- 1School of Public Health, China Medical University, Shenyang, Liaoning province, China
- 2Department of Emergency Response, Chinese PLA Center for Disease Control and Prevention, Beijing, China
- 3Department of Information, Chinese PLA Center for Disease Control and Prevention, Beijing, China
- 4Department of Clinical Laboratory, Seventh Medical Center of Chinese PLA General Hospital, Beijing, China
- 5College of Public Health, Zhengzhou University, Zhengzhou, Henan province, China
Klebsiella michiganensis is an increasingly important bacterial pathogen causing nosocomial infections in clinical patients. In this study, we described the molecular and genomic characteristics of a carbapenem-resistant K. michiganensis strain KM166 cultured from a one-month premature baby’s blood sample. KM166 showed lower biofilm forming ability in optical density (OD) than K. pneumoniae NTUH-K2044 (0.271 ± 0.027 vs. 0.595 ± 0.054, p = 0.001), and the median lethal dose (0.684 lg CFU/mL) was lower than K. pneumoniae strain NTUH-K2044 (6.679 lg CFU/mL). A IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in KM166 was identified carrying three replicon types. It has low homology to Escherichia coli pMRY09-581ECO_1 and the highest homology similarity to the INcFIA/INcFII(p14)-type plasmid in K. michiganensis strain fxq plasmid pB_KPC, suggesting that this multiple replicon plasmid was unlikely to have been transmitted from E. coli and probably a transfer of repFIB replicon genes from other K. michiganensis strains into the INcFIA/INcFII(p14)-type plasmid of KM166 had occurred. Mapping of the gene environment revealed that blaKPC-2 in KM166 plasmid 3 had high identity and same Tn3-tnpR-IS481-blaKPC-2-klcA_1 genomic context structure with K. pneumoniae strain JKP55, plasmid pKPC-J5501, and blaKPC-2-carrying plasmid proved to be autonomously transferred under the help of mobile genetic elements into Escherichia coli 600 by plasmid conjugation experiment. In conclusion, we have characterized a K. michiganensis strain carrying multi-replicon IncFII/IncFIA(HI1)/IncFIB(K) plasmid and blaKPC-2-carrying IncFII(p14)/IncFIA plasmid in this study, which provided insights about the evolutionary diversity of plasmids carried by K. michiganensis.
1. Introduction
Klebsiella michiganensis is one of the bacterial pathogen of the Klebsiella genus (Wyres et al., 2020), it seldom appeared in clinical hospitals at the past, but has been gradually increasing in recent years, for several K. michiganensis strains have been found to be colonized in patients. Carbapenemase production is the common carbapenemase resistance mechanism in clinical infections, such as Acinetobacter baumannii (Oinuma et al., 2021), K. pneumoniae (Pitout et al., 2015) and Pseudomonas aeruginosa (Nordmann et al., 2011; Doi and Paterson, 2015; Iovleva and Doi, 2017). The main carbapenemase genes carried by K. michiganensis include blaKPC-2, blaVIM-1 (Campos-Madueno et al., 2021), blaNDM-1 (Li et al., 2021),blaNDM-5 (Zheng et al., 2018), blaKPC-3 (Seiffert et al., 2019), blaIMP-4 (Li et al., 2021; Zhang et al., 2021), blaOXA-181 (Founou et al., 2018),blaSIM-1, blaOXA-1, blaCTX-M-14 (Xu et al., 2022), etc.
Klebsiella michiganensis has appeared in parts of the world (Ccorahua-Santo et al., 2017; King et al., 2021; Luo et al., 2022), there are highly homology and similarities in the molecular characteristics between K. michiganensis and K. pneumoniae. Some K. michiganensis strains have been found in plants or ecosystems (Kang et al., 2020; Salgueiro et al., 2020; Liu, L. et al., 2021), but the majority have been found in patients (Abed et al., 2021). Due to the cadmium resistance and plant growth promoting properties of K. michiganensis, many studies have been conducted to examine the effects of physical and biochemical stimuli such as PH, cadmium and environmental changes on the growth and genome sequence of K. michiganensis, with many positive results obtained (Mitra et al., 2018, 2019; Oliveira et al., 2020; Yan et al., 2021). However, more research is needed to examine the genomic characteristics of K. michiganensis strains from clinical patients, as well as its drug resistance and virulence profile (Zhang et al., 2021).
2. Materials and methods
2.1. Sample sources and strain identification
A total of 359 strains isolated from a tertiary hospital in Beijing, China were identified to be K. pneumoniae through biochemical indicators during 2017–2020, and 147 strains were resistant to carbapenem antibiotics. One of these 147 carbapenem resistant strain (KM166) was identified as K. michiganensis after whole genome sequencing. KM166 strain was cultured from a one-month premature baby’s blood on July 31, 2018, with a utilization history of penicillin, monocyclic β-lactams and cephalosporins. The patient was admitted on June 23, 2018 and discharged on September 5, 2018, with a hospitalization period of 75 days and had been treated with central venous cannulation, ventilator and blood transfusion. KM166 was cultured in a Luria-Bertani (LB) solid medium for 18–24 h in a constant temperature incubator at 37°C, the single colony was picked into LB liquid medium and incubated in a constant temperature shaker at 37°C for 18 h before being washed.
2.2. Antibiotic sensitivity and hypermucoviscous test
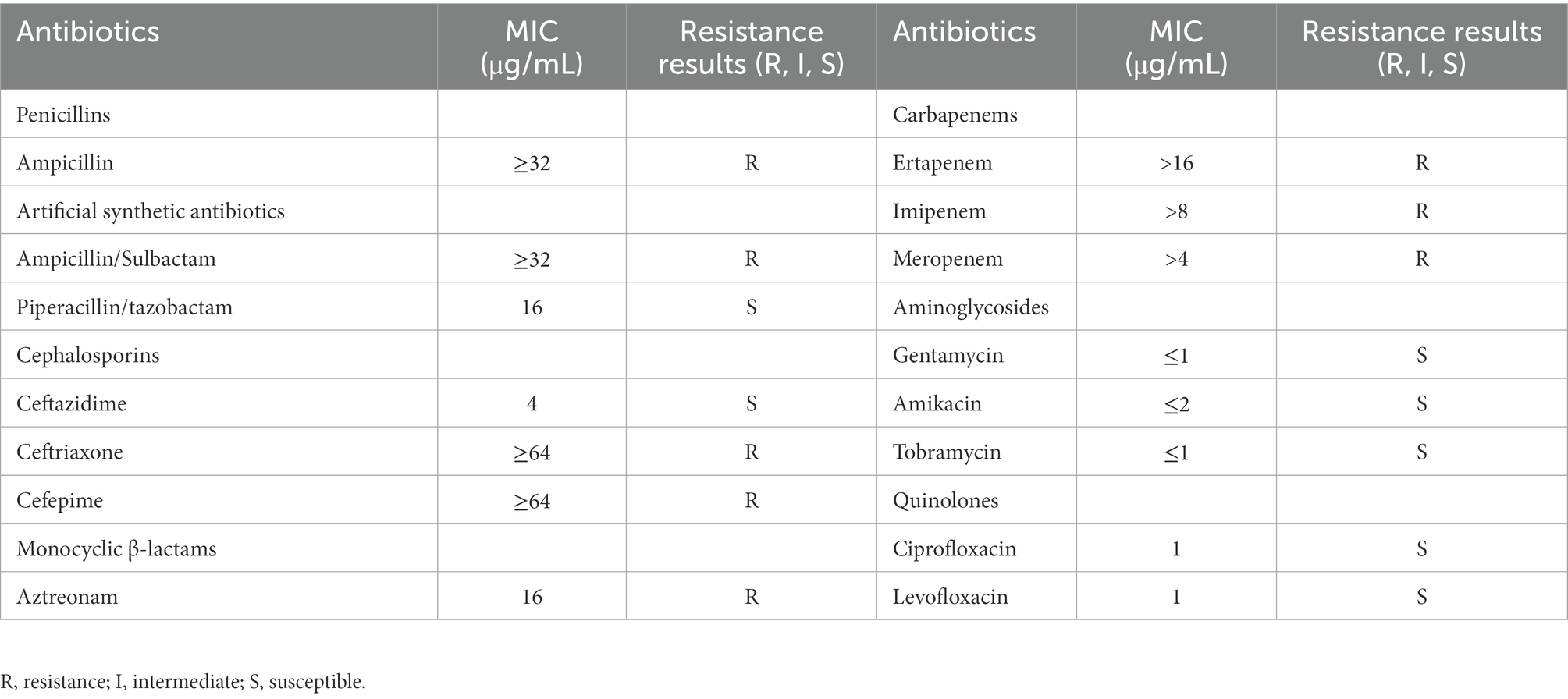
The sensitivity of common clinical antibiotics was tested using the Vitek-2 compact system and agar dilution method. KM166 strain were incubated on Columbia blood agar medium (Oxoid, Basingstoke, England) at 37°C for 18–24 h. Control strains Pseudomonas aeruginosa ATCC 27853 and Escherichia coli ATCC 25922 were used for antimicrobial susceptibility testing. The MICs were verified according to the Clinical and Laboratory Standards Institute (M100, 29th edition, 2019) guidelines. Standard bacteriological rings were used to assess hypermucoviscous phenotype, and the formation of slimy lines >5 mm in length was considered a positive string test (Catalán-Nájera et al., 2017).
