- Department of Biology, Stanford University, Stanford, CA, USA
A major tool in the evolutionary biologist's kit is to study the repeated emergence of certain biological traits. Employment of this tool has allowed substantial recent advances to be made in understanding the adaptive molecular basis of certain key biological traits. However, behavior, one life's most pervasive, and complex traits, is not one. Here we review the concepts of repeated evolution and how they apply to behavior. We assess the distribution and evolutionary dynamics of known cases of repeated behavioral evolution and examine their prospects for success in identifying the genetic and mechanistic bases of behavior. We propose that studying adaptive radiations, such as that seen amongst the cichlids of Lake Malawi, will likely yield results quickly due to the tractability of genetic and comparative analyses. Finally we suggest some possible scenarios that might be observed in the pursuit of the adaptive molecular basis of behavior and advocate for research on a diverse number of case studies of behavioral evolution, allowing for a knowledge base from which general principles of behavioral evolution might be gleaned.
Introduction
Biologists have long appreciated that the repeated emergence of traits can act as a window into the workings of natural selection and the evolutionary process (Simpson, 1953; Mayr, 1963; Endler, 1986). When similar features occur in unrelated species it suggests a form of regularity in the actions of evolution that are capable of overcoming the unique constraints imposed by each species' unique history (Vermeij, 2006). Accordingly, recurrence in evolution is becoming an increasingly popular tool for analyzing how evolution can act at different levels (e.g., genome, development, molecular pathways, cellular functions) to converge on similar phenotypes. Cases of recurrent evolution recently been used to make inferences about molecular and developmental architectures of certain traits, giving insights into the “predictability” of genetic and phenotypic evolution (Stern and Orgogozo, 2008; Stern, 2013). There are now numerous examples in which lineages possessing recurrently evolved traits have been exploited to identify the genetic basis of morphological, life history, and physiological straits (Gompel and Prud'homme, 2009; Martin and Orgogozo, 2009; Stern and Orgogozo, 2009). We argue that it is time for behavioral biologists to join in.
Researchers are increasingly recognizing that complex behaviors too undergo recurrent evolution, both within closely related adaptive radiations and across distant taxa (Johnson et al., 2009), and presumably involving the action of selection acting one or several of many possible biological pathways, some of which are unique to behavioral traits (e.g., nervous system form and function). We contend that the first steps toward this goal will require identifying and evaluating instances of behavioral evolution that are amenable to integrative study incorporating, molecular, developmental, neurobiological, behavioral, and ecological insights. To this end we examine here a number of reported cases of repeated behavioral evolution and propose that early successes are likely to be found in the study of closely-related, yet highly variable, radiations of species. We next focus on an example from our lab-the evolution of bower building in Lake Malawi cichlid fish-to demonstrate the surprising amount of convergence natural behaviors are capable of achieving amongst these radiations. Finally, we explore biological scenarios possibly unique to behavioral traits that might arise from the study of these systems, with a specific focus on theoretical relationships between the genome, nervous systems, and phenotypes in the repeated evolution of behavior.
Repeated Evolution: Varieties
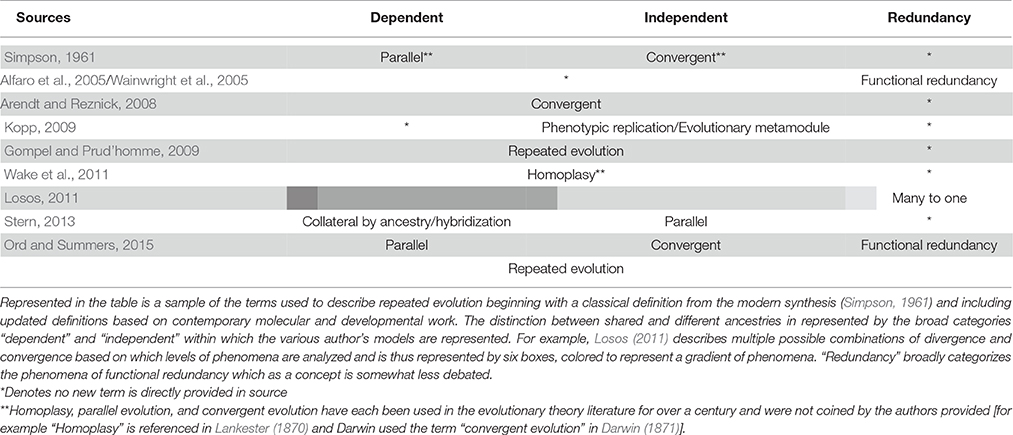
The repeated evolution of traits (“homoplasy”), traditionally construed (Lankester, 1870), can arise through several channels: convergent evolution, parallel evolution, and functional redundancy. Convergent and parallel evolution have been historically discussed together while functional redundancy is a related, but relatively newer concept. To start, the distinction between convergent and parallel evolution historically depends on ancestry and its role in adaptation. A classic outline of this distinction comes from Simpson's Principles of Animal Taxonomy (1961). Here parallel evolution is defined as “independent occurrence of similar changes in groups with a common ancestry and because they had a common ancestry.” In contrast, “convergence is the development of similar characteristics separately in two or more lineages without a common ancestry related to the similarity but involving adaptation to similar ecological status” (Simpson, 1961; Wood et al., 2005). Modern interpretations of this distinction define convergent evolution as occurring when species employ different genetic processes (due to divergent ancestry) to arrive at the same phenotype while parallel evolution produces share phenotypes from the same genetic processes (likely due to shared ancestry).
KEY CONCEPT 1. Repeated evolution
The evolution of traits shared among species that are not possessed by their common ancestor. As used here repeated evolution encompasses a variety of processes and definitions outlined in the main text and Table 1.
KEY CONCEPT 2. Functional redundancy
Convergence on a similar functional outcome through different phenotypic strategies.
