- 1Institute of Biology, Department of Genetics, Martin Luther University Halle-Wittenberg, Halle, Germany
- 2Leibniz Institute of Plant Biochemistry, Halle, Germany
Pathogenicity of the Gram-negative plant-pathogenic bacterium Xanthomonas campestris pv. vesicatoria depends on a type III secretion (T3S) system which translocates effector proteins into plant cells. Effector protein delivery is controlled by the T3S chaperone HpaB, which presumably escorts effector proteins to the secretion apparatus. One intensively studied effector is the transcription activator-like (TAL) effector AvrBs3, which binds to promoter sequences of plant target genes and activates plant gene expression. It was previously reported that type III-dependent delivery of AvrBs3 depends on the N-terminal protein region. The signals that control T3S and translocation of AvrBs3, however, have not yet been characterized. In the present study, we show that T3S and translocation of AvrBs3 depend on the N-terminal 10 and 50 amino acids, respectively. Furthermore, we provide experimental evidence that additional signals in the N-terminal 30 amino acids and the region between amino acids 64 and 152 promote translocation of AvrBs3 in the absence of HpaB. Unexpectedly, in vivo translocation assays revealed that AvrBs3 is delivered into plant cells even in the absence of HrpF, which is the predicted channel-forming component of the T3S translocon in the plant plasma membrane. The presence of HpaB- and HrpF-independent transport routes suggests that the delivery of AvrBs3 is initiated during early stages of the infection process, presumably before the activation of HpaB or the insertion of the translocon into the plant plasma membrane.
Introduction
Many Gram-negative bacterial pathogens translocate effector proteins into eukaryotic host cells to modulate host cellular pathways such as defense responses to their own benefit (Raymond et al., 2013; Ashida et al., 2015; Santos and Finlay, 2015; Büttner, 2016; Ensminger, 2016; Grabowski et al., 2017). Effector protein translocation often depends on the type III secretion (T3S) system, which is an essential pathogenicity factor of many bacterial species and is related to the bacterial flagellum (Büttner, 2012; Diepold and Armitage, 2015). At least eight components of T3S systems are conserved in different bacterial species and were designated Sct (secretion and cellular translocation) proteins in animal-pathogenic bacteria followed by a letter, which indicates the corresponding T3S system components from Yersinia species (Hueck, 1998; Deng et al., 2017). Sct proteins are mainly involved in the assembly of the membrane-spanning part of the secretion apparatus, which consists of ring structures in the inner and outer bacterial membrane and a predicted periplasmic inner rod (Büttner, 2012). The inner membrane rings associate with the export apparatus, which is assembled by five transmembrane proteins and presumably forms a transport channel for secreted proteins (Diepold et al., 2011; Büttner, 2012; Dietsche et al., 2016; Deng et al., 2017). Components of the export apparatus are connected with the cytoplasmic ATPase complex, which provides the energy for secretion and/or unfolds secreted proteins during transport (Büttner, 2012; Deng et al., 2017). The ATPase presumably interacts with members of the SctQ family, which assemble as cytoplasmic ring or pod-like structures and are potential docking sites for T3S substrates (Büttner, 2012; Deng et al., 2017; Hu et al., 2017).
Extracellular components of translocation-associated T3S systems from animal-pathogenic bacteria include the T3S needle, which serves as a conduit for T3S substrates to the host-pathogen interface. T3S needles are 40–80 nm long and thus significantly shorter than the T3S pilus from plant-pathogenic bacteria, which reaches a length of up to 2 μm and spans the plant cell wall (Büttner, 2012). Needle and pilus are directly or indirectly connected to the T3S translocon, which inserts as a channel-like complex into the eukaryotic plasma membrane and mediates effector protein translocation (Mueller et al., 2008; Mattei et al., 2011; Galan et al., 2014). T3S translocons from animal-pathogenic bacteria often consist of two conserved hydrophobic translocators, which form the transmembrane channel, and a third hydrophilic translocator, which presumably provides an assembly platform for the translocon channel at the tip of the needle (Mattei et al., 2011). Notably, translocon-dependent protein delivery into eukaryotic cells does not appear to be restricted to proteins that travel inside the T3S system but was also reported for external proteins. Thus, the effector protein YopH from Yersinia spp. is delivered by the T3S translocon when present on the bacterial surface (Akopyan et al., 2011). Similarly, translocon-dependent delivery was reported for the type V-secreted autotransporter EspC from enteropathogenic Escherichia coli, which interacts with translocon proteins, suggesting that the T3S translocon also transports T3S-unrelated proteins (Vidal and Navarro-Garcia, 2008; Tejeda-Dominguez et al., 2017). The translocon is usually essential for type III-dependent effector protein delivery, however, several type III effector proteins from animal-pathogenic bacteria including SspH1 from Salmonella spp. and YopM from Yersinia spp. can also enter eukaryotic cells independently of the translocon, presumably via endocytosis or direct transport across the plasma membrane (Rüter et al., 2010; Scharnert et al., 2013; Lubos et al., 2014). A similar mechanism has not yet been reported for effector proteins from plant-pathogenic bacteria.
In contrast to T3S translocons from animal-pathogenic bacteria, the predicted translocon from plant-pathogenic bacteria is less well conserved among different species and the mechanisms underlying effector protein translocation are not yet understood. In Xanthomonas campestris pv. vesicatoria (also designated Xanthomonas euvesicatoria), the secreted HrpF protein was identified as putative channel-forming translocon protein (Büttner et al., 2002). Mutant studies and in vivo translocation assays suggest that HrpF is essential for pathogenicity of X. campestris pv. vesicatoria and effector protein translocation (Büttner et al., 2002; Hotson et al., 2003; Jiang et al., 2009; Teper et al., 2016). Homologous proteins are present in many Xanthomonas species and Ralstonia solanacearum. Furthermore, HrpF shares limited sequence similarity with the putative translocon protein HrpK1 from Pseudomonas syringae (Kvitko et al., 2007). HrpK1 contributes to but is not essential for pathogenicity of P. syringae as well as for efficient effector protein delivery and HR induction on tobacco plants (Petnicki-Ocwieja et al., 2005; Kvitko et al., 2007). Accessory hydrophilic proteins, designated harpins, presumably contribute to effector protein translocation in P. syringae and were also identified in other bacterial species (Kvitko et al., 2007; Choi et al., 2013; Ji and Dong, 2015).
X. campestris pv. vesicatoria, which causes bacterial spot disease in pepper and tomato plants, is one of the model systems for the analysis of T3S systems in plant-pathogenic bacteria (Jones et al., 2004; Potnis et al., 2015). The T3S system from X. campestris pv. vesicatoria is essential for pathogenicity and is encoded by the chromosomal hrp (hypersensitive response and pathogenicity) gene cluster (Büttner and Bonas, 2002). Eleven hrp gene products, referred to as Hrc (Hrp conserved), are conserved in animal- and/or plant-pathogenic bacteria and presumably constitute the core components of the secretion apparatus (Büttner and Bonas, 2002). Additional components are encoded by non-conserved hrp genes and include for instance the predicted inner rod proteins HrpB1 and HrpB2, which presumably form an assembly platform for the T3S pilus, the pilus protein HrpE and the translocon protein HrpF (Büttner et al., 2002; Weber et al., 2005; Hartmann et al., 2012; Hausner et al., 2013). An additional protein, which might contribute to effector delivery, is the secreted XopA protein, which is encoded in the flanking region of the hrp gene cluster (Noël et al., 2002).
The hrp gene cluster also encodes Hpa (Hrp-associated) proteins which contribute to but are not essential for type III-dependent effector protein translocation. One example is the general T3S chaperone HpaB, which is essential for pathogenicity and promotes effector protein translocation (Büttner et al., 2004). HpaB presumably targets effectors to the ATPase HrcN of the T3S system and is involved in the recognition of translocation signals (Büttner et al., 2006; Lorenz and Büttner, 2009; Scheibner et al., 2017). The activity of HpaB is controlled by its secreted regulator HpaA, which interacts with and likely inhibits HpaB. Secretion of HpaA presumably liberates HpaB and thus activates effector protein secretion (Lorenz et al., 2008).
The T3S system from X. campestris pv. vesicatoria translocates more than 30 effector proteins into plant cells (Büttner et al., 2006, 2007; Szczesny et al., 2010a; Schulze et al., 2012). In many cases, the precise biochemical functions and plant targets of effectors are still unknown (Büttner and Bonas, 2010). One of the best studied type III effectors from X. campestris pv. vesicatoria is AvrBs3, which is a member of the transcription activator-like (TAL) effector family and acts as a transcription factor in plant cells (Boch and Bonas, 2010). TAL effectors contain C-terminal nuclear localization signals (NLSs) and are imported into the plant cell nucleus (Boch and Bonas, 2010). The central region of TAL effectors consists of a variable number of amino acid repeats with a predominant length of 34 amino acids and mediates DNA binding (Boch and Bonas, 2010). The repeats are almost sequence-identical with the exception of two variable residues (RVDs, repeat variable diresidues) at amino acid positions 12 and 13 of each repeat (Boch and Bonas, 2010). The RVDs allow the base-specific binding of TAL effectors to sequences in the promoter regions of plant target genes (Boch et al., 2009; Moscou and Bogdanove, 2009; Deng et al., 2012; Mak et al., 2012). The subsequent modulation of plant gene expression by TAL effectors depends on their C-terminal acidic activation domain (AAD) (Boch and Bonas, 2010). Among the plant target genes is the Bs3 resistance gene, which is present in AvrBs3-responsive pepper plants and encodes a flavin monooxygenase. Bs3 initiates the induction of a hypersensitive response (HR), which is a local rapid cell death at the infection site and presumably restricts bacterial multiplication (Römer et al., 2009; Wu et al., 2014).
Delivery of AvrBs3 by the T3S system depends on the N-terminal 152 amino acids which presumably contain the export signal (Szurek et al., 2002; Noël et al., 2003). We previously reported that the N-terminal 50 amino acids of AvrBs3 are sufficient for T3S (Büttner et al., 2004). Yet, the precise location of T3S and translocation signals in AvrBs3 has not been determined. Secretion and translocation signals are usually located in the N-terminal regions of T3S substrates and are not conserved on the amino acid level (Sory et al., 1995; Schesser et al., 1996; Rüssmann et al., 2002; Schechter et al., 2004; Arnold et al., 2009). However, they often contain specific amino acid compositions or patterns such as a high content of polar amino acids as was reported for effector proteins from the plant-pathogenic bacterium P. syringae (Guttman et al., 2002; Petnicki-Ocwieja et al., 2002; Greenberg and Vinatzer, 2003; Schechter et al., 2004, 2006; Arnold et al., 2009; Löwer and Schneider, 2009; Samudrala et al., 2009; Buchko et al., 2010). Evidence for an essential role of characteristic amino acids for T3S or translocation, however, is still missing (Schechter et al., 2012).
In the present study, we localized T3S and translocation signals in the TAL effector AvrBs3. The results of in vitro T3S and in vivo translocation assays revealed that AvrBs3 contains separate signals, which control T3S and translocation in the presence or absence of the T3S chaperone HpaB. Furthermore, the analysis of AvrBs3 translocation by HR- and fluorescence based translocation assays showed that AvrBs3 enters plant cells in the absence of a functional translocon, suggesting that it is translocated during early stages of the infection process prior to the assembly of the translocon.
Materials and Methods
Bacterial Strains and Growth Conditions
Bacterial strains and plasmids used in this study are listed in Table 1. Escherichia coli strains were cultivated at 37°C in lysogeny broth (LB) medium and X. campestris pv. vesicatoria strains at 30°C in nutrient-yeast extract-glycerol (NYG) medium (Daniels et al., 1984) or minimal medium A (Ausubel et al., 1996) at pH 7.0 supplemented with 10 mM sucrose and 0.3% casamino acids.
Plant Material and Plant Inoculations
X. campestris pv. vesicatoria strains were inoculated into leaves of the near-isogenic pepper cultivars Early Cal Wonder (ECW), ECW-10R and ECW-30R, and Bs3- or gfp-transgenic Nicotiana benthamiana plants at concentrations of 4 × 108 colony-forming units (CFU) ml−1 in 1 mM MgCl2 if not stated otherwise (Minsavage et al., 1990; Bonas et al., 1991; Kousik and Ritchie, 1998; Scheibner et al., 2016). Agrobacterium tumefaciens strains were infiltrated in 1 mM MgCl2 at a density of 8 × 108 CFU ml−1. Infected pepper plants were incubated for 16 h of light at 28°C and 65% humidity, and 8 h of darkness at 22°C and 65% humidity. N. benthamiana plants were incubated for 16 h of light at 20°C and 75% humidity, and 8 h of darkness at 18°C and 70% humidity. The appearance of plant reactions was scored over a period of one to 12 dpi. For the better visualization of the HR, leaves were destained in 70% ethanol. Experiments were repeated at least twice; representative plant reactions are shown.
