- Department of Neuroscience, University of Wisconsin-Madison, Madison, WI, United States
Axon regeneration is limited in the adult mammalian central nervous system (CNS) due to both intrinsic and extrinsic factors. Rodent studies have shown that developmental age can drive differences in intrinsic axon growth ability, such that embryonic rodent CNS neurons extend long axons while postnatal and adult CNS neurons do not. In recent decades, scientists have identified several intrinsic developmental regulators in rodents that modulate growth. However, whether this developmentally programmed decline in CNS axon growth is conserved in humans is not yet known. Until recently, there have been limited human neuronal model systems, and even fewer age-specific human models. Human in vitro models range from pluripotent stem cell-derived neurons to directly reprogrammed (transdifferentiated) neurons derived from human somatic cells. In this review, we discuss the advantages and disadvantages of each system, and how studying axon growth in human neurons can provide species-specific knowledge in the field of CNS axon regeneration with the goal of bridging basic science studies to clinical trials. Additionally, with the increased availability and quality of ‘omics datasets of human cortical tissue across development and lifespan, scientists can mine these datasets for developmentally regulated pathways and genes. As there has been little research performed in human neurons to study modulators of axon growth, here we provide a summary of approaches to begin to shift the field of CNS axon growth and regeneration into human model systems to uncover novel drivers of axon growth.
1. Introduction
Damage to adult central nervous system (CNS) neuronal projections called axons, in spinal cord injury (SCI) or optic nerve injury for example, results in little to no regeneration following damage. SCI is a devastating injury that results in life-long disability with high rates of morbidity and mortality, resulting in reduced quality of life, and a lifetime economic burden of approximately 2–4 billion dollars (Bennett and Emmady, 2023). The lack of regeneration in adult CNS axons is due to both an inhibitory environment and a lack of intrinsic axon growth in the neurons themselves (Curcio and Bradke, 2018; Zheng and Tuszynski, 2023). Interestingly, there is an age-dependent decrease in axon growth and regeneration ability in rodent CNS neurons in vitro and in vivo around the time of birth (Bregman et al., 1989; Chen et al., 1995; Dusart et al., 1997; Goldberg et al., 2002; Gwak et al., 2004; Blackmore and Letourneau, 2006; Genovese et al., 2006; Siegenthaler et al., 2008; Moore et al., 2009; Mar et al., 2014). This largely has been considered to be due to a downregulation of pro-regenerative networks and an upregulation of inhibitory gene expression networks in addition to developmental changes in the environmental niche (Gao et al., 2004; Arlotta et al., 2005; Wang et al., 2007; Moore et al., 2009; Geoffroy et al., 2016; Lu et al., 2020). Despite immense knowledge gained through identifying modulators of axon regeneration in rodents, there is currently no FDA-approved treatment for SCI (Perrin, 2014; Dietz et al., 2022). The lack of concordance of results between preclinical rodent studies and human clinical trials could be the result of the divergence of not only the human and mouse transcriptome, but also cis-regulatory regions (e.g., promoters, enhancers; Yue et al., 2014). Thus, it is important to establish human studies to determine the conservation across species of critical regulators of axon growth. Until recently, our ability to ask these questions was limited by the inability to acquire and culture human neurons of all ages. In this review, we summarize our current knowledge of developmentally-regulated axon growth pathways in the rodent CNS, human in vitro and in vivo models to study axon regeneration, and finally human cortical tissue transcriptomic and proteomic datasets available for mining developmentally-regulated genes in human CNS neurons.
2. An age-dependent decline in rodent CNS axon regeneration
It has been well-established that developmental age alters the response of rodent CNS neurons to extrinsic and intrinsic factors that limit adult but not embryonic axon growth and regeneration (Chen et al., 1995; Bandtlow and Loschinger, 1997; Dusart et al., 1997; Fawcett, 1997; Goldberg et al., 2002; Blackmore and Letourneau, 2006; Blackmore et al., 2012; Venkatesh et al., 2016; Curcio and Bradke, 2018; Venkatesh et al., 2018; Wang Z. et al., 2018; Zheng and Tuszynski, 2023). Extrinsically, CNS myelin contains inhibitory proteins, such as NOGO, myelin-associated glycoprotein (MAG), and oligodendrocyte myelin glycoprotein (OMGP) that limit adult axon CNS regeneration through collapsing growth cones following injury, but not embryonic CNS neurons (Waxman and Foster, 1980; Foran and Peterson, 1992; Filbin, 2003; Yiu and He, 2006). Further, with aging, there is an attenuated ability for local microglia and macrophages to remove myelin debris resulting in prolonged inflammation (Safaiyan et al., 2016; Cantuti-Castelvetri et al., 2018). After the inflammation has stabilized, a glial scar forms, which is both a mechanical barrier to axon regeneration, and also contains proteins inhibitory to axon growth such as chondroitin sulfate proteoglycans (CSPGs) released by reactive astrocytes and other cell types locally (Snow et al., 1990; McKeon et al., 1999; Becker and Becker, 2002; Jones et al., 2003; Tang et al., 2003; Yiu and He, 2006; Li et al., 2020). Further, in rodent SCI models there is increased inflammation in older animals as represented by an increase in monocyte-derived macrophages at the lesion site (Stewart et al., 2021).
Intrinsically, development and age influence various pathways in the neurons, from epigenome to transcriptome to translatome. Originally, developmentally-regulated transcription factors (TFs) were attractive targets because they can regulate an abundance of genes. For example, Krüppel-like factors (KLF) 6 and KLF7, which promote embryonic axon growth, are downregulated postnatally (Moore et al., 2009; Blackmore et al., 2012; Wang Z. et al., 2018; Kramer et al., 2021), whereas other members of the KLF family, such as KLF4 and 9, are upregulated developmentally, and inhibit axon growth and regeneration (Moore et al., 2009; Apara et al., 2017; Galvao et al., 2018; Trakhtenberg et al., 2018; Avila-Mendoza et al., 2020; Xu et al., 2021). Knockdown of Klf4 and 9 can increase axon growth in adult corticospinal tract (CST) neurons and retinal ganglion cells (RGCs; Moore et al., 2009; Apara et al., 2017; Galvao et al., 2018; Trakhtenberg et al., 2018). KLF9’s inhibitory function is partially mediated through the MAPK pathway as the inhibition or inactivation of c-Jun N-terminal kinase 3 (JNK3) and Dual-specificity phosphatase 14 (DUSP14) abolish KLF9’s inhibitory role in RGC axon regeneration (Apara et al., 2017; Galvao et al., 2018). KLF4 acts through binding signal transducer and activator of transcription 3 (STAT3) to block STAT3’s DNA-binding activity, limiting expression of its downstream regeneration-associated genes (RAGs) (Qin et al., 2013). In contrast to KLF4, the pro-regenerative KLF6 has cooperative roles with STAT3 to promote regeneration through the co-occupancy of similar regulatory regions of DNA (Wang Z. et al., 2018; Kramer et al., 2021). These studies suggest that in the CNS, both developmentally regulated growth-promoting and growth-repressing TFs act to drive changes in axon growth and regenerative abilities.
In addition to developmentally regulated TFs, cell signaling proteins such as insulin-like growth factor, cytochrome P450, and cyclic adenosine monophosphate (cAMP) levels are all downregulated during development (Gao et al., 2004; Arlotta et al., 2005; Wang et al., 2007). cAMP is an example where intrinsic and extrinsic factors collide, as cAMP downregulation in cerebellar and dorsal root ganglion (DRG) neurons at postnatal day (P)3–4 leads to increased inhibition of axon growth by myelin (Cai et al., 2002; Gao et al., 2004), due to the reduced phosphorylation of cAMP- response element-binding protein, and thus its effect on downstream genes such as arginase I and interleukin 6 (Cai et al., 2002; Redmond et al., 2002; Gao et al., 2004; Cao et al., 2006; Deng et al., 2009). In these examples, modulation of these age-dependent factors has been shown to improve CNS axon growth and regeneration in rodent models.
The mammalian target of rapamycin (mTOR) pathway is a central pathway that promotes growth through translational regulation (Ma and Blenis, 2009; Saxton and Sabatini, 2017). During CNS neuronal development, there is a downregulation of mTOR pathway activity, detected through a decrease in phospho-S6 signal, a marker of mTOR pathway activation (Liu et al., 2010; Teotia et al., 2019). The deletion of negative regulators of the mTOR pathway such as phosphatase and tensin homolog (Pten) and tuberous sclerosis 1 (Tsc1), promotes robust axon regeneration in RGCs and CST neurons (Park et al., 2008; Liu et al., 2010). However, later studies found that whereas Pten deletion in young animals (6 weeks) resulted in a strong regenerative phenotype, the same deletion in middle-aged animals (12–18 months) resulted in limited CST neuron axon regeneration (Geoffroy et al., 2016, 2017). This is thought to be due to the upregulation of eukaryotic translation initiation factor 4E-binding protein 1 (4E-BP) during aging which promotes cap-dependent translation of downstream mRNAs. mTORC1 is a repressor of 4E-BP, thus Pten deletion should lead to increased activation of mTORC1, and increased repression of 4E-BP. However, because 4E-BP expression is upregulated with age, this prevents the downstream effects of Pten deletion, leading to the lack of regeneration in older animals (Yang et al., 2014; Geoffroy et al., 2016). Thus, Pten deletion, one of the hallmark gene manipulations that promotes robust axon regeneration either singly (Park et al., 2008, 2010; Liu et al., 2010; Du et al., 2015; Geoffroy et al., 2016) or in combination with other treatments (Kurimoto et al., 2010; Sun et al., 2011; de Lima et al., 2012; Lewandowski and Steward, 2014; O’Donovan et al., 2014; Ohtake et al., 2014; Geoffroy et al., 2015; Jin et al., 2015; Lim J. H. et al., 2016; Weng et al., 2018; Huang et al., 2019; Xie et al., 2022), is hindered in the aged rodent CNS due to age-dependent changes. Other mTOR pathway regulators are developmentally regulated such as LIN28A, an RNA-binding protein, which is downregulated by early postnatal development. Its upregulation in both CST neurons and RGCs results in robust axon regeneration and improved motor function in the injured mice (Nathan et al., 2020).