Since Simpson much debate has occurred surrounding the relationship between convergent and parallel evolution. This owes largely to the fact that the original definitions predated the advent of modern molecular methods (Losos, 2011; summarized in Table 1). Recently authors have suggested replacing the terms with others, including repeated evolution (Gompel and Prud'homme, 2009; Ord and Summers, 2015), homoplasy (Wake et al., 2011), and phylogenetic replication (Kopp, 2009). Arendt and Reznick (2008), on the other hand, collapse the distinctions altogether under one banner- convergent evolution-arguing that the convergent vs. parallel split represents a false dichotomy. They contend that since closely related species can converge on phenotypes using divergent genetic mechanisms and vice versa it is often difficult, if not impossible, to assign an arrow of causality in assessing convergent and parallel evolution (Arendt and Reznick, 2008). Similarly Losos (2011) demonstrates at least six varieties of convergence/divergence that can occur depending on the level of phenomena analyzed (i.e., developmental pathway, gene function, phenotype, etc.).
This movement toward consideration of recurrence at multiple levels reflects has benefited from advances in whole genome sequencing, phylogenomics, and molecular biology. For example Stern (2013) updates the definitions based on results from modern genetic mapping of convergent traits, suggesting that repeated evolution occurs either through convergence on the same genetic variant through de novo mutation (“parallel evolution”), shared inheritance of an ancestral polymorphism (“collateral evolution”), or through the introduction of an allele into a population through hybrid breeding (“hybridization”) and provides a number of examples from recent work for each. Similarly, other authors have begun to highlight numerous instances of recurrent evolution of genomic features, such as gene loss and duplication, related to certain traits (Maeso et al., 2012). We believe that it is desirable to incorporate such information into our understanding of recurrent evolution and in the development of models describing it. Yet, since very few convergent behaviors have been mapped to causal genetic loci, it is at the moment difficult to discuss repeated behavioral evolution using updated definitions such as these. We instead here opt to use “repeated evolution” to refer to the genetic and phenotypic phenomena that constitute the gradient between parallel and convergent evolution represented by these recent models (but in the final section will discuss the prospects for refining this usage).
Cases from morphology exemplify the diversity of repeated evolution. A prime example is image forming eyes where six types of eye, three with chambers and three that are compound have evolved (Fernald, 2006). The strong constraints imposed by collecting and focusing light have limited the solutions yet the outcomes serve their owners well. Another example is reduced pigmentation that has occurred several times across diverse vertebrate taxa and has led to the occurrence of dark and light morphs within natural populations (reviewed in Gompel and Prud'homme, 2009). Several of these populations have distinct genetic mechanisms driving pigmentation reduction have been identified including Oculocutaneous albinism type 2 (Oca2) in cave fish (Protas et al., 2007), Kit ligand (Kitlg) in sticklebacks, and humans (Miller et al., 2007), and Melanocortin 1 receptor (Mc1r) in a variety of mammals, reptiles, birds, and fish (Hubbard et al., 2010). Here unrelated taxa have arrived at common adaptive phenotypes that have arisen from different genomic contexts but use similar genetic mechanisms. In contrast, the recurrence of pelvic reduction in threespine sticklebacks, a very recent postglacial radiation of freshwater fish species (<12,000 years) that have repeatedly colonized similar niches within North American lakes (Schluter, 2000). Recurrent deletions of the same genomic element underlie the adaptive loss pelvic armor within disparate populations of this radiation (Chan et al., 2010). Here the same genetic mechanism is used to create a common adaptive phenotype.
The final concept, functional redundancy, represents cases in which species can use a variety of different mechanisms to achieve the same functional outcome. In these instances the trait of concern may have been arrived at using totally divergent genetic, molecular, or physiological mechanisms. This is well represented by studies of variation in the skeletal morphology of fish from the family Labridae. Labrid fish species use a complex network of facial muscles for feeding that vary widely and are associated with adaptive patters of prey use (Wainwright et al., 2004). The mechanical output of these muscles is measured by the Maxillary Kinematic Transmission Coefficient (Maxillary KT). Species with high Maxillary KT tend to feed on zooplankton and elusive fishes and shrimps while species with low Maxillary KT tend to feed on more stationary prey (Wainwright et al., 2005). Strikingly, high and low Maxillary KT values, and their associated feeding styles, can be arrived at through very different musclulatures, thus demonstrating that “many” phenotypes might be leveraged to achieved “one” functional outcome (Alfaro et al., 2005; Wainwright et al., 2005). In this sense functional redundancy differs from convergence and parallelism in that the traits and the genetic mechanisms of concern need not be the same but rather the functional consequence of their actions.
Future work will illuminate the relative contributions of each form of repeated evolution in the context of behavior. The first step in this process, though, will be identifying clades that possess substantial variation in behaviors that show evidence of independent evolution. In the next section we assess a number of known examples that seem to fit this bill.
The Landscape of Repeated Behavioral Evolution
If, as we suggest, behavioral biologists are to leverage the repeated evolution of behavior as a research tool what do they have to work with? We surveyed the literature for reports of repeated behavioral evolution, looking for cases that were backed by a phylogeny and in which transitions between behaviors could be identified with some confidence. Assessing repeated behavioral evolution as a function of Phenotypic transitions is useful as it allows inferences to be made about the genetic and adaptive landscapes within which traits have evolved. For example, a behavior that has arisen 30 times in a short time period presumably has very different molecular and evolutionary bases than one that arose only twice in distantly related taxa (the study of both being valuable in the developing a body of evidence of behavioral evolution). We sought to address two questions: First, to what extent has repeated behavioral evolution occurred taxonomically and, second, how is its occurrence distributed across evolutionary time? Answers to these questions could provide insight into the relative impact of different types of repeated evolution (e.g., within closely related species or amongst distantly related taxa) and potentially when and where to expect it.
KEY CONCEPT 3. Phenotypic transitions
An instance of repeated evolution within a phylogenetic context.
In total we collected 49 reported instances of repeated behavioral evolution across 39 taxonomic groups (Table 2). For each instance we recorded the estimated clade size, the hypothesized mechanism of repeated evolution, and the divergence time between representative species. The divergence times and taxonomic distances of the collected traits span wide distances, from ecological time scales within populations (e.g., Trinidadian guppies; Schwartz and Hendry, 2007) to >800 MYA splits between kingdoms (e.g., Mussels/tunicates; Koehl, 1996). In addition repeated evolution has occurred for a variety of behaviors, from social organization in fire and alpine silver ants (Purcell et al., 2014) to individual motor ability in Hawaiian gobies (Blob et al., 2006), and can be seen across animal phyla.