Generation of Expression Constructs
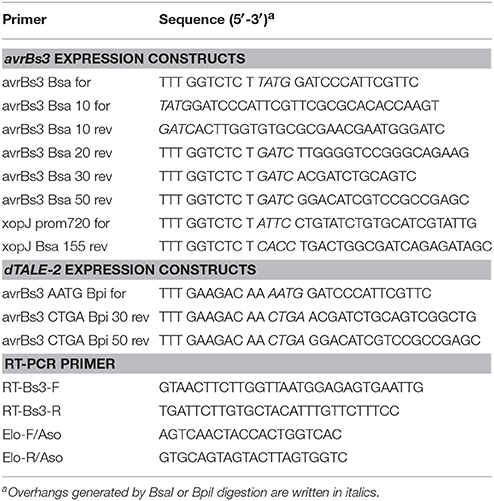
For the generation of expression constructs encoding AvrBs3Δ2 fusion proteins under control of the lac promoter, avrBs3 gene fragments were amplified by PCR from X. campestris pv. vesicatoria and cloned into the BsaI sites of plasmid pBR356, which contains the avrBs3Δ2 reporter gene (Scheibner et al., 2016). For the generation of constructs encoding AvrBs3Δ63 or AvrBs3Δ63 fusion proteins, modules encoding N-terminal, central and C-terminal regions of AvrBs3Δ63 and the respective fusion partners were cloned into the BsaI sites of plasmid pBRM. The stop codon of avrBs3 was included in these constructs. xopJ1-155-avrBs3Δ2 and xopJ1-155-avrBs3Δ63 were expressed under control of the native xopJ promoter in plasmid pBRM-P. For this, fragments containing the first 155 codons of xopJ and 720 bp of the upstream region including the xopJ promoter (Noël et al., 2003) were ligated with fragments encoding the N-terminal, central and C-terminal regions of AvrBs3Δ63.
For the generation of binary constructs for A. tumefaciens-mediated transient gene expression, avrBs31-10 and avrBs31-20 were amplified by PCR and ligated with modules encoding N-terminal, central and C-terminal regions of AvrBs3Δ2 into the BsaI sites of the binary vector pGGA3 downstream of the 35S promoter. Similarly, modules encoding N-terminal, central and C-terminal regions of AvrBs3 were cloned into pGGA3. The module encoding the C-terminal region of AvrBs3 lacked the native stop codon and was therefore expressed in fusion with a FLAG epitope-encoding sequence provided by vector pGGA3.
Expression constructs encoding dTALE-2 and derivatives thereof were generated by Golden-Gate assembly of individual DNA modules encoding N-terminal, central and C-terminal regions of dTALE-2, dTALE-2ΔN as well as N-terminal regions of the respective fusion partners. All primers used in this study are listed in Table 2. Plasmids were introduced into X. campestris pv. vesicatoria by electroporation or by conjugation using pRK2013 as a helper plasmid in triparental matings (Figurski and Helinski, 1979).
Generation of X. campestris pv. vesicatoria Deletion Mutants
For the generation of X. campestris pv. vesicatoria deletion mutants (see Table 1), derivatives of the suicide vector pOK1, which contained the flanking regions of the deleted genes, were transferred to X. campestris pv. vesicatoria recipient strains by triparental conjugation. Double crossovers resulted in deletion mutants which were selected as described previously (Huguet et al., 1998).
Analysis of Protein Extracts and in Vitro Secretion Assays
For protein analysis, bacteria were cultivated over-night in liquid NYG medium and cells were harvested by centrifugation. Equal amounts of proteins adjusted according to the optical densities of the cultures were analyzed by immunoblotting, using AvrBs3- or HrpF-specific antibodies. For the analysis of in vitro T3S, bacteria were incubated in MA medium at pH 5.3 for 1.5 h in the presence of thiamine and bovine serum albumin as described previously (Rossier et al., 1999). Secreted proteins were separated from bacterial cells by filtration and precipitated by trichloroacetic acid. Equal amounts of bacterial total cell extracts and culture supernatants (adjusted according to the optical densities of the cultures) were analyzed by SDS-PAGE and immunoblotting using antibodies specific for AvrBs3, the translocon protein HrpF, the inner membrane ring protein HrcJ and the predicted periplasmic inner rod protein HrpB1, respectively (Knoop et al., 1991; Rossier et al., 2000; Büttner et al., 2002). Horseradish peroxidase-labeled anti-rabbit antibodies (GE Healthcare) were used as secondary antibodies. Experiments were repeated twice.
RNA Analysis
For transcript analysis via semiquantitative reverse-transcription PCR (RT-PCR), strain 85* and derivatives thereof ectopically expressing avrBs3 were inoculated at concentrations of 4 × 108 CFU ml−1 and derivatives of strain 82-8 at concentrations of 8 × 108 CFU ml−1 into leaves of ECW-30R pepper plants or Bs3-transgenic N. benthamiana plants. 16 leaf discs (2 mm diameter) were harvested 8 h (for derivatives of strain 85*) or 24 h (for derivatives of strain 82-8) post infiltration, pooled and immediately frozen in liquid nitrogen. RNA was isolated with the QIAGEN RNeasy Plant Mini Kit, and samples were treated with DNase I (Roche) for 30 min. cDNA was synthesized from 2 μg of total RNA with the RevertAid H Minus First Strand cDNA synthesis kit (Thermo Scientific) using random hexamer primers. Four microlitres of a 1:50 diluted cDNA solution were used as template in an RT-PCR with up to 40 cycles of denaturation, annealing and elongation. The constitutively expressed gene Elongation factor 1α (EF1α) was amplified as control (Römer et al., 2007). To exclude contaminations by genomic DNA, all reactions were also performed in the absence of reverse transcriptase. The experiments were repeated at least twice with similar results.
Results
The N-Terminal 10 Amino Acids of AvrBs3 Contain a T3S Signal
To localize the T3S signal in AvrBs3, we generated fusion proteins consisting of the first 10, 20, or 30 amino acids, respectively, of AvrBs3 and AvrBs3Δ2, which is an N-terminal deletion derivative of AvrBs3. AvrBs3Δ2 lacks amino acids 2–152 and thus the T3S and translocation signal. However, AvrBs3Δ2 contains the effector domain and induces the HR in AvrBs3-responsive plants when delivered as fusion partner of a functional translocation signal (Szurek et al., 2002; Noël et al., 2003; Drehkopf et al., 2017). AvrBs3Δ2 fusion proteins were analyzed in X. campestris pv. vesicatoria strain 85*, which is a derivative of the wild-type strain 85-10 and contains HrpG*, a constitutively active version of the hrp gene regulator HrpG (Rossier et al., 1999; Wengelnik et al., 1999). When bacteria were cultivated in secretion medium, AvrBs31−10-, AvrBs31−20- and AvrBs31−30-AvrBs3Δ2 fusion proteins were detected in the culture supernatants (Figure 1A). Similar results were obtained for strain 85*ΔhpaB, which lacks the T3S chaperone HpaB (Figure 1A; Büttner et al., 2004). As controls, we analyzed the inner membrane-associated protein HrcJ and the predicted periplasmic inner rod protein HrpB1. Both proteins were only detected in cell extracts of strains 85* and 85*ΔhpaB, suggesting that no cell lysis had occurred (Figure 1A). We conclude from these data that the N-terminal 10–30 amino acids of AvrBs3 target AvrBs3Δ2 for secretion, even in the absence of the T3S chaperone HpaB. This is in contrast to the full-length AvrBs3 protein, which depends on HpaB for efficient secretion (Büttner et al., 2004).
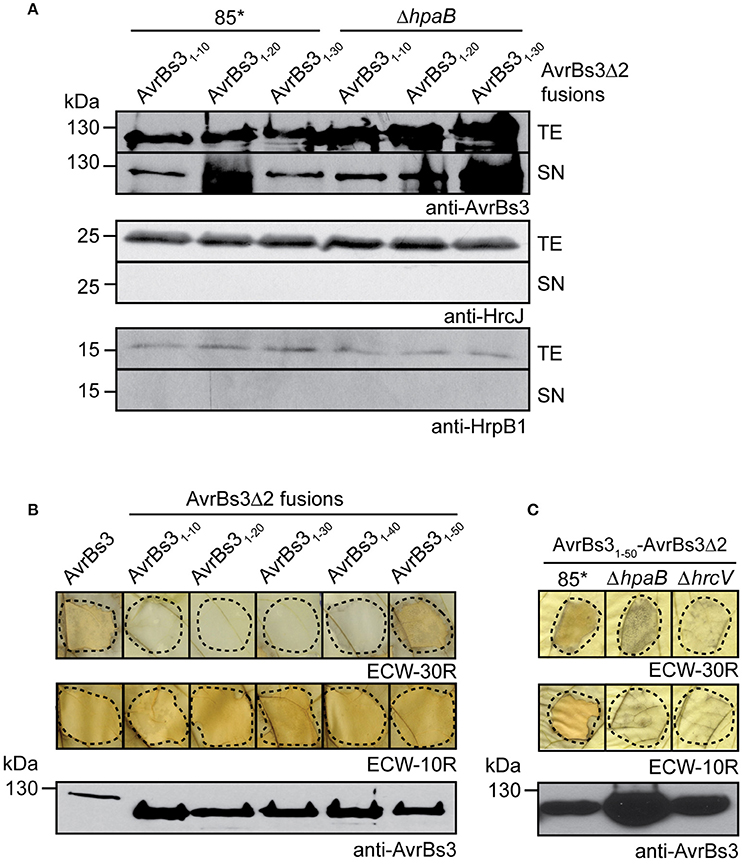
Figure 1. Localization of T3S and translocation signals in AvrBs3. (A) The N-terminal 10 amino acids of AvrBs3 target AvrBs3Δ2 for secretion in wild-type and hpaB deletion mutant strains. X. campestris pv. vesicatoria strains 85* and 85*ΔhpaB (ΔhpaB) ectopically expressing avrBs3Δ2 fusions as indicated were incubated in secretion medium. Total cell extracts (TE) and culture supernatants (SN) were analyzed by immunoblotting, using AvrBs3-specific antibodies. All AvrBs3Δ2 fusions were reproducibly secreted by both strains, the relative secretion levels, however, varied in different experiments. As control, the blots were reprobed with antibodies against the inner membrane protein HrcJ and the periplasmic predicted inner rod protein HrpB1. (B) The N-terminal 50 amino acids of AvrBs3 contain the translocation signal. Strain 85* ectopically expressing avrBs3 or avrBs3Δ2 fusions as indicated was infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. For the better visualization of the HR, leaves were destained in ethanol 3 dpi. Dashed lines indicate the infiltrated areas. Equal amounts of cell extracts were analyzed by immunoblotting using an AvrBs3-specific antiserum. (C) Translocation of AvrBs31−50-AvrBs3Δ2 depends on the T3S system. Strains 85*, 85*ΔhpaB (ΔhpaB) and the T3S-deficient strain 85*ΔhrcV (ΔhrcV) ectopically expressing avrBs31−50-avrBs3Δ2 were infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. Plant reactions and protein synthesis were analyzed as described in (B).
The N-Terminal 50 Amino Acids of AvrBs3 Contain a Translocation Signal
For the analysis of translocation signals in AvrBs3, we performed in vivo translocation assays with AvrBs3Δ2 fusion proteins. For this, strain 85* ectopically expressing individual avrBs3Δ2 fusions was infiltrated into leaves of AvrBs3-responsive ECW-30R pepper plants. AvrBs31−50-AvrBs3Δ2 induced the HR in AvrBs3-responsive ECW-30R pepper plants when delivered by strain 85*, suggesting that the N-terminal 50 amino acids contain a functional translocation signal (Figure 1B). HR induction by AvrBs31−50-AvrBs3Δ2 was dependent on the T3S system because it was macroscopically not detectable when AvrBs31−50-AvrBs3Δ2 was analyzed in the T3S-deficient strain 85*ΔhrcV (Figure 1C). In contrast to AvrBs31−50-AvrBs3Δ2, AvrBs3Δ2 fusions containing the N-terminal 10, 20, 30, or 40 amino acids of AvrBs3 did not induce a visible HR when analyzed in strain 85*, suggesting that they were not or not efficiently translocated (Figure 1B). Immunoblot analysis of bacterial cell extracts revealed that all fusion proteins were stably synthesized (Figure 1B).
As control, bacteria were infiltrated into leaves of ECW-10R pepper plants, which contain the resistance gene Bs1 and initiate the HR upon recognition of the effector protein AvrBs1 (Minsavage et al., 1990). All strains induced the AvrBs1-specific HR in leaves of ECW-10R plants, indicating that AvrBs1 was efficiently translocated. This suggests that the AvrBs3Δ2 fusion proteins did not interfere with the activity of the T3S system (Figure 1B).