2.3. Biofilm formation capacity test
Biofilm formation was measured by the absorbance at 570 nm of crystal violet staining. Hypervirulence K. pneumoniae strain NTUH-K2044 was served as the control and negative control was the group without the addition of bacterial solution. The strain was incubated in LB liquid medium until it reached a logarithmic growth cycle and then washed twice, followed by adjustment of the bacterial liquid to 1.0 McFarland’s concentration solution, then 200 μl LB culture solution and 10 μl bacterial solution were added to each well of the 96-well cell culture plate, and the 96-well cell culture plate was placed in a wet box at 35°C for 48 h. Afterwards, 200 μl of 1% crystalline violet staining solution was added to each well waiting for 10 min, and the plate was washed twice with distilled water and dried. The solution was transferred to a new 96-well plate and the absorbance value at 570 nm (A570) was observed using an enzyme marker (Liu, Y. et al., 2019).
2.4. The neutrophil killing assay
Serum from healthy people (#9193-10ML, purchased from Lablead, Beijing) was collected. The strain was cultured on Columbia blood agar medium (Oxoid, Basingstoke, England) for 15–18 h overnight and cultured in LB liquid medium for 18 h, washed twice with PBS buffer and then diluted to 1 × 106 CFU/ml. Mixtures of 75 μl serum and 25 μl diluted bacterial solution were collected at 37°C on a constant temperature shaker for 0 h, 1 h, 2 h, 3 h, and 4 h (Siu et al., 2011). The classical K. pneumoniae strain NTUH-K2044 was used as a control in the experiment, and observations were made to assess the bacterial killing effect of the serum by counting the average number of viable bacteria after taking 10 μl bacterial solution and applying it to three LB agar medium at different observation times.
2.5. The Galleria mellonela model
The larvae of Galleria mellonella (purchased from Tianjin Huiyude Biotechnology Co., Ltd.) that met the test standards were purchased, each weighing about 300–400 mg. There were two control groups: one received the same amount of PBS and the other was not treated. The strains were grown on LB agar medium for 18–20 h before the bacterial solution was diluted to 104–107 CFU/ml with PBS. A group of 10 G. mellonella was chosen at random, each K. michiganensis strain was injected into three groups and four different dilution concentrations (totally 120 G. mellonella). The hypervirulence K. pneumoniae strain NTUH-K2044 served as the control. A total of 20.0 μl bacterial suspension was injected into the first abdominal segment of G. mellonella using microinjector, then the death number of each group was observed and recorded every 6 h until 72 h. The median lethal dose (1og10 CFU/mL) of strains was calculated using the probit probability method of regression analysis (Gu et al., 2018).
2.6. Plasmid conjugation experiments and strain identification
KM166 carrying blaKPC-2 as donor and E. coli EC600 as recipient (Cai et al., 2008) were co-inoculated on dual resistant BHI solid medium (Meropenem 4 ug/mL and Rifampicin 2 mg/mL) and incubated for 48 h at 37°C. Colonies were grown and amplified using blaKPC-2 primers (F: CGGGTTGGACTCAAGACG; R: TGATGCGGTATTTTCTCC) to determine the blaKPC-2 in the conjugates. The conjugation bacterial species were subsequently identified using Vitek-2 compact system.
2.7. Genomic DNA extraction, sequencing and assembly
Strain DNA was extracted using four steps for long fragment DNA (lysis, purification, precipitation and elution). Pacbio sequel II and the DNBSEQ platform were used to sequence the genome of KM166. An insert DNA of 350 bp was identified to perform genome sequencing and the original PacBio long reads were error-corrected, trimmed and assembled using the program Canu v1.5 (Koren et al., 2017). The raw data was treated in order to obtain more accurate and reliable results in subsequent bioinformatics analysis: (1) remove reads containing a certain proportion of low quality (≤20) bases (40% as default); (2) removal of reads with a total number of bases containing N up to 10%; (3) remove adapter contamination; (4) remove duplication contamination.
2.8. Genome component prediction, gene annotation and protein classification
Species identification was performed using Kleborate software (Lam et al., 2021) to obtain multilocus sequence types (MLST). Genome annotation was performed using prokka v1.14.6 (Seemann, 2014). tRNA, rRNA and sRNA recognition were made use of tRNAscan-SE (Chan et al., 2021), Barrnap and the Rfam database. Prophage regions were predicted using the PHAgeSearch Tool (PHAST) web server1 and CRISPR identification using CRISPRCasFinder.2 Virulence and resistance genes were identified based on the database in VFDB (Virulence Factor Database; Liu, B. et al., 2019) and CARD (The Comprehensive Antibiotic Research Database; Alcock et al., 2020) using ABRicate.
2.9. Comparative genomics and phylogenetic analysis
The K. michiganensis strain K.m.2654 (accession no. GCA_003011775.1) isolated from Denmark in 2017 was selected for comparative genomic analysis with KM166. The comparison of K.m.2654 and KM166 was performed using blast and the average nucleotide identity comparisons between genomes were calculated using FastANI. The phylogenetic tree was constructed by maximum likelihood method based on single nucleotide polymorphism (SNP) using ClonalFrameML v1.12 (Didelot and Wilson, 2015) and roary v3.13.0. The tree building model was calculated by ModelTest-NG v0.1.7 (Flouri et al., 2015; Darriba et al., 2020) and plotted by RaxML-NG v1.1.0 (Kozlov et al., 2019), and the phylogenetic tree was finally visualized by FigTree v1.4.3 and iTOL (Letunic and Bork, 2021).
3. Results
3.1. Antimicrobial resistance and virulence phenotype
The in vitro antimicrobial susceptibility testing showed that K. michiganensis KM166 was resistant to Ampicillin, Aztreonam, carbapenems, and so on (Table 1). The string test showed negative result, which suggested that KM166 was not a hypermucoviscous strain. Both KM166 and classical K. pneumoniae strain NTUH-K2044 showed positive biofilm-forming ability (Figure 1A), but KM166 showed lower biofilm forming ability in optical density (OD) than K. pneumoniae NTUH-K2044 (0.271 ± 0.027 Vs. 0.595 ± 0.054, p = 0.001). The results of the serum killing experiment showed that human serum had a weak killing capacity against KM166, and KM166 has the stronger serum bactericidal capacity after 1 hour of serum and bacterial solution were mixed (Figure 1B). The median lethal dose of KM166 was 0.684 lg CFU/mL, which was much lower than the K. pneumoniae NTUH-K2044 strain (median lethal dose: 6.679 lg CFU/mL).
