- 1Key Laboratory of Neuroregeneration of Jiangsu and Ministry of Education, Co-innovation Center of Neurogeneration, NMPA Key Laboratory for Research and Evaluation of Tissue Engineering Technology Products, Nantong University, Nantong, China
- 2Department of Endocrinology, Changshu No.2 People’s Hospital, Changshu, Jiangsu, China
Gluconeogenesis is the main process for endogenous glucose production during prolonged fasting, or certain pathological conditions, which occurs primarily in the liver. Hepatic gluconeogenesis is a biochemical process that is finely controlled by hormones such as insulin and glucagon, and it is of great importance for maintaining normal physiological blood glucose levels. Dysregulated gluconeogenesis induced by obesity is often associated with hyperglycemia, hyperinsulinemia, and type 2 diabetes (T2D). Long noncoding RNAs (lncRNAs) are involved in various cellular events, from gene transcription to protein translation, stability, and function. In recent years, a growing number of evidences has shown that lncRNAs play a key role in hepatic gluconeogenesis and thereby, affect the pathogenesis of T2D. Here we summarized the recent progress in lncRNAs and hepatic gluconeogenesis.
1 Introduction
Gluconeogenesis is an important biochemical process for maintaining glucose homeostasis in mammals, in which glucose is produced from non-carbohydrate substrates including lactate, pyruvate, propionate, glycerol, and amino acids. In general, gluconeogenesis will be activated when the blood glucose level is very low, often under fasting or starvation conditions. In the body of mammals, gluconeogenesis mainly occurs in the liver, though it may also take place in fewer amounts in the kidney and small intestine. In normal conditions, hepatic gluconeogenesis is finely tuned by hormones including insulin and glucagon and thus keeps blood glucose within physiological concentrations (1, 2). Insulin is a negative regulator of gluconeogenesis, while glucagon is a positive regulator (1, 2). Chronic ectopic increased gluconeogenesis is often associated with metabolic syndromes such as hyperglycemia, hyperinsulinemia, insulin resistance, and type 2 diabetes (T2D) (3). On the contrary, impaired gluconeogenesis may cause hypoglycemia and a shortage of energy supply, leading to dizziness, memory loss, or coma (4).
Human genomics research data revealed that 84% of the human genome could be transcribed, only 2% of which can encode proteins after transcription (5). Therefore, RNAs not only act as carriers of genetic information but also play a variety of regulatory functions. RNAs without protein coding capacity are called non-coding RNAs (ncRNAs), including microRNAs, long-chain noncoding RNAs (lncRNAs), small nucleolar RNAs (snoRNAs), and circular RNAs (circRNAs) (6, 7). Traditionally, noncoding RNA molecules with a length greater than 200 nt have been defined as lncRNAs (8). Accumulating evidence has shown that lncRNAs play a pivotal role in many cellular events, such as cell division, differentiation, migration, and apoptosis (9, 10). Each lncRNA has its own tissue-specific expression pattern, which defines its respective unique function (10, 11). For example, lncRNA-Bvht is a heart-associated lncRNA in mouse, which is enriched in embryonic stem cells (ESCs) and plays a key role in cardiomyocyte differentiation (12). Upon starvation or cold stimulation, some lncRNAs in adipose tissues are transcribed and participate in lipolysis and thermogenesis, which may hold therapeutic potential for treating metabolic diseases such as obesity and diabetes (13). Similarly, hepatocytes also have a unique lncRNA expression profile, which changes during liver development and regulates liver maturation (14). RNA-seq revealed 104 differentially expressed lncRNAs in the liver of T2D rats and bioinformatics analysis suggest these lncRNAs may correlate with the pathogenesis of T2D by affecting lipid metabolism, gluconeogenesis, inflammation, and/or endoplasmic reticulum stress (15). Moreover, metformin treatment induced a number of differentially expressed lncRNAs in the liver of mice, implying lncRNAs are involved in hepatic gluconeogenesis since the promising inhibitory effect of metformin on gluconeogenesis (16). LincIRS2 is an obesity-repressed lncRNA in the liver, its deficiency elevated blood glucose, promoted insulin resistance, and induced glucose output in mice (17). All these data strongly indicate the potential impacts of lncRNAs on hepatic gluconeogenesis. In this review, therefore, we summarized the recent progress regarding the roles of lncRNAs on hepatic gluconeogenesis. Meanwhile, we also discussed the therapeutic potentials of lncRNAs in ectopic gluconeogenesis-associated metabolic disorders such as insulin resistance and T2D.
2 Hepatic gluconeogenesis in health and diseases
As the main energy source of mammals, glucose is necessary for maintaining normal physiological functions for the central nervous system (CNS), retina, and red blood cells (18–20). Adults with normal body weight consume 160 g of glucose per day. Of them, the most glucose (∼120 g) was consumed by the brain (21). In addition to exogenous glucose obtained from food, most the endogenous glucose is stored as glycogen in various organs such as the liver and muscle. In humans, only approximately 15 g of glucose is available for consumption in extracellular fluid, termed blood glucose. The balance between endogenous glucose production and peripheral glucose uptake helps to maintain systemic glucose homeostasis (22). Under normal physiological conditions, the concentration of blood glucose is dynamically constant. Low blood glucose concentration causes fatigue, dizziness, and irreversible damage to the CNS (23, 24). Likewise, long-term high blood glucose is often associated with metabolic dysfunctions, including hyperglycemia, insulin resistance, and T2D. Therefore, blood glucose concentration is an essential indicator of glucose homeostasis, and the dynamic balance of exogenous glucose supply and endogenous glucose production is a key node for maintaining normal physiological blood glucose (25). The liver is an essential organ for keeping blood sugar balance. Hepatic glucose metabolism includes multiple pathways of decomposition and anabolism. Glycogen synthesis, glycogenolysis, gluconeogenesis, and glycolysis jointly determine the stability of blood glucose. Hepatic cells regulate the dynamic balance of glucose metabolism in response to environmental and nutritional changes in an autonomous or involuntary manner. Hepatic gluconeogenesis, as the main way of endogenous glucose production during starvation, plays an important role in maintaining blood glucose balance.
