- 1Vector Behavior and Pathogen Interaction Group, Centro de Pesquisas René Rachou, Fundação Oswaldo Cruz, Belo Horizonte, Brazil
- 2Chemical Ecology Unit, Department of Plant Protection Biology, Swedish University of Agricultural Sciences, Alnarp, Sweden
- 3Laboratory of Termitology, Department of Entomology, Federal University of Viçosa, Viçosa, Brazil
Olfaction is fundamental for most animals and critical for different aspects of triatomine biology, including host-seeking, reproduction, avoidance of predators, and aggregation in shelters. Ethological and physiological aspects of these olfactory-mediated behaviors are well-understood, but their molecular bases are still largely unknown. Here we investigated changes in the molecular mechanisms at the peripheral olfactory level in response to different physiological and developmental conditions. For this, the antennal expression levels of the odorant (Orco) and ionotropic (IR8a, IR25a, and IR76b) coreceptor genes were determined in Rhodnius prolixus by means of quantitative real-time PCR (qRT-PCR) analysis. Gene expression changes were analyzed to test the effect of feeding and imaginal molt for both sexes. Moreover, we analyzed whether expression of these genes changed during the early life of adult bugs. Under these conditions bugs display distinct behavioral responses to diverse chemical stimuli. A significantly decreased expression was induced by blood feeding on all coreceptor genes. The expression of all genes was significantly increased following the imaginal molt. These results show that olfactory coreceptor genes have their expression altered as a response to physiological or developmental changes. Our study suggests that olfactory coreceptor genes confer adaptability to the peripheral olfactory function, probably underlying the known plasticity of triatomine olfactory-mediated behavior.
Introduction
Chagas disease, caused by the protozoan parasite Trypanosoma cruzi, is transmitted to humans and other mammals by hematophagous insects of the subfamily Triatominae (Reduviidae). This zoonosis is endemic to 22 countries in Central and South America, where 90–100 million people live in endemic areas, 8 million people are estimated to be infected, and 12,000–14,000 deaths are reported annually (Senior, 2007; Coura and Viñas, 2010; Rassi et al., 2010; Schmunis and Yadon, 2010). Rhodnius prolixus Stål, 1859 (Hemiptera: Reduviidae) is the second most important vector of Chagas disease, and the main species transmitting T. cruzi to humans in Colombia and Venezuela (Fitzpatrick et al., 2008; Guhl et al., 2009; Rassi et al., 2010; Hashimoto and Schofield, 2012). In the absence of a vaccine and an effective drug treatment, vector control is central to prevent the disease (Rassi et al., 2010). Due to insecticide resistance in triatomine populations in Bolivia (Lardeux et al., 2010), and parts of Argentina and Venezuela (Vassena et al., 2000; González Audino et al., 2004), there is a dire need for the improvement or development of vector control strategies for sustained control of Chagas disease.
The olfactory system plays an essential role in several aspects of the biology of triatomines, such as shelter location (Lorenzo and Lazzari, 1996), food search (Núñez, 1982; Barrozo and Lazzari, 2004a,b), reproduction (Pontes et al., 2008; Vitta et al., 2009; Zacharias et al., 2010; Pontes and Lorenzo, 2012), and avoidance of predators (Ward, 1981; Manrique et al., 2006). In addition, R. prolixus locate hosts mainly through olfactory cues; hence their sense of smell directly regulates disease transmission (Guerenstein and Lazzari, 2009). An increased understanding of the olfactory system is therefore crucial for sustainable control of this disease vector. In other insects, considerable progress has been made in the understanding of the molecular basis of olfaction, which has fostered the development of novel olfactory-based strategies against agricultural pests and disease vectors (Kain et al., 2013; Tauxe et al., 2013). Two molecular components have been shown to be central for the detection of odorant stimuli in insects: the odorant receptors (ORs) (Clyne et al., 1999; Vosshall et al., 2000) and the ionotropic receptors (IRs) (Benton et al., 2009). Genes encoding for these proteins are expressed in olfactory sensory neurons (OSNs), primarily on the insect antennae (Carey and Carlson, 2011). ORs and IRs function as heteromeric odor-gated ion channels composed of one, or in the case of IRs up to five, variable subunits and one, or in the case of IRs up to three, obligate coreceptors: Orco (Vosshall et al., 2000; Larsson et al., 2004; Vosshall and Hansson, 2011), and IR8a, IR25a and IR76b, respectively (Benton et al., 2009; Abuin et al., 2011). These coreceptor proteins are also required for the trafficking of the heteromeric OR and IR complexes to the cilia of the OSNs (Larsson et al., 2004; Benton et al., 2006; Abuin et al., 2011).
Changes in the behavioral responsiveness to host signals and reproductive mates have been reported for bugs of this subfamily and correlated with the ingestion of a blood meal and adult maturation (Bodin et al., 2009b; Vitta and Lorenzo, 2009). Similar changes in vector behavior have been found to be correlated with alterations in gene expression in mosquitoes (Rinker et al., 2013; Omondi et al., 2015a). Regulation of gene transcription tentatively underlies the observed functional changes of the peripheral (Jang, 1995; Siju et al., 2010; Saveer et al., 2012; Omondi et al., 2015a) and central olfactory systems (Anton et al., 2007; Barrozo et al., 2011). The main objective of this report was to analyze ontogenetic and blood-meal induced changes in the transcript levels of OR and IR coreceptor genes in both sexes of R. prolixus. Based on behavioral observations we hypothesize that coreceptor gene expression is decreased in recently fed bugs. Moreover, we hypothesize that imaginal molting induces an increase in gene expression. We observed that RproOrco, RproIR8a, RproIR25a, and RproIR76b transcript levels are altered, in ways correlated with the significantly decreased behavioral responsiveness known for fed insects, as well as the acquisition of sexual signal detection capabilities in adults.
