- 1Biochemistry Department, Faculty of Science, King Abdulaziz University, Jeddah, Saudi Arabia
- 2Cancer Metabolism and Epigenetic Unit, Biochemistry Department, Faculty of Science, King Abdulaziz University, Jeddah, Saudi Arabia
- 3Cancer and Mutagenesis Research Unit, King Fahd Medical Research Centre, King Abdulaziz University, Jeddah, Saudi Arabia
Most of the cancer related deaths are caused mainly by metastasis. Therefore, it is highly important to unfold the major mechanisms governing metastasis process in cancer. Throughout the metastatic cascade, cells need the ability to survive without attachment to neighboring cells and the original Extra Cellular Matrix (ECM). Recent reports showed that loss of ECM attachment shifts cancer cell metabolism towards glycolysis mostly through hypoxia. However, AMPK, a master metabolic regulator was also found to be upregulated under ECM detached conditions. Therefore, in this work we aimed to understand the consequences of targeting AMPK and other metabolic kinases by a broad kinase inhibitor namely Compound C in ECM detached cancer cells. Results showed that Compound C impacts glycolysis as evident by increased levels of pyruvate, but reduces its conversion to lactate thereby negatively regulating the Warburg effect. Simultaneously, Compound C induces block at multiple levels in TCA cycle as evident from accumulation of various TCA metabolites. Interestingly Compound C significantly reduces glutamine and reduced glutathione levels, suggesting loss of antioxidant potential of ECM detached cancer cells. Further, we found increased in metabolites associated with nucleotide synthesis, one carbon metabolism and PPP pathway during Compound C treatment of ECM detached cells. Finally, we also found induction in metabolites associated with DNA damage in ECM detached cancer cells during Compound C treatment, suggesting DNA damage regulatory role of metabolic kinases. Overall, our results showed that Compound C represses pyruvate to lactate conversion, reduces antioxidant potential and invokes DNA damage in ECM detached cancer cells. Our data provides a comprehensive metabolic map of ECM detached cancer cells that can be targeted with a broad kinase inhibitor, is Compound C. The data can be used for designing new combinational therapies to eradicate ECM detached cancer cells.
Introduction
Cancer is the deadliest disease worldwide; it cost nearly 9.6 million lives in 2018. Most cancer deaths associated with the metastasis stage occur during cellular detachment from the primary site of cancer tumour and get attached to secondary metastatic site for growth (1, 2). Cell adhesion to the ECM-extracellular matrix that helps in cell survival and proliferation signal. In the absence of ECM or cell to cell adhesion leads cells to undergo programmed cell death (apoptosis), termed as anoikis (3). Metastatic ECM detached cells develop resistance to anoikis for its survival and re-attach to the distal secondary site to develop metastasis tumor. The signaling pathway involved in regulating cells during anoikis resistance is little known. PKB- The serine/threonine-protein kinase Akt regulates several cellular processes in tumors, including proliferation and cell metabolism (4). the AMP-activated protein kinase (AMPK) is regulated under metabolic stress conditions to maintain cell homeostasis by switching energy metabolism. AMPK inhibits anabolic pathway to minimize ATP consumption, sterol and lipid biosynthesis, glycogen synthesis, and cell cycle progression. AMPK promotes a catabolic pathway to restore ATP and increase glucose uptake, autophagy, lipid utilization, and mitochondrial biogenesis (5–7). AMPK is regulated positively by activation of Try172 residue phosphorylation by LKB1 and CaMKKβ kinase. Recent studies showed the AMPK pro-tumorigenic activity under glucose deprivation and hypoxic condition (7–10).
Glucose deprivation reduces NADPH/GSH level and increases H2O2 due to impaired PPP. It increases the non-metabolizable 2deoxyglucose(2DG) analogy of glucose, 2DG inhibits glycolysis by mimic glucose starvation and induce cell death (11). AMPK activation protects cancer cells from chemotherapy-induced apoptosis and metabolic stress. Dorsomorphin, also known as Compound C, has the property of inhibiting AMPK that directly effect on blocking metabolic action of AMPK. Dorsomorphin has been effectively anti-apoptotic action of AMPK and causes programmed cell death in many cancer cells type (12, 13).
In this study, we demonstrated the metabolic alteration of dorsomorphin (Compound C) in ECM detached different cancer cells. Interestingly we found dorsomorphin modulates metabolic fingerprinting in cancer cells.
Materials and Methods
Cell Culture
The human cancer cell line HCT116 and 22RV1 were kindly gifted from Dr. Hani (Cancer and the epigenetic unit, King Abdulaziz University, Saudi Arabia). Cells were grown at 37˚C with 5% CO2, in a Dulbecco’s Modified Eagle Medium (DMEM) with 10% fetal bovine serum (Sigma).
Matrix Detachment Model
The matrix detachment was done in cell suspension culture. 1 × 106 cells were cultured in ultra-low attachment plate (14) for at 37°C for various time points, which results in formation of spheroids. The spheroids were treated with either vehicle control or with different concentration of compound-C (sigma- P5499) for 5 days. The images were captured by using Nikon (USA) inverted light microscope. Images were analyzed for size measurement using image J software (https://imagej.net/Invasion_assay).
Apoptosis Assay
The apoptosis was detected by using Annexin V-FITC labelled and propidium iodide followed by a flow cytometer. The cells were grown in an ultra-low attachment plate for 6 days with and without treatment with Compound C. After treatment spheroids are harvested and washed with PBS (ice cold) three times. The spheroids were breakdown by multiple pipetting and resuspended in 100 μl 1X binding buffer and 10 μl Annexin V-FITC and 5 μl PI. After incubation for 20 min in RT (dark condition). The cells were analyzed by flow The Guava® easyCyte 5 Flow Cytometer, and the percentage of cells went apoptosis was calculated (15).
Metabolites Extraction
Metabolites were extracted from ECM attached and detached cells and detached cells were treated with dorsomorphin (Compound C, Sigma P5499).ECM detached spheroids were collected and crushed immediately using tissue homogenizer using a combination of ice-cold methanol: acetonitrile: water at a ratio of (2:1:1 v/v) and vortexed for 30s and incubated for 60 min at −20°C, and spin for 15 min at 13,000 rpm at 4°C.
The supernatant was collected, dried in a vacuum concentrator, and reconstituted in 200 µl of acetonitrile in 0.1% formic acid, vortexed for 5 min and centrifuge for 10 min at 13,000 rpm at 4°C. Finally, the samples were taken for LC-MS/MS analysis (15–17).
Mass Spectrometry
Samples were analyzed in LC-MS/MS LTQ XL™ linear ion trap instrument (ThermoFisher Scientific). MSn settings, full scan mode scanning range from 100 to 1000 m/z. Helium was used as buffer gas and Nitrogen was used as sheath gas for run 40 arbitrary units were set as flow rate. The capillary temperature was set at 270°C and voltage 4.0 V; spray voltage was set at −3.0 kV.
Data Analysis
The raw data file was processed using open accesses online XCMS online database. Peaks were searched against human metabolites in the Human Metabolome database. Pathway analysis and statics were Metaboanalyst (15–17).