3 Overview of lncRNAs
3.1 Biogenesis and classification of lncRNAs
LncRNAs are mainly transcribed by RNA polymerase II and generally contain a 5’-m7G cap and a 3’-poly (A) tail (26, 27). lncRNAs were traditionally thought not to possess the ability to encode proteins; however, some lncRNAs, such as LINC00998, LINC00961, LINC00467, and LINC-PINT, have been found to encode specific small polypeptides (28–32). Human GENCODE database shows that more than 173,000 lncRNA transcripts were identified in the human genome (33). Of them, only a small part of lncRNAs has been functionally annotated, while a large number of lncRNAs remain to be determined (34, 35). Most lncRNAs are localized in the nucleus, while some lncRNAs are located in the cytoplasm or other sub-organelles, such as ribosomes and mitochondria (10). Compared with protein-coding genes, lncRNA genes are generally less conserved with lower expression profiles (36, 37). Despite sharing similar patterns of splicing, export, and quality control with mRNAs, most lncRNAs are trapped in the nucleus. In comparison to mRNAs, lncRNAs have fewer and longer exons, and for this reason, lncRNAs prefer the NXF1/NXT1 pathway for nuclear export (38). Furthermore, lncRNAs have lower splicing efficiency but higher splicing frequencies to increase their numbers (39).
Based on genomic location and functioning mechanism, lncRNAs are divided into five groups, intergenic lncRNAs (lincRNAs), intronic lncRNAs, sense lncRNAs, antisense lncRNAs, and bidirectional lncRNAs (40). A large number of non-coding regions are distributed among the coding regions of the human genome, accounting for 98-99%. The lncRNAs transcribed from these non-coding regions are called intergenic lncRNAs. LncRNAs transcribed from introns in the coding region are named intronic lncRNAs. Sense and antisense lncRNAs are transcribed from sense and antisense strands coding proteins, respectively. Currently, most studies are focused on lincRNAs and antisense lncRNAs. LincRNA shows functional importance due to its high active transcription, a certain degree of domain conservation, tissue-specific expression, and stability, whereas antisense lncRNA accounts for a large amount of human lncRNA (40).
3.2 Functioning mechanisms for lncRNAs
By various regulation models, lncRNAs can positively or negatively control coding gene expression, which could be occurred at different stages of eukaryotic gene expression (41). At the chromatin level, lncRNAs induce chromatin epigenetic modification to affect conformational structures of chromatin and thereby control gene expression. LncRNAs can regulate DNA methylation by recruiting DNMTs/TETs, sequestering DNMTs, or regulating the expression of DNMTs/TETs (42). Alternatively, lncRNAs act as decoys to sequester chromatin modifiers from specific genomic sites to induce chromatin remodeling (43). At the transcriptional level, lncRNAs can mediate gene silence or activation. For example, Airn, an antisense transcript of the Iinsulin-like growth factor 2 receptor (gf2r) gene, whose transcription causes Pol II to detach from the Igf2r promoter, resulting in transcriptional pause and gene silencing (44). At the post-transcriptional level, LncRNAs are involved in the post-transcriptional splicing of mRNAs. LncRNA Ctcflos mediates the selective splicing of PRDM16 to generate short isomers with a preference for thermogenesis, thereby promoting fat thermogenesis (45). Moreover, lncRNAs may also regulate gene expression by other means. For example, lncRNAs fold into higher-order structure to bind nucleoprotein and assemble ribonucleoprotein complex to participate in protein nuclear localization, or lncRNAs pair with other RNAs to recruit protein complexes or adsorb microRNAs to regulate gene silencing (46).
4 LncRNAs in hepatic gluconeogenesis
4.1 LncH19
H19 is the first lncRNA originally found in the liver extract, which is 2.3 kb in length and located on chromosome 11. After transcription, lncH19 is exported to the cytoplasm after a similar modification process as mRNAs, such as splicing, capping, and polyadenylation (47). It is enriched in embryonic stem cells and remains highly expressed in the adrenal gland, liver, and adipose tissue after birth (48). LncH19 loses the ability to translate into small peptides due to the special structure of the 5’-terminal (49). Therefore, lncH19 plays a role as an independent functional unit. A clinical study of obese women showed that human linH19 transcription levels were negatively correlated with body mass index (BMI) and homeostatic model assessment of insulin resistance (HOMA-IR) (50). Moreover, H19 has been shown to regulate glucose homeostasis and β cell function (51).
By RNA-seq, Goyal N et al. found that H19 was largely decreased in the liver of diabetic db/db mice, suggesting its potential role in glucose metabolism. In the following experiments, their functional studies showed that the knockdown of H19 stimulates gluconeogenic gene expression and hepatic glucose output in HepG2 cells and primary mouse hepatocytes (52). In vivo studies have shown that, in healthy mice, H19 absence results in dysregulated glucose metabolism including hyperglycemia, hyperinsulinemia, and intolerant insulin, glucose, and pyruvate tests (53). Mechanistically, H19 silencing increases the occupancy of P53 in the promoter of Foxo1, which promotes the transcription of Foxo1, a master regulator of gluconeogenic gene expression (52, 53). However, this view regarding the roles of H19 in gluconeogenesis has been challenged. Deng J et al. have shown that overexpression of H19 in a human liver cell line activates the gluconeogenic program, which is likely due to increased expression of HNF4α (54). Most recently, one report confirmed that overexpression of H19 in Hepa1-6 cells increases Pck1 expression and gluconeogenesis by inducing the nuclear retention of FOXO1 (55). Also, in this study, H19 was identified as an imprinted gene for transducing hyperglycemia from paternal obesity to female offspring (55). Therefore, due to these inconsistent findings, the precise functions of H19 on gluconeogenesis are not clear. More studies are required to clarify this issue.
4.2 LncSHGL
Mouse lncSHGL is located on chromosome 17, and its homologous in humans is lncRNA B4GALT1-AS1. LncSHGL was low expressed in the liver of obese mice, and similarly, lncRNA B4GALT1-AS1 was significantly decreased in patients with nonalcoholic fatty liver disease (56). It has been shown that restoration of hepatic lncSHGL plays a beneficial role against hyperglycemia, insulin resistance, and hepatic steatosis in diabetic mice, while inhibition of lncSHGL worsens hyperglycemia and lipid deposition in livers (56). Mechanistic studies revealed that lncSHGL increases calmodulin (CaM) mRNA translation by recruiting heterogeneous nuclear ribonucleoprotein A1 (hnRNPA1). As a result, increased CaM suppresses gluconeogenic and lipogenic pathways in hepatocytes (56).
