- 1Breast Center of the First Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China
- 2MOE Laboratory of Biosystem Homeostasis and Protection, College of Life Sciences, Zhejiang University, Hangzhou, China
- 3Reproductive and Developmental Biology Laboratory, National Institute of Environmental Health Sciences, Durham, NC, United States
Long noncoding RNAs (lncRNAs) play crucial roles in various aspects of cellular functions. Recent studies have revealed that lncRNAs are critical players in the immune system by modulating immune cell differentiation and functions, particularly in cancer immunity. Here we systematically summarize how lncRNAs are involved in different processes of the cancer immunity cycle, including immune cell differentiation, proliferation, trafficking, and infiltration. Moreover, the limitations of the current understanding of lncRNA’s functions in cancer immunity are described, such as the complexity of the cancer immunity system, the inclusive functions of lncRNAs in this system, and the associated immune response. In sum, the comprehensive investigation of the roles of lncRNAs in cancer immunity aids in cancer diagnosis and therapies.
Introduction
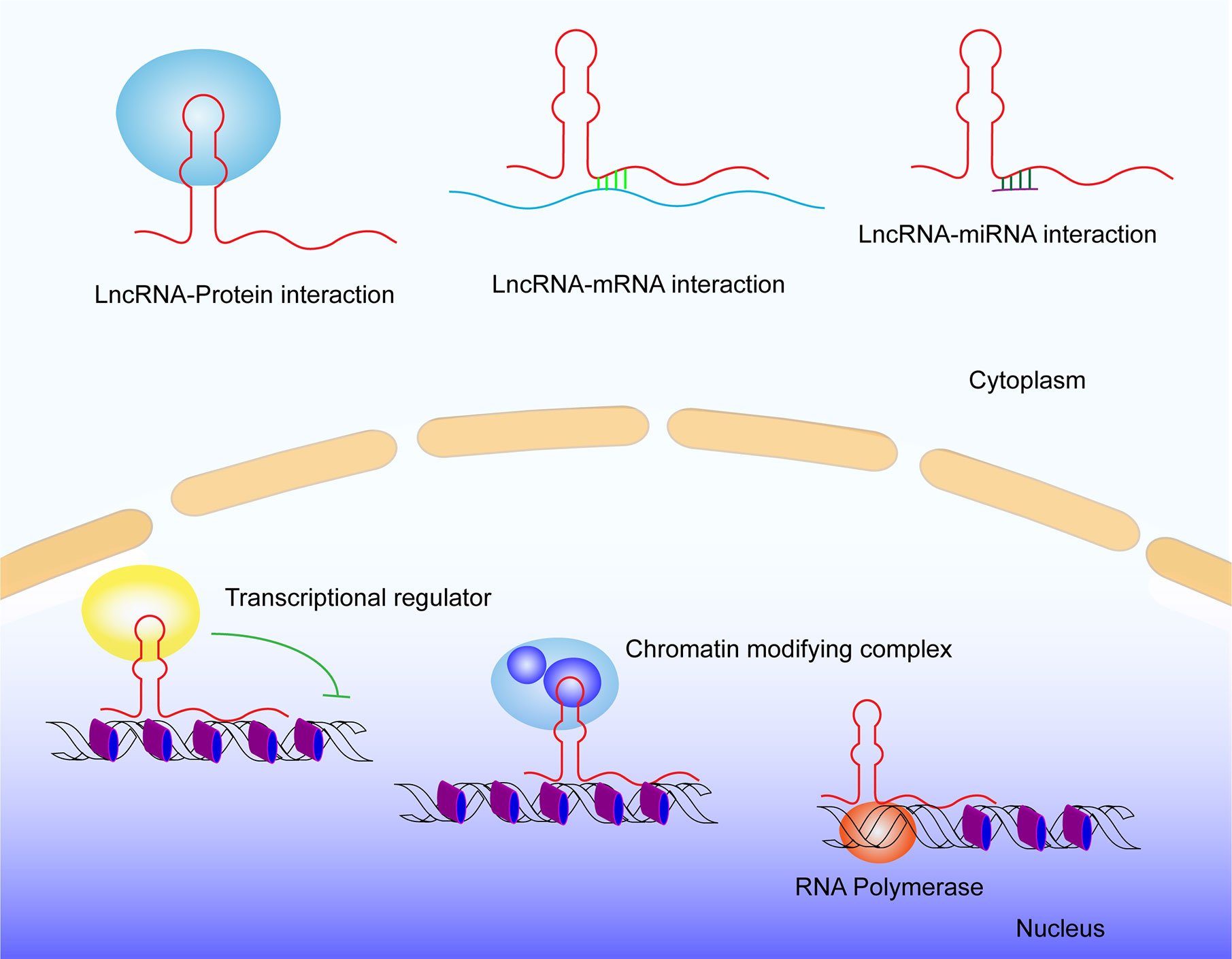
With the advance of the human genome sequence, more than 90% of the human genome transcribed RNAs lack protein-coding functions. Nearly 75% among them are noncoding RNAs (ncRNAs), which are divided into two groups: small ncRNAs (<200nt) and long ncRNAs (lncRNAs) (≥200nt) (1). Although lncRNAs cannot directly encode proteins due to the lack of open reading frames, they can control gene transcription and expression by regulating transcriptional activators or repressors, different components of transcription reaction (e.g., RNA polymerase II), and DNA duplex (2). LncRNAs also regulate post-transcription, such as mRNA splicing (3, 4) and translation (5). What is more, they regulate epigenetic modification, including DNA methylation (6, 7), histone acetylation (8), and ubiquitination (9). LncRNAs powerfully function in various cellular activities. For example, lncRNAs play indispensable roles in the progression of various cancers in past years, such as breast cancer (10, 11), prostate cancer (12), gastric cancer (13), and lung cancer (14–16). Of note, lncRNAs have been found to be the key players in regulating cancer immunity. In this review, we aim to summarize the findings of lncRNAs functions in cancer immunity as well as their clinical impacts.