Materials and Methods
Insects
Experimental insects were obtained from the R. prolixus colony held at the Centro de Pesquisas René Rachou (CPqRR), which was established more than 20 years ago from a batch of domiciliary insects captured during field work in Honduras (donated by Dr. Carlos Ponce, Ministerio de Salud Pública, Honduras). Through the years, this colony has been kept as large as possible (ca. 20,000 insects) in order to preserve as much diversity as possible. Experimental insects were reared under controlled conditions at 26 ± 1°C, 65 ± 10% relative humidity, and at a 12 h:12 h light/dark cycle provided by artificial lights (4 fluorescent lamps, cold white light, 6400 K, 40 W). All experiments were performed with 5th instar larvae or adults, and all tests were developed separately for female and male insects. For experiments with immature insects, a group of 4th instar larvae of similar age was sorted and fed ad libitum with citrated rabbit blood (2.5% buffered sodium citrate, provided by Centro de Criação de Animais de Laboratório-CECAL, FIOCRUZ), using an artificial membrane feeder. After molting to the 5th instar, half of these insects were kept unfed, while the remaining bugs were offered blood ad libitum at day 16 after ecdysis. To obtain adult bugs for the remaining experiments, 5th instar larvae of similar age were sorted by sex and offered an ad libitum blood meal to induce their imaginal molt. As in the case of larvae, the feeding procedure was performed 16 days after the ecdysis of adult bugs. Transcript abundances for RproOrco and each IR coreceptor genes were analyzed separately for male and female bugs as follows: (i) unfed 21 day-old 5thinstar larvae; (ii) blood fed 21 day-old 5th instar larvae; (iii) unfed 1 day-old adult bugs; (iv) unfed 21 day-old adult bugs; and (v) blood fed 21 day-old adult bugs. All bug antennae were dissected between 10 am and 4 pm, and in the case of fed insects, antennae were cut 5 days after the ingestion of the blood meal. Each of the 5 treatments was replicated 6 times using pools of 60 antennae (i.e., 30 bugs) per sample.
Reference Genes and R. prolixus OR and IR Coreceptors
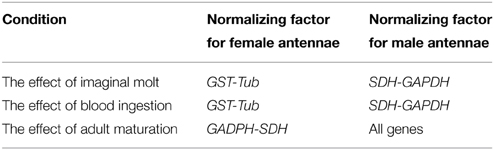
A set of candidate reference genes (Table 1) was selected because they were all previously used for qPCR normalization in triatomines (Majerowicz et al., 2011; Paim et al., 2012) and other insect species (Scharlaken et al., 2008; Lord et al., 2010; Ling and Salvaterra, 2011; Ponton et al., 2011). Table 2 lists all reference factors calculated as the geometric means of the most stable combinations of these genes (Omondi et al., 2015b), used to evaluate changes in gene expression in the antennae of R. prolixus. The sequences of reference and target genes (RproOrco, RproIR8a, RproIR25a, and RproIR76b) were identified in the R. prolixus genome (available on www.vectorbase.org/organisms/rhodnius-prolixus) using a local tBLASTn algorithm (Altschul et al., 1997). Orthologous sequences were obtained from the Swiss Institute of Bioinformatics (Table S1 in Supplementary Material). Primers were designed using Primer3 4.0.0 (http://primer3.ut.ee/) (Rozen and Skaletsky, 2000) and compatibilities tested with Oligoanalyser (Integrated DNA Technologies, Inc. IA, USA) softwares. The melting temperature was set at 60°C. The specificity for each primer was tested in silico using BLASTn (Altschul et al., 1990) in the R. prolixus genome database.
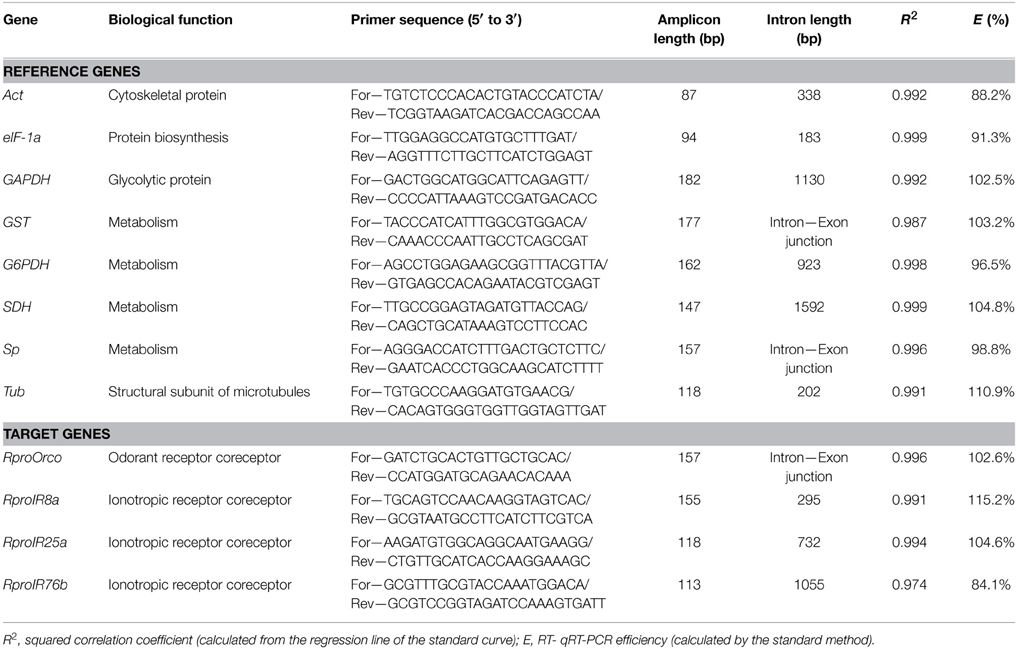
Table 1. Reference and target genes, biological function, primer sequences, amplicon and intron lengths, squared correlation coefficient, and qRT-PCR efficiency.