It has been hypothesized that the reduced effectiveness of known pro-regenerative transcription factors in aging rodent models may be due to changes in chromatin accessibility. For example, overexpression of pro-regenerative transcription factors such as JUN and STAT3 have attenuated effects on adult CST axon regeneration, despite their regenerative phenotypes when overexpressed in early postnatal neurons in vitro (Lerch et al., 2014; Mehta et al., 2016; Venkatesh et al., 2018). Similarly, with chromatin accessibility analysis, the pro-regenerative TF KLF7 had reduced DNA binding in adult cortical neurons due to changes in chromatin structure, and thus decreased DNA accessibility (Blackmore et al., 2012; Venkatesh et al., 2018; Pita-Thomas et al., 2021). These examples of reduced effectiveness of TFs in adult neurons may be due to a general developmental decrease in promoter accessibility of RAGs (Gaub et al., 2011; Venkatesh et al., 2016). A possible mechanism underlying these changes in DNA accessibility is through an age-dependent decline in histone acetyltransferase CBP/p300 activity, which relaxes chromatin and allows for transcriptional initiation (Gaub et al., 2010, 2011). Overexpression of p300 in adult injured RGCs, and to a lesser extent in CNS upper motor neurons, promotes axon regeneration through increased p300 interaction and acetylation of the promoter regions of RAGs, leading to increased DNA accessibility and transcription (Gaub et al., 2010, 2011; Muller et al., 2022). On a global scale, transient pulsing of 3 out of the 4 Yamanaka reprogramming factors (octamer-binding transcription factor 4 (OCT4), sex-determining region Y box 2 (SOX2), and KLF4) in adult RGCs reverted these neurons to an embryonic state, rejuvenating the chromatin landscape, and resulting in increased optic nerve regeneration post-injury (Lu et al., 2020). This increased regeneration following transient reprogramming was mediated by tet methylcytosine dioxygenase 1 (TET1) and TET2-dependent DNA demethylation shifting the epigenetic landscape from an adult to an embryonic state. Thus, a developmental switch in DNA accessibility may act to limit CNS axon regenerative ability in the adult.
Tremendous progress has been made in identifying intrinsic and extrinsic rodent regulators of CNS axon regeneration, yet we do not know if these are conserved in human CNS neurons. For example, do pathways like mTOR and cAMP mediate similar effects on human axon growth? Is CNS axon growth and regeneration ability also developmentally-regulated in humans, and if so, what are the underlying changes that drive this phenotype? These are open questions where little has been studied in the context of human model systems.
3. Models to study human axon growth
In the last two decades, with the advent of human pluripotent stem cell (hPSC) technology which includes both human embryonic stem cells (hESCs) and induced pluripotent stem cells (iPSCs), and direct reprogramming technologies, we have seen an increase in human disease and therapeutic models that can now be applied to studies of axon growth and regeneration.
3.1. Human pluripotent stem cell (hESC and iPSC) models
HESCs are embryonic stem cells isolated from the inner cell mass of a human blastocyst which can be cultured in vitro, and differentiated into a wide variety of cells, including neurons (Figure 1; Shamblott et al., 1998; Thomson et al., 1998; Keller and Snodgrass, 1999). To avoid the ethical concerns of the source of hESCs, iPSCs have become a strong alternative for creating human neurons (Figure 1). IPSCs are generated from somatic cells using the overexpression of specific proteins resulting in cellular, epigenetic, and transcriptomic rejuvenation, and returning the original adult cell to a prenatal epigenetic and cellular age.
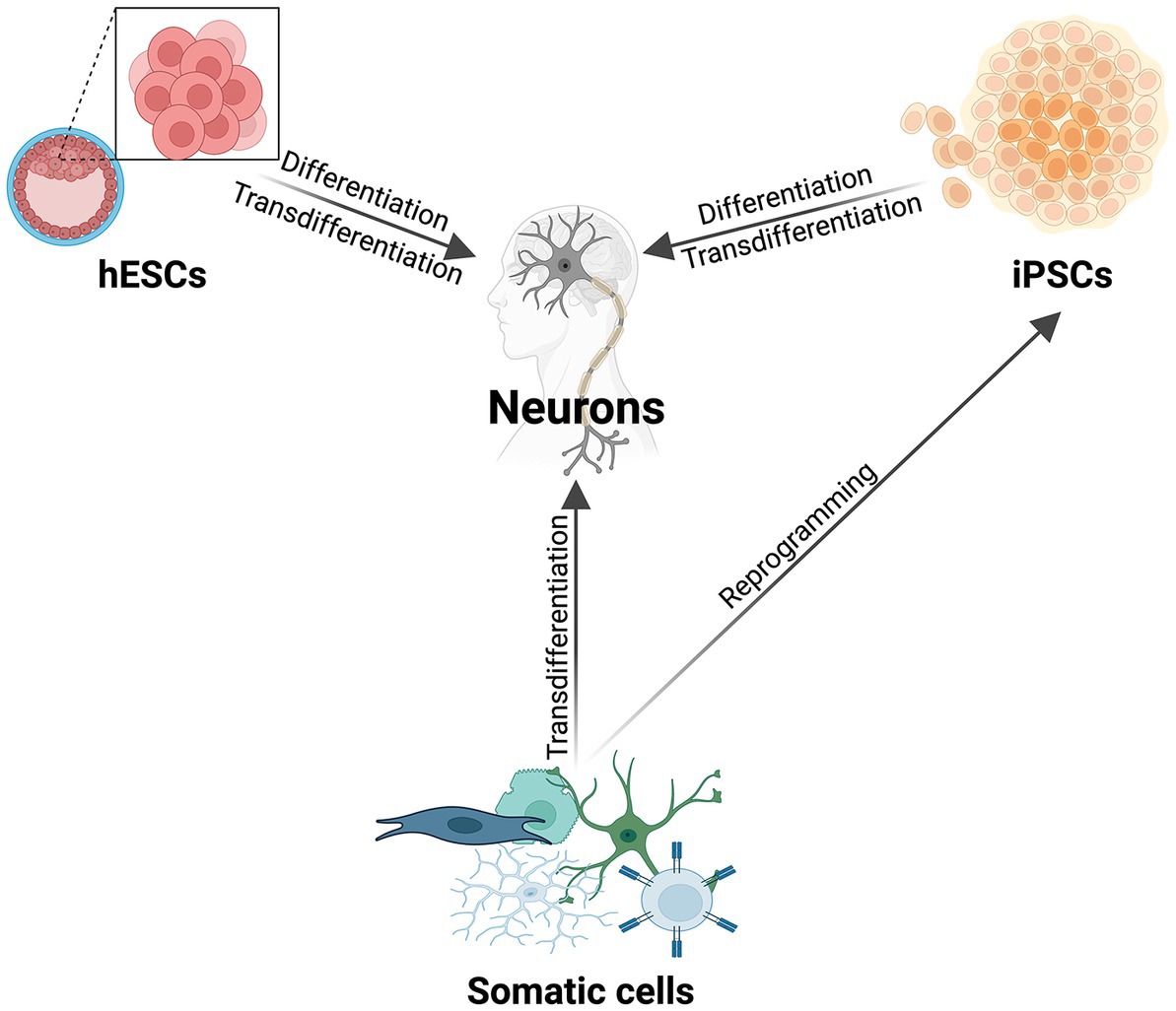
Figure 1. The most commonly used methods to make human neurons. Schematic illustrating the major methods of differentiating and direct reprogramming of various cell types to human neurons (hESCs, iPSCs, somatic cells). In addition to the listed cell types for transdifferentiation, there are a multitude of donor cells that can be directly reprogrammed to human neurons [reviewed in Kim et al. (2021)].
HPSCs can be differentiated into diverse neural precursors, neurons, and glial subtypes following the addition of various combinations of small molecules and growth factors that mirror developmental patterning [reviewed in (Kramer et al., 2013; Chuang et al., 2015; Tao and Zhang, 2016)]. Both hPSCs and hPSC-derived neural progenitor cells (NPCs) are highly proliferative, allowing for the constant expansion of the starting population, and the generation of large numbers of neurons. However, depending on the desired final cell type, it can take weeks to months for neurons to fully differentiate and mature because this differentiation paradigm mimics development. A shorter process to generate these subtypes of neurons is the forced overexpression (direct reprogramming) of neurogenic transcription factors or microRNAs in hPSCs without the natural progression of development (Mertens et al., 2015, 2018; Tang et al., 2017).
HPSC-derived cells can be cultured in 2D or 3D, with 2D systems allowing for easy analysis of individual neuronal morphology, whereas 3D models have the advantage of mirroring the in vivo environment (Seo et al., 2022). A small number of studies have used hPSC-derived neurons in 2D culture for high content screens to test for compounds that could modulate intrinsic axon growth, as well as for the impact of various substrates and scaffolds on axon growth [hESC-derived neurons: (Lam et al., 2010; Shin et al., 2010), iPSC-derived neurons: (Sirenko et al., 2014; Hancock et al., 2015; Sherman and Bang, 2018)]. Recently, researchers have generated 3D motor column organoids to model amyotrophic lateral sclerosis (ALS). These 3D organoids contain a mixture of motor neurons and interneurons, with key elements of spinal cord organization replicated in this model such as motor neuron axons bundles exiting the organoid together, similar to a motor nerve fascicle, and ventral interneuron localization (Seo et al., 2022). Further, assembloids, generated from organoids and spheroids of different structures such as cerebral cortex, hindbrain/spinal cord, and skeletal muscle can be assembled together, in efforts to model the entire cortex to muscle functional circuitry (Giandomenico et al., 2019; Andersen et al., 2020).
While all hPSC-derived models do a good job of modeling developmental ages due to the rejuvenated age of hPSC-differentiated progeny, this may be a disadvantage if one wants to study adult human neuron axon growth and regeneration.
3.2. Human cells as treatment: in vivo use of hESC- and iPSC-derived neurons
The most common application of hPSC-differentiated neurons is therapeutic transplantation for neurodegenerative diseases. In Parkinson’s disease (PD), for example, there has been great progress with midbrain dopaminergic progenitor transplantation to replace the endogenous degenerated neurons in animal models (Wang Y. K. et al., 2018; Piao et al., 2021; Tao et al., 2021; Xiong et al., 2021), and now in clinical trials (Kim et al., 2022). These hPSC-derived midbrain dopaminergic progenitors transplanted into the midbrain of a PD mouse not only extend their axons to different targets in the brain, but are also capable of integrating into the local neuronal circuitry, resulting in moderate recovery of PD-induced motor deficits (Xiong et al., 2021). Similar results were seen with autologous transplantation of iPSCs in non-human primates (Tao et al., 2021).
Transplantation has also been used in efforts to treat SCIs. Early studies transplanted rodent embryonic cortical neurons with high intrinsic growth potential into the adult rodent cortex, and observed neuron grafts extending long-range projections and forming synapses with cortical and subcortical structures (Fricker-Gates et al., 2002; Gaillard et al., 2007; Gaillard and Sauve, 2007; Falkner et al., 2016). With the advent of hPSC technology, scientists have now generated chimeric animals through transplantation of human cells in rodent and non-human primate spinal cords (Nori et al., 2011; Lu et al., 2012, 2014a,b; Kadoya et al., 2016; Li and Chen, 2016; Nagoshi and Okano, 2017; Dulin et al., 2018; Okubo et al., 2018; Rosenzweig et al., 2018; Kumamaru et al., 2018a,b; Kitagawa et al., 2022). For example, Wertheim and colleagues generated hPSC-derived motor neurons and created hydrogels from porcine extracellular matrix (ECM) for transplantation into rodent spinal cord at the site of a SCI. This transplant resulted in enhanced cell survival, reduced inflammation and gliosis at the lesion site, and overall improved motor outcomes (Wertheim et al., 2022). Transplantation of hPSC-derived progenitors and neurons to the injury site not only alters the local environment to be growth-permissive, but also produces neurons that can integrate into the host neural circuitry, acting as a relay to improve recovery post-SCI (Cizkova et al., 2007; Usvald et al., 2010; van Gorp et al., 2013; Lu et al., 2014b, 2017; Okubo et al., 2018; Rosenzweig et al., 2018; Poplawski et al., 2020). These are exciting prospects, however, additional longitudinal studies need to be performed to evaluate how the transplantation of hPSC-derived NPCs alter the local environment and circuitry over time. HPSC-derived NPCs recently have moved into ongoing clinical trials for treatment of patients with SCIs through transplantation at the site of injury (Sugai et al., 2021; Kim et al., 2022), yet a key limitation in moving from animal models to human models is the need to generate clinical grade hPSCs. This process is extremely laborious and expensive, especially when generating autologous hPSCs from patient cells (Kim et al., 2022). As a result, scientists have been developing HLA-compatible iPSC lines to cover most of the world’s population to reduce the need to generate autologous hPSCs to prevent immune rejection (Rim et al., 2018; Jang et al., 2019; Yoshida et al., 2023). The potential for large-scale production of hPSC-derived neurons in combination with their high intrinsic axon growth potential and plasticity supports these cells as a good model for therapeutic treatments.