Plotting phenotypic transitions and divergence time reveals the landscape of repeated behavioral evolution (Figure 1). The first major feature of the landscape is that there are several of instances or repeated evolution with more than 10 conversions including the evolution of bower building in cichlid fish (likely >10 conversions; York et al., 2015) and eusociality across inverterbrates and vertebrate taxa (17 conversions; Duffy et al., 2000). A density distribution of repeated behavioral evolution instances across evolutionary time (Figure 1B) shows a skew toward more recent evolutionary events with the majority of <100 MYA (39/49; 79.59%). Furthermore, more than 80% (40/49) of cases occur at sub-family taxonomic units (species/genus/subfamily/family). Previous observations of repeated evolutionary events across other traits including morphology, physiology, and life history shows a similar trend, indicating that behaviors are not subject to evolutionary processes distinct from these other phenotypes (Vermeij, 2006; Ord and Summers, 2015).
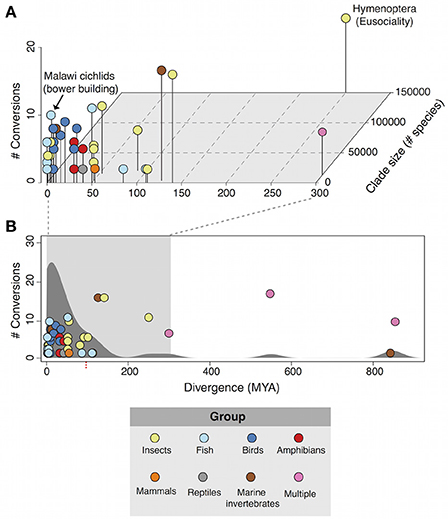
Figure 1. The landscape of repeated behavioral evolution. (A) 3D scatterplot of the number of transitions, divergence time (millions of years), and clade size (number of species) for cases of repeated behavioral evolution that have occurred within 300 MYA. Points are colored by taxonomic grouping. (B) The total distribution of all 49 cases of repeated behavioral evolution identified in this review. A kernel density distribution of the behaviors is plotted behind scatterplot, highlighting a strong skew toward behaviors that have evolved in <100 million years.
In addition we find a significant positive correlation between clade size and conversion number (r2 = 0.46; p = 0.001) but not between evolutionary divergence time and conversion (r2 = 0.11; p = 0.46). This suggests that the probability of repeated behavioral evolution occurring may be driven more by clade size rather than evolutionary time. This point is may have some bearing on the consideration of when and where repeated behavioral evolution is expected to occur since evolution can produce speciose clades in a variety of ways and on a number of time scales, a particularly striking example of which is the case of adaptive radiations (discussed below). If a major determinant of repeated behavior evolution is the sheer number of species a related group then it may be reasonable to expect an enrichment of repeated evolution amongst rapidly radiating groups of species.
It is important to note, though, that at the current moment it is difficult to disassociate the observed patterns from the influence of ascertainment bias and the complexity of identifying behavioral homologies between distantly related taxa. For example, it is possible that the current known cases of repeated behavioral evolution were arrived at due to their ease of study, be it through a ready ability to identify behavioral homologs or through rapid observation and measurement of the traits. Furthermore, there is a presumable multitude of other repeated behavioral evolution instances that we have not included here due to a current lack of knowledge regarding phylogenetic relationships and behavioral definitions. Notable in this list is the perceived repeated evolution of complex social behaviors across animals, especially in relation to the structures and strategies of parental care (Hofmann et al., 2014). Future work resolving the phylogenetic, molecular, and phenotypic bases of complex behaviors such as these will be illuminating and will aid in identifying the extent to which behavioral homologies may exist across taxa. To this end, and influenced by the history of identifying such phenomena in morphology, we propose that early successes may come if a focus is placed on clades that are speciose, diverse, and closely related, qualifications that are well met by the occurrence of adaptive radiations.
Adaptive Radiations as a First Step
Adaptive radiations (ARs) provide examples of the extreme “parallel” end of the repeated evolution spectrum. ARs are rapidly evolving lineages that arose from a common ancestral source and display substantial ecological and phenotypic diversification (Simpson, 1953; Schluter, 2000). Repeated evolution within adaptive radiations is presumably often due to convergence on a particular allele of an ancestral polymorphism, as is the case in the evolution of stickleback body armor (Colosimo et al., 2005). This need not always be the case and indeed examples exist of closely related populations using different genetic and developmental pathways to reach the same phenotype as has occurred in differentially pigmented populations of oldfield mice (Hoekstra et al., 2006) and cave fish (Wilkens and Strecker, 2003). A common method for conceptualizing macroevolutionary change in ARs is the “radiation in stages” model (Streelman and Danley, 2003; Gavrilets and Losos, 2009). The model proposes that species diversification in ARs occurs in the order of (i) macrohabitat, (ii) microhabitat or trophic (feeding) specializations, and (iii) species-specific traits that lead to nonrandom mating (i.e., signals such as nuptial coloration and courtship behavior).
KEY CONCEPT 4. Adaptive radiations (ARs)
The rapid diversification of a species arising from a common ancestral source, often due to novel environmental pressures and opportunities.
In York et al. (2015) we provide evidence that amongst the cichlid fish of Lake Malawi, Africa adaptions arising from stages (i) (macrohabitat) and (ii) (trophic style) have influenced the evolution of a courtship signal: the construction of sand mating nests or “bowers.” Males of more than 100 species within the sand-dwelling lineage of Malawi cichlids seasonally build bowers in large aggregations (“leks”) solely in order to attract and mate with females, a purpose almost identical to that of the eponymous bowerbirds of Oceania (Diamond, 1986; McKaye et al., 2001; Magalhaes et al., 2013). Bowers are extended phenotypes that can exist in two basis form: “pits”-depressions dug out of the sand substrate-and “castles”-aggregations of sand collected from around the bower area and deposited to create a mound (Figure 2A; Dawkins, 1992; York et al., 2015). Phylogenetic analysis shows that pit and castle type bowers are evolutionarily labile. A partial phylogeny of 75 species from the sand-dwelling clade over 20 conversions between pit and castle are present. The number of phenotypic transitions of bower type is currently speculative as the current tree only represents a subset of the sand-dwelling lineage (200+ species in total). Whole-genome phylogenetic analyses will help in the resolution of within-genera variation of bower type. Nevertheless, the pervasiveness of bower building in this clade and its phylogenetic labiality is a striking example of the extremes of courtship behavior diversity.