Real-Time qPCR Analysis for mRNA Expression
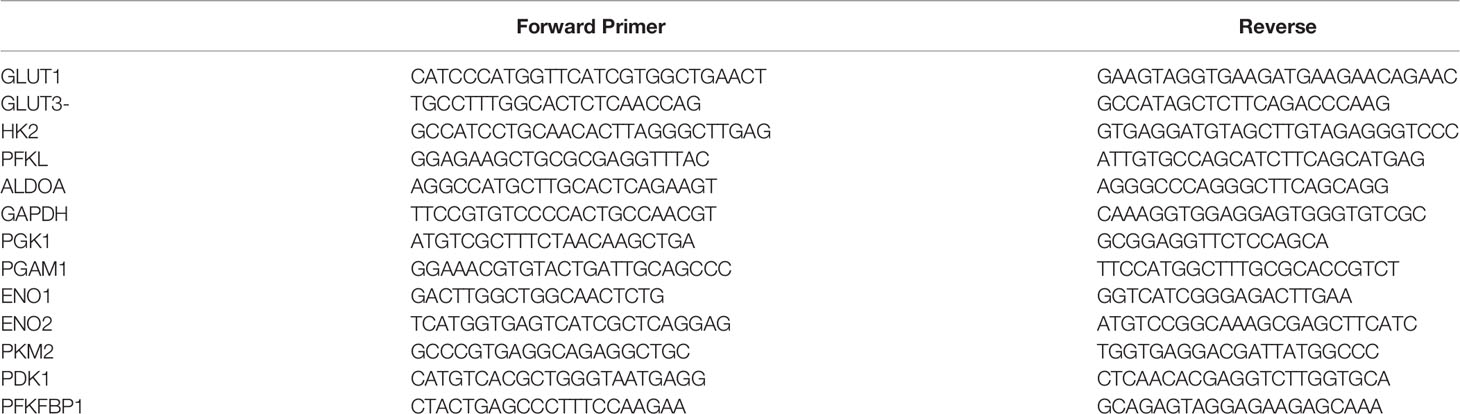
Briefly, RNA was extracted from all the cell lines at the end of different experimental conditions by using RNeasy kit (Qiagen), and reverse transcribed a High capacity cDNA Reverse Transcription kit (applied biosystems). cDNA (1–100 ng) was amplified in triplicate using gene specific primers (Supplementary Table 2). Threshold cycle (CT) values obtained from the instrument’s software were used to calculate the fold change of the respective mRNAs. ΔCT was calculated by subtracting the CT value of the housekeeping gene from that of the mRNA of interest. ΔΔCT for each mRNA was then calculated by subtracting the CT value of the control from the experimental value. Fold change was calculated by the formula 2−ΔΔCT.
Protein Extraction and Western Blot Analysis
Various cancer cells (HCT116 and 22RV1) were cultured in T75 flask (1 × 106/flask). After 24 h cells of cell plating in ultra-low attachment plates cells were treated with Compound C for indicated dose for consecutive 5 days. The fresh treatment was added every 48 h, following completion of treatment, media was aspirated and cells were washed with cold PBS (pH 7.4) and pelleted in 15ml falcon tubes. Ice-cold lysis buffer (RIPA buffer) was added to the pellet, with freshly added protease inhibitor cocktail (Protease Inhibitor Cocktail Set III, Calbiochem, La Jolla, CA). Then cells were passed through needle of the syringe to break up the cell aggregates. The lysate was cleared by centrifugation at 14000 g for 30 min at 4°C and the supernatant was used or immediately stored at −80°C. For western blotting 12% poly acrylamide gels were used to resolve 30μg of protein, transferred on to a nitrocellulose membrane, Equal loading were confirmed with ponceau staining of the membrane. Then probed with appropriate monoclonal primary antibodies and detected by chemiluminescence scanner by LI-COR after incubation with specific secondary antibodies (18). The primary antibodies as follows as follows Phospho-AMPKα (Thr172) (D4D6D), AMPKα (D5A2), LDHA (C4B5)
Results
Compound C Effectively Reduces Proliferation of ECM Detached Cancer Cells
AMPK is a key regulatory during energy deprivation and activated during ECM detached cancer cells. We examined the toxic effect of Compound C on ECM detached cancer cell proliferation. Results showed that low doses of Compound C i.e. 10 µM in HCT116 and 15µM in 22RV1 Compound C significantly reduced cell proliferation (Figure 1A). Prolong treatment of Compound C significantly reduces the spheroid formation and size (Figure 1B) and simultaneously induces apoptosis in spheroids formed due to ECM detachment (Figure 1C)
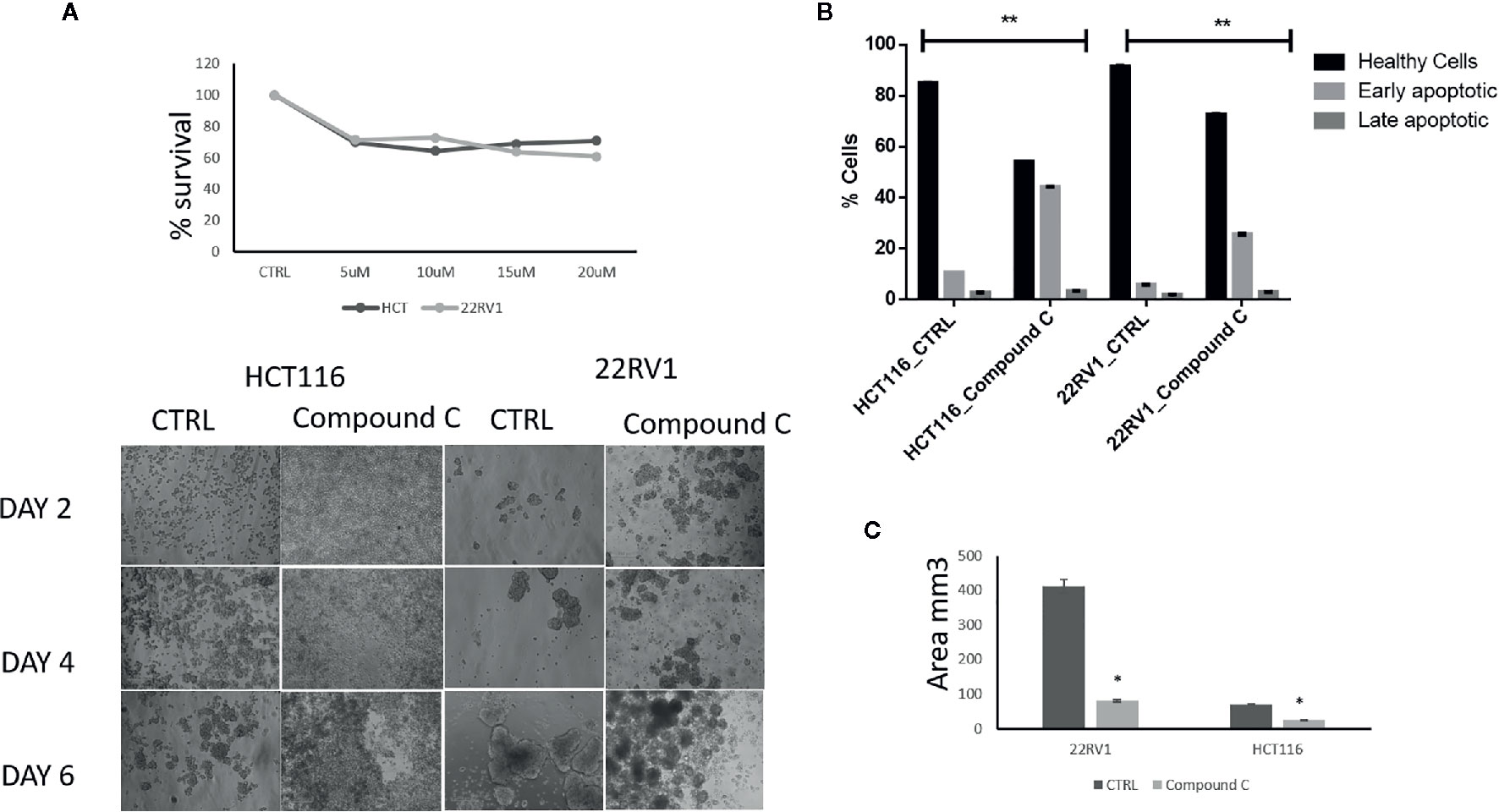
Figure 1 Compound C inhibition reduce cell proliferation and size reduction. (A) Cells are grown in ultra-low attachment 96-well plate and treated with different concentration of Compound C and MTT assay were performed. (B) Cells are grown in ultra-low attachment plate and treated with Compound C, images were acquired, and size were measured using image J software. (C) Cells are grown in ultra-low attachment plat and treated with Compound C; apoptosis assay were performed using Annexin V. * is p-value ≤ 0.05 ** is p-value ≤ 0.01
Metabolic Impact of Compound C in ECM Detached Cancer Cells