RNA Extraction and cDNA Synthesis
Total RNA was extracted from pools of 60 antennae with 500 μL of TRIzol® Reagent (Life Technologies, Carlsbad, CA, USA) according to the manufacturer's instructions. Then, extracted RNA was resuspended in 30 μL of DEPC-treated water (Life Technologies), and its concentration was determined using a Qubit® 2.0 Fluorometer (Life Technologies). RNA integrity was analyzed by visualizing bands on agarose electrophoresis gels. Extraction of RNA was followed by a treatment using RQ1 RNase-Free DNase (Promega, Fitchburg, WI, USA). All treated RNA (11 μL per sample) was immediately used to synthesize cDNA using SuperScript III Reverse Transcriptase (Life Technologies) and a 1:1 mix of Random Hexamer and 10 μM Oligo(dT)20 primers in a final volume of 20 μL.
Quantitative Real-time PCR
For quantitative real time PCR (qPCR) analysis, 10 μL of SYBR Green PCR Master Mix® (Life Technologies) were used in the reaction mixture that also contained 0.8 μL of a 10 μM primer solution and 1 μL of cDNA sample diluted two-fold in a final volume of 20 μL. The reactions were conducted using an ABIPRISM 7500 Sequence Detection System (Life Technologies) under the following conditions: one 10 min cycle at 95°C, followed by 40 cycles of 15 s at 95°C, 20 s at 60°C, and 30 s at 72°C. Following the amplification step, a melting curve analysis and an agarose gel electrophoresis were performed to confirm the specificity of the reaction. In all qPCR experiments, no-template controls (NTC) were included in triplicate for each primer set to verify the absence of exogenous DNA. For each experimental condition, six biological replicates were made, with three technical replicates performed for each of them. The PCR efficiencies (E) and repeatability (R2) for each primer were determined using the slope of a linear regression model (Pfaffl, 2001). Information about primers, PCR amplicons and calibration curves is presented in Table 1. Besides, the output of melt curve analysis for all primers is displayed in Figure S1 of Supplementary Material.
RT-PCR and Sequencing
Pure cDNA was used as a template for PCR reactions of the reference and target gene amplicons which were performed for 35 cycles (94°C for 30 s, 60°C for 30 s, and 72°C for 30 s) with 2 μL of cDNA, 2.2 μL of a 1 mM dNTP solution, 1.2 μL of a 10 μM primer solution and 1 U of Taq polymerase (Promega) in a final volume of 20 μL. The size of the resulting PCR products was visualized by means of electrophoresis in agarose gels. These PCR products were purified using the Wizard Genomic DNA Purification Kit (Promega). The sequencing reactions for the purified products were performed with both primers using an ABI Prism BigDye V 3.1 Terminator Cycle Sequencing kit and an ABI 3730 DNA sequencing system (Life Technologies). The consensus sequences were obtained using the Staden Package 2.0 (Staden et al., 2000) and verified by comparing with the R. prolixus genomic database, using the basic local alignment search tool (BLASTn).
Gene Expression and Statistical Analysis
Data treatment for quantification cycle (Cq) values obtained from technical replicates followed standard procedures for qPCR (Livak and Schmittgen, 2001; Sengul and Tu, 2008). Briefly, readings from each set of technical replicates were checked for consistency using GenEx software (MultiD Analyses AB, Sweden) and then used to calculate mean Cq-values for each biological replicate. To determine the gene expression measures, the Cq-values were normalized to those of reference genes, and then to mean Cq-values obtained with a control treatment (Livak and Schmittgen, 2001; Sengul and Tu, 2008). By using such procedures, we have prioritized a data processing method that allows comparison to most qPCR analyses available in the literature. All raw Cq-values are presented in Table S2 of Supplementary Material.
The relative expression of RproOrco, RproIR8a, RproIR25a, and RproIR76b in female and male antennae was calculated in GenEx software (MultiD Analyses AB, Sweden) using the 2 –ΔΔCt method (Livak and Schmittgen, 2001). First, the expression levels of the four genes were normalized to the reference factors selected for each sex. Then, expression levels of each gene were normalized to the expression levels of unfed female and male larvae, respectively.
Fold-change values were subjected to statistical analysis to determine the effect of treatment on transcript abundance. All tests were performed separately for data obtained from female and male antennae and no comparisons were performed between sexes due to our experimental design. In order to inspect whether gene expression (y-var) was affected by developmental instar (larvae × adult) or feeding status (unfed × fed) (x-vars), data were subjected to Generalized Linear Modeling (GLM) under normal errors. Posterior residual analyses confirmed the choice of the error distribution and the suitability of the model. Modeling proceeded by building a full model, including all of the above parameters and their first order interactions and comparing this with a null model built without any of the above factors. In finding significant differences between null and full models, model simplification was performed on the latter by backward term extraction, removing one term at a time. Terms returned to the model if their removal provoked a change of deviance with P < 0.05. The minimum adequate model was defined as the one holding only significant terms. The procedure above was applied independently for male and female bugs for each of the genes under study (RproOrco, RproIR8a, RproIR25a, and RproIR76b), each new test using a distinct subset of data. All tests were performed using R version 3.2.0 (R Core Team, 2015).
Results
Results from statistical analyses are summarized in Table 3 and Figure 1. Regardless of gender, the expression of all studied genes was enhanced in adults compared to larvae. Similarly, feeding depressed the expression of all coreceptor genes in larvae and adults, irrespective of their gender.
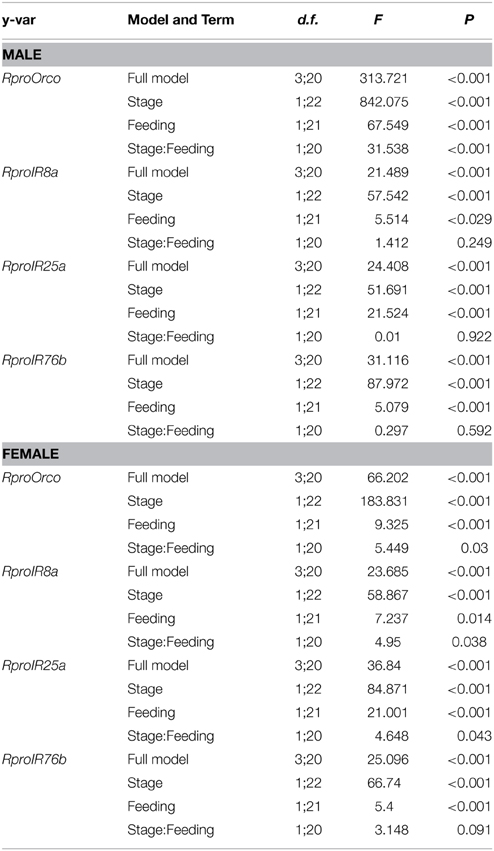
Table 3. Generalized Linear Modeling for olfactory coreceptor gene expression in male and female antennae of Rhodnius prolixus.