The N-Terminal 30 Amino Acids of AvrBs3 Target the AvrBs3Δ2 Reporter for Translocation in the Absence of the T3S Chaperone HpaB
Next, we investigated the influence of the T3S chaperone HpaB on the translocation of AvrBs3Δ2 fusion proteins. Notably, AvrBs31−30-AvrBs3Δ2 was translocated by strain 85*ΔhpaB but did not induce a macroscopically visible HR in AvrBs3-responsive pepper plants when analyzed in strain 85* or the T3S-deficient strain 85*ΔhpaBΔhrcV (Figures 2A,B; see above). Similar data were obtained for the hrpG wild-type strain 85-10ΔhpaB, suggesting that the translocation of AvrBs31−30-AvrBs3Δ2 by hpaB deletion mutants was not caused by the overexpression of the T3S genes in the presence of HrpG* (Figure S1). Hence, the N-terminal 30 amino acids of AvrBs3 contain a translocation signal which is recognized in hpaB deletion mutants but is not sufficient for efficient translocation in the wild-type strain. This signal will hereafter be referred to as minimal translocation signal. Translocation in the absence of HpaB was also observed for AvrBs31−40-AvrBs3Δ2. In contrast, translocation of AvrBs31−50-AvrBs3Δ2 was significantly reduced in strain 85*ΔhpaB (Figure 2A) and not detectable in the hrpG wild-type strain 85-10ΔhpaB (Figure S1), suggesting that the N-terminal 50 amino acids of AvrBs3 depend on HpaB for efficient translocation.
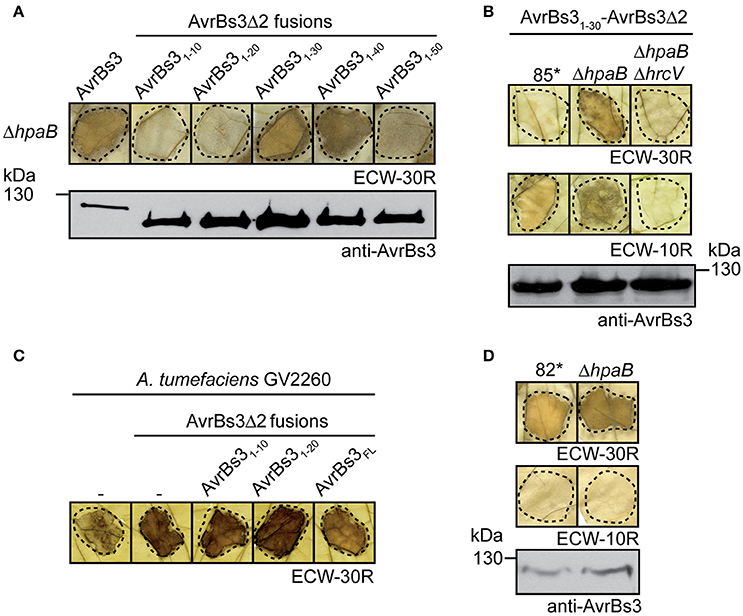
Figure 2. Influence of HpaB on the translocation of AvrBs3 reporter fusions. (A) The N-terminal 30 amino acids of AvrBs3 target the AvrBs3Δ2 reporter for translocation in the absence of HpaB. Strain 85*ΔhpaB (ΔhpaB) ectopically expressing avrBs3 or avrBs3Δ2 fusions as indicated was infiltrated into leaves of AvrBs3-responsive ECW-30R pepper plants. For the better visualization of the HR, leaves were destained in ethanol 3 dpi. Dashed lines indicate the infiltrated areas. Equal amounts of cell extracts were analyzed by immunoblotting using an AvrBs3-specific antiserum. (B) Translocation of AvrBs31−30-AvrBs3Δ2 by hpaB mutants depends on the T3S system. Strains 85*, 85*ΔhpaB (ΔhpaB), and 85*ΔhpaBΔhrcV (ΔhpaBΔhrcV) ectopically expressing avrBs31-30-avrBs3Δ2 were infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. Plant reactions and protein synthesis were analyzed as described in (A). The HR in ECW-30R plants was specifically induced by AvrBs31−30-AvrBs3Δ2 and was not observed after infiltration of X. campestris pv. vesicatoria strains without expression constructs (Figure S1). (C) Transient expression of avrBs3Δ2 fusions induces the HR in AvrBs3-responsive pepper plants. A. tumefaciens without expression construct (−) or ectopically expressing avrBs3 or avrBs3Δ2 fusions as indicated was infiltrated at a density of 8 × 108 CFU ml−1 into leaves of AvrBs3-responsive pepper plants. Leaves were destained in ethanol 4 dpi. (D) AvrBs3 is translocated in the absence of the T3S chaperone HpaB. Strains 82* and 82*ΔhpaB (ΔhpaB) were infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. Plant reactions and protein synthesis were analyzed as described in (A).
AvrBs3Δ2 fusion proteins containing the N-terminal 10 or 20 amino acids of AvrBs3 did not induce a visible HR when analyzed in strain 85*ΔhpaB (Figure 2A). To exclude that the N-terminal 10 or 20 amino acids of AvrBs3 interfere with the activity of the AvrBs3Δ2 reporter, both gene fusions were expressed under control of the 35S promoter in AvrBs3-responsive pepper plants after A. tumefaciens-mediated gene delivery. Transient expression of avrBs31-10- and avrBs31-20-avrBs3Δ2 led to the induction of the Bs3-specific HR, indicating that the AvrBs3Δ2 reporter was functional when present as a fusion partner of the N-terminal 10 or 20 amino acids of AvrBs3 (Figure 2C). Thus, the lack of HR induction by AvrBs31−10- and AvrBs31−20-AvrBs3Δ2 fusion proteins was presumably not caused by a misfolding of the reporter. Taken together, we conclude from these data that the N-terminal 50 amino acids of AvrBs3 contain a translocation signal whereas the N-terminal 30 amino acids promote translocation of AvrBs3 in the absence of HpaB. The N-terminal 10 or 20 amino acids did not target the AvrBs3Δ2 reporter for detectable translocation, however, it cannot be excluded that this region contains a translocation signal which is inactive in the context of the AvrBs3Δ2 fusion.
Amino Acids 64–152 Promote Translocation of AvrBs3
In contrast to AvrBs31−50-AvrBs3Δ2, the full-length AvrBs3 protein induced the HR when analyzed in strains 85*ΔhpaB and 85-10ΔhpaB, suggesting that it was efficiently translocated in the absence of HpaB (Figure 2A; Figure S1). Similar findings were observed for strain 82*ΔhpaB, which is a derivative of strain 82-8 and naturally expresses avrBs3 (Figure 2D). This indicates that the HpaB-independent translocation of AvrBs3 was not caused by the overexpression of T3S genes or of avrBs3. We, therefore, wondered whether the differences in HR induction by AvrBs3 and AvrBs31−50-AvrBs3Δ2 were caused by differences in protein activities. Previous reporter assays revealed that the N-terminal deletion derivative AvrBs3Δ2 led to reduced activation of an AvrBs3-responsive promoter when compared with the full-length AvrBs3 protein (Schreiber et al., 2015). In contrast, transcription activation by an AvrBs3 derivative, which was deleted in the N-terminal 63 amino acids, was like wild type, suggesting that the region between amino acids 63 and 152 of AvrBs3 contributes to protein activity (Schreiber et al., 2015). To investigate whether different activities of AvrBs3 derivatives result in different HR intensities, we compared the HR induction by AvrBs3Δ2 and AvrBs3Δ63 fusion proteins. As fusion partner, which provides the T3S and translocation signal, we chose the N-terminal 155 amino acids of the effector protein XopJ (Noël et al., 2003). When strain 85* or 85-10 delivering XopJ1−155-AvrBs3Δ2 or XopJ1−155-AvrBs3Δ63 was infiltrated into leaves of AvrBs3-responsive pepper plants, no differences in the HR intensities were detected, even when bacteria were infiltrated at lower optical densities (Figure 3A). As both fusion proteins were stably synthesized at comparable levels (Figure 3A), we conclude that the previously observed reduced transcription activation activity of AvrBs3Δ2 did not lead to a macroscopically detectable reduction of the AvrBs3-induced HR. The efficient translocation of AvrBs3 by hpaB deletion mutants might, therefore, be caused by the presence of additional export signals which promote translocation in the absence of HpaB and are absent in AvrBs3Δ2 fusion proteins.
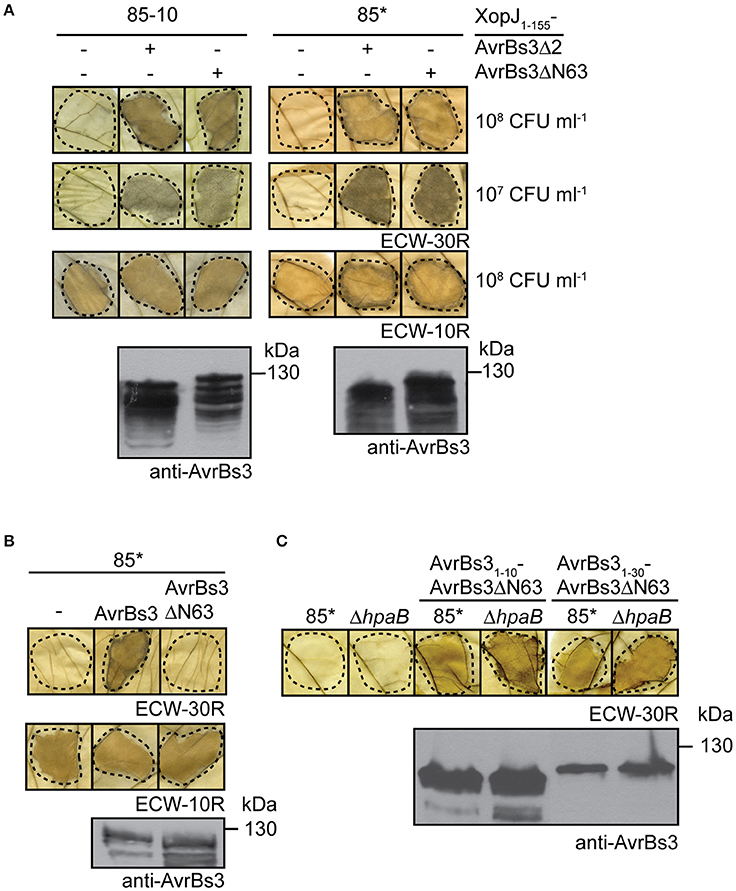
Figure 3. AvrBs3ΔN63 contains a signal that promotes translocation. (A) Translocation assays with AvrBs3Δ2 and AvrBs3ΔN63 fusion proteins. Strains 85-10 and 85* without expression construct (−) or ectopically expressing xopJ1-155-avrBs3Δ2 or xopJ1-155-avrBs3ΔN63 were infiltrated at densities of 108 and 107 CFU ml−1 into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants as indicated. Leaves were destained in ethanol 3 dpi. Dashed lines indicate the infiltrated areas. Equal amounts of cell extracts were analyzed by immunoblotting using an AvrBs3-specific antiserum. (B) AvrBs3ΔN63 does not induce a macroscopic HR when analyzed in strain 85*. Strain 85* without expression construct (−) or ectopically expressing avrBs3 or avrBs3ΔN63 as indicated was infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. Plant reactions and protein synthesis were analyzed as described in (A). (C) The N-terminal 10 amino acids of AvrBs3 target AvrBs3ΔN63 for translocation in wild-type and hpaB mutant strains. Strains 85* and 85*ΔhpaB (ΔhpaB) without expression construct or ectopically expressing avrBs31-10-avrBs3ΔN63 or avrBs31-30-avrBs3ΔN63 as indicated were infiltrated at a density of 108 CFU ml−1 into leaves of AvrBs3-responsive pepper plants. Leaves were destained in ethanol 2 dpi. Dashed lines indicate the infiltrated areas. Protein synthesis was analyzed as described in (A).
To investigate the presence of additional export signals outside the N-terminal 50 amino acids of AvrBs3, we analyzed the translocation of fusion proteins between the N-terminal 10 or 30 amino acids of AvrBs3 and AvrBs3Δ63, which lacks the T3S and translocation signal (Schreiber et al., 2015; Scheibner et al., 2016; Figure 3B). Notably, AvrBs31−10- and AvrBs31−30-AvrBs3Δ63 induced the AvrBs3-specific HR when delivered by strains 85* and 85*ΔhpaB (Figure 3C). This is in contrast to the corresponding AvrBs3Δ2 fusions and suggests that the region between amino acids 64 and 152 of AvrBs3 contains a signal which promotes translocation even in the absence of HpaB.
