- 1Department of Bioinformatics, Hans Raj Mahila Maha Vidyalaya, Punjab, India
- 2Bioclues.org, Hyderabad, India
- 3Centre for Bioinformatics, School of Life Sciences, Pondicherry University, Puducherry, India
- 4Department of Biological Sciences, Indian Institute of Science Education and Research, Kolkata, India
- 5Department of Biotechnology and Bioinformatics, Birla Institute of Scientific Research, Jaipur, India
- 6Vignan’s Foundation for Science, Technology & Research (Deemed to be University), Guntur, India
- 7Department of Chemistry, School of Basic Sciences, Manipal University Jaipur, Jaipur, India
- 8Functional Genomics Unit, Council of Scientific and Industrial Research- Institute of Genomics & Integrative Biology (CSIR-IGIB), Delhi, India
- 9Department of Biotechnology, Haldia Institute of Technology, West Bengal, India
- 10Department of Microbiology, All India Institute of Medical Sciences, Bibinagar, Hyderabad, India
- 11Genomix CARL Pvt. Ltd, Pulivendula, India
- 12Department of Computer Science, Flame University, Pune, India
- 13Cytogenetics Laboratory, Department of Zoology, Benaras Hindu University, Varanasi, India
- 14Department of Genetics, University of Alabama, Birmingham, AL, United States
- 15German Cancer Research Centre (DKFZ), Heidelberg, Germany
- 16Department of Applied Biology, Council of Scientific and Industrial Research-Indian Institute of Chemical Technology (CSIR-IICT), Hyderabad, India
- 17Amrita School of Biotechnology, Amrita Vishwa Vidyapeetham, Kerala, India
The year 2019 has seen an emergence of the novel coronavirus named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) causing coronavirus disease of 2019 (COVID-19). Since the onset of the pandemic, biological and interdisciplinary research is being carried out across the world at a rapid pace to beat the pandemic. There is an increased need to comprehensively understand various aspects of the virus from detection to treatment options including drugs and vaccines for effective global management of the disease. In this review, we summarize the salient findings pertaining to SARS-CoV-2 biology, including symptoms, hosts, epidemiology, SARS-CoV-2 genome, and its emerging variants, viral diagnostics, host-pathogen interactions, alternative antiviral strategies and application of machine learning heuristics and artificial intelligence for effective management of COVID-19 and future pandemics.
Origin, Taxonomy, and Hosts of Coronaviruses
Coronaviruses (CoVs) are single-stranded RNA viruses of size around 65-125 nm in diameter. Due to the presence of crown-like spike structure on the viral outer surface, they were named Coronaviruses (1). The CoVs are known to cause respiratory and gastrointestinal tract infections in humans, poultry, and animals (2, 3). These are a large group of viruses that belong to the order Nidovirales, family Coronaviridae, and subfamily Orthocoronavirinae, which in turn is divided into four genera: Alphacoronavirus (α-CoV), Betacoronavirus (β-CoV), Gammacoronavirus (γ-CoV) and Deltacoronavirus (δ-CoV) (4). Among seven humans CoVs identified to date including SARS-CoV-2, two of them belong to α-CoV (HCoV-NL63 and HCoV-229E) and the remaining five belong to β-CoV (HKU1, HCoV-OC43, SARS-CoV, MERS-CoV and SARS-CoV-2) (5, 6).
Coronaviruses are known for their ability to rapidly mutate, effectively cross the species barriers, and adapt to novel host environments (7). Out of the four genera, α-CoV and β-CoV are known to infect only mammals, while γ-CoV and δ-CoV primarily infect birds, with some reports indicating infections in mammals (8). Several studies have shown that bats, rodents, and avian species are natural reservoirs of diverse CoVs (9–12). For example, bats (Rhinolophus spp.) were identified as reservoirs of more than 30 CoVs including SARS-CoV (1, 13, 14). Currently, SARS-CoV-2 has been speculated to be transmitted to humans from bats through an unknown intermediate, which still needs to be conclusively proven (15). It is also possible that humans might have contracted an avirulent strain or a strain with lesser virulence directly or indirectly and then the virus might have undergone virulence-enhancing mutations resulting in human-to-human transmission (3). Whatever may be the causes for the origin of SARS-CoV-2, it remains to be proven with more supporting data.
Phylogenetic studies of SARS-CoV-2 revealed 79% nucleotide homology with SARS-CoV (16, 17) and 89% and 96% homologies with the two other SARS-like CoVs isolated from Chinese horseshoe bats Rhinolophus sinicus and Rhinolophus affinis, respectively (16, 18). However, these two SARS-like CoVs differ significantly in their receptor-binding domains. Upon comparison, SARS-like CoVs isolated from Malayan pangolins (Manis javanica) exhibited 85-92% nucleotide homology and stronger similarity in receptor binding domain with SARS-CoV-2 (19). Based on these findings, pangolins have also been considered as possible hosts in the emergence of SARS-CoV-2 (19). Besides pangolins, multiple species of wild or domestic animals like camels, mink, may also carry SARS-CoV-2 (8, 17, 20). Whether or not the suspected hosts are naturally infected by SARS-CoV-2 from humans remains to be proven.
Diseased Phenotypes Associated With SARS-CoV-2
The SARS-CoV-2 infection causes multiple disease phenotypes such as pulmonary dysfunction, hematological alterations, inflammation, electrolyte imbalance, coagulation dysfunction, liver and kidney dysfunctions, cardiac muscle injuries (21). It has been surmised that due to coagulation dysfunctions, COVID-19 patients are at increased risk of venous and arterial thromboembolism (TE) and consequential mortality. These findings highlight the need to implement thromboprophylaxis protocols, while treating COVID-19 patients (22). However, more symptoms are still being discovered, some of which appear to be rare. Due to the diverse symptoms, it has been a challenging task to diagnose and manage the disease on time (23). Moreover, the impact of comorbidities (e.g., cardiomyopathies, hypertension, diabetes, etc.) on SARS-CoV-2 infection and disease progression makes the disease management even more challenging (24). Typically, symptomatic individuals with high body temperature pose a greater risk for transmitting the virus to others. However, such symptomatic individuals can be easily identified and isolated. On the other hand, the asymptomatic individuals may continue to remain unidentified through the regular screening procedures, thus jeopardizing the efforts to minimize viral transmission through identification and isolation of the virus-carrying individuals (25). Therefore, the existing precarious situation demands the identification and deployment of reliable serological markers to enable the identification of asymptomatic individuals (26). Application of susceptible-exposed-infectious-removed (SEIR) models to understand the COVID-19 epidemiology has estimated the R0 value of 2.03 for SARS-CoV-2, highlighting the importance of factors like hygiene, social distancing, and wearing PPE in reducing human-to-human and environment-to-human transmission of the virus (27). A recent meta-analysis by Heneghan et al. (28) indicated that firm conclusions cannot be drawn in the absence of recoverable viral culture samples of SARS-CoV-2. On the contrary, Greenhalgh et al. discuss reasons with supporting evidence to argue in the support of airborne transmission of SARS-CoV-2 (29). However, this is beyond the scope of this review as we limit the dissensions.
Over the past year, several studies have identified the presence of SARS-CoV-2 RNA in anal/rectal swabs and stool specimens of COVID-19 patients, even after the clearance of the virus in the upper respiratory tract (14, 30–34). With SARS-CoV-2 angiotensin-converting enzyme 2 receptor (ACE2) reported to be highly expressed in gastrointestinal epithelial cells (35, 36), it is suggested that the virus can actively infect and replicate in the gastrointestinal (GI) tract and thus has critical implications to the disease management, transmission, and infection control. In the past, symptoms such as diarrhea, nausea, vomiting and abdominal pain have been observed in patients infected with other coronaviruses such as SARS and MERS (37–39), indicating that the presence of the virus in the GI tract may be a common feature of the CoV infections. These studies collectively highlight the need for screening multiple clinical specimens from a single patient other than nasopharyngeal swabs, such as lung and tracheal aspirate, blood, pleural fluid, and fecal samples (40). Although different clinical specimens are of particular interest, whether or not this is done during the course of treatment of patients to ensure full recovery and no viral shedding is a subject of debate (41). Some evidence shows that SARS-CoV-2 can be vertically transmitted to the fetus or neonates from the infected mothers (42).
SARS-CoV-2 Genome and Its Variants
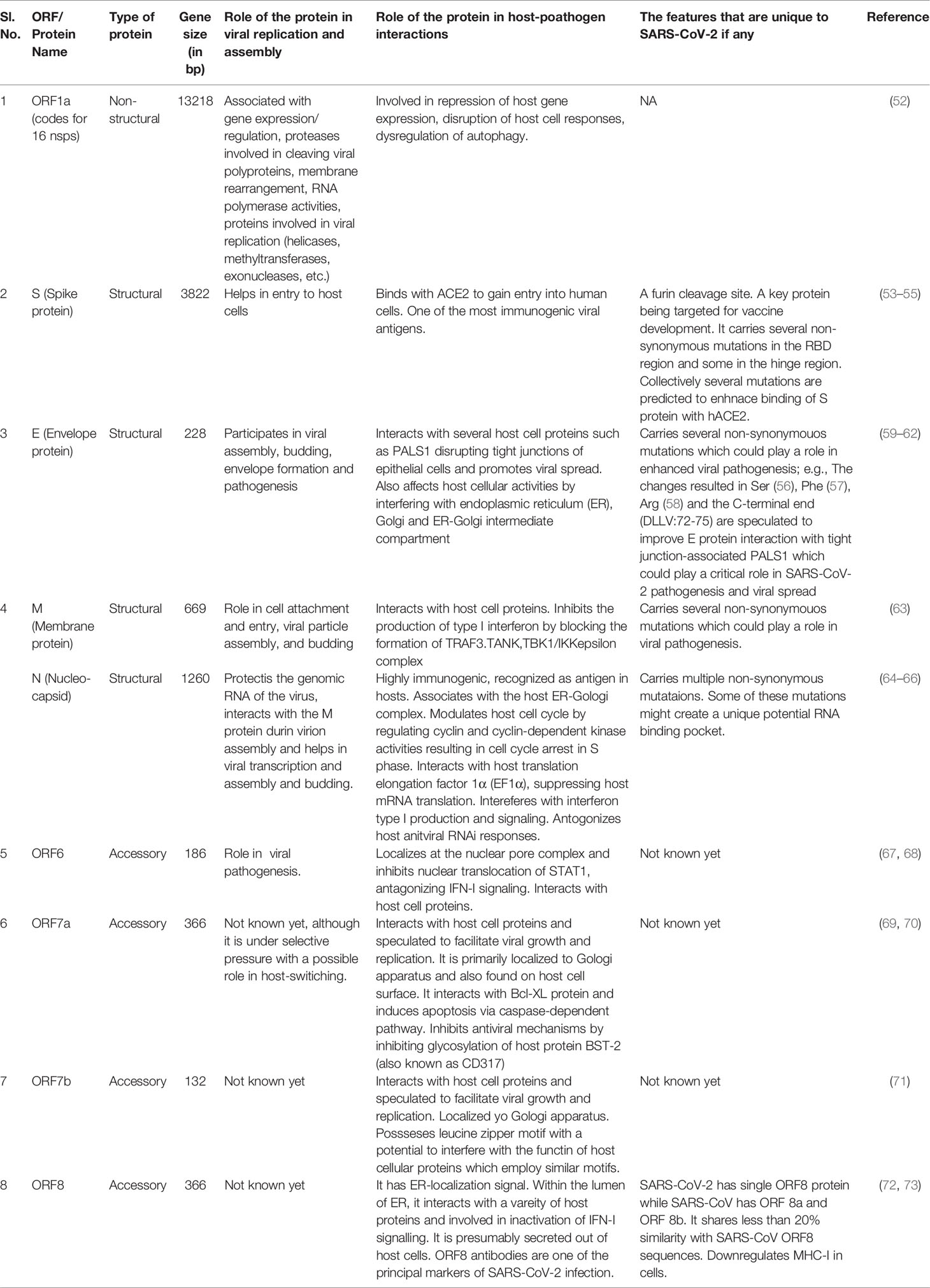
The SARS-CoV-2 carries the largest genome size of ~ 29.7 kb, which shares similarities with the genomes of other β-CoVs that have caused many epidemics in the past (43–50) (Figure 1). The SARS-CoV-2 consists of at least 16 non-structural and 4 structural proteins (51). The list of the genes encoded by SARS-CoV-2 genome and their known/predicted functions in the virus and the host is given in Table 1. After entering the host cell, the positive (sense) strand of viral RNA undergoes translation to synthesize non-structural proteins (nsps) from two protein-coding genes ORF1a and ORF1b (Figure 1). The CoV genome consists of six open reading frames (ORFs) of which the first ORF (ORF1a) encompasses about 2/3rd of the genome and produces polypeptide 1a, which is further cleaved into 11 nsps. Due to ribosomal frameshift occurring upstream of ORF1a stop codon, translation of ORF1b yields another polypeptide called ORF1ab, which is further cleaved into 16 nsps. Cleavage of ORF1a and ORF1ab polypeptides is mediated by the virus-encoded proteases nsp3 and nsp5, which harbor papain-like domain and 3C-like domain, respectively. In addition, diverse forms of CoVs encode structural and accessory proteinS for example orf3 a/b protein, translated from the sub-genomic part of CoVs (52). Apart from the accessory proteins that have been found to play an important role in the viral-host interactions, several viral structural and accessory proteins, and a number of host proteins have been shown to interact with the viral RNA (74). Examples of host proteins that interact with the viral RNA include poly-A-binding protein, mitochondrial aconitase, pyrimidine-binding protein, and nuclear ribonucleoprotein which were known to be shown for SARS-CoV (75) and it is anticipated that that a similar interaction could be seen in SARS-CoV-2. However, this study is in infancy and beyond the scope of this review.