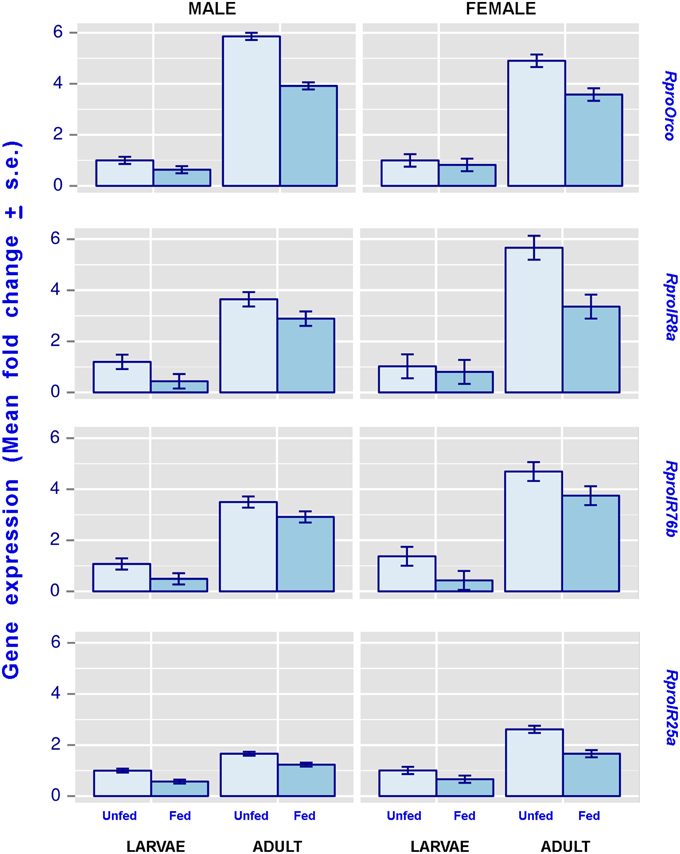
Figure 1. The effect of imaginal molt and nutrition on male and female olfactory coreceptor gene expression levels, as seen in the antennae of 5th instar larvae and adults. Both 5th instar larvae and adults included in this figure were 21 day-old. Error bars represent the standard deviation generated from 6 replicates per condition. All statistical comparisons, i.e., the effect of the imaginal molt and the effect of feeding, were significantly different (more details about these comparisons can be observed in Table 3).
In males, developmental stage (i.e., larvae × adults) and feeding status (i.e., unfed × fed) acted independently from each other on the expression of all genes, except for RproOrco. This could be confirmed by looking at the non-significant interaction terms (stage:feeding) for the three ionotropic coreceptors (RproIR8a, RproIR76b, and RproIR25a), and the significant interaction term for RproOrco (Table 3). That is, unfed larvae differed from fed larvae in the same proportion as unfed adults differed from fed adults for the three ionotropic coreceptors (Table 3). For RproOrco expression, however, the effect of feeding was different from larvae and adults, as revealed by the significant interaction term for this specific case (Table 3).
As for females, there was a distinct pattern: an interdependence of developmental stage and feeding status affected the expression of all genes except for RproIR76b, as revealed by the interaction terms (stage:feeding) in Table 3. That is, the proportion by which unfed larvae differed from fed larvae was distinct from the proportion by which unfed adults differed from fed ones for RproOrco, RproIR8a, and RproIR25a (Table 3). Conversely, such proportions did not differ for RproIR76b (Table 3). In summary, the effects of developmental stage and feeding status tended to affect gene expression independently in males, the opposite occurring in females.
The results obtained with unfed 1-day-old adults are presented in Figure S2 in order to allow their comparison to those of unfed 21-day-old larvae and unfed 21-day-old adults.
Discussion
The abundance of antennal transcripts of the olfactory coreceptor genes of R. prolixus changes in response to development and blood feeding, and can also be affected by interactions between these factors. The results of this study reveal that the expression of olfactory coreceptor genes is a plastic process, closely linked to the observed changes in olfactory-mediated behaviors in these insects. Proper olfactory function requires the obligatory presence of coreceptors in a hypothetical fixed stoichiometry together with olfactory receptors (Vosshall et al., 2000; Benton et al., 2009; Abuin et al., 2011). This would mean that alterations in coreceptor expression levels may reflect changes in the sensitivity of the olfactory system. Nonetheless, variations in coreceptor abundance may not reflect specific states of particular ORs or IRs. In fact, the changes in coreceptor expression reported here may have been the overall outcome of up or down regulation, or even absence of alteration, in specific receptors co-expressed with them. The expression of some olfactory receptors might be expected to remain unchanged in triatomines, e.g., those responsible for alarm or aggregation pheromone detection, as bugs are responsive to these stimuli irrespective of their nutritional or developmental state (Figueiras and Lazzari, 2000). Alternatively, other receptors related to functions that are dependent on good nutritional status may be anticipated to show an increase in expression, e.g., those that detect sexual pheromones (Vitta and Lorenzo, 2009).
