- 1Faculty of Health and Medical Sciences, Department of Drug Design and Pharmacology, University of Copenhagen, Copenhagen, Denmark
- 2Danish Cancer Society Research Center, Danish Cancer Society, Copenhagen, Denmark
Prostate cancer (PCa) is one of the most frequently diagnosed cancer among men in the western societies. Many PCa patients bear tumors that will not threat their lives if left untreated or if treatment is delayed. Our inability for early identification of these patients has resulted in massive overtreatment. Therefore, there is a great need of finding biomarkers for patient stratification according to prognostic risk; as well as there is a need for novel targets that can allow the development of effective treatments for patients that progress to castration-resistant PCa. Most biomarkers in cancer are proteins, including the widely-used prostate-specific antigen (PSA). Recent developments in mass spectrometry allow the identification and quantification of thousands of proteins and posttranslational modifications from small amounts of biological material, including formalin-fixed paraffin-embedded tissues, and biological fluids. Novel diagnostic and prognostic biomarkers have been identified in tissue, blood, urine, and seminal plasma of PCa patients, and new insights in the ethology and progression of this disease have been achieved using this technology. In this review, we summarize these findings and discuss the potential of this technology to pave the way toward the clinical implementation of precision medicine in PCa.
Introduction
Prostate cancer (PCa) leads the statistics in cancer diagnosis and cancer-related death among men in the western societies (1). Current therapies, which are based on radiation or surgery for prostate-confined tumors and on androgen-deprivation therapy (ADT) for locally advanced or metastatic presentations have demonstrated to be very effective in the management of the disease. Unfortunately, the positive effects of these therapies are only temporary and most patients relapse after ADT into the so-called castration-resistant prostate cancer (CRPC), against which no curative therapy exists (2).
Overtreatment is a major concern in the management of PCa. Due to the slow progressing nature of PCa and the advanced age of the patient population, it is estimated that many of the surgically treated patients would not die of the disease or experience major morbidities if therapeutic action was not taken (3). However, our inability to distinguish between indolent PCa tumors and those which are life threatening has led to the problem of overtreatment (3). Thus, two current major areas of research in the PCa field are (i) the identification of biomarkers that can accurately predict the virulence of a prostate localized tumors and (ii) the development of effective treatments against lethal CRPC. It has now become clear that the application of state-of-the-art mass spectrometry-based proteomics to PCa research can contribute to address these and other questions related the clinical management of PCa (Table 1).
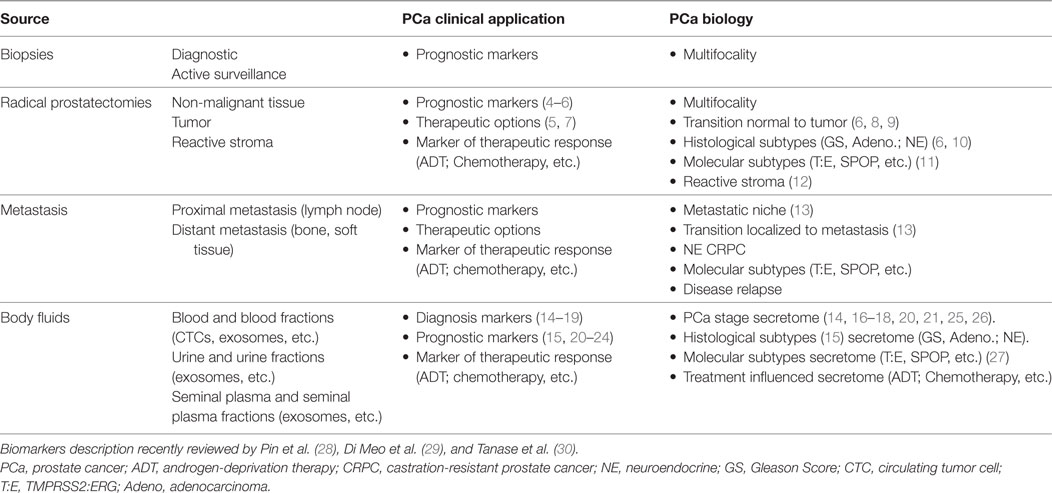
Table 1. Open questions in PCa research susceptible to be addressed by applying mass spectrometry-based proteomics to a range of biological samples.
Proteins are the effectors of most cellular reactions and constitute the cellular targets for a majority of therapeutic drugs. Over the years, many protein biomarkers and potential drug targets have been identified using mass spectrometry techniques [recently reviewed in Ref (28–30).]. This pioneer studies used proteomic methodologies with limited capacity to achieve great depth and unfortunately most of these markers have not been further validated or implemented in the clinical setting. In-depth quantitative proteomic profiling, long-time hampered by the complexity of the proteome and the lack of appropriate technology, is now possible from both frozen and formalin-fixed-paraffin-embedded (FFPE) tissues as well as from body fluids (Figure 1) (6, 31–36). The use of mass spectrometry-based proteomic profiling to well-characterized cohorts of PCa patients with detailed clinical information have the potential to provide effective stratification of PCa patients as well as targets for novel treatments (37). In this review, we aim to summarize the achievements regarding the use of mass spectrometry-based proteomics for large-scale profiling of PCa and discuss the potential use of this technology for the identification of diagnostic and prognostic biomarkers, as well as therapeutic targets.