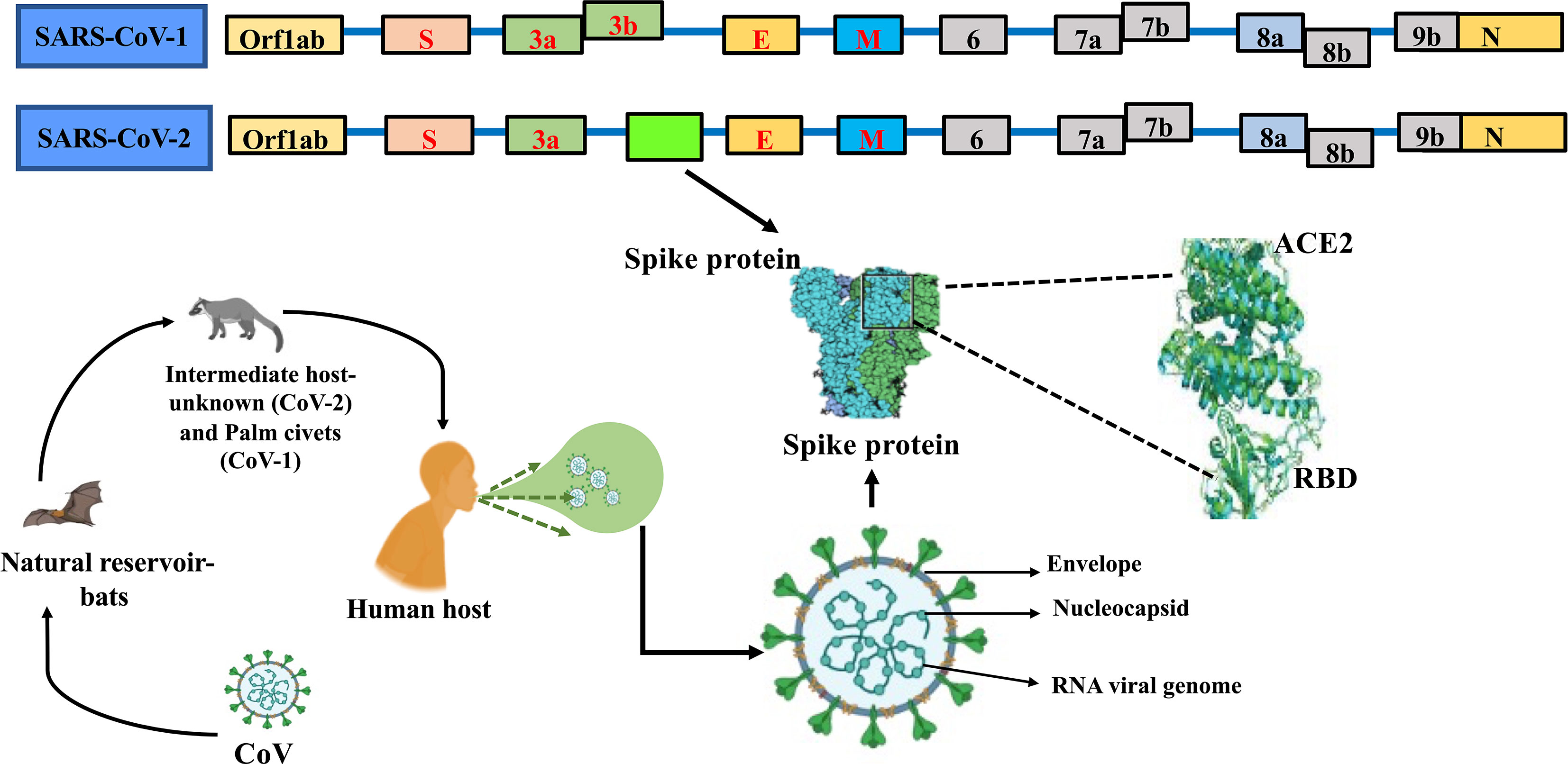
Figure 1 Genome comparison of SARS-CoV and SARS-CoV-2 along with ORFs depicting the common ancestry.
Genome studies revealed that while SARS-CoV-2, SARS-CoV, and MERS-CoV share many similarities, they have multiple differences in their genomic and phenotypic structure which influence their pathogenesis (76). The RNA polymerases of most of the RNA viruses either lack or have a poor proof-reading activity (77). As a consequence, single-stranded RNA viruses mutate at a faster rate than DNA viruses, and as a result, genetic variants emerged as quasispecies (78). For instance, it was estimated that SARS-CoV can mutate its genome at the rate of 0.80-2.38 × 10-3 nucleotide substitutions per site per year which is similar to other RNA viruses (79). It is therefore anticipated that mutations at such rates allow the viruses to evolve faster, enabling them to escape host immune surveillance, develop resistance to drugs, vaccines and switch hosts. Moreover, the rapid rate of mutations could also increase the frequency of false negatives during nucleic acid-based viral diagnostics. Gustine and Jones have highlighted the role of a dysregulated innate immune response associated with hyperinflammatory syndrome in severe COVID-19 patients (80).
Several researchers have identified mutations in the SARS-CoV-2 genome with mutations identified in the SARS-CoV-2 genome that cause variants to become hotspots. Oster et al. have reviewed the trends in impending mutational hotspots ever since the covid infection rate has drastically been accounted for in various patients (81). Pachetti et al. identified eight novel recurrent mutations of SARS-CoV-2 in addition to 5 previously identified hotspots. Of the novel mutation hotspots, one was in the RdRp (RNA dependent RNA polymerase) gene involved in proof-reading machinery. This mutation was associated with a higher number of point mutations in Europe than viral genomes from Asia. The authors speculated that observed RdRp mutation could result in an enhanced viral replication, influencing mortality rates (56). analyzed 220 genome sequences from the GISAID database (www.gisaid.org) derived from patients infected by SARS-CoV-2 worldwide from December 2019 to mid-March 2020. The SARS-CoV-2 genome sequence available from the GenBank database was used as the reference in their study. They identified eight novel recurrent mutations of SARS-CoV-2, in addition to previously identified 5 hotspots. Of these 13 hotspots, 5 were predominantly present in the European isolates, and 3 were found exclusively in the North American isolates. Among the 13 hotspots, 6 were not observed in Asian isolates. It is assumed that the non-synonymous mutation (proline to leucine) in the RdRp gene could be affecting its proof-reading ability presumably by disrupting its interaction with the other protein cofactors such as the Exon domain of nsp14, nsp7, or nsp8, further alter the mutation rate of the virus (56). Further, they speculate that the observed RdRp mutation could result in an enhanced viral replication, influencing mortality rates. Several polymerase inhibitors are currently being tested in clinical studies targeting the RdRp protein of the virus. Some of these drugs have a predicted binding site in the SARS-CoV-2 RdRp hydrophobic cleft, which is adjacent to the identified mutation at the 14408 positions. Henceforth, it is possible that the observed mutation might also impart drug resistance to the virus, however, the proposed hypothesis needs to be experimentally proven.
As algebraic topology-based machine learning models were introduced to evaluate the SARS-CoV-2 spike glycoprotein (S protein) and host ACE2 receptor binding free changes, a number of mutations were identified (44). From the cluster analysis and the transmission dynamics, it is assumed that future SARS-CoV-2 mutations tend to have higher chances to mutate into significantly more infectious COVID-19 strains than the original one from Wuhan. Given the fact that the “infectivity-strengthening” mutations spread faster than “infectivity-weakening” mutations, the study concluded that the proportion of asymptomatic cases have drastically increased despite impending cases in South Korea (82). In another study, phylogenetic analysis of ~1,400 SARS-CoV-2 genomes isolated from India yielded 7 clusters, of which one was unique to India. This unique cluster (Clade I/A3i) included three variants; C6312A, C13730T, and C28311T, which resulted in amino acid changes T2016K (orf1a), A97V (RdRp) or A88V (orf1b) and P13L (N protein), respectively (83). These changes are hypothesized to enhance the virulence and/or infectivity of the virus. The mutation A97V in the RdRp protein is located in its NiRAN domain, suggested to be relevant in RNA binding and nucleotidylation activity (84). Mutations have also been identified in other viral proteins with potential functional consequences (57, 85, 86). However, the impact of all these mutations on the functions of the respective proteins and their consequences on viral pathogenesis needs to be experimentally tested. Among the recently identified variants of SARS-CoV-2, the one carrying D614G mutation in the spike protein of the virus has emerged as the most prevalent variant in the global pandemic, possibly due to its greater fitness advantage (87). This variant was found to be more infectious resulting in a higher viral titer in patients. In addition to viral variants, a recent genome-wide association study (GWAS) in the UK identified eight human genetic variants consisting of a combination of alleles at multiple loci that are predicted to increase the risk of mortality among COVID-19 patients (88). Lately, some of the SARS-CoV-2 variants with mutations in the S protein are behind the surge in the second wave of SARS-CoV-2 infections in many countries including India. Among the three sub lineages of B.1.617, namely B.1.617.1, B.1.617.2, and B.1.617.3, the variant named B.1.617.1.2 has been found to rapidly spread in many countries including India. Recently, B.1.617.1.2 was classified by WHO as a ‘variant of concern’ based on evidence showing higher transmission rates and reduced neutralization by antibodies obtained from the convalescent serum of infected or vaccinated individuals (89). Similarly, many other variants are being identified globally and being monitored for their epidemiology.
Variants of Interest/Concern
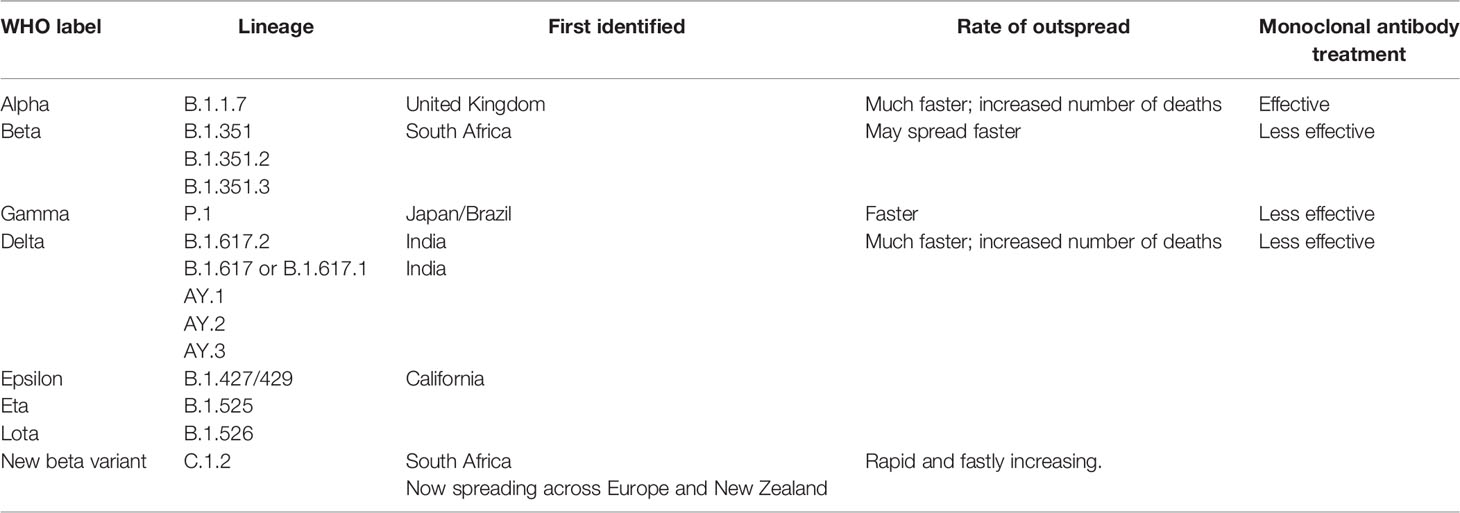
Several variants have been identified globally, which appear to spread more quickly, potentially increasing COVID-19 infections. Based on the factors, including the severity of the virus and their ability to spread, four variants viz. B.1.1.7, B.1.351, P.1 and B.1.617.2, also known as alpha, beta, gamma and delta, respectively, have been characterized as the variants of concern (Table 2). An alpha variant, identified initially in the United Kingdom and delta variant, first identified in India, is outspread with a much faster rate than beta and gamma variants, first determined in South Africa and Japan/Brazil, respectively. Hence, alpha and delta variants may potentially cause more sickness and increase the number of deaths globally. On the other hand, treatment with monoclonal antibodies is less effective against beta, gamma, and delta variants, while treatment for alpha variants is known to be effective (101). These variants have different alterations in the spike protein leading to increased susceptibility, virulence and transmission that may affect viral replication and host immune response. More recently, a novel South African variant C.1.2 was identified which would escape antibody response. BNT162b2 (Pfizer/BioNTech) and mRNA-1273 (Moderna) use a formulated vaccine which is used to elicit potential response against the aforementioned SARS-CoV-2 variants thereby combating rapid diagnosis (102).
Host-Pathogen Interactions, Co-Morbidities and Immune Response
The infection starts with the binding of virus elements to the host cell surface receptors, followed by viral entry and multiplication. Moreover, the binding of virus to the host receptor is primarily required for viral transmission to other host species as well (103). Most of the human CoVs (HCoVs) have been ascertained to recognize protein peptidases as their receptors except HCoV-OC43 and HKU1 which utilize sugar molecules for cellular attachment (104). For example, the HCoV-229E binds to human aminopeptidase N (105), while MERS-CoV binds to human dipeptidyl peptidase 4 (58). On the other hand, SARS-CoV and HCoV-NL63 interact with ACE2 for viral entry into the host (106). In CoVs, the passage of viral entry into the host is mediated by the spike (S) glycoprotein, situated at the viral surface 71. The S protein is cleaved by human proteases into S1 and S2 subunits, which are involved in receptor identification and cell membrane arrangement, respectively (53). The N-terminal and the C-terminal domains (NTD and CTD) of the S1 subunit play an important role in the functioning of receptor-binding domains (RBD) in SARS-CoVs and MERS-CoVs (58, 107). The crystal structures of HCoV-NL63 and SARS-CoV RBD interactions with human ACE2 (hACE2) receptors are already known (107, 108). The composite crystal structure of interaction between S1 CTD of SARS-CoV-2 and hACE2 receptor has revealed that most of the binding residues of SARS-CoV-2 in hACE2 show similarity with SARS-CoV binding sites (90). Furthermore, the crystal structures of trimeric S protein of SARS-CoV-2 published recently with buried and exposed RBD regions have also been found to be consistent with structural characteristics of S proteins in MERS-CoV and SARS-CoV (91). While receptor recognition allows us to identify hCOV pathogenic determinants, the ACE-2 Receptor recognition is an important determinant of hCoVs infection and pathogenesis. Comparable to SARS-CoV RBD, it was assumed that SARS-CoV-2 RBD is less potent and exposed given the spike proteins’ switching between lying down and standing up positions (109). The former buried positions were in turn larger and so the masking regions are favored by immune evasion of spike proteins’ through ACE-2. In summary, SARS-CoV-2 has diffident buried and exposed surfaces affecting the chemistry of pathogenic determinants besides host protease activation.