Cancer Immunity Cycle
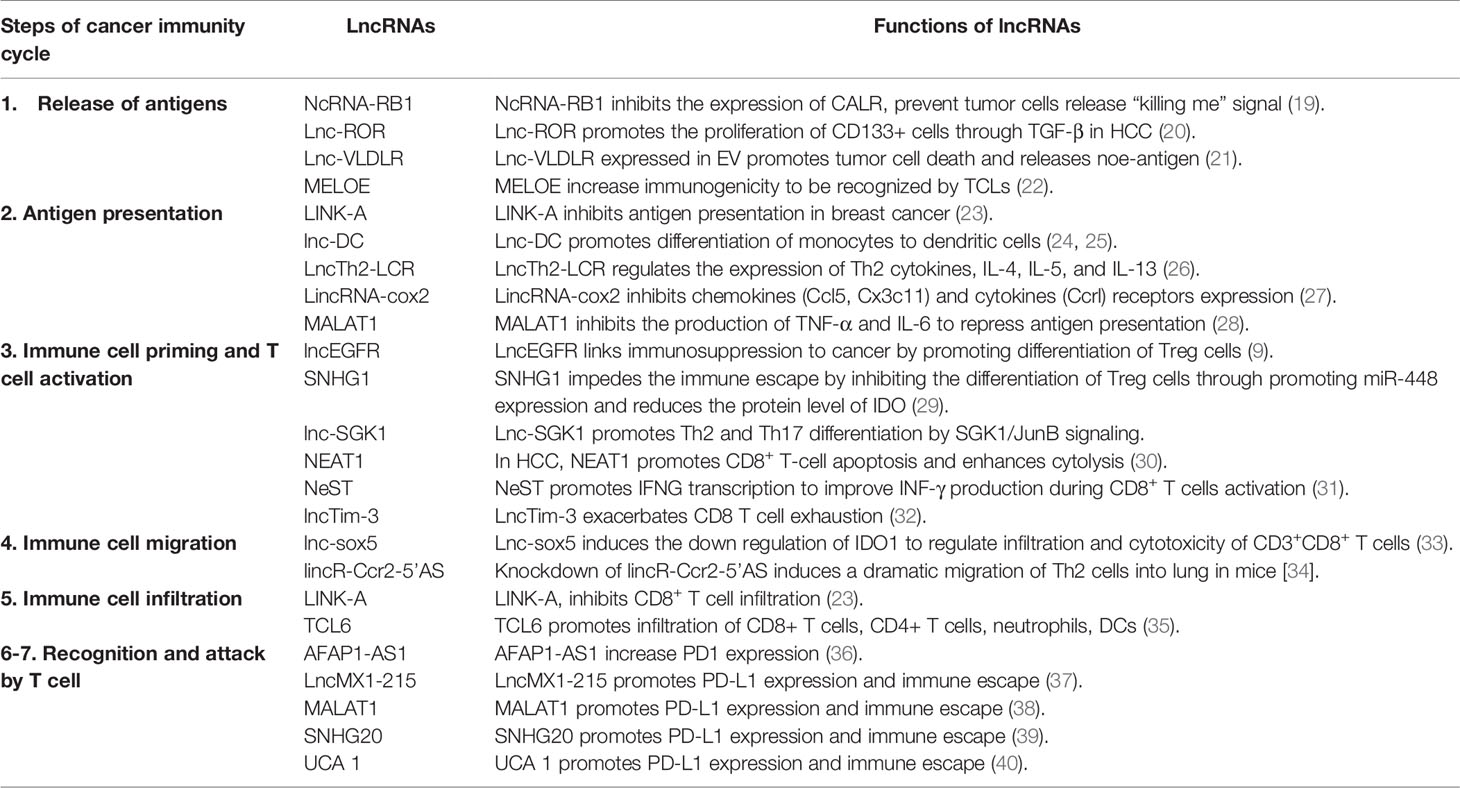
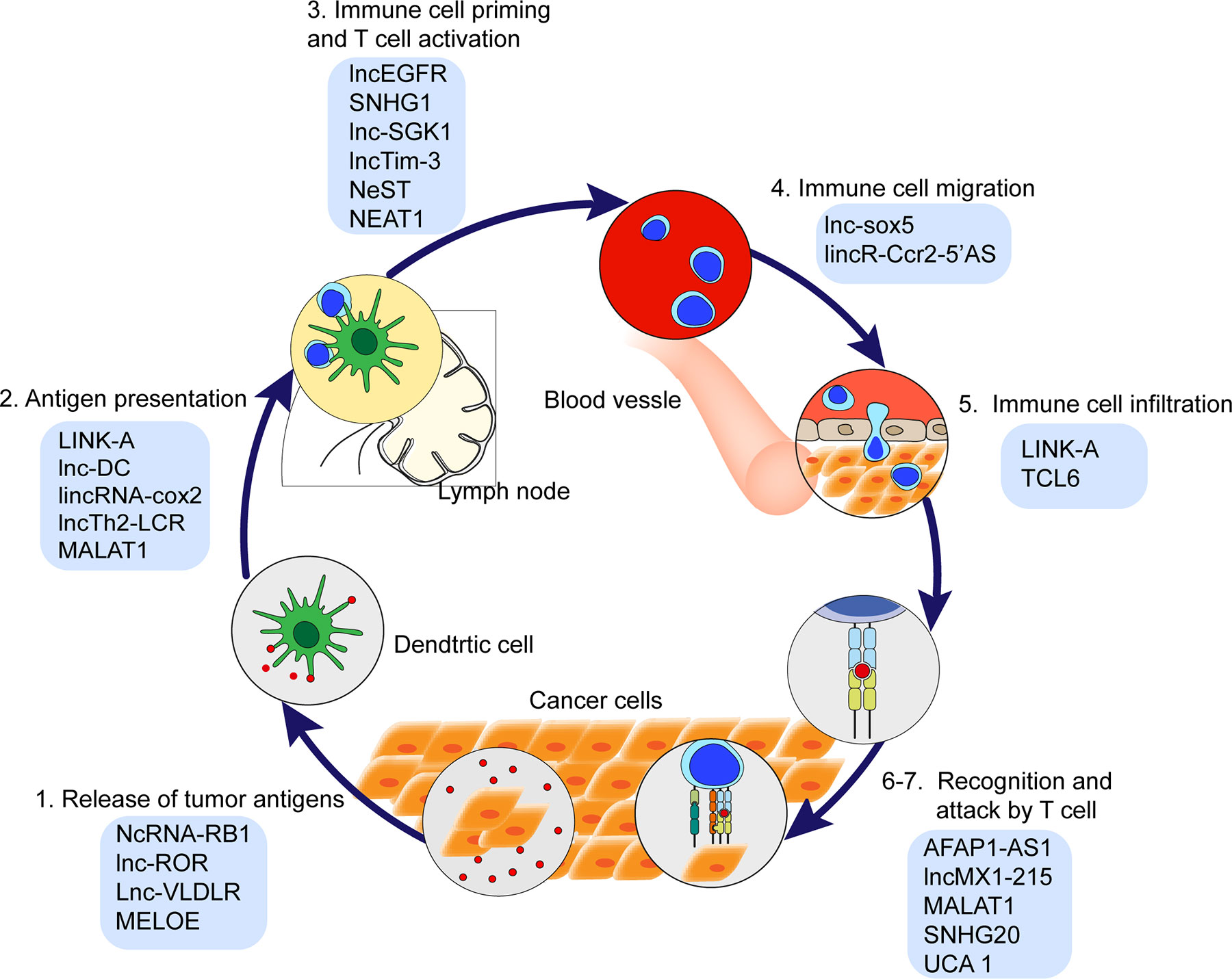
The cancer immunity cycle is defined as stepwise events effectively controlling cancer cell growth by immune system. This cycle is arbitrarily divided into seven steps (Figure 1): cancer cells release antigens (Step 1); antigens are captured by dendritic cells (DCs) (Step 2); DCs with captured antigens migrate to lymph node and prime with T cells to activate tumor-specific cytolytic CD8+ T cells (Step 3); the cytolytic T cells migrate from lymph node into blood vessels (Step 4); immune cells infiltrate into tumor stroma (Step 5); the cytolytic T cells recognize tumor cells (Step 6); T cells kill cancer cells (Step 7) (17, 18). Recently, lncRNAs have been revealed as key regulators in the cancer immunity cycle (Figure 2 and Table 1), including the impact on the epigenetic and transcriptional profiles of immune cells.
lncRNAs in Tumor Antigen Release
Tumor antigen is an antigenic substance produced by tumor cells, especially dying tumor cells. It triggers the immune system to attack it (41, 42). Calreticulin (CALR), which is a calcium-binding chaperone and influences antigen presentation to cytotoxic T cells (43), is expressed in many cancer cells and promotes macrophages to engulf hazardous cancerous cells (44). In lung cancer, NcRNA-RB1, a lncRNA expressed from the RB1 promoter, inhibits the expression of CALR. It shows a tumor-inhibitor mechanism through impairing the “kill me” signal (19). Meanwhile, the efficiency of chemotherapy is associated with the expression of some lncRNAs besides the death of immune cells or released neoantigens. For example, Lnc-ROR, a stress-responsive lncRNA highly