AvrBs3 Is Delivered into Plant Cells in the Absence of the Translocon Protein HrpF
It was previously reported that the translocation of effector proteins from Xanthomonas spp. depends on the translocon protein HrpF (Büttner et al., 2002; Szurek et al., 2002; Hotson et al., 2003; Thieme et al., 2007; Jiang et al., 2009; Teper et al., 2016). To confirm the contribution of HrpF to the translocation of AvrBs3, we performed translocation studies with hrpF deletion mutants. Unexpectedly, strain 85*ΔhrpF ectopically expressing avrBs3 induced a browning of the infected leaf tissue, which was visible after destaining of the leaves in ethanol 3 dpi (Figure 4A). A similar phenotype was observed with strain 85*ΔhrpFΔxopA, which additionally lacks the secreted XopA protein (Figure 4A; Noël et al., 2002). XopA contributes to pathogenicity and is homologous to harpins, which are proposed to be involved in effector protein translocation (Noël et al., 2002; Kim et al., 2004). No browning of the leaf tissue was visible when avrBs3 was ectopically expressed in strains 85*ΔhrpE and 85*ΔhrcN, which are deleted in the pilus gene hrpE and the ATPase gene hrcN, respectively (Figure 4A).
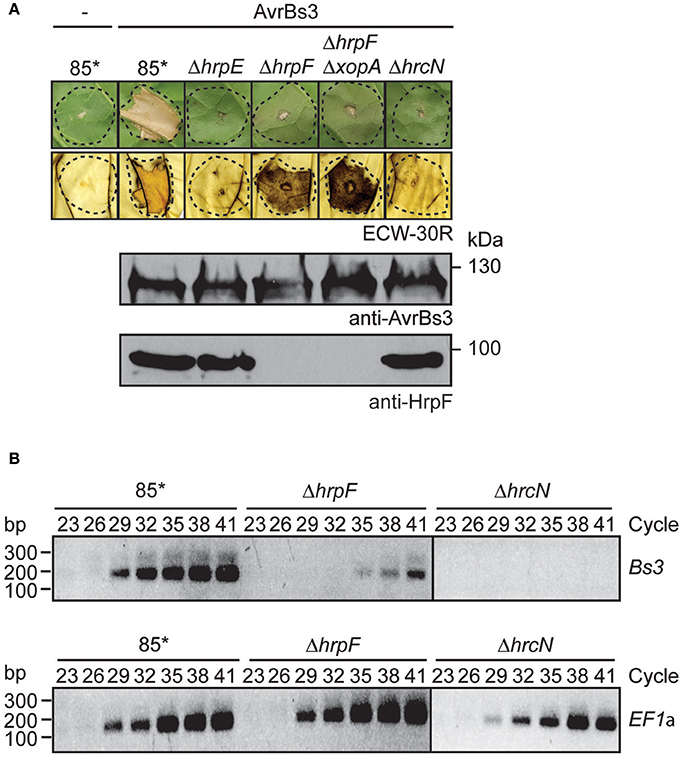
Figure 4. AvrBs3 enters plant cells in the absence of a functional translocon. (A) HrpF-independent entry of AvrBs3 into pepper cells. Strains 85*, 85*ΔhrpE (ΔhrpE), 85*ΔhrpF (ΔhrpF), 85*ΔhrpFΔxopA (ΔhrpFΔxopA), and 85*ΔhrcN (ΔhrcN) without expression construct (−) or ectopically expressing avrBs3 as indicated were infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. Leaves were photographed and destained in ethanol 3 dpi. Dashed lines indicate the infiltrated areas. For the analysis of protein synthesis, equal amounts of cell extracts were analyzed by immunoblotting using AvrBs3- or HrpF-specific antibodies. (B) HrpF-independent delivery of AvrBs3 induces the expression of Bs3 in AvrBs3-responsive pepper plants. Strains 85*, 85*ΔhrpF (ΔhrpF), and 85*ΔhrcN (ΔhrcN) ectopically expressing avrBs3 were infiltrated into leaves of AvrBs3-responsive pepper plants. Eight hours post inoculation, RNA was isolated from infected leaf material and transcribed into cDNA. Fragments corresponding to the Bs3 transcript and the constitutively expressed gene EF1α were amplified for 23–41 PCR cycles as indicated and amplicons were analyzed by agarose gel electrophoresis.
To investigate whether the browning of the infected leaf tissue could have resulted from the induction of the R gene Bs3, we performed RT-PCR studies. The Bs3 transcript was amplified from leaf tissue infected with strain 85* ectopically expressing avrBs3 but not from tissue infected with the corresponding hrcN deletion mutant (Figure 4B). Reduced amounts of the Bs3 transcript were detected in leaf material infected with the hrpF deletion mutant, suggesting that AvrBs3 entered the plant cells in the absence of the translocon protein HrpF and induced the expression of Bs3, albeit in reduced amounts (Figure 4B). We assume that the HrpF-independent entry of AvrBs3 into plant cells was not sufficient for the induction of a macroscopically visible HR reaction in Bs3 pepper plants but resulted in a browning of the infected leaf tissue. The phenotypes were not caused by the overexpression of avrBs3 or the T3S genes, because browning of the leaf tissue was also observed after inoculation of Bs3 pepper plants with the hrpF deletion mutant strain 82-8ΔhrpF, which contains the hrpG wild-type gene and naturally expresses avrBs3 (Figure S2). In agreement with these observations, the Bs3 transcript was detectable in leaf material infected with strain 82-8ΔhrpF but not with the T3S-deficient strain 82-8ΔhrcV (Figure S2).
Translocon-Independent Entry of a TAL Effector into Cells of the Non-Host Plant N. benthamiana
To investigate whether the translocon-independent entry of AvrBs3 into plant cells also occurs in non-host plants, we performed infection assays with Bs3-transgenic N. benthamiana plants. As reported previously, strain 85* induced a non-host HR on N. benthamiana plants, which is visible as necrotic area (Figure 5A; Metz et al., 2005; Scheibner et al., 2016). No macroscopic HR was observed after infection with the hrpF deletion mutant strain 85*ΔhrpF, suggesting that the induction of the non-host HR in N. benthamiana depends on the T3S translocon (Figure 5A). Strain 85*ΔhrpF ectopically expressing avrBs3, however, induced the HR in Bs3-transgenic N. benthamiana plants, suggesting that AvrBs3 entered the plant cells in the absence of HrpF (Figure 5A). The reaction was dependent on the T3S system because AvrBs3 did not induce a visible HR when analyzed in strain 85*ΔhrcN, which lacks the T3S ATPase (Figure 5A).
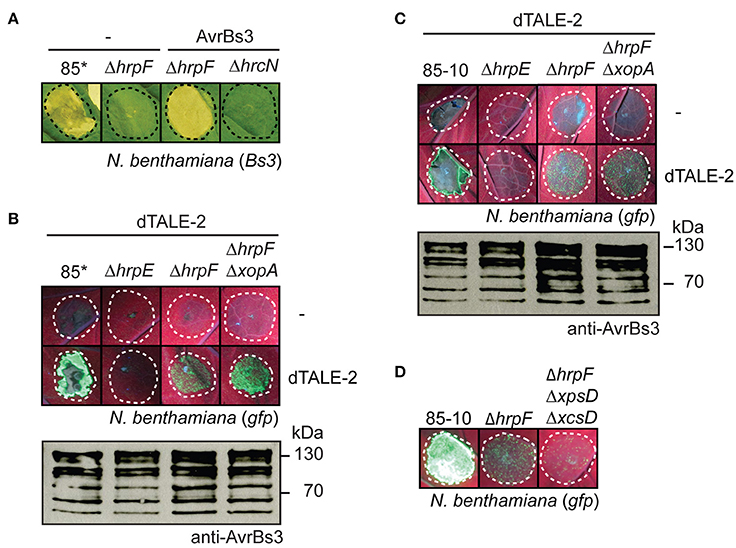
Figure 5. AvrBs3 and dTALE-2 are delivered into N. benthamiana in the absence of a functional translocon. (A) HrpF-independent entry of AvrBs3 into plant cells induces the HR in Bs3 N. benthamiana plants. Strains 85*, 85*ΔhrpF (ΔhrpF) and 85*ΔhrcN (ΔhrcN) without expression construct (−) or ectopically expressing avrBs3 as indicated were infiltrated at a density of 8 × 108 CFU ml−1 into leaves of Bs3-transgenic N. benthamiana plants. Leaves were photographed 8 dpi. Dashed lines indicate the infiltrated areas. (B) Translocon-independent delivery of dTALE-2 into gfp-transgenic N. benthamiana plants. Strains 85*, 85*ΔhrpE (ΔhrpE), 85*ΔhrpF (ΔhrpF), and 85*ΔhrpFΔxopA (ΔhrpFΔxopA) without expression construct (−) or ectopically expressing dTALE-2 as indicated were infiltrated at a density of 5 × 108 CFU ml−1 into leaves of gfp-transgenic N. benthamiana plants. Fluorescence of infected leaf areas was photographed 6 dpi. Dashed lines indicate the infiltrated areas. Fluorescence is reduced in plant tissue infiltrated with strain 85* because of the induction of the non-host HR. For the analysis of protein synthesis, equal amounts of cell extracts were analyzed by immunoblotting using AvrBs3-specific antibodies. (C) Analysis of dTALE-2 in derivatives of strain 85-10. Strains 85-10, 85-10ΔhrpE (ΔhrpE), 85-10ΔhrpF (ΔhrpF) and 85-10ΔhrpFΔxopA (ΔhrpFΔxopA) without expression construct (−) or ectopically expressing dTALE-2 were infiltrated at a density of 5 × 108 CFU ml−1 into leaves of gfp-transgenic N. benthamiana plants. Fluorescence was photographed 13 dpi. Protein synthesis was analyzed as described in B. (D) HrpF-independent entry of dTALE-2 into plant cells is reduced in the absence of functional T2S systems. Strains 85-10, 85-10ΔhrpF (ΔhrpF) and 85-10ΔhrpFΔxpsDΔxcsD (ΔhrpFΔxpsDΔxcsD), which is deficient in the Xps- and Xcs-T2S systems, ectopically expressing dTALE-2 were infiltrated into leaves of gfp-transgenic N. benthamiana plants. Fluorescence was photographed 10 dpi.
To confirm these results, we performed translocation assays with X. campestris pv. vesicatoria strains ectopically expressing dTALE-2 (designer TAL effector-2), which encodes an artificial derivative of AvrBs3 with modified RVDs (Weber et al., 2011b). To analyse the translocation of dTALE-2, bacteria were infiltrated into leaves of transgenic N. benthamiana plants, which encode the green fluorescent protein (GFP) under control of a dTALE-2-responsive promoter on a stably integrated viral vector construct (Werner et al., 2011; Scheibner et al., 2016). dTALE-2 induced GFP fluorescence when delivered by strain 85* (Figure 5B; Scheibner et al., 2016). Reduced fluorescence was observed when dTALE-2 was analyzed in strains 85*ΔhrpF and 85*ΔhrpFΔxopA whereas no fluorescence was detectable after infiltration of the T3S-deficient strain 85*ΔhrpE (Figure 5B). Similar data were obtained for derivatives of the wild-type strain 85-10, suggesting that the HrpF-independent entry of dTALE-2 into plant cells was not caused by the overexpression of the T3S genes (Figure 5C). Taken together, these data suggest that dTALE-2 can enter the plant cell in a translocon-independent manner.
We also performed infection experiments with strain 85-10ΔhrpFΔxpsDΔxcsD, which lacks hrpF and functional T2S systems. Type II-secreted cell wall-degrading enzymes were proposed to facilitate T3S pilus assembly and thus presumably contribute to the HrpF-independent passage of AvrBs3 across the plant plasma membrane (Szczesny et al., 2010b). In agreement with this hypothesis, dTALE-2 induced reduced GFP fluorescence when analyzed in strain 85-10ΔhrpFΔxpsDΔxcsD (Figure 5D).
The Translocation Signal Contributes to the HrpF-Independent Entry of AvrBs3 into Plant Cells
Next, we investigated the contribution of the translocation signal to the translocon-independent entry of AvrBs3 into plant cells. Infection experiments showed that the N-terminal deletion derivative dTALE-2ΔN did not induce detectable GFP fluorescence in gfp-transgenic N. benthamiana plants when analyzed in strains 85* and 85*ΔhrpF (Figure 6A). As dTALE-2ΔN is deleted in the T3S and translocation signal (Schreiber et al., 2015; Scheibner et al., 2016), the lack of GFP fluorescence suggests that the translocon-independent delivery of dTALE-2 depends on its passage through the T3S system. In contrast to dTALE-2ΔN, AvrBs31−50-dTALE-2ΔN induced GFP fluorescence when analyzed in strains 85*, 85*ΔhpaB and 85*ΔhrpF (Figure 6B). Notably, GFP fluorescence induced by strain 85* was reduced because of the induction of the non-host HR by strain 85* which leads to tissue necrosis (see above; Figure 5A). Our observations confirm the results obtained for the AvrBs31−50-AvrBs3Δ2 fusion (see above) and suggest that the N-terminal 50 amino acids of AvrBs3 contain a translocation signal, which can target dTALE-2 for translocon-independent translocation.