Furthermore, the newly evolved delta mutations in SARS-CoV-2 may impact disease severity and become immunocompromised. As it is known that the variant replication is manifold when compared to its predecessor variants, whether or not vaccinated people have the highest chance to skip the infection and transmission is under evaluation (110). This could be due to neutralizing antibodies that might have evaded the infection. These delta variants host multiple mutations in the S1 subunit, including the RBD that seem to have concurrent epitope tags. Therefore, the knowledge of the crystal structure of interactions between host and viral proteins will not only allow repurposing of the existing drugs but also the discovery of new antiviral agents. As a prelude to this, we have developed a bona fide network of HCoV-host interactome networks which could be used to identify key putative candidate interacting pairs responsible for the viral pathogenesis (Figure 2). Among HCoVs, different viral proteins are being used for viral entry. For example, β-CoVs use their S protein for cell binding and entry. SARS‐CoV binds to ACE2, MERS‐CoV utilizes dipeptidyl peptidase 4 (DPP4) of the host. Recent modelling of the structure of SARS‐CoV‐2 S protein predicts that it can bind to both human DPP4 and hACE2 (112).
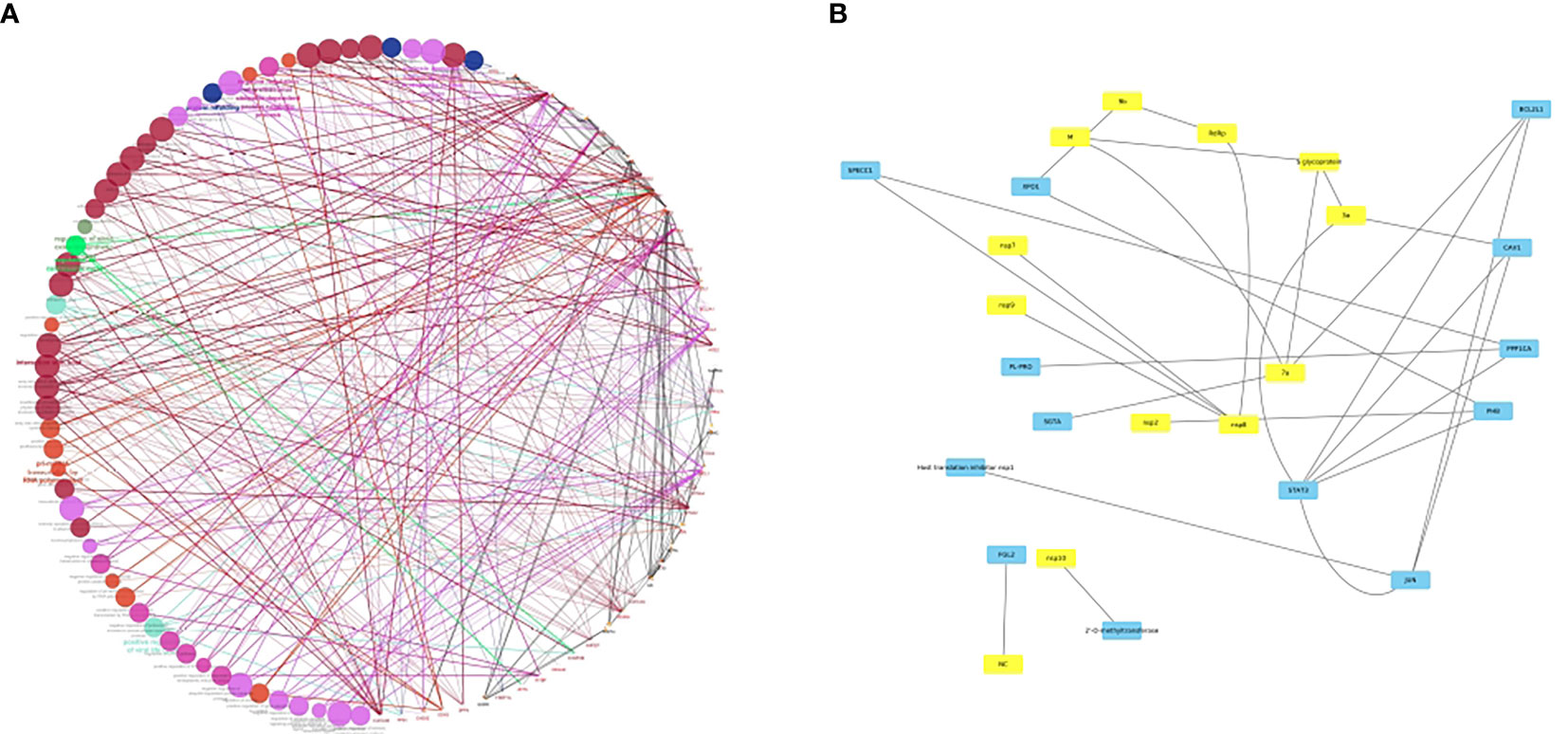
Figure 2 (A) bona fide network analysis of HCoV-host interactome network. A subnetwork illustrating the HCoV-host interactome where colored nodes represent diverse enriched pathways in host proteins and edge colors represent evidence collected from protein-protein interactions. The Figure was generated using Cytoscape (111) with the host proteins shown to be interacting with the viral conglomerate proteins. The pink edges indicate the experimentally validated interactions, with green, grey, orange, and maroon edges representing fusion, subcellular location, coexpression and associations from text mining. (B) A sub-figure with all the viral proteins in yellow colored nodes and the host proteins in blue colored nodes. This figure is depicted keeping in view of the interplay of various host-pathogen interactions. A Supplementary Table 1 is added which shows the list of interactants based on the text mining, co-expression network and overall scores. However, as no co-expression network studies have been deciphered yet between virus and the host, the network is distributed largely based on the text mining interactions. In conclusion, the network largely explores how the SARS-CoV-2 proteins have an interplay with a lot of host proteins, viz. SPECC1, a mitochondrial like protein, PHB, a prohibitin associated with cancer and FGL2/fibroleukin prothrombinase, a protein associated with clotting factors and alveolar macrophage activation.
The diabetic patients appear to be highly vulnerable to SARS-CoV-2 infection. Given the administration of angiotensin-directed medications for diabetics, a new treatment regimen is being suggested for COVID-19 patients with diabetes (113, 114). Lately, diabetic patients have shown an alarming increase in a post-covid complication called mucormycosis, which is a highly invasive opportunistic fungal infection affecting several vital organs of the body including, the brain, eyes, ear, nose, throat and mouth (115). Mucormycosis is caused by a group of fungi called Mucormycetes (also popularly known as black fungus) and the infection has become one of the factors involved in increasing COVID-19-associated morbidity and mortality (116). The use of steroids to reduce inflammation in COVID-19 patients leads to a further increase in blood sugar levels and reduction in immunity, further increasing the susceptibility to opportunistic black fungus infections (115). In addition to diabetes, other factors which further enhance sensitivity to infection are smoking, age, and obesity (117). Similarly, hospital-acquired nosocomial infections pose additional challenges in treating COVID-19 patients, underscoring the need for early diagnosis of these opportunistic pathogens. Pulmonary dysfunction, asthma, and bronchitis due to air pollution have also been reported to increase the chances of SARS-CoV-2-associated morbidity and mortality (118). In this context, mathematical modeling and artificial intelligence can serve as valuable tools in predicting the impact of various environmental variables and comorbid conditions on the progression of SARS-CoV-2 infection and vice versa (119). Given the multifaceted involvement of ACE-2 in SARS-CoV-2 biology and its role in comorbid conditions, there is a need to understand the association of ACE-2 expression with patient fatality rate.
Over the last year, multiple studies reported an association between the ABO blood group and COVID-19 (120–124). All these studies concur that individuals with blood group A exhibit a higher risk of SARS-CoV-2 infection, morbidity and mortality. On the other hand, individuals with blood group O exhibit a lower risk for the same parameters. In vitro studies by Guillon et al. (54) demonstrated inhibition of adhesion of SARS-CoV-expressing cells to ACE2-expressing cells by anti-A antibodies, which might explain why individuals with blood group A showed higher susceptibility to COVID-19 infection. However, these findings need to be further validated before blood groups can be used as prognostic markers.
Lower levels of certain vitamins such as vitamin D, A, and K also appear to influence SARS-CoV-2 infection and disease progression. Given the multifarious roles of these vitamins in the maintenance of mucosal membrane barriers, innate and adaptive immunity, maintenance of blood pressure and blood glucose levels, integrity and functioning of skeletal and non-skeletal tissues among other functions, their deficiency is expected to be a potential risk for SARS-CoV-2 infection and disease severity (125–131).
Does SARS-CoV-2 Hijack Mitochondria to Induce Infections?
Mitochondria have been known to be involved in inducing innate immune responses primarily against viral attacks (132). Upon viral infection, damage to mitochondrial membrane results in leakage of the mitochondrial DNA into the cytosol. In addition to the viral genome, the presence of mitochondrial genome in the cytosol is sensed by cytosolic surveillance systems that recognize these cytosolic DNA called Danger Associated Molecular Patterns (DAMPS) (133). Recognition of DNA by cytosolic DNA receptors triggers immune response and inflammation leading to the recruitment of macrophages and dendritic cells. Hence, any alteration with the mitochondrial membrane during infection may lead to accelerated immune response and explain the observed clinical symptoms during COVID-19. The ACE2 receptor and TMPRSS2, a transmembrane serine protease, both used by hCoVs to enter host cells, can alter mitochondrial function, which might enable the virus to hijack mitochondria to its advantage in facilitating its spread to the neighboring cells. We have earlier shown that mutations in the SARS-CoV-2 may manipulate mitochondria even as ACE2 is regulated by its inherent mitochondrial function (132). Moreover, the 5’ and 3’ UTR regions of the viral transcript have been found to have mitochondria localization signals (51). Several ORFs like ORF9b, ORF3b, ORF7a, and ORF8b of the SARS-CoV are also shown to be localized in the host mitochondria. These sequences have high similarity with those found in SARS-CoV-2, indicating that it is highly likely that SARS-CoV-2 transcripts may possess the ability to localize in host mitochondria too. Several SARS-CoV proteins have been known to interact with host mitochondrial proteins and yet, the knowledge remains limited regarding the cellular significance of these interactions. CoVs have been known to reside in ER-derived double membrane vesicles to avert host immune responses and therefore CoVs might have been adopted to reside in mitochondria-derived vesicles for similar purposes. These observations suggest that mitochondrial hijacking may be an essential mechanism in SARS-CoV-2 infection and therefore, drugs that selectively restore mitochondrial function and promote its biogenesis may prove to be effective anti-inflammatory agents in the treatment of COVID-19. Infection of host cells by SARS-CoV-2 induces a strong immune response which could be linked to the ability of the virus to use the host organelles such as endoplasmic reticulum (ER) for its replication. These observations further emphasize the need to understand the crosstalk between the viral proteins and the proteins of mitochondria and ER and its influence on the formation of mitochondria-derived double-vesicles (MDV) and mitochondrial antiviral-signaling proteins (MAVS), which induce apoptosis. Studies are underway to know whether or not the non-structural proteins (nsps) are an integral part of this mechanism (51). While it is also not clear how the RNA from the virus enters the mitochondria in human cells, it is hypothesized that it may interact with ACE-2 receptors in regulating mitochondrial function. Higher levels of ACE-2 have been shown to restore impaired mitochondrial function (134). As SARS-CoV-2 infection leads to mitochondrial hijacking, it negatively impacts cellular bioenergetics leading to asphyxiation, further causing fatalities. It will be interesting to know how the mutations in mitochondrial genomes in humans might contribute to diverse responses to SARS-CoV-2 infections, akin to the GWAS study conducted in the UK which identified a set of alleles associated with increased risk of mortality among COVID-19 patients (88). An epilog to this, we have also recently hypothesized how SARS-CoV-2 transgressing non-coding RNAs esp lncRNAs of the host may allow us to understand the known unknown regions of the viral genome (110).
Disease Management of COVID-19: Potential Treatment Options and Repurposing the Existing Drugs for COVID-19 Treatment
Since the onset of COVID-19, several attempts have been made to identify or develop new antiviral agents or to repurpose existing drugs to antagonize the virus. Among the antiparasitic agents, hydroxychloroquine (HCQ), chloroquine, and ivermectin were extensively tested to repurpose them against SARS-CoV-2. While some studies showed promising results, others were inconclusive (135). Several antiviral compounds that are known to act against viruses including CoVs have been explored to repurpose them against SARS-CoV-2. Among them, some of them with the most promising outcomes include, Remdesivir (GS-5734), Favipiravir, Lopinavir, and Ritonavir. Remdesivir (GS-5734), an analog of adenosine, is a broad-spectrum antiviral agent that gets incorporated into nascent RNA, resulting in premature termination of RNA synthesis (136–138). Favipiravir is another FDA-approved drug for the influenza virus that showed antiviral activity against SARS-CoV-2 via the inhibition of viral RNA polymerase (139). The HIV protease inhibitors Lopinavir and Ritonavir have been under clinical focus for their known roles in inhibiting 3C-like proteases [3CLpro;a cysteine protease that hydrolyses the Viral proteins (hCoVs)] of CoVs with which SARS-CoV-2 shares up to 96% sequence identity (140). However, recent clinical trials failed to show any significant benefits of these anti-HIV drugs against SARS-Co V-2 (141).
Additionally, some of the herbal medicines are being used or evaluated to treat COVID-19 based on findings from clinical trials or in vitro studies. For example, the traditional Chinese medicine made up of Qingfei Paidu Decoction (QFPD) accelerated the recovery from COVID-19 symptoms and reduced the mortality rates (142–144). Earlier, QFPD had also been shown to be effective against SARS-CoV (145). Therefore, QFPD is currently being used as one of the adjunct treatments against COVID-19 in China. Similarly, tryptanthrin, a compound isolated from the leaf of the Chinese herb, Strobilanthes cusia, has been explored as another treatment option since it displayed an antiviral activity against human coronavirus NL63 in a cell-type independent manner in vitro (146). In addition, several active phytoconstituents obtained from medicinal plant species such as Curcuma longa (turmeric), Withania somnifera (Ashwagandha), Tinospora cordifolia (Giloy), and Ocimum sanctum (Tulsi) have been subjected to molecular docking studies to identify compounds with a potential to interact with and inhibit SARS-CoV-2 proteins (147–149). Some of these compounds have shown promising results and are being pursued further.