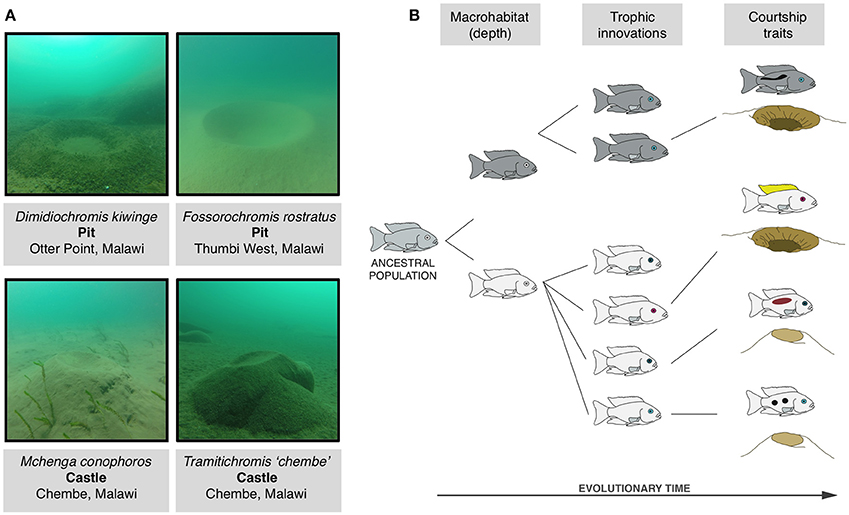
Figure 2. Behavioral evolution amongst Malawi cichlids. (A) Photos of example pit and castle type bowers taken at Lake Malawi during behavioral observations using SCUBA. (B) The evolution of bower building in the context of the radiation in stages model (Streelman and Danley, 2003; York et al., 2015). First the ancestral population diverges in macrohabitat-here depth of occurrence-leading to associated adaptations (represented by body color). Next innovations associated with feeding and life history arise such as variations in jaw and facial morphology and visual acuity (e.g., eye opsin expression, coded here by differences in eye color). Finally species diverge in signals used for courtship and reproduction, graphically represented by male coloration and differences in bower type.
Why then did pit and castle type bowers repeatedly evolve? Like all cases of repeated evolution there are at least two levels at which this question can be addressed. The first is concerned with the genetic and molecular mechanisms that produced the repeated evolution. The second is concerned with the ecological and evolutionary pressures that facilitated it. Addressing the role of genetic constraints on bower type is currently difficult (though efforts are underway to clarify this issue). On the other hand there is some clarification regarding the roles of ecology and phenotype in the evolution of bower building. First, bower type correlates with macrohabitat (depth of occurrence) wherein castle-building species tend to occur at significantly shallower depths than pit-digging species. Second, bower type is associated with differences in functional morphology and eye opsin expression, suggesting that adaptations for specific diets, and trophic environments influence the propensity to build a pit or a castle (York et al., 2015). Broadly speaking castle-building species tend be shallow-living, possess vision acuities sensitive to UV-rich light environments, and are adapted to extracting mobile food sources from the water column while pit-digging species are distributed at deeper depths, do not have UV vision, and tend to extract food through biting and scraping (Figure 2B).
Given the observed convergence in diverse traits across macrohabitat, morphology, and behavior amongst Malawi cichlids it is tempting to consider their evolution as having occurred on a macroevolutionary adaptive landscape with multiple peaks, each influencing and constraining the overall phenotypic makeup of individual species (Gavrilets and Vose, 2005). In such a case constraints imposed by the landscape will influence which species or lineages may or may not evolve certain traits. This has been observed in the evolution of the entire East African cichlid radiation variation in which both sexually selected traits and a variety of ecological factors predict whether or not species and lineages will radiate, indicating that certain clades may more prone to diversification and the evolution of specific traits than others (Wagner et al., 2012). This pattern suggests that similar adaptive landscapes exist across the lakes of Eastern Africa and has led to a process of replicated adaptive radiations in which common ecological pressures have led to phenotypic convergence across a number of them.
Other well-documented cases of such replicated ARs featuring behavioral convergence specifically have occurred in North American sticklebacks (Rundle et al., 2000), Caribbean Anolis lizards (Mahler et al., 2013), and Hawaiian spiders (Blackledge and Gillespie, 2004). Ecomorphs (phenotypes associated with distinct ecological niches) of Hawaiian orb-weaving spiders of the genus Tetragnatha show convergence of web architecture across the Hawaiian islands. Replicated ARs may hold great promise for the study of the behavioral evolution since they allow inferences to be made about the relative importance of non-behavioral factors (e.g., morphology, environment) on the phenotype of interest. Furthermore, ARs with cases of repeated evolution may offer unique opportunities to leverage population genetic methods for identifying the genomic basis of these traits, as for example has occurred for cichlid egg spots (Santos et al., 2014) and butterfly wing pattern (Reed et al., 2011) but has so far largely neglected physiological, life history, and behavioral traits (Berner and Salzburger, 2015).
Future Prospects for Behavior Evolution Research
Using the systems such as the ones described above researchers may soon begin to identify the mechanistic (genomic, molecular, phenotypic) bases of behavioral evolution. It will be illuminating to examine how these results compare to those obtained for other traits and to assay the extent to which behaviors are unique in their evolutionary patterns. We anticipate that one such idiosyncrasy will be the unique evolutionary relationship between the nervous system and the behaviors it produces. This is because, when selection acts on a behavioral phenotype, the genetic variants associated with that behavior most likely have their effects through nervous system structure and function. Barring future studies directly linking a gene with a specific behavioral phenotype it appears likely that this trend will become the rule.