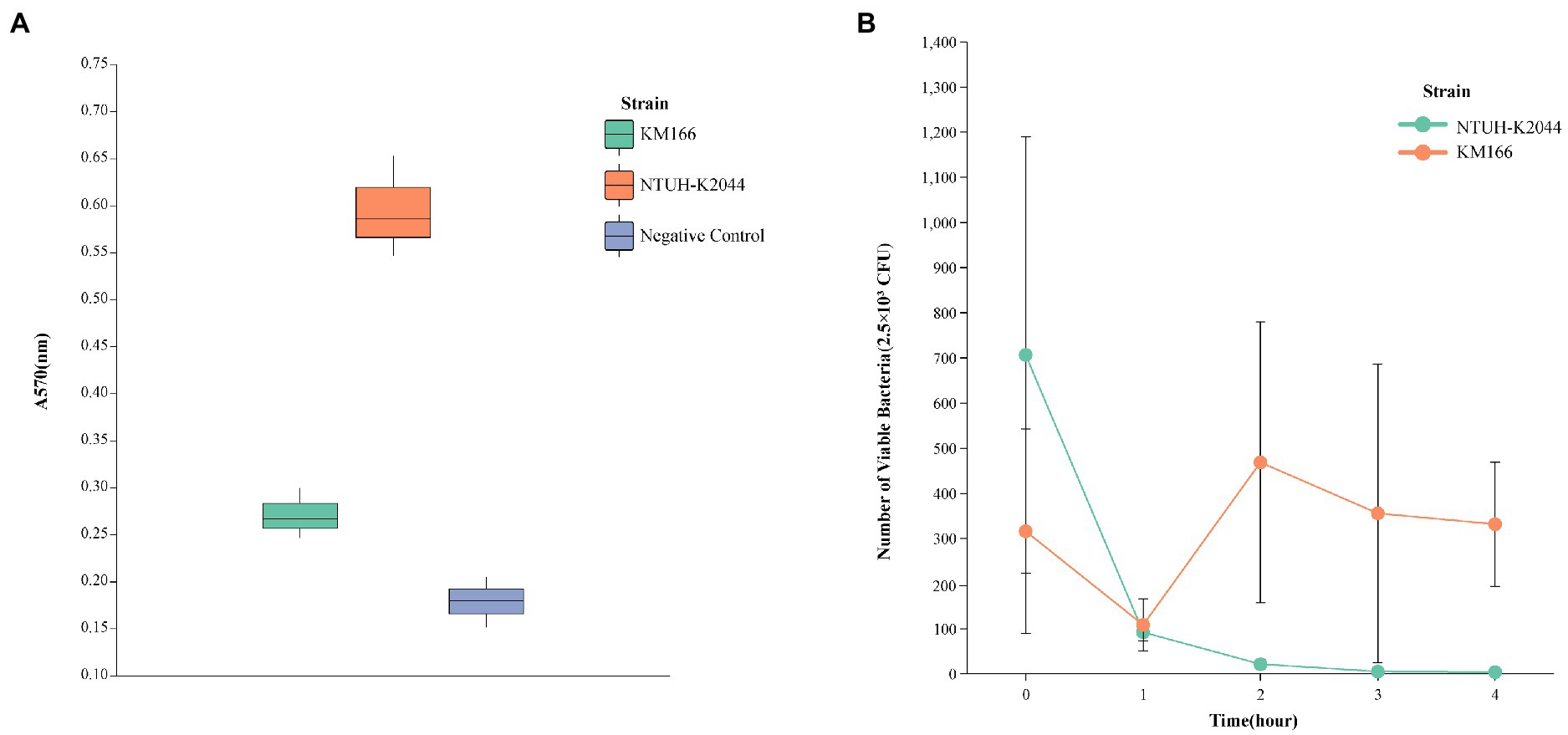
Figure 1. Defensive capability test of Klebsiella michiganensis KM166. (A) biofilm formation capacity test. (B) killing experiment with human neutrophils.
3.2. Genomic characterization
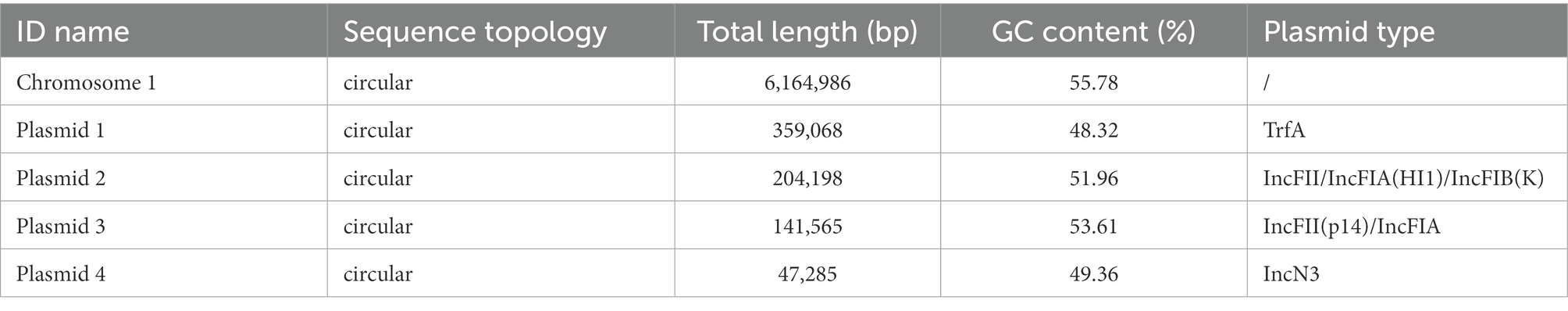
A total of 1,297 Mb whole-genome sequencing reads (187X) and 5,159 Mb (745X) PacBio platform reads were generated. The total genome of K. michiganensis KM166 has a size of 6,917,102 bp and contains 6,765 genes, with a GC content of 55.19 percent. The complete genome consists of a 6,164,986-bp single chromosome and four plasmids including Plasmid 1(359,068 bp), Plasmid 2(204,198 bp), Plasmid 3(141,565 bp) and Plasmid 4(47,285 bp). The G + C contents of the chromosome and the four plasmids were 55.78, 48.32, 51.96, 53.61 and 49.36%, respectively (Table 2). The plasmid types identified in KM166 were TrfA, IncFII/IncFIA(HI1)/IncFIB(K), IncFII(p14)/IncFIA and IncN3.
A total of 168 ncRNA (85 tRNAs, 25 rRNAs and 58 sRNAs) and 290 repeats were identified. In addition, three CRISPRs were identified in the chromosome of KM166, with lengths of 90 bp, 97 bp and 111 bp, respectively. Eleven prophages were identified on chromosomes and one on plasmid 3. It was found that the serotype of KM166 was KL41, three resistance genes blaOXY-1, blaCTX-M-37 and blaKPC-2 were present in chromosome, IncN3 plasmid and IncFII(p14)/IncFIA plasmid, respectively.
3.3. Comparison of the nucleic acid identity
Two clinical K. pneumoniae strains BSIKPN-9 and BSIKPN-11 collected from the same hospital as KM166 were randomly selected for comparison, and the average nucleotide identity (ANI) for K. michiganensis KM166 and K. pneumoniae (BSIKPN-9 and BSIKPN-11) were 84.64 and 84.65%, the range of which was between 83 and 95% of interspecific relationships. The ANI for KM166 and K. michiganensis strain K.m.2654 (accession no.GCF_003011775.1), on the other hand, was 99.81%, indicating intraspecific variation. Amino acid level synteny analysis of K.m.2654 and KM166 revealed that KM166 had 6,765 genes and K.m.2654 had 5,531 genes, with 5,052 aligned genes (74.68%, 5052/6765). Chromosomes of K.m.2654 and KM166 were found to be very similar, and the variants were mostly found in four plasmids (Figure 2).
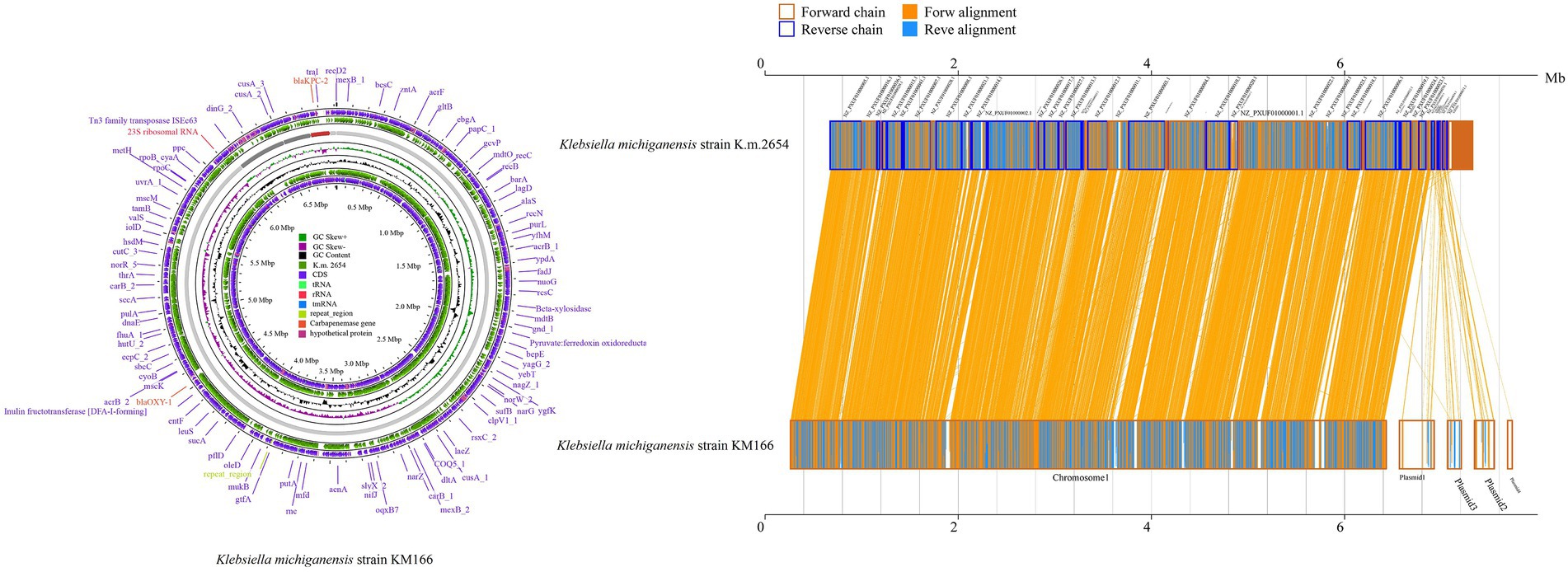
Figure 2. Amino acid level synteny analysis of Klebsiella michiganensis K.m.2654 and K. michiganensis KM166. Comparative genomic circle map was created using CGview (https://proksee.ca/). The yellow line in the middle region of the two sequences in the amino acid level synteny map indicates the forward alignment, and the blue line indicates the reverse complementary alignment.