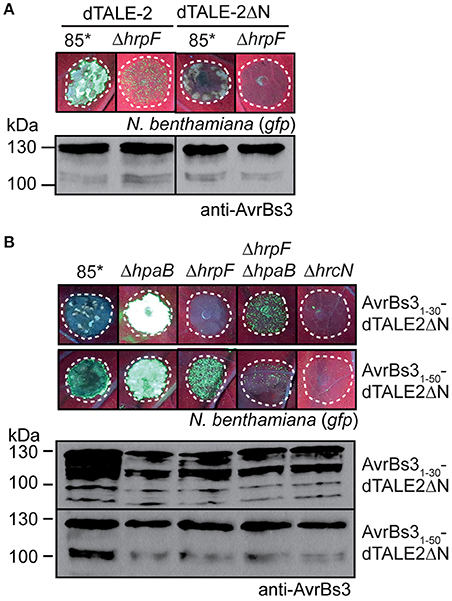
Figure 6. The N-terminal translocation signal is required for translocon-independent delivery of dTALE-2. (A) dTALE-2ΔN does not induce detectable GFP fluorescence when analyzed in strain 85*. Strains 85* and 85*ΔhrpF (ΔhrpF) ectopically expressing dTALE-2 or dTALE-2ΔN as indicated were infiltrated at a density of 8 × 108 CFU ml−1 into leaves of gfp-transgenic N. benthamiana plants. Fluorescence was photographed 7 dpi. Dashed lines indicate the infiltrated areas. For the analysis of protein synthesis, equal amounts of cell extracts were analyzed by immunoblotting using AvrBs3-specific antibodies. (B) Translocation assays with dTALE-2ΔN fusion proteins. Strains 85*, 85*ΔhpaB (ΔhpaB), 85*ΔhrpF (ΔhrpF), 85*ΔhrpFΔhpaB (ΔhrpFΔhpaB), and 85*ΔhrcN (ΔhrcN) ectopically expressing dTALE-2ΔN fusions as indicated were infiltrated at a density of 8 × 108 CFU ml−1 into leaves of gfp-transgenic N. benthamiana plants. Fluorescence was photographed 11 dpi. Dashed lines indicate the infiltrated areas. Fluorescence is reduced in plant tissue infiltrated with strain 85* because of the induction of the non-host HR. Protein synthesis was analyzed as described in (A).
Translocation assays with AvrBs31−30-dTALE-2ΔN revealed that the N-terminal 30 amino acids of AvrBs3 targeted dTALE-2ΔN for translocation in strain 85*ΔhpaB whereas no fluorescence was observed with the wild-type strain or the hrpF deletion mutant (Figure 6B). This confirms the finding that the N-terminal 30 amino acids of AvrBs3 contain a minimal translocation signal, which is recognized in the absence of HpaB. Reduced GFP fluorescence, however, was detected when AvrBs31−30-dTALE-2ΔN was analyzed in strain 85*ΔhrpFΔhpaB (Figure 6B). Thus, AvrBs31−30-dTALE-2ΔN can enter the plant cell in the absence of the translocon when delivered by the hpaB deletion mutant (Figure 6B). Taken together, these data suggest that the translocon-dependent and -independent delivery of AvrBs3 into plant cells depends on similar targeting signals.
Discussion
In the present study, we localized T3S and translocation signals in the N-terminal region of the TAL effector AvrBs3. The analysis of AvrBs3-reporter fusion proteins revealed that the N-terminal 10 amino acids of AvrBs3 are sufficient to target the reporter for T3S (Figure 1). This is in agreement with previous findings that amino acids 6–10 of T3S substrates often contain essential features of T3S signals as was also shown for the effector proteins XopE2 and XopJ from X. campestris pv. vesicatoria (Wang et al., 2013; Scheibner et al., 2017). The N-terminal 30 amino acids of AvrBs3 contain a minimal translocation signal, which promotes translocation of the reporter in the absence of the T3S chaperone HpaB but not in the wild-type strain (Figures 1, 2). Minimal translocation signals were previously also identified in the N-terminal regions of the effectors XopE2 and XopJ (Scheibner et al., 2017).
Translocation of AvrBs3 in the wild-type strain depends on a signal in the N-terminal 50 amino acids. Notably, however, in contrast to the minimal translocation signal, the N-terminal 50 amino acids did not efficiently target the AvrBs3Δ2 reporter for translocation in the absence of HpaB (Figure 2), suggesting that the function of the minimal translocation signal depends on the neighboring protein regions. In contrast to AvrBs31−50-AvrBs3Δ2, however, the full-length AvrBs3 protein was efficiently translocated by both wild-type and hpaB mutant strains (Figures 1, 2). AvrBs3 likely contains additional signals outside the N-terminal region, which promote translocation in the absence of HpaB. Thus, the comparative analysis of AvrBs3Δ2 and AvrBs3ΔN63 fusion proteins revealed that the region between amino acids 64 and 152 promotes translocation of AvrBs3 in both wild-type and hpaB deletion mutant strains (Figures 3, 7A), suggesting that AvrBs3 contains multiple signals that control secretion and translocation in the presence and absence of HpaB. HpaB-independent translocation was already previously observed for other effectors and is indicative of a hierarchy in effector protein delivery (Büttner et al., 2006; Schulze et al., 2012). Thus, in X. campestris pv. vesicatoria, HpaB-independent effectors might be translocated during the early stages of the plant-pathogen interaction, prior to the activation of HpaB (Figure 7B). Differences in the timing of effector protein delivery were previously reported for animal-pathogenic bacteria but have not yet been investigated in plant-pathogenic bacteria (Enninga et al., 2005; Schlumberger et al., 2005; Mills et al., 2008; Van Engelenburg and Palmer, 2008; Winnen et al., 2008). It is also still unknown whether the timing of effector protein translocation is controlled by the N-terminal export signals.
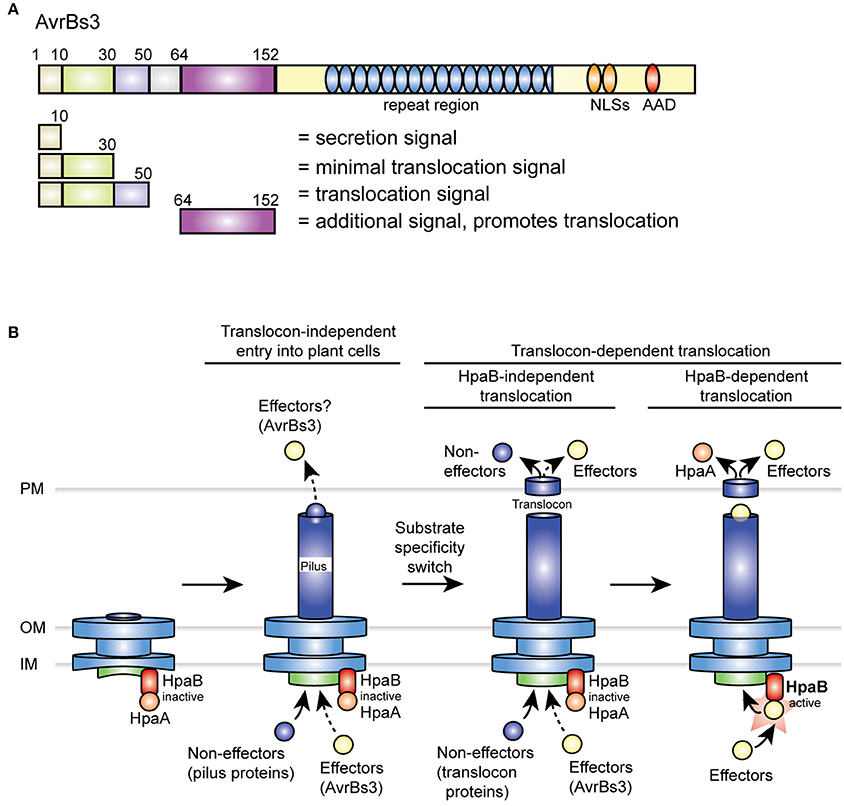
Figure 7. Overview on type III export signals in AvrBs3 and model of the T3S hierarchy in X. campestris pv. vesicatoria. (A) Overview on secretion and translocation signals in AvrBs3. N-terminal signals are indicated, numbers refer to amino acid positions. NLS, nuclear localization signal; AAD, acidic activation domain. (B) Predicted T3S hierarchy in X. campestris pv. vesicatoria. After formation of the membrane-spanning part of the secretion apparatus, the T3S pilus is assembled. HpaA presumably associates with the T3S chaperone HpaB and thus inactivates HpaB to prevent secretion of effector proteins prior to the insertion of the translocon. As suggested by data in the present study, AvrBs3 might already enter the plant cell in the absence of a functional translocon (indicated by a dashed arrow). A possible translocon-independent entry of other effectors into the plant cell (indicated by a question mark) remains to be investigated. A predicted switch in T3S substrate specificity after pilus formation leads to the secretion of translocon proteins and thus to the assembly of the T3S translocon in the plant plasma membrane. While HpaB is still inactive during this stage of the T3S process, HpaB-independent effectors including AvrBs3 and non-effectors such as XopA and HrpF are already translocated. However, when compared to the wild-type strain, hpaB mutants presumably translocate reduced amounts of effectors (indicated by a dashed arrow; Büttner et al., 2006; Schulze et al., 2012). After insertion of the translocon, a yet unknown signal triggers the translocation of HpaA. This leads to the liberation of HpaB and thus activates the translocation of HpaB-dependent effectors. AAD, acidic activation domain; IM, inner membrane; NLS, nuclear localization signal; OM, outer membrane; PM, plant plasma membrane.
Comparative sequence analysis revealed that the N-terminal region of AvrBs3 does not share homology with corresponding regions of other effector proteins, indicating that T3S and translocation signals are not conserved in T3S substrates from X. campestris pv. vesicatoria. Similar findings were previously reported for T3S substrates from other plant- and animal-pathogenic bacteria (Schechter et al., 2004; Arnold et al., 2009; Löwer and Schneider, 2009; Samudrala et al., 2009; Buchko et al., 2010; McDermott et al., 2011). The analysis of the amino acid composition of AvrBs3 revealed that the N-terminal 50 amino acids of AvrBs3 contain slightly higher levels of arginine and aspartate residues than the remainder of the protein (Figure S3). Furthermore, the N-terminal 50 amino acids of AvrBs3 and the region between amino acids 64 and 152, which both promote translocation, contain increased amounts of proline residues (Figure S3). Notably, elevated proline levels are also present in the N-terminal regions of the effectors XopE2 and XopJ, which contain T3S and minimal translocation signals (Scheibner et al., 2017). The contribution of N-terminal proline, arginine and aspartate residues to the translocation of AvrBs3 remains to be investigated in future studies.
The precise roles of N-terminal T3S and translocation signals as well as the molecular mechanisms underlying their recognition are still largely unknown in both plant- and animal-pathogenic bacteria. Potential docking sites for type III effectors in X. campestris pv. vesicatoria include the cytoplasmic putative C ring protein HrcQ and the cytoplasmic domain of the inner membrane protein HrcV, which both interact with T3S substrates in vitro (Lorenz et al., 2012; Hartmann and Büttner, 2013). In case of AvrBs3, however, the N-terminal protein region, which contains the T3S and translocation signal, is dispensable for the interaction with HrcQ and the cytoplasmic domain of HrcV (Lorenz et al., 2012; Hartmann and Büttner, 2013). Similar findings were observed for N-terminal deletion derivatives of XopE2 and XopJ, which interact with HrcQ (Scheibner et al., 2017). The specific recognition of the N-terminal export signal might, therefore, depend on additional components of the T3S system or might occur after the initial docking of T3S substrates to the secretion apparatus.
Given the anticipated essential role of the translocon for type III-dependent protein translocation, it was assumed that the insertion of the translocon precedes effector protein translocation. In agreement with this hypothesis, previous studies suggested that the translocation of effector-reporter fusions as well as the in situ detection of AvrBs3 in plant nuclei is abolished in HrpF-deficient strains (Büttner et al., 2002; Szurek et al., 2002; Hotson et al., 2003; Thieme et al., 2007; Jiang et al., 2009; Teper et al., 2016). In the present study, however, we observed that AvrBs3 enters plant cells even in the absence of the translocon protein HrpF and the secreted XopA protein, which might contribute to effector protein translocation (Figure 4; Noël et al., 2002). Translocon-independent entry of AvrBs3 into plant cells was significantly reduced when compared to the translocation by the wild-type strain and only led to a browning of the infected leaf tissue in Bs3 pepper plants, suggesting a much weaker HR. This was accompanied by a detectable increase in the Bs3 transcript level (Figure 4) and has not yet been investigated for the wild-type AvrBs3 protein in previous studies. In Bs3-transgenic N. benthamiana plants, AvrBs3 induces a macroscopically visible Bs3-dependent HR even when delivered by hrpF deletion mutants (Figure 5). This was not observed in Bs3 pepper plants and might be caused by an increased activation of Bs3 expression, an increased HR induction or a more efficient translocon-independent entry of AvrBs3 in Bs3-transgenic N. benthamiana plants. HrpF-independent entry into plant cells was confirmed for dTALE-2 in gfp-transgenic N. benthamiana plants. No detectable fluorescence was observed with mutants lacking the T3S ATPase HrcN or the pilus protein HrpE, suggesting that the HrpF-independent delivery of AvrBs3 depends on the T3S system (Figure 5). Given that the transport of AvrBs3 through the T3S pilus is essential for its passage across the plant cell wall, we did not investigate a possible autonomous entry of recombinant AvrBs3 into plant cells.
