- 1Shanxi Key Laboratory of Birth Defect and Cell Regeneration, MOE Key Laboratory of Coal Environmental Pathogenicity and Prevention, Department of Biochemistry and Molecular Biology, Shanxi Medical University, Taiyuan, China
- 2Key Laboratory of Biological Resources and Ecology of Pamirs Plateau in Xinjiang Uygur Autonomous Region, College of Life and Geographic Sciences, Kashi University, Kashi, China
- 3Department of Marine Biology, Institute of Evolution and Marine Biodiversity, Ocean University of China, Qingdao, China
- 4Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology, Qingdao, China
In the post-COVID-19 era, people are increasingly concerned about microbial infections, including fungal infections that have risen in recent years. However, the currently available antifungal agents are rather limited. Worse still, the widespread use of the antifungal agents has caused the emergence of antifungal resistance in Candida, Cryptococcus, and Aspergillus species. Therefore, the development of novel antifungals is urgently needed. Antimicrobial peptides (AMPs), as components of the first-line defense of the host, are found to exhibit broad antimicrobial activity against bacteria, fungi, parasites, viruses, and protozoa. AMPs with antifungal activity are specifically referred to as antifungal peptides (AFPs). AFPs are currently regarded as the most promising alternative to conventional antifungal agents due to the fact that they are highly selective and less prone to facilitate the selection of drug resistance. In this review, we present an overview of the origin and classification of natural AFPs as well as their modes of action. Additionally, the production of natural, semisynthetic, and synthetic AFPs with a view to greater levels of exploitation is discussed. Finally, we evaluate the current and potential applications of AFPs in clinics and in the food industry.
1 Introduction
Fungi are eukaryotic microorganisms ranging from giant mushrooms to tiny multicellular molds and unicellular yeasts. It is estimated that there are approximately 2–11 million fungi on Earth, of which only 150,600 are officially categorized (Phukhamsakda et al., 2022; Lücking et al., 2021; Baldrian et al., 2021). Fungi are widely distributed in the soil and the air, in lakes, rivers, and oceans, on plants and animals, and in food and clothing. In recent years, fungi have been found to be a part of the commensal microbiota at different sites of human bodies (e.g., oral cavity, intestine, skin, lung, and vagina), although it remains still controversial over what constitutes the standard mycobiome composition (Auchtung et al., 2018; Huffnagle and Noverr, 2013; Kapitan et al., 2018).
Fungi are beneficial to many aspects of our daily life, notably the production of bread, wine, beer, soy sauce, and certain cheeses. Fungi are also used as a source of food; for example, some mushrooms, morels, and truffles are epicurean delicacies (Campbell-Platt and Cook, 2008; Money, 2016; Mukherjee et al., 2018). However, fungi have a harmful side too. It has been reported that at least 300 species of fungi can cause infections in both human beings and animals (Gupta et al., 2017). Recently, a list of fungal priority pathogens has been presented by the World Health Organization (2022) to guide research, development, and public health action. The list includes 19 fungal pathogens that are ranked and categorized into three priority (critical, high, and medium priority) groups based on their mortality rate, infection rate, and difficulty in diagnosis and treatment. The critical group includes Candida albicans, Aspergillus fumigatus, Candida auris, and Cryptococcus neoformans; the high group contains Candida glabrata, Candida parapsilosis, Candida tropicalis, Fusarium spp., Histoplasma spp., Mucorales, and eumycetoma causative agents; and the medium group comprises Candida krusei, Pneumocystis jirovecii, Scedosporium spp., Cryptococcus gattii, Lomentospora prolificans, Coccidioides spp., Talaromyces marneffei, and Paracoccidioides spp. Fungal infections have become a serious threat to human health, especially for people with weakened immune systems and potential health problems, such as diabetes mellitus, cancer, and HIV. It is estimated that fungal infections annually affect approximately 25% of the general population globally, causing high morbidity and mortality rates (Brown et al., 2012; Gamaletsou et al., 2018). Unfortunately, there are limited effective antifungal agents to treat fungal infections (Kathiravan et al., 2012). Currently, only four classes of antifungal drugs, i.e., polyenes (e.g., amphotericin B), triazoles (e.g., fluconazole), echinocandins (e.g., caspofungin), and fluorinated pyrimidines (e.g., 5-flucytosine), are available for the choice of systemic therapy of fungal diseases, and most of them, especially amphotericin B, can induce nephrotoxicity and hematotoxicity (Wang et al., 2024; Turcu et al., 2009). Worse still, the restricted spectrum and widespread use of the antifungal agents have caused the emergence of antifungal resistance in Candida, Cryptococcus, and Aspergillus species (Fisher et al., 2018; Lestrade et al., 2019; Pfaller et al., 2019). Furthermore, some fungal pathogens, such as Mucorales, C. auris, and some molds, are intrinsically resistant to the drugs above and difficult to treat at present. These all prompt an urgent need for the development of new antifungal agents with high efficiency and low toxicity. Of note, fungal cells are eukaryotic, and the development of selective antifungals is thus a particularly great challenge to identify pathogen-specific targets that are not present in human cells.
In addition to infection of humans and animals, fungi can also cause food spoilage, which leads to economic losses and may affect human health. Food-spoiling fungi and their mycotoxins released contaminate approximately 25% of raw materials produced by agriculture worldwide (World Health Organization (WHO), 1999). Therefore, the control and prevention of fungal pathogens and foodborne poisoning is one of the most important public health challenges that we are facing today. A number of physical, chemical, and biological methods have been applied to control fungal pathogens and mycotoxin contamination, including green and emerging technologies such as ionizing and non-ionizing radiation, ultrasound, pulsed electric field and high-pressure processing, and biological preservation. Among them, the use of antifungal compounds is regarded as an alternative environmentally friendly strategy. However, the antifungal compounds currently available for this use are quite limited (Thery et al., 2019). Antimicrobial peptides (AMPs) can really be a good candidate, given their lower likelihood of selecting resistance.
AMPs, first described by Dubos (1939) from Bacillus brevis, are short peptides with rapid microbicidal effects that are typically composed of <100 amino acids. Albeit highly diverse in amino acid sequence, AMPs usually possess a net positive charge and hydrophobic regions and facilitate interactions with membranes. AMPs kill microbes via several mechanisms, including binding to or inserting into microbial membranes (which has fatal depolarization of the normally polarized membrane), forming physical pores, disrupting the usual distribution of lipids between the bilayer leaflets, and damaging critical intracellular targets (Gennaro and Zanetti, 2000; Hancock, 2000; Nawrocki et al., 2014). Because of their multi-point and multi-level mechanisms of action, the likelihood of developing resistance to AMPs is relatively low.
AMPs, as components of the first-line defense of the host, are produced by all organisms, from bacteria to humans (Faruck et al., 2016; Kang et al., 2015; Shishido et al., 2015; Silva et al., 2014; Tam et al., 2015), and exhibit broad antimicrobial activity against bacteria, fungi, parasites, viruses, and protozoa (Hancock and Chapple, 1999). Currently, there are 1,479 peptides with antifungal properties documented in the Antimicrobial Peptide Database (APD3). In the majority of cases, the classification of AFPs is based on the peptide origin: natural, semisynthetic, or synthetic (De Lucca, 2000). In this study, we present an overview of the origin and classification of natural AFPs and their modes of action. In addition, the production of natural, semisynthetic, and synthetic AFPs with a view to greater levels of exploitation is discussed. Finally, we evaluate the current and potential applications of AFPs in clinics and in the food industry.
2 Origin and classification of AFPs
The innate immunological components including endogenic peptides of organisms could rapidly respond to invading pathogens to avoid their adverse effects on the host. Natural AFPs are produced by a number of species of bacteria, archaea, and eukarya isolated from natural sources (De Lucca and Walsh, 1999). They typically adopt an α-helix structure, β-hairpin or sheet (containing two cysteine residues) structure, or mixed α-helix/β-sheet structure upon interaction with membranes. Some natural AFPs are rich in specific amino acids such as glycine, proline, arginine, histidine, and tryptophan, and accordingly, they are often classified as glycine-rich, proline-rich, arginine-rich, histidine-rich, and tryptophan-rich AFPs (Bondaryk et al., 2017).
2.1 AFPs from microorganisms
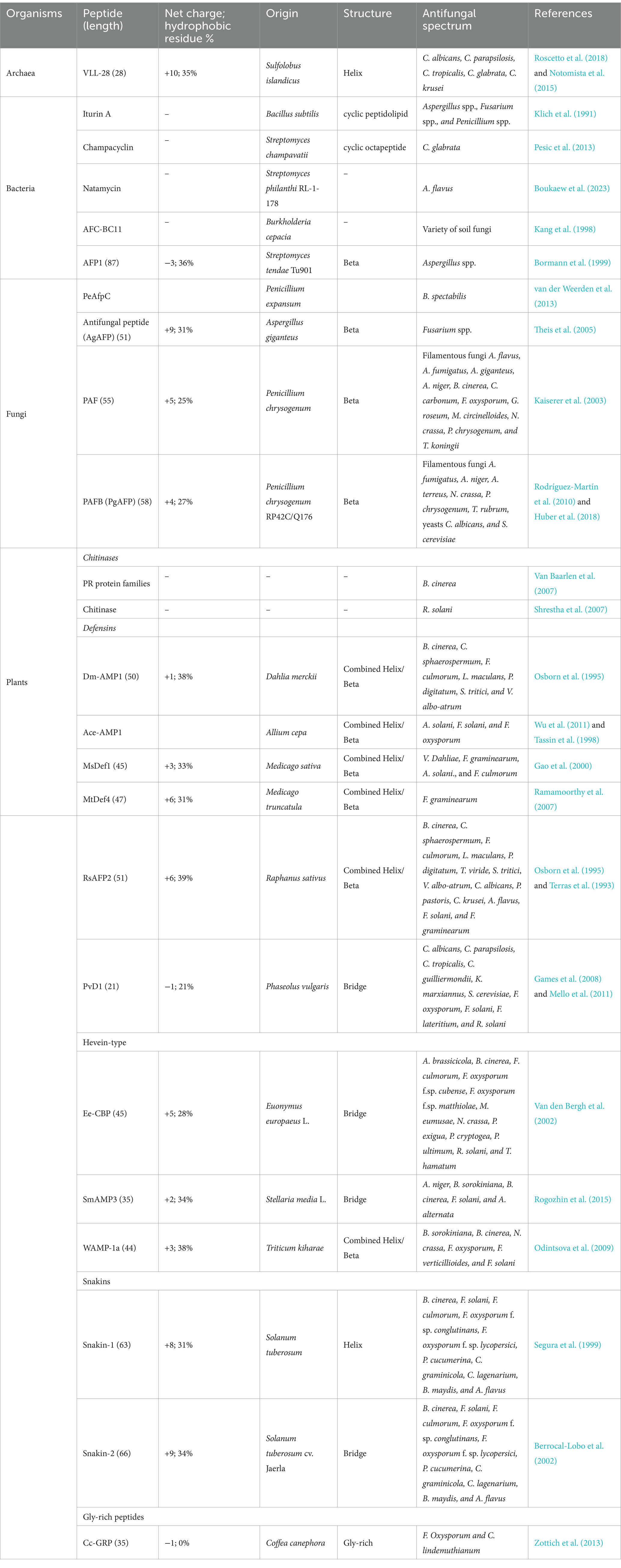
The AFPs produced by microorganisms, including bacteria and archaea (both prokaryotes) as well as fungi (eukaryotes), can be secreted into extracellular surroundings and offer a competitive advantage in ecological niches. Bacteria generate a number of different AFPs (Table 1). The first example of an archaeal antimicrobial peptide with antifungal activity is VLL-28, isolated from the archaeon Sulfolobus islandicus, which showed antifungal activity against 10 clinical isolates of Candida spp. (Roscetto et al., 2018). The well-known AFP-producing bacteria include the genera Bacillus, Lactobacillus, Streptomyces, and Burkholderia. For example, the iturin A produced by Bacillus subtilis exhibits a conspicuous antifungal activity against Aspergillus spp., Fusarium spp., and Penicillium spp. (Klich et al., 1991); the peptide mixture generated by Lactobacillus plantarum TE10 suppresses Aspergillus flavus in maize seeds, displaying a considerable potential for the development of bio-control agents (Muhialdin et al., 2020); the champacyclin, a head-to-tail cyclic octapeptide obtained from Streptomyces champavatii, inhibits the growth of the yeast C. glabrata (Pesic et al., 2013); the natamycin from Streptomyces philanthi RL-1-178 possesses a fungicidal activity against A. flavus (Boukaew et al., 2023); and the AFC-BC11, a lipopeptide isolated from Burkholderia cepacia, exerts an antifungal activity toward a variety of soil fungi (Kang et al., 1998). Similarly, eukaryotic microorganisms such as filamentous fungi and yeasts also produce a variety of AFPs (Table 1). For instance, the peptide PeAfpC produced by the filamentous fungus Penicillium expansum can effectively inhibit the growth of Byssochlamys spectabilis, which is capable of causing the spoilage of pasteurized juices and canned foods (van der Weerden et al., 2013). In addition, the two peptides, namely, PAF and PAFB, generated by the filamentous fungus Penicillium chrysogenum, are capable of exerting antifungal activity against a variety of filamentous fungi, with PAFB suppressing the growth of some toxigenic molds (Kaiserer et al., 2003; Rodríguez-Martín et al., 2010; Huber et al., 2018).