To explore the metabolic profile of ECM detached cancer cells treated with broad kinase inhibitor namely Compound C, we performed untargeted metabolomics to comprehend the difference in ECM attached cancer cells with ECM detached cancer cells, and Compound C treated ECM detached cancer cells. Metabolites were acquired using HPLC-MS/MS in DDA- Data Dependent mode, and spectrum raw files were processed using XCMS online database. The total 649 features were identified using ESI positive mode and 546 metabolites (Figure 2A) with (p-value ≤ 0.01) statistically significant. Principal component analysis (PCA) models score plots of all samples showed a significant difference in metabolomics between sample groups (Figure 2B). The correlation coefficient of the metabolomics data showed metabolites were self-correlations. However, in the Heatmap for the metabolites showed variation in metabolic expression between ECM attached, ECM detached, and ECM detached cells treated with Compound C (Figure 2C). The top highly significant metabolite in terms of PCA analysis with expression index showed the variation in metabolite accumulation in Compound C treated ECM detached cancer cells (Figure 2D), Top 275 metabolite with expression value shown in Table 1. and their raw table with p-value replicates details given in the Supplementary Table 1. The pathway enrichment analysis in ECM detached and Compound C treated ECM detached cancer cells compared with ECM attached cancer cells, pathway enrichment, and pathway linkage analysis were obtained through MetaboAnalyst 4.0. A total of 93 metabolic pathways were shown to be enriched (Figure 2E), mainly involved in energy metabolisms like Warburg Effect, Gluconeogenesis, Glycolysis, and TCA Cycle. As well as some other metabolic pathways, such as Methyl histidine Metabolism, Pentose Phosphate Pathway, Mitochondrial Electron Transport Chain, urea cycle, and Lactose Degradation, etc. (Figure 2F),
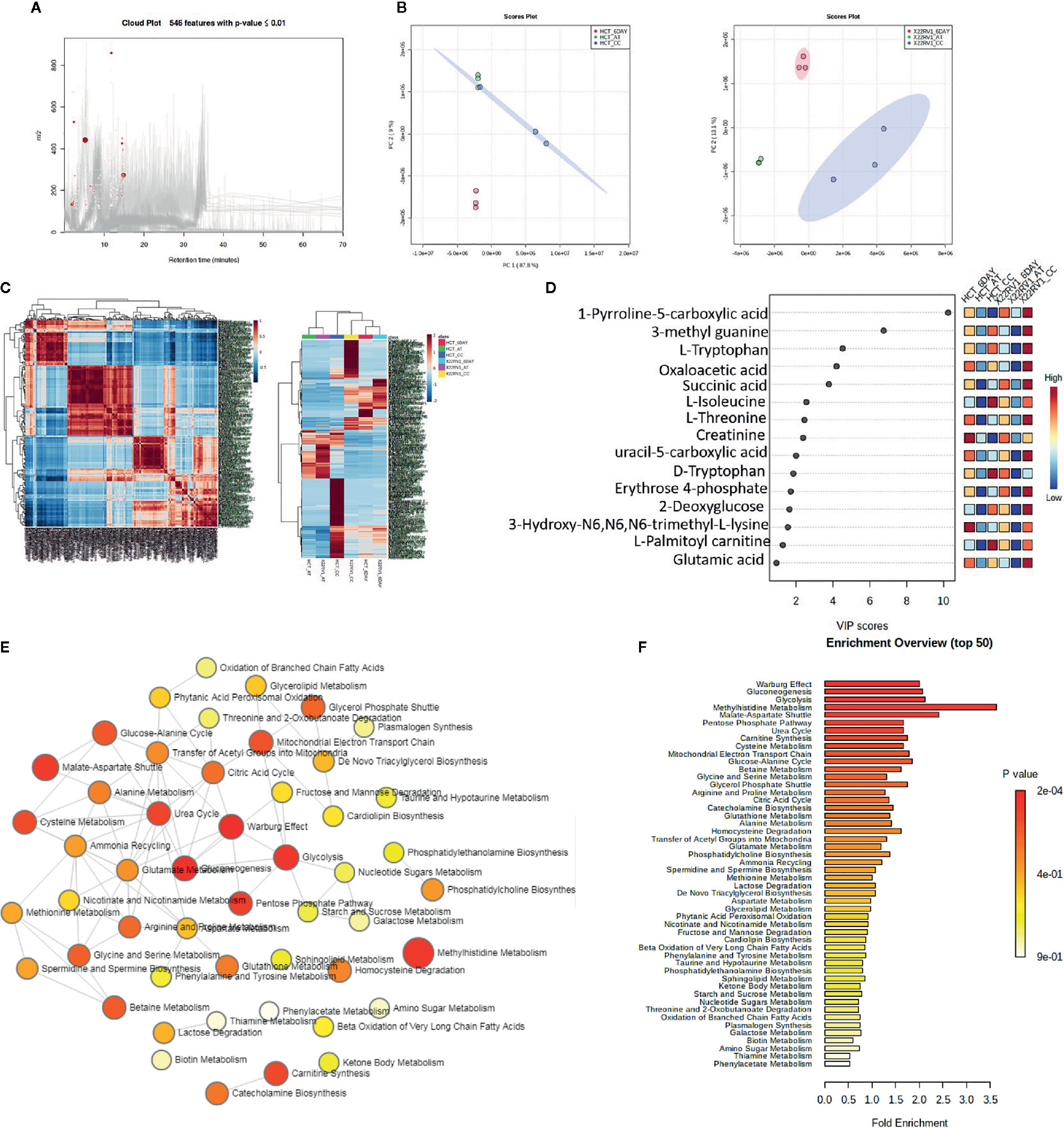
Figure 2 Metabolomic analysis of ECM detached cancer cells and Compound C treated in ECM detached cancer cells, (A) Metabolites were extracted and run in LTQ-XL linear ion trap LC-MS and its total ion chromatogram with significate features. (B) PCA analysis of total metabolite of HCT 116 and 22RV1 with three different condition. (C) Correlation heat map and expression heat map of differential metabolite expressed between ECM attached, ECM detached, and ECM detached cells treated with Compound C. (D) VIP score based on PCA analysis of top metabolites. (E) Pathway network analysis. (F) Top pathway enriched in metabolome analysis between ECM attached, ECM detached, and ECM detached cells treated with Compound C.
Compound C Reduces Lactate Formation
Lactate production in aerobic glycolysis is a prevalent energy metabolism in cancer cells metabolic pathway. We evaluate mRNA expression and metabolites involved in different steps of glycolysis. Data showed that the mRNA expression of many glycolytic gene namely GLUT1 and 3, Hexokinase 2 (HK2), PFKL (6-phosphofructokinase), PFKFB1 (fructose-2,6-biphosphatase 1), ENO2 (Enolase 2), PGAM1 (Phosphoglycerate Mutase 1), PDK1 (Pyruvate Dehydrogenase Kinase 1), LDHA (Lactate dehydrogenase A), PKM2 (pyruvate kinase M2) and their associated metabolites were high in ECM detached conditions. Compound C reduces both transcripts and associated metabolites of the above-mentioned enzymes in ECM detached cancer cells (Figures 3A, B). We further noticed that some enzyme transcript levels i.e. ALDOA (aldolase, fructose-bisphosphate A) GAPDH (glyceraldehyde 3-phosphate dehydrogenase), and PGK1- (Phosphoglycerate Kinase 1) failed to get induced in ECM detached conditions (Figures 3A, B).