expressed in hepatocellular carcinoma (HCC) cells, promotes the proliferation of CD133+ cells by activating TGF-β pathway to weaken the effectiveness of chemotherapy drugs in HCC (20). Lnc-VLDLR (very low-density lipoprotein-receptor-VLDLR) is expressed in extracellular vesicles (EV) and promotes tumor cell death during chemotherapy, leading to increased chemotherapy efficiency (21). Taken together, the roles of lncRNAs in tumor antigen release are starting to be recognized.
lncRNAs in Antigen Presentation
Antigen presentation is a vital immune process to trigger effective immune response. Activation of T cells depends on effective antigen presentation, including immunogenic signals and specific antigen presentation cells (APCs). LINK-A inhibits antigen presentation by specifically inhibiting antigen-presenting cell (APC) and decreasing CD8+ T cell abundance in basal-like breast cancer (45). DCs are antigen-presenting cells that act as the messengers between the innate and adaptive immune system. The lncRNA that is expressed exclusively in human DC (lnc-DC) has been identified as an lncRNA involved in the differentiation of monocytes to DCs. Lnc-DC mainly exists in the cytoplasm and activates DC transcription factor STAT3by binding with STAT3 and promoting STAT3 tyrosine-705 phosphorylation. During the differentiation of DCs, the transcript of lnc-DC is upregulated (24). Knockdown of lnc-DC impairs the expression of DC-specific genes regulating antigen presentation, T cell activation, and cell migration, such as CD40, CD80, CD86, and CCR7 (24, 25). Moreover, deletion of lnc-DC in DC cells leads to impaired priming CD4+ T cells or secreting inflammatory cytokines after pathogen stimulation (24).