4.3 LncMEG3
Maternal expression gene 3 (MEG3) is an imprinted gene located on the human chromosome 14q32. It is the ortholog of the gene trap locus 2 (Gtl2) on mouse chromosome 12. LncMEG3 is generally considered to be a tumor suppressor, which is expressed in a variety of tissues and encodes lncRNAs associated with liver disease. Different from lncH19, the transcript of MEG3 is positively correlated with obesity index and HOMA-IR in humans. In accordance, lncMEG3 is highly expressed in high-fat diet-induced obese mice and ob/ob mice (57). In primary hepatocytes, overexpression of lncMEG3 results in increased expression of Foxo1, G6pc, Pck1; meanwhile, insulin-stimulated glycogen synthesis was suppressed by lncMEG3 (57). These alterations could be reversed by lncMEG3 interference (57). In another study, lncMEG3 is found to be a glucagon-inducible lncRNA in mouse primary hepatocytes, where it interacts with miR-302a-3p as a competing endogenous RNA (ceRNA) (58). By this way, lncMEG3 increases CREB-regulated transcriptional coactivator 2 (CRTC2), which is a target of miR-302a-3p. Consequently, upregulated CRTC2 stimulates gluconeogenesis by activating the axis of PGC-1α/Pck1/G6pc in hepatocytes (58). Furthermore, as a ceRNA, miR-214 is another substrate of lncMEG3. In hepatocytes, lncMEG3 sequesters miR-214 to favor transcription factor 4 (ATF4) expression (59). ATF4 is capable of inducing the gluconeogenic program by affecting the transcriptional activity of FOXO1 (60). Therefore, lncMEG3 promotes gluconeogenesis in hepatocytes by targeting the axis of miR-302a-3p/CRTC2 or miR-214/ATF4.
4.4 LncBhmt-AS
Betaine homocysteine methyltransferase (BHMT) is an enzyme that catalyzes the synthesis of methionine from homocysteine and is associated with insulin resistance and diabetes (61, 62). BHMT is highly expressed in the liver of rodents, which may play a role in gluconeogenesis by interacting with L-serine dehydratase/L-threonine deaminase to affect the use of the amino acid for gluconeogenesis (63). Recently, a new lncRNA was discovered during fasting, which is an antisense RNA of Bhmt, therefore, named lncBhmt-AS (64). LncBhmt-AS is located on chromosome 13 in mice with 1464 bp in length. Deficiency of lncBhmt-AS restricts gluconeogenesis in primary hepatocytes and inhibits liver glucose production and gluconeogenic gene expression in vivo (64). In contrast, Bhmt overexpression restores gluconeogenesis induced by lncBhmt-AS knockdown (64). These evidences indicate that lncBhmt-AS plays an important role in regulating hepatic gluconeogenesis by targeting Bhmt.
4.5 LncGm10768
Gm10768 is a lncRNA specifically enriched in the liver. Cui et al. found an abnormal increase of lncGm10768 in mouse livers after fasting by RNA-seq (65). In addition, lncGm10768 is positively correlated with glucose production in mouse primary hepatocytes (65). Liver-specific knockout of lncGm10768 alleviates hyperglycemia and insulin resistance in db/db mice (65). LncGm10768 is localized in the nucleus and cytoplasm, therefore, lncGm10768 may regulate gene expression at both transcriptional and post-transcriptional levels. As endogenous competitive suppressors of microRNAs, LncRNAs can reverse gene silencing induced by microRNAs. miR-214 has a high affinity binding site with lncGm10768, and it decreases in response to lncGm10768 overexpression (65). As mentioned above, miR-214 can target and activate transcription factor 4 (ATF4) to inhibit the expression of G6pc and Pck1 (59). Therefore, the positive impact of lncGm10768 on hepatic gluconeogenesis is likely due to the interaction with miR-214 (65). In this regard, lncGm10768 and lcnMEG3 play a similar role in hepatic gluconeogenesis by targeting miR-214, indicating different lncRNAs may have synergistic effects to regulate gluconeogenesis jointly.
4.6 LncGomafu
LncGomafu is a conserved lncRNA in mammalian species, which was localized in the nucleus in most cases. It has been well documented that lncGomafu plays a key role in neuronal development and involves in the pathogenesis of neuropsychiatric disorders (66, 67). Similar to lncMEG3, lncGomafu is highly expressed in the livers of ob/ob mice and mice on a high-fat diet (HFD) (68). Knockdown of lncGomafu in the liver inhibits hepatic glucose production and improves insulin sensitivity in obese mice. On the contrary, overexpression of lncGomafu increases blood glucose levels in lean mice. Mechanistically, lncGomafu competitively sponge miR-139 to increase Foxo1 expression, increasing gluconeogenic gene expression and hepatic gluconeogenesis (68).
4.7 LncMALAT1
Metastasis associated lung adenocarcinoma transcript 1(MALAT1) is a conserved lncRNA located on human chromosome 11q13 with a length of 8.5 kb (26). LncMALAT1 is considered a biomarker for tumor diagnosis and has been proven to be involved in the regulation of several signaling pathways, including PI3K/AKT, NF-kB, MAPK/ERK (69). Knockdown of lncMALAT1 in HepG2 and FLC4 cells leads to increased glucose secretion and expression of gluconeogenic genes such as G6pc and Pck1 (70). Meanwhile, this study revealed that the negative regulation of lncMALAT1 on gluconeogenesis is due to the upregulation of TCF7L2 (70). TCF7L2 has been shown to interact with the promoters of G6pc and Pck1, this interaction impedes the transcriptional activities of CREB/CRTC2 and FOXO1, thereby repressing gluconeogenic gene expression (71).
4.8 LncGm10804
LncGm10804 is highly enriched in high glucose-treated hepatocytes and livers of non-alcoholic fatty liver disease (NAFLD) model mice. Both in vitro and in vivo studies have shown that the knockdown of lncGm10804 reduces the expression of Pck1 and G6pc in cultured hepatocytes and NAFLD mice. Meanwhile, lncGm10804 silencing alleviates hepatic steatosis and lipid accumulation by decreasing the expression of sterol regulatory element-binding protein-1c (SREBP-1c) and fatty acid synthase (FAS) in NAFLD mouse livers (72).