3.4. Global evolutionary analysis of Klebsiella michiganensis
A total of 446 complete genomic assemblies classified as K. michiganensis were used to construct the phylogenetic tree, of which 445 downloaded from the Pathosystems Resource Integration Center (PATRIC), as well as one KM166 we collected in this study. Genomic information of 445 strains is provided in Supplementary Table S1. All 446 K. michiganensis strains were isolated from 27 countries, Switzerland was the main country source (52.24%, 233/446), followed by China with 50 strains (11.21%, 50/446). Five K. michiganensis strains were isolated from Beijing, China, but they did not carry blaKPC-2. The phylogenetic tree revealed a single major group, and the homology between strains was extremely high, with no significant evolutionary variation. KM166 was found at the end of an evolutionary branch that included some of the most recent strains, with the highest homology to a K. michiganensis strain collected in Switzerland (ANI: 99.93%). KM166 was collected from a hospital in Beijing on July 31, 2018, and several K. michiganensis strains with the highest homology were collected in 2017 (Figure 3).
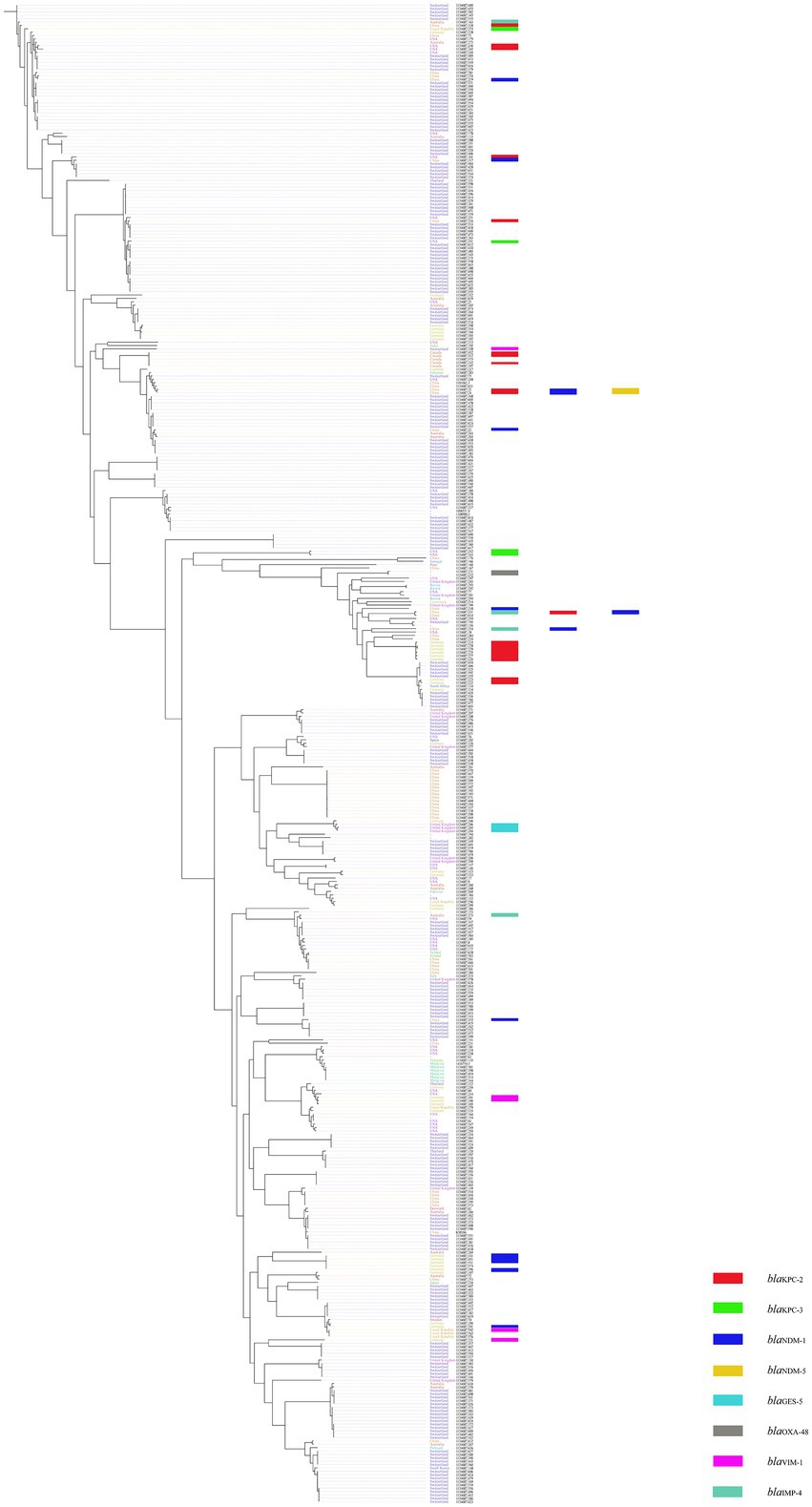

Figure 3. Phylogenetic tree of 445 Klebsiella michiganensis strains downloaded from Pathosystems Resource Integration Center and one Klebsiella michiganensis strain KM166 we collected. Plotted using ClonalFrameML and roray based on SNP. “-” means the collection of strains from unknown cities.
3.5. Comparative analysis of the blaKPC-2-harbouring Klebsiella michiganensis
Twenty-one blaKPC-2-carrying K. michiganensis strains were identified from 445 K. michiganensis strains downloaded from the PATRIC database, and they were then combined with KM166 for phylogenetic analysis in order to investigate their evolutionary relationships (Figure 4). It showed that two strains (1134687.279 and 1134687.221) isolated from the Czech Republic in 2018 and Germany in 2017 that shared high homology with KM166, particularly 1134687.221 (ANI: 99.15%). According to the phylogenetic tree, KM166 was located at a small branch and showed highly homologous to two strains isolated from geographically distant countries.
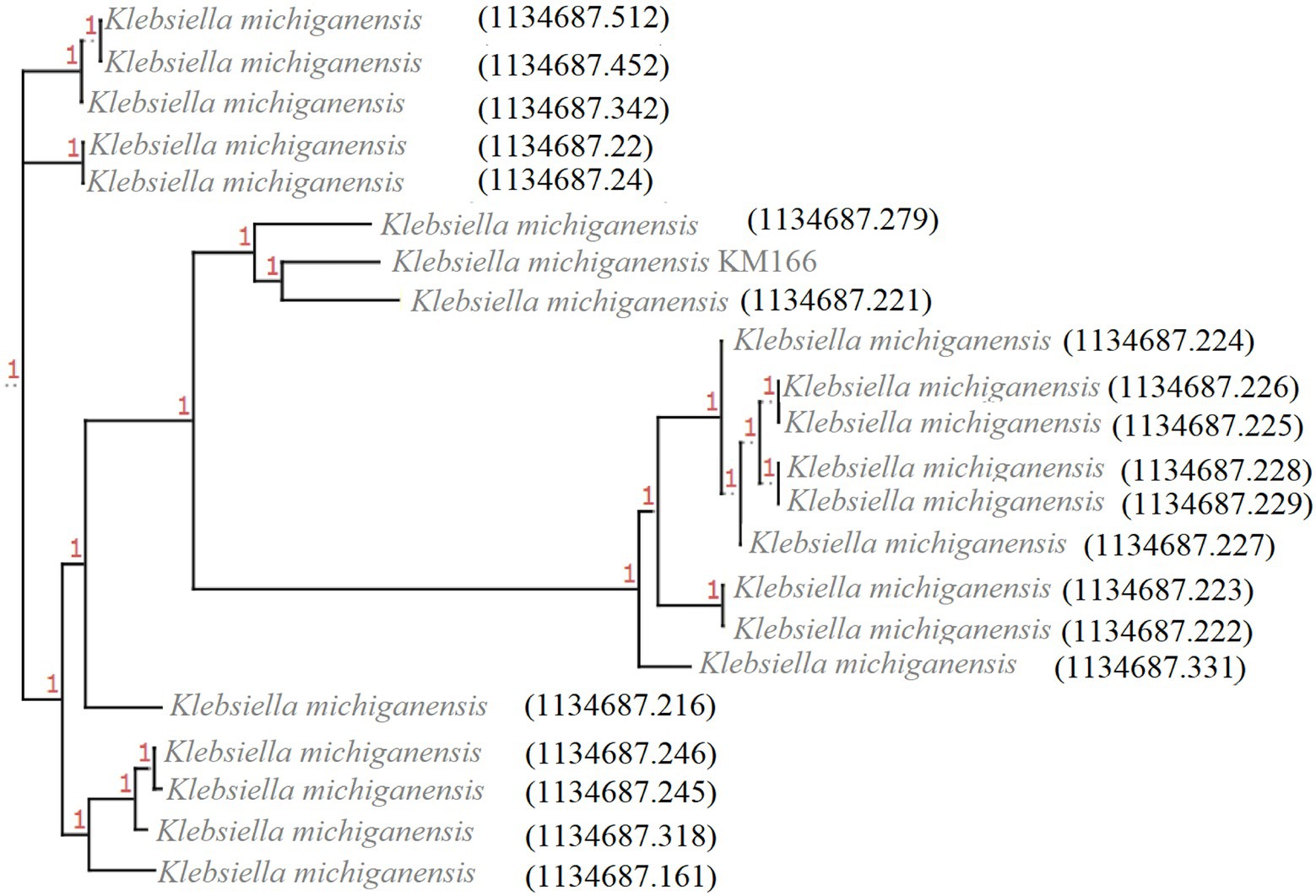
Figure 4. Phylogenetic tree of 22 Klebsiella michiganensis strains carrying blaKPC-2. The phylogenetic tree was generated using ClonalFrameML and roray based on SNP, plotted by treeview (http://etetoolkit.org/treeview/). 21 Klebsiella michiganensis strains carrying blaKPC-2 was identified from 445 K. michiganensis strains downloaded from Pathosystems Resource Integration Center and one K. michiganensis strain KM166 carrying blaKPC-2 was collected in this study.