2.2 AFPs from plants
Plants have developed various mechanisms in their innate immune systems to protect themselves against fungal attacks, including soluble peptides and proteins released from plants with antifungal activities (Chiu et al., 2022). These peptides/proteins that are constitutively synthesized are able to trigger defense responses in plants. On the basis of sequence, cysteine residues, and function, the plant-sourced AFPs can be divided into different families (Table 1), including chitinases, defensins, and snakins, as well as hevein-type and gly-rich peptides (Tam et al., 2015; Yan et al., 2015). Chitinases are among the best-known and most-studied plant AFPs. They display strong antifungal activity against a wide range of phytopathogenic fungi, including Botrytis cinerea (Van Baarlen et al., 2007) and Rhizoctonia solani (Shrestha et al., 2007). Plant chitinases have been used to treat fungal infections as exogenously applied pest control agents (Karasuda et al., 2003). The Dm-AMP1 is a defensin peptide found in Dahlia merckii that shows inhibitory activity against a variety of fungi, such as B. cinerea and Leptosphaeria maculans in the presence of CaCl2 and KCl (Osborn et al., 1995). Another defensin Ace-AMP1, a potent AFP found in onion (Allium cepa) seeds, has been applied to control the tomato early blight disease caused by the pathogen Alternaria solani (Wu et al., 2011). The peptide Ee-CBP containing five disulfide bridges obtained from the bark of spindle tree (Euonymus europaeus) is a hevein-type AFP, exhibiting antifungal activity against various fungi including Alternaria brassicicola (50% growth inhibition IC50 = 3 μg/mL), Phoma exigua (IC50 = 33 μg/mL), and Fusarium oxysporum f.sp. cubense (IC50 = 15 μg/mL) (Van den Bergh et al., 2002). The WAMP-1a, another hevein-type AFP from seeds of Triticum kiharae, shows high broad-spectrum inhibitory activity against a wide range of chitin-containing and non-chitin-containing pathogens including Bipolaris sorokiniana, Neurospora crassa, Fusarium verticillioides, and Fusarium solani (Odintsova et al., 2009). The snakins 1 and 2 are representative AFPs of snakin family identified from Solanum tuberosum, exerting antifungal activity against fungi such as B. cinerea, F. solani, A. flavus, Colletotrichum graminicola, and Bipolaris maydis (Segura et al., 1999; Berrocal-Lobo et al., 2002). The Cc-GRP is a gly-rich AFP identified from Coffea canephora that can combat fungi such as F. oxysporum and Colletotrichum lindemuthianum (Zottich et al., 2013).
2.3 AFPs from animals
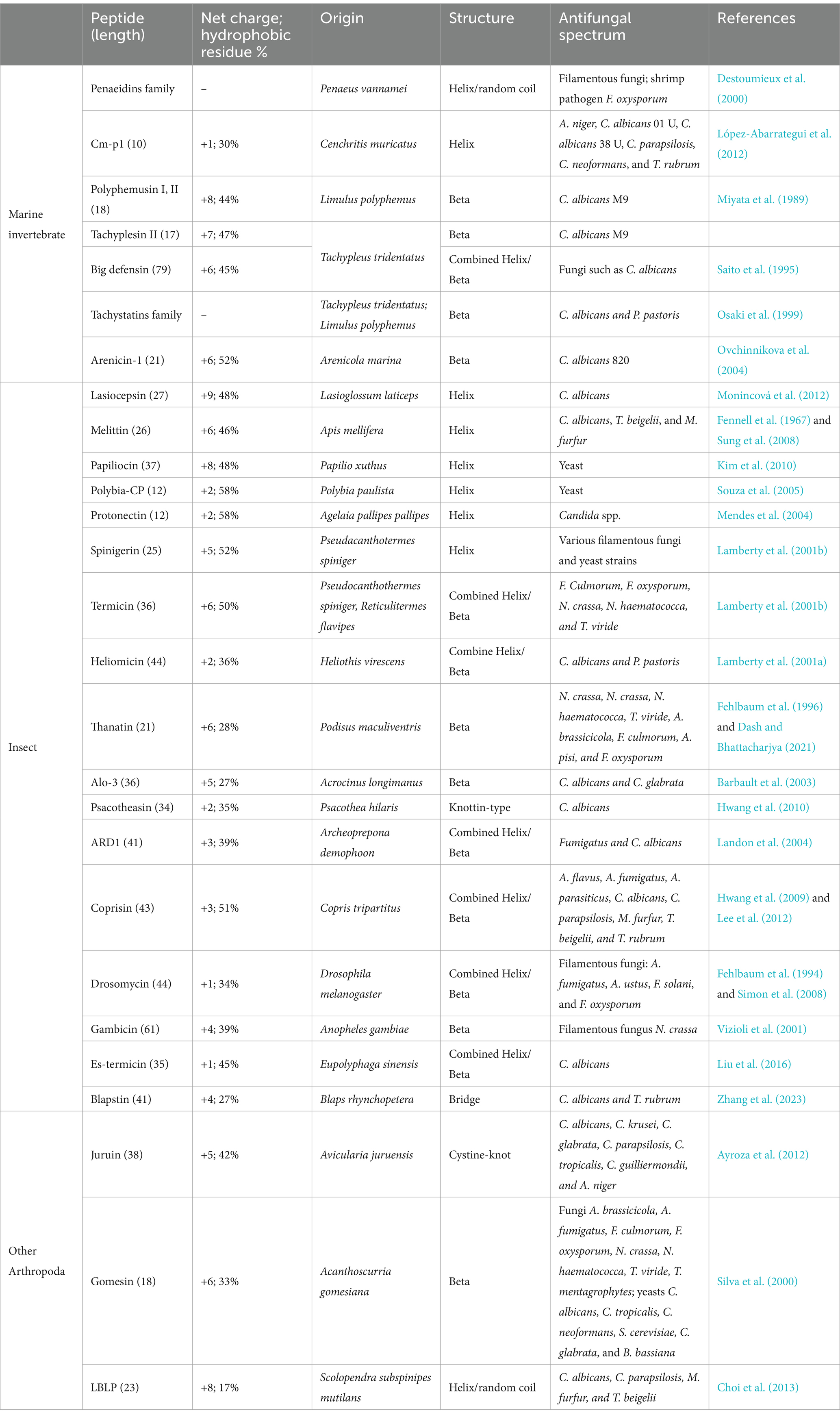
Many animal-sourced AFPs have been found to be part of the innate immune responses of both invertebrates and vertebrates (Tables 2, 3). As invertebrates lack adaptive immunity, AMPs play a significant role in their immune response and comprise an essential source of AFPs (Table 2). The AFPs obtained from marine invertebrates include penaeidins, Cm-p1, and tachystatins. The penaeidin family originates from shrimp, and currently, there are nine members available for this family in the ADP3 database (Destoumieux et al., 2000; Cuthbertson et al., 2002; An et al., 2016). The members of the penaeidin family show broad-spectrum fungicidal activity. For example, both penaeidins 2 and 3a are fungicidal against filamentous fungi and shrimp pathogen F. oxysporum (Destoumieux et al., 2000). Cm-p1 is a 10-mer short peptide isolated from marine snails. Cm-P1 has the ability to inhibit the growth of yeasts and filamentous fungi, while it shows little toxic effects on mammalian cells (López-Abarrategui et al., 2012). The tachystatin family consists of four members: tachystatin A, tachystatin B1, tachystatin B2, and tachystatin C, all of which have been identified in horseshoe crabs. All four members of the tachystatin family contain three disulfide bridges and have sequences similar to spider neurotoxins. Since horseshoe crabs are close relatives of spiders, tachystatins and neurotoxins may have evolved from a common ancestral peptide gene. Tachystatins are capable of binding to chitin and then exert their antifungal activity against C. albicans and Pichia pastoris (Osaki et al., 1999). AFPs are also found in insects. Representative AFPs from insects include melittin and thanatin. Melittin was isolated from bee venom by Neuman et al. (1952), which existed in hemolysin phospholipase A (Habermann, 1972). Melittin shows strong antifungal activity against various strains of fungi, including Aspergillus sp., Candida sp., Malassezia sp., Penicillium sp., and Trichoderma sp. (Memariani and Memariani, 2020). It is found that melittin exerts inhibitory activity against fungi via a series of combined mechanisms of inhibition of (1,3)-β-D-glucan synthase, membrane permeabilization, apoptosis induction by reactive oxygen species (ROS), and alterations in gene expression (Memariani and Memariani, 2020). Thanatin, a 21-residue peptide, was first isolated from the insect Podisus maculiventris (Fehlbaum et al., 1996). It is highly potent in inhibiting the growth of fungi at considerably low concentrations (Fehlbaum et al., 1996; Dash and Bhattacharjya, 2021). Protonectin was originally isolated from the venom of the neotropical social wasp Agelaia pallipes pallipes (Mendes et al., 2004). Later, protonectin was shown to have potent antifungal and fungicidal activity against C. glabrata, C. albicans, C. parapsilosis, C. tropicalis, and C. krusei by disturbing membrane integrity and inducing ROS production in yeast cells (Wang et al., 2015). Recently, a 41-amino acid peptide called blapstin, isolated from the Chinese medicinal beetle Blaps rhynchopetera, was shown to possess antifungal activity against C. albicans and Trichophyton rubrum. Cryo-scanning electron microscope (Cryo-SEM) observations showed that blapstin directly resulted in disruption in the cell structure of C. albicans and T. rubrum (Zhang et al., 2023).
Spiders and centipedes are also known to produce AFPs, such as juruin, gomesin, and lactoferricin B-like peptide (LBLP), especially in venoms. Juruin isolated from the venom of the Amazonian pink toe spider Avicularia juruensis has antifungal activity against filamentous fungi and yeasts, including Aspergillus niger, Beauveria bassiana, C. albicans, C. krusei, C. glabrata, C. parapsilosis, C. tropicalis, and Candida guilliermondii (Ayroza et al., 2012). Gomesin, an 18-amino acid AMP isolated from the hemolymph of the tarantula spider Acanthoscurria gomesiana, inhibits the development of filamentous fungus and yeast (Silva et al., 2000). LBLP, a 23-mer AMP derived from the centipede Scolopendra subspinipes mutilans, has been found to have antifungal and fungicidal activity against C. albicans, C. parapsilosis, Malassezia furfur, and Trichosporon beigelii by forming pores in the membrane, eventually leading to fungal cell death (Choi et al., 2013). Recently, LBLP has been reported to trigger mitochondrial disruption-mediated apoptosis by inhibiting respiration under nitric oxide accumulation in C. albicans (Kim et al., 2020).
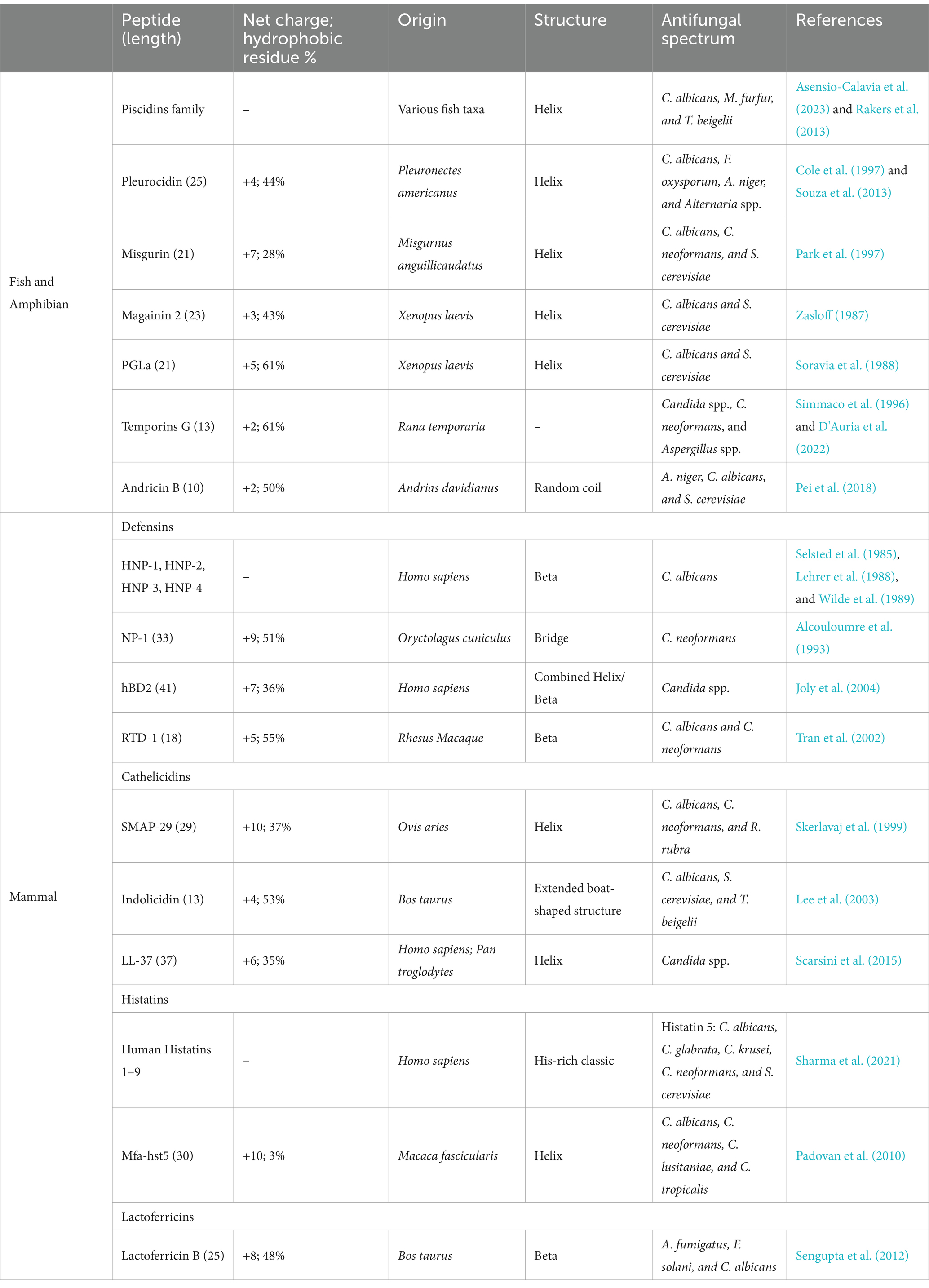
In vertebrates (Table 3), the immune system is divided into innate immunity and adaptive immunity. Adaptive immunity provides an effective and specific immune response against pathogens, while innate immunity consisting of the first line of defense is much quicker to respond to initial attacks. AMPs produced in response to pathogenic attacks form part of the first line of defense and are a source of AFPs. AFPs derived from vertebrates are usually produced on the skin, mucous membranes, and other areas that are easily exposed to microbial environments (López-Meza et al., 2011). It has been shown that the piscidins synthesized in the epithelia of gills, skin, stomach, and gut of a variety of teleost species exhibit antifungal activity against C. albicans, M. furfur, and T. beigelii through membrane disruption mode (Asensio-Calavia et al., 2023; Rakers et al., 2013). In addition, pleurocidin secreted by the skin of winter flounder inhibits the growth of Alternaria spp., C. albicans, F. Oxysporum, and A. niger (Cole et al., 1997; Souza et al., 2013). Recently, we have shown that AP10W, a short peptide derived from AP-2 complex subunit mu-A of zebrafish, displays conspicuous antifungal activities against the main fungal pathogens of human infections C. albicans and A. fumigatus. We also show that AP10W inhibits fungal biofilm formation and decrease pre-established fungal biofilms (Gong et al., 2022).