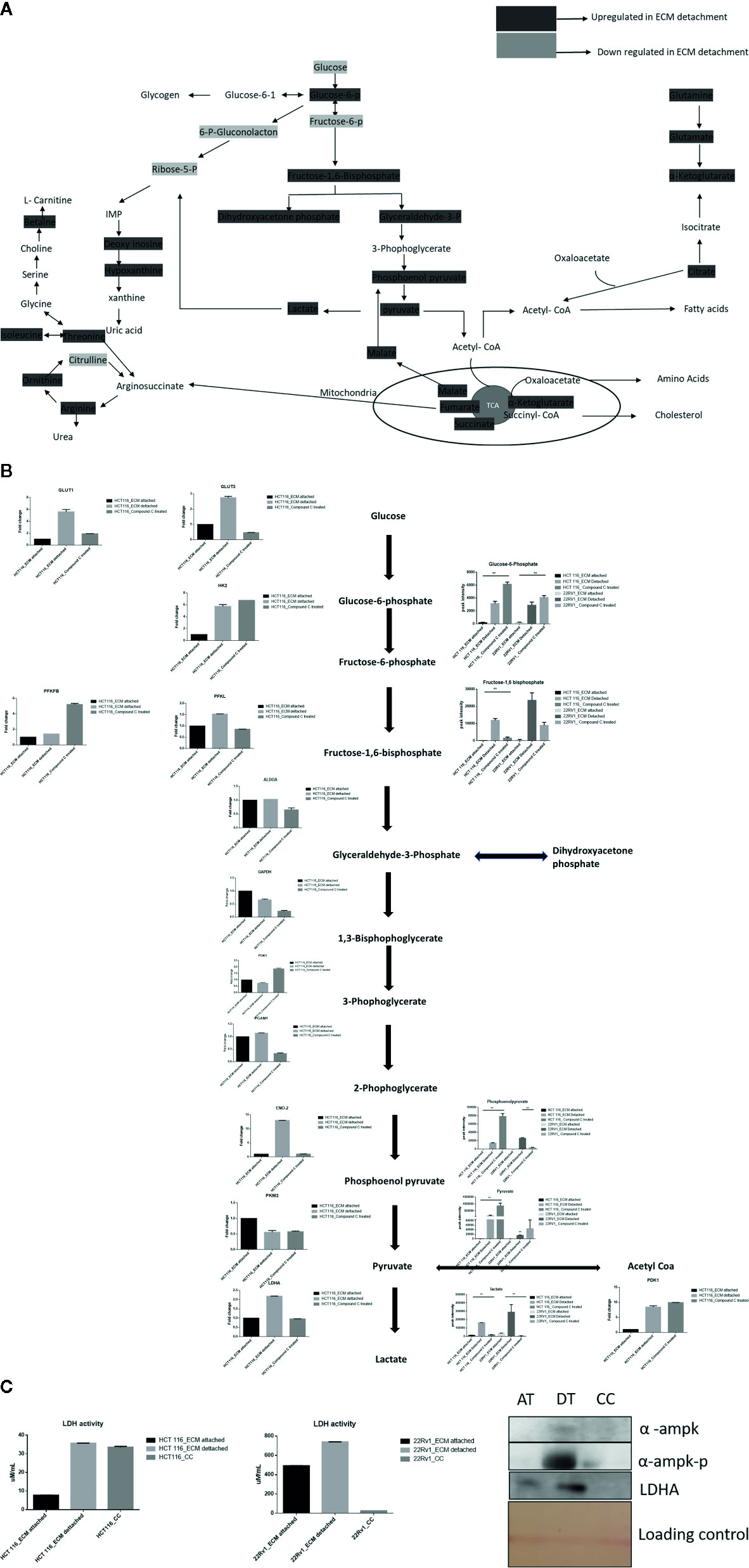
Figure 3 Compound C alter energy enrichment pathway in ECM detachment. (A) Overall energy enrichment pathway with metabolite expression index. (B) Expression of transcript (HCT116) and metabolite involved in glycolysis during ECM detachment and AMPK inhibited ECM detached cells, P-value p < 0.01. (C) LDHA assay and Western blot analysis of protein involved in AMPK activation and glycolysis(AT- ECM attached, DT- ECM detached and CC- Compound C treated).
The reduction of LDHA transcript was well corroborated with lower lactate dehydrogenase activity in Compound C treated cells (Figure 3C). These findings suggest that the end product of glycolysis i.e. pyruvate was preferred to enter oxidative phosphorylation rather than lactate formation in ECM detached cancer cells during Compound c treatment. We also observed reduction in phosphorylated-AMPK levels well-known to be targeted by Compound C in treated cells (Figure 3C).
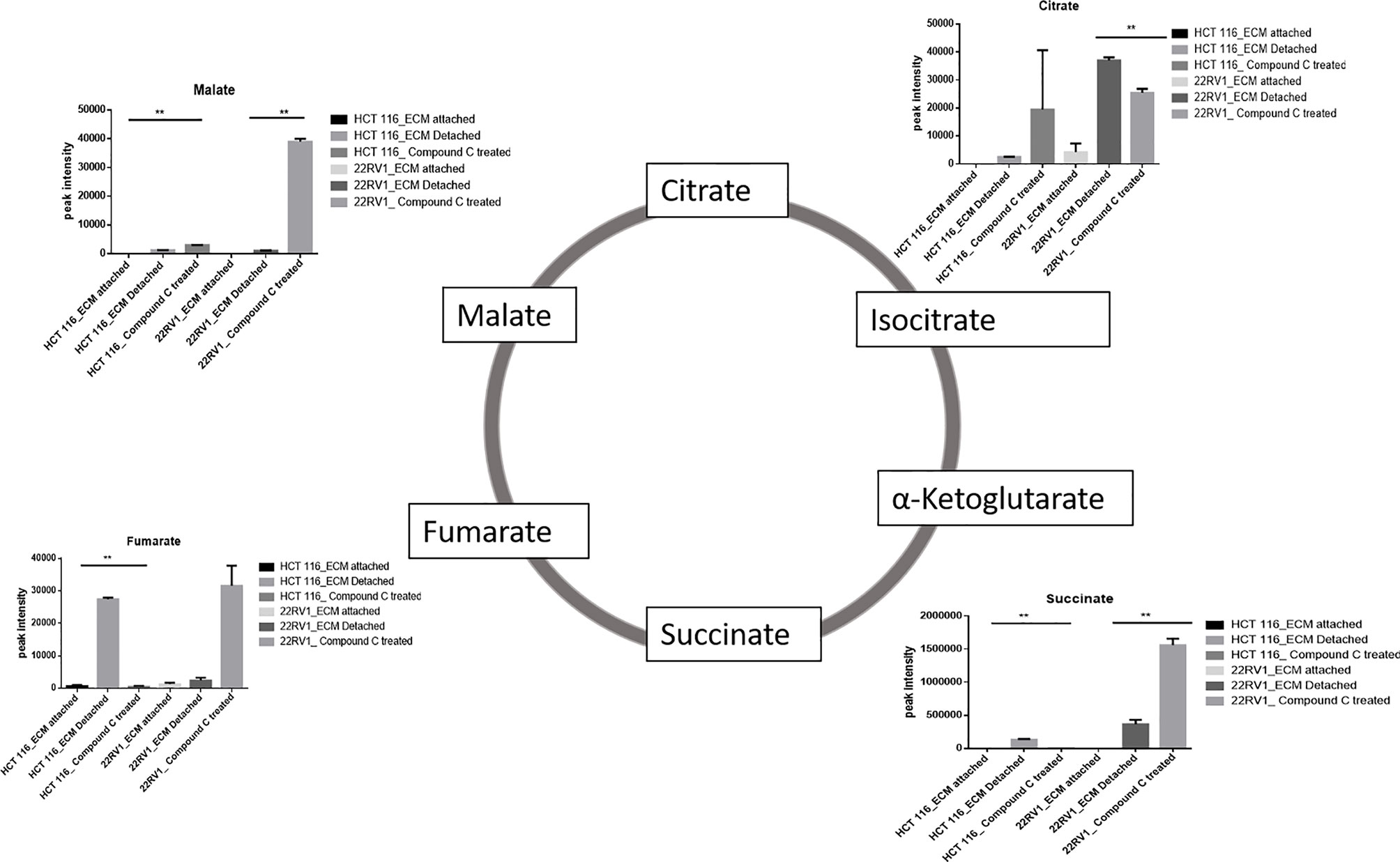
Compound C Treatment Results in Malic Acid Accumulation in ECM Detached Cells
ECM detached cancer cells need metabolic reprogramming to survive and within the known metabolic kinases, AMP-activated protein kinase (AMPK) is a primary regulator of metabolism (19). We found that ECM detached cancer cells have accumulation of various TCA intermediates such as malate and fumarate, suggesting their potential role in ECM detached cancer cell metabolism. Compound C treatment induces block at multiple steps of TCA cycle as evident by accumulation of TCA metabolites. We observed a further increase in both malate and fumarate in both cell lines treated with Compound C (Figure 4).