The blaKPC-2 gene of KM166 was located in a 140 kb IncFII(p14)/IncFIA type plasmid 3, blasting with the nucleic acid database (NT) found that a plasmid named K. pneumoniae strain JKP55, plasmid pKPC-J5501(GenBank accession no. NZ_OL891656) showed 17% coverage and 99.97% identity to KM166 plasmid 3. Mapping of the gene environment using easyfig v2.2.5 (Sullivan et al., 2011) software revealed that blaKPC-2 in KM166 plasmid 3 had high identity and same Tn3-tnpR-IS481-blaKPC-2-klcA_1 genomic context structure with K. pneumoniae strain JKP55, plasmid pKPC-J5501, which carried two blaKPC-2 genes (Figure 5). Plasmid conjugation experiment demonstrated that the plasmid carrying blaKPC-2 could transfer autonomously into Escherichia coli 600.
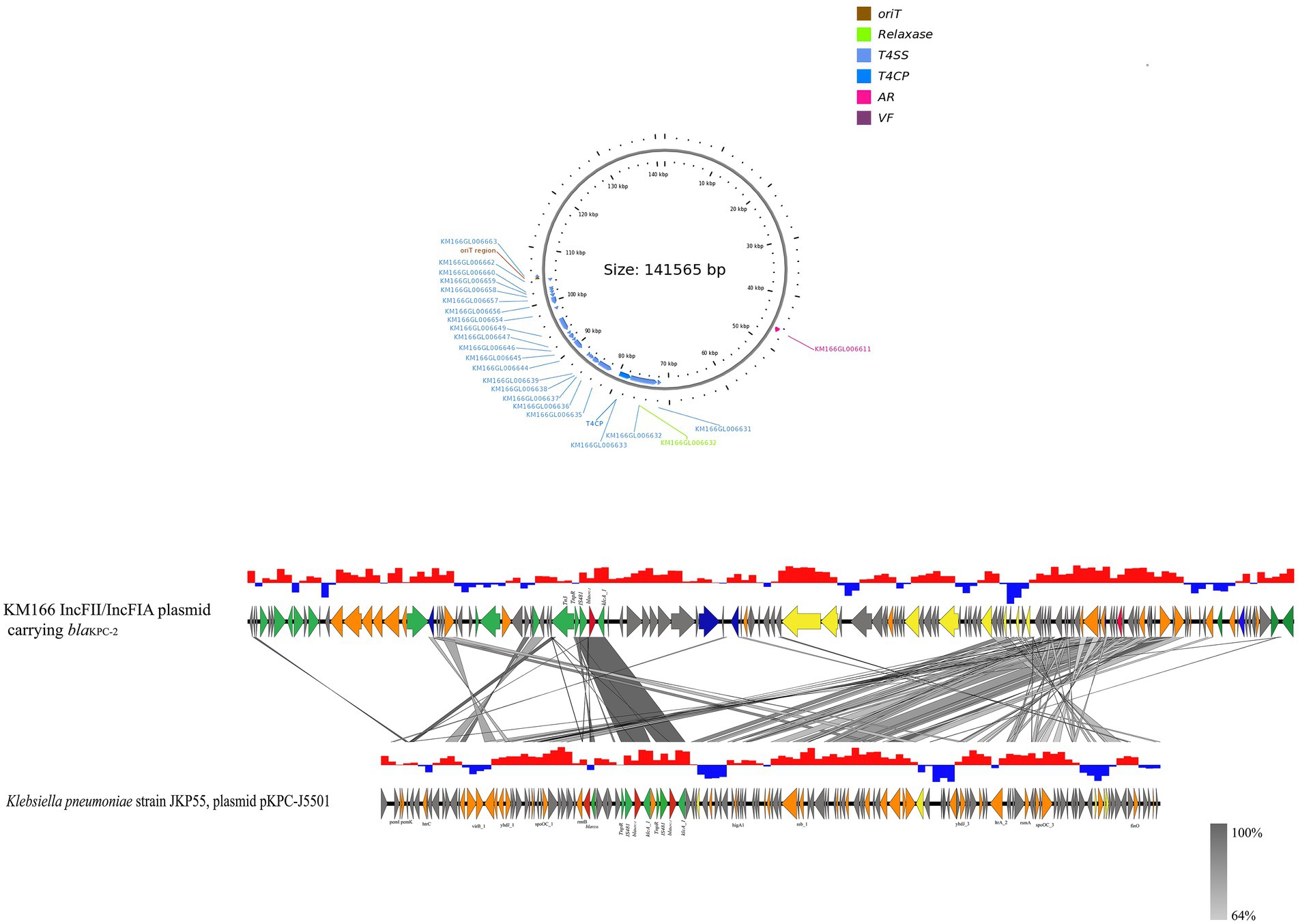
Figure 5. Sequencing comparison among the regions surrounding blaKPC-2 from KM166 IncFII(p14)/IncFIA plasmid and Klebsiella pneumoniae strain JKP55, plasmid pKPC- J5501, the most closely genetically matched complete plasmid sequences from NCBI. High homology shared sections are shaded in gray. An arrow is colored in accordance with its ostensible function. The function of genes about coupling transfer, plasmid stability, replication, mobile elements, antimicrobial resistance are yellow, orange, blue, green and red arrows, respectively. Genes that encode putative proteins and proteins with unknown functions are indicated by grey arrows.
3.6. Gene environment characteristics of IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid
We detected a 202 kb IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in KM166 Plasmid 2 (Figure 6A), which was only previously detected in E. coli strains from Italy and United States. A study had detected IncFII/IncFIA/IncFIB multiple replicon plasmid in 24 E. coli strains, the most complete sequence of which was pMRY09-581ECO_1 (accession no. AP018456), the genomic environment comparison between pMRY09-581ECO_1 and KM166 Plasmid 2 showed that the IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in KM166 was larger than pMRY09-581ECO_1, little similarities of genomic environment between them were found and many resistance genes were only detected in pMRY09-581ECO_1 (Figure 6B). Comparison of plasmid sequence with GenBank data using BLAST3 revealed a remarkable homology (12% coverage and 99.99% identity) to K. michiganensis strain fxq plasmid pB_KPC (GenBank accession no. NZ_CP095001) isolated from Zhengzhou, China in 2022 and K. michiganensis strain K516 plasmid pK516_KPC (15% coverage and 99.99% identity) isolated from Zhejiang, China in 2017 (GenBank accession no. NZ_CP022349). The presence of many mobile genetic elements in hha-ymoA-IS110-Tn3-pinR-Tn3 structure region of KM166 Plasmid 2 was revealed by genetic environment analysis (Figure 6C). The IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in KM166 has low homology to pMRY09-581ECO_1 and the highest homology similarity to the INcFIA/INcFII(p14)-type plasmid in K. michiganensis strain fxq plasmid pB_KPC, suggesting that this multiple replicon plasmid was unlikely to have been transmitted from E. coli and probably a transfer of repFIB replicon genes from other K. michiganensis strains into the INcFIA/INcFII(p14)-type plasmid of KM166 had occurred, resulting in a strain with multiple plasmid types co-existing.