Moreover, lncRNAs regulate pro-inflammatory cytokines that are crucial to antigen presentation, such as IFN-γ and TNF-α (46–48). LincRNA-cox2 is an lncRNA located adjacent to Cox2. The study reported that silencing lincRNA-cox2 leads to increased expression of chemokines (Ccl5, Cx3c11) and cytokine receptors (Ccrl) (27). Moreover, lncRNA Th2 locus control region (lncTh2-LCR) regulates expression of Th2 cytokines, IL-4, IL-5, and IL-13 (26), and MALAT1 (Metastasis Associated Lung Adenocarcinoma Transcript 1) inhibits the production of TNF-α and IL-6 by inhibiting NF-kB DNA binding to gene promoters (28). More interestingly, lncRNAs also improve antigen presentation after being translated into short polypeptides. In melanoma, lncRNA MELOE, which is translated into MELOE-1, MELOE-2, and MELOE-3 by different translational approaches, shows the highest immunogenicity and can be recognized by tumor-infiltrating lymphocytes. MELOE is currently considered as a targeted specific antigen to improve the efficacy of melanoma immunotherapy (22). These findings establish the key roles of lncRNAs in antigen presentation.
lncRNAs in Immune Cell Priming and Activation
Effective immune priming and activation are determined by the ratio of T effector cells and T regulatory (Treg) cells (49). Some lncRNAs have been demonstrated to regulate lymphocyte differentiation and activation (50–54). In HCC, lnc-epidermal growth factor receptor (lncEGFR) promotes Treg cells differentiation and suppresses cytotoxic T lymphocytes (CTLs) activity (9, 55). Mechanistically, lncEGFR specifically binds to EGFR and blocks its ubiquitination by c-CBL, leading to EGFR stabilization and to augment its activation and its downstream AP-1/NF-AT1 axis. Indoleamine 2, 3-dioxygenase (IDO) is encoded by IDO1 gene and a heme-containing enzyme physiologically expressed in a number of tissues. IDO not only suppresses anti-tumor immune response in the body (56, 57), it also limits T-cell function and engages in mechanisms of immune tolerance (58). In breast cancer, small nucleolar RNA host gene 1 (SNHG1) negatively regulates the protein level of IDO to inhibit Treg cells differentiation and impede immune escape (29). In addition, lnc-SGK1 (Serine/threonine-protein kinase-SGK1) promotes Th2 and Th17 differentiation by SGK1/JunB signaling in gastric cancer (59). In HCC, Nuclear Enriched Abundant Transcript 1 (NEAT1) promotes CD8+ T cell apoptosis and enhances cytolysis through miR-155/Tim-3 pathway (30). Lnc-Tim-3 specifically binds to Tim-3 and blocks its interaction with Bat3, thus suppressing downstream Lck/NEAT1/AP-1 signaling and exacerbating CD8+ T cell exhaustion (32). NeST (Nettoie salmonella pas Theiler’s) is a lncRNA expressed in different immune cells including CD8+ T cells, CD4+ T TH1 cells, and natural killer cells (34, 60, 61). NeST promotes IFNG transcription to improve INF-γ production during CD8+ T cell activation (31). These findings indicate lncRNAs modulate immune cell priming and activation by different mechanisms. The emerging roles of lncRNAs in immune cell priming and activation indicate that these lncRNAs can be potential targets for cancer therapies.
Role of lncRNAs in Immune Cell Migration
Recent studies have shown that some lncRNAs (e.g., lnc-sox5 and lincR-Ccr2-5’AS) are involved in immune cell trafficking. Knockdown of lnc-sox5 induces down-regulation of IDO1 in xenograft mouse model, leading to the modulation of CD3+CD8+ T cells infiltration and cytotoxicity (33). lincR-Ccr2-5’AS is a lncRNA specifically expressed in Th2 cells and transcribed in the opposite direction of the Ccr2 gene. Knockdown of lincR-Ccr2-5’AS induces a dramatic Th2 cells migration into the lung in mice (61). These findings spike the notion that lncRNAs can directly modulate the migration of immune cells.
lncRNAs in Immune Cell Infiltration
Effective response of cancer immunity is also indispensable of effective T cells infiltrating into tumor stroma. By far, there are limited pieces of evidence about lncRNAs in the process of T cell infiltration in the cancer immunity cycle. In triple-negative breast cancer (TNBC) of patients receiving pembrolizumab (anti-PD-1) treatment, the pembrolizumab responders exhibited a relatively lower expression of LINK-A and a higher CD8+ T cell infiltration compared with non-responders (45), concluding that the CD8+ T cell infiltration in TNBC is negatively correlated with LINK-A expression (45). In breast cancer, lncRNA T-cell leukemia/lymphoma 6 (TCL6) is correlated with the infiltration of CD8+ T cells, CD4+ T cells, neutrophils, DCs, and the frequency of tumor-infiltrating lymphocytes (TILs) (35). Thus, more efforts to show direct evidence of whether and how lncRNAs function in immune cell infiltration are in urgent need.
lncRNAs in the Recognition and Attack of Cancer Cells
Cell-cell recognition ability helps cells distinguish one type of neighboring cells from another. This ability requires complementary molecules of the different cellular surfaces to recognize each other, such as a receptor and its specific ligand. For example, programmed cell death 1 (PD-1) is an immune checkpoint receptor on the T cell surface. PD-1 binds to its ligands, programmed cell death 1 ligand 1 (PD-L1) and PD-L2 on cancer cell surface, functionally causing immune inhibition (17, 62). Overexpression of PD-L1 associates with poor prognosis and enables cancer cells to escape from immune surveillance in various cancer types. Of note, lncRNAs, including lncRNA AFAP1-AS1 and lncMX1-215, have been found to regulate PD-1/PD-L1 signaling. LncRNA AFAP1-AS1 can induce increased expression of PD1 in tumor microenvironment of nasopharyngeal carcinoma (36). Moreover, there is a positively expressing correlation between lncRNA AFAP1-AS1 and PD1 in nasopharyngeal carcinoma (NPC) (36). LncMX1-215, a novel IFNα-induced long noncoding RNA, significantly inhibits IFNα-induced expression of PD-L1 and galectin-9 in head and neck squamous cell carcinoma (37). In addition, in diffuse large B-cell lymphoma, MALAT1 sponges miR-195 to upregulate PD-L1 expression to promote cancer cells migration and immune escape. Inhibition of MALAT1 can rescue these events and attenuate epithelial-mesenchymal transition-like process (38). In esophageal squamous cell carcinoma, Small nucleolar RNA host gene 20 (SNHG20) promotes PD-L1 expression via ataxia telangiectasia mutated kinase (ATM)/JAK-PD-L1 pathway (39). In gastric cancer, UCA1 (Urothelial Cancer-Associated 1) promotes PD-L1 expression by repressing miR-26a/b, miR-193a, and miR-214, which contributes to immune escape of gastric cancer cells (40). These findings point out that lncRNAs are involved in the regulation of immune checkpoints in cancer immunity cycle, indicating that these lncRNAs may be synergistic targets for immunotherapy.