Similarly, the skin and secretory glands of amphibian frogs are also a rich source of AFPs, such as magainins and peptide glycine-leucine-amide (PGLa) identified in the clawed frog Xenopus laevis (Zasloff, 1987; Soravia et al., 1988), and temporins A, B, and L identified in the red frog Rana temporaria (Simmaco et al., 1996; Marcocci et al., 2018; Roscetto et al., 2021). Temporin G, recently isolated from the skin of Rana temporaria, is demonstrated to exert fungicidal ability against C. neoformans, Candida spp., and Aspergillus spp. In addition, temporin G reduces the metabolic activity of C. albicans cells, induces moderate membrane perturbation, and is effective against virulence factors of C. albicans (D'Auria et al., 2022).
In mammals, both neutrophils and epithelial cells are known to produce AFPs, including defensins, cathelicidins, histatins, and lactoferricins. Defensins are widely present in eukaryotes (fungi, plants, and animals), with four types of human defensins, known as HNP-1, HNP-2, HNP − 3, and HNP-4. All human defensins have been shown to possess candidacidal ability and are capable of influencing the ionic environment and the metabolic state of C. albicans cells (Selsted et al., 1985; Lehrer et al., 1988; Wilde et al., 1989). Cathelicidins have been identified in both humans and chimpanzees. LL-37, a 37-mer peptide derived from the N-terminal 37 residues of human cathelicidins, inhibits the growth of fungi including Aspergillus, Candida, Colletotrichum, Fusarium, Malassezia, Pythium, and Trichophyton. LL-37 exerts its fungal inhibition through several mechanisms, including cell wall integrity disruption, membrane permeabilization, and intracellular effects such as formation of autophagy-like structures, disturbance of endoplasmic reticulum homeostasis, induction of oxidative stress, inhibition of cell cycle progression, and alterations in gene expression (Memariani and Memariani, 2023). Histatins are histidine-rich peptides abundantly present in human saliva and the oral cavity (Oppenheim et al., 1988). Histatins exhibit a broad spectrum of antifungal activities and play an important role in controlling periodontal and oral fungal infections (Kavanagh and Dowd, 2004; Pólvora et al., 2018). Human histatins are composed of nine classes (human histatins 1–9), and histatin 5 is found to have strong fungicidal activity against C. albicans, C. glabrata, C. krusei, C. neoformans, and Saccharomyces cerevisiae (Sharma et al., 2021). Lactoferrin is a glycoprotein with AMP activity, found in saliva, milk, vaginal secretions, tears, and other exocrine secretions of mammals (cows, pigs, mice, and humans) (Moreno-Expósito et al., 2018; Rascón-Cruz et al., 2021; Drago-Serrano et al., 2018). Lactoferricin is found to have antifungal activity against a variety of fungi including Clavispora lusitaniae, Pichia kudriavzevii, Kluyveromyces marxianus, Meyerozyma guilliermondii, S. cerevisiae, Candida spp., and Cryptococcus spp.(Fernandes et al., 2020). Moreover, lactoferricin also shows inhibitory activity against the fungal biofilm formed by keratitis-associated fungal pathogens, such as A. fumigatus, F. solani, and C. albicans (Sengupta et al., 2012).
3 Modes of action of AFPs
Exploration of the modes of action of AFPs is a significant aspect of AFP research as the understanding of antifungal mechanisms can be a great help for researchers to further develop and design novel AFPs. In general, AFPs are known to function via three modes of action (Figure 1), i.e., inhibition of biosynthesis of cell wall components, interaction with membrane components, and interference with intracellular targets (Zhang et al., 2021). It is notable that some AFPs often function via multiple modes of action. For example, the fish-sourced peptide AP10W has been shown to exert its fungicidal activity through modes of combined actions, including interaction with the fungal cell walls via laminarin, mannan, and chitin, enhancement of cell wall permeabilization, induction of membrane depolarization, and increase in intracellular ROS generation (Gong et al., 2022).
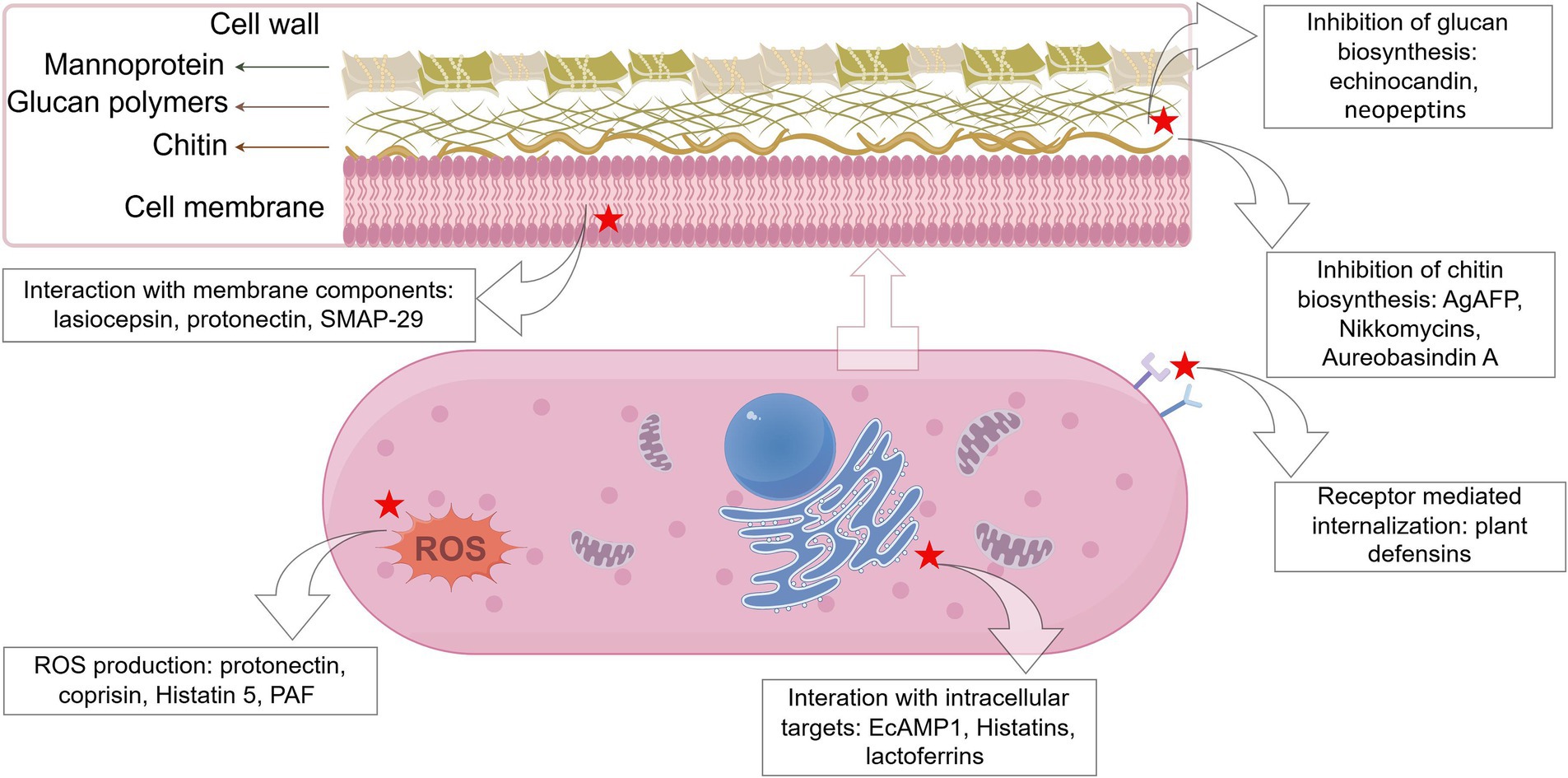
Figure 1. Schematic representation of the targets of some representative antifungal peptides. The red star marks the targets of AFPs (created by figdraw).
3.1 Inhibition of biosynthesis of cell wall components
The fungal cell wall, the first barrier of the cell to effectively resist the influence of the external environment, is mainly composed of chitin, mannans, glycoproteins, and glucans (β − 1,3-glucan, β − 1,6-glucan, α-1,3-glucan, α-1,4-glucan, and mixed β-1,3−/β-1,4-glucan) (Bowman and Free, 2006). It protects the fungal cell from the pressure of the external environment and maintains normal cell metabolism, ion exchange, and osmotic pressure (Cabib and Arroyo, 2013).
Chitin and β-1,3-glucan are both the major structural components of the cell walls of many fungi and play a key role in maintaining the structural integrity of fungal cell walls (Bowman and Free, 2006; Lenardon et al., 2010; Fleet, 1991). Some AFPs are inhibitors of chitin synthase or (1–3)-β-D-glucan synthase, capable of blocking the synthesis of cell wall components, which then disrupts the normal cell morphology and diminishes the cell’s capacity to regulate osmotic pressure. For example, the fungus-sourced AgAFP suppresses the activity of chitin synthases III and V, which are important for chitin biosynthesis (Hagen et al., 2007). The echinocandin, a cyclic hexapeptide isolated from several species of Aspergillus, inhibits β-1,3-D-glucan synthase necessary for glucan biosynthesis (Ciociola et al., 2016; Thorn et al., 2024). Neopeptins have been reported against some plant pathogenic fungi by inhibiting proteoheteroglycan and β-1,3-glucan synthesis, which could affect cell wall biosynthesis (Ubukata et al., 1984). The nikkomycins (a complex of nucleoside-peptides), an analog of chitin synthase natural substrate N-acetylglucosamine, also block the synthesis of chitin in C. albicans (McCarthy et al., 1985). Analogously, the aureobasidin A, a cyclic depsipeptide produced by Aureobasidium pullulans, interferes with fungal cell wall integrity by affecting actin assembly, chitin delocalization, and synthesis of sphingolipids (Endo et al., 1997; Nagiec et al., 1997).
3.2 Interaction with membrane components
AFPs, as part of AMPs, are usually small (< 100 amino acids) cationic polypeptides, with amphiphilic structures with hydrophobic domains capable of binding to lipids and positively charged hydrophilic domains capable of binding to water or negatively charged residues (Hollmann et al., 2018). This property of AFPs renders them able to form strong binding to the amphiphilic part of the cell membrane, which lays the structural basis for the interaction of AFPs with fungal cell membranes. Several models have been proposed to explain the action of membrane disruption caused by AFPs, such as the barrel-stave, toroidal pore, and carpet models (Zhang et al., 2021; Yeaman and Yount, 2003). According to the barrel-stave model, AFPs bind to lipid membranes and recognize each other to form a transmembrane pore (Theis and Stahl, 2004; Soltani et al., 2007). Ultimately, the cell membrane components are penetrated by AFPs, resulting in cell collapse and death. The toroidal model suggests that AFPs insert into the hydrophobic center of the cell membrane (Soltani et al., 2007; Le et al., 2017), triggering the phospholipid molecular layer to curve inward and form a mixed cavity randomly (Yang et al., 2001). As a consequence, the membrane structures are disordered, eventually leading to cell death. In the carpet model, AFPs interact only with the lipid head groups and are oriented parallel to the surface, forming a carpet-like pattern. Once AFPs on the membrane surface accumulate to a certain concentration threshold, they act as detergents by distorting the phospholipid bilayer, reducing the stability of the cell membrane and causing cell membrane disintegration and cell lysis (Lohner and Prenner, 1999; Zhang et al., 2020). For example, plant defensins Dm-AMP1 from dahlia (Dahlia merckii), RsAFP2 from radish (Raphanus sativus), and HsAFP1 from coral bells (Heuchera sanguinea) target specific binding sites on fungal cells (e.g., mannosyl diinositolphosphoryl ceramide from S. cerevisiae for Dm-AMP1 and glucosylceramide from P. pastoris for RsAFP2), leading to membrane permeabilization and eventually decreasing viability (Thevissen et al., 2003; Thevissen et al., 2004; Aerts et al., 2008). Similarly, protonectin, isolated from the venom of the neotropical social wasp Agelaia pallipes pallipes, exerts antifungal/fungicidal activities against the tested fungal cells via interaction with lipid membranes, disruption of the membrane integrity, and induction of the production of intracellular ROS (Wang et al., 2015). MAF-1A, a linear 26-amino acid peptide, is derived from the carboxy-terminal functional domain of the antifungal peptide-1 (MAF-1) isolated from the hemolymph of Musca domestica larvae (Zhou et al., 2016). It has been shown that MAF-1A disrupts the cell membrane of C. albicans and then enters the cell where it binds and interacts with nucleic acids (Cheng et al., 2021). However, it must be pointed out that although the interaction of AMPs (including AFPs) with biological membranes plays an important role in the entire process of their antimicrobial actions, membrane disruption is a rather complex and dynamic process, which still needs detailed study to refine the specific mechanisms (Haney et al., 2019).
3.3 Interference with intracellular targets
AFPs can also interact with fungal intracellular targets, including nucleotides (DNA and RNA), proteins, and organelles. For example, CGA-N9, an antifungal peptide derived from human chromogranin A (CGA), has been found to exert antifungal activity toward C. tropicalis by attenuating mitochondrial function (Li et al., 2019). Psd1 defensin from pea (Pisum sativum) has been found to enter N. crassa cells, localize to the nuclei, and interfere with the cell cycle, probably via interacting with cyclin F (Lobo et al., 2007). Similarly, histatin 5 binds to mitochondria after crossing fungal membranes via transmembrane potentials or receptors (without damaging the plasma membrane) and then induces non-lytic release of adenosine triphosphate (ATP) into the cytoplasm. The released ATP binds to purinergic receptors on the cell surface, leading to the inhibition of mitochondrial respiration and the generation of ROS, which then causes damage to nucleic acids and organelles (Kavanagh and Dowd, 2004; Helmerhorst et al., 2001). The antifungal peptide EcAMP1 isolated from barnyard grass shows strong antifungal action toward species of the Fusarium genus. It has been found that EcAMP1 first binds to one or several abundant components of the fungal cell surface and is then internalized by the fungal cell and accumulates in a vesicular structure in the cytoplasm without disturbing the integrity of the membrane (Nolde et al., 2011).