4.9 LincIRS2
LncRNA 4833411C07Rik was named LincIRS2 by Marta Pradas-Juni et al. for its location at 80 kb of 5’ Irs2 (17). LincIRS2 is induced upon fasting or glucagon stimulation and responds to cAMP signaling (17), suggesting lincIRS2 might be involved in hepatic gluconeogenesis. Indeed, in lean mice, the knockdown of lincIRS2 in the liver induces enhanced blood glucose, insulin resistance and ectopic glucose output. Meanwhile, deficiency of Mafg in hepatocytes evokes a fasting-like gene expression profile as evidenced by elevated expression of Fbp1, G6pc and Pck1 (17). Later, they found that MAFG controls the expression of lincIRS2 and thereby regulates glucose metabolism in the liver (17).
5 Conclusion and perspectives
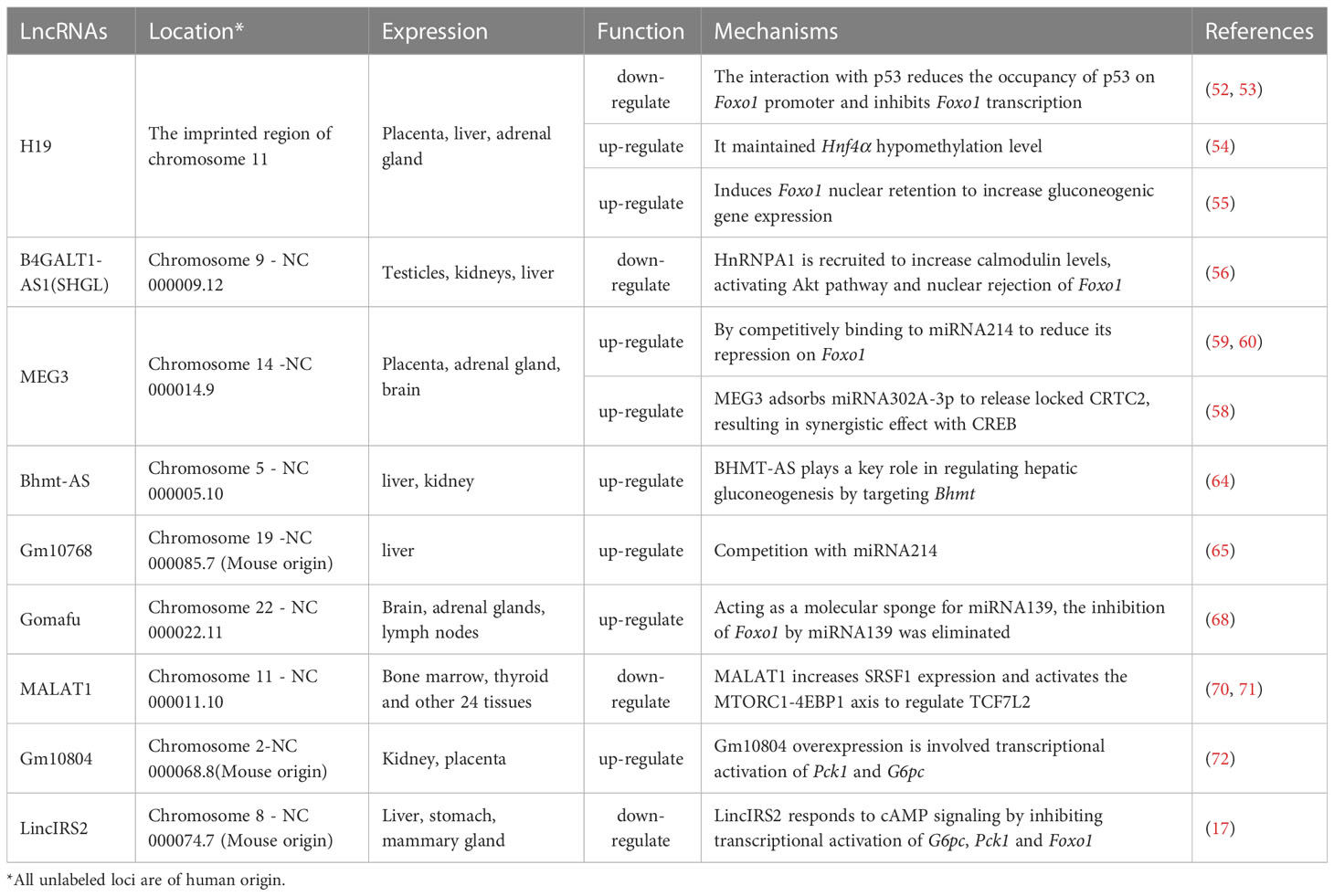
Hepatic gluconeogenesis is an essential bio-process to keep blood glucose in normal physiological scope. Dysregulated hepatic gluconeogenesis may course various metabolic disorders. For instance, ectopic upregulated gluconeogenesis is a causative factor for inducing hyperglycemia, hyperinsulinemia, insulin resistance, and T2D. In this regard, gluconeogenesis also is a target for developing anti-T2D drugs. Metformin is such an anti-T2D drug which has achieved great success in clinical. Therefore, in-depth studies on hepatic gluconeogenesis regulation and its molecular mechanisms are important for developing novel strategies for treating disorders induced by malfunctioned gluconeogenesis. The current evidences have shown that lncRNAs play a crucial role in hepatic gluconeogenesis, although just 9 lncRNAs have been examined in this field to date. Of note, these lncRNAs exhibited different functions on hepatic gluconeogenesis, lncH19, lncMEG3, lncGm10768, lncGomafu, and lncBhmt-AS function as positive regulators, whereas lncMALAT1 and lncSHGL act as negative regulators (Table 1). As for involved mechanisms, each lncRNA has its own working model (Figure 1).