The fact that behaviors are associated with variation in both the nervous system and the specific behavior trait itself differentiates them from other traits and may be unique to their evolutionary process. There may be several possible avenues through which adaptive variants in the genome may sculpt behavior. Using models derived from known cases of the molecular evolution of morphological and life history traits, one can imagine hypothetical of cases parallel and collateral evolution (terminology from Stern, 2013) within a closely related group of species as examples. It is possible to schematize these scenarios (Figure 3). The first example demonstrates a case where two species have evolved a common behavioral phenotype arising from an identical genetic mutation (parallel evolution) that has its effect through the same neural mechanism (Figure 3A). It is also theoretically possible that, while both species share the same genotype and behavior, due to other evolutionary processes the neural mechanism associated with the behavior differs (Figure 3B). This case of parallel evolution may be seen as also incorporating an aspect of functional redundancy much like that seen in the feeding styles of Labrid fish discussed above. The same dichotomy between nervous system/behavioral convergence and divergence with functional redundancy can also be considered for collateral evolution, either arising from shared ancestry or hybridization (Figures 3C,D; Stern, 2013).
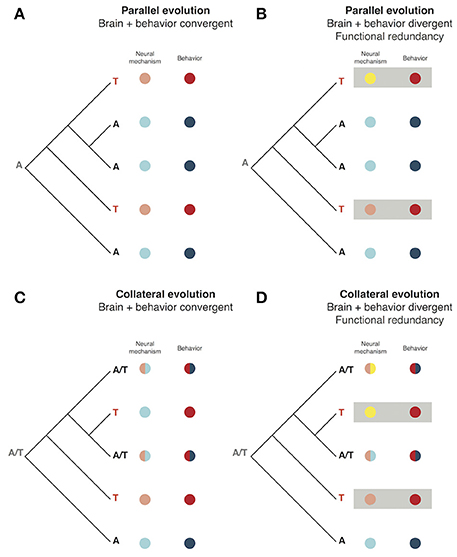
Figure 3. The phenotypic interface of brain and behavior in evolution. (A) A cartoon phylogeny of 5 related species in which a variant producing a novel behavior (“T”) has independently arisen twice through parallel evolution. This variant also leads to homologous changes in the neural mechanism used to produce the behavior. (B) Parallel evolution of a behavior with the same phylogeny as (A) but in this example the neural mechanisms associated with the independently evolved behavior differ. This produces an instance of functional redundancy where the same behavioral output is achieved by divergent mechanisms. (C) Collateral evolution of a behavior amongst 5 closely related species. An ancestral polymorphism (“A/T”) leads to variation in the 5 descents wherein some have an intermediate genotype, neural mechanism, and behavior while others are homozygous for either A or T, leading to independent evolution of the same novel behavior as in (A). (D) Functional redundancy of brain/behavior in the context of collateral evolution.
When taking into account other biological processes such as development and gene function it is likely that the evolution of behaviors will not fall into this clean of a dichotomy (see Losos, 2011). What will be needed is the aggregation of case studies that tract the evolution of diverse behaviors across taxa and through time, from the genome through development to brain function and ultimately behavior. With these empirical data in hand researchers may begin outline general integrative principles of how and when behaviors evolve, thus allowing this trait to take its place alongside others in the pantheon of evolutionary analysis.
Author Contributions
RY conceived of the manuscript idea and performed meta-analyses. RY and RF wrote the manuscript.
Funding
Supported by a Stanford Bio-X Bowes Fellowship (RY), NIGMS 101095, and NINDS 034950 (RF).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The reviewer ED and handling Editor declared their shared affiliation, and the handling Editor states that the process nevertheless met the standards of a fair and objective review.
Acknowledgments
We thank members of the Fernald lab, Austin Hilliard, Joshua Lillvis, and David Stern for useful comments.
Author Biography

References
Alfaro, M. E., Bolnick, D. I., and Wainwright, P. C. (2005). Evolutionary dynamics of complex biomechanical systems: an example using the four-bar mechanism. Evolution 58, 495–503. doi: 10.1111/j.0014-3820.2004.tb01673.x
Aliabadian, A., Kaboli, M., Förschler, M. I., Nijman, V., Chamani, A., Tillier, A., et al. (2012). Convergent evolution of morphological and ecological traits in the open-habitat chat complex (Aves, Muscicapidae: Saxicolinae). Mol. Phylogenet. Evol. 65, 35–45. doi: 10.1016/j.ympev.2012.05.011
Arendt, J., and Reznick, D. (2008). Convergence and parallelism reconsidered: what have we learned about the genetics of adapation? Trends. Ecol. Evol. 23, 26–32. doi: 10.1016/j.tree.2007.09.011
Berner, D., and Salzburger, W. (2015). The genomics of organismal diversification illuminated by adaptive radiations. Trends Genet. 31, 491–499. doi: 10.1016/j.tig.2015.07.002
Blackledge, T. A., and Gillespie, R. G. (2004). Convergent evolution of behavior in an adaptive radiation of Hawaiian web-building spiders. Proc. Natl. Acad. Sci. U.S.A. 101, 16228–16233. doi: 10.1073/pnas.0407395101
Blob, R. W., Rai, R., Julius, M. L., and Schoenfuss, H. L. (2006). Functional diversity in extremem environments: effects of locomotor style and substrate texture on the waterfall-climbing performance of Hawaiian gobiid fishes. J. Zool. 288, 315–324. doi: 10.1111/j.1469-7998.2005.00034.x
Bullock, T. H., Behrend, K., and Heiligenberg, W. (1975). Comparison of the jamming avoidance responses in gymnotid and gymnarchid electric fish: a case of convergent evolution of behavior and its sensory basis. J. Comp. Physiol. 103, 97–121.