The mechanisms underlying the HrpF-independent passage of AvrBs3 into plant cells as well as its biological significance remain to be elucidated. Notably, translocon-independent transport into eukaryotic cells was previously reported for effector proteins from animal-pathogenic bacteria including SspH1 from Salmonella spp. and YopM from Yersinia spp. (Rüter et al., 2010; Scharnert et al., 2013; Lubos et al., 2014). Both proteins cross the host plasma membrane when added as recombinant proteins to cultured cells. Autonomously translocating effectors present a novel class of cell-penetrating peptides (CPPs), which are often used as vehicles for the transport of cargo molecules into target cells (Rüter and Schmidt, 2017). CPPs include basic/amphiphilic or hydrophobic peptides which consist of usually less than 30 amino acids and autonomously translocate across biological membranes, even when present as fusion partner of a large cargo protein (Takeuchi and Futaki, 2016; Radis-Baptista et al., 2017). CPPs are either directly transported across the membrane or are delivered by endocytosis (Scharnert et al., 2013; Radis-Baptista et al., 2017). Both transport pathways were observed for YopM from Yersinia spp. (Scharnert et al., 2013). Our preliminary infection experiments with X. campestris pv. vesicatoria translocon mutants in the presence of the endocytosis inhibitors bafilomycin A1, cytochalasin D, or wortmannin did not reveal a contribution of endocytosis to the delivery of AvrBs3. It, therefore, remains to be investigated whether AvrBs3 can directly cross biological membranes. The direct transport of CPPs across the membrane presumably involves the interaction of positively charged amino acids with negatively charged lipids of the membrane and was for instance reported for arginine-rich CPPs (Scharnert et al., 2013; Rüter and Schmidt, 2017). Interestingly, the N-terminal region of AvrBs3 is enriched in arginine residues (Figure S3), however, AvrBs3 does not contain sequence motifs with homology to typical CPP sequences or prolonged stretches of positively charged amino acids, which could mediate the interaction with membranes (Radis-Baptista et al., 2017). In future studies, we will investigate which protein regions of AvrBs3 are required for the translocon-independent entry into plant cells.
Author Contributions
FS, DB, and SM conceived the study. FS performed secretion assays, infection studies, immunoblot analyses and RT-PCR experiments. DB performed infection assays and immunoblot analyses. All authors analyzed the data. DB and FS wrote the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by grants from the Deutsche Forschungsgemeinschaft (BU2145/5-1 and CRC 648 “Molecular mechanisms of information processing in plants”) to DB. We acknowledge the financial support of the open access publication fund of the Martin Luther university Halle-Wittenberg.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We are grateful to U. Bonas for comments on the manuscript and for providing the AvrBs3-specific antibody. We thank M. Jordan for technical assistance, S. Thieme for providing vector pGGA3 and J. Uding for generating constructs pBRM-PxopJ1−155-356 and pBRM-PxopJ1−155-avrBs3ΔN63.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2017.02180/full#supplementary-material
Figure S1. Translocation of AvrBs3Δ2 and AvrBs3ΔN63 fusion proteins by derivatives of strain 85-10. (A) The HR induction in ECW-30R pepper plants is specific for AvrBs3Δ2 fusion proteins. Strains 85* and 85*ΔhpaB (ΔhpaB) without expression construct or ectopically expressing avrBs31-30-avrBs3Δ2 were infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. For the better visualization of the HR, leaves of ECW-10R and ECW-30R plants were destained in ethanol 2 and 3 dpi, respectively. Dashed lines indicate the infiltrated areas. (B) Translocation assays with strains 85-10 and 85-10ΔhpaB. Strains 85-10 and 85-10ΔhpaB without expression construct (−) or encoding AvrBs3, AvrBs31−X-AvrBs3Δ2 or AvrBs31−X-AvrBs3ΔN63 fusion proteins (1-X = amino acids 1–30 or 1–50 of AvrBs3) on corresponding expression constructs as indicated were infiltrated into leaves of AvrBs3-responsive ECW-30R and AvrBs1-responsive ECW-10R pepper plants. Plant reactions were analyzed as described in (A). (C) Detection of AvrBs3Δ2 and AvrBs3ΔN63 fusion proteins. Equal amounts of cell extracts from strains described in (B) were analyzed by immunoblotting using an AvrBs3-specific antiserum.
Figure S2. Translocon-independent delivery of the native AvrBs3 protein. (A) Infection assays with derivatives of strain 82-8. Strains 82-8, 82-8ΔhrpF (ΔhrpF), 82-8ΔhrpFΔxopA (ΔhrpFΔxopA), and 82-8ΔhrcV (ΔhrcV) were infiltrated at a density of 8 × 108 CFU ml−1 into leaves of AvrBs3-responsive ECW-30R pepper plants and Bs3-transgenic N. benthamiana plants. Leaves were destained in ethanol 3 and 5 dpi, respectively. Dashed lines indicate the infiltrated areas. Equal amounts of cell extracts were analysed by immunoblotting using an AvrBs3-specific antiserum. (B) Analysis of Bs3 transcript levels. Strains 82-8, 82-8ΔhrpF (ΔhrpF), 82-8ΔhrpFΔxopA (ΔhrpFΔxopA), and 82-8ΔhrcV (ΔhrcV) were infiltrated at a density of 8 × 108 CFU ml−1 into leaves AvrBs3-responsive ECW-30R pepper plants and Bs3-transgenic N. benthamiana plants. RNA was isolated from infected leaf material and transcribed into cDNA. Fragments corresponding to Bs3 and the constitutively expressed EF1α gene were amplified for 40 cycles by PCR and the amplification products were analyzed by agarose gel electrophoresis.
Figure S3. Amino acid composition of AvrBs3. The percentage of proline, polar and charged amino acid residues in different regions of AvrBs3 is shown.
References
Akopyan, K., Edgren, T., Wang-Edgren, H., Rosqvist, R., Fahlgren, A., Wolf-Watz, H., et al. (2011). Translocation of surface-localized effectors in type III secretion. Proc. Natl. Acad. Sci. U.S.A. 108, 1639–1644. doi: 10.1073/pnas.1013888108
Arnold, R., Brandmaier, S., Kleine, F., Tischler, P., Heinz, E., Behrens, S., et al. (2009). Sequence-based prediction of type III secreted proteins. PLoS Pathog. 5:e1000376. doi: 10.1371/journal.ppat.1000376
Ashida, H., Mimuro, H., and Sasakawa, C. (2015). Shigella manipulates host immune responses by delivering effector proteins with specific roles. Front. Immunol. 6:219. doi: 10.3389/fimmu.2015.00219
Ausubel, F. M., Brent, R., Kingston, R. E., Moore, D. D., Seidman, J. G., Smith, J. A., et al. (1996). Current Protocols in Molecular Biology. New York, NY: John Wiley & Sons Inc.
Boch, J., and Bonas, U. (2010). Xanthomonas AvrBs3 family-type III effectors: discovery and function. Annu. Rev. Phytopathol. 48, 419–436. doi: 10.1146/annurev-phyto-080508-081936
Boch, J., Scholze, H., Schornack, S., Landgraf, A., Hahn, S., Kay, S., et al. (2009). Breaking the code of DNA-binding specificity of TAL-type III effectors. Science 326, 1509–1512. doi: 10.1126/science.1178811
Bonas, U., Schulte, R., Fenselau, S., Minsavage, G. V., Staskawicz, B. J., and Stall, R. E. (1991). Isolation of a gene-cluster from Xanthomonas campestris pv. vesicatoria that determines pathogenicity and the hypersensitive response on pepper and tomato. Mol. Plant Microb. Interact. 4, 81–88. doi: 10.1094/MPMI-4-081
Buchko, G. W., Niemann, G., Baker, E. S., Belov, M. E., Smith, R. D., Heffron, F., et al. (2010). A multi-pronged search for a common structural motif in the secretion signal of Salmonella enterica serovar Typhimurium type III effector proteins. Mol. Biosyst. 6, 2448–2458. doi: 10.1039/c0mb00097c
Büttner, D. (2012). Protein export according to schedule – architecture, assembly and regulation of type III secretion systems from plant and animal pathogenic bacteria. Microbiol. Mol. Biol. Rev. 76, 262–310. doi: 10.1128/MMBR.05017-11
Büttner, D. (2016). Behind the lines-actions of bacterial type III effector proteins in plant cells. FEMS Microbiol. Rev. 40, 894–937. doi: 10.1093/femsre/fuw026
Büttner, D., and Bonas, U. (2002). Getting across-bacterial type III effector proteins on their way to the plant cell. EMBO J. 21, 5313–5322. doi: 10.1093/emboj/cdf536
Büttner, D., and Bonas, U. (2010). Regulation and secretion of Xanthomonas virulence factors. FEMS Microbiol. Rev. 34, 107–133. doi: 10.1111/j.1574-6976.2009.00192.x
Büttner, D., Gürlebeck, D., Noël, L. D., and Bonas, U. (2004). HpaB from Xanthomonas campestris pv. vesicatoria acts as an exit control protein in type III-dependent protein secretion. Mol. Microbiol. 54, 755–768. doi: 10.1111/j.1365-2958.2004.04302.x
Büttner, D., Lorenz, C., Weber, E., and Bonas, U. (2006). Targeting of two effector protein classes to the type III secretion system by a HpaC- and HpaB-dependent protein complex from Xanthomonas campestris pv. vesicatoria. Mol. Microbiol. 59, 513–527. doi: 10.1111/j.1365-2958.2005.04924.x
Büttner, D., Nennstiel, D., Klüsener, B., and Bonas, U. (2002). Functional analysis of HrpF, a putative type III translocon protein from Xanthomonas campestris pv. vesicatoria. J. Bacteriol. 184, 2389–2398. doi: 10.1128/JB.184.9.2389-2398.2002
Büttner, D., Noël, L., Stuttmann, J., and Bonas, U. (2007). Characterization of the non-conserved hpaB-hrpF region in the hrp pathogenicity island from Xanthomonas campestris pv. vesicatoria. Mol. Plant Microbe Interact. 20, 1063–1074. doi: 10.1094/MPMI-20-9-1063
Canteros, B. I. (1990). Diversity of Plasmids and Plasmid-Encoded Phenotypic Traits in Xanthomonas campestris pv. vesicatoria. Ph.D. thesis, University of Florida, Gainesville, FL.
Choi, M. S., Kim, W., Lee, C., and Oh, C. S. (2013). Harpins, multifunctional proteins secreted by gram-negative plant-pathogenic bacteria. Mol. Plant Microbe Interact. 26, 1115–1122. doi: 10.1094/MPMI-02-13-0050-CR
Daniels, M. J., Barber, C. E., Turner, P. C., Cleary, W. G., and Sawczyc, M. K. (1984). Isolation of mutants of Xanthomonas campestris pathovar campestris showing altered pathogenicity. J. Gen. Microbiol. 130, 2447–2455.