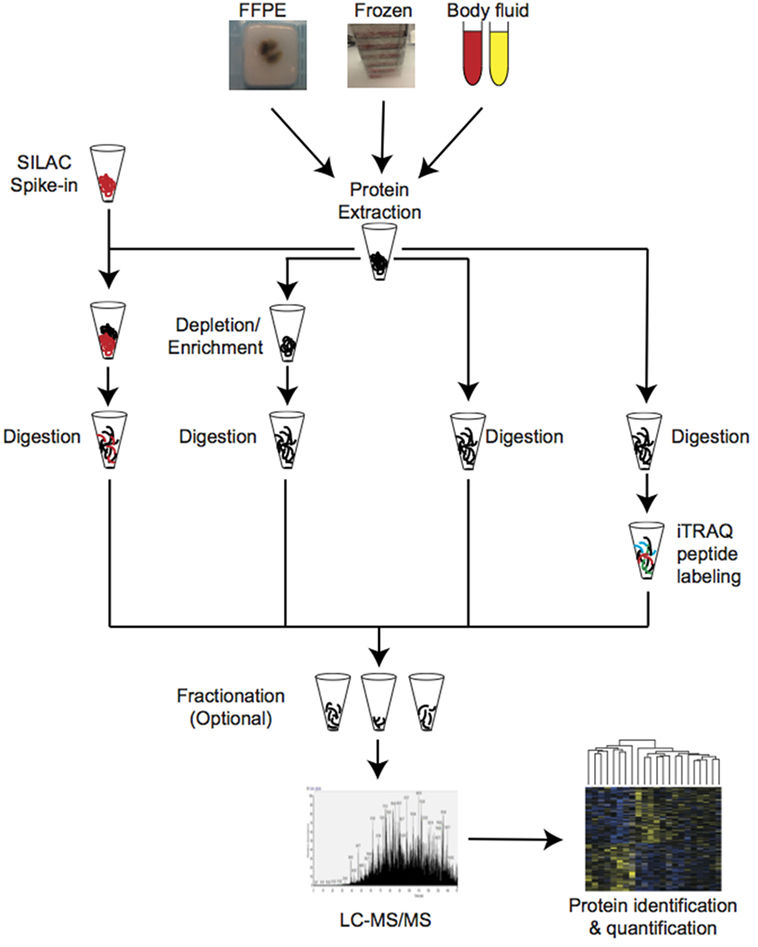
Figure 1. Schematic representation of the commonly used approaches for mass spectrometry-based proteomic profiling in clinical cancer research.
Genomic Characterization Revealed Novel Molecular Subtypes in PCa
In recent years, several large-scale genomic studies have been performed with the aim of defining molecular subtypes in PCa that could possibly improve or complement the histopathological classification currently used in medical practice (38–45). The major genomic alterations occurring in PCa, including TMPRSS2:ERG translocations and mutations in PTEN, P53, CHD1, and SPOP, have been defined (42, 43, 46). Moreover, gene expression profiles associated to these genomic subtypes have advanced our knowledge of the molecular mechanisms driving these tumors, but our understanding of PCa progression remains limited. Although both microarray and RNA-sequencing techniques provide high degree of information with broad coverage, the correlation between mRNA and protein expression is only partial (47–49), therefore our understanding of PCa proteome remains incomplete.
Proteomic Profiling of PCa Tissues
The utilization of several proteomic techniques in PCa profiling has been recently reviewed (29, 30). Bottom-up approaches, that is, protein identification from the analysis of their peptide components, has so far resulted in better proteome coverage. Tryptic digestion followed by peptide fractionation, peptide separation through liquid chromatography, and on-line electrospray ionization (50) coupled to a MS/MS orbitrap-based analyzers (51) is the most extended choice for in-depth proteomic analysis1 (52) (Figure 1). Improvements in sample preparation, the sensitivity of mass analyzers and computational developments now allows to quantify what is considered to be close to a full proteome (over 10,000 proteins expressed per cell) (48, 53, 54). This depth of coverage is state-of-the-art today but will become standard in the near future. Alternative approaches such as the SWATH-MS methodology developed by Aebersold’s laboratory, based on data-independent acquisition, can provide large coverage with high degree of reproducibility (55). However, this methodology has rarely been applied to PCa research so far (4).
Of all the proteomic profiles performed on PCa tumors, two studies stand out for the extensive coverage achieved (6, 13). Drake et al. reported the phosphoproteomic profile of 16 metastatic castration-resistant samples obtained from 13 patients, together with 6 prostate localized tumors and 5 benign prostate tissue obtained from autopsy programs (13). They identify and quantify over 8,000 phospho-peptides using label-free mass spectrometry. Integration of these data with other genome-wide approaches allowed them to identify kinase-regulated signaling pathways activated in castration-resistant PCa compared to untreated tumors including DNA repair, PI3K–AKT–MTOR, and cell cycle progression related processes. The use of drugs targeting some of the kinases involved in the pathways that are active in metastatic PCa are currently being tested in clinical trials2 (clinical trial IDs: NCT02407054 and NCT02091531).
While Drake and coworkers used frozen sample for their studies, mass spectrometry quality protein extracts can also be obtained from formalin-fixed paraffin-embedded (FFPE) tissues (6, 31–33). Due to the reduce maintenance cost and long-lasting preservation, FFPE tissue archives are the preferred option for long-term tumor storage. The use of FFPE samples opens the possibility of retrospective proteomic analysis of samples from clinical trials where long-term clinical follow-up of patients is available. This is especially important when investigating diseases such as PCa, with a disease course that sometimes expands over 25 years.
Recently, we analyzed the proteome of 36 samples obtained from FFPE preserved radical prostatectomy specimens. Twenty-eight corresponded to tumors with different histological patterns (Gleason score) and eight to neighboring non-malignant tissue (6). An average of 5,580 (±515) proteins were identified for a total sum of over 9,000. Accurate relative quantification was achieved by the use of extracts from SILAC (Stable-Isotope Labeling by Amino acids in Cell culture)-labeled prostate cell lines as spike-in standard for each sample (56). In addition to earlier described changes in the early progression of prostate tumors (increased expression of androgen receptor regulated proteins, reduced levels of proteins involved in cell adhesion, etc.), we showed that prostate localized tumors exhibit increased expression of mitochondrial proteins, which come accompanied by increased mitochondrial activity. This seems to be a particularity of prostate tumors compared to other types of cancer that commonly depend on glycolytic pathways for energy production. The targeting of mitochondrial function with agents like Metformin or Phenformin in combined regimens with Enzalutamide (AR inhibitor) is currently being tested in clinical trials for the treatment of metastatic PCa [(57), see text footnote 2 for clinical trial IDs: NCT01620593; NCT02339168; NCT02176161; NCT02640534; NCT00881725; NCT01864096, among others]. As part of this study, by comparing PCa tumors of different Gleason grades, we identified pro-NPY as a novel prognostic biomarker in early PCa and verify its performance in two independent large cohorts of minimally treated PCa patients. The potential role of pro-NPY as an early prognostic biomarker must now be further validated using a modern cohort of PCa patients, preferable those enrolled in active surveillance programs.