Various computational tools have been employed to rapidly identify potential new drugs and the existing drugs for treatment of COVID-19 (150). Artificial intelligence (AI) has been deployed for predicting the structure of SARS-CoV-2 proteins which will help in identifying new drugs besides repurposing the existing drugs to treat the virus. For example, Beck et al. (151) developed a deep learning-based pretrained drug-interaction model called molecule transformer-drug target interaction (MT-DTI) to shortlist commercially available drugs for their potential to target SARS-CoV-2 viral proteins. Their findings predicted that Atazanavir, an anti- HIV drug to possess the highest inhibitory potency. Their study also predicted other drugs including Remdesivir, Efavirenz, Ritonavir, and Dolutegravir to have anti-SARS-CoV-2.
Molecular docking studies have been extensively used to identify molecules that can bind to the SARS-CoV-2 machinery and inhibit its replication (152). Docking studies of nigelledine and hederin from Nigella sativa with SARS-CoV-2 main protease called 3CLpro (Mpro), showed improved docking score for ligand binding free energies compared to Chloroquine, HCQ and Favipiravir. Therefore, these drugs have been provisionally considered as potential therapeutic agents for treating SARS-CoV-2 (153). In another study, a chemographic analysis of anti-CoV structure-activity information from a public database (ChEMBL) containing a vast pool of 800 million organic compounds, about 380 potential anti-CoV agents were identified, which needs to be experimentally tested and validated for their antiviral activity against SARS-CoV-2 (154).
Elfiky (155) built a model for the viral RdRp to test its binding affinity to some of the clinically approved drugs and drug candidates by carrying out molecular modeling, docking, and molecular dynamics simulations for the viral protein RNA-dependent RNA polymerase (RdRp). In addition to Sofosbuvir, Ribavirin, Galidesivir, Remdesivir, Favipiravir, Cefuroxime, Tenofovir, and Hydroxychloroquine showing effectiveness, new derivatives were shown to have promising results for the attachment to the SARS-CoV-2 RdRp. Recently, the crystal structure of SARS-CoV-2 3CLpro (Liu et al., 10.2210/pdb6LU7/pdb) was used to screen several approved drugs or drugs in clinical trials via virtual docking (156). The findings from this study predicted several promising drugs to inhibit the 3CLpro. The list includes Carfilzomib (a proteasome inhibitor being used as an anticancer drug), Eravacycline (a florocycline- a synthetic analog of tetracycline), Valrubicin (an analog of doxorubicin which is an inhibitor of nucleic acid metabolism, being used to treat bladder cancer), Lopinavir (HIV-1 protease inhibitor), Elbasvir (anti Hepatitis C viral drug, which targets nsp 5A), and Streptomycin (a known antibiotic that inhibits bacterial protein synthesis). In a similar docking study, along with HIV protease inhibitors (Lopinavir, Asunaprevir, Indivavir, and Ritonavir), new molecules including Methisazone (an antiviral drug that inhibits mRNA and protein synthesis in poxviruses), CGP42112A (an angiotensin AT2 receptor agonist), and ABT450 (Paritaprevir: an antiviral drug that inhibits nsp 3-4A serine protease of HCV) were predicted to bind and inhibit 3CLpro of SARS-CoV-2 (157).
Furthermore, synergistic studies carried out on Lopinavir, Oseltamivir, and Ritonavir showed that when used together these drugs exhibited greater binding affinity and inhibition of the viral protease proteins than when used alone (158). In another docking study, ~ 1.3 billion compounds from the ZINC15 database were docked against the active site of the SARS-CoV-2 3CLpro using the Deep Docking (DD) platform, which provides a fast prediction of docking scores (159). The authors identified the top 1000 potential ligands for SARS CoV-2 3CLpro. In a similar study, the potential drugs from the ZINC15 database were screened for their binding affinity for S protein and 3CLproo of SARS-CoV-2. The findings from this study identified Zanamivir (an anti-influenza drug), Indinavir and Saquinavir (anti-HIV drugs), Remdesivir (anti-SARS-CoV and an anti-ebola virus drug) as compounds with high binding affinities for 3CLpro. In addition, flavin adenine dinucleotide (FAD) adeflavin, coenzyme A, tiludronate, and Dpnh (NADH) were predicted to bind the S protein with high affinity (160). Recently, 2 deoxy glucose (2DG), an anticancer drug, has been approved for emergency use in India as an adjunct therapy to treat COVID-19. 2DG inhibits host glycolytic pathway, anti-inflammatory action and interacts with viral proteins (161, 162). An exhaustive list of repurposed drugs targeting various stages of virus life cycle and their current status of clinical trials has been listed in Almasi and Mohammadipanah (163).
Machine Learning Heuristics and Artificial Intelligence for COVID-19 Management
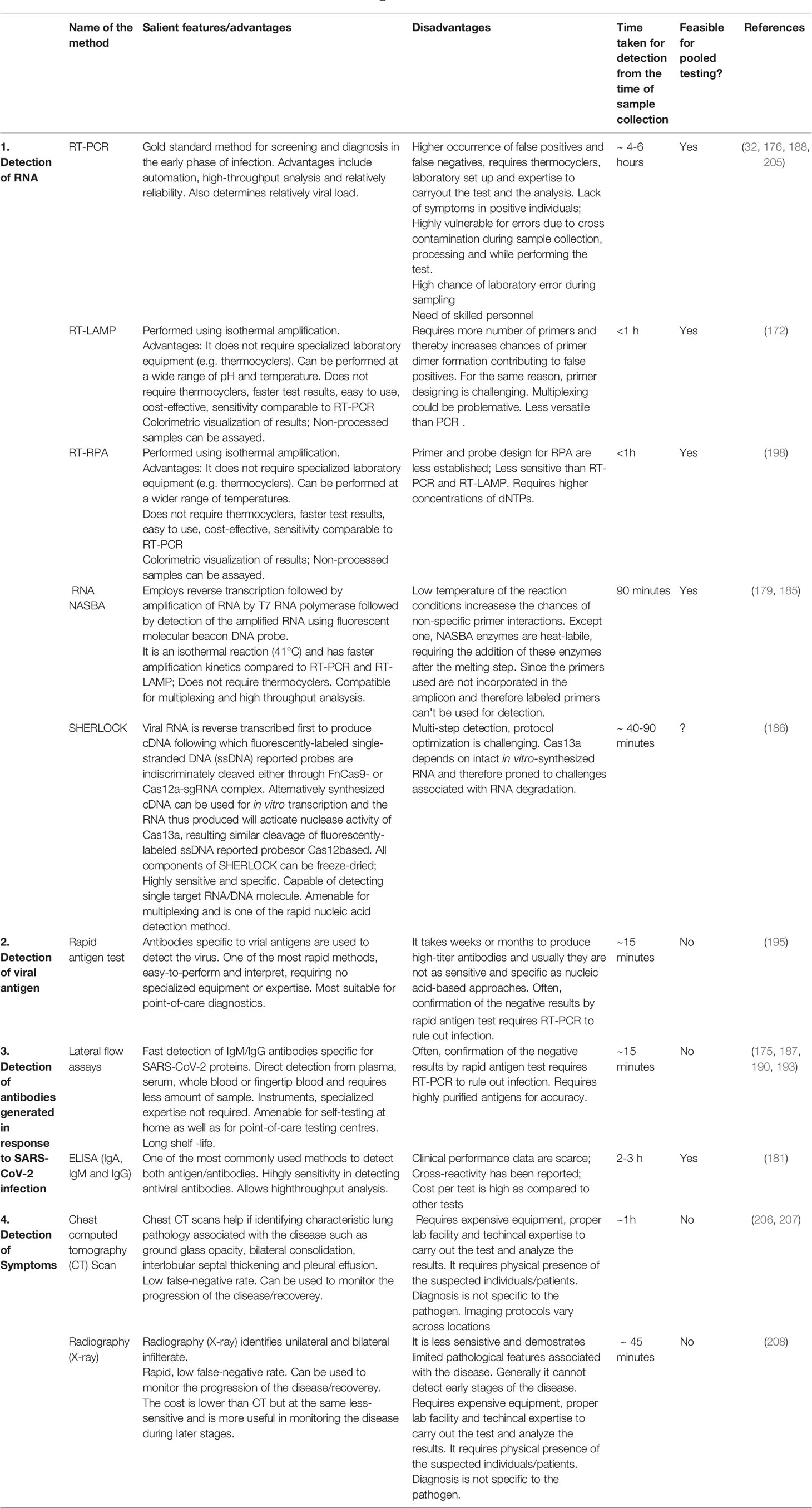
The artificial intelligence (AI) and machine learning (ML) paradigms have offered effective tools and algorithms to combat COVID-19 pandemic. AI is being successfully applied for disease cluster identification, monitoring of COVID-19 patients, in determining mortality risk, disease diagnosis and management, contact tracing through geotagging, resource allocation, facilitating training of health care personnel, data management, and in predicting future pandemic outbreaks and the disease trend (164–166). The power of AI-ML in viral diagnostics and the management of COVID-19 have been summarized in various reviews (167–170) (Table 3). Clinically, computed tomography (CT), positron emission tomography-CT (PET/CT), lung ultrasound, and magnetic resonance imaging (MRI) are being used in COVID-19 diagnosis. The AI can supplement medical imaging-based COVID-19 diagnosis particularly reducing the diagnosis time (167, 206). Application of deep learning to X-ray and CT scan imaging has resulted in detection of COVID-19 with high accuracy, sensitivity, and specificity (207, 208). Such applications also effectively differentiated symptoms due to COVID-19 from bacterial pneumonia. In another novel study, AI was applied to predict the outcome of the RT-PCR-based diagnosis of COVID-19 on the basis of 16 simple parameters derived from complete blood profile (205).
Randhawa et al. (209) identified an intrinsic SARS-CoV-2 virus genomic signature in combination with a machine learning-based alignment-free approach for a fast, scalable, and highly accurate classification of whole SARS-CoV-2 genomes. They coupled supervised machine learning with digital signal processing (MLDSP) for genome analyses. Moreover, the authors claimed highly accurate real-time taxonomic classification of SARS-CoV-2 genomes simply based on raw DNA sequence, without any requirement of specialized biological knowledge or training or gene/genome annotations. Their method is extremely rapid; for example, analysis of a data set consisting of 5538 unique viral genomes with a total of ~ 62 Mb was carried out within a few minutes with high classification accuracy. Other applications of ML and AI in protein structure prediction, identification of potential drugs using molecular docking studies, reverse vaccinology, automating the data collection and transmission using cyber physical systems (CPS) from the current and futuristic diagnostic devices (e.g., lab-on-chip) are described in previous sections.
Conclusions
COVID-19 undoubtedly has turned out to be one of the most destructive pandemics in recent history having a negative impact on all aspects of human life. The repetitive global resurgence of apparently more infectious strains of SARS-CoV-2 has made the pandemic unrelenting. We have attempted to summarize the various critical findings pertaining to COVID-19 biology from detection to treatment. Given the fatic paradigm, there is a need for the scientific community to apply Thoughts-Action-Debate (TAD) applying Trikarna shuddhi, a Sanskrit adage of learning as we say “ome” to prepare for future pandemics.
Author Contributions
PSu and PBK conceptualized the idea for the review. The authors with + sign contributed equally, performed the literature search, analyzed cited references, and wrote the original article. PSu, ORB, PBK, AKS, HKP, KKS, GC critically revised the article. All authors contributed to the article and approved the submitted version.