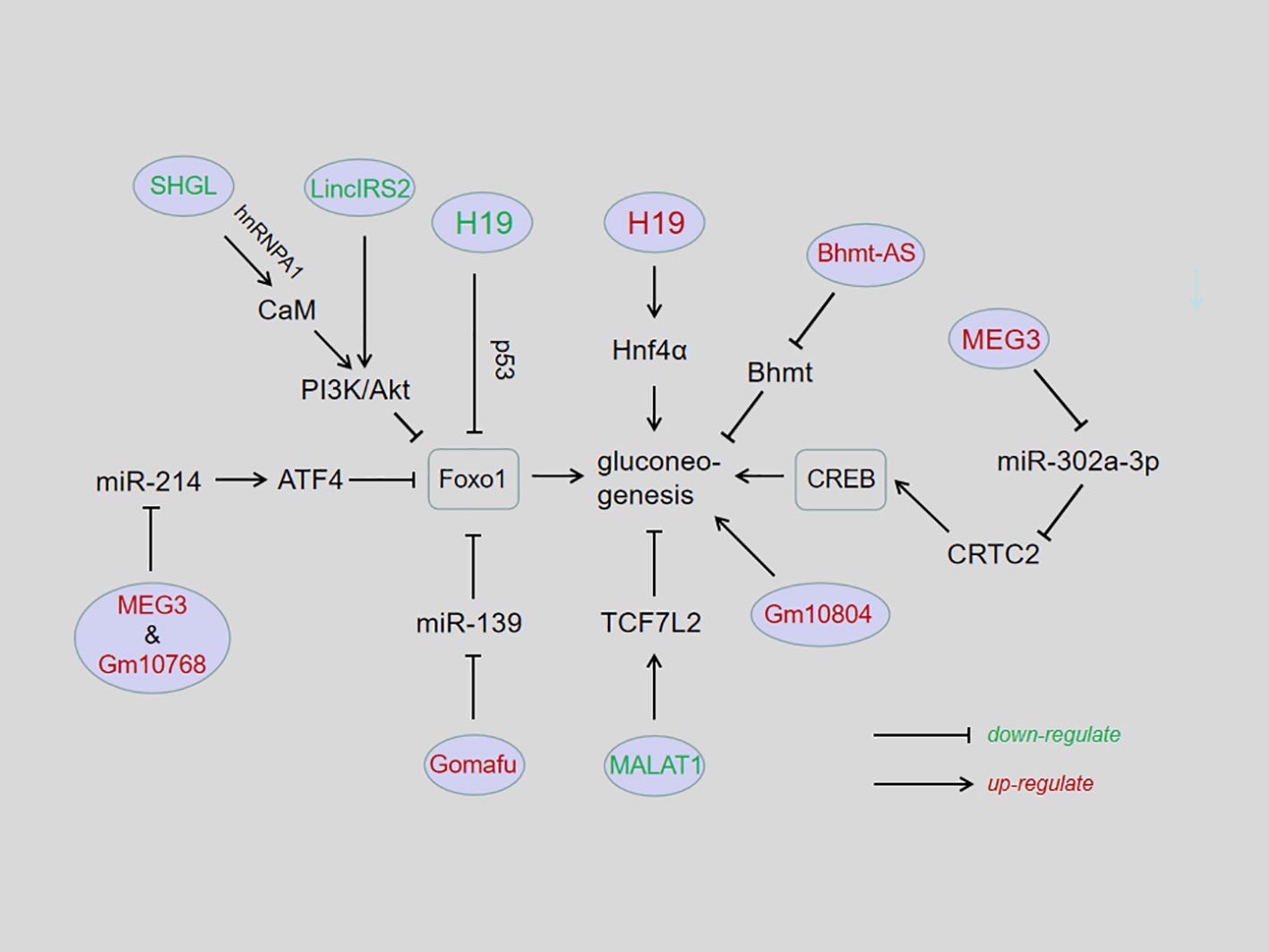
Figure 1 Influences of lncRNAs in hepatic gluconeogenesis. In general, lncRNAs affect hepatic gluconeogenesis in the following ways: lncRNAs directly regulate the expression of gluconeogenic genes, such as lncH19, lncSHGL, lncBhmt-AS, lcnMALAT1, lincIRS2, and lncGm10804; lncRNAs interact with miRNAs as competitive endogenous RNAs, such as lncMEG3/miR214; lncGm10768/miR214; lncRNAs act as microRNA molecular sponges to block subsequent pathways, such as lncH19/miRlet-7; lncMEG3/miR302-a-3p; lncGomafu/miR139; or lncRNAs bind to the promoter to affect gene transcription, such as lncH19/p53.
It should be mentioned that these gluconeogenesis-associated lncRNAs are mainly enriched in the liver. It is reasonable to predict that other tissues-derived lncRNAs may also be involved in hepatic gluconeogenesis, although no direct evidences support this prediction. It has been shown that lncRNAs are often carried by exosomes (73). Hence, lncRNAs derived from other tissues, such as muscle, pancreas, and fat, could be transferred into the liver via exosomes, where they might affect the gluconeogenic program. For this reason, exosomal lncRNA-mediated crosstalk between other tissues and hepatic gluconeogenesis could be investigated as a future direction. Therefore, we predicted that more lncRNAs potentially involved in the hepatic gluconeogenesis will be identified by RNA-seq technologies in non-liver tissues under stress, such as fasting or a high-fat diet. Engineered exosomes with specific lncRNAs with inhibitory effects on hepatic gluconeogenesis might hold great therapeutic potential for treating T2D. In addition, exosomal lncRNAs in blood might be diagnostic markers of dysregulated hepatic gluconeogenesis.