These studies constitute a great leap forward toward the complete characterization of PCa proteome and have unraveled the most general proteomic features that characterize PCa progression. However, they all show a similar limitation, which concerns to the relatively small number of samples analyzed in each study. In order to validate the utility of proteomic profiling toward personalized treatment of PCa, more tissue samples need to be analyzed, including different genetic and histological subtypes, tumors subjected to different treatment options, and tumors growing in different metastatic niches (Table 1). These questions are only starting to be investigated. For instance, proteome differences between tumors bearing or not the TMPRSS2:ERG translocation have been studied (11). Despite the small number of clinical samples analyzed and limited proteome coverage, some proteins like MYO6 were found as differentially regulated between ERG+ and ERG− tumors (11). Another example of initial proteomic study tackling a relevant PCa question is the analysis of epithelial and microenvironment tissue that were micro-dissected from tumors graded with different Gleason scores (12). Despite similar limitations as the previously described study, this investigation served as a proof of principle for the existence of major differences in tumor microenvironment. The composition of the tumor microenvironment is critical in the development of PCa (58–60). Therefore, deep proteomic characterization of this compartment is warranted in order to understand its influence in PCa progression. Many aspects of PCa tissue proteome dynamics remain to be explored. The analysis of the proteome changes occurring during metastatic dissemination of PCa (primarily to bone), in combination with the effects of androgen deprivation and chemo-therapies, including neuroendocrine differentiation, will potentially contribute to identify targets for novel therapeutic approaches and contribute to patient stratification toward personalized treatments (Table 1).
Quantitative Proteomic Profile of Biofluids from PCa Patients
Blood and urine are ideal sources for biomarker identification due to the minimally invasive procedures of acquisition and the plentiful availability. In prostatic diseases, direct evidence of prostatic alteration can be obtained from the analysis of the seminal plasma. All these fluids are composed by less complex mix of proteins compared to tissues (61–63) but present other challenges, primarily: the dynamic range of the proteome; the variation in composition related to storage time; the intra- and inter patient variability and the relationship between the fluid components and the tumor.
In blood, the dynamic range, that is, the difference in concentration among the different proteins, spans for over 10 orders of magnitude (64). Highly abundant proteins such as albumins and globulins account from most of the serum protein content (64). Depletion of these proteins to enrich for low abundant ones is a common pre-requisite during sample preparation (35) although novel methodologies have been recently proposed to identify and quantify hundreds of plasma proteins using direct measurements (34).
Measurement of Prostate-Specific Antigen (PSA) blood levels is a routine tool for PCa diagnosis and screening (65). However, PSA levels are not informative of cancer aggressiveness, leading to overdiagnose of PCa patients with asymptomatic disease (66) and cannot discriminate between PCa and other prostate pathologies (67). Finding serum biomarkers that precisely identify PCa and show correlation with tumor aggressiveness have been investigated using mass-spectrometry-based approaches (30). Additionally, identification of prognostic biomarkers or indicators of treatment response has also been tested proteomic profiling of PCa patients’ blood (29, 30). From those studies, the identification of the pigment epithelium-derived factor (PEDF) stands out as common serum marker with altered expression during PCa progression (19–21). PEDF seems to promote macrophage recruitment to the tumors and to inhibit the expression of metalloproteases involved in PCa cells invasion (68–70), supporting a role as tumors suppressor in PCa. Therefore, the potential of PEDF as diagnostic and prognostic biomarker for PCa should be validated in larger cohorts.
Other approaches aiming at evaluating tumor characteristics through non-invasive protocols is to purify tumor-derived components from blood. This is the case of exosomes and circulating tumor cells (CTCs). Exosomes are small vesicles of 50–100 nm in diameter secreted from most tissues including the prostate (71–73). Proteomic characterization of these microvesicles has the advantage of being directly produced from the prostatic tissue both normal and cancerous, and being composed by a relatively low complex proteome, which make them potentially interesting as a source of blood biomarkers. However, isolation and enrichment of exosomes is a time-consuming process that requires larger amounts of initial material due to the relative low number of these vesicles presents in the blood (27, 74). Moreover, most methods applied today for exosome isolation from blood are based on centrifugation protocols with limited specificity for PCa cell derived vesicles (27). This limited specificity can be ignored when analyzing genetic changes specifics to PCa such as the presence of AR receptor variants, whose detection on exosomes from PCa patient’s blood can predict resistance to hormonal therapy (27). However, for proteomic profiling more specific methodologies of PCa cells derived exosome isolation would be required.
Circulating tumor cells analysis constitutes a direct source of information regarding tumor genetics. Proteomic characterization of CTCs could additionally reveal signaling pathways activated in the tumors and be used to direct personalized treatments. This is particularly relevant during the metastatic onset of the disease due to the difficulties and ethical concerns related to the collection of metastatic biopsies. However, proteomic profiling of CTCs remains extraordinarily challenging due to the limited amount available per sample. Development of single-cell mass spectrometry protocols (75) would be critical for the future proteome profile of this type of clinical material.