Discussion
Immunotherapy has gained a lot of interest from researchers and clinicians due to its great potential for effectively treating cancer. The immune checkpoint inhibitors in the clinic, like PD1, PD-L1, and CTAL-4, have made great clinical achievements (63, 64). But some of the cancer patients do not respond to these immune checkpoint inhibitors, pointing out the need for more investigation of the cancer immune system. LncRNAs exhibit multiple and powerful functions in the tumor microenvironment, including tumorigenesis, tumor metastases, and immune disorders (10, 14, 33, 37). LncRNAs interact with DNA, mRNA, ncRNAs, and proteins to alter cellular physiology. LncRNAs have high tissue- and cell-specificity indicating a wholely different range of diagnosis and treatment of cancer (65). Some combination therapies, including combination of immunotherapy with chemotherapy, combination of surgery with chemotherapy, and even drug combination therapy, have attained obvious clinal progress (18, 62, 64, 66). Thus, consideration of a combination of targeting on lncRNAs and immunotherapy is attractive for cancer treatment. With the rapid development of RNA-targeting technologies, there are many approaches targeting lncRNAs emerging, like aptamers, small molecule inhibitors, natural antisense transcripts (NATs), an antisense ligonucleotide (ASO), synthetic lncRNA mimics, and other constructs (67). The off-target effect in systemic nucleic acid based therapies is a main problem in lncRNA therapies. CRISPR-Cas13a, a programmable RNA editing system that allows for temporal modulation of genetic variants in transcripts, may be a potential candidate for manipulation RNA expression with low off-target efficiency in lncRNA therapies. These approaches enable us to target lncRNAs to modulate their functions for clinical purposes. However, as for utilizing lncRNAs to aid in cancer diagnosis and therapy, the biggest challenge in lncRNAs research is the lack of conservation for many lncRNA species. Some cancer-related lncRNAs are well conserved, but most human cancer-related lncRNAs do not exhibit sequence conservation across mammalian species. This will impede preclinical research for therapeutic targeting in animals. By far, the patient-derived tumor organoids serve as a powerful model to explore the role of lncRNA in a patient-specific manner. Moreover, due to the complexity of the immune system, the functional investigation of lncRNAs in cancer immunity is just beginning, and their roles as both biomarkers and targets in cancer immunity cycle need further study.
Author contributions
AL contributed to the study design and data analysis, and edited the manuscript. MW and PF wrote the manuscript. MW and LQ contributed to the figure and table design. AL and JL edited the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was supported in part by the National Natural Science Foundation of China (81672791, 81872300), Zhejiang Provincial Natural Science Fund for Distinguished Young Scholars of China (LR18C060002) to AL. AL is a scholar of Thousand Youth Talents-China, a scholar of Thousand Talents-Zhejiang, a scholar of Hundred Talents-Zhejiang University.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The manuscript was edited by JL in his private capacity. No official support or endorsement by the National Institutes of Health, National Institute of Environmental Health is intended or should be inferred. Due to space limitations, we apologize to the authors of the publications that could not be cited in the scope of this review.
Abbreviations
LncRNA, long noncoding RNA; APC, antigen presenting cell; CTL, cytotoxic T lymphocyte; MHC I, major histocompatibility complex class I; NK, natural killer; NSCLC, non-small-cell lung cancer; PD1, programmed cell death protein 1; PD-L1, programmed cell death ligand 1; TCR, T cell receptor; TH, T Helper; TNBC, triple negative breast cancer; Treg, T-regulatory; CALR, calreticulin; VLDLR, very low-density lipoprotein-receptor; HCC, hepatocellular carcinoma; DC, dendritic cell; IDO, indoleamine 2, 3-dioxygenase; SGK1, serine/threonine-protein kinase; SNHG1, small nucleolar RNA host gene 1; NEAT1, nuclear Enriched Abundant Transcript 1; NeST, nettoie salmonella pas Theiler’s; TILs, tumor-infiltrating lymphocytes; NATs, natural antisense transcripts; ASO, antisense oligonucleotide.
References
1. Chen L, Dzakah EE, Shan G. Targetable long non-coding RNAs in cancer treatments. Cancer Lett (2018) 418:119–24. doi: 10.1016/j.immuni.2013.07.012
2. Goodrich JA, Kugel JF. Non-coding-RNA regulators of RNA polymerase II transcription. Nat Rev Mol Cell Biol (2006) 7:612–6. doi: 10.1038/nrm1946
3. Beltran M, Puig I, Pena C, Garcia JM, Alvarez AB, Pena R, et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev (2008) 22:756–69. doi: 10.1101/gad.455708
4. Munroe SH, Lazar MA. Inhibition of c-erbA mRNA splicing by a naturally occurring antisense RNA. J Biol Chem (1991) 266:22083–6.