4 Production of AFPs
To effectively explore the structure–activity relationships, efficacy, and safety, especially in clinical treatments, it is necessary to produce sufficient amounts of highly pure AFPs. There are currently three major approaches to achieving this goal, i.e., direct isolation from various organisms, recombinant expression, and chemical synthesis (Fernández de Ullivarri et al., 2020).
4.1 Natural production
AFPs are naturally isolated from different species of organisms. Currently, only a limited number of AFPs are obtained from their natural sources for clinical use as the isolation of these natural peptides is time-consuming and expensive due to their relatively low abundance (Vriens et al., 2014). The methods commonly used now for industrial-scale AFP production from natural sources are microbial fermentation and proteolysis.
The natural echinocandins, echinocandin B, pneumocandin B0, and FR901379, are produced for commercial purposes from Aspergillus rugulosus, Glarea lozoyensis, and Coleophoma empetri, respectively. The production of echinocandins is an exceptional example of AFPs produced by microbial fermentation and further chemical modification in the case of the semisynthetic variants. The fermentation process is critical to obtain a competitive product and thus needs optimizing to increase the amount of natural echinocandins and control their purification costs. Temperature is a key factor in fermentation production and influences the overall production costs of AFPs (Emri et al., 2013). The optimal temperatures for the production of echinocandin B, pneumocandin B 0, and FR901379 are similar, all below 30°C. However, the optimal growth temperatures of the strains of A. rugulosus, G. lozoyensis, and C. empetri exhibited considerable variation. The optimal growth temperatures of G. lozoyensis and C. empetri were observed to be lower than 30°C, whereas the optimal growth temperatures of A. rugulosus were found to be higher than 30°C, reaching 37°C; it exhibited poor growth at the temperatures utilized for the production of echinocandin B. The addition of complex nitrogen sources and plant oils also influences the generation of echinocandins (Emri et al., 2013).
Production of pharmaceutical-grade AFPs can be achieved through enzymatic hydrolysis of proteins, resulting in the release of encrypted peptides. The process generally involves three steps, i.e., acquisition of raw materials, protein hydrolysis, and fractionation and isolation. By-products from dairy, fish, and meat industries are all suitable sources of proteins (Sibel Akalin, 2014; Ryder et al., 2016). For proteolysis, the utilization of immobilized enzymes possesses several advantages over the conventional soluble enzymes, such as milder and controlled conditions and recycling of enzymes used (Sewczyk et al., 2018). The final step of fractionation and isolation of AFPs includes ultrafiltration, precipitation with solvents, and liquid chromatography techniques, which are usually expensive (Brady et al., 2008). Fortunately, an alternative and cost-effective method, electro-membrane filtration, which combines electrophoresis with conventional membrane filtration, has been established (Bazinet and Firdaous, 2013) and is increasingly being applied for the fractionation and isolation of AFPs.
4.2 Recombinant production
Recombinant expression presents a solid option for producing AFPs at low cost and high efficiency. In addition, sequencing technologies have generated a vast amount of genomic and transcriptomic data, providing valuable resources for discovering and designing new and more active AFPs (Amaral et al., 2012; Porto et al., 2012; Tracanna et al., 2017). Bacteria (mainly Escherichia coli), yeasts (mainly P. pastoris), and plants (e.g., tobacco plant Nicotiana tabacum) are the most common expression platforms for recombinant proteins.
E. coli BL21 (DE3), deficient in proteases that may lead to protein degradation, is by far the most commonly used bacterial species as a host for recombinant production of proteins (Li, 2011). Many E. coli strains are unable to export proteins across their outer membrane. As a result, in the majority of cases, proteins are secreted into the cytoplasm or periplasm, leading to the formation of inclusion bodies (Singh et al., 2015). Examples of AFPs produced in E. coli include lactoferricin B, nikkomycin, magainin-2, and cecropin (Fernández de Ullivarri et al., 2020).
In contrast to prokaryotic E. coli, eukaryotic yeasts are capable of implementing certain post-translational modifications to heterologous recombinant proteins. Yeasts commonly used as hosts for recombinant proteins include S. cerevisiae, P. pastoris, Kluyveromyces lactis, Yarrowia lipolytica, Schizosaccharomyces pombe, and Hansenula polymorpha. For example, the recombinant antifungal proteins serum albumin and hen lysozyme were produced by K. lactis and S. cerevisiae, respectively (Vieira Gomes et al., 2018). In addition, filamentous fungi such as A. pullulans, P. Chrysogenum, and P. digitatum have also been used to generate AFPs. Some examples of AFPs produced by yeasts or filamentous fungi are protegrin-1, porcine lactoferrin, aureobasidin A, and NFAP2 (Fernández de Ullivarri et al., 2020; Slightom et al., 2009).
Plants have been explored as hosts for recombinant expression of AFPs due to their capacity for large-scale production and their cost-effectiveness. The advantages of plants as expression systems are their capability to perform appropriate glycosylation, folding, and disulfide bond formation of recombinant AFPs. Different genetic approaches have been employed to produce AMPs in plants including using whole plants, tissue-specific expression, tissue culture, or transient expression (Holaskova et al., 2015). Whole tobacco plants have been utilized to produce lactoferrin and dermaseptin with a higher yield (Chahardoli et al., 2018; Shams et al., 2019).
4.3 Chemical synthesis
The chemical synthesis of peptides allows scientists to design and produce specific sequences of AFPs on demand. Ideally, an AFP should be short. De novo peptide design may help reduce production costs, potential toxicity, and lability and increase bioactivity in vivo (Steckbeck et al., 2014). Peptide chemical synthesis is divided into two types: solid- (SPPS) or liquid (solution)-phase peptide synthesis (LPPS). Currently, fluorenylmethyloxycarbonyl (Fmoc) SPPS is the preferred method for chemical synthesis of AFPs due to the versatility and low cost of very high-quality building blocks. We have recently designed and synthesized a peptide named AP10W that shows increased antifungal activity (Gong et al., 2022).
5 Applications and prospects
A primary focus of natural AFP research is the development of novel specific antifungal drugs against fungal infections (Matejuk et al., 2010). A number of AFPs, such as human lactoferricin-based hLF1-11, novexatin, and pexiganan, are in preclinical development, but few of them reach the clinical stage (Ciociola et al., 2016; Duncan and O’Neil, 2013; Koo and Seo, 2019). The hLF1-11 is proposed for intravenous usage in immunocompromised recipients of stem cell transplants for treating both bacterial and fungal infections; novexatin, a cationic peptide generated from defensins, is suggested for treating fungal toe infections; and pexiganan, an analog of peptide magainin (extracted from the skin of Xenopus laevis) with 22 amino acid residues, shows robust antimicrobial activity against bacterial and fungal pathogens. In addition, human histatin-based PAC-113 is also under clinical trials as a mouth rinse for oral candidiasis in patients with human immunodeficiency virus (HIV). AFPs undergoing clinical trials are generally designed for topical use because topical administration of peptides can overcome the inherent limitation related to poor stability in physiological fluids, due to their susceptibility to proteases. In addition, in certain areas of the body, such as skin, oral cavity, and vagina, where fungal infections may occur, physiological pH values and salt concentrations are compatible with the optimum activity of AFPs (Duncan and O’Neil, 2013; Wei et al., 2007; Lombardi et al., 2015).
Emerging resistance to conventional antifungals and serious side effects of drugs currently available demand urgent development of novel strategies for protection against fungal pathogens. This goal can be achieved with combination therapy, in which conventional antifungals are used together with other antifungal drugs or AFPs to increase the treatment efficacy compared to single-drug therapy. For instance, lactoferrin-derived peptides Lf (1–11) and bLfcin both exhibit synergy with azole and amphotericin B, which reduce the minimal inhibitory concentrations against Candida spp. (Wakabayashi et al., 1996; Lupetti et al., 2003; Fernandes and Carter, 2017). AFPs may also be conjugated with virus-like particles, such as rotavirus VP6 inner capsid protein, to deliver the peptides at the site of infection (Bugli et al., 2014). Likewise, nanoparticles consisting of self-assembled amphiphilic peptides can be generated. Nanotechnology can offer a better delivery system for targeted therapy (Kovalainen et al., 2015). For example, the histatin 5-conjugated polymer-based AmB-delivery carrier system, which is redox-sensitive and pH-responsive, acts both as a synergistic molecule and as a targeting ligand against C. albicans (Park et al., 2017).
Fungal growth and consequent mycotoxin release in food and feed pose a serious risk to human health, which may lead to death in acute cases. Therefore, control and prevention of fungal pathogens and foodborne poisoning are among the main tasks of public health we face today. AFPs have been shown to possess both antifungal and anti-mycotoxin biosynthesis activities and thus meet the desired requirements to fight fungal contaminations. Several AFPs have been approved for applications in the food industry as preservatives, such as lactoferrin certified by the European Food Safety Authority (EFSA) since 2012 (EFSA Panel on Dietetic Products, Nutrition and Allergies (NDA), 2012). Mycotoxins commonly found in the food industry include aflatoxins (AFs), deoxynivalenol (DON), ochratoxin A (OTA), zearalenone (ZEA), fumonisins (FUM), patulin (PAT), and citrinin (CIT) (Marin et al., 2013). A few AFPs have been shown to inhibit mycotoxin biosynthesis. For example, peptide cyclo-L-leucyl-L-prolyl from Achromobacter xylosoxidans inhibits AF production by suppressing the expression of the AF biosynthesis regulatory gene aflR (Yan et al., 2004). Similarly, peptides cyclo-L-Val-L-Pro and cyclo-L-Ala-L-Pro both can inhibit AF biosynthesis of Aspergillus parasiticus and A. flavus by reducing the mRNA level of aflR and blocking the production of norsorolinic acid, a biosynthetic intermediate involved in an early step of AF biosynthetic pathway (Jermnak et al., 2013). In addition, iturin A from B. subtilis has been shown to inhibit OTA production by Aspergillus carbonarius (Jiang et al., 2020), and AgAFP peptide produced by Aspergillus giganteus has been shown to decrease DON production by the genus Fusarium (Barakat et al., 2010). Lipopeptides, such as surfactins and fengycins from Bacillus species, also have the capacity to inhibit mycotoxin synthesis (Martínez-Culebras et al., 2021).
Given the huge clinical and market needs, the development of novel AFPs and detailed studies on existing AFPs are necessary and urgent. Computer-aided drug design has become an integral part of AFP discovery and development efforts in the pharmaceutical and biotechnology fields. Sequencing has produced a large number of databases including genomic, transcriptomic, proteomic, and functional information. Progresses in the field of molecular biology, analysis of whole genomes, and high-throughput screening of natural and synthetic compounds have resulted in the identification and characterization of new targets, novel scaffolds, and leading structures for potential AFP candidates. Definitely, modern in silico molecular modeling techniques will make the screening and discovery of new AFPs more efficient and faster. At the same time, continuing efforts are required to develop and improve natural/modified AFPs, or their analogs/mimics, with high efficiency and a low risk of resistant pathogen emergence. In addition, more efforts should focus on combination therapy, where the synergy between AFPs and conventional antifungal drugs is the main objective. This approach can promote their effectiveness while reducing their toxicity to the host. Last but not least, the modes of action of AFPs still demand further investigation, especially in combination with animal models (Capilla et al., 2007; Hohl, 2014; Van Dijck et al., 2018).
Author contributions
YG: Conceptualization, Writing – original draft, Writing – review & editing. QX: Writing – original draft. JL: Writing – original draft. SZ: Conceptualization, Writing – review & editing.
Funding
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This study was supported by the Shanxi Scholarship Council of China (2024-078), the Basic Research Program of Shanxi Province (202303021222151), and the Initiative Research Fund of Kashi University (022023184).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Aerts, A. M., François, I. E., Cammue, B. P., and Thevissen, K. (2008). The mode of antifungal action of plant, insect and human defensins. CMLS 65, 2069–2079. doi: 10.1007/s00018-008-8035-0
Alcouloumre, M. S., Ghannoum, M. A., Ibrahim, A. S., Selsted, M. E., and Edwards, J. E. Jr. (1993). Fungicidal properties of defensin NP-1 and activity against Cryptococcus neoformans in vitro. Antimicrob. Agents Chemother. 37, 2628–2632. doi: 10.1128/AAC.37.12.2628
Amaral, A. C., Silva, O. N., Mundim, N. C., de Carvalho, M. J., Migliolo, L., Leite, J. R., et al. (2012). Predicting antimicrobial peptides from eukaryotic genomes: in silico strategies to develop antibiotics. Peptides 37, 301–308. doi: 10.1016/j.peptides.2012.07.021
An, M. Y., Gao, J., Zhao, X. F., and Wang, J. X. (2016). A new subfamily of penaeidin with an additional serine-rich region from kuruma shrimp (Marsupenaeus japonicus) contributes to antimicrobial and phagocytic activities. Dev. Comp. Immunol. 59, 186–198. doi: 10.1016/j.dci.2016.02.001
Asensio-Calavia, P., González-Acosta, S., Otazo-Pérez, A., López, M. R., Morales-delaNuez, A., and Pérez de la Lastra, J. M. (2023). Teleost Piscidins-in silico perspective of natural peptide antibiotics from marine sources. Antibiotics 12:855. doi: 10.3390/antibiotics12050855
Auchtung, T. A., Fofanova, T. Y., Stewart, C. J., Nash, A. K., Wong, M. C., Gesell, J. R., et al. (2018). Investigating colonization of the healthy adult gastrointestinal tract by fungi. mSphere 3:e00092–18. doi: 10.1128/mSphere.00092-18
Ayroza, G., Ferreira, I. L., Sayegh, R. S., Tashima, A. K., and da Silva Junior, P. I. (2012). Juruin: an antifungal peptide from the venom of the Amazonian pink toe spider, Avicularia juruensis, which contains the inhibitory cystine knot motif. Front. Microbiol. 3:324. doi: 10.3389/fmicb.2012.00324
Baldrian, P., Větrovský, T., Lepinay, C., and Kohout, P. (2021). High-throughput sequencing view on the magnitude of global fungal diversity. Fungal Divers. 19, 1–9.