Urine is a plentiful source of biomarkers. In prostatic diseases, direct evidence of the prostate secretome can be obtained from urine samples, especially when prostatic massage is performed before the sample collection. Thus, clinical tests based on the measurement of the non-coding RNA PCA3 levels in urine, have been approved by the FDA to evaluate the necessity of prostate re-biopsy in men with previously negative biopsy (76–78). Similar to the analysis of blood proteome, challenges regarding wide dynamic range, and inter- and intra- individual heterogeneity apply to proteomic studies performed on urinary samples (36). In an attempt to identify novel diagnostic and prognostic biomarker for PCa, several mass spectrometry-based proteomic profiles have been performed either by direct measurements or after exosomes isolation [reviewed in Ref (79).]. Especially relevant is the identification of CD14 as urine marker to discriminate between benign prostatic hyperplasia (BPH) and PCa with high sensitivity and specificity (16). A similar attempt to discriminate between BPH and PCa using urine proteins found β2M, PGA3, and MUC3, whose levels, in combination with PSA concentration in blood, achieved a better predictive accuracy that PSA alone with a receiving operating characteristic (ROC), area under the curve (AUC) of 0.812 (17). Additionally, altered expression in urine of the proteins serotransferrin (TF), haptoglobin (HP), and AMBP was retrospectively found in men diagnosed or not with PCa (18), with an ROC AUC of 0.848. Finally, Li and coworkers described that lower concentration of peptides from osteopontin (SPP1) and prothrombin (F2) in PCa that in BPH (14). Unfortunately, despite being similar approaches, the results of the different studies do not overlap and, therefore, the clinical utility of any of these biomarkers would require independent replication in larger cohorts of PCa patients.
Non-invasive biomarkers of disease progression would help to monitor patients managed with active surveillance programs. These patients are periodically monitored for changes in blood PSA concentration and other signs of disease progression (80, 81). After proteomic profiling of urinary extracellular vesicles using mass spectrometry, Fujita et al. identified FABP5 protein differentially expressed among tumors with different histological features (25). All these findings require further validation but serve as a proof of principle that despite the common sample size limitation, deep proteomic profile of urinary samples has the potential to identify novel PCa diagnostic and prognostic biomarkers.
In addition to blood and urine, analysis of seminal fluid is particularly interesting to prostate diseases as it may provide direct evidence of alterations in the prostate gland, including the development of PCa. Early studies by Drake and coworkers identified 916 proteins from prostatic fluids obtained from PCa patients after prostatic massage. Identified proteins were enriched for those of prostatic origin, therefore, validating their approach (26). Then, they applied this methodology to discriminate between patients bearing organ-confined or extracapsular prostate tumors and identified a series of proteins and validated 3 proteins (MME, PARK7, and TIMP1) as differentially expressed between both conditions using a selective reaction monitoring (SRM)-MS targeted approach (23). Proteomics on seminal fluid has also been used to try to discriminate between indolent and advance PCa. Thus, Neuhaus et al. proposed a signature of peptides isolated from seminal fluid able to retrospectively identify patients with advance and localize prostate tumors histologically scored with similar Gleason grade (24). As for most of these studies, replication of the results in larger cohorts of patients is needed in order to conclude about the utility of the individual markers.
Future Perspectives
Clinical management of PCa can benefit from the fast-paced development within the mass spectrometry field (Table 1). Understanding PCa natural progression and identifying signaling pathways that turn deregulated during this evolution is critical to find novel and more personalized treatments. Proteomic characterization of metastatic PCa has the potential to increase our understanding of the lethal stage of the disease. The identification of prognostic biomarkers that can predict tumor aggressiveness or response to therapies at early stages are needed to limit the unnecessary treatment of patients bearing low-risk tumors, while offering the most adequate treatment to each patient at the earliest possible time during disease progression.
Current mass spectrometry-based proteomic techniques allow determination of numerous posttranslational modifications, including phosphorylation, glycosylation, and ubiquitination (82–84). However, application of these methodologies to clinical PCa samples is scarce. Deep profiling of these protein modifications will provide new insights regarding the cellular pathways activated during disease progression and would help to develop novel therapeutic approaches and identification of novel diagnostic and prognostic biomarker in PCa. Moreover, the use of targeted proteomic technologies, such as SRM and MRM, which can provide accurate and reproducible measurement of biomarkers of interest (85–89), may be critical for the clinical use of the identified biomarkers. An example is the development of such targeted approach to identify different isoforms of SPOP, a commonly mutated gene in PCa (90). In addition, improved protocols and technical developments are still necessary to allow for the proteomic characterization of, for instance, CTCs.
Finally, integration of mass spectrometry-based proteomic profiling with other state-of-the-art technologies, especially with next-generation sequencing, will provide specific information of the signaling pathways and processes activated in a given patient with its unique tumor genetic background, helping to elucidate the functional consequences of somatic mutations, define driver proteins, and identify therapeutic targets. Integration of these preoteogenomic approaches (91) has already been applied to other cancer types (49, 92) and will be critical to achieve full personalized management in PCa.
Author Contributions
Collection of data and drafting the manuscript: AF-M and DI-G.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
This work was supported by grants from the Danish Research Council (DFF 4004-00450), the Movember Foundation and the Danish Cancer Society (R90-A6060-14-S2) awarded to AF-M.