Author contributions
ZW: Validation, writing-original draft. JM: Writing-original draft. RW: Writing-original draft. YK: Project administration, writing-review and editing. CS: Conceptualization, funding acquisition, writing-review and editing. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the National Key Research and Development Program of China (Grant No. 2017YFA0701304), the National Natural Science Foundation of China (Nos. 81970747; 32271193), and the Scientific Program of Changshu City (csws202009).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
1. Kraus-Friedmann N. Hormonal regulation of hepatic gluconeogenesis. Physiol Rev (1984) 64:170–259. doi: 10.1152/physrev.1984.64.1.170
2. Pilkis SJ, el-Maghrabi MR, Claus TH. Hormonal regulation of hepatic gluconeogenesis and glycolysis. Annu Rev Biochem (1988) 57:755–83. doi: 10.1146/annurev.bi.57.070188.003543
3. Gerich JE, Nurjhan N. Gluconeogenesis in type 2 diabetes. Adv Exp Med Biol (1993) 334:253–8. doi: 10.1007/978-1-4615-2910-1_18
4. Cherrington AD, Frizzell RT, Biggers DW, Connolly CC. Hypoglycemia, gluconeogenesis and the brain. Adv Exp Med Biol (1991) 291:197–211. doi: 10.1007/978-1-4684-5931-9_15
5. Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature (2012) 489:57–74. doi: 10.1038/nature11247
6. Cech TR, Steitz JA. The noncoding RNA revolution-trashing old rules to forge new ones. Cell (2014) 157:77–94. doi: 10.1016/j.cell.2014.03.008
7. Mattick JS, Makunin IV. Non-coding RNA. Hum Mol Genet (2006) 15 Spec No 1:R17–29. doi: 10.1093/hmg/ddl046
8. Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in development and disease: Background, mechanisms, and therapeutic approaches. Physiol Rev (2016) 96:1297–325. doi: 10.1152/physrev.00041.2015
9. Palazzo AF, Koonin EV. Functional long non-coding RNAs evolve from junk transcripts. Cell (2020) 183:1151–61. doi: 10.1016/j.cell.2020.09.047
10. Bridges MC, Daulagala AC, Kourtidis A. LNCcation: lncRNA localization and function. J Cell Biol (2021) 220:e202009045. doi: 10.1083/jcb.202009045
11. Grammatikakis I, Lal A. Significance of lncRNA abundance to function. Mamm Genome (2022) 33:271–80. doi: 10.1007/s00335-021-09901-4
12. Klattenhoff CA, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML, et al. Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell (2013) 152:570–83. doi: 10.1016/j.cell.2013.01.003
13. Ghafouri-Fard S, Taheri M. The expression profile and role of non-coding RNAs in obesity. Eur J Pharmacol (2021) 892:173809. doi: 10.1016/j.ejphar.2020.173809
14. Goldfarb CN, Waxman DJ. Global analysis of expression, maturation and subcellular localization of mouse liver transcriptome identifies novel sex-biased and TCPOBOP-responsive long non-coding RNAs. BMC Genomics (2021) 22:212. doi: 10.1186/s12864-021-07478-5
15. Xia C, Zhang X, Cao T, Wang J, Li C, Yue L, et al. Hepatic transcriptome analysis revealing the molecular pathogenesis of type 2 diabetes mellitus in zucker diabetic fatty rats. Front Endocrinol (Lausanne) (2020) 11:565858. doi: 10.3389/fendo.2020.565858
16. Wang Y, Tang H, Ji X, Zhang Y, Xu W, Yang X, et al. Expression profile analysis of long non-coding RNAs involved in the metformin-inhibited gluconeogenesis of primary mouse hepatocytes. Int J Mol Med (2018) 41:302–10. doi: 10.3892/ijmm.2017.3243
17. Pradas-Juni M, Hansmeier NR, Link JC, Schmidt E, Larsen BD, Klemm P, et al. A MAFG-lncRNA axis links systemic nutrient abundance to hepatic glucose metabolism. Nat Commun (2020) 11:644. doi: 10.1038/s41467-020-14323-y
18. López-Gambero AJ, Martínez F, Salazar K, Cifuentes M, Nualart F. Brain glucose-sensing mechanism and energy homeostasis. Mol Neurobiol (2019) 56:769–96. doi: 10.1007/s12035-018-1099-4
19. Kumagai AK. Glucose transport in brain and retina: implications in the management and complications of diabetes. Diabetes/metabolism Res Rev (2015) 15:261–73. doi: 10.1002/(SICI)1520-7560(199907/08)15:4<261::AID-DMRR43>3.0.CO;2-Z
20. Chrościcki P, Usarek M, Bryla J. The role of biological clock in glucose homeostasis. Postpy Higieny i Medycyny Dowiadczalnej (Advances Hygiene Exp Medicine) (2013) 67:569–83. doi: 10.5604/17322693.1053906
21. Gerich JE. Physiology of glucose homeostasis. Diabetes Obes Metab (2010) 2:345–50. doi: 10.1046/j.1463-1326.2000.00085.x
22. Al-Yousif N, Rawal S, Jurczak M, Mahmud H, Shah FA. Endogenous glucose production in critical illness. Nutr Clin Pract (2021) 36:344–59. doi: 10.1002/ncp.10646
23. Baruh S, Sherman L, Kolodny HD, Jit A, Singh. Fasting hypoglycemia. Med Clinics North America (1973) 57:1441–62. doi: 10.1016/S0025-7125(16)32200-3
24. Oddo M, Schmidt JM, Carrera E, Badjatia N, Connolly ES, Presciutti M, et al. Impact of tight glycemic control on cerebral glucose metabolism after severe brain injury: A microdialysis study*. Crit Care Med (2008) 36:3233–8. doi: 10.1097/CCM.0b013e31818f4026
25. Petersen MC, Vatner DF, Shulman GI. Regulation of hepatic glucose metabolism in health and disease. Nat Rev Endocrinol (2017) 13:572–87. doi: 10.1038/nrendo.2017.80