Compound C Depletes Glutamine and Alters Cellular Antioxidant Levels
Glutamine have multipurpose roles in cellular metabolism. Glutamine feds TCA cycle along with nucleotide biosynthesis, GSH production and production of nonessential amino acids. We found a significant increase in glutamine and hydroxyglutarate (2HG) in ECM detached cells. 2HG is an oncometabolite. 2HG is known to be produced by mutant IDH ½ enzymes and can also be generated from glutamine derived from αKG (20, 21). Compound C significantly reduces the cellular glutamine levels of ECM detached cancer cells. Since glutamine feeds for reduced glutathione (GSH) production, we measured the GSH levels and as expected ECM detachment showed increased GSH levels which were significantly reduced by Compound C (Figure 5A). Next, we measured activity of super oxide dismutase (SOD), reduces ROS, and found that ECM detached cancer cells have high activity and Compound C reduces its activity (Figure 5B). Overall, these results showed ECM detached cancer cells possess increase glutamine, GSH and antioxidant capacity and Compound C reduces glutamine, GSH and antioxidant capacity.
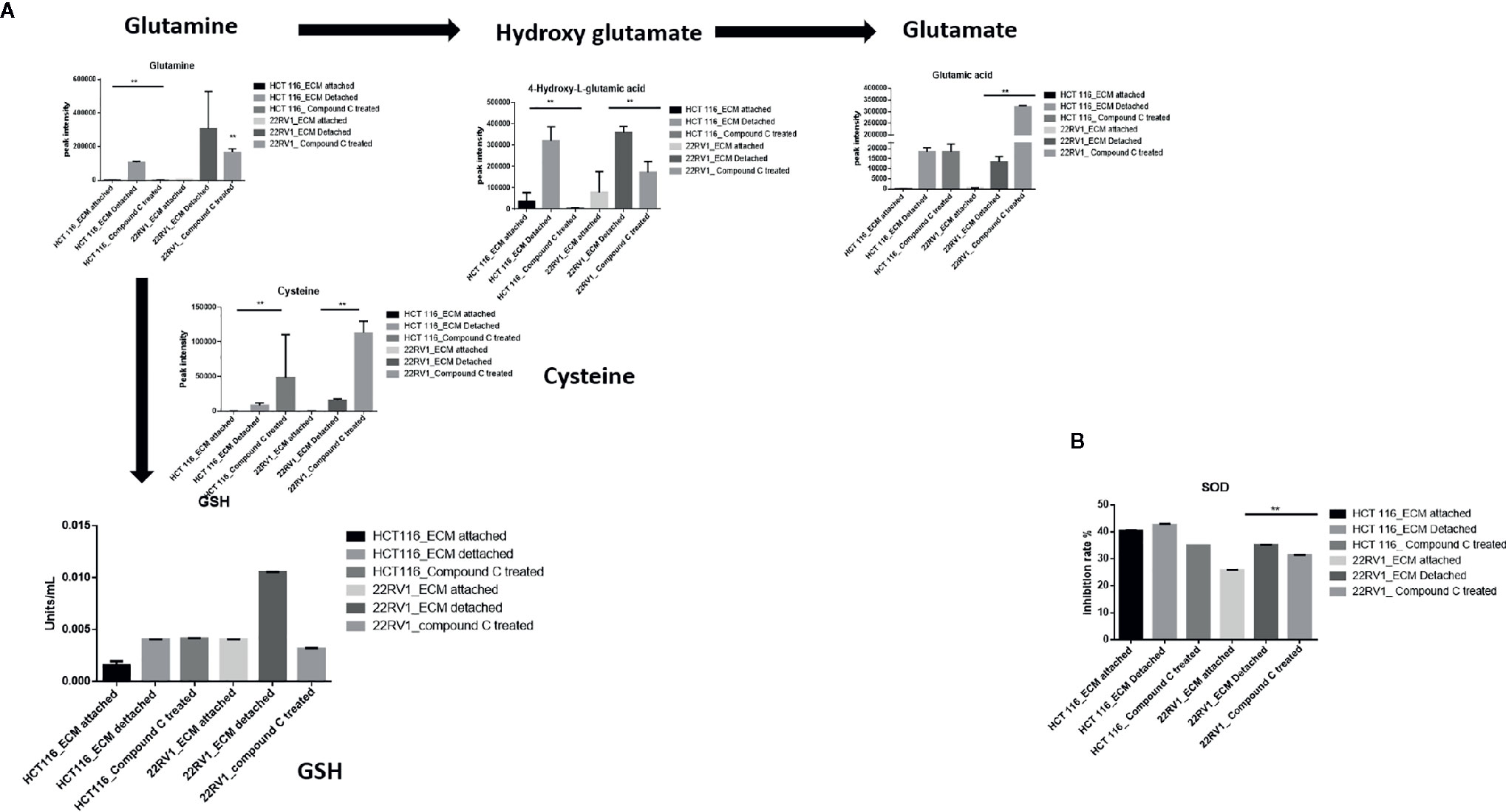
Figure 5 Compound C alters GSH level. (A) Overall GSH enrichment pathway with metabolite expression index and GSH assay. (B) SOD assay during ECM detachment and AMPK inhibited ECM detached cells, P-value **p < 0.01.
Compound C Favors Ribosugar Synthesis
Hexose monophosphate shunt (HMP), also known as Pentose phosphate pathway (PPP) or phosphogluconate pathway, starts from the branch of glycolysis and the first step of glucose metabolism. HMP activation is demonstrated in different types of cancer and in their role in association with metastasis, invasion, and angiogenesis (22, 23). Our studies showed the ECM detached cells increased intermediate metabolite of HMP shunt in both cell lines. Compound C treatment increases ribose, erythrose and xylose levels. However, there were no significant changes observed in nucleotide levels of adenosine and guanosine (Figure 6) (Table 2).