3.3. Direct reprogramming to create age-specific human neurons
To ask if developmental and age-specific regulators drive changes in human CNS axon regeneration requires the use of a human neuronal model system that maintains age. Direct reprogramming is the conversion of one somatic cell type into another, or transdifferentiation (Figure 1; Mertens et al., 2015; Tang et al., 2017; Xu et al., 2020; Vasan et al., 2021). This process preserves both the epigenetic and transcriptomic age signature, and cellular aging components of the donor cell (Mertens et al., 2015; Tang et al., 2017). This is dissimilar to differentiated cells from hPSCs, as both hESCs and iPSCs have epigenetic signatures consistent with a prenatal age (Takahashi et al., 2007; Yu et al., 2007; Horvath, 2013; Kramer et al., 2013; Mertens et al., 2015, 2021; Chen et al., 2019; Piao et al., 2021).
In recent years, in vitro directly reprogrammed neurons have become a powerful tool to study the molecular underpinnings of aging in neurons, and to model neurodegenerative diseases (Liu et al., 2014; Fiesel et al., 2015; Lim S. M. et al., 2016; Liu M. L. et al., 2016; Victor et al., 2018; Mertens et al., 2021; Vasan et al., 2021; Oh et al., 2022). Direct reprogramming can be achieved through overexpression of key transcription factors (El Wazan et al., 2019), overexpression of microRNAs (Yoo et al., 2011; Victor et al., 2014; Huh et al., 2016; Abernathy et al., 2017; Cates et al., 2021), genome editing with CRISPR/Cas9 (Chakraborty et al., 2014; Chavez et al., 2015; Savell et al., 2019), and small molecules (Cheng et al., 2015; Dai et al., 2015; Li et al., 2015; Zhang et al., 2015; Gao et al., 2017; Wan et al., 2018; Yang et al., 2019). Additionally, in vitro direct reprogramming can transdifferentiate various cell types, such as fibroblasts, astrocytes, and even T-cells (Zhang et al., 2015; Gao et al., 2017; Tanabe et al., 2018; Wang et al., 2021) into diverse neuronal types, such as glutamatergic (Ambasudhan et al., 2011; Pang et al., 2011; Yoo et al., 2011; Aydin et al., 2019), dopaminergic (Caiazzo et al., 2011; Pfisterer et al., 2011), spinal lower motor (Son et al., 2011; Liu L. et al., 2016; Tang et al., 2017), cholinergic (Liu et al., 2013), serotonergic neurons (Vadodaria et al., 2016; Xu et al., 2016), and RGCs (Wang et al., 2020). One of the strengths of this system is that the age of the original cell is retained, making it a better model to study age-specific effects. For example, Huntington’s disease (HD) patient fibroblast-induced neurons showed mutant huntingtin (HTT) aggregates and associated mitochondrial and DNA defects, but patient fibroblasts reprogrammed first to iPSCs, then transdifferentiated into neurons, did not show any abnormalities (Victor et al., 2018). This was also recapitulated in another HD study with autophagy (Oh et al., 2022). In the context of axon growth, this reprogramming strategy enables the study of adult human neurons, however, a major limitation of this system is the low yield of neurons compared to hPSC-derived, and the limited starting material due to passage exhaustion of somatic cells.
Each model discussed above has pros and cons based on the specific scientific question being asked. HPSC-derived neurons may be the best model for SCI and optic neuropathy transplantation, or high content screens for modulators of axon growth. However, direct reprogramming is a better strategy for studying age-specific regulation of axon outgrowth.
3.4. Human cortical tissue datasets
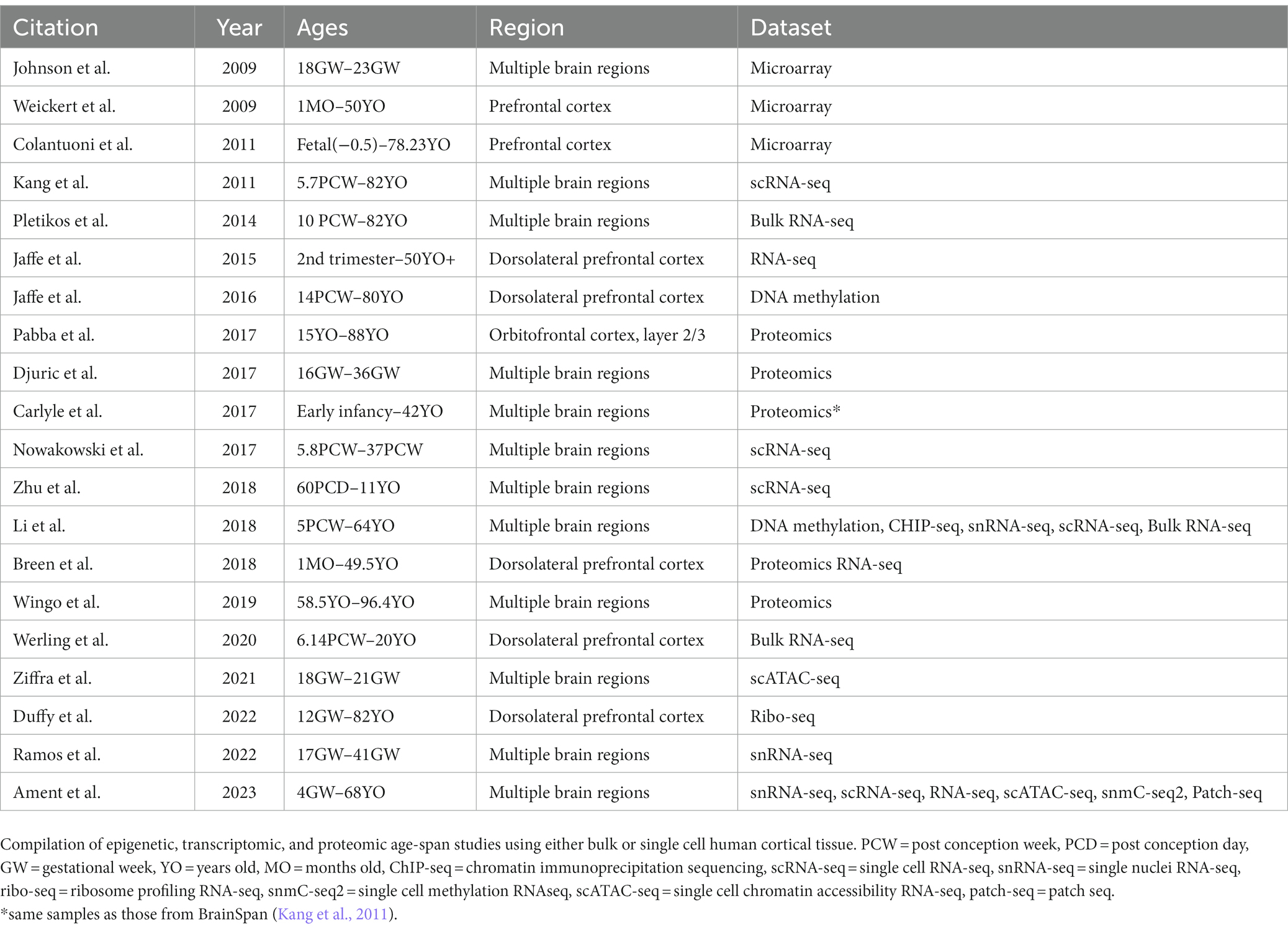
Large sequencing datasets from human brain are useful repositories for identifying gene expression changes in human CNS neurons during development and throughout aging. Global consortiums have been formed with the goal of identifying transcriptomic and proteomic changes across development and pathological states (PsychENCODE, BrainSpan, Brainseq, SpaceTx, Human cell atlas, etc.). Using bulk RNA sequencing (RNA-seq) and microarrays, multiple studies have published developmentally-regulated transcripts and their expression trajectories across time in various brain regions such as the dorsolateral prefrontal cortex (Breen et al., 2018; Werling et al., 2020), prefrontal cortex (Weickert et al., 2009; Colantuoni et al., 2011; Jaffe et al., 2015), and other brain regions (Johnson et al., 2009; Pletikos et al., 2014). These studies can be useful for correlating key genes and pathways that change across lifespan. However, the cell composition of bulk-dissected regions can vary greatly during brain development and across regions, limiting our understanding of cell type-specific contributions to gene expression. Single cell RNA-seq (scRNA-seq) and single nuclei RNA-seq (snRNA-seq), while often lacking depth and resolution of gene expression, can provide neuronal type-specific transcriptome and spatial localization data (Kang et al., 2011; Nowakowski et al., 2017; Li et al., 2018; Zhu et al., 2018; Ramos et al., 2022; Ament et al., 2023). Recent studies have demonstrated that mismatches of proteomic data when superimposed on transcriptomic data can occur, demonstrating post-transcriptional regulation (Carlyle et al., 2017; Breen et al., 2018). This has supported the need to generate human proteomics data across aging. One method of addressing this knowledge gap is through tissue proteomics as seen in studies on postmortem tissue, spanning from gestational to adult brain donors (Carlyle et al., 2017; Djuric et al., 2017; Pabba et al., 2017; Breen et al., 2018; Wingo et al., 2019). Another method is through ribosome profiling followed by sequencing (Ribo-seq), which provides insight on the translational regulation that leads to proteome diversity across human lifespan (Duffy et al., 2022).
Transcription can be regulated at the epigenetic level through controlling DNA accessibility. It is well known that during development and aging, the chromatin landscape can reflect and control gene regulation (Ziffra et al., 2021; Ament et al., 2023). DNA methylation is one aspect of epigenetic regulation and has been studied in the human brain across development, aging, and in diseased states (Jaffe et al., 2016; Li et al., 2018). Chromatin accessibility can also be examined through assay for transposase-accessible chromatin sequencing (ATAC-seq). A recent study performing single cell ATAC-seq of the developing human forebrain revealed cell-type and region-specific changes during corticogenesis (Ziffra et al., 2021). Further, rapid advancements in sequencing technology have led scientists to collaborate in generating multi-omics datasets, integrating various types of transcriptomic data with epigenomic data to provide a comprehensive molecular view of human brain development and aging (Li et al., 2018; Ament et al., 2023). By superimposing epigenomic datasets on transcriptomic datasets, it is possible to identify meaningful gene clusters that are similarly regulated epigenetically, potentially revealing developmental changes in DNA accessibility and master drivers of transcriptional programs in human neurons.