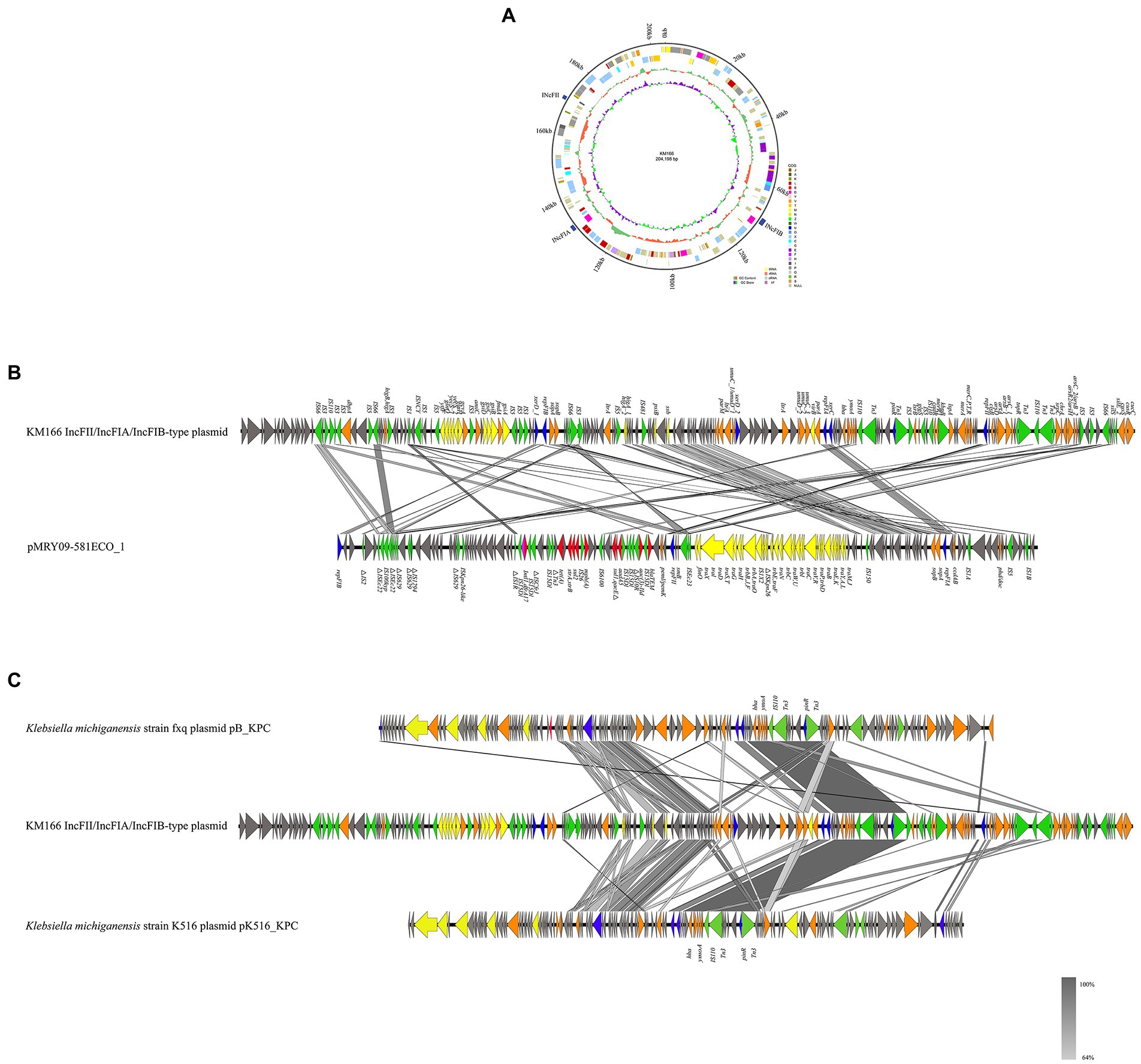
Figure 6. Mapping of the gene environment of a IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in Klebsiella michiganensis KM166. (A) Genome circle map of the KM166 IncFII/IncFIA(HI1)/IncFIB(K) plasmid. (B) Comparison of the genetic environment between KM166 IncFII/IncFIA(HI1)/IncFIB(K) plasmid and pMRY09-581ECO_1. (C) comparison of KM166 IncFII/IncFIA(HI1)/IncFIB(K) plasmid with the two highest homology genome sequences from NCBI. Genes involved in coupling transfer are indicated by yellow arrows, genes associated with plasmid stability and replication are highlighted in orange and blue arrows, respectively. Red and green arrows indicate antimicrobial resistance genes and mobile element genes, respectively.
4. Discussion
It is an important clinical case that a blaKPC-2-carrying IncFII(p14)/IncFIA plasmid and an IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid were both identified in the K. michiganensis. In terms of clinical detection and current research condition, K. michiganensis is not as mature as K. pneumoniae, but several studies (Zheng et al., 2018; Li et al., 2021) have shown that K. michiganensis strains already carry multiple drug resistance genes, especially carbapenem resistance genes. K. michiganensis has been shown to be highly resistant and multidrug-resistant like K. pneumoniae (Chapman et al., 2020; Campos-Madueno et al., 2021).
There are 445 K. michiganensis strains in the public database, mainly from Switzerland and China. Despite the fact that the patient had not traveled abroad, it is reasonable to suspect that KM166 was transmitted from other patients with a history of international travel within the same hospital. Phylogenetic tree showed that all strains were located in one evolutionary branch, indicating that K. michiganensis strains have been relatively conservative in the evolution and have not undergone much evolutionary variation. The fact that K. michiganensis strains carry multiple drug-resistant genes serves as a reminder to us to strictly control antibiotic use in order to prevent and control healthcare associated infections.
A blaKPC-2-carrying IncFII(p14)/IncFIA plasmid carried on the K. michiganensis strain was rarely reported around the world. There was a 117.8-kb IncFIA-IncFII plasmid carrying the β-lactamase-encoding gene blaOXA-926 identified in a carbapenem-resistant K. pneumoniae strain isolated from China in 2021, which could be transferred with the facilitation of the blaKPC-2-carrying plasmid (Liu, S. et al., 2021). The IncFII(p14)/IncFIA plasmid identified in our study carried blaKPC-2 and possessed the complete conjugative regions of the self-transmissible mobile genetic elements (oriT, T4CP, T4SS), which can transfer autonomously, and the resistance genes in this plasmid could transfer with the help of mobile genetic elements around. The detection rate of carbapenem-resistant K. pneumoniae carrying blaKPC-2 in blood samples between 2017 and 2020 in hospitals where KM166 was isolated was 40.67% (146/359), and we therefore suspect that the blaKPC-2-carrying IncFII(p14)/IncFIA plasmid carried by KM166 may have been transferred from K. pneumoniae carrying blaKPC-2. Besides, it is important to recognize the significance of the K. michiganensis strain we isolated from a patient’s blood sample in a Beijing hospital with a total of 38 virulence genes and 41 resistance genes.
The IncFII/IncFIA(HI1)/IncFIB(K)-type plasmid has rarely been reported globally. Two blaNDM-5-carrying plasmids with IncFIA/IncFIB/IncFII/IncQ1 replicons have been found in previous studies (Hirabayashi et al., 2021; Mohamed et al., 2022). Meanwhile, some researchers discovered multiple resistance genes in an IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in E. coli (Hayashi et al., 2020; Marchetti et al., 2020; Simner et al., 2022). We discovered an IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid in K. michiganensis strain KM166, implying that plasmid heterozygous transfer may be an evolutionary trend through genetic rearrangement via sequence insertion and homologous recombination. The IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid did not contain any factors associated with autonomous transfer, nor was a resistance gene discovered, indicating that the multiple replicon plasmid we identified is still in a primitive state.
Some limitations were found in this study. Firstly, of the 445 available K. michiganensis strains, a relatively low percentage of isolates were from China, and the most recent source in the evolutionary analysis was a foreign strain. Expanding the number of strains collected may discover a local Chinese near-source strain. Secondly, we verified that the plasmid could be transferred from K. michiganensis to E. coli, but further validation is needed to see if the plasmid is indeed transferred from K. pneumoniae to K. michiganensis. Only 22 of the 446 K. michiganensis strains collected carried the blaKPC-2 gene, indicating that the blaKPC-2 carriage rate was low. The acquisition of blaKPC-2 by K. michiganensis strain from other K. pneumoniae strains may influenced by the characteristics of the strain itself and the specific environment.