26. Amodio N, Raimondi L, Juli G, Stamato MA, Caracciolo D, Tagliaferri P, et al. MALAT1: A druggable long non-coding RNA for targeted anti-cancer approaches. J Hematol Oncol (2018) 11:63. doi: 10.1186/s13045-018-0606-4
27. Nojima T, Proudfoot NJ. Mechanisms of lncRNA biogenesis as revealed by nascent transcriptomics. Nat Rev Mol Cell Biol (2022) 23:389–406. doi: 10.1038/s41580-021-00447-6
28. Matsumoto A, Pasut A, Matsumoto M, Yamashita R, Fung J, Monteleone E, et al. mTORC1 and muscle regeneration are regulated by the LINC00961-encoded SPAR polypeptide. Nature (2017) 541:228–32. doi: 10.1038/nature21034
29. Zhang M, Zhao K, Xu X, Yang Y, Yan S, Wei P. A peptide encoded by circular form of LINC-PINT suppresses oncogenic transcriptional elongation in glioblastoma. Nat Commun (2018) 9:4475. doi: 10.1038/s41467-018-06862-2
30. Pang Y, Liu Z, Han H, Wang B, Li W, Mao C, et al. Peptide SMIM30 promotes HCC development by inducing SRC/YES1 membrane anchoring and MAPK pathway activation. J Hepatol (2020) 73:1155–69. doi: 10.1016/j.jhep.2020.05.028
31. Zhu J, Liu C, Wang D, Cao X, Wang S, Liu Y, et al. The emerging landscapes of long noncoding RNA in thyroid carcinoma: Biological functions and clinical significance. Front Oncol (2021) 11:706011. doi: 10.3389/fonc.2021.706011
32. Ge Q, Jia D, Cen D, Qi Y, Shi C, Li J, et al. Micropeptide ASAP encoded by LINC00467 promotes colorectal cancer progression by directly modulating ATP synthase activity. J Clin Invest (2021) 131:e152911. doi: 10.1172/JCI152911
33. Zhao L, Wang J, Li Y, Song T, Wu Y, Fang S, et al. NONCODEV6: an updated database dedicated to long non-coding RNA annotation in both animals and plants. Nucleic Acids Res (2020) 49:D1. doi: 10.1093/nar/gkaa1046
34. Ma L, Cao J, Liu L, Du Q, Li Z, Zou D, et al. LncBook: a curated knowledgebase of human long non-coding RNAs. Nucleic Acids Res (2019) 47:5. doi: 10.1093/nar/gkz073
35. Yip CW, Sivaraman DM, Prabhu AV, Shin JW. Functional annotation of lncRNA in high-throughput screening. Essays Biochem (2021) 65:761–73. doi: 10.1042/EBC20200061
36. Chen L, Zhang YH, Pan X, Liu M, Wang S, Huang T, et al. Tissue expression difference between mRNAs and lncRNAs. Int J Mol Sci (2018) 19:3416. doi: 10.3390/ijms19113416
37. Ali T, Grote P. Beyond the RNA-dependent function of LncRNA genes. eLife Sci (2020) 9:e60583. doi: 10.7554/eLife.60583
38. Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol (2021) 22:96–118. doi: 10.1038/s41580-020-00315-9
39. Staněk D. Long non-coding RNAs and splicing. Essays Biochem (2021) 65:723–9. doi: 10.1042/EBC20200087
40. Ma L, Bajic VB, Zhang Z. On the classification of long non-coding RNAs. RNA Biol (2013) 10:924–33. doi: 10.4161/rna.24604
41. Kornienko AE, Guenzl PM, Barlow DP, Pauler FM. Gene regulation by the act of long non-coding RNA transcription. BMC Biol (2013) 11:59. doi: 10.1186/1741-7007-11-59
42. Huang W, Li H, Yu Q, Xiao W, Wang DO. LncRNA-mediated DNA methylation: an emerging mechanism in cancer and beyond. J Exp Clin Cancer Res (2022) 41:100. doi: 10.1186/s13046-022-02319-z
43. Han P, Chang CP, Long non-coding RNA. And chromatin remodeling. RNA Biol (2015) 12:00–0. doi: 10.1080/15476286.2015.1063770
44. Latos PA, Pauler FM, Koerner MV, Senergin HB, Hudson QJ, Stocsits RR, et al. Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science (2012) 338:1469–72. doi: 10.1126/science.1228110
45. Bast-Habersbrunner A, Kiefer C, Weber P, Fromme T, Schiessl A, Schwalie PC, et al. LncRNA ctcflos orchestrates transcription and alternative splicing in thermogenic adipogenesis. EMBO Rep (2021) 22:e51289. doi: 10.15252/embr.202051289
46. Schmitz SU, Grote P, Herrmann BG. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci (2016) 73:2491–509. doi: 10.1007/s00018-016-2174-5
47. Yang J, Qi M, Fei X, Wang X, Wang K. LncRNA H19: A novel oncogene in multiple cancers. Int J Biol Sci (2021) 17:3188–208. doi: 10.7150/ijbs.62573
48. Goshen R, Rachmilewitz J, Schneider T, de-Groot N, Ariel I. The expression of the h-19 and IGF-2 genes during human embryogenesis and placental development. Mol Reprod Dev (1993) 34:374–9. doi: 10.1002/mrd.1080340405
49. Juan V, Crain C, Wilson C. Evidence for evolutionarily conserved secondary structure in the H19 tumor suppressor RNA. Nucleic Acids Res (2000) 28:1221–7. doi: 10.1093/nar/28.5.1221
50. Daneshmoghadam J, Omidifar A, Akbari Dilmaghani N, Karimi Z, Emamgholipour S, Shanaki M. The gene expression of long non-coding RNAs (lncRNAs): MEG3 and H19 in adipose tissues from obese women and its association with insulin resistance and obesity indices. J Clin Lab Anal (2021) 35:e23741. doi: 10.1002/jcla.23741
51. Bi Y, Wang Y, Sun X. Recent advances of LncRNA H19 in diabetes LncRNA H19 in diabetes. Hormone Metab Res (2022) 54:212–9. doi: 10.1055/a-1769-5032
52. Goyal N, Sivadas A, Shamsudheen KV, Jayarajan R, Verma A, Sivasubbu S, et al. RNA Sequencing of db/db mice liver identifies lncRNA H19 as a key regulator of gluconeogenesis and hepatic glucose output. Sci Rep (2017) 7:8312. doi: 10.1038/s41598-017-08281-7
53. Goyal N, Tiwary S, Kesharwani D, Datta M. Long non-coding RNA H19 inhibition promotes hyperglycemia in mice by upregulating hepatic FoxO1 levels and promoting gluconeogenesis. J Mol Med (Berl) (2019) 97:115–26. doi: 10.1007/s00109-018-1718-6