Carlson, B. A., Hasan, S. M., Hollmann, M., Miller, D. B., Harmon, L. J., and Anergard, M. E. (2011). Brain evolution triggers increased diversification of electric fishes. Science 332, 583–585. doi: 10.1126/science.1201524
Cervo, R., Macninai, V., Dechigi, F., and Turillazzi, S. (2004). Fast growth of immature brood in a social parasite wasp: a convergent evolution between avian and insect cuckoos. Am. Nat. 164, 814–820. doi: 10.1086/425987
Chan, Y. F., Marks, M. E., Jones, F. C., Villarreal, G. Jr., Shapiro, M. D., Brady, S. D., et al. (2010). Adaptive evolution of pelvic reduction in sticklebacks by recurrent deletion of a Pitx1 enhancer. Science 327, 302–305. doi: 10.1126/science.1182213
Colosimo, P. F., Hosemann, K. E., Balabhadra, S., Villarreal, G. Jr., Dickson, M., Grimwood, J., et al. (2005). Widespread parallel evolution in Sticklebacks by repeated fixation of Ectodysplasin alleles. Science 307, 1928–1933. doi: 10.1126/science.1107239
Danforth, B. N., Sipes, S., Fang, J., and Brady, S. G. (2006). Recent and simultaneous origins of eusociality in halictid bees. Proc. Natl. Acad. Sci. U.S.A. 103, 15118–15123. doi: 10.1073/pnas.0604033103
Diamond, J. (1986). Animal art: variation in bower decorating style among male bowerbirds Amblyornis inornatus. Proc. Natl. Acad. Sci. U.S.A. 83, 3042–3046.
Duboué, E. R., Keene, A. C., and Borowsky, R. L. (2011). Evolutionary convergence on sleep loss in cavefish populations. Curr. Biol. 21, 671–676. doi: 10.1016/j.cub.2011.03.020
Duffy, J. E., Morrison, C. L., and Ríos, R. (2000). Multiple origins of eusociality among sponge-dwelling shrimps (Synalpheus). Evolution 54, 503–516. doi: 10.1111/j.0014-3820.2000.tb00053.x
Elipot, Y., Hinaux, H., Callebert, J., and Rétaux, S. (2013). Evolutionary shift from fighting to foraging in blind cavefish through changes in the serotonin network. Curr. Biol. 23, 1–10. doi: 10.1016/j.cub.2012.10.044
Fernald, R. D. (2006). Casting a genetic light on the evolution of eyes. Science 313, 1914–1918. doi: 10.1126/science.1127889
Frédérich, B., Sorenson, L., Santini, F., Slater, G. J., and Alfaro, M. E. (2013). Iterative ecological radiation and convergence during the evolution history of damselfishes (Pomacentridae). Am. Nat. 181, 94–113. doi: 10.1086/668599
Gavrilets, S., and Losos, J. B. (2009). Adaptive radiation: contrasting theory with data. Science 323, 732–737. doi: 10.1126/science.1157966.
Gavrilets, S., and Vose, A. (2005). Dynamics patterns of adaptive radiation. Proc. Natl. Acad. Sci. U.S.A. 102, 18040–18045. doi: 10.1073/pnas.0506330102
Ge, D., Chesters, D., Gómez-Zurita, J., Zhang, L., Yang, X., and Vogler, A. P. (2011). Anti-predator defence drive parallel morphological evolution in flea beetles. Proc. Biol. Sci. 278, 2133–2141. doi: 10.1098/rspb.2010.1500
Gompel, N., and Prud'homme, B. (2009). The causes of repeated genetic evolution. Dev. Biol. 332, 36–47. doi: 10.1016/j.ydbio.2009.04.040
Haase, M. (2005). Rapid and convergent evolution of parental care in hydrobiid gastropods from New Zealand. J. Evol. Biol. 18, 1076–1086. doi: 10.1111/j.1420-9101.2005.00894.x
Hall, Z. J., Street, S. E., Auty, S., and Healy, S. D. (2015). The coevolution of building nests on the ground and domed nests in Timaliidae. Auk 132, 584–593. doi: 10.1642/AUK-15-23.1
Hedges, S. B., Dudley, J., and Kumar, S. (2006). TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22, 2971–2972. doi: 10.1093/bioinformatics/btl505
Henry, C. S., Wells, M. L. M., and Simon, C. M. (1999). Convergent evolution of courtship songs among cryptic species of the Carnea group of green lacewings (Neuroptera: Chrysopidae: Chrysoperla). Evolution 53, 1165–1179.
Hoekstra, H. E., Hirschmann, R. J., Bundey, R. A., Insel, P. A., and Crossland, J. P. (2006). A single amino acid mutation contributes to adaptive beach mouse color pattern. Science 313, 101–104. doi: 10.1126/science.1126121
Hofmann, H. A., Beery, A. K., Blumstein, D. T., Couzin, I. D., Earley, R. L., Hayes, D. L., et al. (2014). An evolutionary framework for studying mechanisms of social behavior. Trends Ecol. Evo. 29, 581–589. doi: 10.1016/j.tree.2014.07.008
Hubbard, J. K., Uy, J. A. C., Hauber, M. E., Hoekstra, H. E., and Safran, R. J. (2010). Vertebrate pigmentation: from underlying genes to adapative function. Trends Genet. 26, 231–239. doi: 10.1016/j.tig.2010.02.002
Irestedt, M., Fjeldså, J., Dalén, L., and Ericson, G. P. (2009a). Convergent evolution, habitat shifts, and variable diversification rates in the ovenbird-woodcreeper family (Furnariidae). BMC Evol. Biol. 9:268. doi: 10.1186/1471-2148-9-268
Irestedt, M., Jønsson, K. A., Fjeldså, J., Christidis, L., and Ericson, G. P. (2009b). An unexpectedly long history of sexual selection in birds-of-paradise. BMC Evol. Biol. 9:235. doi: 10.1186/1471-2148-9-235
Jarvis, J. U. M., O'Riain, M. J., Bennett, N. C., and Sherman, P. W. (1994). Mammalian eusociality: a family affair. Tree 9, 47–51.