Footnotes
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin (2017) 67(1):7–30. doi:10.3322/caac.21387
2. Pienta KJ, Bradley D. Mechanisms underlying the development of androgen-independent prostate cancer. Clin Cancer Res (2006) 12(6):1665–71. doi:10.1158/1078-0432.CCR-06-0067
3. Schroder FH, Hugosson J, Roobol MJ, Tammela TL, Ciatto S, Nelen V, et al. Screening and prostate-cancer mortality in a randomized European study. N Engl J Med (2009) 360(13):1320–8. doi:10.1056/NEJMoa0810084
4. Liu Y, Chen J, Sethi A, Li QK, Chen L, Collins B, et al. Glycoproteomic analysis of prostate cancer tissues by SWATH mass spectrometry discovers N-acylethanolamine acid amidase and protein tyrosine kinase 7 as signatures for tumor aggressiveness. Mol Cell Proteomics (2014) 13(7):1753–68. doi:10.1074/mcp.M114.038273
5. Endoh K, Nishi M, Ishiguro H, Uemura H, Miyagi Y, Aoki I, et al. Identification of phosphorylated proteins involved in the oncogenesis of prostate cancer via Pin1-proteomic analysis. Prostate (2012) 72(6):626–37. doi:10.1002/pros.21466
6. Iglesias-Gato D, Wikstrom P, Tyanova S, Lavallee C, Thysell E, Carlsson J, et al. The proteome of primary prostate cancer. Eur Urol (2016) 69(5):942–52. doi:10.1016/j.eururo.2015.10.053
7. Ummanni R, Mundt F, Pospisil H, Venz S, Scharf C, Barett C, et al. Identification of clinically relevant protein targets in prostate cancer with 2D-DIGE coupled mass spectrometry and systems biology network platform. PLoS One (2011) 6(2):e16833. doi:10.1371/journal.pone.0016833
8. Garbis SD, Tyritzis SI, Roumeliotis T, Zerefos P, Giannopoulou EG, Vlahou A, et al. Search for potential markers for prostate cancer diagnosis, prognosis and treatment in clinical tissue specimens using amine-specific isobaric tagging (iTRAQ) with two-dimensional liquid chromatography and tandem mass spectrometry. J Proteome Res (2008) 7(8):3146–58. doi:10.1021/pr800060r
9. Han ZD, Zhang YQ, He HC, Dai QS, Qin GQ, Chen JH, et al. Identification of novel serological tumor markers for human prostate cancer using integrative transcriptome and proteome analysis. Med Oncol (2012) 29(4):2877–88. doi:10.1007/s12032-011-0149-9
10. Lexander H, Palmberg C, Hellman U, Auer G, Hellstrom M, Franzen B, et al. Correlation of protein expression, Gleason score and DNA ploidy in prostate cancer. Proteomics (2006) 6(15):4370–80. doi:10.1002/pmic.200600148
11. Tan SH, Furusato B, Fang X, He F, Mohamed AA, Griner NB, et al. Evaluation of ERG responsive proteome in prostate cancer. Prostate (2014) 74(1):70–89. doi:10.1002/pros.22731
12. Staunton L, Tonry C, Lis R, Espina V, Liotta L, Inzitari R, et al. Pathology-driven comprehensive proteomic profiling of the prostate cancer tumor microenvironment. Mol Cancer Res (2017) 15(3):281–93. doi:10.1158/1541-7786.MCR-16-0358
13. Drake JM, Paull EO, Graham NA, Lee JK, Smith BA, Titz B, et al. Phosphoproteome integration reveals patient-specific networks in prostate cancer. Cell (2016) 166(4):1041–54. doi:10.1016/j.cell.2016.07.007
14. Li C, Zang T, Wrobel K, Huang JT, Nabi G. Quantitative urinary proteomics using stable isotope labelling by peptide dimethylation in patients with prostate cancer. Anal Bioanal Chem (2015) 407(12):3393–404. doi:10.1007/s00216-015-8569-6
15. Rehman I, Evans CA, Glen A, Cross SS, Eaton CL, Down J, et al. iTRAQ identification of candidate serum biomarkers associated with metastatic progression of human prostate cancer. PLoS One (2012) 7(2):e30885. doi:10.1371/journal.pone.0030885
16. Cheng HL, Huang HJ, Ou BY, Chow NH, Chen YW, Tzai TS, et al. Urinary CD14 as a potential biomarker for benign prostatic hyperplasia – discovery by combining MALDI-TOF-based biostatistics and ESI-MS/MS-based stable-isotope labeling. Proteomics Clin Appl (2011) 5(3–4):121–32. doi:10.1002/prca.201000011
17. Jedinak A, Curatolo A, Zurakowski D, Dillon S, Bhasin MK, Libermann TA, et al. Novel non-invasive biomarkers that distinguish between benign prostate hyperplasia and prostate cancer. BMC Cancer (2015) 15:259. doi:10.1186/s12885-015-1284-z
18. Davalieva K, Kiprijanovska S, Komina S, Petrusevska G, Zografska NC, Polenakovic M. Proteomics analysis of urine reveals acute phase response proteins as candidate diagnostic biomarkers for prostate cancer. Proteome Sci (2015) 13(1):2. doi:10.1186/s12953-014-0059-9
19. Bergamini S, Bellei E, Reggiani Bonetti L, Monari E, Cuoghi A, Borelli F, et al. Inflammation: an important parameter in the search of prostate cancer biomarkers. Proteome Sci (2014) 12:32. doi:10.1186/1477-5956-12-32
20. Qingyi Z, Lin Y, Junhong W, Jian S, Weizhou H, Long M, et al. Unfavorable prognostic value of human PEDF decreased in high-grade prostatic intraepithelial neoplasia: a differential proteomics approach. Cancer Invest (2009) 27(7):794–801. doi:10.1080/07357900802175617
21. Byrne JC, Downes MR, O’Donoghue N, O’Keane C, O’Neill A, Fan Y, et al. 2D-DIGE as a strategy to identify serum markers for the progression of prostate cancer. J Proteome Res (2009) 8(2):942–57. doi:10.1021/pr800570s
22. Rosenzweig CN, Zhang Z, Sun X, Sokoll LJ, Osborne K, Partin AW, et al. Predicting prostate cancer biochemical recurrence using a panel of serum proteomic biomarkers. J Urol (2009) 181(3):1407–14. doi:10.1016/j.juro.2008.10.142
23. Kim Y, Ignatchenko V, Yao CQ, Kalatskaya I, Nyalwidhe JO, Lance RS, et al. Identification of differentially expressed proteins in direct expressed prostatic secretions of men with organ-confined versus extracapsular prostate cancer. Mol Cell Proteomics (2012) 11(12):1870–84. doi:10.1074/mcp.M112.017889