Barakat, H., Spielvogel, A., Hassan, M., El-Desouky, A., El-Mansy, H., Rath, F., et al. (2010). The antifungal protein AFP from aspergillus giganteus prevents secondary growth of different fusarium species on barley. Appl. Microbiol. Biotechnol. 87, 617–624. doi: 10.1007/s00253-010-2508-4
Barbault, F., Landon, C., Guenneugues, M., Meyer, J. P., Schott, V., Dimarcq, J. L., et al. (2003). Solution structure of Alo-3: a new knottin-type antifungal peptide from the insect Acrocinus longimanus. Biochemistry 42, 14434–14442. doi: 10.1021/bi035400o
Bazinet, L., and Firdaous, L. (2013). Separation of bioactive peptides by membrane processes: technologies and devices. Recent Pat. Biotechnol. 7, 9–27. doi: 10.2174/1872208311307010003
Berrocal-Lobo, M., Segura, A., Moreno, M., López, G., García-Olmedo, F., and Molina, A. (2002). Snakin-2, an antimicrobial peptide from potato whose gene is locally induced by wounding and responds to pathogen infection. Plant Physiol. 128, 951–961. doi: 10.1104/pp.010685
Bondaryk, M., Staniszewska, M., Zielińska, P., and Urbańczyk-Lipkowska, Z. (2017). Natural antimicrobial peptides as inspiration for design of a new generation antifungal compounds. J. Fungi 3:46. doi: 10.3390/jof3030046
Bormann, C., Baier, D., Hörr, I., Raps, C., Berger, J., Jung, G., et al. (1999). Characterization of a novel, antifungal, chitin-binding protein from Streptomyces tendae Tü901 that interferes with growth polarity. J. Bacteriol. 181, 7421–7429. doi: 10.1128/JB.181.24.7421-7429.1999
Boukaew, S., Zhang, Z., Prasertsan, P., and Igarashi, Y. (2023). Antifungal and antiaflatoxigenic mechanism activity of freeze-dried culture filtrate of Streptomyces philanthi RL-1-178 on the two aflatoxigenic fungi and identification of its active component. J. Appl. Microbiol. 134:lxac091. doi: 10.1093/jambio/lxac091
Bowman, S. M., and Free, S. J. (2006). The structure and synthesis of the fungal cell wall. BioEssays 28, 799–808. doi: 10.1002/bies.20441
Brady, R., Woonton, B., Gee, M. L., and O’Connor, A. J. (2008). Hierarchical mesoporous silica materials for separation of functional food ingredients — a review. Innov. Food Sci. Emerg. Technol. 9, 243–248. doi: 10.1016/j.ifset.2007.10.002
Brown, G. D., Denning, D. W., Gow, N. A., Levitz, S. M., Netea, M. G., and White, T. C. (2012). Hidden killers: human fungal infections. Sci. Transl. Med. 4:165rv13. doi: 10.1126/scitranslmed.3004404
Bugli, F., Caprettini, V., Cacaci, M., Martini, C., Paroni Sterbini, F., Torelli, R., et al. (2014). Synthesis and characterization of different immunogenic viral nanoconstructs from rotavirus VP6 inner capsid protein. Int. J. Nanomedicine 9, 2727–2739. doi: 10.2147/IJN.S60014
Cabib, E., and Arroyo, J. (2013). How carbohydrates sculpt cells: chemical control of morphogenesis in the yeast cell wall. Nat. Rev. Microbiol. 11, 648–655. doi: 10.1038/nrmicro3090
Campbell-Platt, G., and Cook, P. E. (2008). Fungi in the production of foods and food ingredients. J. Appl. Microbiol. 67, 117s–131s. doi: 10.1111/j.1365-2672.1989.tb03776.x
Capilla, J., Clemons, K. V., and Stevens, D. A. (2007). Animal models: an important tool in mycology. Med. Mycol. 45, 657–684. doi: 10.1080/13693780701644140
Chahardoli, M., Fazeli, A., and Ghabooli, M. (2018). Recombinant production of bovine lactoferrin-derived antimicrobial peptide in tobacco hairy roots expression system. Plant Physiol. Biochem. 123, 414–421. doi: 10.1016/j.plaphy.2017.12.037
Cheng, R., Xu, Q., Hu, F., Li, H., Yang, B., Duan, Z., et al. (2021). Antifungal activity of MAF-1A peptide against Candida albicans. Int. Microbiol. Off. J. Spanish Soc. Microbiol. 24, 233–242. doi: 10.1007/s10123-021-00159-z
Chiu, T., Poucet, T., and Li, Y. (2022). The potential of plant proteins as antifungal agents for agricultural applications. Synth. Syst. Biotechnol. 7, 1075–1083. doi: 10.1016/j.synbio.2022.06.009
Choi, H., Hwang, J. S., and Lee, D. G. (2013). Antifungal effect and pore-forming action of lactoferricin B like peptide derived from centipede Scolopendra subspinipes mutilans. Biochim. Biophys. Acta 1828, 2745–2750. doi: 10.1016/j.bbamem.2013.07.021
Ciociola, T., Giovati, L., Conti, S., Magliani, W., Santinoli, C., and Polonelli, L. (2016). Natural and synthetic peptides with antifungal activity. Future Med. Chem. 8, 1413–1433. doi: 10.4155/fmc-2016-0035
Cole, A. M., Weis, P., and Diamond, G. (1997). Isolation and characterization of pleurocidin, an antimicrobial peptide in the skin secretions of winter flounder. J. Biol. Chem. 272, 12008–12013. doi: 10.1074/jbc.272.18.12008
Cuthbertson, B. J., Shepard, E. F., Chapman, R. W., and Gross, P. S. (2002). Diversity of the penaeidin antimicrobial peptides in two shrimp species. Immunogenetics 54, 442–445. doi: 10.1007/s00251-002-0487-z
Dash, R., and Bhattacharjya, S. (2021). Thanatin: An emerging host defense antimicrobial peptide with multiple modes of action. Int. J. Mol. Sci. 22:1522. doi: 10.3390/ijms22041522
D'Auria, F. D., Casciaro, B., De Angelis, M., Marcocci, M. E., Palamara, A. T., Nencioni, L., et al. (2022). Antifungal activity of the frog skin peptide Temporin G and its effect on Candida albicans virulence factors. Int. J. Mol. Sci. 23:6345. doi: 10.3390/ijms23116345
De Lucca, A. J. (2000). Antifungal peptides: potential candidates for the treatment of fungal infections. Expert Opin. Investig. Drugs 9, 273–299. doi: 10.1517/13543784.9.2.273
De Lucca, A. J., and Walsh, T. J. (1999). Antifungal peptides: novel therapeutic compounds against emerging pathogens. Antimicrob. Agents Chemother. 43, 1–11. doi: 10.1128/AAC.43.1.1
Destoumieux, D., Munoz, M., Bulet, P., and Bachère, E. (2000). Penaeidins, a family of antimicrobial peptides from penaeid shrimp (Crustacea, Decapoda). Cell. Mol. Life Sci. 57, 1260–1271. doi: 10.1007/pl00000764
Drago-Serrano, M. E., Campos-Rodriguez, R., Carrero, J. C., and de la Garza, M. (2018). Lactoferrin and peptide-derivatives: antimicrobial agents with potential use in nonspecific immunity modulation. Curr. Pharm. Des. 24, 1067–1078. doi: 10.2174/1381612824666180327155929
Dubos, R. J. (1939). Studies on a bactericidal agent extracted from a soil bacillus: II. Protective effect of the bactericidal agent against experimental pneumococcus infections in mice. J. Exp. Med. 70, 11–17. doi: 10.1084/jem.70.1.11
Duncan, V. M. S., and O’Neil, D. A. (2013). Commercialization of antifungal peptides. Fungal Biol. Rev. 26, 156–165. doi: 10.1016/j.fbr.2012.11.001
EFSA Panel on Dietetic Products, Nutrition and Allergies (NDA) (2012). Scientific opinion on bovine lactoferrin. EFSA J. 10:2811. doi: 10.2903/j.efsa.2012.2811
Emri, T., Majoros, L., Tóth, V., and Pócsi, I. (2013). Echinocandins: production and applications. Appl. Microbiol. Biotechnol. 97, 3267–3284. doi: 10.1007/s00253-013-4761-9
Endo, M., Takesako, K., Kato, I., and Yamaguchi, H. (1997). Fungicidal action of aureobasidin a, a cyclic depsipeptide antifungal antibiotic, against Saccharomyces cerevisiae. Antimicrob. Agents Chemother. 41, 672–676. doi: 10.1128/AAC.41.3.672
Faruck, M. O., Yusof, F., and Chowdhury, S. (2016). An overview of antifungal peptides derived from insect. Peptides 80, 80–88. doi: 10.1016/j.peptides.2015.06.001
Fehlbaum, P., Bulet, P., Chernysh, S., Briand, J. P., Roussel, J. P., Letellier, L., et al. (1996). Structure-activity analysis of thanatin, a 21-residue inducible insect defense peptide with sequence homology to frog skin antimicrobial peptides. Proc. Natl. Acad. Sci. USA 93, 1221–1225. doi: 10.1073/pnas.93.3.1221
Fehlbaum, P., Bulet, P., Michaut, L., Lagueux, M., Broekaert, W. F., Hetru, C., et al. (1994). Insect immunity. Septic injury of Drosophila induces the synthesis of a potent antifungal peptide with sequence homology to plant antifungal peptides. J. Biol. Chem. 269, 33159–33163. doi: 10.1016/S0021-9258(20)30111-3
Fennell, J. F., Shipman, W. H., and Cole, L. J. (1967). Antibacterial action of a bee venom fraction (melittin) against a penicillin-resistant staphylococcus and other microorganisms. USNRDL-TR-67-101. Res. Dev. Tech. Rep., 1–13. doi: 10.21236/ad0658324
Fernandes, K. E., and Carter, D. A. (2017). The antifungal activity of lactoferrin and its derived peptides: mechanisms of action and synergy with drugs against fungal pathogens. Front. Microbiol. 8:2. doi: 10.3389/fmicb.2017.00002
Fernandes, K. E., Weeks, K., and Carter, D. A. (2020). Lactoferrin is broadly active against yeasts and highly synergistic with amphotericin B. Antimicrob. Agents Chemother. 64, e02284–e02219. doi: 10.1128/AAC.02284-19
Fernández de Ullivarri, M., Arbulu, S., Garcia-Gutierrez, E., and Cotter, P. D. (2020). Antifungal peptides as therapeutic agents. Front. Cell. Infect. Microbiol. 10:105. doi: 10.3389/fcimb.2020.00105
Fisher, M. C., Hawkins, N. J., Sanglard, D., and Gurr, S. J. (2018). Worldwide emergence of resistance to antifungal drugs challenges human health and food security. Science 360, 739–742. doi: 10.1126/science.aap7999
Fleet, G. H. (1991). “Cell walls” in The yeasts. eds. A. H. Rose and J. S. Harrison. 2nd ed (New York, NY: Academic Press), 199–277.
Gamaletsou, M. N., Walsh, T. J., and Sipsas, N. V. (2018). Invasive fungal infections in patients with hematological malignancies: emergence of resistant pathogens and new antifungal therapies. Turk. J. Haematol. 35, 1–11. doi: 10.4274/tjh.2018.0007
Games, P. D., Dos Santos, I. S., Mello, E. O., Diz, M. S., Carvalho, A. O., and de Souza-Filho, G. A. (2008). Isolation, characterization and cloning of a cDNA encoding a new antifungal defensin from Phaseolus vulgaris L. seeds. Peptides 29, 2090–2100. doi: 10.1016/j.peptides.2008.08.008
Gao, A.-G., Hakimi, S. M., Mittanck, C. A., Wu, Y., Woerner, B. M., Stark, D. M., et al. (2000). Fungal pathogen protection in potato by expression of a plant defensin peptide. Nat. Biotechnol. 18, 1307–1310. doi: 10.1038/82436
Gennaro, R., and Zanetti, M. (2000). Structural features and biological activities of the cathelicidin-derived antimicrobial peptides. Biopolymers 55, 31–49. doi: 10.1002/1097-0282(2000)55:1<31::AID-BIP40>3.0.CO;2-9
Gong, Y., Li, H., Wu, F., Li, Y., and Zhang, S. (2022). Fungicidal activity of AP10W, a short peptide derived from AP-2 complex subunit mu-a, in vitro and in vivo. Biomolecules 12:965. doi: 10.3390/biom12070965
Gupta, A., Gupta, R., and Singh, R. L. (2017). Microbes and environment. Principles and applications of environmental biotechnology for a sustainable future. Cham: Springer, 43–84.