Johnson, M. A., Revell, L. J., and Losos, J. B. (2009). Behavior convergence and adaptive radiation: effects of habitat use on territorial behavior in Anolis lizards. Evolution 64, 1151–1159. doi: 10.1111/j.15558-5646.2009.00881.x
Jones, G., and Teeling, E. C. (2006). The evolution of echolocation in bats. Trends Ecol. Evol. 21, 149–156. doi: 10.1016/j.tree.2006.01.001
Köhler, F., and Günther, R. (2008). The radiation of microhlyid frogs (Amphibia: Anura) on New Guinea: a mitochondrial phylogeny reveals parallel evolution of morphological and life history traits and disproves the current morphology-based classification. Mol. Phylogenet. Evol. 47, 353–365. doi: 10.1016/j.ympev.2007.11.032
Kopp, A. (2009). Metamodels and phylogenetic replication: a systematic approach to the evolution of developmental pathways. Evolution. 63, 2771–2789. doi: 10.1111/j.1558-5646.2009.00761.x
Kowalko, J. E., Rohner, N., Rompani, S. B., Peterson, B. K., Linden, T., Yoshizawa, M., et al. (2013). Loss of schooling behavior in cavefish through sight-dependent and sight-independent mechanisms. Curr. Biol. 23, 1874–1883. doi: 10.1016/j.cub.2013.07.056
Laiolo, P. (2012). Interspecific interactions drive cultural co-evolution and acoustic donvergence in syntopic species. J. Anim. Ecol. 81, 594–604. doi: 10.1111/j.1365-2656.2011.01946.x
Lankester, E. R. (1870). On the use of the term homology in modern zoology and the distinction between homogenetic and homoplastic agreements. Ann. Mag. Nat. Illist. 4, 34–43.
Lillvis, J. L., and Katz, P. S. (2013). Parallel evolution of serotonergic neuromodulation underlies independent evolution of rhythmic motor behavior. J. Neurosci. 33, 2709–2717. doi: 10.1523/JNEUROSCI.4196-12.2013
Losos, J. B. (2011). Convergence, adaptation, and constraint. Evolution 65, 1827–1850. doi: 10.1111/j.1558-5646.2011.01289.x
Maeso, I., Roy, S. W., and Irimia, M. (2012). Widespread recurrent evolution of genomic features. Genome Bio Evol. 4, 486–500. doi: 10.1093/gbe/evs022. Epub 2012 Mar 13.
Magalhaes, I. S., Croft, G. E., and Joyce, A. J. (2013). Altering an extended phenotype reduces intraspecific male aggression and can maintain diversity in cichlid fish. PeerJ. 1:e209. doi: 10.7717/peerj.209
Mahler, D. L., Ingram, T., Revell, L. J., and Losos, J. B. (2013). Execptional convergence on the macroevolutionary landscape in island lizard radiations. Science 341, 292–295. doi: 10.1126/science.1232392
Markow, T., and O'Grady, P. M. (2005). Evolutionary genetics of reproductive behavior in Drosophila: connecting the dots. Annu. Rev. Genet. 39, 263–291. doi: 10.1146/annurev.genet.39.073003.112454
Martin, A., and Orgogozo, V. (2009). The loci of repeated evolution: a catalog of genetic hotspots of phenotypic variation. Evolution 67, 1235–1250. doi: 10.1111/evo.12081
McKaye, K. R., Stauffer, J. R., Turner, G. F., Konings, A., and Sato, T. (2001). Fishes, as well as birds, build bowers. J. Aquaricult. Aquat. Sci. 9, 121–133.
Miller, C. T., Beleza, S., Polen, A. A., Schluter, D., Kittles, R. A., Shriver, M. D., et al. (2007). cis-Regulatory changes in Kit ligand expression and parallel evolution of pigmentation in sticklebacks and humans. Cell 131, 1179–1189. doi: 10.1016/j.cell.2007.10.055
Miller, J. A. (2007). Repeated evolution of male sacrifice behavior in spiders correlated with genital mutilation. Evolution 61, 1301–1315. doi: 10.1111/j.1558-5646.2007.00115.x
Ödeen, A., and Björklund, M. (2003). Dynamics in the evolution of sexual traits: losses and gains, radiation and convergence in yellow wagtails (Motacilla flava). Mol. Ecol. 12, 2113–2130. doi: 10.1046/j.1365-294X.2003.01883.x
Ord, T. J., Stamps, J. A., and Losos, J. B. (2013). Convergent evolution in the territorial communication of a class adaptive radiation: Carribbean Anolis lizards. Anim. Behav. 85, 1415–1426. doi: 10.1016/j.anbehav.2013.03.037
Ord, T. J., and Summers, T. C. (2015). Repeated evolution and the impact of evolutionary history on adaptation. BMC Evol. Biol. 15:137. doi: 10.1186/s12862-015-0424-z
Palkovacs, E. P., Dion, K. B., Post, D. M., and Caccone, A. (2008). Independent evolution of landlocked alewife populations and rapid, parallel evolution of phenotypic traits. Mol. Ecol. 17, 582–597. doi: 10.1111/j.1365-294X.2007.03593.x
Petkov, C. I., and Jarvis, E. D. (2012). Birds, primates, and spoken language origins: behavioral phenotypes and neurobiological substrates. Front. Evol. Neurosci. 4:12. doi: 10.3389/fnevo.2012.00012
Plath, M., and Schlupp, I. (2008). Parallel evolution leads to reduced shoaling behavior in two cave dwelling populations of Atlantic mollies (Poecilia Mexicana, Poeciliidae, Teleostei). Environ. Biol. Fishes 82, 289–297. doi: 10.1007/s10641-007-9291-9
Price, J. J., Friedman, N. R., and Omland, K. E. (2007). Song and plumage evolution in the new world orioles (Icterus) show similar lability and convergence in patterns. Evolution 61, 850–863. doi: 10.1111/j.1558-5646.2007.00082.x
Protas, M., Conrad, M., Gross, J. B., Tabin, C., and Borowsky, R. (2007). Regressive evolution of the Mexican cave tetra, Astyanax mexicanus. Curr. Biol. 17, 452–454. doi: 10.1016/j.cub.2007.01.051
Prum, R. O. (1990). Phylogenetic analysis of the evolution of display behavior in the Neotropical manakins (Aves: Pipridae). Ethology 84, 202–231.