24. Neuhaus J, Schiffer E, von Wilcke P, Bauer HW, Leung H, Siwy J, et al. Seminal plasma as a source of prostate cancer peptide biomarker candidates for detection of indolent and advanced disease. PLoS One (2013) 8(6):e67514. doi:10.1371/journal.pone.0067514
25. Fujita K, Kume H, Matsuzaki K, Kawashima A, Ujike T, Nagahara A, et al. Proteomic analysis of urinary extracellular vesicles from high Gleason score prostate cancer. Sci Rep (2017) 7:42961. doi:10.1038/srep42961
26. Drake RR, Elschenbroich S, Lopez-Perez O, Kim Y, Ignatchenko V, Ignatchenko A, et al. In-depth proteomic analyses of direct expressed prostatic secretions. J Proteome Res (2010) 9(5):2109–16. doi:10.1021/pr1001498
27. Del Re M, Biasco E, Crucitta S, Derosa L, Rofi E, Orlandini C, et al. The detection of androgen receptor splice variant 7 in plasma-derived exosomal RNA strongly predicts resistance to hormonal therapy in metastatic prostate cancer patients. Eur Urol (2017) 71(4):680–7. doi:10.1016/j.eururo.2016.08.012
28. Pin E, Fredolini C, Petricoin EF III. The role of proteomics in prostate cancer research: biomarker discovery and validation. Clin Biochem (2013) 46(6):524–38. doi:10.1016/j.clinbiochem.2012.12.012
29. Di Meo A, Pasic MD, Yousef GM. Proteomics and peptidomics: moving toward precision medicine in urological malignancies. Oncotarget (2016) 7(32):52460–74. doi:10.18632/oncotarget.8931
30. Tanase CP, Codrici E, Popescu ID, Mihai S, Enciu AM, Necula LG, et al. Prostate cancer proteomics: current trends and future perspectives for biomarker discovery. Oncotarget (2017) 8(11):18497–512. doi:10.18632/oncotarget.14501
31. Ostasiewicz P, Zielinska DF, Mann M, Wisniewski JR. Proteome, phosphoproteome, and N-glycoproteome are quantitatively preserved in formalin-fixed paraffin-embedded tissue and analyzable by high-resolution mass spectrometry. J Proteome Res (2010) 9(7):3688–700. doi:10.1021/pr100234w
32. Maes E, Valkenborg D, Mertens I, Broeckx V, Baggerman G, Sagaert X, et al. Proteomic analysis of formalin-fixed paraffin-embedded colorectal cancer tissue using tandem mass tag protein labeling. Mol Biosyst (2013) 9(11):2686–95. doi:10.1039/c3mb70177h
33. Wisniewski JR, Dus K, Mann M. Proteomic workflow for analysis of archival formalin-fixed and paraffin-embedded clinical samples to a depth of 10 000 proteins. Proteomics Clin Appl (2013) 7(3–4):225–33. doi:10.1002/prca.201200046
34. Geyer PE, Kulak NA, Pichler G, Holdt LM, Teupser D, Mann M. Plasma proteome profiling to assess human health and disease. Cell Syst (2016) 2(3):185–95. doi:10.1016/j.cels.2016.02.015
35. Keshishian H, Burgess MW, Specht H, Wallace L, Clauser KR, Gillette MA, et al. Quantitative, multiplexed workflow for deep analysis of human blood plasma and biomarker discovery by mass spectrometry. Nat Protoc (2017) 12(8):1683–701. doi:10.1038/nprot.2017.054
36. Zhao M, Li M, Yang Y, Guo Z, Sun Y, Shao C, et al. A comprehensive analysis and annotation of human normal urinary proteome. Sci Rep (2017) 7(1):3024. doi:10.1038/s41598-017-03226-6
37. Gregorc V, Novello S, Lazzari C, Barni S, Aieta M, Mencoboni M, et al. Predictive value of a proteomic signature in patients with non-small-cell lung cancer treated with second-line erlotinib or chemotherapy (PROSE): a biomarker-stratified, randomised phase 3 trial. Lancet Oncol (2014) 15(7):713–21. doi:10.1016/S1470-2045(14)70162-7
38. Lee AK, D’Amico AV. Utility of prostate-specific antigen kinetics in addition to clinical factors in the selection of patients for salvage local therapy. J Clin Oncol (2005) 23(32):8192–7. doi:10.1200/JCO.2005.03.0007
39. Baca SC, Garraway LA. The genomic landscape of prostate cancer. Front Endocrinol (2012) 3:69. doi:10.3389/fendo.2012.00069
40. Grasso CS, Wu YM, Robinson DR, Cao X, Dhanasekaran SM, Khan AP, et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature (2012) 487(7406):239–43. doi:10.1038/nature11125
41. Rajan P, Sudbery IM, Villasevil ME, Mui E, Fleming J, Davis M, et al. Next-generation sequencing of advanced prostate cancer treated with androgen-deprivation therapy. Eur Urol (2014) 66(1):32–9. doi:10.1016/j.eururo.2013.08.011
42. Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell (2015) 163(4):1011–25. doi:10.1016/j.cell.2015.10.025
43. Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM, et al. Integrative clinical genomics of advanced prostate cancer. Cell (2015) 162(2):454. doi:10.1016/j.cell.2015.06.053
44. Magi-Galluzzi C, Montironi R, Epstein JI. Contemporary Gleason grading and novel grade groups in clinical practice. Curr Opin Urol (2016) 26(5):488–92. doi:10.1097/MOU.0000000000000320
45. Fraser M, Sabelnykova VY, Yamaguchi TN, Heisler LE, Livingstone J, Huang V, et al. Genomic hallmarks of localized, non-indolent prostate cancer. Nature (2017) 541(7637):359–64. doi:10.1038/nature20788
46. Barbieri CE, Baca SC, Lawrence MS, Demichelis F, Blattner M, Theurillat JP, et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat Genet (2012) 44(6):685–9. doi:10.1038/ng.2279
47. Chen G, Gharib TG, Huang CC, Taylor JM, Misek DE, Kardia SL, et al. Discordant protein and mRNA expression in lung adenocarcinomas. Mol Cell Proteomics (2002) 1(4):304–13. doi:10.1074/mcp.M200008-MCP200