5. Wang H, Iacoangeli A, Lin D, Williams K, Denman RB, Hellen CU, et al. Dendritic BC1 RNA in translational control mechanisms. J Cell Biol (2005) 171:811–21. doi: 10.1083/jcb.200506006
6. Tang B. Inference of Crosstalk Effects between DNA Methylation and lncRNA Regulation in NSCLC. BioMed Res Int (2018) 2018:7602794. doi: 10.1155/2018/7602794
7. Zhou J, Yang L, Zhong T, Mueller M, Men Y, Zhang N, et al. H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase. Nat Commun (2015) 6:10221. doi: 10.1038/ncomms10221
8. Jain AK, Xi Y, McCarthy R, Allton K, Akdemir KC, Patel LR, et al. LncPRESS1 Is a p53-Regulated LncRNA that Safeguards Pluripotency by Disrupting SIRT6-Mediated De-acetylation of Histone H3K56. Mol Cell (2016) 64:967–81. doi: 10.1016/j.molcel.2016.10.039
9. Li Z, Hou P, Fan D, Dong M, Ma M, Li H, et al. The degradation of EZH2 mediated by lncRNA ANCR attenuated the invasion and metastasis of breast cancer. Cell Death Differentiation (2017) 24:59–71. doi: 10.1038/cdd.2016.95
10. Lin A, Hu Q, Li C, Xing Z, Ma G, Wang C, et al. The LINK-A lncRNA interacts with PtdIns(3,4,5)P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Nat Cell Biol (2017) 19:238–51. doi: 10.1038/ncb3473
11. Sang LJ, Ju HQ, Liu GP, Tian T, Ma GL, Lu YX, et al. LncRNA CamK-A Regulates Ca(2+)-Signaling-Mediated Tumor Microenvironment Remodeling. Mol Cell (2018) 72:601. doi: 10.1016/j.molcel.2018.08.014
12. Jin G, Sun J, Isaacs SD, Wiley KE, Kim ST, Chu LW, et al. Human polymorphisms at long non-coding RNAs (lncRNAs) and association with prostate cancer risk. Carcinogenesis (2011) 32:1655–9. doi: 10.1093/carcin/bgr187
13. Li L, Jia F, Bai P, Liang Y, Sun R, Yuan F, et al. Association between polymorphisms in long non-coding RNA PRNCR1 in 8q24 and risk of gastric cancer. Tumour Biol J Int Soc Oncodevelopmental Biol Med (2016) 37:299–303. doi: 10.1007/s13277-015-3750-2
14. Li L, Wang Y, Song G, Zhang X, Gao S, Liu H. HOX cluster-embedded antisense long non-coding RNAs in lung cancer. Cancer Lett (2019) 450:14–21. doi: 10.1016/j.canlet.2019.02.036
15. Teng C, Huang G, Luo Y, Pan Y, Wang H, Liao X, et al. Differential long noncoding RNAs expression in cancer-associated fibroblasts of non-small-cell lung cancer. Pharmacogenomics (2019) 20:143–53. doi: 10.2217/pgs-2018-0102
16. Yuan H, Liu H, Liu Z, Owzar K, Han Y, Su L, et al. A Novel Genetic Variant in Long Non-coding RNA Gene NEXN-AS1 is Associated with Risk of Lung Cancer. Sci Rep (2016) 6:34234. doi: 10.1038/srep34234
17. Chen DS, Mellman I. Oncology meets immunology: the cancer-immunity cycle. Immunity (2013) 39:1–10. doi: 10.1016/j.immuni.2013.07.012
18. Gajiwala S, Torgeson A, Garrido-Laguna I, Kinsey C, Lloyd S. Combination immunotherapy and radiation therapy strategies for pancreatic cancer-targeting multiple steps in the cancer immunity cycle. J Gastrointest Oncol (2018) 9:1014–26. doi: 10.21037/jgo.2018.05.16
19. Musahl AS, Huang X, Rusakiewicz S, Ntini E, Marsico A, Kroemer G, et al. A long non-coding RNA links calreticulin-mediated immunogenic cell removal to RB1 transcription. Oncogene (2015) 34:5046–54. doi: 10.1038/onc.2014.424
20. Takahashi K, Yan IK, Kogure T, Haga H, Patel T. Extracellular vesicle-mediated transfer of long non-coding RNA ROR modulates chemosensitivity in human hepatocellular cancer. FEBS Open Bio (2014a) 4:458–67. doi: 10.1016/j.fob.2014.04.007
21. Takahashi K, Yan IK, Wood J, Haga H, Patel T. Involvement of extracellular vesicle long noncoding RNA (linc-VLDLR) in tumor cell responses to chemotherapy. Mol Cancer Res (2014b) 12:1377–87. doi: 10.1158/1541-7786.MCR-13-0636
22. Charpentier M, Croyal M, Carbonnelle D, Fortun A, Florenceau L, Rabu C, et al. IRES-dependent translation of the long non coding RNA meloe in melanoma cells produces the most immunogenic MELOE antigens. Oncotarget (2016) 7:59704–13. doi: 10.18632/oncotarget.10923
23. Hu Q, Ye Y, Chan L, Li Y, Liang K, Lin A, et al. Oncogenic lncRNA downregulates cancer cell antigen presentation and intrinsic tumor suppression. Nat Immunol (2019) 20:835–51. doi: 10.1038/s41590-019-0400-7
24. Wang P, Xue Y, Han Y, Lin L, Wu C, Xu S, et al. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science (2014) 344:310–3. doi: 10.1126/science.1251456
25. Tsuiji H, Yoshimoto R, Hasegawa Y, Furuno M, Yoshida M, Nakagawa S. Competition between a noncoding exon and introns: Gomafu contains tandem UACUAAC repeats and associates with splicing factor-1. Genes Cells (2011) 16:479–90. doi: 10.1111/j.1365-2443.2011.01502.x
26. Spurlock 3rd CF, Tossberg JT, Guo Y, Collier SP, Crooke 3rd PS, Aune TM. Expression and functions of long noncoding RNAs during human T helper cell differentiation. Nat Commun (2015) 6:6932. doi: 10.1038/ncomms7932