Purcell, J., Brelsford, A., Wurm, Y., Perrin, N., and Chapuisat, M. (2014). Convergent genetic architecture underlies social organization in ants. Curr. Biol. 24, 2728–2732. doi: 10.1016/j.cub.2014.09.071
Reed, R. D., Papa, R., Martin, A., Hines, H. M., Counterman, B. A., Pardo-Diaz, C., et al. (2011). optix drives the repeated convergent evolution of butterfly wing pattern mimicry. Science 333, 1136–1141. doi: 10.1126/science.1208227
Rundle, H. D., Nagel, L., Boughman, J. W., and Schluter, D. (2000). Natural selection and parallel speciation in sympatric sticklebacks. Science 287, 306–308. doi: 10.1126/science.287.5451.306
Santos, M. E., Braasch, I., Boileau, N., Meyer, B. S., Sauteur, L., Bohne, A., et al. (2014). The evolution of cichlid fish egg-spots is linked with a cis-regulatory change. Nat. Commun. 5:5149. doi: 10.1038/ncomms6149
Schluter, D. (2000). Ecological character displacement in adaptive radiation. Am. Natural. 156, S4–S16. doi: 10.1086/303412
Schwartz, A. K., and Hendry, A. P. (2007). A test for the parallel co-evolution of male colour and female preference in Trinidadian guppies (Poecilia reticulata). Evol. Ecol. Res. 9, 71–90.
Soria-Carrasco, V., Gompert, Z., Comeault, A. A., Farkas, T. E., Parchman, T. L., Johnston, J. S., et al. (2014). Stick insect genomes reveal natural selection's role in parallel speciation. Science 344, 738–742. doi: 10.1126/science.1252136
Stern, D. L. (1994). A phylogenetic analysis of soldier evolution in the aphid family Hormaphididae. Proc. Biol. Sci. 256, 203–209.
Stern, D. L. (2013). The genetic causes of convergent evolution. Nat. Rev. Genet. 14, 751–764. doi: 10.1038/nrg3483
Stern, D. L., and Orgogozo, V. (2008). The loci of evolution: how predictable is genetic evolution? Evolution 62, 2155–2177. doi: 10.1111/j.1558-5646.2008.00450.x
Stern, D. L., and Orgogozo, V. (2009). Is genetic evolution predictable? Science 323, 746–751. doi: 10.1126/science.1158997
Stoks, R., Nystrom, J. L., May, M. L., and McPeek, M. A. (2005). Parallel evolution in ecological and reproductive traits to produce cryptic damselfly species across the Holarctic. Evolution 59, 1976–1988. doi: 10.1111/j.0014-3820.2005.tb01067.x
Streelman, J. T., and Danley, P. D. (2003). The stages of vertebrate evolutionary radiation. Trends Ecol. Evol. 18, 126–131. doi: 10.1016/S0169-5347(02)00036-8
Summers, K., and Tumulty, J. (2014). “Parental care, sexual selection, and mating systems in neotropical poison frogs,” in Sexual Selection. Perspectives and Models from the Neotropics, eds G. Machado and R. H. Macedo (London: Academic Press), 289–320.
Vermeij, G. J. (2006). Historical contingency and the purported uniqueness of evolutionary innovations. Proc. Natl. Acad. Sci. U.S.A. 103, 1804–1809. doi: 10.1073/pnas.0508724103
Wainwright, P. C., Alfaro, M. E., Bolnick, D. I., and Hulsey, C. D. (2005). Many-to-one mapping of form to function: a general principle in organismal design? Integr. Comp. Biol. 45, 256–262. doi: 10.1093/icb/45.2.256
Wainwright, P. C., Bellwood, D. R., Westneat, M. W., Grubich, J. R., and Hoey, A. S. (2004). A functional morphospace for the skull of labrid fishes: Patterns of diversity in a complex biomechanical system. Biol. J. Linnean. Soc. 82, 1–25. doi: 10.1111/j.1095-8312.2004.00313.x
Wagner, C. E., Harmon, L. J., and Seehausen, O. (2012). Ecological opportunity and sexual selection together predict adaptive radiation. Nature 487, 366–369. doi: 10.1038/nature11144
Wake, D. C., Wake, M. H., and Specht, C. D. (2011). Homoplasy: from detecting pattern to determining process and mechanisms of evolution. Science 331, 1032–1035. doi: 10.1126/science.1188545
Wilkens, H., and Strecker, U. (2003). Convergent evolution of the cavefish Astyanax (Characidae: Teleostei): Genetic evidence from reduced eye-size and pigmentation. Biol. J. Linn. Soc. 80, 545–554. doi: 10.1111/j.1095-8312.2003.00230.x
Wood, H. M., Parkinson, D. Y., Griswold, C. E., Gillespie, R. G., and Elias, D. O. (2016). Repeated evolution of power-amplified predatory strikes in trap-jaw spiders. Curr. Biol. 26, 1057–1061. doi: 10.1016/j.cub.2016.02.029
Wood, T. E., Burke, J. M., and Rieseberg, L. H. (2005). Parallel genotypic adaptation: when evolution repeats itself. Genetica 123, 157–170. doi: 10.1007/s10709-003-2738-9
Woodward, S. H., Fischman, B. J., Venkat, A., Hudson, M. E., Varala, K., Cameron, S. A., et al. (2011). Genes involved in convergent evolution of eusociality in bees. Proc. Natl. Acad. Sci. U.S.A. 108, 7472–7477. doi: 10.1073/pnas.1103457108
Keywords: animal behavior, behavioral evolution, malawi cichlids, adaptive radiations, repeated evolution
Citation: York RA and Fernald RD (2017) The Repeated Evolution of Behavior. Front. Ecol. Evol. 4:143. doi: 10.3389/fevo.2016.00143
Received: 18 May 2016; Accepted: 15 December 2016;
Published: 31 January 2017.
Edited by:
Jordi Figuerola, Estación Biológica de Doñana, SpainReviewed by:
Elena Gómez-Díaz, Estación Biológica de Doñana, SpainRaphaël Boulay, Université François-Rabelais, France
Copyright © 2017 York and Fernald. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: cnlhbnlvcmtAc3RhbmZvcmQuZWR1
 Ryan A. York
Ryan A. York Russell D. Fernald
Russell D. Fernald