The antennae of triatomines show a three-to-five-fold increase in the number of olfactory sensilla after their imaginal molt (Catalá, 1997; Gracco and Catalá, 2000; Akent'eva, 2008). Information about the ultrastructure of triatomines chemosensory sensilla is scarce, but available data suggest that triatomine trichoid sensilla may house up to 15 sensory neurons (Wigglesworth and Gillett, 1934). Rough estimates suggest that adult R. prolixus have approximately 1700 olfactory sensilla (Gracco and Catalá, 2000). Therefore, a concomitant increase in olfactory receptor expression would be expected when adult bug antennae are compared to those of fifth instar larvae. Consistently our results showed that the antennal expression of all coreceptors studied presented a significant increase in 21-day-old adults (Figure 1). This indicates that both the OR and IR based olfactory subsystems (Silbering et al., 2011) seem to undergo a significant expansion in the adult phase of these hemimetabolous insects. Immature triatomines share several chemosensory mediated behaviors with adult bugs (Ward, 1981; Lorenzo Figueiras et al., 1994; Manrique et al., 2006; Guerenstein and Lazzari, 2009). These include the orientation to hosts, alarm and aggregation responses. Nevertheless, adult triatomines make use of sexual pheromones to find mates for reproduction (Pontes et al., 2008; Vitta et al., 2009; May-Concha et al., 2013) and the observed increase in coreceptor expression in adult antennae seems to support the hypothesis that a significant expansion is taking place on OR and IR subsystems to cope with sexual functions. Further experiments need to be performed to determine whether this proposal is indeed correct.
The increase in coreceptor expression observed for adults could be hypothesized to be originated either during the imaginal molt or at the initial phase of adult life. Newly molted bugs do not respond to cues associated with their vertebrate hosts and recently molted adults show a low behavioral responsiveness toward mates, unlike older ones (Bodin et al., 2009b; Vitta and Lorenzo, 2009). The latter happens despite the fact that the antennae of triatomines show an increase in the number of olfactory sensilla after the imaginal molt (Catalá, 1997; Gracco and Catalá, 2000; Akent'eva, 2008). Combined, previous behavioral studies and our gene expression analyses suggest that the peripheral olfactory system of R. prolixus undergoes a post-eclosion maturation process in adult bugs (Figure S2 in Supplementary Material). Similar maturation has been reported in female mosquitoes (Omondi et al., 2015a), which at early imaginal life do not express proper host-seeking behavior, have a decreased neural sensitivity to host volatiles and a lower expression level of olfactory receptor genes (Davis, 1984; Grant and O'Connell, 2007; Bohbot et al., 2013). Since proper olfactory function requires the obligatory presence of coreceptors (Vosshall et al., 2000; Benton et al., 2009; Abuin et al., 2011), alterations in coreceptor expression levels may induce changes in the sensitivity of the olfactory system of R. prolixus, ultimately leading to an increased behavioral responsiveness toward vertebrate host volatiles and pheromones in mature adults.
Larval R. prolixus display reduced electrophysiological responses to ammonia after ingesting a blood meal (Reisenman, 2014). Moreover, engorged triatomine larvae are refractory to host odor stimulation for a prolonged time after feeding (Bodin et al., 2009a) and remain hidden in shelters for several days while their molting is completed. A similar refractory period has been observed in blood fed mosquitoes (Klowden and Lea, 1979; Takken et al., 2001). In both R. prolixus (our study) and the mosquito Anopheles gambiae (Rinker et al., 2013), blood feeding induces a reduction in chemosensory gene transcript production. Moreover, both A. gambiae and Aedes aegypti mosquitoes have reduced electrophysiological responses to host odors during the refractory period post-blood meal (Takken et al., 2001; Siju et al., 2010). We suggest that coreceptor down-regulation would represent a way to shut down the system and save energy. It is interesting to note that the decrease induced by the blood meal tended to be more significant for adult bug antennae, when compared to larval expression. This was the case for RproOrco (in both sexes), RproIr8a and RproIr25a (only for females). Further experiments would be necessary to clarify the functional bases of the observed differences.
Our results show that changes in olfactory coreceptor gene transcripts seem to be linked with the observed plasticity in behavioral responsiveness of larval and adult R. prolixus to host volatiles and mates. How these changes are reflected in the functional characteristics of the peripheral and central olfactory systems requires further analysis. This report is the first in line for understanding the molecular basis of neurophysiological modulation of triatomine olfactory driven behaviors.
Author Contributions