48. Schwanhausser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, et al. Global quantification of mammalian gene expression control. Nature (2011) 473(7347):337–42. doi:10.1038/nature10098
49. Zhang B, Wang J, Wang X, Zhu J, Liu Q, Shi Z, et al. Proteogenomic characterization of human colon and rectal cancer. Nature (2014) 513(7518):382–7. doi:10.1038/nature13438
50. Mann M, Hendrickson RC, Pandey A. Analysis of proteins and proteomes by mass spectrometry. Annu Rev Biochem (2001) 70:437–73. doi:10.1146/annurev.biochem.70.1.437
51. Makarov A, Denisov E, Kholomeev A, Balschun W, Lange O, Strupat K, et al. Performance evaluation of a hybrid linear ion trap/orbitrap mass spectrometer. Anal Chem (2006) 78(7):2113–20. doi:10.1021/ac0518811
52. Vizcaino JA, Deutsch EW, Wang R, Csordas A, Reisinger F, Rios D, et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat Biotechnol (2014) 32(3):223–6. doi:10.1038/nbt.2839
53. Beck M, Schmidt A, Malmstroem J, Claassen M, Ori A, Szymborska A, et al. The quantitative proteome of a human cell line. Mol Syst Biol (2011) 7:549. doi:10.1038/msb.2011.82
54. Nagaraj N, Wisniewski JR, Geiger T, Cox J, Kircher M, Kelso J, et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol Syst Biol (2011) 7:548. doi:10.1038/msb.2011.81
55. Gillet LC, Navarro P, Tate S, Rost H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics (2012) 11(6):O111016717. doi:10.1074/mcp.O111.016717
56. Geiger T, Wisniewski JR, Cox J, Zanivan S, Kruger M, Ishihama Y, et al. Use of stable isotope labeling by amino acids in cell culture as a spike-in standard in quantitative proteomics. Nat Protoc (2011) 6(2):147–57. doi:10.1038/nprot.2010.192
57. Rothermundt C, Hayoz S, Templeton AJ, Winterhalder R, Strebel RT, Bartschi D, et al. Metformin in chemotherapy-naive castration-resistant prostate cancer: a multicenter phase 2 trial (SAKK 08/09). Eur Urol (2014) 66(3):468–74. doi:10.1016/j.eururo.2013.12.057
58. Ganguly SS, Li X, Miranti CK. The host microenvironment influences prostate cancer invasion, systemic spread, bone colonization, and osteoblastic metastasis. Front Oncol (2014) 4:364. doi:10.3389/fonc.2014.00364
59. Casey SC, Amedei A, Aquilano K, Azmi AS, Benencia F, Bhakta D, et al. Cancer prevention and therapy through the modulation of the tumor microenvironment. Semin Cancer Biol (2015) 35(Suppl):S199–223. doi:10.1016/j.semcancer.2015.02.007
60. Msaouel P, Galeas JN, Boiles AR, Ruiz RR, Koutsilieris M. Targeting the bone microenvironment in metastatic castration-resistant prostate cancer. Curr Drug Targets (2016) 17(3):276–89. doi:10.2174/1389450116666150420143932
61. Batruch I, Lecker I, Kagedan D, Smith CR, Mullen BJ, Grober E, et al. Proteomic analysis of seminal plasma from normal volunteers and post-vasectomy patients identifies over 2000 proteins and candidate biomarkers of the urogenital system. J Proteome Res (2011) 10(3):941–53. doi:10.1021/pr100745u
62. Kagedan D, Lecker I, Batruch I, Smith C, Kaploun I, Lo K, et al. Characterization of the seminal plasma proteome in men with prostatitis by mass spectrometry. Clin Proteomics (2012) 9(1):2. doi:10.1186/1559-0275-9-2
63. Nanjappa V, Thomas JK, Marimuthu A, Muthusamy B, Radhakrishnan A, Sharma R, et al. Plasma proteome database as a resource for proteomics research: 2014 update. Nucleic Acids Res (2014) 42(Database issue):D959–65. doi:10.1093/nar/gkt1251
64. Hortin GL, Sviridov D. The dynamic range problem in the analysis of the plasma proteome. J Proteomics (2010) 73(3):629–36. doi:10.1016/j.jprot.2009.07.001
65. Carlsson S, Assel M, Ulmert D, Gerdtsson A, Hugosson J, Vickers A, et al. Screening for prostate cancer starting at age 50–54 years. A population-based cohort study. Eur Urol (2017) 71(1):46–52. doi:10.1016/j.eururo.2016.03.026
66. Hugosson J, Carlsson S. Overdetection in screening for prostate cancer. Curr Opin Urol (2014) 24(3):256–63. doi:10.1097/MOU.0000000000000054
67. De Luca S, Passera R, Fiori C, Bollito E, Cappia S, Mario Scarpa R, et al. Prostate health index and prostate cancer gene 3 score but not percent-free prostate specific antigen have a predictive role in differentiating histological prostatitis from PCa and other nonneoplastic lesions (BPH and HG-PIN) at repeat biopsy. Urol Oncol (2015) 33(10):424.e17–23. doi:10.1016/j.urolonc.2015.05.032
68. Halin S, Rudolfsson SH, Doll JA, Crawford SE, Wikstrom P, Bergh A. Pigment epithelium-derived factor stimulates tumor macrophage recruitment and is downregulated by the prostate tumor microenvironment. Neoplasia (2010) 12(4):336–45. doi:10.1593/neo.92046
69. Filiz G, Dass CR. Reduction in tumour cell invasion by pigment epithelium-derived factor is mediated by membrane type-1 matrix metalloproteinase downregulation. Pharmazie (2012) 67(12):1010–4. doi:10.1691/ph.2012.2067
70. Nelius T, Samathanam C, Martinez-Marin D, Gaines N, Stevens J, Hickson J, et al. Positive correlation between PEDF expression levels and macrophage density in the human prostate. Prostate (2013) 73(5):549–61. doi:10.1002/pros.22595
71. Duijvesz D, Luider T, Bangma CH, Jenster G. Exosomes as biomarker treasure chests for prostate cancer. Eur Urol (2011) 59(5):823–31. doi:10.1016/j.eururo.2010.12.031
72. Zijlstra C, Stoorvogel W. Prostasomes as a source of diagnostic biomarkers for prostate cancer. J Clin Invest (2016) 126(4):1144–51. doi:10.1172/JCI81128