Conflict of Interest
RP was employed by Genomix CARL Pvt. Ltd.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Acknowledgments
PK is thankful to VFSTR for providing Emeritus fellowship. All authors are life members of Bioclues.org, India’s largest bioinformatics professional society.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2021.724914/full#supplementary-material
References
1. Shereen MA, Khan S, Kazmi A, Bashir N, Siddique R. COVID-19 Infection: Origin, Transmission, and Characteristics of Human Coronaviruses. J Adv Res (2020) 24:91–8. doi: 10.1016/j.jare.2020.03.005
2. Xie M, Chen Q. Insight Into 2019 Novel Coronavirus - An Updated Interim Review and Lessons From SARS-CoV and MERS-CoV. Int J Infect Dis (2020) 94:119–24. doi: 10.1016/j.ijid.2020.03.071
3. Zhang G, Li B, Yoo D, Qin T, Zhang X, Jia Y, et al. Animal Coronaviruses and SARS-CoV-2. Transbound Emerg Dis (2021) 68(3):1097–110. doi: 10.1111/tbed.13791
4. Gabutti G, d’Anchera E, Sandri F, Savio M, Stefanati A. Coronavirus: Update Related to the Current Outbreak of COVID-19. Infect Dis Ther (2020) 9(2):241–53. doi: 10.1007/s40121-020-00295-5
5. Ciotti M, Ciccozzi M, Terrinoni A, Jiang WC, Wang CB, Bernardini S. The COVID-19 Pandemic. Crit Rev Clin Lab Sci (2020) 57(6):365–88. doi: 10.1080/10408363.2020.1783198
6. Helmy YA, Fawzy M, Elaswad A, Sobieh A, Kenney SP, Shehata AA. The COVID-19 Pandemic: A Comprehensive Review of Taxonomy, Genetics, Epidemiology, Diagnosis, Treatment, and Control. J Clin Med (2020) 9(4):1225. doi: 10.3390/jcm9041225
7. Dhama K, Patel SK, Sharun K, Pathak M, Tiwari R, Yatoo MI, et al. SARS-CoV-2 Jumping the Species Barrier: Zoonotic Lessons From SARS, MERS and Recent Advances to Combat This Pandemic Virus. Travel Med Infect Dis (2020) 37:101830. doi: 10.1016/j.tmaid.2020.101830
8. Cui J, Li F, Shi ZL. Origin and Evolution of Pathogenic Coronaviruses. Nat Rev Microbiol (2019) 17(3):181–92. doi: 10.1038/s41579-018-0118-9
9. Chu DKW, Leung CYH, Gilbert M, Joyner PH, Ng EM, Tse TM, et al. Avian Coronavirus in Wild Aquatic Birds. J Virol (2011) 85(23):12815–20. doi: 10.1128/JVI.05838-11
10. Rabi FA, Al Zoubi MS, Kasasbeh GA, Salameh DM, Al-Nasser AD. SARS-CoV-2 and Coronavirus Disease 2019: What We Know So Far. Pathogens (2020) 9(3):231. doi: 10.3390/pathogens9030231
11. Vijaykrishna D, Smith GJD, Zhang JX, Peiris JSM, Chen H, Guan Y. Evolutionary Insights Into the Ecology of Coronaviruses. J Virol (2007) 81(8):4012–20. doi: 10.1128/JVI.02605-06
12. Woo PCY, Lau SKP, Lam CSF, Lau CCY, Tsang AKL, Lau JHN, et al. Discovery of Seven Novel Mammalian and Avian Coronaviruses in the Genus Deltacoronavirus Supports Bat Coronaviruses as the Gene Source of Alphacoronavirus and Betacoronavirus and Avian Coronaviruses as the Gene Source of Gammacoronavirus and Deltacoronavirus. J Virol (2012) 86(7):3995–4008. doi: 10.1128/JVI.06540-11
13. Song Z, Xu Y, Bao L, Zhang L, Yu P, Qu Y, et al. From SARS to MERS, Thrusting Coronaviruses Into the Spotlight. Viruses (2019) 11(1):59. doi: 10.3390/v11010059
14. Xu X-W, Wu X-X, Jiang X-G, Xu K-J, Ying L-J, Ma C-L, et al. Clinical Findings in a Group of Patients Infected With the 2019 Novel Coronavirus (SARS-Cov-2) Outside of Wuhan, China: Retrospective Case Series. BMJ (2020) 368:m606. doi: 10.1136/bmj.m606
15. Turcios-Casco MA, Cazzolla Gatti R. Do Not Blame Bats and Pangolins! Global Consequences for Wildlife Conservation After the SARS-CoV-2 Pandemic. Biodivers Conserv (2020) 29(13):3829–33. doi: 10.1007/s10531-020-02053-y
16. Zhou P, Yang X-L, Wang X-G, Hu B, Zhang L, Zhang W, et al. A Pneumonia Outbreak Associated With a New Coronavirus of Probable Bat Origin. Nature (2020) 579(7798):270–3. doi: 10.1038/s41586-020-2012-7
17. Wu F, Zhao S, Yu B, Chen Y-M, Wang W, Song Z-G, et al. A New Coronavirus Associated With Human Respiratory Disease in China. Nature (2020) 579(7798):265–9. doi: 10.1038/s41586-020-2008-3
18. Ren L-L, Wang Y-M, Wu Z-Q, Xiang Z-C, Guo L, Xu T, et al. Identification of a Novel Coronavirus Causing Severe Pneumonia in Human: A Descriptive Study. Chin Med J (Engl) (2020) 133(9):1015–24. doi: 10.1097/CM9.0000000000000722
19. Lam TT-Y, Jia N, Zhang Y-W, Shum MH-H, Jiang J-F, Zhu H-C, et al. Identifying SARS-CoV-2-Related Coronaviruses in Malayan Pangolins. Nature (2020) 583(7815):282–5. doi: 10.1038/s41586-020-2169-0
20. Zhou P, Shi Z-L. SARS-CoV-2 Spillover Events. Science (2021) 371(6525):120–2. doi: 10.1126/science.abf6097
21. Ciaccio M, Agnello L. Biochemical Biomarkers Alterations in Coronavirus Disease 2019 (COVID-19). Diagnosis (Berl) (2020) 7(4):365–72. doi: 10.1515/dx-2020-0057
22. Malas MB, Naazie IN, Elsayed N, Mathlouthi A, Marmor R, Clary B. Thromboembolism Risk of COVID-19 Is High and Associated With a Higher Risk of Mortality: A Systematic Review and Meta-Analysis. EClinicalMedicine (2020) 29-30:100639. doi: 10.1016/j.eclinm.2020.100639
23. Sanyaolu A, Okorie C, Marinkovic A, Patidar R, Younis K, Desai P, et al. Comorbidity and Its Impact on Patients With COVID-19. SN Compr Clin Med (2020) 2(8):1069–76. doi: 10.1007/s42399-020-00363-4
24. Bordallo B, Bellas M, Cortez AF, Vieira M, Pinheiro M. Severe COVID-19: What Have We Learned With the Immunopathogenesis? Adv Rheumatol (2020) 60(1):50. doi: 10.1186/s42358-020-00151-7
25. Sun P, Lu X, Xu C, Sun W, Pan B. Understanding of COVID-19 Based on Current Evidence. J Med Virol (2020) 92(6):548–51. doi: 10.1002/jmv.25722
26. Lipsitch M, Swerdlow DL, Finelli L. Defining the Epidemiology of Covid-19 — Studies Needed. N Engl J Med (2020) 382(13):1194–6. doi: 10.1056/NEJMp2002125
27. Mwalili S, Kimathi M, Ojiambo V, Gathungu D, Mbogo R. SEIR Model for COVID-19 Dynamics Incorporating the Environment and Social Distancing. BMC Res Notes (2020) 13:352. doi: 10.1186/s13104-020-05192-1
28. Heneghan CJ, Spencer EA, Brassey J, Plüddemann A, Onakpoya IJ, Evans DH, et al. SARS-CoV-2 and the Role of Airborne Transmission: A Systematic Review. F1000Res (2021) 10:232. doi: 10.12688/f1000research.52091.1
29. Greenhalgh T, Jimenez JL, Prather KA, Tufekci Z, Fisman D, Schooley R. Ten Scientific Reasons in Support of Airborne Transmission of SARS-CoV-2. Lancet (2021) 397(10285):1603–5. doi: 10.1016/S0140-6736(21)00869-2
30. Crespo J, Iglesias-García J, Hinojosa Del Val JE, García García F, Gil de Miguel Á, Fernández Carrillo C, et al. COVID-19 and the Digestive System: Protection and Management During the SARS-CoV-2 Pandemic. Rev Esp Enferm Dig (2020) 112:389–96. doi: 10.17235/reed.2020.7128/2020
31. Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, et al. First Case of 2019 Novel Coronavirus in the United States. N Engl J Med (2020) 382(10):929–36. doi: 10.1056/NEJMoa2001191
32. Tang A, Tong ZD, Wang HL, Dai Y-X, Li K-F, Liu J-N, et al. Detection of Novel Coronavirus by RT-PCR in Stool Specimen From Asymptomatic Child, China. Emerg Infect Dis (2020) 26(6):1337–9. doi: 10.3201/eid2606.200301
33. Wong SH, Lui RN, Sung JJ. Covid-19 and the Digestive System. J Gastroenterol Hepatol (2020) 35(5):744–8. doi: 10.1111/jgh.15047
34. Young BE, Ong SWX, Kalimuddin S, Low JG, Tan SY, Loh J, et al. Epidemiologic Features and Clinical Course of Patients Infected With SARS-CoV-2 in Singapore. JAMA (2020) 323(15):1488–94. doi: 10.1001/jama.2020.3204
35. Harmer D, Gilbert M, Borman R, Clark KL. Quantitative mRNA Expression Profiling of ACE 2, a Novel Homologue of Angiotensin Converting Enzyme. FEBS Lett (2002) 532(1-2):107–10. doi: 10.1016/s0014-5793(02)03640-2
36. Fei X, Tang M, Zheng X, Liu Y, Li X, Shan H. Evidence for Gastrointestinal Infection of SARS-CoV-2. Gastroenterology (2020) 158(6):1831–3.e3. doi: 10.1053/j.gastro.2020.02.055
37. Chan JF, Lau SK, To KK, Cheng VC, Woo PC, Yuen KY. Middle East Respiratory Syndrome Coronavirus: Another Zoonotic Betacoronavirus Causing SARS-Like Disease. Clin Microbiol Rev (2015) 28(2):465–522. doi: 10.1128/CMR.00102-14
38. Lee N, Hui D, Wu A, Chan P, Cameron P, Joynt GM, et al. A Major Outbreak of Severe Acute Respiratory Syndrome in Hong Kong. N Engl J Med (2003) 348(20):1986–94. doi: 10.1056/NEJMoa030685
39. Leung CW, Kwan YW, Ko PW, Chiu SS, Loung P, Fong N, et al. Severe Acute Respiratory Syndrome Among Children. Pediatrics (2004) 113(6):e535–43. doi: 10.1542/peds.113.6.e535
40. Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, et al. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA (2020) 323(18):1843–4. doi: 10.1001/jama.2020.3786
41. He X, Lau EHY, Wu P, Deng X, Wang J, Hao X, et al. Temporal Dynamics in Viral Shedding and Transmissibility of COVID-19. Nat Med (2020) 26(5):672–5. doi: 10.1038/s41591-020-0869-5
42. Caparros-Gonzalez RA. Consecuencias Maternas Y Neonatales De La Infección Por Coronavirus COVID-19 Durante El Embarazo: Una Scoping Review [Maternal and Neonatal Consequences of Coronavirus COVID-19 Infection During Pregnancy: A Scoping Review]. Rev Esp Salud Publica (2020) 94:e202004033. doi: 10.4321/s1135-57272020000100025
43. Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The Proximal Origin of SARS-CoV-2. Nat Med (2020) 26(4):450–2. doi: 10.1038/s41591-020-0820-9
44. Chen J, Wang R, Wang M, Wei GW. Mutations Strengthened SARS-CoV-2 Infectivity. J Mol Biol (2020) 432(19):5212–26. doi: 10.1016/j.jmb.2020.07.009
45. Fung TS, Liu DX. Human Coronavirus: Host-Pathogen Interaction. Annu Rev Microbiol (2019) 73(1):529–57. doi: 10.1146/annurev-micro-020518-115759
46. Khailany RA, Safdar M, Ozaslan M. Genomic Characterization of a Novel SARS-CoV-2. Gene Rep (2020) 19:100682. doi: 10.1016/j.genrep.2020.100682
47. Gorbalenya AE, Enjuanes L, Ziebuhr J, Snijder EJ. Nidovirales: Evolving the Largest RNA Virus Genome. Virus Res (2006) 117(1):17–37. doi: 10.1016/j.virusres.2006.01.017
48. Peiris JSM, Lai ST, Poon LLM, Guan Y, Yam LYC, Lim W, et al. Coronavirus as a Possible Cause of Severe Acute Respiratory Syndrome. Lancet (2003) 361(9366):1319–25. doi: 10.1016/s0140-6736(03)13077-2
49. Woo PCY, Huang Y, Lau SKP, Yuen K-Y. Coronavirus Genomics and Bioinformatics Analysis. Viruses (2010) 2(8):1804–20. doi: 10.3390/v2081803
50. Wang C, Liu Z, Chen Z, Huang X, Xu M, He T, et al. The Establishment of Reference Sequence for SARS-CoV-2 and Variation Analysis. J Med Virol (2020) 92(6):667–74. doi: 10.1002/jmv.25762
51. Wu KE, Fazal FM, Parker KR, Zou J, Chang HY. RNA-GPS Predicts SARS-CoV-2 RNA Residency to Host Mitochondria and Nucleolus. Cell Syst (2020) 11(1):102–8. doi: 10.1016/j.cels.2020.06.008