JML, Provided most experimental data, analyzed data and wrote the manuscript; BO, Advised with experimental procedures, analyzed data, wrote, and provided comments on the manuscript; OD, Analyzed data and wrote the manuscript; IO, Provided experimental data; RI, Wrote and provided comments on the manuscript; ML, Conceived the project, help and advised on experiments, wrote, and provided comments on the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors wish to thank the Program for Technological Development in Tools for Health-PDTIS-FIOCRUZ for having facilitated the use of its facilities. We wish to thank Mercedes Carolina Soares da Silva for her help in processing samples used in our experiments and Dra. Maria Victoria Periago for her support on statistical procedures. Authors are indebted to INCTEM (Project number: 573959/2008-0), FAPEMIG (Project number: APQ-01359-11), PROEP-FIOCRUZ (Project number: 401973/2012-3), CNPq (Project number: 483805/2013-0), FIOCRUZ Visiting researcher fellowship program (fellowship JMLE 550017/2012-7), the Linnaeus initiative “Insect Chemical Ecology, Ethology and Evolution” IC-E3 (Formas, SLU) and OS is supported by a Research Fellowship from CNPq (PQ 305736/2013-2)
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fevo.2015.00074
References
Abuin, L., Bargeton, B., Ulbrich, M., Isacoff, E., Kellenberger, S., and Benton, R. (2011). Functional architecture of olfactory ionotropic glutamate receptors. Neuron 69, 44–60. doi: 10.1016/j.neuron.2010.11.042
Akent'eva, N. A. (2008). The formation of the antenna sensory apparatus in some bug (Heteroptera) species in the course of their postembryonic development. Entomol. Rev. 88, 381–390. doi: 10.1134/s0013873808040015
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local alignment search tool. J. Mol. Biol. 215, 403–410. doi: 10.1016/S0022-2836(05)80360-2
Altschul, S. F., Madden, T. L., Schaffer, A. A., Zhang, J., Zhang, Z., Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402. doi: 10.1093/nar/25.17.3389
Anton, S., Dufour, M. C., and Gadenne, C. (2007). Plasticity of olfactory guided behaviour and its neurobiological basis: lessons from moths and locusts. Entomol. Exp. Appl. 123, 1–11. doi: 10.1111/j.1570-7458.2007.00516.x
Barrozo, R. B., Jarriault, D., Deisig, N., Gemeno, C., Monsempes, C., Lucas, P., et al. (2011). Mating induced differential coding of plant odour and sex pheromone in a male moth. Eur. J. Neurosci. 33, 1841–1850. doi: 10.1111/j.1460-9568.2011.07678.x
Barrozo, R. B., and Lazzari, C. R. (2004a). The response of the blood-sucking bug Triatoma infestans to carbon dioxide and other host odours. Chem. Senses 29, 319. doi: 10.1093/chemse/bjh035
Barrozo, R. B., and Lazzari, C. R. (2004b). Orientation behaviour of the blood-sucking bug Triatoma infestans to short-chain fatty acids: synergistic effect of L-lactic acid and carbon dioxide. Chem. Senses 29, 833–841. doi: 10.1093/chemse/bjh249
Benton, R., Sachse, S., Michnick, S. W., and Vosshall, L. B. (2006). Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol. 4:e20. doi: 10.1371/journal.pbio.0040020
Benton, R., Vannice, K., Gomez-Diaz, C., and Vosshall, L. B. (2009). Variant ionotropic glutamate receptors as chemosensory receptors in Drosophila. Cell 136, 149–162. doi: 10.1016/j.cell.2008.12.001
Bodin, A., Vinauger, C., and Lazzari, C. R. (2009a). Behavioural and physiological state dependency of host seeking in the blood-sucking insect Rhodnius prolixus. J. Exp. Biol. 212, 2386–2393. doi: 10.1242/jeb.030668
Bodin, A., Vinauger, C., and Lazzari, C. R. (2009b). State-dependency of host-seeking in Rhodnius prolixus: the post-ecdysis time. J. Insect Physiol. 55, 574–579. doi: 10.1016/j.jinsphys.2009.02.004
Bohbot, J. D., Durand, N. F., Vinyard, B. T., and Dickens, J. C. (2013). Functional development of the octenol response in Aedes aegypti. Front. Physiol. 4:39. doi: 10.3389/fphys.2013.00039
Carey, A. F., and Carlson, J. R. (2011). Insect olfaction from model systems to disease control. Proc. Natl. Acad. Sci. U.S.A. 108, 12987–12995. doi: 10.1073/pnas.1103472108
Catalá, S. S. (1997). Antennal sensilla of triatominae (Hemiptera, Reduviidae): a comparative study of five genera. Int. J. Insect Morphol. Embryol. 26, 67–73. doi: 10.1016/S0020-7322(97)00014-7
Clyne, P., Warr, C., Freeman, M., Lessing, D., Kim, J., and Carlson, J. R. (1999). A novel family of divergent seven-transmembrane proteins: candidate odorant receptors in Drosophila. Neuron 22, 327–338. doi: 10.1016/S0896-6273(00)81093-4
Coura, J. R., and Viñas, P. A. (2010). Chagas disease: a new worldwide challenge. Nature 465(7301), S6–S7. doi: 10.1038/nature09221
Davis, E. E. (1984). Development of lactic acid-receptor sensitivity and host-seeking behaviour in newly emerged female Aedes aegypti mosquitoes. J. Insect Physiol. 30, 211–215. doi: 10.1016/0022-1910(84)90005-2
Figueiras, A. N. L., and Lazzari, C. R. (2000). Temporal change of the aggregation response in Triatoma infestans. Mem. Inst. Oswaldo Cruz 95, 889–892. doi: 10.1590/S0074-02762000000600026
Fitzpatrick, S., Feliciangeli, M. D., Sanchez-Martin, M. J., Monteiro, F. A., and Miles, M. A. (2008). Molecular genetics reveal that silvatic Rhodnius prolixus do colonise rural houses. PLoS Negl. Trop. Dis. 2:e210. doi: 10.1371/journal.pntd.0000210
González Audino, P., Vassena, C., Barrios, S., Zerba, E., and Picollo, M. I. (2004). Role of enhanced detoxication in a deltamethrin-resistant population of Triatoma infestans (Hemiptera, Reduviidae) from Argentina. Mem. Inst. Oswaldo Cruz 99, 335–339. doi: 10.1590/S0074-02762004000300018
Gracco, M., and Catalá, S. S. (2000). Inter-specific and developmental differences on the array of antennal chemoreceptors in four species of Triatominae (Hemiptera: reduviidae). Mem. Inst. Oswaldo Cruz 95, 67–74. doi: 10.1590/S0074-02762000000100010
Grant, A. J., and O'Connell, R. J. (2007). Age-related changes in female mosquito carbon dioxide detection. J. Med. Entomol. 44, 617–623. doi: 10.1603/0022-2585(2007)44[617:ACIFMC]2.0.CO;2
Guerenstein, P. G., and Lazzari, C. R. (2009). Host-seeking: how triatomines acquire and make use of information to find blood. Acta Trop. 110, 148–158. doi: 10.1016/j.actatropica.2008.09.019