73. Soekmadji C, Corcoran NM, Oleinikova I, Jovanovic L; Australian Prostate Cancer Collaboration BioResource; Ramm GA, et al. Extracellular vesicles for personalized therapy decision support in advanced metastatic cancers and its potential impact for prostate cancer. Prostate (2017) 77:1416–23. doi:10.1002/pros.23403
74. Turay D, Khan S, Diaz Osterman CJ, Curtis MP, Khaira B, Neidigh JW, et al. Proteomic profiling of serum-derived exosomes from ethnically diverse prostate cancer patients. Cancer Invest (2016) 34(1):1–11. doi:10.3109/07357907.2015.1081921
75. Sun B, Kovatch JR, Badiong A, Merbouh N. Optimization and modeling of quadrupole orbitrap parameters for sensitive analysis toward single-cell proteomics. J Proteome Res (2017) 16:3711–21. doi:10.1021/acs.jproteome.7b00416
76. Gittelman MC, Hertzman B, Bailen J, Williams T, Koziol I, Henderson RJ, et al. PCA3 molecular urine test as a predictor of repeat prostate biopsy outcome in men with previous negative biopsies: a prospective multicenter clinical study. J Urol (2013) 190(1):64–9. doi:10.1016/j.juro.2013.02.018
77. Ochiai A, Okihara K, Kamoi K, Oikawa T, Shimazui T, Murayama S, et al. Clinical utility of the prostate cancer gene 3 (PCA3) urine assay in Japanese men undergoing prostate biopsy. BJU Int (2013) 111(6):928–33. doi:10.1111/j.1464-410X.2012.11683.x
78. Leyten GH, Hessels D, Jannink SA, Smit FP, de Jong H, Cornel EB, et al. Prospective multicentre evaluation of PCA3 and TMPRSS2-ERG gene fusions as diagnostic and prognostic urinary biomarkers for prostate cancer. Eur Urol (2014) 65(3):534–42. doi:10.1016/j.eururo.2012.11.014
79. Wood SL, Knowles MA, Thompson D, Selby PJ, Banks RE. Proteomic studies of urinary biomarkers for prostate, bladder and kidney cancers. Nat Rev Urol (2013) 10(4):206–18. doi:10.1038/nrurol.2013.24
80. Tosoian JJ, Carter HB, Lepor A, Loeb S. Active surveillance for prostate cancer: current evidence and contemporary state of practice. Nat Rev Urol (2016) 13(4):205–15. doi:10.1038/nrurol.2016.45
81. Garisto JD, Klotz L. Active surveillance for prostate cancer: how to do it right. Oncology (Williston Park) (2017) 31(5):333–40.
82. Porras-Yakushi TR, Hess S. Recent advances in defining the ubiquitylome. Expert Rev Proteomics (2014) 11(4):477–90. doi:10.1586/14789450.2014.926223
83. Thaysen-Andersen M, Packer NH. Advances in LC-MS/MS-based glycoproteomics: getting closer to system-wide site-specific mapping of the N- and O-glycoproteome. Biochim Biophys Acta (2014) 1844(9):1437–52. doi:10.1016/j.bbapap.2014.05.002
84. von Stechow L, Francavilla C, Olsen JV. Recent findings and technological advances in phosphoproteomics for cells and tissues. Expert Rev Proteomics (2015) 12(5):469–87. doi:10.1586/14789450.2015.1078730
85. Morrissey B, O’Shea C, Armstrong J, Rooney C, Staunton L, Sheehan M, et al. Development of a label-free LC-MS/MS strategy to approach the identification of candidate protein biomarkers of disease recurrence in prostate cancer patients in a clinical trial of combined hormone and radiation therapy. Proteomics Clin Appl (2013) 7(5–6):316–26. doi:10.1002/prca.201300004
86. Shi T, Fillmore TL, Gao Y, Zhao R, He J, Schepmoes AA, et al. Long-gradient separations coupled with selected reaction monitoring for highly sensitive, large scale targeted protein quantification in a single analysis. Anal Chem (2013) 85(19):9196–203. doi:10.1021/ac402105s
87. Shi T, Gao Y, Quek SI, Fillmore TL, Nicora CD, Su D, et al. A highly sensitive targeted mass spectrometric assay for quantification of AGR2 protein in human urine and serum. J Proteome Res (2014) 13(2):875–82. doi:10.1021/pr400912c
88. Ebhardt HA, Root A, Sander C, Aebersold R. Applications of targeted proteomics in systems biology and translational medicine. Proteomics (2015) 15(18):3193–208. doi:10.1002/pmic.201500004
89. Percy AJ, Yang J, Hardie DB, Chambers AG, Tamura-Wells J, Borchers CH. Precise quantitation of 136 urinary proteins by LC/MRM-MS using stable isotope labeled peptides as internal standards for biomarker discovery and/or verification studies. Methods (2015) 81:24–33. doi:10.1016/j.ymeth.2015.04.001
90. Wang H, Barbieri CE, He J, Gao Y, Shi T, Wu C, et al. Quantification of mutant SPOP proteins in prostate cancer using mass spectrometry-based targeted proteomics. J Transl Med (2017) 15(1):175. doi:10.1186/s12967-017-1276-7
91. Ruggles KV, Krug K, Wang X, Clauser KR, Wang J, Payne SH, et al. Methods, tools and current perspectives in proteogenomics. Mol Cell Proteomics (2017) 16(6):959–81. doi:10.1074/mcp.MR117.000024
Keywords: mass spectrometry, prostate cancer proteomics, proteome, FFPE, biofluid proteomics, prognostic biomarker, diagnostic biomarker
Citation: Flores-Morales A and Iglesias-Gato D (2017) Quantitative Mass Spectrometry-Based Proteomic Profiling for Precision Medicine in Prostate Cancer. Front. Oncol. 7:267. doi: 10.3389/fonc.2017.00267
Received: 08 September 2017; Accepted: 23 October 2017;
Published: 07 November 2017
Edited by:
Stéphane Terry, Institut Gustave Roussy, FranceReviewed by:
Harald Mischak, University of Glasgow, United KingdomNaser Uddin Höti, Johns Hopkins School of Medicine, United States
Katarina Davalieva, Macedonian Academy of Sciences and Arts, Macedonia
Copyright: © 2017 Flores-Morales and Iglesias-Gato. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Diego Iglesias-Gato, ZGllZ28uaWdsZXNpYXNAc3VuZC5rdS5kaw==
 Amilcar Flores-Morales
Amilcar Flores-Morales Diego Iglesias-Gato
Diego Iglesias-Gato