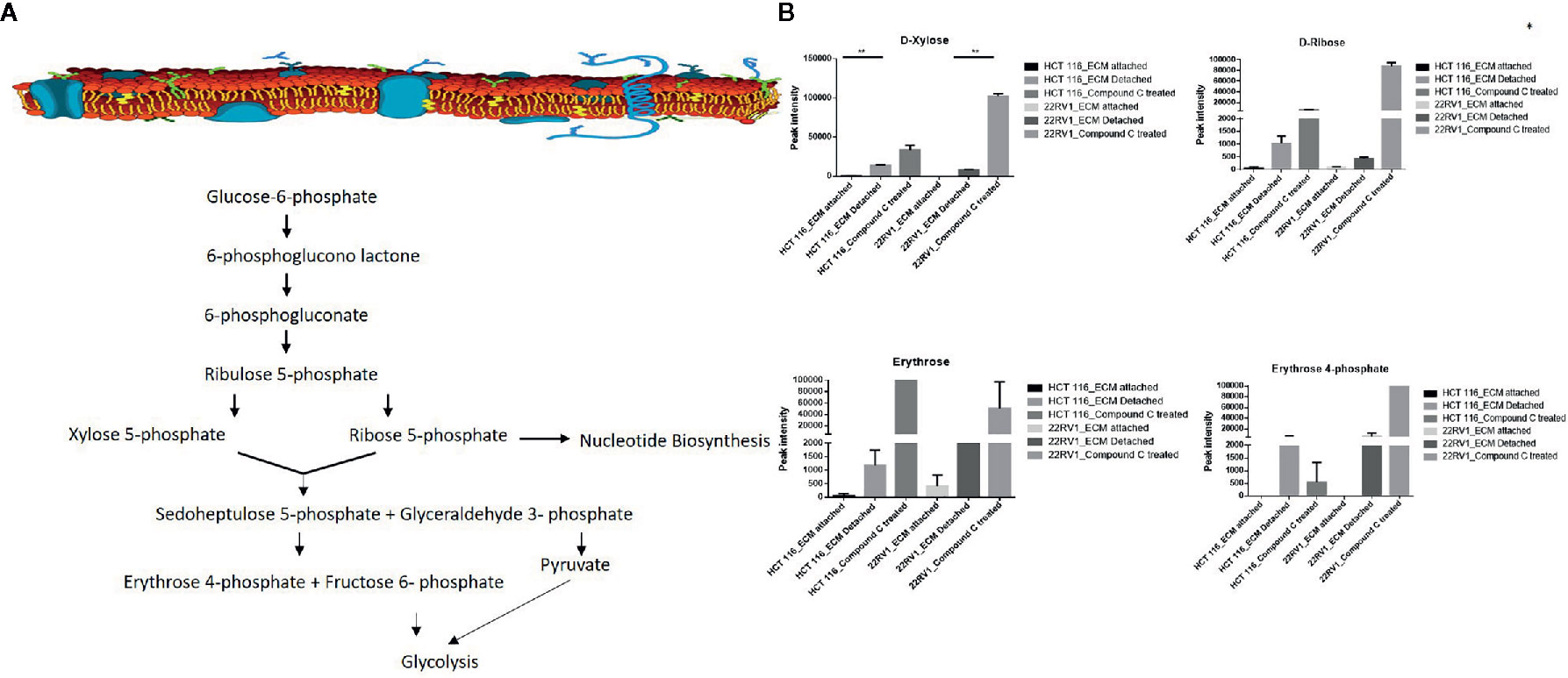
Figure 6 Compound C alter the metabolite involved pentose phosphate pathway. (A) Metabolite involved in PPP pathway. (B) Expression of metabolite involved in PPP during ECM detachment and AMPK inhibited ECM detached cells, P-value p < 0.01. * is p-value ≤ 0.05 ** is p-value ≤ 0.01
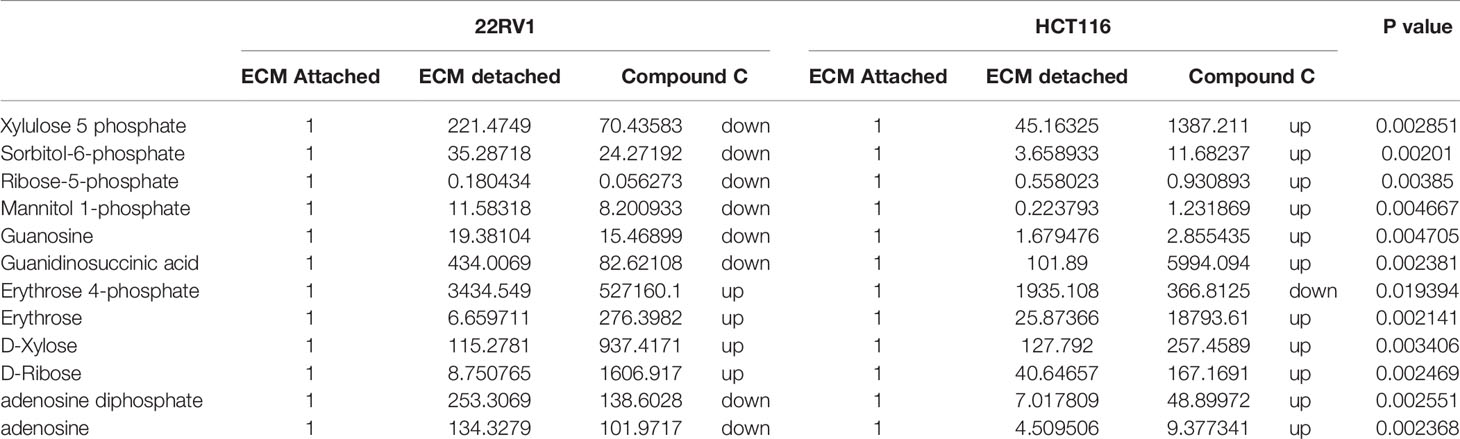
Table 2 Fold change of metabolite involved in PPP- fold change was calculated with normalized value of ECM attached and up and down regulation was mention with proposition to ECM detached and Compound C treated.
Compound C Modulates One Carbon Metabolites
One-carbon (1C) metabolism, which sets a broader set of transformation from folate metabolism. 0ne-carbon metabolism is regulatory in cellular physiology. One-carbon metabolism is mostly derived from glycine and non-essential amino acid serin. One- carbon metabolism generate output metabolites that serve as essential building blocks for a redox reaction, biosynthesis, and methylation (24). We sought to verify the functional effect on one-carbon metabolism by Compound C in detached cells; data showed increased one-carbon metabolite intermediates in ECM detached cells such as SAM, L-cystathionine, methyl-cysteine, homocysteine and cysteine suggesting that ECM detached cancer cells might have more methylation phenotype. Coherent with this observation, Compound C treatment in ECM detached cells showed further increase in S-adenosyl-methionine (SAM), homocysteine, and cysteine in both cell lines, but other intermediate metabolites showed different trends between two cell lines (Figure 7) (Table 3).
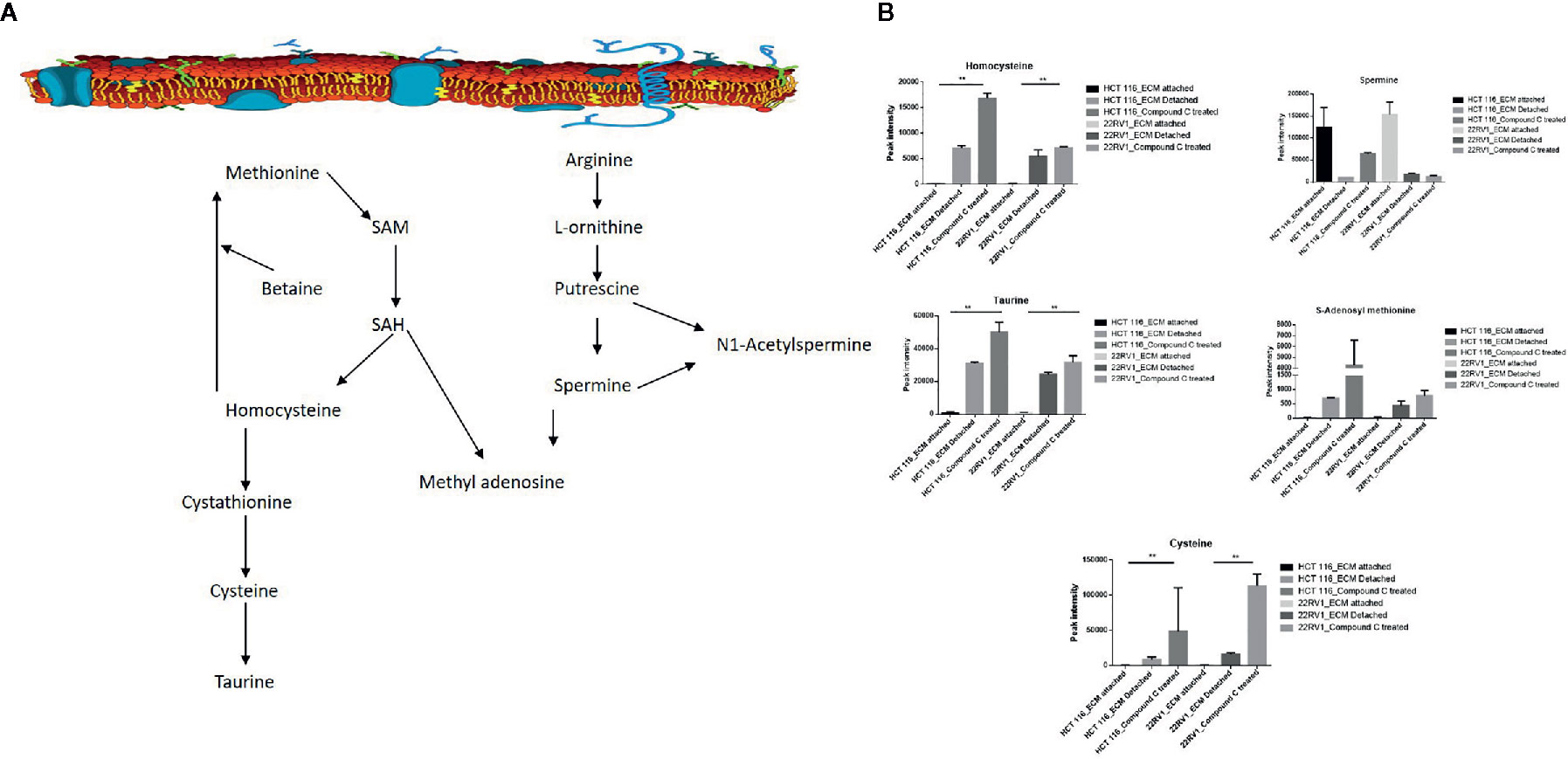
Figure 7 Compound C the modulate metabolite expression of one carbon metabolism (A) Metabolite involved in one carbon metabolism pathway. (B) Expression of metabolite involved in One carbon metabolite during ECM detachment and AMPK inhibited ECM detached cells, P-value p < 0.01. ** is p-value ≤ 0.01.