Guhl, F., Pinto, N., and Aguilera, G. (2009). Sylvatic triatominae: a new challenge in vector control transmission. Mem. Inst. Oswaldo Cruz 104 Suppl. 1, 71–75. doi: 10.1590/S0074-02762009000900012
Hashimoto, K., and Schofield, C. J. (2012). Elimination of Rhodnius prolixus in Central America. Parasit. Vectors 5:45. doi: 10.1186/1756-3305-5-45
Jang, E. B. (1995). Effects of mating and accessory gland injections on olfactory-mediated behavior in the female mediterranean fruit fly Ceratitis capitata. J. Insect Physiol. 41, 705–710. doi: 10.1016/0022-1910(95)00015-M
Kain, P., Boyle, S. M., Tharadra, S. K., Guda, T., Pham, C., Dahanukar, A., et al. (2013). Odour receptors and neurons for DEET and new insect repellents. Nature 502, 507–512. doi: 10.1038/nature12594
Klowden, M. J., and Lea, A. O. (1979). Humoral inhibition of host-seeking in Aedes aegypti during oocyte maturation. J. Insect Physiol. 25, 231–235. doi: 10.1016/0022-1910(79)90048-9
Lardeux, F., Depickère, S., Duchon, S., and Chavez, T. (2010). Insecticide resistance of Triatoma infestans (Hemiptera, Reduviidae) vector of Chagas disease in Bolivia. Trop. Med. Int. Health 15, 1037–1048. doi: 10.1111/j.1365-3156.2010.02573.x
Larsson, M. C., Domingos, A. I., Jones, W. D., Chiappe, M. E., Amrein, H., and Vosshall, L. B. (2004). Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron 43, 703–714. doi: 10.1016/j.neuron.2004.8.019
Ling, D., and Salvaterra, P. (2011). Robust RT-qPCR data normalization: validation and selection of internal reference genes during post-experimental data analysis. PLoS ONE 6:e17762. doi: 10.1371/journal.pone.0017762
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Lord, J., Hartzer, K., Toutges, M., and Oppert, B. (2010). Evaluation of quantitative PCR reference genes for gene expression studies in Tribolium castaneum after fungal challenge. J. Microbiol. Methods 80, 219–221. doi: 10.1016/j.mimet.2009.12.007
Lorenzo, M. G., and Lazzari, C. R. (1996). The spatial pattern of defecation in Triatoma infestans and the role of faeces as a chemical mark of the refuge. J. Insect Physiol. 42, 903–907. doi: 10.1016/0022-1910(96)00008-X
Lorenzo Figueiras, A. N., Kenigsten, A., and Lazzari, C. R. (1994). Aggregation in the haematophagous bug Triatoma infestans: chemical signals and temporal pattern. J. Insect Physiol. 40, 311–316. doi: 10.1016/0022-1910(94)90071-X
Tauxe, G. M., MacWilliam, D., Boyle, S. M., Guda, T., and Ray, A. (2013). Targeting a dual detector of skin and CO2 to modify mosquito host seeking. Cell 155, 1365–1379. doi: 10.1016/j.cell.2013.11.013
Majerowicz, D., Alves-Bezerra, M., Logullo, R., Fonseca-De-Souza, A. L., Meyer-Fernandes, J. R., Braz, G. R., et al. (2011). Looking for reference genes for real-time quantitative PCR experiments in Rhodnius prolixus (Hemiptera: Reduviidae). Insect Mol. Biol. 20, 713–722. doi: 10.1111/j.1365-2583.2011.01101.x
Manrique, G., Vitta, A. C. R., Ferreira, R. A., Zani, C. L., Unelius, C. R., Lazzari, C. R., et al. (2006). Chemical communication in Chagas disease vectors. Source, identity, and potential function of volatiles released by the metasternal and brindley's glands of Triatoma infestans adults. J. Chem. Ecol. 32, 2035–2052. doi: 10.1007/s10886-006-9127-7
May-Concha, I., Rojas, J. C., Cruz-López, L., Millar, J. G., and Ramsey, J. M. (2013). Volatile compounds emitted by Triatoma dimidiata, a vector of Chagas disease: chemical analysis and behavioural evaluation. Med. Vet. Entomol. 27, 165–174. doi: 10.1111/j.1365-2915.2012.01056.x
Núñez, J. (1982). Food source orientation and activity in Rhodnius prolixus Stål (Hemiptera: Reduviidae). Bull. Entomol. Res. 72, 253–262. doi: 10.1017/S0007485300010555
Omondi, B. A., Latorre-Estivalis, J. M., Oliveira, I. H. R., Ignell, R., and Lorenzo, M. G. (2015b). Evaluation of reference genes for insect olfaction studies. Parasit. Vectors 8, 243. doi: 10.1186/s13071-015-0862-x
Omondi, B. A., Majeed, S., and Ignell, R. (2015a). Functional development of carbon dioxide detection in the maxillary palp of Anopheles gambiae. J. Exp. Biol. doi: 10.1242/jeb.116798. [Epub ahead of print].
Paim, R. M., Pereira, M. H., Di Ponzio, R., Rodrigues, J. O., Guarneri, A. A., Gontijo, N. F., et al. (2012). Validation of reference genes for expression analysis in the salivary gland and the intestine of Rhodnius prolixus (Hemiptera, Reduviidae) under different experimental conditions by quantitative real-time PCR. BMC Res. Notes 5:128. doi: 10.1186/1756-0500-5-128
Pfaffl, M. W. (2001). A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29:e45. doi: 10.1093/nar/29.9.e45
Pontes, G. B., Bohman, B., Unelius, C., and Lorenzo, M. G. (2008). Metasternal gland volatiles and sexual communication in the triatomine bug, Rhodnius prolixus. J. Chem. Ecol. 34, 450–457. doi: 10.1007/s10886-008-9431-5
Pontes, G. B., and Lorenzo, M. G. (2012). Female metasternal gland odours mediate male aggregation in Rhodnius prolixus, a triatomid bug. Med. Vet. Entomol. 26, 33–36. doi: 10.1111/j.1365-2915.2011.00983.x
Ponton, F., Chapuis, M.-P., Pernice, M., Sword, G., and Simpson, S. (2011). Evaluation of potential reference genes for reverse transcription-qPCR studies of physiological responses in Drosophila melanogaster. J. Insect Physiol. 57, 840–850. doi: 10.1016/j.jinsphys.2011.03.014
Rassi, A. Jr., Rassi, A., and Marin-Neto, J. A. (2010). Chagas disease. Lancet 375, 1388–1402. doi: 10.1016/S0140-6736(10)60061-X
R Core Team. (2015). R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing.