All the datasets described above have been used primarily to understand neurodevelopmental disorders, however, they are also suitable for mining developmentally-regulated genes for human axon growth and regeneration studies (Table 1).
4. Discussion: what do these human model systems mean for the future of axon regeneration studies?
The establishment of numerous techniques to make human CNS neurons allows us to ask species-specific questions about regulators of human axon growth. Each in vitro system has its unique positive attributes and limitations suitable for answering specific types of questions. While in vitro studies can provide a wealth of knowledge, the inability to study human neurons in an intact in vivo nervous system, which has a dynamic endogenous environment and age-specific signaling, cellular, and structural changes, has limited the adaptation of human model systems for studies of axon growth. However, developments in chimeric transplantation of human cells into the rodent CNS to dynamically visualize adult human axon growth in a systemic environment could take us one step closer to an improved human in vivo model. Harnessing the knowledge gained from these diverse human model systems and datasets may reveal novel human-specific, cell type-specific regulators of axon growth, as well as potentially identifying developmentally-regulated factors that could influence this growth. Ultimately, adding human systems to our studies of CNS axon growth and regeneration may accelerate and increase the success of the transition of preclinical studies to clinical trials.
4.1. Permission to reuse and copyright
Permission must be obtained for use of copyrighted material from other sources (including the web). Please note that it is compulsory to follow figure instructions.
Author contributions
BL and DM co-designed, interpreted the relevant literature, and wrote the manuscript. All authors contributed to the article and approved the submitted version.
Funding
The authors thank our funding sources: 1F30NS122478 (to BL), 1R21NS111192–01 (to DM), T32GM140935 (to University of Wisconsin-Madison Medical Scientist Training Program).
Acknowledgments
The authors thank A. Bhattacharyya for comments on the manuscript.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fnins.2023.1198041/full#supplementary-material
References
Abernathy, D. G., Kim, W. K., McCoy, M. J., Lake, A. M., Ouwenga, R., Lee, S. W., et al. (2017). MicroRNAs induce a permissive chromatin environment that enables neuronal subtype-specific reprogramming of adult human fibroblasts. Cell Stem Cell 21:e9, 332–348.e9. doi: 10.1016/j.stem.2017.08.002
Ambasudhan, R., Talantova, M., Coleman, R., Yuan, X., Zhu, S., Lipton, S. A., et al. (2011). Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell 9, 113–118. doi: 10.1016/j.stem.2011.07.002
Ament, S. A., Adkins, R. S., Carter, R., Chrysostomou, E., Colantuoni, C., Crabtree, J., et al. (2023). The neuroscience multi-Omic archive: a BRAIN initiative resource for single-cell transcriptomic and epigenomic data from the mammalian BRAIN. Nucleic Acids Res. 51, D1075–D1085. doi: 10.1093/nar/gkac962
Andersen, J., Revah, O., Miura, Y., Thom, N., Amin, N. D., Kelley, K. W., et al. (2020). Generation of functional human 3D Cortico-motor Assembloids. Cells 183:e26
Apara, A., Galvao, J., Wang, Y., Blackmore, M., Trillo, A., Iwao, K., et al. (2017). KLF9 and JNK3 interact to suppress axon regeneration in the adult CNS. J. Neurosci. 37, 9632–9644. doi: 10.1523/JNEUROSCI.0643-16.2017
Arlotta, P., Molyneaux, B. J., Chen, J., Inoue, J., Kominami, R., and Macklis, J. D. (2005). Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221. doi: 10.1016/j.neuron.2004.12.036
Avila-Mendoza, J., Subramani, A., and Denver, R. J. (2020). Kruppel-like factors 9 and 13 Block axon growth by transcriptional repression of key components of the cAMP Signaling pathway. Front. Mol. Neurosci. 13:602638. doi: 10.3389/fnmol.2020.602638
Aydin, B., Kakumanu, A., Rossillo, M., Moreno-Estelles, M., Garipler, G., Ringstad, N., et al. (2019). Proneural factors Ascl1 and Neurog2 contribute to neuronal subtype identities by establishing distinct chromatin landscapes. Nat. Neurosci. 22, 897–908. doi: 10.1038/s41593-019-0399-y
Bandtlow, C. E., and Loschinger, J. (1997). Developmental changes in neuronal responsiveness to the CNS myelin-associated neurite growth inhibitor NI-35/250. Eur. J. Neurosci. 9, 2743–2752. doi: 10.1111/j.1460-9568.1997.tb01703.x
Becker, C. G., and Becker, T. (2002). Repellent guidance of regenerating optic axons by chondroitin sulfate glycosaminoglycans in zebrafish. J. Neurosci. 22, 842–853. doi: 10.1523/JNEUROSCI.22-03-00842.2002
Blackmore, M., and Letourneau, P. C. (2006). Changes within maturing neurons limit axonal regeneration in the developing spinal cord. J. Neurobiol. 66, 348–360. doi: 10.1002/neu.20224
Blackmore, M. G., Wang, Z., Lerch, J. K., Motti, D., Zhang, Y. P., Shields, C. B., et al. (2012). Kruppel-like factor 7 engineered for transcriptional activation promotes axon regeneration in the adult corticospinal tract. Proc. Natl. Acad. Sci. U. S. A. 109, 7517–7522. doi: 10.1073/pnas.1120684109
Breen, M. S., Ozcan, S., Ramsey, J. M., Wang, Z., Ma’ayan, A., Rustogi, N., et al. (2018). Temporal proteomic profiling of postnatal human cortical development. Transl. Psychiatry 8:267. doi: 10.1038/s41398-018-0306-4
Bregman, B. S., Kunkel-Bagden, E., McAtee, M., and O’Neill, A. (1989). Extension of the critical period for developmental plasticity of the corticospinal pathway. J. Comp. Neurol. 282, 355–370. doi: 10.1002/cne.902820304
Cai, D., Deng, K., Mellado, W., Lee, J., Ratan, R. R., and Filbin, M. T. (2002). Arginase I and polyamines act downstream from cyclic AMP in overcoming inhibition of axonal growth MAG and myelin in vitro. Neuron 35, 711–719. doi: 10.1016/S0896-6273(02)00826-7
Caiazzo, M., Dell’Anno, M. T., Dvoretskova, E., Lazarevic, D., Taverna, S., Leo, D., et al. (2011). Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nature 476, 224–227. doi: 10.1038/nature10284
Cantuti-Castelvetri, L., Fitzner, D., Bosch-Queralt, M., Weil, M. T., Su, M., Sen, P., et al. (2018). Defective cholesterol clearance limits remyelination in the aged central nervous system. Science 359, 684–688. doi: 10.1126/science.aan4183
Cao, Z., Gao, Y., Bryson, J. B., Hou, J., Chaudhry, N., Siddiq, M., et al. (2006). The cytokine interleukin-6 is sufficient but not necessary to mimic the peripheral conditioning lesion effect on axonal growth. J. Neurosci. 26, 5565–5573. doi: 10.1523/JNEUROSCI.0815-06.2006
Carlyle, B. C., Kitchen, R. R., Kanyo, J. E., Voss, E. Z., Pletikos, M., Sousa, A. M. M., et al. (2017). A multiregional proteomic survey of the postnatal human brain. Nat. Neurosci. 20, 1787–1795. doi: 10.1038/s41593-017-0011-2
Cates, K., McCoy, M. J., Kwon, J. S., Liu, Y., Abernathy, D. G., Zhang, B., et al. (2021). Deconstructing stepwise fate conversion of human fibroblasts to neurons by MicroRNAs. Cell Stem Cell 28:e9, 127–140.e9. doi: 10.1016/j.stem.2020.08.015
Chakraborty, S., Ji, H., Kabadi, A. M., Gersbach, C. A., Christoforou, N., and Leong, K. W. (2014). A CRISPR/Cas9-based system for reprogramming cell lineage specification. Stem Cell Rep. 3, 940–947. doi: 10.1016/j.stemcr.2014.09.013
Chavez, A., Scheiman, J., Vora, S., Pruitt, B. W., Tuttle, M., P R Iyer, E., et al. (2015). Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328. doi: 10.1038/nmeth.3312
Chen, D. F., Jhaveri, S., and Schneider, G. E. (1995). Intrinsic changes in developing retinal neurons result in regenerative failure of their axons. Proc. Natl. Acad. Sci. U. S. A. 92, 7287–7291. doi: 10.1073/pnas.92.16.7287
Chen, S., Zhang, J., Zhang, D., and Jiao, J. (2019). Acquisition of functional neurons by direct conversion: switching the developmental clock directly. J. Genet. Genomics 46, 459–465. doi: 10.1016/j.jgg.2019.10.003
Cheng, L., Gao, L., Guan, W., Mao, J., Hu, W., Qiu, B., et al. (2015). Direct conversion of astrocytes into neuronal cells by drug cocktail. Cell Res. 25, 1269–1272. doi: 10.1038/cr.2015.120
Chuang, J. H., Tung, L. C., and Lin, Y. (2015). Neural differentiation from embryonic stem cells in vitro: An overview of the signaling pathways. World J. Stem Cells 7, 437–447. doi: 10.4252/wjsc.v7.i2.437
Cizkova, D., Kakinohana, O., Kucharova, K., Marsala, S., Johe, K., Hazel, T., et al. (2007). Functional recovery in rats with ischemic paraplegia after spinal grafting of human spinal stem cells. Neuroscience 147, 546–560. doi: 10.1016/j.neuroscience.2007.02.065
Colantuoni, C., Lipska, B. K., Ye, T., Hyde, T. M., Tao, R., Leek, J. T., et al. (2011). Temporal dynamics and genetic control of transcription in the human prefrontal cortex. Nature 478, 519–523. doi: 10.1038/nature10524
Curcio, M., and Bradke, F. (2018). Axon regeneration in the central nervous system: facing the challenges from the inside. Annu. Rev. Cell Dev. Biol. 34, 495–521. doi: 10.1146/annurev-cellbio-100617-062508
Dai, P., Harada, Y., and Takamatsu, T. (2015). Highly efficient direct conversion of human fibroblasts to neuronal cells by chemical compounds. J. Clin. Biochem. Nutr. 56, 166–170. doi: 10.3164/jcbn.15-39