52. Hussain S, Ja P, Chen Y, Yang Y, Xu J, Peng Y, et al. Identification of Novel Subgenomic RNAs and Noncanonical Transcription Initiation Signals of Severe Acute Respiratory Syndrome Coronavirus. J Virol (2005) 79(9):5288–95. doi: 10.1128/JVI.79.9.5288-5295.2005
53. Du L, Kao RY, Zhou Y, He Y, Zhao G, Wong C, et al. Cleavage of Spike Protein of SARS Coronavirus by Protease Factor Xa Is Associated With Viral Infectivity. Biochem Biophys Res Commun (2007) 359(1):174–9. doi: 10.1016/j.bbrc.2007.05.092
54. Guillon P, Clément M, Sébille V, Rivain JG, Chou CF, Ruvoen-Clouet N, et al. Inhibition of the Interaction Between the SARS-CoV Spike Protein and Its Cellular Receptor by Anti-Histo-Blood Group Antibodies. Glycobiology (2008) 18(12):1085–93. doi: 10.1093/glycob/cwn093
55. Xie Y, Karki CB, Du D, Li H, Wang J, Sobitan A, et al. Spike Proteins of SARS-CoV and SARS-CoV-2 Utilize Different Mechanisms to Bind With Human ACE2. Front Mol Biosci (2020) 7:591873. doi: 10.3389/fmolb.2020.591873
56. Pachetti M, Marini B, Benedetti F, Giudici F, Mauro E, Storici P, et al. Emerging SARS-CoV-2 Mutation Hot Spots Include a Novel RNA-Dependent-RNA Polymerase Varian. J Transl Med (2020) 18(1):179. doi: 10.1186/s12967-020-02344-6
57. Phan LT, Nguyen TV, Luong QC, Nguyen TV, Nguyen HT, Le HQ, et al. Importation and Human-to-Human Transmission of a Novel Coronavirus in Vietnam. N Engl J Med (2020) 382(9):872–4. doi: 10.1056/NEJMc2001272
58. Lu G, Hu Y, Wang Q, Qi J, Gao F, Li Y, et al. Molecular Basis of Binding Between Novel Human Coronavirus MERS-CoV and its Receptor CD26. Nature (2013) 500(7461):227–31. doi: 10.1038/nature12328
59. Li Y, Surya W, Claudine S, Torres J. Structure of a Conserved Golgi Complex-Targeting Signal in Coronavirus Envelope Proteins. J Biol Chem (2014) 289(18):12535–49. doi: 10.1074/jbc.M114.560094
60. Schoeman D, Fielding BC. Coronavirus Envelope Protein: Current Knowledge. Virol J (2019) 16(1):69. doi: 10.1186/s12985-019-1182-0
61. Maio FD, Lo Cascio EL, Babini G, Sali M, Longa SD, Tilocca B, et al. Improved Binding of SARS-CoV-2 Envelope Protein to Tight Junction-Associated PALS1 Could Play a Key Role in COVID-19 Pathogenesis. Microbes Infect (2020) 22(10):592–7. doi: 10.1016/j.micinf.2020.08.006
62. Rahman MS, Hoque MN, Islam MR, Islam I, Mishu ID, Rahman MM, et al. Mutational Insights Into the Envelope Protein of SARS-CoV-2. Gene Rep (2021) 22:100997. doi: 10.1016/j.genrep.2020.100997
63. Bianchi M, Benvenuto D, Giovanetti M, Angeletti S, Ciccozzi M, Pascarella S. Sars-CoV-2 Envelope and Membrane Proteins: Structural Differences Linked to Virus Characteristics? BioMed Res Int (2020) 2020:4389089. doi: 10.1155/2020/4389089
64. Dutta NK, Mazumdar K, Gordy JT. The Nucleocapsid Protein of SARS–CoV-2: A Target for Vaccine Development. J Virol (2020) 94(13)::e00647–20. doi: 10.1128/jvi.00647-20
65. Mu J, Xu J, Zhang L, Shu T, Wu D, Huang M, et al. SARS-CoV-2-Encoded Nucleocapsid Protein Acts as a Viral Suppressor of RNA Interference in Cells. Sci China Life Sci (2020) 63(9):1413–6. doi: 10.1007/s11427-020-1692-1
66. Kang S, Yang M, Hong Z, Zhang L, Huang Z, Chen X, et al. Crystal Structure of SARS-CoV-2 Nucleocapsid Protein RNA Binding Domain Reveals Potential Unique Drug Targeting Sites. Acta Pharm Sin B (2020) 10(7):1228–38. doi: 10.1016/j.apsb.2020.04.009
67. Miorin L, Kehrer T, Sanchez-Aparicio MT, Zhang K, Cohen P, Patel RS, et al. SARS-CoV-2 Orf6 Hijacks Nup98 to Block STAT Nuclear Import and Antagonize Interferon Signaling. Proc Natl Acad Sci USA (2020) 117(45):28344–54. doi: 10.1073/pnas.2016650117
68. Lee JG, Huang W, Lee H, van de Leemput J, Kane MA, Han Z. Characterization of SARS-CoV-2 Proteins Reveals Orf6 Pathogenicity, Subcellular Localization, Host Interactions and Attenuation by Selinexor. Cell Biosci (2021) 11:58. doi: 10.1186/s13578-021-00568-7
69. Nelson CA, Pekosz A, Lee CA, Diamond MS, Fremont DH. Structure and Intracellular Targeting of the SARS-Coronavirus Orf7a Accessory Protein. Structure (2005) 13(1):75–85. doi: 10.1016/j.str.2004.10.010
70. Taylor JK, Coleman CM, Postel S, Sisk JM, Bernbaum JG, Venkataraman T, et al. Severe Acute Respiratory Syndrome Coronavirus ORF7a Inhibits Bone Marrow Stromal Antigen 2 Virion Tethering Through a Novel Mechanism of Glycosylation Interference. J Virol (2015) 89(23):11820–33. doi: 10.1128/JVI.02274-15
71. Fogeron ML, Montserret R, Zehnder J, Nguyen MH, Dujardin M, Brigandat L, et al. SARS-CoV-2 ORF7b: Is a Bat Virus Protein Homologue a Major Cause of COVID-19 Symptoms? bioRxiv (2021) 428650. doi: 10.1101/2021.02.05.428650
72. Flower TG, Buffalo CZ, Hooy RM, Allaire M, Ren X, Hurley JH. Structure of SARS-CoV-2 ORF8, a Rapidly Evolving Immune Evasion Protein. Proc Natl Acad Sci USA (2021) 118(2):e2021785118. doi: 10.1073/pnas.2021785118
73. Zhang Y, Zhang J, Chen Y, Luo B, Yuan Y, Huang F, et al. The ORF8 Protein of SARS-CoV-2 Mediates Immune Evasion Through Potently Downregulating MHC-I. bioRxiv (2020). doi: 10.1101/2020.05.24.111823
74. Liu DX, Fung TS, Chong KK-L, Shukla A, Hilgenfeld R. Accessory Proteins of SARS-CoV and Other Coronaviruses. Antiviral Res (2014) 109:97–109. doi: 10.1016/j.antiviral.2014.06.013
75. Shi ST, Lai MM. Viral and Cellular Proteins Involved in Coronavirus Replication. Curr Top Microbiol Immunol (2005) 287:95–131. doi: 10.1007/3-540-26765-4_4
76. Mousavizadeh L, Ghasemi S. Genotype and Phenotype of COVID-19: Their Roles in Pathogenesis. J Microbiol Immunol Infect (2021) 54(2):159–63. doi: 10.1016/j.jmii.2020.03.022
77. Ziebuhr J. The Coronavirus Replicase. Curr Top Microbiol Immunol (2005) 287:57–94. doi: 10.1007/3-540-26765-4_3
78. Sanjuán R, Domingo-Calap P. Mechanisms of Viral Mutation. Cell Mol Life Sci (2016) 73(23):4433–48. doi: 10.1007/s00018-016-2299-6
79. Zhao Z, Li H, Wu X, Zhong Y, Zhang K, Zhang YP, et al. Moderate Mutation Rate in the SARS Coronavirus Genome and its Implications. BMC Evol Biol (2004) 4:21. doi: 10.1186/1471-2148-4-21
80. Gustine JN, Jones D. Immunopathology of Hyperinflammation in COVID-19. Am J Pathol (2021) 191(1):4–17. doi: 10.1016/j.ajpath.2020.08.009
81. Oster AM, Kang GJ, Cha AE, Beresovsky V, Rose CE, Rainisch G, et al. Trends in Number and Distribution of COVID-19 Hotspot Counties - United States, March 8-July 15, 2020. MMWR Morb Mortal Wkly Rep (2020) 69(33):1127–32. doi: 10.15585/mmwr.mm6933e2
82. Ryu S, Ali ST, Noh E, Kim D, Lau EH, Cowling BJ. Transmission Dynamics and Control of Two Epidemic Waves of SARS-CoV-2 in South Korea. BMC Infect Dis (2021) 21(1):1–9. doi: 10.1186/s12879-021-06204-6
83. Banu S, Jolly B, Mukherjee P, Singh P, Khan S, Zaveri L, et al. A Distinct Phylogenetic Cluster of Indian SARS-CoV-2 Isolates. BioRxiv (2020) 2020:5. doi: 10.1101/2020.05.31.126136
84. Gao Y, Yan L, Huang Y, Liu F, Zhao Y, Cao L, et al. Structure of the RNA-Dependent RNA Polymerase From COVID-19 Virus. Science (2020) 368(6492):779–82. doi: 10.1126/science.abb7498
85. Yin C. Genotyping Coronavirus SARS-CoV-2: Methods and Implications. Genomics (2020) 112(5):3588–96. doi: 10.1016/j.ygeno.2020.04.016
86. Yu F, Du L, Ojcius DM, Pan C, Jiang S. Measures for Diagnosing and Treating Infections by a Novel Coronavirus Responsible for a Pneumonia Outbreak Originating in Wuhan, China. Microbes Infect (2020) 22(2):74–9. doi: 10.1016/j.micinf.2020.01.003
87. Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, et al. Tracking Changes in SARS-CoV-2 Spike: Evidence That D614G Increases Infectivity of the COVID-19 Virus. Cell (2020) 182:812–27. doi: 10.1016/j.cell.2020.06.043
88. Hu J, Li C, Wang S, Li T, Zhang H. Genetic Variants are Identified to Increase Risk of COVID-19 Related Mortality From UK Biobank Data. medRxiv (2020) 15(1):10. doi: 10.1101/2020.11.05.2026761
89. Vaidyanathan G. Coronavirus Variants Are Spreading in India -What Scientists Know So Far. Nature (2021) 593(7859):321–2. doi: 10.1038/d41586-021-01274-7
90. Wang Q, Zhang Y, Wu L, Niu S, Song C, Zhang Z, et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human Ace2. Cell (2020) 181(4):894–904. doi: 10.1016/j.cell.2020.03.045
91. Walls AC, Park Y-J, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell (2020) 181(2):281–292.e6. doi: 10.1016/j.cell.2020.02.058
92. Wu J, Yuan X, Wang B, Gu R, Li W, Xiang X, et al. Severe Acute Respiratory Syndrome Coronavirus 2: From Gene Structure to Pathogenic Mechanisms and Potential Therapy. Front Microbiol (2020) 11:1576. doi: 10.3389/fmicb.2020.01576
93. Yoshimoto FK. The Proteins of Severe Acute Respiratory Syndrome Coronavirus-2 (SARS CoV-2 or N-COV19), the Cause of COVID-19. Protein J (2020) 39(3):198–216. doi: 10.1007/s10930-020-09901-4
94. Zhao J, Falcón A, Zhou H, Netland J, Enjuanes L, Breña PP, et al. Severe Acute Respiratory Syndrome Coronavirus Protein 6 is Required for Optimal Replication. J Virol (2009) 83(5):2368–73. doi: 10.1016/j.virol.2015.08.010
95. Randhawa PK, Scanlon K, Rappaport J, Gupta MK. Modulation of Autophagy by SARS-CoV-2: A Potential Threat for Cardiovascular System. Front Physiol (2020) 11:611275. doi: 10.3389/fphys.2020.611275
96. Siu KL, Kok KH, Ng MHJ, Poon VKM, Yuen KY, Zheng BJ, et al. Severe Acute Respiratory Syndrome Coronavirus M Protein Inhibits Type I Interferon Production by Impeding the Formation of TRAF3.TANK.TBK1/IKK Epsilon Complex. J Biol Chem (2009) 284(24):16202–9. doi: 10.1074/jbc.M109.008227
97. Mu J, Fang Y, Yang Q, Shu T, Wang A, Huang M, et al. SARS-CoV-2 N Protein Antagonizes Type I Interferon Signaling by Suppressing Phosphorylation and Nuclear Translocation of STAT1 and STAT2. Cell Discovery (2020) 6:65. doi: 10.1038/s41421-020-00208-3
98. Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, et al. A SARS-CoV-2 Protein Interaction Map Reveals Targets for Drug Repurposing. Nature (2020) 583(7816):459–68. doi: 10.1038/s41586-020-2286-9
99. Tang X, Li G, Vasilakis N, Zhang Y, Shi Z, Zhong Y, et al. Differential Stepwise Evolution of SARS Coronavirus Functional Proteins in Different Host Species. BMC Evol Biol (2009) 9:52. doi: 10.1186/1471-2148-9-52
100. Tan YX, Tan THP, Lee MJR, Tham PY, Gunalan V, Druce J, et al. Induction of Apoptosis by the Severe Acute Respiratory Syndrome Coronavirus 7a Protein Is Dependent on its Interaction With the Bcl-XL Protein. J Virol (2007) 81(12):6346–55. doi: 10.1128/JVI.00090-07
101. Center for Disease Control . Available at: https://www.cdc.gov/coronavirus/2019-ncov/variants/variant.html (Accessed August 24,2021).