54. Deng J, Mueller M, Geng T, Shen Y, Liu Y, Hou P, et al. H19 lncRNA alters methylation and expression of Hnf4alpha in the liver of metformin-exposed fetuses. Cell Death Dis (2017) 8:e3175. doi: 10.1038/cddis.2017.392
55. Wu HY, Cheng Y, Jin LY, Zhou Y, Pang HY, Zhu H, et al. Paternal obesity impairs hepatic gluconeogenesis of offspring by altering Igf2/H19 DNA methylation. Mol Cell Endocrinol (2021) 529:111264. doi: 10.1016/j.mce.2021.111264
56. Wang J, Yang W, Chen Z, Chen J, Meng Y, Feng B, et al. Long noncoding RNA lncSHGL recruits hnRNPA1 to suppress hepatic gluconeogenesis and lipogenesis. Diabetes (2018) 67:581–93. doi: 10.2337/db17-0799
57. Zhu X, Wu YB, Zhou J, Kang DM. Upregulation of lncRNA MEG3 promotes hepatic insulin resistance via increasing FoxO1 expression. Biochem Biophys Res Commun (2016) 469:319–25. doi: 10.1016/j.bbrc.2015.11.048
58. Zhu X, Li H, Wu Y, Zhou J, Yang G, Wang W, et al. CREB-upregulated lncRNA MEG3 promotes hepatic gluconeogenesis by regulating miR-302a-3p-CRTC2 axis. J Cell Biochem (2019) 120:4192–202. doi: 10.1002/jcb.27706
59. Zhu X, Li H, Wu Y, Zhou J, Yang G, Wang W. lncRNA MEG3 promotes hepatic insulin resistance by serving as a competing endogenous RNA of miR-214 to regulate ATF4 expression. Int J Mol Med (2019) 43:345–57. doi: 10.3892/ijmm.2018.3975
60. Li K, Zhang J, Yu J, Liu B, Guo Y, Deng J, et al. MicroRNA-214 suppresses gluconeogenesis by targeting activating transcriptional factor 4. J Biol Chem (2015) 290:8185–95. doi: 10.1074/jbc.M114.633990
61. Teng YW, Ellis JM, Coleman RA, Zeisel SH. Mouse betaine-homocysteine s-methyltransferase deficiency reduces body fat via increasing energy expenditure and impairing lipid synthesis and enhancing glucose oxidation in white adipose tissue. J Biol Chem (2012) 287:16187–98. doi: 10.1074/jbc.M111.303255
62. Wijekoon EP, Brosnan ME, Brosnan JT. Homocysteine metabolism in diabetes. Biochem Soc Trans (2007) 35:1175–9. doi: 10.1042/BST0351175
63. Garrido F, Pacheco M, Vargas-Martinez R, Velasco-Garcia R, Jorge I, Serrano H, et al. Identification of hepatic protein-protein interaction targets for betaine homocysteine s-methyltransferase. PloS One (2018) 13:e0199472. doi: 10.1371/journal.pone.0199472
64. Shen X, Zhang Y, Zhang X, Yao Y, Zheng Y, Cui X, et al. Long non-coding RNA bhmt-AS attenuates hepatic gluconeogenesis via modulation of bhmt expression. Biochem Biophys Res Commun (2019) 516:215–21. doi: 10.1016/j.bbrc.2019.06.008
65. Cui X, Tan J, Shi Y, Sun C, Li Y, Ji C, et al. The long non-coding RNA Gm10768 activates hepatic gluconeogenesis by sequestering microRNA-214 in mice. J Biol Chem (2018) 293:4097–109. doi: 10.1074/jbc.M117.812818
66. Barry G, Briggs JA, Vanichkina DP, Poth EM, Beveridge NJ, Ratnu VS, et al. The long non-coding RNA gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing. Mol Psychiatry (2014) 19:486–94. doi: 10.1038/mp.2013.45
67. Spadaro PA, Flavell CR, Widagdo J, Ratnu VS, Troup M, Ragan C, et al. Long noncoding RNA-directed epigenetic regulation of gene expression is associated with anxiety-like behavior in mice. Biol Psychiatry (2015) 78:848–59. doi: 10.1016/j.biopsych.2015.02.004
68. Yan C, Li J, Feng S, Li Y, Tan L. Long noncoding RNA gomafu upregulates Foxo1 expression to promote hepatic insulin resistance by sponging miR-139-5p. Cell Death Dis (2018) 9:289. doi: 10.1038/s41419-018-0321-7
69. Li ZX, Zhu QN, Zhang HB, Hu Y, Wang G, Zhu YS. MALAT1: a potential biomarker in cancer. Cancer Manag Res (2018) 10:6757–68. doi: 10.2147/CMAR.S169406
70. Malakar P, Stein I, Saragovi A, Winkler R, Stern-Ginossar N, Berger M, et al. Long noncoding RNA MALAT1 regulates cancer glucose metabolism by enhancing mTOR-mediated translation of TCF7L2. Cancer Res (2019) 79:2480–93. doi: 10.1158/0008-5472.CAN-18-1432
71. Oh KJ, Park J, Kim SS, Oh H, Choi CS, Koo SH, et al. TCF7L2 modulates glucose homeostasis by regulating CREB- and FoxO1-dependent transcriptional pathway in the liver. PloS Genet (2012) 8:e1002986. doi: 10.1371/journal.pgen.1002986
72. Li T, Huang X, Yue Z, Meng L, Hu Y. Knockdown of long non-coding RNA Gm10804 suppresses disorders of hepatic glucose and lipid metabolism in diabetes with non-alcoholic fatty liver disease. Cell Biochem Funct (2020) 38:839–46. doi: 10.1002/cbf.3495
Keywords: long non-coding RNAs, hepatic gluconeogenesis, blood glucose, type 2 diabetes, PEPCK, G6Pase
Citation: Wang Z, Ma J, Wu R, Kong Y and Sun C (2023) Recent advances of long non-coding RNAs in control of hepatic gluconeogenesis. Front. Endocrinol. 14:1167592. doi: 10.3389/fendo.2023.1167592
Received: 16 February 2023; Accepted: 21 March 2023;
Published: 31 March 2023.
Edited by:
Xinyu Xu, Nanjing Medical University, ChinaReviewed by:
Yunli Zhou, Massachusetts General Hospital and Harvard Medical School, United StatesCopyright © 2023 Wang, Ma, Wu, Kong and Sun. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yinghong Kong, a29uZ3lpbmdob25nQDE2My5jb20=; Cheng Sun, c3VuY2hlbmcxOTc1QG50dS5lZHUuY24=
†These authors have contributed equally to this work
 Zhe Wang1†
Zhe Wang1† Cheng Sun
Cheng Sun