de Lima, S., Koriyama, Y., Kurimoto, T., Oliveira, J. T., Yin, Y., Li, Y., et al. (2012). Full-length axon regeneration in the adult mouse optic nerve and partial recovery of simple visual behaviors. Proc. Natl. Acad. Sci. U. S. A. 109, 9149–9154. doi: 10.1073/pnas.1119449109
Deng, K., He, H., Qiu, J., Lorber, B., Bryson, J. B., and Filbin, M. T. (2009). Increased synthesis of spermidine as a result of upregulation of arginase I promotes axonal regeneration in culture and in vivo. J. Neurosci. 29, 9545–9552. doi: 10.1523/JNEUROSCI.1175-09.2009
Dietz, V. A., Roberts, N., Knox, K., Moore, S., Pitonak, M., Barr, C., et al. (2022). Fighting for recovery on multiple fronts: the past, present, and future of clinical trials for spinal cord injury. Front. Cell. Neurosci. 16:977679. doi: 10.3389/fncel.2022.977679
Djuric, U., Rodrigues, D. C., Batruch, I., Ellis, J., Shannon, P., and Diamandis, P. (2017). Spatiotemporal proteomic profiling of human cerebral development. Mol. Cell. Proteomics 16, 1548–1562. doi: 10.1074/mcp.M116.066274
Du, K., Zheng, S., Zhang, Q., Li, S., Gao, X., Wang, J., et al. (2015). Pten deletion promotes regrowth of corticospinal tract axons 1 year after spinal cord injury. J. Neurosci. 35, 9754–9763. doi: 10.1523/JNEUROSCI.3637-14.2015
Duffy, E. E., Finander, B., Choi, G., Carter, A. C., Pritisanac, I., Alam, A., et al. (2022). Developmental dynamics of RNA translation in the human brain. Nat. Neurosci. 25, 1353–1365. doi: 10.1038/s41593-022-01164-9
Dulin, J. N., Adler, A. F., Kumamaru, H., Poplawski, G. H. D., Lee-Kubli, C., Strobl, H., et al. (2018). Injured adult motor and sensory axons regenerate into appropriate organotypic domains of neural progenitor grafts. Nat. Commun. 9:84. doi: 10.1038/s41467-017-02613-x
Dusart, I., Airaksinen, M. S., and Sotelo, C. (1997). Purkinje cell survival and axonal regeneration are age dependent: an in vitro study. J. Neurosci. 17, 3710–3726. doi: 10.1523/JNEUROSCI.17-10-03710.1997
El Wazan, L., Urrutia-Cabrera, D., and Wong, R. C. (2019). Using transcription factors for direct reprogramming of neurons in vitro. World J. Stem Cells 11, 431–444. doi: 10.4252/wjsc.v11.i7.431
Falkner, S., Grade, S., Dimou, L., Conzelmann, K. K., Bonhoeffer, T., Gotz, M., et al. (2016). Transplanted embryonic neurons integrate into adult neocortical circuits. Nature 539, 248–253. doi: 10.1038/nature20113
Fawcett, J. W. (1997). Astrocytic and neuronal factors affecting axon regeneration in the damaged central nervous system. Cell Tissue Res. 290, 371–377. doi: 10.1007/s004410050943
Fiesel, F. C., Ando, M., Hudec, R., Hill, A. R., Castanedes-Casey, M., Caulfield, T. R., et al. (2015). (Patho-)physiological relevance of PINK1-dependent ubiquitin phosphorylation. EMBO Rep. 16, 1114–1130. doi: 10.15252/embr.201540514
Filbin, M. T. (2003). Myelin-associated inhibitors of axonal regeneration in the adult mammalian CNS. Nat. Rev. Neurosci. 4, 703–713. doi: 10.1038/nrn1195
Foran, D. R., and Peterson, A. C. (1992). Myelin acquisition in the central nervous system of the mouse revealed by an MBP-lac Z transgene. J. Neurosci. 12, 4890–4897. doi: 10.1523/JNEUROSCI.12-12-04890.1992
Fricker-Gates, R. A., Shin, J. J., Tai, C. C., Catapano, L. A., and Macklis, J. D. (2002). Late-stage immature neocortical neurons reconstruct interhemispheric connections and form synaptic contacts with increased efficiency in adult mouse cortex undergoing targeted neurodegeneration. J. Neurosci. 22, 4045–4056. doi: 10.1523/JNEUROSCI.22-10-04045.2002
Gaillard, A., Prestoz, L., Dumartin, B., Cantereau, A., Morel, F., Roger, M., et al. (2007). Reestablishment of damaged adult motor pathways by grafted embryonic cortical neurons. Nat. Neurosci. 10, 1294–1299. doi: 10.1038/nn1970
Gaillard, F., and Sauve, Y. (2007). Cell-based therapy for retina degeneration: the promise of a cure. Vis. Res. 47, 2815–2824. doi: 10.1016/j.visres.2007.06.018
Galvao, J., Iwao, K., Apara, A., Wang, Y., Ashouri, M., Shah, T. N., et al. (2018). The Kruppel-like factor gene target Dusp14 regulates axon growth and regeneration. Invest. Ophthalmol. Vis. Sci. 59, 2736–2747. doi: 10.1167/iovs.17-23319
Gao, Y., Deng, K., Hou, J., Bryson, J. B., Barco, A., Nikulina, E., et al. (2004). Activated CREB is sufficient to overcome inhibitors in myelin and promote spinal axon regeneration in vivo. Neuron 44, 609–621. doi: 10.1016/j.neuron.2004.10.030
Gao, L., Guan, W., Wang, M., Wang, H., Yu, J., Liu, Q., et al. (2017). Direct generation of human neuronal cells from adult astrocytes by small molecules. Stem Cell Rep. 8, 538–547. doi: 10.1016/j.stemcr.2017.01.014
Gaub, P., Joshi, Y., Wuttke, A., Naumann, U., Schnichels, S., Heiduschka, P., et al. (2011). The histone acetyltransferase p300 promotes intrinsic axonal regeneration. Brain 134, 2134–2148. doi: 10.1093/brain/awr142
Gaub, P., Tedeschi, A., Puttagunta, R., Nguyen, T., Schmandke, A., and Di Giovanni, S. (2010). HDAC inhibition promotes neuronal outgrowth and counteracts growth cone collapse through CBP/p300 and P/CAF-dependent p53 acetylation. Cell Death Differ. 17, 1392–1408. doi: 10.1038/cdd.2009.216
Genovese, T., Mazzon, E., Di Paola, R., Crisafulli, C., Muia, C., Bramanti, P., et al. (2006). Increased oxidative-related mechanisms in the spinal cord injury in old rats. Neurosci. Lett. 393, 141–146. doi: 10.1016/j.neulet.2005.09.060
Geoffroy, C. G., Hilton, B. J., Tetzlaff, W., and Zheng, B. (2016). Evidence for an age-dependent decline in axon regeneration in the adult mammalian central nervous system. Cell Rep. 15, 238–246. doi: 10.1016/j.celrep.2016.03.028
Geoffroy, C. G., Lorenzana, A. O., Kwan, J. P., Lin, K., Ghassemi, O., Ma, A., et al. (2015). Effects of PTEN and Nogo Codeletion on corticospinal axon sprouting and regeneration in mice. J. Neurosci. 35, 6413–6428. doi: 10.1523/JNEUROSCI.4013-14.2015
Geoffroy, C. G., Meves, J. M., and Zheng, B. (2017). The age factor in axonal repair after spinal cord injury: a focus on neuron-intrinsic mechanisms. Neurosci. Lett. 652, 41–49. doi: 10.1016/j.neulet.2016.11.003
Giandomenico, S. L., Mierau, S. B., and Gibbons, G. M (2019). Cerebral organoids at the air–liquid interface generate diverse nerve tracts with functional output. Nat Neurosci, 22, 669–679. doi: 10.1038/s41593-019-0350-2
Goldberg, J. L., Klassen, M. P., Hua, Y., and Barres, B. A. (2002). Amacrine-signaled loss of intrinsic axon growth ability by retinal ganglion cells. Science 296, 1860–1864. doi: 10.1126/science.1068428
Gwak, Y. S., Hains, B. C., Johnson, K. M., and Hulsebosch, C. E. (2004). Effect of age at time of spinal cord injury on behavioral outcomes in rat. J. Neurotrauma 21, 983–993. doi: 10.1089/0897715041650999
Hancock, M. K., Kopp, L., Kaur, N., and Hanson, B. J. (2015). A facile method for simultaneously measuring neuronal cell viability and neurite outgrowth. Curr. Chem. Genom. Trans. Med. 9, 6–16. doi: 10.2174/2213988501509010006
Horvath, S. (2013). DNA methylation age of human tissues and cell types. Genome Biol. 14:R115. doi: 10.1186/gb-2013-14-10-r115
Huang, H., Miao, L., Yang, L., Liang, F., Wang, Q., Zhuang, P., et al. (2019). AKT-dependent and -independent pathways mediate PTEN deletion-induced CNS axon regeneration. Cell Death Dis. 10:203. doi: 10.1038/s41419-018-1289-z
Huh, C. J., Zhang, B., Victor, M. B., Dahiya, S., Batista, L. F., Horvath, S., et al. (2016). Maintenance of age in human neurons generated by microRNA-based neuronal conversion of fibroblasts. elife 5. doi: 10.7554/eLife.18648
Jaffe, A. E., Gao, Y., Deep-Soboslay, A., Tao, R., Hyde, T. M., Weinberger, D. R., et al. (2016). Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nat. Neurosci. 19, 40–47. doi: 10.1038/nn.4181
Jaffe, A. E., Shin, J., Collado-Torres, L., Leek, J. T., Tao, R., Li, C., et al. (2015). Developmental regulation of human cortex transcription and its clinical relevance at single base resolution. Nat. Neurosci. 18, 154–161. doi: 10.1038/nn.3898
Jang, Y., Choi, J., Park, N., Kang, J., Kim, M., Kim, Y., et al. (2019). Development of immunocompatible pluripotent stem cells via CRISPR-based human leukocyte antigen engineering. Exp. Mol. Med. 51, 1–11. doi: 10.1038/s12276-018-0190-2
Jin, D., Liu, Y., Sun, F., Wang, X., Liu, X., and He, Z. (2015). Restoration of skilled locomotion by sprouting corticospinal axons induced by co-deletion of PTEN and SOCS3. Nat. Commun. 6:8074. doi: 10.1038/ncomms9074
Johnson, M. B., Kawasawa, Y. I., Mason, C. E., Krsnik, Z., Coppola, G., Bogdanovic, D., et al. (2009). Functional and evolutionary insights into human brain development through global transcriptome analysis. Neuron 62, 494–509. doi: 10.1016/j.neuron.2009.03.027
Jones, L. L., Sajed, D., and Tuszynski, M. H. (2003). Axonal regeneration through regions of chondroitin sulfate proteoglycan deposition after spinal cord injury: a balance of permissiveness and inhibition. J. Neurosci. 23, 9276–9288. doi: 10.1523/JNEUROSCI.23-28-09276.2003
Kadoya, K., Lu, P., Nguyen, K., Lee-Kubli, C., Kumamaru, H., Yao, L., et al. (2016). Spinal cord reconstitution with homologous neural grafts enables robust corticospinal regeneration. Nat. Med. 22, 479–487. doi: 10.1038/nm.4066
Kang, H. J., Kawasawa, Y. I., Cheng, F., Zhu, Y., Xu, X., Li, M., et al. (2011). Spatio-temporal transcriptome of the human brain. Nature 478, 483–489. doi: 10.1038/nature10523