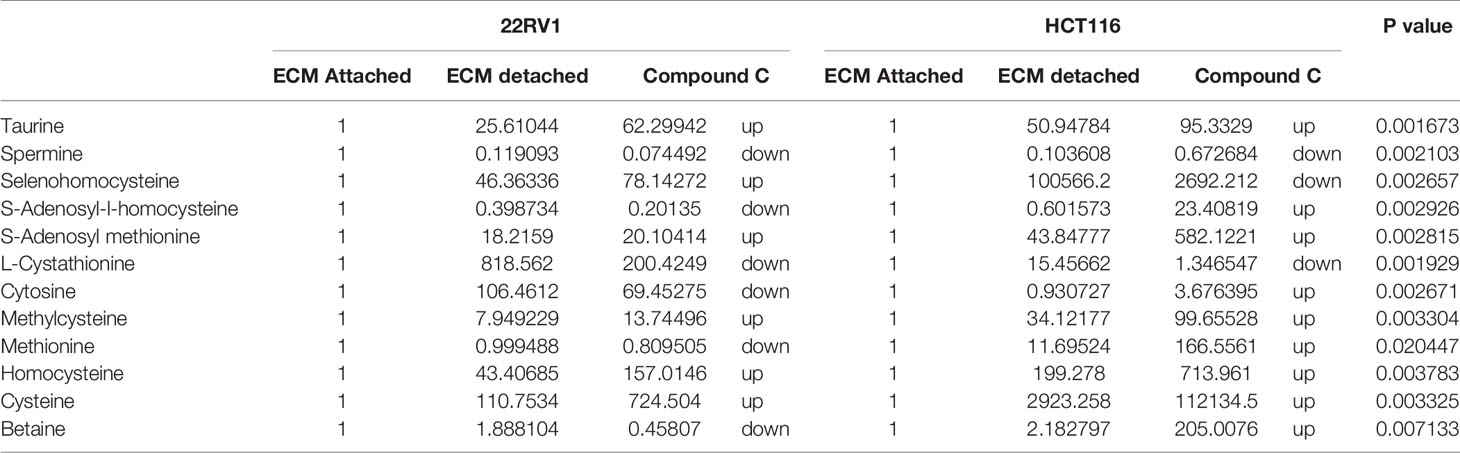
Table 3 Fold change of metabolite involved in one carbon metabolism- fold change was calculated with normalized value of ECM attached and up and down regulation was mention with proposition to ECM detached and Compound C treated.
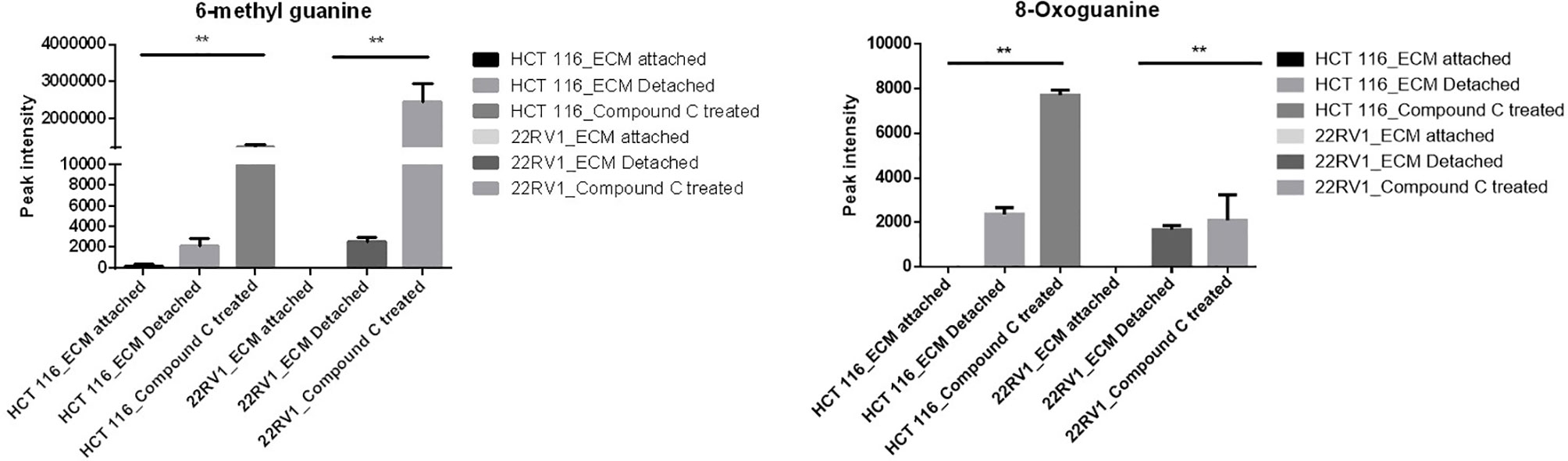
Compound C Increase Levels of Metabolites Associated With DNA Damage
Genomic DNA damage is instigated by reactive oxygen species (ROS)generated by aerobic metabolism. Both cytotoxic and mutagenic DNA lesions are associated with oxidative DNA damage (25–27). We noticed increased levels of metabolites namely 8-Oxoguanine, 3’-O-Methylguanosine, 6-methyl guanine and 3-Methylcytosine, associated with DNA damage in ECM detached cancer cells. Compound C further increased their levels suggesting a protective role of AMPK and other metabolic kinases in DNA damage conditions [Figure 8, (Table 4)].