102. Cyranoski D. Alarming COVID Variants Show Vital Role of Genomic Surveillance. Nature (2021) 589(7842):337–8. doi: 10.1038/d41586-021-00065-4
104. Liu DX, Liang JQ, Fung TS. Human Coronavirus-229e, -OC43,-NL63, and -HKU1 (Coronaviridae). Encyclopedia Virol (2021) 2021:428–40. doi: 10.1016/b978-0-12-809633-8.21501-x
105. Li Z, Tomlinson AC, Wong AHM, Zhou D, Desforges M, Talbot PJ, et al. The Human Coronavirus HCoV-229e S-Protein Structure and Receptor Binding. eLife (2019) 8. doi: 10.7554/elife.51230
106. Hofmann H, Pyrc K, van der Hoek L, Geier M, Berkhout B, Pöhlmann S. Human Coronavirus NL63 Employs the Severe Acute Respiratory Syndrome Coronavirus Receptor for Cellular Entry. Proc Natl Acad Sci USA (2005) 102(22):7988–93. doi: 10.1073/pnas.0409465102
107. Li F. Receptor Recognition Mechanisms of Coronaviruses: A Decade of Structural Studies. J Virol (2015) 89(4):1954–64. doi: 10.1128/jvi.02615-14
108. Wu K, Li W, Peng G, Li F. Crystal Structure of NL63 Respiratory Coronavirus Receptor-Binding Domain Complexed With Its Human Receptor. Proc Natl Acad Sci USA (2009) 106(47):19970–4. doi: 10.2210/pdb3kbh/pdb
109. Rossi GA, Sacco O, Mancino E, Cristiani L, Midulla F. Differences and Similarities Between SARS-CoV and SARS-CoV-2: Spike Receptor-Binding Domain Recognition and Host Cell Infection With Support of Cellular Serine Proteases. Infection (2020) 48(5):665–9. doi: 10.1007/s15010-020-01486-5
110. Shukla N, Prasad A, Kanga U, Suravajhala R, Nigam VK, Kishor PK, et al. SARS-CoV-2 Transgressing LncRNAs Uncovers the Known Unknowns. Physiol Genomics (2021). doi: 10.1152/physiolgenomics.00075.2021
111. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res (2003) 13(11):2498–504. doi: 10.1101/gr.1239303
112. Bassendine MF, Bridge SH, McCaughan GW, Gorrell MD. COVID-19 and Comorbidities: A Role for Dipeptidyl Peptidase 4 (DPP4) in Disease Severity? J Diabetes (2020) 12(9):649–58. doi: 10.1111/1753-0407.13052
113. Jeong I-K, Yoon KH, Lee MK. Diabetes and COVID-19: Global and Regional Perspectives. Diabetes Res Clin Pract (2020) 166:108303. doi: 10.1016/j.diabres.2020.108303
114. Yu B, Li C, Sun Y, Wang DW. Insulin Treatment Is Associated With Increased Mortality in Patients With COVID-19 and Type 2 Diabetes. Cell Metab (2021) 33(1):65–77.e2. doi: 10.1016/j.cmet.2020.11.014
115. Song G, Liang G, Liu W. Fungal Co-Infections Associated With Global COVID-19 Pandemic: A Clinical and Diagnostic Perspective From China. Mycopathologia (2020) 185(4):599–606. doi: 10.1007/s11046-020-00462-9
116. Sharma S, Grover M, Bhargava S, Samdani S, Kataria T. Post Coronavirus Disease Mucormycosis: A Deadly Addition to the Pandemic Spectrum. J Laryngol Otol (2021) 135(5):442–7. doi: 10.1017/S0022215121000992
117. Tadic M, Cuspidi C, Sala C. COVID-19 and Diabetes: Is There Enough Evidence? J Clin Hypertens (Greenwich) (2020) 22(6):943–8. doi: 10.1111/jch.13912
118. Sharma AK, Balyan P. Air Pollution and COVID-19: Is the Connect Worth Its Weight? Indian J Public Health (2020) 64(6):132. doi: 10.4103/ijph.ijph_466_20
119. Mohamadou Y, Halidou A, Kapen PT. A Review of Mathematical Modeling, Artificial Intelligence and Datasets Used in the Study, Prediction and Management of COVID-19. Appl Intell (2020) 50(11):3913–25. doi: 10.1007/s10489-020-01770-9
120. Li J, Wang X, Chen J, Cai Y, Deng A, Yang M. Association Between ABO Blood Groups and Risk of SARS-CoV-2 Pneumonia. Br J Haematol (2020) 190(1):24–7. doi: 10.1111/bjh.16797
121. Wu B-B, Gu D-Z, Yu J-N, Yang J, Shen W-Q. Association Between ABO Blood Groups and COVID-19 Infection, Severity and Demise: A Systematic Review and Meta-Analysis. Infect Genet Evol (2020) 84:104485. doi: 10.1016/j.meegid.2020.104485
122. Zietz M, Zucker J, Tatonetti NP. Associations Between Blood Type and COVID-19 Infection, Intubation, and Death. Nat Commun (2020) 11(1):5761. doi: 10.1038/s41467-020-19623-x
123. Muñiz-Diaz E, Llopis J, Parra R, Roig I, Ferrer G, Grifols J, et al. Relationship Between the ABO Blood Group and COVID-19 Susceptibility, Severity and Mortality in Two Cohorts of Patients. Blood Transfus (2021) 19(1):54–63. doi: 10.2450/2020.0256-20
124. Zhao J, Yang Y, Huang H, Li D, Gu D, Lu X, et al. Relationship Between the ABO Blood Group and the COVID-19 Susceptibility. Clin Infect Dis (2021) 73(2):328–31. doi: 10.1093/cid/ciaa1150
125. Biesalski HK, Nohr D. New Aspects in Vitamin A Metabolism: The Role of Retinyl Esters as Systemic and Local Sources for Retinol in Mucous Epithelia. J Nutr (2004) 134(12 Suppl):3453S–7S. doi: 10.1093/jn/134.12.3453S
126. Dofferhoff AS, Piscaer I, Schurgers LJ, Visser MPJ, van den Ouweland JMW, de Jong PA, et al. Reduced Vitamin K Status as a Potentially Modifiable Risk Factor of Severe Coronavirus Disease 2019. Clin Infect Dis (2020), ciaa1258. doi: 10.1093/cid/ciaa1258
127. Engelsen O. The Relationship Between Ultraviolet Radiation Exposure and Vitamin D Status. Nutrients (2010) 2(5):482–95. doi: 10.3390/nu2050482
128. MacLaughlin J, Holick MF. Aging Decreases the Capacity of Human Skin to Produce Vitamin D3. J Clin Invest (1985) 76(4):1536–8. doi: 10.1172/JCI112134
129. Martineau AR, Forouhi NG. Vitamin D for COVID-19: A Case to Answer? Lancet Diabetes Endocrinol (2020) 8(9):735–6. doi: 10.1016/S2213-8587(20)30268-0
130. Prietl B, Treiber G, Pieber TR, Amrein K. Vitamin D and Immune Function. Nutrients (2013) 5(7):2502–21. doi: 10.3390/nu5072502
131. Zhang D, Peng C, Zhao H, Xia Y, Zhang D, Dong H, et al. Induction of Thymic Stromal Lymphopoietin Expression in 16-HBE Human Bronchial Epithelial Cells by 25-Hydroxyvitamin D3 and 1, 25-Dihydroxyvitamin D3. Int J Mol Med (2013) 32(1):203–10. doi: 10.3892/ijmm.2013.1353
132. Singh KK, Chaubey G, Chen JY, Suravajhala P. Decoding SARS-CoV-2 Hijacking of Host Mitochondria in COVID-19 Pathogenesis. Am J Physiol Cell Physiol (2020) 319(2):C258–67. doi: 10.1152/ajpcell.00224.2020
133. Roh JS, Sohn DH. Damage-Associated Molecular Patterns in Inflammatory Diseases. Immune Netw (2018) 18(4):e27. doi: 10.4110/in.2018.18.e27
134. Shi TT, Yang FY, Liu C, Cao X, Lu J, Zhang XL, et al. Angiotensin-Converting Enzyme 2 Regulates Mitochondrial Function in Pancreatic β-Cells. Biochem Biophys Res Commun (2018) 495(1):860–6. doi: 10.1016/j.bbrc.2017.11.055
135. Rakedzon S, Khoury Y, Rozenberg G, Neuberger A. Hydroxychloroquine and Coronavirus Disease 2019: A Systematic Review of a Scientific Failure. Rambam Maimonides Med J (2020) 11(3):e0025. doi: 10.5041/RMMJ.10416
136. Martinez MA. Compounds With Therapeutic Potential Against Novel Respiratory 2019 Coronavirus. Antimicrob Agents Chemother (2020) 64(5):e00399–20. doi: 10.1128/AAC.00399-20
137. Sheahan TP, Sims AC, Graham RL, Menachery VD, Gralinski LE, Case JB, et al. Broad-Spectrum Antiviral GS-5734 Inhibits Both Epidemic and Zoonotic Coronaviruses. Sci Transl Med (2017) 9(396):eaal3653. doi: 10.1126/scitranslmed.aal3653
138. Siegel D, Hui HC, Doerffler E, Clarke MO, Chun K, Zhang L, et al. Discovery and Synthesis of a Phosphoramidate Prodrug of a Pyrrolo [2, 1-F][Triazin-4-Amino] Adenine C-Nucleoside (GS-5734) for the Treatment of Ebola and Emerging Viruses. J Med Chem (2017) 60(5):1648–61. doi: 10.1021/acs.jmedchem.6b01594
139. Furuta Y, Komeno T, Nakamura T. Favipiravir (T-705), a Broad Spectrum Inhibitor of Viral RNA Polymerase. Proc Jpn Acad Ser B Phys Biol Sci (2017) 93(7):449–63. doi: 10.2183/pjab.93.027
140. Chu C, Cheng V, Hung I, Wong M, Chan K, Chan K, et al. Role of Lopinavir/Ritonavir in the Treatment of SARS: Initial Virological and Clinical Findings. Thorax (2004) 59(3):252–6. doi: 10.1136/thorax.2003.012658
141. Cao B, Wang Y, Wen D, Liu W, Wang J, Fan G, et al. A Trial of Lopinavir–Ritonavir in Adults Hospitalized With Severe Covid-19. N Engl J Med (2020) 382(19):1787–99. doi: 10.1056/NEJMoa2001282
142. Ren W, Ma Y, Wang R, Liang P, Sun Q, Pu Q, et al. Research Advance on Qingfei Paidu Decoction in Prescription Principle, Mechanism Analysis and Clinical Application. Front Pharmacol (2021) 11:589714. doi: 10.3389/fphar.2020.589714
143. Ren JL, Zhang AH, Wang XJ. Traditional Chinese Medicine for COVID-19 Treatment. Pharmacol Res (2020) 155:104743. doi: 10.1016/j.phrs.2020.104743
144. Zhang Y, Xie H, Li Y, Li T, Yuan H, Fu X, et al. Qingfei Paidu Decoction for Treating COVID-19: A Protocol for a Meta-Analysis and Systematic Review of Randomized Controlled Trials. Med (Baltimore) (2020) 99(36):e22040. doi: 10.1097/MD.0000000000022040
145. Leung PC. The Efficacy of Chinese Medicine for SARS: A Review of Chinese Publications After the Crisis. Am J Chin Med (2007) 35(4):575–81. doi: 10.1142/S0192415X07005077
146. Tsai YC, Lee CL, Yen HR, Chang YS, Lin YP, Huang SH, et al. Antiviral Action of Tryptanthrin Isolated From Strobilanthes Cusia Leaf Against Human Coronavirus NL63. Biomolecules (2020) 10(3):366. doi: 10.3390/biom10030366
147. Shree P, Mishra P, Selvaraj C, Singh SK, Chaube R, Garg N, et al. Targeting COVID-19 (SARS-CoV-2) Main Protease Through Active Phytochemicals of Ayurvedic Medicinal Plants - Withania Somnifera (Ashwagandha), Tinospora Cordifolia (Giloy) and Ocimum Sanctum (Tulsi) - A Molecular Docking Study. J Biomol Struct Dyn (2020) 1–14. doi: 10.1080/07391102.2020.1810778
148. Shanmugarajan D, Prabitha P, Kumar BRP, Suresh B. Curcumin to Inhibit Binding of Spike Glycoprotein to ACE2 Receptors: Computational Modelling, Simulations, and ADMET Studies to Explore Curcuminoids Against Novel SARS-CoV-2 Targets. RSC Adv (2020) 10(52):31385–99. doi: 10.1039/D0RA03167D
149. Patel A, Rajendran M, Shah A, Patel H, Pakala SB, Karyala P. Virtual Screening of Curcumin and Its Analogs Against the Spike Surface Glycoprotein of SARS-CoV-2 and SARS-CoV. J Biomol Struct Dyn (2020) 5:1–9. doi: 10.1080/07391102.2020.1868338
150. Baig MH, Ahmad K, Roy S, Ashraf JM, Adil M, Siddiqui MH, et al. Computer Aided Drug Design: Success and Limitations. Curr Pharm Des (2016) 22:572–81. doi: 10.2174/1381612822666151125000550
151. Beck BR, Shin B, Choi Y, Park S, Kang K. Predicting Commercially Available Antiviral Drugs That may Act on the Novel Coronavirus (SARS-CoV-2) Through a Drug-Target Interaction Deep Learning Model. Comput Struct Biotechnol J (2020) 18:784–90. doi: 10.1016/j.csbj.2020.03.025
152. Yu R, Chen L, Lan R, Shen R, Li P. Computational Screening of Antagonists Against the SARS-CoV-2 (COVID-19) Coronavirus by Molecular Docking. Int J Antimicrob Agents (2020) 56(2):106012. doi: 10.1016/j.ijantimicag.2020.106012
153. Bouchentouf S, Missoum N. Identification of Compounds From Nigella Sativa as New Potential Inhibitors of 2019 Novel Coronavirus (COVID-19). Mol Docking Study (2020). doi: 10.20944/preprints202004.0079.v1
154. Horvath D, Orlov A, Osolodkin DI, Ishmukhametov AA, Marcou G, Varnek A. A Chemographic Audit of Anti-Coronavirus Structure-Activity Information From Public Databases (ChEMBL). Mol Inform (2020) 39(12):e2000080. doi: 10.1002/minf.202000080
155. Elfiky AA. SARS-CoV-2 RNA Dependent RNA Polymerase (RdRp) Targeting: An in Silico Perspective. J Biomol Struct Dyn (2021) 39(9):3204–12. doi: 10.1080/07391102.2020.1761882
156. Wang J. Fast Identification of Possible Drug Treatment of Coronavirus Disease-19 (COVID-19) Through Computational Drug Repurposing Study. J Chem Inf Model (2020) 60(6):3277–86. doi: 10.1021/acs.jcim.0c00179
157. Shah B, Modi P, Sagar SR. In Silico Studies on Therapeutic Agents for COVID-19: Drug Repurposing Approach. Life Sci (2020) 252:117652. doi: 10.1016/j.lfs.2020.117652
158. Muralidharan N, Sakthivel R, Velmurugan D, Gromiha MM. Computational Studies of Drug Repurposing and Synergism of Lopinavir, Oseltamivir and Ritonavir Binding With SARS-CoV-2 Protease Against COVID-19. J Biomol Struct Dyn (2021) 39(7):2673–8. doi: 10.1080/07391102.2020.1752802
159. Ton AT, Gentile F, Hsing M, Ban F, Cherkasov A. Rapid Identification of Potential Inhibitors of SARS-CoV-2 Main Protease by Deep Docking of 1.3 Billion Compounds. Mol Inform (2020) 39(8):2000028. doi: 10.1002/minf.202000028
160. Hall DC Jr., Ji H-F. A Search for Medications to Treat COVID-19 via in Silico Molecular Docking Models of the SARS-CoV-2 Spike Glycoprotein and 3CL Protease. Travel Med Infect Dis (2020) 35:101646. doi: 10.1016/j.tmaid.2020.101646
161. Choi SY, Heo MJ, Lee C, Choi YM, An IS, Bae S, et al. 2-Deoxy-D-Glucose Ameliorates Animal Models of Dermatitis. Biomedicines (2020) 8(2):20. doi: 10.3390/biomedicines8020020