Keller, G., and Snodgrass, H. R. (1999). Human embryonic stem cells: the future is now. Nat. Med. 5, 151–152. doi: 10.1038/5512
Kim, J. Y., Nam, Y., Rim, Y. A., and Ju, J. H. (2022). Review of the current trends in clinical trials involving induced pluripotent stem cells. Stem Cell Rev. Rep. 18, 142–154. doi: 10.1007/s12015-021-10262-3
Kim, K. M., Thaqi, M., Peterson, D. A., and Marr, R. A. (2021). Induced neurons for disease Modeling and repair: a focus on non-fibroblastic cell sources in direct reprogramming. Front. Bioeng. Biotechnol. 9:658498. doi: 10.3389/fbioe.2021.658498
Kitagawa, T., Nagoshi, N., Kamata, Y., Kawai, M., Ago, K., Kajikawa, K., et al. (2022). Modulation by DREADD reveals the therapeutic effect of human iPSC-derived neuronal activity on functional recovery after spinal cord injury. Stem Cell Reports 17, 127–142. doi: 10.1016/j.stemcr.2021.12.005
Kramer, A. S., Harvey, A. R., Plant, G. W., and Hodgetts, S. I. (2013). Systematic review of induced pluripotent stem cell technology as a potential clinical therapy for spinal cord injury. Cell Transplant. 22, 571–617. doi: 10.3727/096368912X655208
Kramer, A. A., Olson, G. M., Chakraborty, A., and Blackmore, M. G. (2021). Promotion of corticospinal tract growth by KLF6 requires an injury stimulus and occurs within four weeks of treatment. Exp. Neurol. 339:113644. doi: 10.1016/j.expneurol.2021.113644
Kumamaru, H., Kadoya, K., Adler, A. F., Takashima, Y., Graham, L., Coppola, G., et al. (2018a). Generation and post-injury integration of human spinal cord neural stem cells. Nat. Methods 15, 723–731. doi: 10.1038/s41592-018-0074-3
Kumamaru, H., Lu, P., Rosenzweig, E. S., and Tuszynski, M. H. (2018b). Activation of intrinsic growth State enhances host axonal regeneration into neural progenitor cell grafts. Stem Cell Rep. 11, 861–868. doi: 10.1016/j.stemcr.2018.08.009
Kurimoto, T., Yin, Y., Omura, K., Gilbert, H. Y., Kim, D., Cen, L. P., et al. (2010). Long-distance axon regeneration in the mature optic nerve: contributions of oncomodulin, cAMP, and pten gene deletion. J. Neurosci. 30, 15654–15663. doi: 10.1523/JNEUROSCI.4340-10.2010
Lam, H. J., Patel, S., Wang, A., Chu, J., and Li, S. (2010). In vitro regulation of neural differentiation and axon growth by growth factors and bioactive nanofibers. Tissue Eng. Part A 16, 2641–2648. doi: 10.1089/ten.tea.2009.0414
Lerch, J. K., Martinez-Ondaro, Y. R., Bixby, J. L., and Lemmon, V. P. (2014). cJun promotes CNS axon growth. Mol. Cell. Neurosci. 59, 97–105. doi: 10.1016/j.mcn.2014.02.002
Lewandowski, G., and Steward, O. (2014). AAVshRNA-mediated suppression of PTEN in adult rats in combination with salmon fibrin administration enables regenerative growth of corticospinal axons and enhances recovery of voluntary motor function after cervical spinal cord injury. J. Neurosci. 34, 9951–9962. doi: 10.1523/JNEUROSCI.1996-14.2014
Li, H., and Chen, G. (2016). In vivo reprogramming for CNS repair: regenerating neurons from endogenous glial cells. Neuron 91, 728–738. doi: 10.1016/j.neuron.2016.08.004
Li, X., Li, M., Tian, L., Chen, J., Liu, R., and Ning, B. (2020). Reactive Astrogliosis: implications in spinal cord injury progression and therapy. Oxidative Med. Cell. Longev. 2020, 1–14. doi: 10.1155/2020/9494352
Li, M., Santpere, G., Imamura Kawasawa, Y., Evgrafov, O. V., Gulden, F. O., Pochareddy, S., et al. (2018). Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362. doi: 10.1126/science.aat7615
Li, X., Zuo, X., Jing, J., Ma, Y., Wang, J., Liu, D., et al. (2015). Small-molecule-driven direct reprogramming of mouse fibroblasts into functional neurons. Cell Stem Cell 17, 195–203. doi: 10.1016/j.stem.2015.06.003
Lim, S. M., Choi, W. J., Oh, K. W., Xue, Y., Choi, J. Y., Kim, S. H., et al. (2016). Directly converted patient-specific induced neurons mirror the neuropathology of FUS with disrupted nuclear localization in amyotrophic lateral sclerosis. Mol. Neurodegener. 11:8. doi: 10.1186/s13024-016-0075-6
Lim, J. H., Stafford, B. K., Nguyen, P. L., Lien, B. V., Wang, C., Zukor, K., et al. (2016). Neural activity promotes long-distance, target-specific regeneration of adult retinal axons. Nat. Neurosci. 19, 1073–1084.
Liu, L., Huang, J. S., Han, C., Zhang, G. X., Xu, X. Y., Shen, Y., et al. (2016). Induced pluripotent stem cells in Huntington’s disease: disease Modeling and the potential for cell-based therapy. Mol. Neurobiol. 53, 6698–6708. doi: 10.1007/s12035-015-9601-8
Liu, K., Lu, Y., Lee, J. K., Samara, R., Willenberg, R., Sears-Kraxberger, I., et al. (2010). PTEN deletion enhances the regenerative ability of adult corticospinal neurons. Nat. Neurosci. 13, 1075–1081. doi: 10.1038/nn.2603
Liu, Y., Xue, Y., Ridley, S., Zhang, D., Rezvani, K., Fu, X. D., et al. (2014). Direct reprogramming of Huntington’s disease patient fibroblasts into neuron-like cells leads to abnormal neurite outgrowth, increased cell death, and aggregate formation. PLoS One 9:e109621. doi: 10.1371/journal.pone.0109621
Liu, M. L., Zang, T., and Zhang, C. L. (2016). Direct lineage reprogramming reveals disease-specific phenotypes of motor neurons from human ALS patients. Cell Rep. 14, 115–128. doi: 10.1016/j.celrep.2015.12.018
Liu, M. L., Zang, T., Zou, Y., Chang, J. C., Gibson, J. R., Huber, K. M., et al. (2013). Small molecules enable neurogenin 2 to efficiently convert human fibroblasts into cholinergic neurons. Nat. Commun. 4:2183. doi: 10.1038/ncomms3183
Lu, Y., Brommer, B., Tian, X., Krishnan, A., Meer, M., Wang, C., et al. (2020). Reprogramming to recover youthful epigenetic information and restore vision. Nature 588, 124–129. doi: 10.1038/s41586-020-2975-4
Lu, P., Ceto, S., Wang, Y., Graham, L., Wu, D., Kumamaru, H., et al. (2017). Prolonged human neural stem cell maturation supports recovery in injured rodent CNS. J. Clin. Invest. 127, 3287–3299. doi: 10.1172/JCI92955
Lu, P., Kadoya, K., and Tuszynski, M. H. (2014a). Axonal growth and connectivity from neural stem cell grafts in models of spinal cord injury. Curr. Opin. Neurobiol. 27, 103–109. doi: 10.1016/j.conb.2014.03.010
Lu, P., Wang, Y., Graham, L., McHale, K., Gao, M., Wu, D., et al. (2012). Long-distance growth and connectivity of neural stem cells after severe spinal cord injury. Cells 150, 1264–1273. doi: 10.1016/j.cell.2012.08.020
Lu, P., Woodruff, G., Wang, Y., Graham, L., Hunt, M., Wu, D., et al. (2014b). Long-distance axonal growth from human induced pluripotent stem cells after spinal cord injury. Neuron 83, 789–796. doi: 10.1016/j.neuron.2014.07.014
Ma, X. M., and Blenis, J. (2009). Molecular mechanisms of mTOR-mediated translational control. Nat. Rev. Mol. Cell Biol. 10, 307–318. doi: 10.1038/nrm2672
Mar, F. M., Bonni, A., and Sousa, M. M. (2014). Cell intrinsic control of axon regeneration. EMBO Rep. 15, 254–263. doi: 10.1002/embr.201337723
McKeon, R. J., Jurynec, M. J., and Buck, C. R. (1999). The chondroitin sulfate proteoglycans neurocan and phosphacan are expressed by reactive astrocytes in the chronic CNS glial scar. J. Neurosci. 19, 10778–10788. doi: 10.1523/JNEUROSCI.19-24-10778.1999
Mehta, S. T., Luo, X., Park, K. K., Bixby, J. L., and Lemmon, V. P. (2016). Hyperactivated Stat3 boosts axon regeneration in the CNS. Exp. Neurol. 280, 115–120. doi: 10.1016/j.expneurol.2016.03.004
Mertens, J., Herdy, J. R., Traxler, L., Schafer, S. T., Schlachetzki, J. C. M., Bohnke, L., et al. (2021). Age-dependent instability of mature neuronal fate in induced neurons from Alzheimer’s patients. Cell Stem Cell 28, 1533–1548.e6. doi: 10.1016/j.stem.2021.04.004
Mertens, J., Paquola, A. C. M., Ku, M., Hatch, E., Bohnke, L., Ladjevardi, S., et al. (2015). Directly reprogrammed human neurons retain aging-associated transcriptomic signatures and reveal age-related nucleocytoplasmic defects. Cell Stem Cell 17, 705–718. doi: 10.1016/j.stem.2015.09.001
Mertens, J., Reid, D., Lau, S., Kim, Y., and Gage, F. H. (2018). Aging in a dish: iPSC-derived and directly induced neurons for studying brain aging and age-related neurodegenerative diseases. Annu. Rev. Genet. 52, 271–293. doi: 10.1146/annurev-genet-120417-031534
Moore, D. L., Blackmore, M. G., Hu, Y., Kaestner, K. H., Bixby, J. L., Lemmon, V. P., et al. (2009). KLF family members regulate intrinsic axon regeneration ability. Science 326, 298–301. doi: 10.1126/science.1175737
Muller, F., De Virgiliis, F., Kong, G., Zhou, L., Serger, E., Chadwick, J., et al. (2022). CBP/p300 activation promotes axon growth, sprouting, and synaptic plasticity in chronic experimental spinal cord injury with severe disability. PLoS Biol. 20:e3001310. doi: 10.1371/journal.pbio.3001310
Nagoshi, N., and Okano, H. (2017). Applications of induced pluripotent stem cell technologies in spinal cord injury. J. Neurochem. 141, 848–860. doi: 10.1111/jnc.13986
Nathan, F. M., Ohtake, Y., Wang, S., Jiang, X., Sami, A., Guo, H., et al. (2020). Upregulating Lin28a promotes axon regeneration in adult mice with optic nerve and spinal cord injury. Mol. Ther. 28, 1902–1917. doi: 10.1016/j.ymthe.2020.04.010
Nori, S., Okada, Y., Yasuda, A., Tsuji, O., Takahashi, Y., Kobayashi, Y., et al. (2011). Grafted human-induced pluripotent stem-cell-derived neurospheres promote motor functional recovery after spinal cord injury in mice. Proc. Natl. Acad. Sci. U. S. A. 108, 16825–16830. doi: 10.1073/pnas.1108077108