Hagen, S., Marx, F., Ram, A. F., and Meyer, V. (2007). The antifungal protein AFP from aspergillus giganteus inhibits chitin synthesis in sensitive fungi. Appl. Environ. Microbiol. 73, 2128–2134. doi: 10.1128/AEM.02497-06
Hancock, R. E. (2000). Cationic antimicrobial peptides: towards clinical applications. Expert Opin. Investig. Drugs 9, 1723–1729. doi: 10.1517/13543784.9.8.1723
Hancock, R. E., and Chapple, D. S. (1999). Peptide antibiotics. Antimicrob. Agents Chemother. 43, 1317–1323. doi: 10.1128/AAC.43.6.1317
Haney, E. F., Straus, S. K., and Hancock, R. E. W. (2019). Reassessing the host defense peptide landscape. Front. Chem. 7:43. doi: 10.3389/fchem.2019.00043
Helmerhorst, E. J., Troxler, R. F., and Oppenheim, F. G. (2001). The human salivary peptide histatin 5 exerts its antifungal activity through the formation of reactive oxygen species. Proc. Natl. Acad. Sci. USA 98, 14637–14642. doi: 10.1073/pnas.141366998
Hohl, T. M. (2014). Overview of vertebrate animal models of fungal infection. J. Immunol. Methods 410, 100–112. doi: 10.1016/j.jim.2014.03.022
Holaskova, E., Galuszka, P., Frebort, I., and Oz, M. T. (2015). Antimicrobial peptide production and plant-based expression systems for medical and agricultural biotechnology. Biotechnol. Adv. 33, 1005–1023. doi: 10.1016/j.biotechadv.2015.03.007
Hollmann, A., Martinez, M., Maturana, P., Semorile, L. C., and Maffia, P. C. (2018). Antimicrobial peptides: interaction with model and biological membranes and synergism with chemical antibiotics. Front. Chem. 6:204. doi: 10.3389/fchem.2018.00204
Huber, A., Hajdu, D., Bratschun-Khan, D., Gáspári, Z., Varbanov, M., Philippot, S., et al. (2018). New antimicrobial potential and structural properties of PAFB: a cationic, cysteine-rich protein from Penicillium chrysogenum Q176. Sci. Rep. 8:1751. doi: 10.1038/s41598-018-20002-2
Huffnagle, G. B., and Noverr, M. C. (2013). The emerging world of the fungal microbiome. Trends Microbiol. 21, 334–341. doi: 10.1016/j.tim.2013.04.002
Hwang, J. S., Lee, J., Hwang, B., Nam, S. H., Yun, E. Y., Kim, S. R., et al. (2010). Isolation and characterization of Psacotheasin, a novel Knottin-type antimicrobial peptide, from Psacothea hilaris. J. Microbiol. Biotechnol. 20, 708–711. doi: 10.4014/jmb.1002.02003
Hwang, J. S., Lee, J., Kim, Y. J., Bang, H. S., Yun, E. Y., Kim, S. R., et al. (2009). Isolation and characterization of a Defensin-like peptide (Coprisin) from the dung beetle, Copris tripartitus. Int. J. Pept. 2009:136284. doi: 10.1155/2009/136284
Jermnak, U., Chinaphuti, A., Poapolathep, A., Kawai, R., Nagasawa, H., and Sakuda, S. (2013). Prevention of aflatoxin contamination by a soil bacterium of Stenotrophomonas sp. that produces aflatoxin production inhibitors. Microbiology 159, 902–912. doi: 10.1099/mic.0.065813-0
Jiang, C., Li, Z., Shi, Y., Guo, D., Pang, B., Chen, X., et al. (2020). Bacillus subtilis inhibits aspergillus Carbonarius by producing Iturin a, which disturbs the transport, energy metabolism, and osmotic pressure of fungal cells as revealed by transcriptomics analysis. Int. J. Food Microbiol. 330:108783. doi: 10.1016/j.ijfoodmicro.2020.108783
Joly, S., Maze, C., McCray, P. B. Jr., and Guthmiller, J. M. (2004). Human beta-defensins 2 and 3 demonstrate strain-selective activity against oral microorganisms. J. Clin. Microbiol. 42, 1024–1029. doi: 10.1128/JCM.42.3.1024-1029.2004
Kaiserer, L., Oberparleiter, C., Weiler-Görz, R., Burgstaller, W., Leiter, E., and Marx, F. (2003). Characterization of the Penicillium chrysogenum antifungal protein PAF. Arch. Microbiol. 180, 204–210. doi: 10.1007/s00203-003-0578-8
Kang, Y., Carlson, R., Tharpe, W., and Schell, M. A. (1998). Characterization of genes involved in biosynthesis of a novel antibiotic from Burkholderia cepacia BC11 and their role in biological control of Rhizoctonia solani. Appl. Environ. Microbiol. 64, 3939–3947. doi: 10.1128/AEM.64.10.3939-3947.1998
Kang, H. K., Seo, C. H., and Park, Y. (2015). Marine peptides and their anti-infective activities. Mar. Drugs 13, 618–654. doi: 10.3390/md13010618
Kapitan, M., Niemiec, M. J., Steimle, A., Frick, J. S., and Jacobsen, I. D. (2018). Fungi as part of the microbiota and interactions with intestinal bacteria. Curr. Top. Microbiol. Immunol. 422, 265–301. doi: 10.1007/82_2018_117
Karasuda, S., Tanaka, S., Kajihara, H., Yamamoto, Y., and Koga, D. (2003). Plant chitinase as a possible biocontrol agent for use instead of chemical fungicides. Biosci. Biotechnol. Biochem. 67, 221–224. doi: 10.1271/bbb.67.221
Kathiravan, M. K., Salake, A. B., Chothe, A. S., Dudhe, P. B., Watode, R. P., Mukta, M. S., et al. (2012). The biology and chemistry of antifungal agents: a review. Bioorg. Med. Chem. 20, 5678–5698. doi: 10.1016/j.bmc.2012.04.045
Kavanagh, K., and Dowd, S. (2004). Histatins: antimicrobial peptides with therapeutic potential. J. Pharm. Pharmacol. 56, 285–289. doi: 10.1211/0022357022971
Kim, S. R., Hong, M. Y., Park, S. W., Choi, K. H., Yun, E. Y., Goo, T. W., et al. (2010). Characterization and cDNA cloning of a cecropin-like antimicrobial peptide, papiliocin, from the swallowtail butterfly, Papilio xuthus. Mol. Cells. 29, 419–424. doi: 10.1007/s10059-010-0050-y
Kim, S., Hwang, J. S., and Lee, D. G. (2020). Lactoferricin B like peptide triggers mitochondrial disruption-mediated apoptosis by inhibiting respiration under nitric oxide accumulation in Candida albicans. IUBMB Life 72, 1515–1527. doi: 10.1002/iub.2284
Klich, M. A., Lax, A. R., and Bland, J. M. (1991). Inhibition of some mycotoxigenic fungi by iturin a, a peptidolipid produced by Bacillus subtilis. Mycopathologia 116, 77–80. doi: 10.1007/BF00436368
Koo, H. B., and Seo, J. (2019). Antimicrobial peptides under clinical investigation. J. Pept. Sci. 111:e24122. doi: 10.1002/pep2.24122
Kovalainen, M., Monkare, J., Riikonen, J., et al. (2015). Novel delivery systems for improving the clinical use of peptides. Pharmacol. Rev. 67, 541–561. doi: 10.1124/pr.113.008367
Lamberty, M., Caille, A., Landon, C., Tassin-Moindrot, S., Hetru, C., Bulet, P., et al. (2001a). Solution structures of the antifungal heliomicin and a selected variant with both antibacterial and antifungal activities. Biochemistry 40, 11995–12003. doi: 10.1021/bi0103563
Lamberty, M., Zachary, D., Lanot, R., Bordereau, C., Robert, A., Hoffmann, J. A., et al. (2001b). Insect immunity. Constitutive expression of a cysteine-rich antifungal and a linear antibacterial peptide in a termite insect. J. Biol. Chem. 276, 4085–4092. doi: 10.1074/jbc.M002998200
Landon, C., Barbault, F., Legrain, M., Menin, L., Guenneugues, M., Schott, V., et al. (2004). Lead optimization of antifungal peptides with 3D NMR structures analysis. Protein Sci. 13, 703–713. doi: 10.1110/ps.03404404
Le, C. F., Fang, C. M., and Sekaran, S. D. (2017). Intracellular targeting mechanisms by antimicrobial peptides. Antimicrob. Agents Chemother. 61, e02340–e02316. doi: 10.1128/AAC.02340-16
Lee, J., Hwang, J. S., Hwang, I. S., Cho, J., Lee, E., Kim, Y., et al. (2012). Coprisin-induced antifungal effects in Candida albicans correlate with apoptotic mechanisms. Free Radic. Biol. Med. 52, 2302–2311. doi: 10.1016/j.freeradbiomed.2012.03.012
Lee, D. G., Kim, H. K., Kim, S. A., Park, Y., Park, S. C., Jang, S. H., et al. (2003). Fungicidal effect of indolicidin and its interaction with phospholipid membranes. Biochem. Biophys. Res. Commun. 305, 305–310. doi: 10.1016/s0006-291x(03)00755-1
Lehrer, R. I., Ganz, T., Szklarek, D., and Selsted, M. E. (1988). Modulation of the in vitro candidacidal activity of human neutrophil defensins by target cell metabolism and divalent cations. J. Clin. Invest. 81, 1829–1835. doi: 10.1172/JCI113527
Lenardon, M. D., Munro, C. A., and Gow, N. A. (2010). Chitin synthesis and fungal pathogenesis. Curr. Opin. Microbiol. 13, 416–423. doi: 10.1016/j.mib.2010.05.002
Lestrade, P. P. A., Meis, J. F., Melchers, W. J. G., and Verweij, P. E. (2019). Triazole resistance in Aspergillus fumigatus: recent insights and challenges for patient management. Clin. Microbiol. Infect. 25, 799–806. doi: 10.1016/j.cmi.2018.11.027
Li, Y. (2011). Recombinant production of antimicrobial peptides in Escherichia coli: a review. Protein Expr. Purif. 80, 260–267. doi: 10.1016/j.pep.2011.08.001
Li, R., Chen, C., Zhang, B., Jing, H., Wang, Z., Wu, C., et al. (2019). The chromogranin A-derived antifungal peptide CGA-N9 induces apoptosis in Candida tropicalis. Biochem. J. 476, 3069–3080. doi: 10.1042/BCJ20190483
Liu, Z., Yuan, K., Zhang, R., Ren, X., Liu, X., Zhao, S., et al. (2016). Cloning and purification of the first termicin-like peptide from the cockroach Eupolyphaga sinensis. J. Venom Anim. Toxins Incl. Trop Dis. 22:5. doi: 10.1186/s40409-016-0058-7
Lobo, D. S., Pereira, I. B., Fragel-Madeira, L., Medeiros, L. N., Cabral, L. M., Faria, J., et al. (2007). Antifungal Pisum sativum defensin 1 interacts with Neurospora crassa cyclin F related to the cell cycle. Biochemistry 46, 987–996. doi: 10.1021/bi061441j
Lohner, K., and Prenner, E. J. (1999). Differential scanning calorimetry and X-ray diffraction studies of the specificity of the interaction of antimicrobial peptides with membrane-mimetic systems. Biochim. Biophys. Acta 1462, 141–156. doi: 10.1016/s0005-2736(99)00204-7
Lombardi, L., Maisetta, G., Batoni, G., and Tavanti, A. (2015). Insights into the antimicrobial properties of hepcidins: advantages and drawbacks as potential therapeutic agents. Molecules 20, 6319–6341. doi: 10.3390/molecules20046319
López-Abarrategui, C., Alba, A., Silva, O. N., Reyes-Acosta, O., Vasconcelos, I. M., Oliveira, J. T. A., et al. (2012). Functional characterization of a synthetic hydrophilic antifungal peptide derived from the marine snail Cenchritis muricatus. Biochimie 94, 968–974. doi: 10.1016/j.biochi.2011.12.016
López-Meza, J., Ochoa-Zarzosa, A., Aguilar, J., and Loeza-Lara, P. (2011). “Antimicrobial peptides: diversity and perspectives for their biomedical application,” in Biomedical Engineering, Trends, Research and Technologies. London: IntechOpen.
Lücking, R., Aime, M. C., Robbertse, B., Miller, A. N., Aoki, T., Ariyawansa, H. A., et al. (2021). Fungal taxonomy and sequence-based nomenclature. Nat. Microbiol. 6, 540–548. doi: 10.1038/s41564-021-00888-x
Lupetti, A., Paulusma-Annema, A., Welling, M. M., Dogterom-Ballering, H., Brouwer, C. P. J. M., Senesi, S., et al. (2003). Synergistic activity of the N-terminal peptide of human lactoferrin and fluconazole against Candida species. Antimicrob. Agents Chemother. 47, 262–267. doi: 10.1128/aac.47.1.262-267.2003
Marcocci, M. E., Amatore, D., Villa, S., Casciaro, B., Aimola, P., Franci, G., et al. (2018). The amphibian antimicrobial peptide Temporin B inhibits in vitro herpes simplex virus 1 infection. Antimicrob. Agents Chemother. 62, e02367–e02317. doi: 10.1128/AAC.02367-17
Marin, S., Ramos, A. J., Cano-Sancho, G., and Sanchis, V. (2013). Mycotoxins: occurrence, toxicology, and exposure assessment. Food Chem. Toxicol. 60, 218–237. doi: 10.1016/j.fct.2013.07.047
Martínez-Culebras, P. V., Gandía, M., Garrigues, S., Marcos, J. F., and Manzanares, P. (2021). Antifungal peptides and proteins to control toxigenic Fungi and mycotoxin biosynthesis. Int. J. Mol. Sci. 22:13261. doi: 10.3390/ijms222413261
Matejuk, A., Leng, Q., Begum, M. D., Woodle, M. C., Scaria, P., Chou, S. T., et al. (2010). Peptide-based antifungal therapies against emerging infections. Drugs Future 35:197. doi: 10.1358/dof.2010.35.3.1452077
McCarthy, P. J., Troke, P. F., and Gull, K. (1985). Mechanism of action of nikkomycin and the peptide transport system of Candida albicans. J. Gen. Microbiol. 131, 775–780. doi: 10.1099/00221287-131-4-775
Mello, E. O., Ribeiro, S. F., Carvalho, A. O., Santos, I. S., Da Cunha, M., Santa-Catarina, C., et al. (2011). Antifungal activity of PvD1 defensin involves plasma membrane permeabilization, inhibition of medium acidification, and induction of ROS in fungi cells. Curr. Microbiol. 62, 1209–1217. doi: 10.1007/s00284-010-9847-3
Memariani, H., and Memariani, M. (2020). Anti-fungal properties and mechanisms of melittin. Appl. Microbiol. Biotechnol. 104, 6513–6526. doi: 10.1007/s00253-020-10701-0
Memariani, M., and Memariani, H. (2023). Antifungal properties of cathelicidin LL-37: current knowledge and future research directions. World J. Microbiol. Biotechnol. 40:34. doi: 10.1007/s11274-023-03852-5
Mendes, M. A., de Souza, B. M., Marques, M. R., and Palma, M. S. (2004). Structural and biological characterization of two novel peptides from the venom of the neotropical social wasp Agelaia pallipes pallipes. Toxicon 44, 67–74. doi: 10.1016/j.toxicon.2004.04.009
Miyata, T., Tokunaga, F., Yoneya, T., Yoshikawa, K., Iwanaga, S., Niwa, M., et al. (1989). Antimicrobial peptides, isolated from horseshoe crab hemocytes, tachyplesin II, and polyphemusins I and II: chemical structures and biological activity. J. Biochem. 106, 663–668. doi: 10.1093/oxfordjournals.jbchem.a122913
Money, N. P. (2016). Fungi and biotechnology. J. Fungi 2016, 401–424. doi: 10.1016/B978-0-12-382034-1.00012-8
Monincová, L., Slaninová, J., Fučík, V., Hovorka, O., Voburka, Z., Bednárová, L., et al. (2012). Lasiocepsin, a novel cyclic antimicrobial peptide from the venom of eusocial bee Lasioglossum laticeps (Hymenoptera: Halictidae). Amino Acids 43, 751–761. doi: 10.1007/s00726-011-1125-6
Moreno-Expósito, L., Illescas-Montes, R., Melguizo-Rodríguez, L., Ruiz, C., Ramos-Torrecillas, J., and de Luna-Bertos, E. (2018). Multifunctional capacity and therapeutic potential of lactoferrin. Life Sci. 195, 61–64. doi: 10.1016/j.lfs.2018.01.002
Muhialdin, B. J., Algboory, H. L., Kadum, H., Mohammed, N. K., Saari, N., Hassan, Z., et al. (2020). Antifungal activity determination for the peptides generated by Lactobacillus plantarum TE10 against aspergillus flavus in maize seeds. Food Control 109:106898. doi: 10.1016/j.foodcont.2019.106898
Mukherjee, D., Singh, S., Kumar, M., Kumar, V., Datta, S., and Dhanjal, D. S. (2018). “Fungal biotechnology: role and aspects” in Fungi and their role in sustainable development: Current perspectives. eds. P. Gehlot and J. Singh (Singapore: Springer Singapore), 91–103.