Deblaere, R., Bytebier, B., De Greve, H., Deboeck, F., Schell, J., Van Montagu, M., et al. (1985). Efficient octopine Ti plasmid-derived vectors for Agrobacterium-mediated gene transfer to plants. Nucleic Acids Res. 13, 4777–4788. doi: 10.1093/nar/13.13.4777
Deng, D., Yan, C., Pan, X., Mahfouz, M., Wang, J., Zhu, J. K., et al. (2012). Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 335, 720–723. doi: 10.1126/science.1215670
Deng, W., Marshall, N. C., Rowland, J. L., McCoy, J. M., Worrall, L. J., Santos, A. S., et al. (2017). Assembly, structure, function and regulation of type III secretion systems. Nat. Rev. Microbiol. 15, 323–337. doi: 10.1038/nrmicro.2017.20
Diepold, A., and Armitage, J. P. (2015). Type III secretion systems: the bacterial flagellum and the injectisome. Philos. Trans. R. Soc. Lond. B Biol. Sci. 5:370. doi: 10.1098/rstb.2015.0020
Diepold, A., Wiesand, U., and Cornelis, G. R. (2011). The assembly of the export apparatus (YscR,S,T,U,V) of the Yersinia type III secretion apparatus occurs independently of other structural components and involves the formation of an YscV oligomer. Mol. Microbiol. 82, 502–514. doi: 10.1111/j.1365-2958.2011.07830.x
Dietsche, T., Tesfazgi Mebrhatu, M., Brunner, M. J., Abrusci, P., Yan, J., Franz-Wachtel, M., et al. (2016). Structural and functional characterization of the bacterial type III secretion export apparatus. PLoS Pathog. 12:e1006071. doi: 10.1371/journal.ppat.1006071
Drehkopf, S., Hausner, J., Jordan, M., Scheibner, F., Bonas, U., and Buttner, D. (2017). A TAL-based reporter assay for monitoring type III-dependent protein translocation in Xanthomonas. Methods Mol. Biol. 1531, 121–139. doi: 10.1007/978-1-4939-6649-3_11
Enninga, J., Mounier, J., Sansonetti, P., and Tran Van Nhieu, G. (2005). Secretion of type III effectors into host cells in real time. Nat. Methods 2, 959–965. doi: 10.1038/nmeth804
Ensminger, A. W. (2016). Legionella pneumophila, armed to the hilt: justifying the largest arsenal of effectors in the bacterial world. Curr. Opin. Microbiol. 29, 74–80. doi: 10.1016/j.mib.2015.11.002
Figurski, D., and Helinski, D. R. (1979). Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc. Natl. Acad. Sci. U.S.A. 76, 1648–1652. doi: 10.1073/pnas.76.4.1648
Galan, J. E., Lara-Tejero, M., Marlovits, T. C., and Wagner, S. (2014). Bacterial type III secretion systems: specialized nanomachines for protein delivery into target cells. Annu. Rev. Microbiol. 68, 415–438. doi: 10.1146/annurev-micro-092412-155725
Grabowski, B., Schmidt, M. A., and Rüter, C. (2017). Immunomodulatory Yersinia outer proteins (Yops)-useful tools for bacteria and humans alike. Virulence doi: 10.1080/21505594.2017.1303588. [Epub ahead of print].
Greenberg, J., and Vinatzer, B. A. (2003). Identifying type III effectors of plant pathogens and analyzing their interaction with plant cells. Curr. Opin. Microbiol. 6, 20–28. doi: 10.1016/S1369-5274(02)00004-8
Guttman, D. S., Vinatzer, B. A., Sarkar, S. F., Ranall, M. V., Kettler, G., and Greenberg, J. T. (2002). A functional screen for the type III (Hrp) secretome of the plant pathogen Pseudomonas syringae. Science 295, 1722–1726. doi: 10.1126/science.295.5560.1722
Hartmann, N., and Büttner, D. (2013). The inner membrane protein HrcV from Xanthomonas is involved in substrate docking during type III secretion. Mol. Plant Microbe Interact. 26, 1176–1189. doi: 10.1094/MPMI-01-13-0019-R
Hartmann, N., Schulz, S., Lorenz, C., Fraas, S., Hause, G., and Büttner, D. (2012). Characterization of HrpB2 from Xanthomonas campestris pv. vesicatoria identifies protein regions that are essential for type III secretion pilus formation. Microbiology 158, 1334–1349. doi: 10.1099/mic.0.057604-0
Hausner, J., Hartmann, N., Lorenz, C., and Büttner, D. (2013). The periplasmic HrpB1 protein from Xanthomonas spp. binds to peptidoglycan and to components of the type III secretion system. Appl. Environ. Microbiol. 79, 6312–6324. doi: 10.1128/AEM.01226-13
Hotson, A., Chosed, R., Shu, H., Orth, K., and Mudgett, M. B. (2003). Xanthomonas type III effector XopD targets SUMO-conjugated proteins in planta. Mol. Microbiol. 50, 377–389. doi: 10.1046/j.1365-2958.2003.03730.x
Hu, B., Lara-Tejero, M., Kong, Q., Galan, J. E., and Liu, J. (2017). In situ molecular architecture of the Salmonella type III secretion machine. Cell 168, 1065–1074. doi: 10.1016/j.cell.2017.02.022
Hueck, C. J. (1998). Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol. Mol. Biol. Rev. 62, 379–433.
Huguet, E., Hahn, K., Wengelnik, K., and Bonas, U. (1998). hpaA mutants of Xanthomonas campestris pv. vesicatoria are affected in pathogenicity but retain the ability to induce host-specific hypersensitive reaction. Mol. Microbiol. 29, 1379–1390. doi: 10.1046/j.1365-2958.1998.01019.x
Ji, H., and Dong, H. (2015). Key steps in type III secretion system (T3SS) towards translocon assembly with potential sensor at plant plasma membrane. Mol. Plant Pathol. 16, 762–773. doi: 10.1111/mpp.12223
Jiang, W., Jiang, B. L., Xu, R. Q., Huang, J. D., Wei, H. Y., Jiang, G. F., et al. (2009). Identification of six type III effector genes with the PIP box in Xanthomonas campestris pv. campestris and five of them contribute individually to full pathogenicity. Mol. Plant Microbe Interact. 22, 1401–1411. doi: 10.1094/MPMI-22-11-1401
Jones, J. B., Lacy, G. H., Bouzar, H., Stall, R. E., and Schaad, N. W. (2004). Reclassification of the xanthomonads associated with bacterial spot disease of tomato and pepper. Syst. Appl. Microbiol. 27, 755–762. doi: 10.1078/0723202042369884
Karimi, M., De Meyer, B., and Hilson, P. (2005). Modular cloning in plant cells. Trends Plant Sci. 10, 103–105. doi: 10.1016/j.tplants.2005.01.008
Kim, J. G., Jeon, E., Oh, J., Moon, J. S., and Hwang, I. (2004). Mutational analysis of Xanthomonas harpin HpaG identifies a key functional region that elicits the hypersensitive response in nonhost plants. J. Bacteriol. 186, 6239–6247. doi: 10.1128/JB.186.18.6239-6247.2004
Knoop, V., Staskawicz, B., and Bonas, U. (1991). Expression of the avirulence gene avrBs3 from Xanthomonas campestris pv. vesicatoria is not under the control of hrp genes and is independent of plant factors. J. Bacteriol. 173, 7142–7150. doi: 10.1128/jb.173.22.7142-7150.1991
Kousik, C. S., and Ritchie, D. F. (1998). Response of bell pepper cultivars to bacterial spot pathogen races that individually overcome major resistance genes. Plant Dis. 82, 181–186. doi: 10.1094/PDIS.1998.82.2.181
Kovach, M. E., Elzer, P. H., Hill, D. S., Robertson, M. A., Farris, M. A., Roop, I. I. R. M., et al. (1995). Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175–176. doi: 10.1016/0378-1119(95)00584-1
Kvitko, B. H., Ramos, A. R., Morello, J. E., Oh, H. S., and Collmer, A. (2007). Identification of harpins in Pseudomonas syringae pv. tomato DC3000, which are functionally similar to HrpK1 in promoting translocation of type III secretion system effectors. J. Bacteriol. 189, 8059–8072. doi: 10.1128/JB.01146-07
Lorenz, C., and Büttner, D. (2009). Functional characterization of the type III secretion ATPase HrcN from the plant pathogen Xanthomonas campestris pv. vesicatoria. J. Bacteriol. 191:1414–1428. doi: 10.1128/JB.01446-08
Lorenz, C., Hausner, J., and Büttner, D. (2012). HrcQ provides a docking site for early and late type III secretion substrates from Xanthomonas. PLoS ONE 7:e51063. doi: 10.1371/journal.pone.0051063
Lorenz, C., Kirchner, O., Egler, M., Stuttmann, J., Bonas, U., and Büttner, D. (2008). HpaA from Xanthomonas is a regulator of type III secretion. Mol. Microbiol. 69, 344–360. doi: 10.1111/j.1365-2958.2008.06280.x
Löwer, M., and Schneider, G. (2009). Prediction of type III secretion signals in genomes of gram-negative bacteria. PLoS ONE 4:e5917. doi: 10.1371/journal.pone.0005917
Lubos, M. L., Norkowski, S., Stolle, A. S., Langel, Ü., Schmidt, A., and Rüter, C. (2014). Analysis of T3SS-independent autonomous internalization of the bacterial effector protein SspH1 from Salmonella typhimurium. Inflamm. Cell. Signal. 1:e423. doi: 10.14800/ics.423
Mak, A. N., Bradley, P., Cernadas, R. A., Bogdanove, A. J., and Stoddard, B. L. (2012). The crystal structure of TAL effector PthXo1 bound to its DNA target. Science 335, 716–719. doi: 10.1126/science.1216211
Mattei, P. J., Faudry, E., Job, V., Izore, T., Attree, I., and Dessen, A. (2011). Membrane targeting and pore formation by the type III secretion system translocon. FEBS J. 278, 414–426. doi: 10.1111/j.1742-4658.2010.07974.x
McDermott, J. E., Corrigan, A., Peterson, E., Oehmen, C., Niemann, G., Cambronne, E. D., et al. (2011). Computational prediction of type III and IV secreted effectors in gram-negative bacteria. Infect. Immun. 79, 23–32. doi: 10.1128/IAI.00537-10
Ménard, R., Sansonetti, P. J., and Parsot, C. (1993). Nonpolar mutagenesis of the ipa genes defines IpaB, IpaC, and IpaD as effectors of Shigella flexneri entry into epithelial cells. J. Bacteriol. 175, 5899–5906. doi: 10.1128/jb.175.18.5899-5906.1993
Metz, M., Dahlbeck, D., Morales, C. Q., Al Sady, B., Clark, E. T., and Staskawicz, B. J. (2005). The conserved Xanthomonas campestris pv. vesicatoria effector protein XopX is a virulence factor and suppresses host defense in Nicotiana benthamiana. Plant J. 41, 801–814. doi: 10.1111/j.1365-313X.2005.02338.x
Mills, E., Baruch, K., Charpentier, X., Kobi, S., and Rosenshine, I. (2008). Real-time analysis of effector translocation by the type III secretion system of enteropathogenic Escherichia coli. Cell. Host Microbe 3, 104–113. doi: 10.1016/j.chom.2007.11.007
Minsavage, G. V., Dahlbeck, D., Whalen, M. C., Kearny, B., Bonas, U., Staskawicz, B. J., et al. (1990). Gene-for-gene relationships specifying disease resistance in Xanthomonas campestris pv. vesicatoria- pepper interactions. Mol. Plant Microbe Interact. 3, 41–47. doi: 10.1094/MPMI-3-041
Morbitzer, R., Elsaesser, J., Hausner, J., and Lahaye, T. (2011). Assembly of custom TALE-type DNA binding domains by modular cloning. Nucleic Acids Res. 39, 5790–5799. doi: 10.1093/nar/gkr151
Moscou, M. J., and Bogdanove, A. J. (2009). A simple cipher governs DNA recognition by TAL effectors. Science 326:1501. doi: 10.1126/science.1178817
Mueller, C. A., Broz, P., and Cornelis, G. R. (2008). The type III secretion system tip complex and translocon. Mol. Microbiol. 68, 1085–1095. doi: 10.1111/j.1365-2958.2008.06237.x
Noël, L., Thieme, F., Gäbler, J., Büttner, D., and Bonas, U. (2003). XopC and XopJ, two novel type III effector proteins from Xanthomonas campestris pv. vesicatoria. J. Bacteriol. 185, 7092–7102. doi: 10.1128/JB.185.24.7092-7102.2003
Noël, L., Thieme, F., Nennstiel, D., and Bonas, U. (2002). Two novel type III system-secreted proteins of Xanthomonas campestris pv. vesicatoria are encoded within the hrp pathogenicity island. J. Bacteriol. 184, 1340–1348. doi: 10.1128/JB.184.5.1340-1348.2002
Petnicki-Ocwieja, T., Schneider, D. J., Tam, V. C., Chancey, S. T., Shan, L., Jamir, Y., et al. (2002). Genomewide identification of proteins secreted by the Hrp type III protein secretion system of Pseudomonas syringae pv. tomato DC3000. Proc. Natl. Acad. Sci. U.S.A. 99, 7652–7657. doi: 10.1073/pnas.112183899