Reisenman, C. E. (2014). Hunger is the best spice: effect of starvation in the antennal responses of the blood-sucking bug Rhodnius prolixus. J. Insect Physiol. 71, 8–13. doi: 10.1016/j.jinsphys.2014.09.009
Rinker, D. C., Pitts, R. J., Zhou, X., Suh, E., Rokas, A., and Zwiebel, L. J. (2013). Blood meal-induced changes to antennal transcriptome profiles reveal shifts in odor sensitivities in Anopheles gambiae. Proc. Natl. Acad. Sci. U.S.A. 110, 8260–8265. doi: 10.1073/pnas.1302562110
Rozen, S., and Skaletsky, H. (2000). Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386. doi: 10.1385/1-59259-192-2:365
Saveer, A. M., Kromann, S. H., Birgersson, G., Bengtsson, M., Lindblom, T., Balkenius, A., et al. (2012). Floral to green: mating switches moth olfactory coding and preference. Proc. Biol. Sci. 279, 2314–2322. doi: 10.1098/rspb.2011.2710
Scharlaken, B., de Graaf, D. C., Goossens, K., Brunain, M., Peelman, L. J., and Jacobs, F. J. (2008). Reference gene selection for insect expression studies using quantitative real-time PCR: the honeybee, Apis mellifera, head after a bacterial challenge. J. Insect Sci. 8:33. doi: 10.1673/031.008.3301
Schmunis, G. A., and Yadon, Z. E. (2010). Chagas disease: a Latin American health problem becoming a world health problem. Acta Trop. 115, 14–21. doi: 10.1016/j.actatropica.2009.11.003
Sengul, M. S., and Tu, Z. (2008). Characterization and expression of the odorant-binding protein 7 gene in Anopheles stephensi and comparative analysis among five mosquito species. Insect Mol. Biol. 17, 631–645. doi: 10.1111/j.1365-2583.2008.00837
Senior, K. (2007). Chagas disease: moving towards global elimination. Lancet Infect. Dis. 7, 572. doi: 10.1016/S1473-3099(07)70194-9
Siju, K., Hill, S. R., Hansson, B. S., and Ignell, R. (2010). Influence of blood meal on the responsiveness of olfactory receptor neurons in antennal sensilla trichodea of the yellow fever mosquito Aedes aegypti. J. Insect Physiol. 56, 659–665. doi: 10.1016/j.jinsphys.2010.02.002
Silbering, A., Rytz, R., Grosjean, Y., Abuin, L., Ramdya, P., Jefferis, G., et al. (2011). Complementary function and integrated wiring of the evolutionarily distinct Drosophila olfactory subsystems. J. Neurosci. 31, 13357–13375. doi: 10.1523/JNEUROSCI.2360-11.2011
Staden, R., Beal, K. F., and Bonfield, J. K. (2000). The Staden package, 1998. Methods Mol. Biol. 132, 115–130. doi: 10.1385/1-59259-192-2:115
Takken, W., Van Loon, J. J., and Adam, W. (2001). Inhibition of host-seeking response and olfactory responsiveness in Anopheles gambiae following blood feeding. J. Insect Physiol. 47, 303–310. doi: 10.1016/S0022-1910(00)00107-4
Vassena, C. V., Picollo, M. I., and Zerba, E. N. (2000). Insecticide resistance in Brazilian Triatoma infestans and Venezuelan Rhodnius prolixus. Med. Vet. Entomol. 14, 51–55. doi: 10.1046/j.1365-2915.2000.00203.x
Vitta, A. C., and Lorenzo, M. G. (2009). Copulation and mate guarding behavior in Triatoma brasiliensis (Hemiptera: Reduviidae). J. Med. Entomol. 46, 789–795. doi: 10.1603/033.046.0409
Vitta, A. C. R., Bohman, B., Unelius, C. R., and Lorenzo, M. G. (2009). Behavioral and electrophysiological responses of Triatoma brasiliensis males to volatiles produced in the metasternal glands of females. J. Chem. Ecol. 35, 1212–1221. doi: 10.1007/s10886-009-9709-2
Vosshall, L. B., and Hansson, B. S. (2011). A unified nomenclature system for the insect olfactory coreceptor. Chem. Senses 36, 497–498. doi: 10.1093/chemse/bjr022
Vosshall, L. B., Wong, A. M., and Axel, R. (2000). An olfactory sensory map in the fly brain. Cell 102, 147–159. doi: 10.1016/S0092-8674(00)00021-0
Ward, J. P. (1981). A comparison of the behavioural responses of the haematophagous bug, Triatoma infestans, to synthetic homologues of two naturally occurring chemicals (n- and iso-butyric acid). Physiol. Entomol. 6, 325–329. doi: 10.1111/j.1365-3032.1981.tb00277.x
Wigglesworth, V. B., and Gillett, J. D. (1934). The function of the antennae in Rhodnius prolixus (Hemiptera) and the mechanism of orientation to the host. J. Exp. Biol. 11, 120–139.
Keywords: olfaction, olfactory coreceptors, triatomines, behavior, physiology
Citation: Latorre-Estivalis JM, Omondi BA, DeSouza O, Oliveira IHR, Ignell R and Lorenzo MG (2015) Molecular basis of peripheral olfactory plasticity in Rhodnius prolixus, a Chagas disease vector. Front. Ecol. Evol. 3:74. doi: 10.3389/fevo.2015.00074
Received: 19 December 2014; Accepted: 23 June 2015;
Published: 10 July 2015.
Edited by:
Astrid T. Groot, University of Amsterdam, NetherlandsReviewed by:
Hong Lei, University of Arizona, USACaroline Marie Nieberding, Université Catholique de Louvain, Belgium
Jonathan Daniel Bohbot, Hebrew University of Jerusalem, Israel
Copyright © 2015 Latorre-Estivalis, Omondi, DeSouza, Oliveira, Ignell and Lorenzo. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jose M. Latorre-Estivalis, Vector Behavior and Pathogen Interaction Group, Centro de Pesquisas René Rachou, Fundação Oswaldo Cruz, Av. Augusto de Lima, 1715, MG CEP: 301900-02, Belo Horizonte, Brazil,am9zbWFudG9ycmVzQGdtYWlsLmNvbQ==
†Present Address: Bonaventure A. Omondi, Consultative Group for International Agricultural Research, Bioversity International, Bujumbura, Burundi
‡These authors have contributed equally to this work.
 Jose M. Latorre-Estivalis
Jose M. Latorre-Estivalis Bonaventure A. Omondi
Bonaventure A. Omondi Og DeSouza
Og DeSouza Ivana H. R. Oliveira1
Ivana H. R. Oliveira1 Rickard Ignell
Rickard Ignell Marcelo G. Lorenzo
Marcelo G. Lorenzo