Nagiec, M. M., Nagiec, E. E., Baltisberger, J. A., Wells, G. B., Lester, R. L., and Dickson, R. C. (1997). Sphingolipid synthesis as a target for antifungal drugs. Complementation of the inositol phosphorylceramide synthase defect in a mutant strain of Saccharomyces cerevisiae by the AUR1 gene. J. Biol. Chem. 272, 9809–9817. doi: 10.1074/jbc.272.15.9809
Nawrocki, K. L., Crispell, E. K., and McBride, S. M. (2014). Antimicrobial peptide resistance mechanisms of gram-positive Bacteria. Antibiotics 3, 461–492. doi: 10.3390/antibiotics3040461
Nolde, S. B., Vassilevski, A. A., Rogozhin, E. A., Barinov, N. A., Balashova, T. A., Samsonova, O. V., et al. (2011). Disulfide-stabilized helical hairpin structure and activity of a novel antifungal peptide EcAMP1 from seeds of barnyard grass (Echinochloa crus-galli). J. Biol. Chem. 286, 25145–25153. doi: 10.1074/jbc.M110.200378
Notomista, E., Falanga, A., Fusco, S., Pirone, L., Zanfardino, A., Galdiero, S., et al. (2015). The identification of a novel Sulfolobus islandicus CAMP-like peptide points to archaeal microorganisms as cell factories for the production of antimicrobial molecules. Microb. Cell Factories 14:126. doi: 10.1186/s12934-015-0302-9
Odintsova, T. I., Vassilevski, A. A., Slavokhotova, A. A., Musolyamov, A. K., Finkina, E. I., Khadeeva, N. V., et al. (2009). A novel antifungal hevein-type peptide from Triticum kiharae seeds with a unique 10-cysteine motif. FEBS J. 276, 4266–4275. doi: 10.1111/j.1742-4658.2009.07135.x
Oppenheim, F. G., Xu, T., McMillian, F. M., Levitz, S. M., Diamond, R. D., Offner, G. D., et al. (1988). Histatins, a novel family of histidine-rich proteins in human parotid secretion. Isolation, characterization, primary structure, and fungistatic effects on Candida albicans. J. Biol. Chem. 263, 7472–7477. doi: 10.1016/S0021-9258(18)68522-9
Osaki, T., Omotezako, M., Nagayama, R., Hirata, M., Iwanaga, S., Kasahara, J., et al. (1999). Horseshoe crab hemocyte-derived antimicrobial polypeptides, tachystatins, with sequence similarity to spider neurotoxins. J. Biol. Chem. 274, 26172–26178. doi: 10.1074/jbc.274.37.26172
Osborn, R. W., de Samblanx, G. W., Thevissen, K., Goderis, I., Torrekens, S., van Leuven, F., et al. (1995). Isolation and characterisation of plant defensins from seeds of Asteraceae, Fabaceae, Hippocastanaceae and Saxifragaceae. FEBS Lett. 368, 257–262. doi: 10.1016/0014-5793(95)00666-w
Ovchinnikova, T. V., Aleshina, G. M., Balandin, S. V., Krasnosdembskaya, A. D., Markelov, M. L., Frolova, E. I., et al. (2004). Purification and primary structure of two isoforms of arenicin, a novel antimicrobial peptide from marine polychaeta Arenicola marina. FEBS Lett. 577, 209–214. doi: 10.1016/j.febslet.2004.10.012
Padovan, L., Segat, L., Pontillo, A., Antcheva, N., Tossi, A., and Crovella, S. (2010). Histatins in non-human primates: gene variations and functional effects. Protein Pept. Lett. 17, 909–918. doi: 10.2174/092986610791306715
Park, S.-C., Kim, Y.-M., Lee, J.-K., Kim, N.-H., Kim, E.-J., Heo, H., et al. (2017). Targeting and synergistic action of an antifungal peptide in an antibiotic drug-delivery system. J. Control. Release 256, 46–55. doi: 10.1016/j.jconrel.2017.04.023
Park, C. B., Lee, J. H., Park, I. Y., Kim, M. S., and Kim, S. C. (1997). A novel antimicrobial peptide from the loach, Misgurnus anguillicaudatus. FEBS Lett. 411, 173–178. doi: 10.1016/s0014-5793(97)00684-4
Pei, J., Feng, Z., Ren, T., Sun, H., Han, H., Jin, W., et al. (2018). Purification, characterization and application of a novel antimicrobial peptide from Andrias davidianus blood. Lett. Appl. Microbiol. 66, 38–43. doi: 10.1111/lam.12823
Pesic, A., Baumann, H. I., Kleinschmidt, K., Ensle, P., Wiese, J., Süssmuth, R. D., et al. (2013). Champacyclin, a new cyclic octapeptide from Streptomyces strain C42 isolated from the Baltic Sea. Mar. Drugs 11, 4834–4857. doi: 10.3390/md11124834
Pfaller, M. A., Huband, M. D., Flamm, R. K., Bien, P. A., and Castanheira, M. (2019). In vitro activity of APX001A (Manogepix) and comparator agents against 1,706 fungal isolates collected during an international surveillance program in 2017. Antimicrob. Agents Chemother. 63, e00840–e00819. doi: 10.1128/AAC.00840-19
Phukhamsakda, C., Nilsson, R. H., Bhunjun, C. S., de Farias, A. R. G., Sun, Y. R., Wijesinghe, S. N., et al. (2022). The numbers of fungi: contributions from traditional taxonomic studies and challenges of metabarcoding. Fungal Divers. 114, 327–386. doi: 10.1007/s13225-022-00502-3
Pólvora, T. L. S., Nobre, Á. V. V., Tirapelli, C., Taba, M. Jr., Macedo, L. D., Santana, R. C., et al. (2018). Relationship between human immunodeficiency virus (HIV-1) infection and chronic periodontitis. Expert Rev. Clin. Immunol. 14, 315–327. doi: 10.1080/1744666X.2018.1459571
Porto, W. F., Silva, O. N., and Franco, O. L. (2012). “Prediction and rational design of antimicrobial peptides,” in Protein Structure Eshel Faraggi (New York, NY: IntechOpen).
Rakers, S., Niklasson, L., Steinhagen, D., Kruse, C., Schauber, J., Sundell, K., et al. (2013). Antimicrobial peptides (AMPs) from fish epidermis: perspectives for investigative dermatology. J. Invest. Dermatol. 133, 1140–1149. doi: 10.1038/jid.2012.503
Ramamoorthy, V., Zhao, X., Snyder, A. K., Xu, J. R., and Shah, D. M. (2007). Two mitogen-activated protein kinase signalling cascades mediate basal resistance to antifungal plant defensins in fusarium graminearum. Cell. Microbiol. 9, 1491–1506. doi: 10.1111/j.1462-5822.2006.00887.x
Rascón-Cruz, Q., Espinoza-Sánchez, E. A., Siqueiros-Cendón, T. S., Nakamura-Bencomo, S. I., Arévalo-Gallegos, S., and Iglesias-Figueroa, B. F. (2021). Lactoferrin: a glycoprotein involved in immunomodulation, anticancer, and antimicrobial processes. Molecules 26:205. doi: 10.3390/molecules26010205
Rodríguez-Martín, A., Acosta, R., Liddell, S., Núñez, F., Benito, M. J., and Asensio, M. A. (2010). Characterization of the novel antifungal protein PgAFP and the encoding gene of Penicillium chrysogenum. Peptides 31, 541–547. doi: 10.1016/j.peptides.2009.11.002
Rogozhin, E. A., Slezina, M. P., Slavokhotova, A. A., Istomina, E. A., Korostyleva, T. V., Smirnov, A. N., et al. (2015). A novel antifungal peptide from leaves of the weed Stellaria media L. Biochimie 116, 125–132. doi: 10.1016/j.biochi.2015.07.014
Roscetto, E., Bellavita, R., Paolillo, R., Merlino, F., Molfetta, N., Grieco, P., et al. (2021). Antimicrobial activity of a Lipidated Temporin L analogue against Carbapenemase-producing Klebsiella pneumoniae clinical isolates. Antibiotics 10:1312. doi: 10.3390/antibiotics10111312
Roscetto, E., Contursi, P., Vollaro, A., Fusco, S., Notomista, E., and Catania, M. R. (2018). Antifungal and anti-biofilm activity of the first cryptic antimicrobial peptide from an archaeal protein against Candida spp. clinical isolates. Sci. Rep. 8:17570. doi: 10.1038/s41598-018-35530-0
Ryder, K., Bekhit, A. E.-D., McConnell, M., and Carne, A. (2016). Towards generation of bioactive peptides from meat industry waste proteins: generation of peptides using commercial microbial proteases. Food Chem. 208, 42–50. doi: 10.1016/j.foodchem.2016.03.121
Saito, T., Kawabata, S., Shigenaga, T., Takayenoki, Y., Cho, J., Nakajima, H., et al. (1995). A novel big defensin identified in horseshoe crab hemocytes: isolation, amino acid sequence, and antibacterial activity. J. Biochem. 117, 1131–1137. doi: 10.1093/oxfordjournals.jbchem.a124818
Scarsini, M., Tomasinsig, L., Arzese, A., D'Este, F., Oro, D., and Skerlavaj, B. (2015). Antifungal activity of cathelicidin peptides against planktonic and biofilm cultures of Candida species isolated from vaginal infections. Peptides 71, 211–221. doi: 10.1016/j.peptides.2015.07.023
Segura, A., Moreno, M., Madueño, F., Molina, A., and García-Olmedo, F. (1999). Snakin-1, a peptide from potato that is active against plant pathogens. Mol. Plant-Microbe Interact. 12, 16–23. doi: 10.1094/MPMI.1999.12.1.16
Selsted, M. E., Harwig, S. S., Ganz, T., Schilling, J. W., and Lehrer, R. I. (1985). Primary structures of three human neutrophil defensins. J. Clin. Invest. 76, 1436–1439. doi: 10.1172/JCI112121
Sengupta, J., Saha, S., Khetan, A., Sarkar, S. K., and Mandal, S. M. (2012). Effects of lactoferricin B against keratitis-associated fungal biofilms. J. Infect. Chemother. 18, 698–703. doi: 10.1007/s10156-012-0398-3
Sewczyk, T., Hoog Antink, M., Maas, M., Kroll, S., and Beutel, S. (2018). Flow rate dependent continuous hydrolysis of protein isolates. AMB Express 8:18. doi: 10.1186/s13568-018-0548-9
Shams, M. V., Nazarian-Firouzabadi, F., Ismaili, A., and Shirzadian-Khorramabad, R. (2019). Production of a recombinant dermaseptin peptide in nicotiana tabacum hairy roots with enhanced antimicrobial activity. Mol. Biotechnol. 61, 241–252. doi: 10.1007/s12033-019-00153-x
Sharma, P., Chaudhary, M., Khanna, G., Rishi, P., and Kaur, I. P. (2021). Envisaging antifungal potential of Histatin 5: a physiological salivary peptide. J. Fungi 7:1070. doi: 10.3390/jof7121070
Shishido, T. K., Humisto, A., Jokela, J., Liu, L., Wahlsten, M., Tamrakar, A., et al. (2015). Antifungal compounds from cyanobacteria. Mar. Drugs 13, 2124–2140. doi: 10.3390/md13042124
Shrestha, C. L., Oña, I., Muthukrishnan, S., and Mew, T. W. (2007). Chitinase levels in rice cultivars correlate with resistance to the sheath blight pathogen Rhizoctonia solani. Eur. J. Plant Pathol. 120, 69–77. doi: 10.1007/s10658-007-9199-4
Sibel Akalin, A. (2014). Dairy-derived antimicrobial peptides: action mechanisms, pharmaceutical uses and production proposals. Trends Food Sci. Technol. 36, 79–95. doi: 10.1016/j.tifs.2014.01.002
Silva, P. M., Gonçalves, S., and Santos, N. C. (2014). Defensins: antifungal lessons from eukaryotes. Front. Microbiol. 5:97. doi: 10.3389/fmicb.2014.00097
Silva, P. I. Jr., Daffre, S., and Bulet, P. (2000). Isolation and characterization of gomesin, an 18-residue cysteine-rich defense peptide from the spider Acanthoscurria gomesiana hemocytes with sequence similarities to horseshoe crab antimicrobial peptides of the tachyplesin family. J. Biol. Chem. 275, 33464–33470. doi: 10.1074/jbc.M001491200
Simmaco, M., Mignogna, G., Canofeni, S., Miele, R., Mangoni, M. L., and Barra, D. (1996). Temporins, antimicrobial peptides from the European red frog Rana temporaria. Eur. J. Biochem. 242, 788–792. doi: 10.1111/j.1432-1033.1996.0788r.x
Simon, A., Kullberg, B. J., Tripet, B., Boerman, O. C., Zeeuwen, P., van der Ven-Jongekrijg, J., et al. (2008). Drosomycin-like defensin, a human homologue of Drosophila melanogaster drosomycin with antifungal activity. Antimicrob. Agents Chemother. 52, 1407–1412. doi: 10.1128/AAC.00155-07
Singh, A., Upadhyay, V., Upadhyay, A. K., Singh, S. M., and Panda, A. K. (2015). Protein recovery from inclusion bodies of Escherichia coli using mild solubilization process. Microb. Cell Factories 14:41. doi: 10.1186/s12934-015-0222-8