162. Verma A, Adhikary A, Woloschak G, Dwarakanath BS, Papineni RVL. A Combinatorial Approach of a Polypharmacological Adjuvant 2-Deoxy-D-Glucose With Low Dose Radiation Therapy to Quell the Cytokine Storm in COVID-19 Management. Int J Radiat Biol (2020) 96(11):1323–8. doi: 10.1080/09553002.2020.1818865
163. Almasi F, Mohammadipanah F. Hypothetical Targets and Plausible Drugs of Coronavirus Infection Caused by SARS-CoV-2. Transbound Emerg Dis (2021) 68(2):318–32. doi: 10.1111/tbed.13734
164. Santosh KC. AI-Driven Tools for Coronavirus Outbreak: Need of Active Learning and Cross-Population Train/Test Models on Multitudinal/Multimodal Data. J Med Syst (2020) 44(5):1–5. doi: 10.1007/s10916-020-01562-1
165. Maghdid HS, Ghafoor KZ. A Smartphone Enabled Approach to Manage COVID-19 Lockdown and Economic Crisis. SN Comput Sci (2020) 1(5):271. doi: 10.1007/s42979-020-00290-0
166. Arora N, Banerjee AK, Narasu ML. The Role of Artificial Intelligence in Tackling COVID-19. Future Virol (2020) 15(11):717–24. doi: 10.2217/fvl-2020-0130
167. Barstugan M, Ozkaya U, Ozturk S. Coronavirus (Covid-19) Classification Using Ct Images by Machine Learning Methods. arXiv Preprint (2003). doi:arXiv:2003.09424v1
168. Bullock J, Luccioni A, Pham KH, Lam CSN, Luengo-Oroz M. Mapping the Landscape of Artificial Intelligence Applications Against COVID-19. J Artif Intell Res (2020) 69:807–45. doi: 10.1613/jair.1.12162
169. Naudé W. Artificial Intelligence vs COVID-19: Limitations, Constraints and Pitfalls. AI Soc (2020) 35(3):761–5. doi: 10.1007/s00146-020-00978-0
170. Vaishya R, Javaid M, Khan IH, Haleem A. Artificial Intelligence (AI) Applications for COVID-19 Pandemic. Diabetes Metab Syndr Clin Res Rev (2020) 14(4):337–9. doi: 10.1016/j.dsx.2020.04.012
171. Abd El Wahed A, Patel P, Heidenreich D, Hufert FT, Weidmann M. Reverse Transcription Recombinase Polymerase Amplification Assay for the Detection of Middle East Respiratory Syndrome Coronavirus. PloS Curr (2013) 5:ecurrents.outbreaks.62df1c7c75ffc96cd59034531e2e8364. doi: 10.1371/currents.outbreaks.62df1c7c75ffc96cd59034531e2e8364
172. Augustine R, Hasan A, Das S, Ahmed R, Mori Y, Notomi T, et al. Loop-Mediated Isothermal Amplification (LAMP): A Rapid, Sensitive, Specific, and Cost-Effective Point-Of-Care Test for Coronaviruses in the Context of COVID-19 Pandemic. Biol (Basel) (2020) 9:182. doi: 10.3390/biology9080182
173. Bernheim A, Mei X, Huang M, Yang Y, Fayad ZA, Zhang N, et al. Chest CT Findings in Coronavirus Disease-19 (COVID-19): Relationship to Duration of Infection. Radiol (2020) 295(3):685–91. doi: 10.1148/radiol.2020200463
174. Brigger D, Horn MP, Pennington LF, Powell AF, Siegrist D, Weber B, et al. Accuracy of Serological Testing for SARS-CoV-2 Antibodies: First Results of a Large Mixed-Method Evaluation Study. Allergy (2021) 76(3):853–65. doi: 10.1111/all.14608
175. Broughton JP, Deng X, Yu G, Fasching CL, Singh J, Streithorst J, et al. Rapid Detection of 2019 Novel Coronavirus SARS-CoV-2 Using a CRISPR-Based DETECTR Lateral Flow Assay. medRxiv (2020). doi: 10.1101/2020.03.06.20032334
176. Bustin SA, Benes V, Nolan T, Pfaffl MW. Quantitative Real-Time RT-PCR–a Perspective. J Mol Endocrinol (2005) 34(3):597–601. doi: 10.1677/jme.1.01755
177. Cai XF, Chen J, Hu JL, Long Q, Deng H, Liu P, et al. A Peptide-Based Magnetic Chemiluminescence Enzyme Immunoassay for Serological Diagnosis of Coronavirus Disease 2019. J Infect Dis (2020) 222(2):189–93. doi: 10.1093/infdis/jiaa243
178. Understanding Long COVID: A Modern Medical Challenge. Lancet 8: 398(10302):725. doi: 10.1016/S0140-6736(21)01900-0
179. Deiman B, van Aarle P, Sillekens P. Characteristics and Applications of Nucleic Acid Sequence-Based Amplification (NASBA). Mol Biotechnol (2002) 20:163–79. doi: 10.1385/MB:20:2:163
180. Dinnes J, Deeks JJ, Berhane S, Taylor M, Adriano A, Davenport C, et al. Rapid, Point-of-Care Antigen and Molecular-Based Tests for Diagnosis of SARS-CoV-2 Infection. Cochrane Database Syst Rev (2020) 3(3):CD013705. doi: 10.1002/14651858.CD013705.pub2
181. Faccini-Martínez ÁA, Rivero R, Garay E, García A, Mattar S, Botero Y, et al. Serological Cross-Reactivity Using a SARS-CoV-2 ELISA Test in Acute Zika Virus Infection, Colombia. Int J Infect Dis (2020) 101:191–3. doi: 10.1016/j.ijid.2020.09.1451
182. Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, et al. Nucleic Acid Detection With CRISPR-Cas13a/C2c2. Science (2017) 356(6336):438–42. doi: 10.1126/science.aam9321
183. Howson ELA, Kurosaki Y, Yasuda J, Takahashi M, Goto H, Gray AR, et al. Defining the Relative Performance of Isothermal Assays That can be Used for Rapid and Sensitive Detection of Foot-and-Mouth Disease Virus. J Virol Methods (2017) 249:102–10. doi: 10.1016/j.jviromet.2017.08.013
184. Ji T, Liu Z, Wang G, Guo X, Akbar Khan S, Lai C, et al. Detection of COVID-19: A Review of the Current Literature and Future Perspectives. Biosens Bioelectron (2020) 166:112455. doi: 10.1016/j.bios.2020.112455
185. Keightley MC, Sillekens P, Schippers W, Rinaldo C, George KST. Real-Time NASBA Detection of SARS-Associated Coronavirus and Comparison With Real-Time Reverse Transcription-PCR. J Med Virol (2005) 77(4):602–8. doi: 10.1002/jmv.20498
186. Kellner MJ, Koob JG, Gootenberg JS, Abudayyeh OO, Zhang F. SHERLOCK: Nucleic Acid Detection With CRISPR Nucleases. Nat Protoc (2019) 14(10):2986–3012. doi: 10.1038/s41596-019-0210-2
187. Koczula KM, Gallotta A. Lateral Flow Assays. Essays Biochem (2016) 60(1):111–20. doi: 10.1042/EBC20150012
188. Li Y, Yao L, Li J, Chen L, Song Y, Cai Z, et al. Stability Issues of RT-PCR Testing of SARS-CoV-2 for Hospitalized Patients Clinically Diagnosed With COVID-19. J Med Virol (2020) 92(7):903–8. doi: 10.1002/jmv.25786
189. Manigandan S, Wu MT, Ponnusamy VK, Raghavendra VB, Pugazhendhi A, Brindhadevi K. A Systematic Review on Recent Trends in Transmission, Diagnosis, Prevention and Imaging Features of COVID-19. Process Biochem (2020) 98:233–40. doi: 10.1016/j.procbio.2020.08.016
190. Nicol T, Lefeuvre C, Serri O, Pivert A, Joubaud F, Dubée V, et al. Assessment of SARS-CoV-2 Serological Tests for the Diagnosis of COVID-19 Through the Evaluation of Three Immunoassays: Two Automated Immunoassays (Euroimmun and Abbott) and One Rapid Lateral Flow Immunoassay (NG Biotech). J Clin Virol (2020) 129:104511. doi: 10.1016/j.jcv.2020.104511
191. Niesters HGM. Clinical Virology in Real Time. J Clin Virol (2002) 25:S3–12. doi: 10.1016/s1386-6532(02)00197-x
192. Pan F, Ye T, Sun P, Gui S, Liang B, Li L, et al. Time Course of Lung Changes at Chest CT During Recovery From Coronavirus Disease 2019 (COVID-19). Radiology (2020) 295(3):715–21. doi: 10.1148/radiol.2020200370
193. Pieri M, Ciotti M, Carlozzi N, Frassanito ML, Meloni A, Cistera A, et al. SARS-CoV-2 Infection Serology Validation of Different Methods: Usefulness of IgA in the Early Phase of Infection. Clin Chim Acta (2020) 511:28–32. doi: 10.1016/j.cca.2020.09.033
194. Ragnesola B, Jin D, Lamb CC, Shaz BH, Hillyer CD, Luchsinger LL. COVID19 Antibody Detection Using Lateral Flow Assay Tests in a Cohort of Convalescent Plasma Donors. BMC Res Notes (2020) 13(1):372. doi: 10.1186/s13104-020-05212-0
195. Scohy A, Anantharajah A, Bodéus M, Kabamba-Mukadi B, Verroken A, Rodriguez-Villalobos H. Low Performance of Rapid Antigen Detection Test as Frontline Testing for COVID-19 Diagnosis. J Clin Virol 129:104455. doi: 10.1016/j.jcv.2020.104455
196. Shao JM, Ayuso SA, Deerenberg EB, Elhage SA, Augenstein VA, Heniford BT. A Systematic Review of CT Chest in COVID-19 Diagnosis and Its Potential Application in a Surgical Setting. Colorectal Dis (2020) 22(9):993–1001. doi: 10.1111/codi.15252
197. Wu Q, Suo C, Brown T, Wang T, Teichmann SA, Bassett AR. INSIGHT: A Population-Scale COVID-19 Testing Strategy Combining Point-of-Care Diagnosis With Centralized High-Throughput Sequencing. Sci Adv (2021) 7(7):eabe5054. doi: 10.1126/sciadv.abe5054
198. Xia S, Chen X. Single-Copy Sensitive, Field-Deployable, and Simultaneous Dual-Gene Detection of SARS-CoV-2 RNA via Modified RT-RPA. Cell Discovery (2020) 6:37. doi: 10.1038/s41421-020-0175-x
199. Xiang J, Yan M, Li H, Liu T, Lin C, Huang S, et al. Evaluation of Enzyme-Linked Immunoassay and Colloidal Gold-Immunochromatographic Assay Kit for Detection of Novel Coronavirus (SARS-Cov-2) Causing an Outbreak of Pneumonia (COVID-19). MedRxiv (2020). doi: 10.1101/2020.02.27.20028787
200. Zhang W, Du RH, Li B, Zheng XS, Yang X-L, Hu B, et al. Molecular and Serological Investigation of 2019-Ncov Infected Patients: Implication of Multiple Shedding Routes. Emerg Microbes Infect (2020) 9(1):386–9. doi: 10.1080/22221751.2020.1729071
201. Rashid ZZ, Othman SN, Samat MN, Ali UK, Wong KK. Diagnostic Performance of COVID-19 Serology Assays. Malays J Pathol (2020) 42(1):13–21.
202. Sahoo PR, Sethy K, Mohapatra S, Panda D. Loop Mediated Isothermal Amplification: An Innovative Gene Amplification Technique for Animal Diseases. Vet World (2016) 9(5):465–9. doi: 10.14202/vetworld.2016.465-469
203. Subsoontorn P, Lohitnavy M, Kongkaew C. The Diagnostic Accuracy of Isothermal Nucleic Acid Point-of-Care Tests for Human Coronaviruses: A Systematic Review and Meta-Analysis. Sci Rep (2020) 10:22349. doi: 10.1038/s41598-020-79237-7
204. Allen JWL, Verkerke H, Owens J, Saeedi B, Boyer D, Shin S, et al. Serum Pooling for Rapid Expansion of Anti-SARS-CoV-2 Antibody Testing Capacity. Transfus Clin Biol (2021) 28(1):51–4. doi: 10.1016/j.tracli.2020.10.008
205. Soares F, Villavicencio A, José Anzanello M, Fogliatto FS, Idiart MAP, Stevenson M. A Novel High Specificity COVID-19 Screening Method Based on Simple Blood Exams and Artificial Intelligence. medRxiv (2020) 2020:4. doi: 10.1101/2020.04.10.20061036
206. Dong D, Tang Z, Wang S, et al. The Role of Imaging in the Detection and Management of COVID-19: A Review. IEEE Rev BioMed Eng (2021). doi: 10.1109/RBME.2020.2990959
207. Li L, Qin L, Xu Z, Yin Y, Wang X, Kong B, et al. Using Artificial Intelligence to Detect COVID-19 and Community-Acquired Pneumonia Based on Pulmonary CT: Evaluation of the Diagnostic Accuracy. Radiology (2020) 296:E65–71. doi: 10.1148/radiol.2020200905
208. Apostolopoulos ID, Mpesiana TA. Covid-19: Automatic Detection From X-Ray Images Utilizing Transfer Learning With Convolutional Neural Networks. Phys Eng Sci Med (2020) 43(2):635–40. doi: 10.1007/s13246-020-00865-4
Keywords: SARS-CoV-2, COVID-19, ORFs, machine learning, co-morbidities
Citation: Kaur A, Chopra M, Bhushan M, Gupta S, Kumari P H, Sivagurunathan N, Shukla N, Rajagopal S, Bhalothia P, Sharma P, Naravula J, Suravajhala R, Gupta A, Abbasi BA, Goswami P, Singh H, Narang R, Polavarapu R, Medicherla KM, Valadi J, Kumar S A, Chaubey G, Singh KK, Bandapalli OR, Kavi Kishor PB and Suravajhala P (2021) The Omic Insights on Unfolding Saga of COVID-19. Front. Immunol. 12:724914. doi: 10.3389/fimmu.2021.724914
Received: 14 June 2021; Accepted: 27 September 2021;
Published: 20 October 2021.
Edited by:
Betsy J. Barnes, Feinstein Institute for Medical Research, United StatesReviewed by:
Wanpen Chaicumpa, Mahidol University, ThailandSiavash Bolourani, Feinstein Institute for Medical Research, United States
Copyright © 2021 Kaur, Chopra, Bhushan, Gupta, Kumari P, Sivagurunathan, Shukla, Rajagopal, Bhalothia, Sharma, Naravula, Suravajhala, Gupta, Abbasi, Goswami, Singh, Narang, Polavarapu, Medicherla, Valadi, Kumar S, Chaubey, Singh, Bandapalli, Kavi Kishor and Suravajhala. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Prashanth Suravajhala, cHJhc2hAYmlvY2x1ZXMub3Jn; Polavarapu Bilhan Kavi Kishor, cGJrYXZpQHlhaG9vLmNvbQ==; Keshav K. Singh, a2tzaW5naEB1YWIuZWR1; Obul Reddy Bandapalli, YmFuZGFwYWxsaUBnbWFpbC5jb20=;
†These authors have contributed equally to this work
 Arvinpreet Kaur
Arvinpreet Kaur Mehak Chopra
Mehak Chopra Mahak Bhushan
Mahak Bhushan Sonal Gupta
Sonal Gupta Hima Kumari P
Hima Kumari P Narmadhaa Sivagurunathan5†
Narmadhaa Sivagurunathan5† Nidhi Shukla
Nidhi Shukla Purnima Sharma
Purnima Sharma Renuka Suravajhala
Renuka Suravajhala Ayam Gupta
Ayam Gupta Bilal Ahmed Abbasi
Bilal Ahmed Abbasi Prittam Goswami
Prittam Goswami Jayaraman Valadi
Jayaraman Valadi Anil Kumar S
Anil Kumar S Gyaneshwer Chaubey
Gyaneshwer Chaubey Keshav K. Singh
Keshav K. Singh Obul Reddy Bandapalli
Obul Reddy Bandapalli Polavarapu Bilhan Kavi Kishor
Polavarapu Bilhan Kavi Kishor Prashanth Suravajhala
Prashanth Suravajhala