27. Carpenter S, Aiello D, Atianand MK, Ricci EP, Gandhi P, Hall LL, et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science (2013) 341:789–92. doi: 10.1126/science.1240925
28. Zhao G, Su Z, Song D, Mao Y, Mao X. The long noncoding RNA MALAT1 regulates the lipopolysaccharide-induced inflammatory response through its interaction with NF-kappaB. FEBS Lett (2016) 590:2884–95. doi: 10.1002/1873-3468.12315
29. Pei X, Wang X, Li H. LncRNA SNHG1 regulates the differentiation of Treg cells and affects the immune escape of breast cancer via regulating miR-448/IDO. Int J Biol Macromolecules (2018) 118:24–30. doi: 10.1016/j.ijbiomac.2018.06.033
30. Yan K, Fu Y, Zhu N, Wang Z, Hong JL, Li Y, et al. Repression of lncRNA NEAT1 enhances the antitumor activity of CD8(+)T cells against hepatocellular carcinoma via regulating miR-155/Tim-3. Int J Biochem Cell Biol (2019) 110:1–8. doi: 10.1016/j.biocel.2019.01.019
31. Gomez JA, Wapinski OL, Yang YW, Bureau JF, Gopinath S, Monack DM, et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-gamma locus. Cell (2013) 152:743–54. doi: 10.1016/j.cell.2013.01.015
32. Ji J, Yin Y, Ju H, Xu X, Liu W, Fu Q, et al. Long non-coding RNA Lnc-Tim3 exacerbates CD8 T cell exhaustion via binding to Tim-3 and inducing nuclear translocation of Bat3 in HCC. Cell Death Dis (2018) 9:478. doi: 10.1038/s41419-018-0528-7
33. Wu K, Zhao Z, Liu K, Zhang J, Li G, Wang L. Long noncoding RNA lnc-sox5 modulates CRC tumorigenesis by unbalancing tumor microenvironment. Cell Cycle (2017) 16:1295–301. doi: 10.1080/15384101.2017.1317416
34. Collier SP, Collins PL, Williams CL, Boothby MR, Aune TM. Cutting edge: influence of Tmevpg1, a long intergenic noncoding RNA, on the expression of Ifng by Th1 cells. J Immunol (2012) 189:2084–8. doi: 10.4049/jimmunol.1200774
35. Zhang Y, Li Z, Chen M, Chen H, Zhong Q, Liang L, et al. lncRNA TCL6 correlates with immune cell infiltration and indicates worse survival in breast cancer. Breast Cancer (2020) 27(4):573–85. doi: 10.1007/s12282-020-01048-5
36. Tang Y, He Y, Shi L, Yang L, Wang J, Lian Y, et al. Co-expression of AFAP1-AS1 and PD-1 predicts poor prognosis in nasopharyngeal carcinoma. Oncotarget (2017) 8:39001–11. doi: 10.18632/oncotarget.16545
37. Ma H, Chang H, Yang W, Lu Y, Hu J, Jin S. A novel IFNalpha-induced long noncoding RNA negatively regulates immunosuppression by interrupting H3K27 acetylation in head and neck squamous cell carcinoma. Mol Cancer (2020) 19:4. doi: 10.1186/s12943-019-1123-y
38. Wang Q-M, Lian G-Y, Song Y, Huang Y-F, Gong Y. LncRNA MALAT1 promotes tumorigenesis and immune escape of diffuse large B cell lymphoma by sponging miR-195. Life Sci (2019) 231:116335–5. doi: 10.1016/j.lfs.2019.03.040
39. Zhang C, Jiang F, Su C, Xie P, Xu L. Upregulation of long noncoding RNA SNHG20 promotes cell growth and metastasis in esophageal squamous cell carcinoma via modulating ATM-JAK-PD-L1 pathway. J Cell Biochem (2019) 20(7):11642–50. doi: 10.1002/jcb.28444
40. Wang CJ, Zhu CC, Xu J, Wang M, Zhao WY, Liu Q, et al. The lncRNA UCA1 promotes proliferation, migration, immune escape and inhibits apoptosis in gastric cancer by sponging anti-tumor miRNAs. Mol Cancer (2019) 18:115. doi: 10.1186/s12943-019-1032-0
41. Wang X, Lu XJ, Sun B. The pros and cons of dying tumour cells in adaptive immune responses. Nat Rev Immunol (2017) 17:591. doi: 10.1038/nri.2017.87
42. Yatim N, Cullen S, Albert ML. Dying cells actively regulate adaptive immune responses. Nat Rev Immunol (2017) 17:262–75. doi: 10.1038/nri.2017.9
43. Raghavan M, Wijeyesakere SJ, Peters LR, Del Cid N. Calreticulin in the immune system: ins and outs. Trends Immunol (2013) 34:13–21. doi: 10.1016/j.it.2012.08.002
44. Chao MP, Jaiswal S, Weissman-Tsukamoto R, Alizadeh AA, Gentles AJ, Volkmer J, et al. Calreticulin is the dominant pro-phagocytic signal on multiple human cancers and is counterbalanced by CD47. Sci Trans Med (2010) 2:63ra94. doi: 10.1126/scitranslmed.3001375
45. Li H, Li CW, Li X, Ding Q, Guo L, Liu S, et al. MET Inhibitors Promote Liver Tumor Evasion of the Immune Response by Stabilizing PDL1. Gastroenterology (2019) 1561849-1861:e1813. doi: 10.1053/j.gastro.2019.01.252
46. Basler M, Kirk CJ, Groettrup M. The immunoproteasome in antigen processing and other immunological functions. Curr Opin Immunol (2013) 25:74–80. doi: 10.1016/j.coi.2012.11.004
47. Lippitz BE. Cytokine patterns in patients with cancer: a systematic review. Lancet Oncol (2013) 14:e218–228. doi: 10.1016/S1470-2045(12)70582-X