Skerlavaj, B., Benincasa, M., Risso, A., Zanetti, M., and Gennaro, R. (1999). SMAP-29: a potent antibacterial and antifungal peptide from sheep leukocytes. FEBS Lett. 463, 58–62. doi: 10.1016/s0014-5793(99)01600-2
Slightom, J. L., Metzger, B. P., Luu, H. T., and Elhammer, A. P. (2009). Cloning and molecular characterization of the gene encoding the Aureobasidin a biosynthesis complex in Aureobasidium pullulans BP-1938. Gene 431, 67–79. doi: 10.1016/j.gene.2008.11.011
Soltani, S., Keymanesh, K., and Sardari, S. (2007). In silico analysis of antifungal peptides. Expert Opin. Drug Discov. 2, 837–847. doi: 10.1517/17460441.2.6.837
Soravia, E., Martini, G., and Zasloff, M. (1988). Antimicrobial properties of peptides from Xenopus granular gland secretions. FEBS Lett. 228, 337–340. doi: 10.1016/0014-5793(88)80027-9
Souza, A. L., Díaz-Dellavalle, P., Cabrera, A., Larrañaga, P., Dalla-Rizza, M., and De-Simone, S. G. (2013). Antimicrobial activity of pleurocidin is retained in Plc-2, a C-terminal 12-amino acid fragment. Peptides 45, 78–84. doi: 10.1016/j.peptides.2013.03.030
Souza, B. M., Mendes, M. A., Santos, L. D., Marques, M. R., César, L. M. M., Almeida, R. N. A., et al. (2005). Structural and functional characterization of two novel peptide toxins isolated from the venom of the social wasp Polybia Paulista. Peptides 26, 2157–2164. doi: 10.1016/j.peptides.2005.04.026
Steckbeck, J. D., Deslouches, B., and Montelaro, R. C. (2014). Antimicrobial peptides: new drugs for bad bugs? Expert. Opin. Biol. Ther. 14, 11–14. doi: 10.1517/14712598.2013.844227
Sung, W. S., Park, S. H., and Lee, D. G. (2008). Antimicrobial effect and membrane-active mechanism of Urechistachykinins, neuropeptides derived from Urechis unicinctus. FEBS Lett. 582, 2463–2466. doi: 10.1016/j.febslet.2008.06.015
Tam, J. P., Wang, S., Wong, K. H., and Tan, W. L. (2015). Antimicrobial peptides from plants. Pharmaceuticals 8, 711–757. doi: 10.3390/ph8040711
Tassin, S., Broekaert, W. F., Marion, D., Acland, D. P., Ptak, M., Vovelle, F., et al. (1998). Solution structure of ace-AMP1, a potent antimicrobial protein extracted from onion seeds. Structural analogies with plant nonspecific lipid transfer proteins. Biochemistry 37, 3623–3637. doi: 10.1021/bi9723515
Terras, F. R., Torrekens, S., Van Leuven, F., et al. (1993). A new family of basic cysteine-rich plant antifungal proteins from Brassicaceae species. FEBS Lett. 316, 233–240. doi: 10.1016/0014-5793(93)81299-f
Theis, T., Marx, F., Salvenmoser, W., Stahl, U., and Meyer, V. (2005). New insights into the target site and mode of action of the antifungal protein of aspergillus giganteus. Res. Microbiol. 156, 47–56. doi: 10.1016/j.resmic.2004.08.006
Theis, T., and Stahl, U. (2004). Antifungal proteins: targets, mechanisms and prospective applications. CMLS 61, 437–455. doi: 10.1007/s00018-003-3231-4
Thery, T., Lynch, K. M., and Arendt, E. K. (2019). Natural antifungal peptides/proteins as model for novel food preservatives. Compr. Rev. Food Sci. Food Saf. 18, 1327–1360. doi: 10.1111/1541-4337.12480
Thevissen, K., Ferket, K. K., François, I. E., and Cammue, B. P. (2003). Interactions of antifungal plant defensins with fungal membrane components. Peptides 24, 1705–1712. doi: 10.1016/j.peptides.2003.09.014
Thevissen, K., Warnecke, D. C., François, I. E., Leipelt, M., Heinz, E., Ott, C., et al. (2004). Defensins from insects and plants interact with fungal glucosylceramides. J. Biol. Chem. 279, 3900–3905. doi: 10.1074/jbc.M311165200
Thorn, H. I., Guruceaga, X., Martin-Vicente, A., Nywening, A. V., Xie, J., Ge, W., et al. (2024). MOB-mediated regulation of septation initiation network (SIN) signaling is required for echinocandin-induced hyperseptation in Aspergillus fumigatus. mSphere 9:e0069523. doi: 10.1128/msphere.00695-23
Tracanna, V., de Jong, A., Medema, M. H., and Kuipers, O. P. (2017). Mining prokaryotes for antimicrobial compounds: from diversity to function. FEMS Microbiol. Rev. 41, 417–429. doi: 10.1093/femsre/fux014
Tran, D., Tran, P. A., Tang, Y. Q., Yuan, J., Cole, T., and Selsted, M. E. (2002). Homodimeric theta-defensins from rhesus macaque leukocytes: isolation, synthesis, antimicrobial activities, and bacterial binding properties of the cyclic peptides. J. Biol. Chem. 277, 3079–3084. doi: 10.1074/jbc.M109117200
Turcu, R., Patterson, M. J., and Omar, S. (2009). Influence of sodium intake on amphotericin B-induced nephrotoxicity among extremely premature infants. Pediatr. Nephrol. 24, 497–505. doi: 10.1007/s00467-008-1050-4
Ubukata, M., Uramoto, M., and Isono, K. (1984). The structure of neopeptins, inhibitors of fungal cell wall biosynthesis. Tetrahedron Lett. 25, 423–426. doi: 10.1016/S0040-4039(00)99901-5
Van Baarlen, P., Legendre, L., Kan, J., and Van, A. L. (2007). Plant defence compounds against botrytis infection. Botrytis Biol. Pathol. Control 22, 143–161. doi: 10.1007/978-1-4020-2626-39
Van den Bergh, K. P., Proost, P., Van Damme, J., Coosemans, J., Van Damme, E. J., and Peumans, W. J. (2002). Five disulfide bridges stabilize a hevein-type antimicrobial peptide from the bark of spindle tree (Euonymus europaeus L.). FEBS Lett. 530, 181–185. doi: 10.1016/s0014-5793(02)03474-9
van der Weerden, N. L., Bleackley, M. R., and Anderson, M. A. (2013). Properties and mechanisms of action of naturally occurring antifungal peptides. Cellular Mol. Life Sci. 70, 3545–3570. doi: 10.1007/s00018-013-1260-1
Van Dijck, P., Sjollema, J., Cammue, B. P., Lagrou, K., Berman, J., d'Enfert, C., et al. (2018). Methodologies for in vitro and in vivo evaluation of efficacy of antifungal and antibiofilm agents and surface coatings against fungal biofilms. Microbial Cell 5, 300–326. doi: 10.15698/mic2018.07.638
Vieira Gomes, A. M., Souza Carmo, T., Silva Carvalho, L., Mendonça Bahia, F., and Parachin, N. S. (2018). Comparison of yeasts as hosts for recombinant protein production. Microorganisms 6:38. doi: 10.3390/microorganisms6020038
Vizioli, J., Bulet, P., Hoffmann, J. A., Kafatos, F. C., Müller, H. M., and Dimopoulos, G. (2001). Gambicin: a novel immune responsive antimicrobial peptide from the malaria vector Anopheles gambiae. Proc. Natl. Acad. Sci. USA 98, 12630–12635. doi: 10.1073/pnas.221466798
Vriens, K., Cammue, B. P. A., and Thevissen, K. (2014). Antifungal plant defensins: mechanisms of action and production. Molecules 19, 12280–12303. doi: 10.3390/molecules190812280
Wakabayashi, H., Abe, S., Okutomi, T., Tansho, S., Kawase, K., and Yamaguchi, H. (1996). Cooperative anti-Candida effects of lactoferrin or its peptides in combination with azole antifungal agents. Microbiol. Immunol. 40, 821–825. doi: 10.1111/j.1348-0421.1996.tb01147.x
Wang, K., Dang, W., Xie, J., Zhu, R., Sun, M., Jia, F., et al. (2015). Antimicrobial peptide protonectin disturbs the membrane integrity and induces ROS production in yeast cells. Biochim. Biophys. Acta 1848, 2365–2373. doi: 10.1016/j.bbamem.2015.07.008
Wang, J., Qiang, J., Li, J., and Wang, D. (2024). Effect of high sodium ion level on the interaction of AmB with a cholesterol-rich phospholipid monolayer. Front. Mol. Biosci. 11:1405383. doi: 10.3389/fmolb.2024.1405383
Wei, G. X., Campagna, A. N., and Bobek, L. A. (2007). Factors affecting antimicrobial activity of MUC7 12-mer, a human salivary mucin-derived peptide. Ann. Clin. Microbiol. Antimicrob. 6:14. doi: 10.1186/1476-0711-6-14
Wilde, C. G., Griffith, J. E., Marra, M. N., Snable, J. L., and Scott, R. W. (1989). Purification and characterization of human neutrophil peptide 4, a novel member of the defensin family. J. Biol. Chem. 264, 11200–11203. doi: 10.1016/S0021-9258(18)60449-1
World Health Organization (2022). WHO fungal priority pathogens list to guide research, development and public health action. World Health Organization. Geneva: World Health Organization.
World Health Organization (WHO) (1999). Basic food safety for health workers. Geneva: World Health Organization (WHO).
Wu, Y., He, Y., and Ge, X. (2011). Functional characterization of the recombinant antimicrobial peptide Trx-ace-AMP1 and its application on the control of tomato early blight disease. Appl. Microbiol. Biotechnol. 90, 1303–1310. doi: 10.1007/s00253-011-3166-x
Yan, P.-S., Song, Y., Sakuno, E., Nakajima, H., Nakagawa, H., and Yabe, K. (2004). Cyclo(l-Leucyl-l-prolyl) produced by Achromobacter Xylosoxidans inhibits aflatoxin production by aspergillus parasiticus. Appl. Environ. Microbiol. 70, 7466–7473. doi: 10.1128/AEM.70.12.7466-7473.2004
Yan, J., Yuan, S. S., Jiang, L. L., Ye, X. J., Ng, T. B., and Wu, Z. J. (2015). Plant antifungal proteins and their applications in agriculture. Appl. Microbiol. Biotechnol. 99, 4961–4981. doi: 10.1007/s00253-015-6654-6
Yang, L., Harroun, T. A., Weiss, T. M., Ding, L., and Huang, H. W. (2001). Barrel-stave model or toroidal model? A case study on melittin pores. Biophys. J. 81, 1475–1485. doi: 10.1016/S0006-3495(01)75802-X
Yeaman, M. R., and Yount, N. Y. (2003). Mechanisms of antimicrobial peptide action and resistance. Pharmacol. Rev. 55, 27–55. doi: 10.1124/pr.55.1.2
Zasloff, M. (1987). Magainins, a class of antimicrobial peptides from Xenopus skin: isolation, characterization of two active forms, and partial cDNA sequence of a precursor. Proc. Natl. Acad. Sci. USA 84, 5449–5453. doi: 10.1073/pnas.84.15.5449
Zhang, D., Lu, Y., Chen, H., Wu, C., Zhang, H., Chen, L., et al. (2020). Antifungal peptides produced by actinomycetes and their biological activities against plant diseases. J. Antibiot. 73, 265–282. doi: 10.1038/s41429-020-0287-4
Zhang, Q. Y., Yan, Z. B., Meng, Y. M., Hong, X. Y., Shao, G., Ma, J. J., et al. (2021). Antimicrobial peptides: mechanism of action, activity and clinical potential. Mil. Med. Res. 8:48. doi: 10.1186/s40779-021-00343-2
Zhang, L. M., Yang, M., Zhou, S. W., Zhang, H., Feng, Y., Shi, L., et al. (2023). Blapstin, a diapause-specific peptide-like peptide from the Chinese medicinal beetle Blaps rhynchopetera. Has Antifungal Funct. Microbiol. Spectr. 11:e0308922. doi: 10.1128/spectrum.03089-22
Zhou, J., Kong, L., Fang, N., Mao, B., and Ai, H. (2016). Synthesis and functional characterization of MAF-1A peptide derived from the larvae of housefly, Musca domestica (Diptera: Muscidae). J. Med. Entomol. 53, 1467–1472. doi: 10.1093/jme/tjw110
Keywords: antifungal peptide, sources, mechanisms, production, application
Citation: Gong Y, Xue Q, Li J and Zhang S (2024) Antifungal peptides from living organisms. Front. Microbiol. 15:1511461. doi: 10.3389/fmicb.2024.1511461
Edited by:
Sunil D. Saroj, Symbiosis International University, IndiaReviewed by:
Lisa Lombardi, University College Dublin, IrelandPratima Gurung, Harvard Medical School, United States
Copyright © 2024 Gong, Xue, Li and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Shicui Zhang, c2N6aGFuZ0BvdWMuZWR1LmNu
 Yi Gong1
Yi Gong1 Shicui Zhang
Shicui Zhang