 77, 2763–2771.
Smith, P. A., and Cronan, D. S. (1983). The geochemistry of metalliferous sediments and waters associated with shallow submarine hydrothermal activity (Santorini, Aegean Sea). Chem. Geol. 39, 241–262.
Soberón-Chávez, G., and Maier, R. M. (2011). “Biosurfactants: a general overview,” in Biosurfactants, Vol. 20, ed G. Soberón-Chávez (Berlin: Springer), 1–11.
Takai, K., Moyer, C. L., Miyazaki, M., Nogi, Y., Hirayama, H., Nealson, K. H., et al. (2005). Marinobacter alkaliphilus sp. nov., a novel alkaliphilic bacterium isolated from subseafloor alkaline serpentine mud from Ocean Drilling Program Site 1200 at South Chamorro Seamount, Mariana Forearc. Extremophiles 9, 17–27.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., and Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739.
Thompson, J. D., Higgins, D. G., and Gibson, T. J. (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680.
Validation List no. 94. (2003). Validation of publication of new names and new combinations previously effectively published outside the IJSEM. Int. J. Syst. Evol. Microbiol. 53, 1701–1702.
Validation List no. 148. (2012). List of new names and new combinations previously effectively, but not validly, published. Int. J. Syst. Evol. Microbiol. 62, 2549–2554.
Wang, C. Y., Ng, C. C., Chen, T. W., Wu, S. J., and Shyu, Y. T. (2007). Microbial diversity analysis of former salterns in southern Taiwan by 16S rRNA-based methods. J. Basic Microbiol. 47, 525–533.
Wang, C. Y., Ng, C. C., Tzeng, W. S., and Shyu, Y. T. (2009). Marinobacter szutsaonensis sp. nov., isolated from a solar saltern. Int. J. Syst. Evol. Microbiol. 59, 2605–2609.
Wang, H., Li, H., Shao, Z., Liao, S., Johnstone, L., Rensing, C., et al. (2012). Genome sequence of deep-sea manganese-oxidizing bacterium Marinobacter manganoxydans MnI7–9. J. Bacteriol. 194, 899–900.
Whitman, W. B., Coleman, D. C., and Wiebe, W. J. (1998). Prokaryotes: the unseen majority. Proc. Natl. Acad. Sci. U.S.A. 95, 6578–6583.
Williams, K. (2009). Biosurfactants for cosmetic application: overcoming production challenges. Basic Biotech. 5, 78–83.
Williams, K. P., Gillespie, J. J., Sobral, B. W., Nordberg, E. K., Snyder, E. E., Shallom, J. M., et al. (2010). Phylogeny of Gammaproteobacteria. J. Bacteriol. 192, 2305–2314.
Xu, X. W., Wu, Y. H., Wang, C. S., Yang, J. Y., Oren, A., and Wu, M. (2008). Marinobacter pelagius sp. nov., a moderately halophilic bacterium. Int. J. Syst. Evol. Microbiol. 58, 637–640.
Yoon, J. H., Lee, M. H., Kang, S. J., and Oh, T. K. (2007). Marinobacter salicampi sp. nov., isolated from a marine solar saltern in Korea. Int. J. Syst. Evol. Microbiol. 57, 2102–2105.
Yoon, J. H., Shin, D. Y., Kim, I. G., Kang, K. H., and Park, Y. H. (2003). Marinobacter litoralis sp. nov., a moderately halophilic bacterium isolated from sea water from the East Sea in Korea. Int. J. Syst. Evol. Microbiol. 53, 563–568.
Yoon, J. H., Yeo, S. H., Kim, I. G., and Oh, T. K. (2004). Marinobacter flavimaris sp. nov. and Marinobacter daepoensis sp. nov., slightly halophilic organisms isolated from sea water of the Yellow Sea in Korea. Int. J. Syst. Evol. Microbiol. 54, 1799–1803.
Zhuang, D. C., Chen, Y. G., Zhang, Y. Q., Tang, S. K., Wu, X. L., Tan, Z. C., et al. (2009). Marinobacter zhanjiangensis sp. nov., a marine bacterium isolated from sea water of a tidal flat of the South China Sea. Antonie Van Leeuwenhoek 96, 295–301.
Keywords: Marinobacter, marine, hydrothermal, biogeochemical cycling, hydrocarbon, iron, arsenic, opportunistic
Citation: Handley KM and Lloyd JR (2013) Biogeochemical implications of the ubiquitous colonization of marine habitats and redox gradients by Marinobacter species. Front. Microbiol. 4: 136. doi: 10.3389/fmicb.2013.00136
Received: 15 February 2013; Accepted: 07 May 2013;
Published online: 22 May 2013.
Edited by:
Andreas Teske, University of North Carolina at Chapel Hill, USAReviewed by:
Purificacion Lopez-Garcia, Centre National de la Recherche Scientifique, FranceJuergen Wiegel, University of Georgia, USA
Copyright © 2013 Handley and Lloyd. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in other forums, provided the original authors and source are credited and subject to any copyright notices concerning any third-party graphics etc.
*Correspondence: Kim M. Handley, Searle Chemistry Laboratory, Computation Institute, University of Chicago, 5735 South Ellis Avenue, Chicago, IL 60637, USA. e-mail:a21oYW5kbGV5QHVjaGljYWdvLmVkdQ==