In conclusion, this study described a clinical K. michiganensis strain carrying multiple resistance genes. The IncFII/IncFIA(HI1)/IncFIB(K) multiple replicon plasmid and blaKPC-2-carrying IncFII(p14)/IncFIA plasmid were both found in a K. michiganensis strain. There was probably a transfer of repFIB replicon genes from other K. michiganensis strains into the INcFIA-INcFII(p14)-type plasmid of KM166 that resulted in the occurrence of the multiple replicon IncFII/IncFIA(HI1)/IncFIB(K) plasmid. Meanwhile, the genetic environment of blaKPC-2 in IncFII(p14)/IncFIA plasmid was same as K. pneumoniae strain JKP55, plasmid pKPC-J5501, suggesting that the blaKPC-2 might have been transferred from K. pneumoniae strains under the help of mobile genetic elements. It provides a reference for understanding the clinical infection and evolutionary characteristics of K. michiganensis.
Data availability statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found at: https://www.ncbi.nlm.nih.gov/genbank/, GCA_025263805.1.
Ethics statement
The study was authorized by The Seven Medical Center of PLA General Hospital’s institutional ethical boards in Beijing, China (2021-82). The institutional ethics committee waived the need for informed consent from the participants because all data were gathered and analyzed anonymously.
Author contributions
NZ and YC: conceptualization. NZ, LQ, XY, MJ, JC, SQ, and FL: data curation. NZ: formal analysis and visualization. YC, XL, and LQ: funding acquisition. NZ and JG: methodology. YC: project administration. NZ and XL: software. CW and YC: supervision. NZ, XL, and LQ: writing—original draft. NZ, XL, and YC: writing—review and editing. All authors contributed to the article and approved the submitted version.
Funding
The work was supported by a grant from National Key Program for Infectious Diseases of China (2018ZX10733-402), Beijing Nova Program (Z181100006218107), National Natural Science Foundation of China (31900151), National Natural Science Foundation of China (81902112).
Acknowledgments
We are grateful to the Pathosystems Resource Integration Center (PATRIC) (http://www.patricbrc.org) for providing the genomic data of Klebsiella michiganensis strains.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2022.1086296/full#supplementary-material
Footnotes
References
Abed, J. Y., Déraspe, M., Bérubé, È., D'Iorio, M., Dewar, K., Boissinot, M., et al. (2021). Complete genome sequences of Klebsiella michiganensis and Citrobacter farmeri, KPC-2-producers serially isolated from a single patient. Antibiotics (Basel). 10:1408. doi: 10.3390/antibiotics10111408
Alcock, B. P., Raphenya, A. R., Lau, T. T. Y., Tsang, K. K., Bouchard, M., Edalatmand, A., et al. (2020). CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 48, D517–D525. doi: 10.1093/nar/gkz935
Cai, J. C., Zhou, H. W., Zhang, R., and Chen, G. X. (2008). Emergence of Serratia marcescens, Klebsiella pneumoniae, and Escherichia coli isolates possessing the plasmid-mediated carbapenem-hydrolyzing beta-lactamase KPC-2 in intensive care units of a Chinese hospital. Antimicrob. Agents Chemother. 52, 2014–2018. doi: 10.1128/AAC.01539-07
Campos-Madueno, E. I., Sigrist, T., Flückiger, U. M., Risch, L., Bodmer, T., and Endimiani, A. (2021). First report of a Bla(VIM-1) metallo-β-lactamase-possessing Klebsiella michiganensis. J Glob Antimicrob Resist. 25, 310–314. doi: 10.1016/j.jgar.2021.03.027
Catalán-Nájera, J. C., Garza-Ramos, U., and Barrios-Camacho, H. (2017). Hypervirulence and hypermucoviscosity: two different but complementary Klebsiella spp. phenotypes? Virulence 8, 1111–1123. doi: 10.1080/21505594.2017.1317412
Ccorahua-Santo, R., Cervantes, M., Duran, Y., Aguirre, M., Marin, C., and Ramírez, P. (2017). Draft genome sequence of Klebsiella michiganensis 3T412C, harboring an arsenic resistance Genomic Island, isolated from mine tailings in Peru. Genome Announc. 5:0611. doi: 10.1128/genomeA.00611-17
Chan, P. A.-O., Lin, B. Y., Mak, A. J., and Lowe, T. A.-O. (2021). tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 49, 9077–9096. doi: 10.1093/nar/gkab688
Chapman, P., Forde, B. M., Roberts, L. W., Bergh, H., Vesey, D., Jennison, A. V., et al. (2020). Genomic investigation reveals contaminated detergent as the source of an extended-Spectrum-β-lactamase-producing Klebsiella michiganensis outbreak in a neonatal unit. J. Clin. Microbiol. 58:980. doi: 10.1128/jcm.01980-19
Darriba, D., Posada, D., Kozlov, A. M., Stamatakis, A., Morel, B., and Flouri, T. (2020). ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 37, 291–294. doi: 10.1093/molbev/msz189
Didelot, X., and Wilson, D. J. (2015). ClonalFrameML: efficient inference of recombination in whole bacterial genomes. PLoS Comput. Biol. 11:e1004041. doi: 10.1371/journal.pcbi.1004041
Doi, Y., and Paterson, D. L. (2015). Carbapenemase-producing Enterobacteriaceae. Semin. Respir. Crit. Care Med. 36, 74–84. doi: 10.1055/s-0035-1544208
Flouri, T., Izquierdo-Carrasco, F., Darriba, D., Aberer, A. J., Nguyen, L. T., Minh, B. Q., et al. (2015). The phylogenetic likelihood library. Syst. Biol. 64, 356–362. doi: 10.1093/sysbio/syu084
Founou, R. C., Founou, L. L., Allam, M., Ismail, A., and Essack, S. Y. (2018). Genomic characterisation of Klebsiella michiganensis co-producing OXA-181 and NDM-1 carbapenemases isolated from a cancer patient in uMgungundlovu district, KwaZulu-Natal Province. South Africa. S Afr Med J. 109, 7–8. doi: 10.7196/SAMJ.2018.v109i1.13696
Gu, D., Dong, N., Zheng, Z., Lin, D., Huang, M., Wang, L., et al. (2018). A fatal outbreak of ST11 carbapenem-resistant hypervirulent Klebsiella pneumoniae in a Chinese hospital: a molecular epidemiological study. Lancet Infect. Dis. 18, 37–46. doi: 10.1016/S1473-3099(17)30489-9
Hayashi, M., Matsui, M., Sekizuka, T., Shima, A., Segawa, T., Kuroda, M., et al. (2020). Dissemination of IncF group F1:A2:B20 plasmid-harbouring multidrug-resistant Escherichia coli ST131 before the acquisition of blaCTX-M in Japan. J Glob Antimicrob Resist. 23, 456–465. doi: 10.1016/j.jgar.2020.10.021
Hirabayashi, A., Yanagisawa, H., Takahashi, H., Yahara, K., Boeing, P., Wolfenden, B., et al. (2021). On-site genomic epidemiological analysis of antimicrobial-resistant bacteria in Cambodia with portable laboratory equipment. Front. Microbiol. 12:675463. doi: 10.3389/fmicb.2021.675463
Iovleva, A., and Doi, Y. (2017). Carbapenem-Resistant Enterobacteriaceae. Clin. Lab. Med. 37, 303–315. doi: 10.1016/j.cll.2017.01.005
Kang, M., Chmara, J., Duceppe, M. O., Phipps-Todd, B., and Huang, H. (2020). Complete genome sequence of a Canadian Klebsiella michiganensis strain, obtained using Oxford Nanopore technologies sequencing. Microbiol Resour Announc. 9:960. doi: 10.1128/mra.00960-20
King, T. L., Schmidt, S., Thakur, S., Fedorka-Cray, P., Keelara, S., Harden, L., et al. (2021). Resistome of a carbapenemase-producing novel ST232 Klebsiella michiganensis isolate from urban hospital effluent in South Africa. J Glob Antimicrob Resist. 24, 321–324. doi: 10.1016/j.jgar.2021.01.004
Koren, S., Walenz, B. P., Berlin, K., Miller, J. R., Bergman, N. H., and Phillippy, A. M. (2017). Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736. doi: 10.1101/gr.215087.116
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B., and Stamatakis, A. (2019). RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455. doi: 10.1093/bioinformatics/btz305