48. Zhou F. Molecular mechanisms of IFN-gamma to up-regulate MHC class I antigen processing and presentation. Int Rev Immunol (2009) 28:239–60. doi: 10.1080/08830180903013034
49. Kishton RJ, Sukumar M, Restifo NP. Metabolic Regulation of T Cell Longevity and Function in Tumor Immunotherapy. Cell Metab (2017) 26:94–109. doi: 10.1016/j.cmet.2017.06.016
50. Kanduri K, Tripathi S, Larjo A, Mannerstrom H, Ullah U, Lund R, et al. Identification of global regulators of T-helper cell lineage specification. Genome Med (2015) 7:122. doi: 10.1186/s13073-015-0237-0
51. Pang KC, Dinger ME, Mercer TR, Malquori L, Grimmond SM, Chen W, et al. Genome-wide identification of long noncoding RNAs in CD8+ T cells. J Immunol (2009) 182:7738–48. doi: 10.4049/jimmunol.0900603
52. Panzeri I, Rossetti G, Abrignani S, Pagani M. Long Intergenic Non-Coding RNAs: Novel Drivers of Human Lymphocyte Differentiation. Front Immunol (2015) 6:175. doi: 10.3389/fimmu.2015.00175
53. Ranzani V, Arrigoni A, Rossetti G, Panzeri I, Abrignani S, Bonnal RJ, et al. Next-Generation Sequencing Analysis of Long Noncoding RNAs in CD4+ T Cell Differentiation. Methods Mol Biol (2017) 1514:173–85. doi: 10.1007/978-1-4939-6548-9_14
54. Xia F, Dong F, Yang Y, Huang A, Chen S, Sun D, et al. Dynamic transcription of long non-coding RNA genes during CD4+ T cell development and activation. PloS One (2014) 9:e101588. doi: 10.1371/journal.pone.0101588 doi: 10.1371/journal.pone.0101588
55. Jiang R, Tang J, Chen Y, Deng L, Ji J, Xie Y, et al. The long noncoding RNA lnc-EGFR stimulates T-regulatory cells differentiation thus promoting hepatocellular carcinoma immune evasion. Nat Commun (2017) 8:15129. doi: 10.1038/ncomms15129
56. Munn DH, Mellor AL. Indoleamine 2,3 dioxygenase and metabolic control of immune responses. Trends Immunol (2013) 34:137–43. doi: 10.1016/j.it.2012.10.001
57. Prendergast GC, Smith C, Thomas S, Mandik-Nayak L, Laury-Kleintop L, Metz R, et al. Indoleamine 2,3-dioxygenase pathways of pathogenic inflammation and immune escape in cancer. Cancer Immunol Immunother CII (2014) 63:721–35. doi: 10.1007/s00262-014-1549-4
58. Munn DH, Mellor AL. IDO in the Tumor Microenvironment: Inflammation, Counter-Regulation, and Tolerance. Trends Immunol (2016) 37:193–207. doi: 10.1016/j.it.2016.01.002
59. Yao Y, Jiang Q, Jiang L, Wu J, Zhang Q, Wang J, et al. Lnc-SGK1 induced by Helicobacter pylori infection and highsalt diet promote Th2 and Th17 differentiation in human gastric cancer by SGK1/Jun B signaling. Oncotarget (2016) 7:20549–60. doi: 10.18632/oncotarget.7823
60. Collier SP, Henderson MA, Tossberg JT, Aune TM. Regulation of the Th1 genomic locus from Ifng through Tmevpg1 by T-bet. J Immunol (2014) 193:3959–65. doi: 10.4049/jimmunol.1401099
61. Hu G, Tang Q, Sharma S, Yu F, Escobar TM, Muljo SA, et al. Expression and regulation of intergenic long noncoding RNAs during T cell development and differentiation. Nat Immunol (2013) 14:1190–8. doi: 10.1038/ni.2712
62. Rotte A. Combination of CTLA-4 and PD-1 blockers for treatment of cancer. J Exp Clin Cancer Res CR (2019) 38:255. doi: 10.1186/s13046-019-1259-z
63. Gubin MM, Zhang X, Schuster H, Caron E, Ward JP, Noguchi T, et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature (2014) 515:577–81. doi: 10.1038/nature13988
64. Marmarelis ME, Aggarwal C. Combination Immunotherapy in Non-small Cell Lung Cancer. Curr Oncol Rep (2018) 20:55. doi: 10.1007/s11912-018-0697-7
65. Karczewski KJ, Dudley JT, Kukurba KR, Chen R, Butte AJ, Montgomery SB, et al. Systematic functional regulatory assessment of disease-associated variants. Proc Natl Acad Sci U States America (2013) 110:9607–12. doi: 10.1073/pnas.1219099110
66. Ahn MJ, Sun JM, Lee SH, Ahn JS, Park K. EGFR TKI combination with immunotherapy in non-small cell lung cancer. Expert Opin Drug Saf (2017) 16:465–9. doi: 10.1080/14740338.2017.1300656
Keywords: cancer, cancer immunity, lncRNA, immunotherapy, combined therapy
Citation: Wu M, Fu P, Qu L, Liu J and Lin A (2020) Long Noncoding RNAs, New Critical Regulators in Cancer Immunity. Front. Oncol. 10:550987. doi: 10.3389/fonc.2020.550987
Received: 07 July 2020; Accepted: 09 September 2020;
Published: 30 October 2020.
Edited by:
Zhi Sheng, Virginia Tech, United StatesReviewed by:
Kevin James Pridham, Virginia Tech Carilion, United StatesRobin T. Varghese, Edward Via College of Osteopathic Medicine, United States
Copyright © 2020 Wu, Fu, Qu, Liu and Lin. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Aifu Lin, bGluYWlmdUB6anUuZWR1LmNu
†These authors have contributed equally to this work
 Minjie Wu
Minjie Wu Peifen Fu
Peifen Fu Lei Qu
Lei Qu Jian Liu
Jian Liu Aifu Lin
Aifu Lin