Nowakowski, T. J., Bhaduri, A., Pollen, A. A., Alvarado, B., Mostajo-Radji, M. A., Di Lullo, E., et al. (2017). Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323. doi: 10.1126/science.aap8809
O’Donovan, K. J., Ma, K., Guo, H., Wang, C., Sun, F., Han, S. B., et al. (2014). B-RAF kinase drives developmental axon growth and promotes axon regeneration in the injured mature CNS. J. Exp. Med. 211, 801–814. doi: 10.1084/jem.20131780
Oh, Y. M., Lee, S. W., Kim, W. K., Chen, S., Church, V. A., Cates, K., et al. (2022). Age-related Huntington’s disease progression modeled in directly reprogrammed patient-derived striatal neurons highlights impaired autophagy. Nat. Neurosci. 25, 1420–1433. doi: 10.1038/s41593-022-01185-4
Ohtake, Y., Park, D., Abdul-Muneer, P. M., Li, H., Xu, B., Sharma, K., et al. (2014). The effect of systemic PTEN antagonist peptides on axon growth and functional recovery after spinal cord injury. Biomaterials 35, 4610–4626. doi: 10.1016/j.biomaterials.2014.02.037
Okubo, T., Nagoshi, N., Kohyama, J., Tsuji, O., Shinozaki, M., Shibata, S., et al. (2018). Treatment with a gamma-secretase inhibitor promotes functional recovery in human iPSC- derived transplants for chronic spinal cord injury. Stem Cell Rep. 11, 1416–1432. doi: 10.1016/j.stemcr.2018.10.022
Pabba, M., Scifo, E., Kapadia, F., Nikolova, Y. S., Ma, T., Mechawar, N., et al. (2017). Resilient protein co-expression network in male orbitofrontal cortex layer 2/3 during human aging. Neurobiol. Aging 58, 180–190. doi: 10.1016/j.neurobiolaging.2017.06.023
Pang, Z. P., Yang, N., Vierbuchen, T., Ostermeier, A., Fuentes, D. R., Yang, T. Q., et al. (2011). Induction of human neuronal cells by defined transcription factors. Nature 476, 220–223. doi: 10.1038/nature10202
Park, K. K., Liu, K., Hu, Y., Kanter, J. L., and He, Z. (2010). PTEN/mTOR and axon regeneration. Exp. Neurol. 223, 45–50. doi: 10.1016/j.expneurol.2009.12.032
Park, K. K., Liu, K., Hu, Y., Smith, P. D., Wang, C., Cai, B., et al. (2008). Promoting axon regeneration in the adult CNS by modulation of the PTEN/mTOR pathway. Science 322, 963–966. doi: 10.1126/science.1161566
Perrin, S. (2014). Preclinical research: make mouse studies work. Nature 507, 423–425. doi: 10.1038/507423a
Pfisterer, U., Kirkeby, A., Torper, O., Wood, J., Nelander, J., Dufour, A., et al. (2011). Direct conversion of human fibroblasts to dopaminergic neurons. Proc. Natl. Acad. Sci. U. S. A. 108, 10343–10348. doi: 10.1073/pnas.1105135108
Piao, J., Zabierowski, S., Dubose, B. N., Hill, E. J., Navare, M., Claros, N., et al. (2021). Preclinical efficacy and safety of a human embryonic stem cell-derived midbrain dopamine progenitor product, MSK-DA01. Cell Stem Cell 28, 217–229.e7. doi: 10.1016/j.stem.2021.01.004
Pita-Thomas, W., Goncalves, T. M., Kumar, A., Zhao, G., and Cavalli, V. (2021). Genome-wide chromatin accessibility analyses provide a map for enhancing optic nerve regeneration. Sci. Rep. 11:14924. doi: 10.1038/s41598-021-94341-y
Pletikos, M., Sousa, A. M., Sedmak, G., Meyer, K. A., Zhu, Y., Cheng, F., et al. (2014). Temporal specification and bilaterality of human neocortical topographic gene expression. Neuron 81, 321–332. doi: 10.1016/j.neuron.2013.11.018
Poplawski, G. H. D., Kawaguchi, R., Van Niekerk, E., Lu, P., Mehta, N., Canete, P., et al. (2020). Injured adult neurons regress to an embryonic transcriptional growth state. Nature 581, 77–82. doi: 10.1038/s41586-020-2200-5
Qin, S., Zou, Y., and Zhang, C. L. (2013). Cross-talk between KLF4 and STAT3 regulates axon regeneration. Nat. Commun. 4:2633. doi: 10.1038/ncomms3633
Ramos, S. I., Mussa, Z. M., Falk, E. N., Pai, B., Giotti, B., Allette, K., et al. (2022). An atlas of late prenatal human neurodevelopment resolved by single-nucleus transcriptomics. Nat. Commun. 13:7671. doi: 10.1038/s41467-022-34975-2
Redmond, L., Kashani, A. H., and Ghosh, A. (2002). Calcium regulation of dendritic growth via CaM kinase IV and CREB-mediated transcription. Neuron 34, 999–1010. doi: 10.1016/S0896-6273(02)00737-7
Rim, Y. A., Park, N., Nam, Y., Ham, D. S., Kim, J. W., Ha, H. Y., et al. (2018). Recent progress of national banking project on homozygous HLA-typed induced pluripotent stem cells in South Korea. J. Tissue Eng. Regen. Med. 12, e1531–e1536. doi: 10.1002/term.2578
Rosenzweig, E. S., Brock, J. H., Lu, P., Kumamaru, H., Salegio, E. A., Kadoya, K., et al. (2018). Restorative effects of human neural stem cell grafts on the primate spinal cord. Nat. Med. 24, 484–490. doi: 10.1038/nm.4502
Safaiyan, S., Kannaiyan, N., Snaidero, N., Brioschi, S., Biber, K., Yona, S., et al. (2016). Age-related myelin degradation burdens the clearance function of microglia during aging. Nat. Neurosci. 19, 995–998. doi: 10.1038/nn.4325
Savell, K. E., Sultan, F. A., and Day, J. J. (2019). A novel dual lentiviral CRISPR-based transcriptional activation system for gene expression regulation in neurons. Biol. Protoc. 9:e3348. doi: 10.21769/BioProtoc.3348
Saxton, R. A., and Sabatini, D. M. (2017). mTOR Signaling in growth, metabolism, and disease. Cells 169, 361–371. doi: 10.1016/j.cell.2017.03.035
Seo, W. M., Yoon, J., Lee, J. H., Lee, Y., Lee, H., Geum, D., et al. (2022). Modeling axonal regeneration by changing cytoskeletal dynamics in stem cell-derived motor nerve organoids. Sci. Rep. 12:2082. doi: 10.1038/s41598-022-05645-6
Shamblott, M. J., Axelman, J., Wang, S., Bugg, E. M., Littlefield, J. W., Donovan, P. J., et al. (1998). Derivation of pluripotent stem cells from cultured human primordial germ cells. Proc. Natl. Acad. Sci. U. S. A. 95, 13726–13731. doi: 10.1073/pnas.95.23.13726
Sherman, S. P., and Bang, A. G. (2018). High-throughput screen for compounds that modulate neurite growth of human induced pluripotent stem cell-derived neurons. Dis. Model. Mech. 11. doi: 10.1242/dmm.031906
Shin, H. S., Kim, H. J., Min, S. K., Kim, S. H., Lee, B. M., and Jeon, N. L. (2010). Compartmental culture of embryonic stem cell-derived neurons in microfluidic devices for use in axonal biology. Biotechnol. Lett. 32, 1063–1070. doi: 10.1007/s10529-010-0280-2
Siegenthaler, M. M., Ammon, D. L., and Keirstead, H. S. (2008). Myelin pathogenesis and functional deficits following SCI are age-associated. Exp. Neurol. 213, 363–371. doi: 10.1016/j.expneurol.2008.06.015
Sirenko, O., Hesley, J., Rusyn, I., and Cromwell, E. F. (2014). High-content high-throughput assays for characterizing the viability and morphology of human iPSC-derived neuronal cultures. Assay Drug Dev. Technol. 12, 536–547. doi: 10.1089/adt.2014.592
Snow, D. M., Lemmon, V., Carrino, D. A., Caplan, A. I., and Silver, J. (1990). Sulfated proteoglycans in astroglial barriers inhibit neurite outgrowth in vitro. Exp. Neurol. 109, 111–130. doi: 10.1016/S0014-4886(05)80013-5
Son, E. Y., Ichida, J. K., Wainger, B. J., Toma, J. S., Rafuse, V. F., Woolf, C. J., et al. (2011). Conversion of mouse and human fibroblasts into functional spinal motor neurons. Cell Stem Cell 9, 205–218. doi: 10.1016/j.stem.2011.07.014
Stewart, A. N., Lowe, J. L., Glaser, E. P., Mott, C. A., Shahidehpour, R. K., McFarlane, K. E., et al. (2021). Acute inflammatory profiles differ with sex and age after spinal cord injury. J. Neuroinflamm. 18:113. doi: 10.1186/s12974-021-02161-8
Sugai, K., Sumida, M., Shofuda, T., Yamaguchi, R., Tamura, T., Kohzuki, T., et al. (2021). First-in-human clinical trial of transplantation of iPSC-derived NS/PCs in subacute complete spinal cord injury: study protocol. Regen. Ther. 18, 321–333. doi: 10.1016/j.reth.2021.08.005
Sun, F., Park, K. K., Belin, S., Wang, D., Lu, T., Chen, G., et al. (2011). Sustained axon regeneration induced by co-deletion of PTEN and SOCS3. Nature 480, 372–375. doi: 10.1038/nature10594
Takahashi, K., Tanabe, K., Ohnuki, M., Narita, M., Ichisaka, T., Tomoda, K., et al. (2007). Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cells 131, 861–872. doi: 10.1016/j.cell.2007.11.019
Tanabe, K., Ang, C. E., Chanda, S., Olmos, V. H., Haag, D., Levinson, D. F., et al. (2018). Transdifferentiation of human adult peripheral blood T cells into neurons. Proc. Natl. Acad. Sci. U. S. A. 115, 6470–6475. doi: 10.1073/pnas.1720273115
Tang, X., Davies, J. E., and Davies, S. J. (2003). Changes in distribution, cell associations, and protein expression levels of NG2, neurocan, phosphacan, brevican, versican V2, and tenascin-C during acute to chronic maturation of spinal cord scar tissue. J. Neurosci. Res. 71, 427–444. doi: 10.1002/jnr.10523
Tang, Y., Liu, M. L., Zang, T., and Zhang, C. L. (2017). Direct reprogramming rather than iPSC-based reprogramming maintains aging hallmarks in human motor neurons. Front. Mol. Neurosci. 10:359. doi: 10.3389/fnmol.2017.00359
Tao, Y., Vermilyea, S. C., Zammit, M., Lu, J., Olsen, M., Metzger, J. M., et al. (2021). Autologous transplant therapy alleviates motor and depressive behaviors in parkinsonian monkeys. Nat. Med. 27, 632–639. doi: 10.1038/s41591-021-01257-1
Tao, Y., and Zhang, S. C. (2016). Neural subtype specification from human pluripotent stem cells. Cell Stem Cell 19, 573–586. doi: 10.1016/j.stem.2016.10.015