Lam, M., Wick, R. R., Watts, S. C., Cerdeira, L. T., Wyres, K. L., and Holt, K. E. (2021). A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 12:4188. doi: 10.1038/s41467-021-24448-3
Letunic, I., and Bork, P. (2021). Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296. doi: 10.1093/nar/gkab301
Li, X., He, J., Jiang, Y., Peng, M., Yu, Y., and Fu, Y. (2021). Genetic characterization and passage instability of a hybrid plasmid co-harboring Bla(IMP-4) and Bla(NDM-1) reveal the contribution of insertion sequences during plasmid formation and evolution. Microbiol Spectr 9:e0157721. doi: 10.1128/Spectrum.01577-21
Liu, L., Feng, Y., Wei, L., Xiao, Y., and Zong, Z. (2021). KPC-2-producing Carbapenem-resistant Klebsiella pneumoniae of the uncommon ST29 type carrying OXA-926, a novel narrow-Spectrum OXA β-lactamase. Front. Microbiol. 12:701513. doi: 10.3389/fmicb.2021.701513
Liu, S., Liu, H., Chen, R., Ma, Y., Yang, B., Chen, Z., et al. (2021). Role of two plant growth-promoting bacteria in remediating cadmium-contaminated soil combined with Miscanthus floridulus (lab.). Plants (Basel). 10:912. doi: 10.3390/plants10050912
Liu, Y., Long, D., Xiang, T. X., Du, F. L., Wei, D. D., Wan, L. G., et al. (2019). Whole genome assembly and functional portrait of hypervirulent extensively drug-resistant NDM-1 and KPC-2 co-producing Klebsiella pneumoniae of capsular serotype K2 and ST86. J. Antimicrob. Chemother. 74, 1233–1240. doi: 10.1093/jac/dkz023
Liu, B., Zheng, D., Jin, Q., Chen, L., and Yang, J. (2019). VFDB 2019: a comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res. 47, D687–D692. doi: 10.1093/nar/gky1080
Luo, X., Zhang, J., Yuan, M., Mou, S., Xu, M., Hu, D., et al. (2022). Epidemiology of Klebsiella michiganensis carrying multidrug-resistant IncHI5 plasmids in the southeast coastal area of China. Infect Drug Resist. 15, 1831–1843. doi: 10.2147/idr.S358839
Marchetti, V. M., Bitar, I., Mercato, A., Nucleo, E., Bonomini, A., Pedroni, P., et al. (2020). Complete nucleotide sequence of plasmids of two Escherichia coli strains carrying blaNDM-5 and blaNDM-5 and blaOXA-181 from the same patient. Front. Microbiol. 10:3095. doi: 10.3389/fmicb.2019.03095
Mitra, S., Pramanik, K., Ghosh, P. K., Soren, T., Sarkar, A., Dey, R. S., et al. (2018). Characterization of cd-resistant Klebsiella michiganensis MCC3089 and its potential for rice seedling growth promotion under cd stress. Microbiol. Res. 210, 12–25. doi: 10.1016/j.micres.2018.03.003
Mitra, S., Purkait, T., Pramanik, K., Maiti, T. K., and Dey, R. S. (2019). Three-dimensional graphene for electrochemical detection of cadmium in Klebsiella michiganensis to study the influence of cadmium uptake in rice plant. Mater. Sci. Eng. C Mater. Biol. Appl. 103:109802. doi: 10.1016/j.msec.2019.109802
Mohamed, N. M., Zakaria, A. S., and Edward, E. A. (2022). Genomic characterization of international high-risk clone ST410 Escherichia coli co-harboring ESBL-encoding genes and blaNDM-5 on IncFIA/IncFIB/IncFII/IncQ1 multireplicon plasmid and carrying a chromosome-borne blaCMY-2 from Egypt. Antibiotics (Basel). 11:1031. doi: 10.3390/antibiotics11081031
Nordmann, P., Naas, T., and Poirel, L. (2011). Global spread of Carbapenemase-producing Enterobacteriaceae. Emerg. Infect. Dis. 17, 1791–1798. doi: 10.3201/eid1710.110655
Oinuma, K. I., Suzuki, M. A.-O., Sakiyama, A., Tsubouchi, T., Saeki, K., Sato, K., et al. (2021). Genomic characterization of triple-carbapenemase-producing Acinetobacter baumannii. JAC Antimicrob Resist. 3:dlab191. doi: 10.1093/jacamr/dlab191
Oliveira, R. A., Ng, K. M., Correia, M. B., Cabral, V., Shi, H., Sonnenburg, J. L., et al. (2020). Klebsiella michiganensis transmission enhances resistance to Enterobacteriaceae gut invasion by nutrition competition. Nat. Microbiol. 5, 630–641. doi: 10.1038/s41564-019-0658-4
Pitout, J. D., Nordmann, P., and Poirel, L. (2015). Carbapenemase-producing Klebsiella pneumoniae, a key pathogen set for global nosocomial dominance. Antimicrob. Agents Chemother. 59, 5873–5884. doi: 10.1128/AAC.01019-15
Salgueiro, V., Manageiro, V., Bandarra, N. M., Reis, L., Ferreira, E., and Caniça, M. (2020). Bacterial diversity and antibiotic susceptibility of Sparus aurata from aquaculture. Microorganisms. 8:343. doi: 10.3390/microorganisms8091343
Seemann, T. (2014). Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069. doi: 10.1093/bioinformatics/btu153
Seiffert, S. N., Wüthrich, D., Gerth, Y., Egli, A., Kohler, P., and Nolte, O. (2019). First clinical case of KPC-3-producing Klebsiella michiganensis in Europe. New Microbes New Infect. 29:100516. doi: 10.1016/j.nmni.2019.100516
Simner, P. J., Mostafa, H. H., Bergman, Y., Ante, M., Tekle, T., Adebayo, A., et al. (2022). Progressive development of Cefiderocol resistance in Escherichia coli during therapy is associated with an increase in blaNDM-5 copy number and gene expression. Clin. Infect. Dis. 75, 47–54. doi: 10.1093/cid/ciab888
Siu, L. K., Fung, C. P., Chang, F. Y., Lee, N., Yeh, K. M., Koh, T. H., et al. (2011). Molecular typing and virulence analysis of serotype K1 Klebsiella pneumoniae strains isolated from liver abscess patients and stool samples from noninfectious subjects in Hong Kong, Singapore, and Taiwan. J. Clin. Microbiol. 49, 3761–3765. doi: 10.1128/JCM.00977-11
Sullivan, M. J., Petty, N. K., and Beatson, S. A. (2011). Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010. doi: 10.1093/bioinformatics/btr039
Wyres, K. L., Lam, M., and Holt, K. E. (2020). Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 18, 344–359. doi: 10.1038/s41579-019-0315-1
Xu, J., Ding, H., Zhao, Y., Chen, J., Zhao, Z., Huang, J., et al. (2022). Characterization of a novel Carbapenem-resistant Klebsiella michiganensis strain Coharboring the Bla(SIM-1), Bla(OXA-1), Bla(CTX-M-14), qnrS, and aac(6′)-Ib-cr genes. Curr. Microbiol. 79:228. doi: 10.1007/s00284-022-02920-7
Yan, X., Yang, J., Wang, Q., and Lin, S. (2021). Transcriptomic analysis reveals resistance mechanisms of Klebsiella michiganensis to copper toxicity under acidic conditions. Ecotoxicol. Environ. Saf. 211:111919. doi: 10.1016/j.ecoenv.2021.111919
Zhang, Y., Gu, D., Yang, X., Wu, Y., Liu, C., Shen, Z., et al. (2021). Emergence and genomic characterization of a KPC-2-, NDM-1-, and IMP-4-producing Klebsiella michiganensis isolate. Front. Microbiol. 12:762509. doi: 10.3389/fmicb.2021.762509
Keywords: Klebsiella michiganensis, genome analysis, evolution, plasmid, carbapenemase
Citation: Zhang N, Liu X, Qi L, Chen J, Qin S, Jin M, Yang X, Liu F, Guo J, Liu J, Wang C and Chen Y (2023) A clinical KPC-producing Klebsiella michiganensis strain carrying IncFII/IncFIA (HI1)/IncFIB (K) multiple replicon plasmid. Front. Microbiol. 13:1086296. doi: 10.3389/fmicb.2022.1086296
Edited by:
Swapnil Prakash Doijad, University Hospital Giessen and Marburg, GermanyReviewed by:
Jess Vergis, Kerala Veterinary and Animal Sciences University, IndiaMangesh Vasant Suryavanshi, Cleveland Clinic, United States
Copyright © 2023 Zhang, Liu, Qi, Chen, Qin, Jin, Yang, Liu, Guo, Liu, Wang and Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yong Chen, ✉ Y2hlbnlvbmdlckAxMjYuY29t; Changjun Wang, ✉ c2NpZW5jZTIwMDhAaG90bWFpbC5jb20=; Jie Liu, ✉ ZmhhYWxqQDE2My5jb20=
†These authors have contributed equally to this work and share first authorship
 Na Zhang1,2†
Na Zhang1,2† Xiong Liu
Xiong Liu Jiali Chen
Jiali Chen Shiyu Qin
Shiyu Qin Fangni Liu
Fangni Liu Changjun Wang
Changjun Wang Yong Chen
Yong Chen