Petnicki-Ocwieja, T., van Dijk, K., and Alfano, J. R. (2005). The hrpK operon of Pseudomonas syringae pv. tomato DC3000 encodes two proteins secreted by the type III (Hrp) protein secretion system: HopB1 and HrpK, a putative type III translocator. J. Bacteriol. 187, 649–663. doi: 10.1128/JB.187.2.649-663.2005
Potnis, N., Timilsina, S., Strayer, A., Shantharaj, D., Barak, J. D., Paret, M. L., et al. (2015). Bacterial spot of tomato and pepper: diverse Xanthomonas species with a wide variety of virulence factors posing a worldwide challenge. Mol. Plant Pathol. 16, 907–920. doi: 10.1111/mpp.12244
Radis-Baptista, G., Campelo, I. S., Morlighem, J. R. L., Melo, L. M., and Freitas, V. J. F. (2017). Cell-penetrating peptides (CPPs): from delivery of nucleic acids and antigens to transduction of engineered nucleases for application in transgenesis. J. Biotechnol. 252, 15–26. doi: 10.1016/j.jbiotec.2017.05.002
Raymond, B., Young, J. C., Pallett, M., Endres, R. G., Clements, A., and Frankel, G. (2013). Subversion of trafficking, apoptosis, and innate immunity by type III secretion system effectors. Trends Microbiol. 21, 430–441. doi: 10.1016/j.tim.2013.06.008
Römer, P., Hahn, S., Jordan, T., Strauss, T., Bonas, U., and Lahaye, T. (2007). Plant pathogen recognition mediated by promoter activation of the pepper Bs3 resistance gene. Science 318, 645–648. doi: 10.1126/science.1144958
Römer, P., Strauss, T., Hahn, S., Scholze, H., Morbitzer, R., Grau, J., et al. (2009). Recognition of AvrBs3-like proteins is mediated by specific binding to promoters of matching pepper Bs3 alleles. Plant Physiol. 150, 1697–1712. doi: 10.1104/pp.109.139931
Rossier, O., Van den Ackerveken, G., and Bonas, U. (2000). HrpB2 and HrpF from Xanthomonas are type III-secreted proteins and essential for pathogenicity and recognition by the host plant. Mol. Microbiol. 38, 828–838. doi: 10.1046/j.1365-2958.2000.02173.x
Rossier, O., Wengelnik, K., Hahn, K., and Bonas, U. (1999). The Xanthomonas Hrp type III system secretes proteins from plant and mammalian pathogens. Proc. Natl. Acad. Sci. U.S.A. 96, 9368–9373. doi: 10.1073/pnas.96.16.9368
Rüssmann, H., Kubori, T., Sauer, J., and Galan, J. E. (2002). Molecular and functional analysis of the type III secretion signal of the Salmonella enterica InvJ protein. Mol. Microbiol. 46, 769–779. doi: 10.1046/j.1365-2958.2002.03196.x
Rüter, C., and Schmidt, M. A. (2017). Cell-penetrating bacterial effector proteins: better tools than targets. Trends Biotechnol. 35, 109–120. doi: 10.1016/j.tibtech.2016.08.002
Rüter, C., Buss, C., Scharnert, J., Heusipp, G., and Schmidt, M. A. (2010). A newly identified bacterial cell-penetrating peptide that reduces the transcription of pro-inflammatory cytokines. J. Cell. Sci. 123, 2190–2198. doi: 10.1242/jcs.063016
Samudrala, R., Heffron, F., and McDermott, J. E. (2009). Accurate prediction of secreted substrates and identification of a conserved putative secretion signal for type III secretion systems. PLoS Pathog. 5:e1000375. doi: 10.1371/journal.ppat.1000375
Santos, A. S., and Finlay, B. B. (2015). Bringing down the host: enteropathogenic and enterohaemorrhagic Escherichia coli effector-mediated subversion of host innate immune pathways. Cell. Microbiol. 17, 318–332. doi: 10.1111/cmi.12412
Scharnert, J., Greune, L., Zeuschner, D., Lubos, M. L., Alexander Schmidt, M., and Rüter, C. (2013). Autonomous translocation and intracellular trafficking of the cell-penetrating and immune-suppressive effector protein YopM. Cell. Mol. Life Sci. 70, 4809–4823. doi: 10.1007/s00018-013-1413-2
Schechter, L. M., Roberts, K. A., Jamir, Y., Alfano, J. R., and Collmer, A. (2004). Pseudomonas syringae type III secretion system targeting signals and novel effectors studied with a Cya translocation reporter. J. Bacteriol. 186, 543–555. doi: 10.1128/JB.186.2.543-555.2004
Schechter, L. M., Valenta, J. C., Schneider, D. J., Collmer, A., and Sakk, E. (2012). Functional and computational analysis of amino acid patterns predictive of type III secretion system substrates in Pseudomonas syringae. PLoS ONE 7:e36038. doi: 10.1371/journal.pone.0036038
Schechter, L. M., Vencato, M., Jordan, K. L., Schneider, S. E., Schneider, D. J., and Collmer, A. (2006). Multiple approaches to a complete inventory of Pseudomonas syringae pv. tomato DC3000 type III secretion system effector proteins. Mol. Plant Microbe Interact. 19, 1180–1192. doi: 10.1094/MPMI-19-1180
Scheibner, F., Hartmann, N., Hausner, J., Lorenz, C., Hoffmeister, A. K., and Büttner, D. (2017). The type III secretion chaperone HpaB controls the translocation of effector and non-effector proteins from Xanthomonas campestris pv. vesicatoria. Mol. Plant Microbe Interact. doi: 10.1094/MPMI-06-17-0138-R. [Epub ahead of print].
Scheibner, F., Schulz, S., Hausner, J., Marillonnet, S., and Büttner, D. (2016). Type III-dependent translocation of HrpB2 by a non-pathogenic hpaABC mutant of the plant-pathogenic bacterium Xanthomonas campestris pv. vesicatoria. Appl. Environ. Microbiol. 82, 3331–3347. doi: 10.1128/AEM.00537-16
Schesser, K., Frithz-Lindsten, E., and Wolf-Watz, H. (1996). Delineation and mutational analysis of the Yersinia pseudotuberculosis YopE domains which mediate translocation across bacterial and eukaryotic cellular membranes. J. Bacteriol. 178, 7227–7233. doi: 10.1128/jb.178.24.7227-7233.1996
Schlumberger, M. C., Muller, A. J., Ehrbar, K., Winnen, B., Duss, I., Stecher, B., et al. (2005). Real-time imaging of type III secretion: Salmonella SipA injection into host cells. Proc. Natl. Acad. Sci. U.S.A. 102, 12548–12553. doi: 10.1073/pnas.0503407102
Schreiber, T., Sorgatz, A., List, F., Bluher, D., Thieme, S., Wilmanns, M., et al. (2015). Refined requirements for protein regions important for activity of the TALE AvrBs3. PLoS ONE 10:e0120214. doi: 10.1371/journal.pone.0120214
Schulze, S., Kay, S., Büttner, D., Egler, M., Eschen-Lippold, L., Hause, G., et al. (2012). Analyses of new type III effectors from Xanthomonas uncover XopB and XopS as suppressors of plant immunity. New Phytol. 195, 894–911. doi: 10.1111/j.1469-8137.2012.04210.x
Sory, M. P., Boland, A., Lambermont, I., and Cornelis, G. R. (1995). Identification of the YopE and YopH domains required for secretion and internalization into the cytosol of macrophages, using the cyaA gene fusion approach. Proc. Natl. Acad. Sci. U.S.A. 92, 11998–12002. doi: 10.1073/pnas.92.26.11998
Szczesny, R., Büttner, D., Escolar, L., Schulze, S., Seiferth, A., and Bonas, U. (2010a). Suppression of the AvrBs1-specific hypersensitive response by the YopJ effector homolog AvrBsT from Xanthomonas depends on a SNF1-related kinase. New Phytol. 187, 1058–1074. doi: 10.1111/j.1469-8137.2010.03346.x
Szczesny, R., Jordan, M., Schramm, C., Schulz, S., Cogez, V., Bonas, U., et al. (2010b). Functional characterization of the Xps and Xcs type II secretion systems from the plant pathogenic bacterium Xanthomonas campestris pv. vesicatoria. New Phytol. 187, 983–1002. doi: 10.1111/j.1469-8137.2010.03312.x
Szurek, B., Rossier, O., Hause, G., and Bonas, U. (2002). Type III-dependent translocation of the Xanthomonas AvrBs3 protein into the plant cell. Mol. Microbiol. 46, 13–23. doi: 10.1046/j.1365-2958.2002.03139.x
Takeuchi, T., and Futaki, S. (2016). Current understanding of direct translocation of arginine-rich cell-penetrating peptides and its internalization mechanisms. Chem. Pharm. Bull. 64, 1431–1437. doi: 10.1248/cpb.c16-00505
Tejeda-Dominguez, F., Huerta-Cantillo, J., Chavez-Duenas, L., and Navarro-Garcia, F. (2017). A novel mechanism for protein delivery by the type 3 secretion system for extracellularly secreted proteins. MBio 8, e00184–e00187. doi: 10.1128/mBio.00184-17
Teper, D., Burstein, D., Salomon, D., Gershovitz, M., Pupko, T., and Sessa, G. (2016). Identification of novel Xanthomonas euvesicatoria type III effector proteins by a machine-learning approach. Mol. Plant Pathol. 17, 398–411. doi: 10.1111/mpp.12288
Thieme, F., Szczesny, R., Urban, A., Kirchner, O., Hause, G., and Bonas, U. (2007). New type III effectors from Xanthomonas campestris pv. vesicatoria trigger plant reactions dependent on a conserved N-myristoylation motif. Mol. Plant Microb. Interact. 20, 1250–1261. doi: 10.1094/MPMI-20-10-1250
Van Engelenburg, S. B., and Palmer, A. E. (2008). Quantification of real-time Salmonella effector type III secretion kinetics reveals differential secretion rates for SopE2 and SptP. Chem. Biol. 15, 619–628. doi: 10.1016/j.chembiol.2008.04.014
Vidal, J. E., and Navarro-Garcia, F. (2008). EspC translocation into epithelial cells by enteropathogenic Escherichia coli requires a concerted participation of type V and III secretion systems. Cell. Microbiol. 10, 1975–1986. doi: 10.1111/j.1462-5822.2008.01181.x
Wang, Y., Sun, M., Bao, H., Zhang, Q., and Guo, D. (2013). Effective identification of bacterial type III secretion signals using joint element features. PLoS ONE 8:e59754. doi: 10.1371/journal.pone.0059754
Weber, E., Engler, C., Gruetzner, R., Werner, S., and Marillonnet, S. (2011a). A modular cloning system for standardized assembly of multigene constructs. PLoS ONE 6:e16765. doi: 10.1371/journal.pone.0016765
Weber, E., Gruetzner, R., Werner, S., Engler, C., and Marillonnet, S. (2011b). Assembly of designer TAL effectors by golden gate cloning. PLoS ONE 6:e19722. doi: 10.1371/journal.pone.0019722
Weber, E., Ojanen-Reuhs, T., Huguet, E., Hause, G., Romantschuk, M., Korhonen, T. K., et al. (2005). The type III-dependent Hrp pilus is required for productive interaction of Xanthomonas campestris pv. vesicatoria with pepper host plants. J. Bacteriol. 187, 2458–2468. doi: 10.1128/JB.187.7.2458-2468.2005
Wengelnik, K., Rossier, O., and Bonas, U. (1999). Mutations in the regulatory gene hrpG of Xanthomonas campestris pv. vesicatoria result in constitutive expression of all hrp genes. J. Bacteriol. 181, 6828–6831.
Werner, S., Breus, O., Symonenko, Y., Marillonnet, S., and Gleba, Y. (2011). High-level recombinant protein expression in transgenic plants by using a double-inducible viral vector. Proc. Natl. Acad. Sci. U.S.A. 108, 14061–14066. doi: 10.1073/pnas.1102928108
Winnen, B., Schlumberger, M. C., Sturm, A., Schupbach, K., Siebenmann, S., Jenny, P., et al. (2008). Hierarchical effector protein transport by the Salmonella Typhimurium SPI-1 type III secretion system. PLoS ONE 3:e2178. doi: 10.1371/journal.pone.0002178
Keywords: transcription activator-like effector, AvrBs3, Xanthomonas, type III secretion, chaperone, translocon
Citation: Scheibner F, Marillonnet S and Büttner D (2017) The TAL Effector AvrBs3 from Xanthomonas campestris pv. vesicatoria Contains Multiple Export Signals and Can Enter Plant Cells in the Absence of the Type III Secretion Translocon. Front. Microbiol. 8:2180. doi: 10.3389/fmicb.2017.02180
Received: 10 August 2017; Accepted: 24 October 2017;
Published: 09 November 2017.
Edited by:
Gail Preston, University of Oxford, United KingdomReviewed by:
Brian H. Kvitko, University of Georgia, United StatesFernando Navarro-Garcia, Center for Research and Advanced Studies of the National Polytechnic Institute (CINVESTAV), Mexico
Copyright © 2017 Scheibner, Marillonnet and Büttner. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Daniela Büttner, ZGFuaWVsYS5idWV0dG5lckBnZW5ldGlrLnVuaS1oYWxsZS5kZQ==
 Felix Scheibner1
Felix Scheibner1 Sylvestre Marillonnet
Sylvestre Marillonnet Daniela Büttner
Daniela Büttner