Teotia, P., Van Hook, M. J., Fischer, D., and Ahmad, I. (2019). Human retinal ganglion cell axon regeneration by recapitulating developmental mechanisms: effects of recruitment of the mTOR pathway. Development 146. doi: 10.1242/dev.178012
Thomson, J. A., Itskovitz-Eldor, J., Shapiro, S. S., Waknitz, M. A., Swiergiel, J. J., et al. (1998). Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147. doi: 10.1126/science.282.5391.1145
Trakhtenberg, E. F., Li, Y., Feng, Q., Tso, J., Rosenberg, P. A., Goldberg, J. L., et al. (2018). Zinc chelation and Klf9 knockdown cooperatively promote axon regeneration after optic nerve injury. Exp. Neurol. 300, 22–29. doi: 10.1016/j.expneurol.2017.10.025
Usvald, D., Vodicka, P., Hlucilova, J., Prochazka, R., Motlik, J., Kuchorova, K., et al. (2010). Analysis of dosing regimen and reproducibility of intraspinal grafting of human spinal stem cells in immunosuppressed minipigs. Cell Transplant. 19, 1103–1122. doi: 10.3727/096368910X503406
Vadodaria, K. C., Mertens, J., Paquola, A., Bardy, C., Li, X., Jappelli, R., et al. (2016). Generation of functional human serotonergic neurons from fibroblasts. Mol. Psychiatry 21, 49–61. doi: 10.1038/mp.2015.161
van Gorp, S., Leerink, M., Kakinohana, O., Platoshyn, O., Santucci, C., Galik, J., et al. (2013). Amelioration of motor/sensory dysfunction and spasticity in a rat model of acute lumbar spinal cord injury by human neural stem cell transplantation. Stem Cell Res Ther 4:57. doi: 10.1186/scrt209
Vasan, L., Park, E., David, L. A., Fleming, T., and Schuurmans, C. (2021). Direct neuronal reprogramming: bridging the gap between basic science and clinical application. Front. Cell Dev. Biol. 9:681087. doi: 10.3389/fcell.2021.681087
Venkatesh, I., Mehra, V., Wang, Z., Califf, B., and Blackmore, M. G. (2018). Developmental chromatin restriction of pro-growth gene networks acts as an epigenetic barrier to axon regeneration in cortical neurons. Dev. Neurobiol. 78, 960–977. doi: 10.1002/dneu.22605
Venkatesh, I., Simpson, M. T., Coley, D. M., and Blackmore, M. G. (2016). Epigenetic profiling reveals a developmental decrease in promoter accessibility during cortical maturation in vivo. Neuroepigenetics 8, 19–26. doi: 10.1016/j.nepig.2016.10.002
Victor, M. B., Richner, M., Hermanstyne, T. O., Ransdell, J. L., Sobieski, C., Deng, P. Y., et al. (2014). Generation of human striatal neurons by microRNA-dependent direct conversion of fibroblasts. Neuron 84, 311–323. doi: 10.1016/j.neuron.2014.10.016
Victor, M. B., Richner, M., Olsen, H. E., Lee, S. W., Monteys, A. M., Ma, C., et al. (2018). Striatal neurons directly converted from Huntington’s disease patient fibroblasts recapitulate age-associated disease phenotypes. Nat. Neurosci. 21, 341–352. doi: 10.1038/s41593-018-0075-7
Wan, X. Y., Xu, L. Y., Li, B., Sun, Q. H., Ji, Q. L., Huang, D. D., et al. (2018). Chemical conversion of human lung fibroblasts into neuronal cells. Int. J. Mol. Med. 41, 1463–1468. doi: 10.3892/ijmm.2018.3375
Wang, J., He, Q., Zhang, K., Sun, H., Zhang, G., Liang, H., et al. (2020). Quick commitment and efficient reprogramming route of direct induction of retinal ganglion cell-like neurons. Stem Cell Rep. 15, 1095–1110. doi: 10.1016/j.stemcr.2020.09.008
Wang, J. T., Kunzevitzky, N. J., Dugas, J. C., Cameron, M., Barres, B. A., and Goldberg, J. L. (2007). Disease gene candidates revealed by expression profiling of retinal ganglion cell development. J. Neurosci. 27, 8593–8603. doi: 10.1523/JNEUROSCI.4488-06.2007
Wang, Z., Mehra, V., Simpson, M. T., Maunze, B., Chakraborty, A., Holan, L., et al. (2018). KLF6 and STAT3 co-occupy regulatory DNA and functionally synergize to promote axon growth in CNS neurons. Sci. Rep. 8:12565. doi: 10.1038/s41598-018-31101-5
Wang, H., Yang, Y., Liu, J., and Qian, L. (2021). Direct cell reprogramming: approaches, mechanisms and progress. Nat. Rev. Mol. Cell Biol. 22, 410–424. doi: 10.1038/s41580-021-00335-z
Wang, Y. K., Zhu, W. W., Wu, M. H., Wu, Y. H., Liu, Z. X., Liang, L. M., et al. (2018). Human clinical-Grade parthenogenetic ESC-derived dopaminergic neurons recover locomotive defects of nonhuman primate models of Parkinson’s disease. Stem Cell Rep. 11, 171–182. doi: 10.1016/j.stemcr.2018.05.010
Waxman, S. G., and Foster, R. E. (1980). Development of the axon membrane during differentiation of myelinated fibres in spinal nerve roots. Proc. R. Soc. Lond. B Biol. Sci. 209, 441–446. doi: 10.1098/rspb.1980.0105
Weickert, C. S., Elashoff, M., Richards, A. B., Sinclair, D., Bahn, S., Paabo, S., et al. (2009). Transcriptome analysis of male-female differences in prefrontal cortical development. Mol. Psychiatry 14, 558–561. doi: 10.1038/mp.2009.5
Weng, Y. L., Wang, X., An, R., Cassin, J., Vissers, C., Liu, Y., et al. (2018). Epitranscriptomic m(6)a regulation of axon regeneration in the adult mammalian nervous system. Neuron 97:e6, 313–325.e6. doi: 10.1016/j.neuron.2017.12.036
Werling, D. M., Pochareddy, S., Choi, J., An, J. Y., Sheppard, B., Peng, M., et al. (2020). Whole-genome and RNA sequencing reveal variation and transcriptomic coordination in the developing human prefrontal cortex. Cell Rep. 31:107489. doi: 10.1016/j.celrep.2020.03.053
Wertheim, L., Edri, R., Goldshmit, Y., Kagan, T., Noor, N., Ruban, A., et al. (2022). Regenerating the injured spinal cord at the chronic phase by engineered iPSCs-derived 3D neuronal networks. Adv. Sci. (Weinh) 9:e2105694. doi: 10.1002/advs.202105694
Wingo, A. P., Dammer, E. B., Breen, M. S., Logsdon, B. A., Duong, D. M., Troncosco, J. C., et al. (2019). Large-scale proteomic analysis of human brain identifies proteins associated with cognitive trajectory in advanced age. Nat. Commun. 10:1619. doi: 10.1038/s41467-019-09613-z
Xie, L., Cen, L. P., Li, Y., Gilbert, H. Y., Strelko, O., Berlinicke, C., et al. (2022). Monocyte-derived SDF1 supports optic nerve regeneration and alters retinal ganglion cells’ response to Pten deletion. Proc. Natl. Acad. Sci. U. S. A. 119:e2113751119. doi: 10.1073/pnas.2113751119
Xiong, M., Tao, Y., Gao, Q., Feng, B., Yan, W., Zhou, Y., et al. (2021). Human stem cell-derived neurons repair circuits and restore neural function. Cell Stem Cell 28, 112–126.e6. doi: 10.1016/j.stem.2020.08.014
Xu, Z., Jiang, H., Zhong, P., Yan, Z., Chen, S., and Feng, J. (2016). Direct conversion of human fibroblasts to induced serotonergic neurons. Mol. Psychiatry 21, 62–70. doi: 10.1038/mp.2015.101
Xu, J. H., Qin, X. Z., Zhang, H. N., Ma, Y. X., Qi, S. B., Zhang, H. C., et al. (2021). Deletion of Kruppel-like factor-4 promotes axonal regeneration in mammals. Neural Regen. Res. 16, 166–171. doi: 10.4103/1673-5374.286978
Xu, Z., Su, S., Zhou, S., Yang, W., Deng, X., Sun, Y., et al. (2020). How to reprogram human fibroblasts to neurons. Cell Biosci. 10:116. doi: 10.1186/s13578-020-00476-2
Yang, Y., Chen, R., Wu, X., Zhao, Y., Fan, Y., Xiao, Z., et al. (2019). Rapid and efficient conversion of human fibroblasts into functional neurons by small molecules. Stem Cell Rep. 13, 862–876. doi: 10.1016/j.stemcr.2019.09.007
Yang, L., Miao, L., Liang, F., Huang, H., Teng, X., Li, S., et al. (2014). The mTORC1 effectors S6K1 and 4E-BP play different roles in CNS axon regeneration. Nat. Commun. 5:5416. doi: 10.1038/ncomms6416
Yiu, G., and He, Z. (2006). Glial inhibition of CNS axon regeneration. Nat. Rev. Neurosci. 7, 617–627. doi: 10.1038/nrn1956
Yoo, A. S., Sun, A. X., Li, L., Shcheglovitov, A., Portmann, T., Li, Y., et al. (2011). MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476, 228–231. doi: 10.1038/nature10323
Yoshida, S., Kato, T. M., Sato, Y., Umekage, M., Ichisaka, T., Tsukahara, M., et al. (2023). A clinical-grade HLA haplobank of human induced pluripotent stem cells matching approximately 40% of the Japanese population. Medicine 4:e10, 51–66.e10. doi: 10.1016/j.medj.2022.10.003
Yu, J., Vodyanik, M. A., Smuga-Otto, K., Antosiewicz-Bourget, J., Frane, J. L., Tian, S., et al. (2007). Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920. doi: 10.1126/science.1151526
Yue, F., Cheng, Y., Breschi, A., Vierstra, J., Wu, W., Ryba, T., et al. (2014). A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355–364. doi: 10.1038/nature13992
Zhang, L., Yin, J. C., Yeh, H., Ma, N. X., Lee, G., Chen, X. A., et al. (2015). Small molecules efficiently reprogram human Astroglial cells into functional neurons. Cell Stem Cell 17, 735–747. doi: 10.1016/j.stem.2015.09.012
Zheng, B., and Tuszynski, M. H. (2023). Regulation of axonal regeneration after mammalian spinal cord injury. Nat. Rev. Mol. Cell Biol. 24, 396–413. doi: 10.1038/s41580-022-00562-y
Zhu, Y., Sousa, A. M. M., Gao, T., Skarica, M., Li, M., Santpere, G., et al. (2018). Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362. doi: 10.1126/science.aat8077
Keywords: axon growth, axon regeneration, direct reprogramming, hPSCs, reprogramming, development
Citation: Lear BP and Moore DL (2023) Moving CNS axon growth and regeneration research into human model systems. Front. Neurosci. 17:1198041. doi: 10.3389/fnins.2023.1198041
Edited by:
Inês Dinis Aires, University of Coimbra, PortugalReviewed by:
Elisa Waxman, Children’s Hospital of Philadelphia, United StatesCopyright © 2023 Lear and Moore. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Darcie L. Moore, ZGFyY2llLm1vb3JlQHdpc2MuZWR1
 Bo P. Lear
Bo P. Lear Darcie L. Moore
Darcie L. Moore