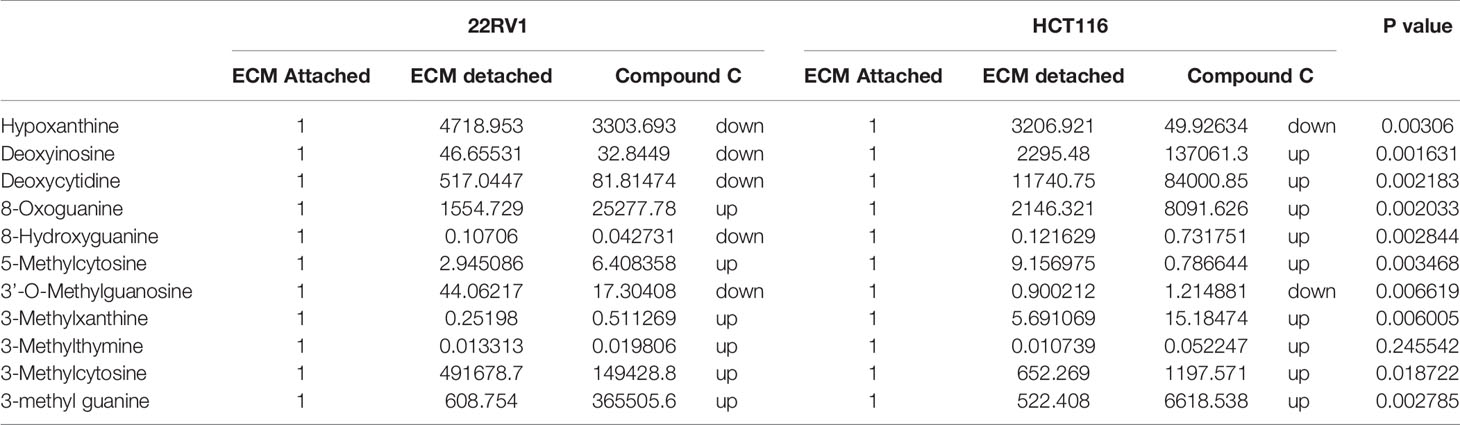
Table 4 Fold change of metabolite involved in DNA lesion - fold change was calculated with normalized value of ECM attached and up and down regulation was mention with proposition to ECM detached and Compound C treated.
Discussion
Numerous studies have shown that the metabolic reprogramming induced by ECM detachment contributes to cell survival and promotes metastasis. AMPK is a nutrient sensor activated in ECM detachment when ATP : ADP ratio is low (27). In this study, we have shown that metabolic adaption during ECM detachment and during the energy crisis. A metabolic switch happened during Compound C treatment in ECM detached cells. ECM detachment is a signature for metastasis and tumorigenesis (28). Cell depend on glucose metabolism for energy production and biosynthesis need for cell survival and proliferation (29). We showed ECM detachment increase in glycolysis in detached cells. AMPKα1, keeps the high level of reduced glutathione to maintain reduction–oxidation reaction (redox) homeostasis, AMPKα1 regulate the glutathione reductase (GSR) phosphorylation possibly through residue Thr507 which enhances its activity of enzyme (30). Our studies have shown that glutamine levels in ECM detached cells increased and supports metabolic switch reductive carboxylation for energy requirements. Reductive carboxylation is known for mitochondrial disfunction and supports cell proliferation. Our results demonstrated an Compound C was able to reduce glutamine, and other intermediate metabolites. ECM detached activates reductive carboxylation and oxidation glutamine with glycolysis, enabling biomass generation and ATP production; and maintained redox homeostasis, AMPK inhibition reduces glutamine and reduces biomass production and energy deprivation.
ECM detached cancer cells that allow glycolytic intermediate to accumulate and switch to alternate pathway HMP or PPP, which can stimulate macromolecular synthesis and combat oxidative stress (31, 32), our studies showed the ECM detachment activates PPP, intermediate metabolite accumulation was increased on ECM detached cells. The PPP is composed of two functionally interrelated branches: the oxidative and the non-oxidative metabolism. In the oxidative arm of the HMP, three major irreversible reactions involved in the reduction of glucose-6-phosphate to ribose-5-phosphate with reduction of nicotinamide adenine dinucleotide phosphate (NADPH) and ribulose-5-phosphate (Ru5P), In the synthesis of nucleotide (33–35). The non-oxidative branch of the HMP generates nucleotide synthesis from glyceraldehyde-3-phosphate and fructose-6-phosphate. Our studies show the glyceraldehyde-3-phosphate increase in 22RV1 and decrease in HCT116, in both cell lines, AMPK inhibition shows an increase in ribose sugar and erythrose involve in the salvage pathway. In this scenario, we observed that the22RV1 might involve non-oxidative and HCT116 oxidative PPP pathways during AMPK inhibition.
Several pathways are intermediate and generate one-carbon metabolism is include serine and glycine metabolism. Cancer cells show alter DNA, RNA, and histone methylation patterns. Epigenetic modification in DNA, RNA, and histone regulates various gene regulation and function. Post-translational modification (PTM) in protein methylation alter protein-protein interaction (36). Our observation showed, ECM detachment responsible for the activation of epigenetic modulator metabolite involved in one-carbon metabolism. S-adenosylmethionine (SAM) is a methyl donor; SAM transfer the methyl group to DNA and RNA, it converted to S-adenosylhomocysteine, which is converted to homocysteine and recycled to methionine. Importantly, SAM and homocysteine levels increased in ECM detached cancer cells. This observation is showing that methylation might be high in ECM detached cancer cells. The recent studies revealed that cysteine regulates the mTOR1 activity, cysteine uptake inhibits the mTOR activation and halts protein synthesis, and mTOR is regulated by AMPK. Our studies have shown the cysteine level in ECM detached and AMPK inhibited ECM detached cancer cells are increased, it is showing Compound C induces the cysteine level in ECM detached cells.
DNA lesion and DNA damage is ingested by effect of ROS production by cellular level, 8-oxoguanine is from oxidative DNA damage, effect of oxidative DNA damage results in pathophysiological changes (37). 6-methyl guanine is example for in what way DNA lesion leads to development of DNA damage. DNA damage may responsible apoptotic effect and mutagenesis properties, insertion of mutated 6-methyl guanine pass through cell cycle and causes the secondary DNA lesions and infer in DNA replication. 6-methyl guanine break the double standard DNA and triggered apoptosis (37). The data reported here reveal ECM detachment enhanced the expression of antioxidant enzyme GSH and SOD levels and inhibition of AMPK reduce antioxidant enzyme. Reduction in antioxidant enzyme activity may be increase in ROS levels and causes the DNA damage.
The significant consequence of data presented here represent advance in our understanding of ECM detached cancer cells switch the metabolic pathways., promotes cancer cell survival during metastasis. Compound C reduces pyruvate to lactate conversion, promotes OXPHOS, reduces glutamines and reduced GSH levels, and increases metabolite levels associated with DNA damage suggesting a broad metabolic impact. Beside Compound C, some other AMPK inhibitors are available like Doxorubicin, GSK6906893 and metformin. Our data also provides a detailed metabolic map of ECM detached cells, can be used to extrapolate and design future studies to identify novel targets to eradicate ECM detached cancer cells.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
Author Contributions
MRSM and RA performed the experiment and wrote the first draft of the manuscript. AA performed a flow cytometry experiment. RA and MZ guidance and proofreading of the manuscript. MRSM and HC supervised and reviewed the manuscript. MK proposed, designed, and supervised the entire study and also wrote and proofread the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This project was funded by the Deanship of Scientific Research (DSR) at King Abdulaziz University (KAU), Jeddah, under grant no. (KEP-15-130-41). The authors, therefore, acknowledge with thanks DSR technical and financial support.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors thank the Core Metabolomics Facility at the Department of Biochemistry, KAU for providing help for metabolomics study.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.612778/full#supplementary-material
Supplementary Table 1 | raw metabolite list from XCMS.
Supplementary Table 2 | Primer list.
References
1. Mehlen P, Puisieux A. Metastasis: a question of life or death. Nat Rev Cancer (2006) 6(6):449–58. doi: 10.1038/nrc1886
2. Giancotti FG, Ruoslahti E. Integrin signaling. Science (1999) 285(5430):1028–33. doi: 10.1126/science.285.5430.1028
3. Frisch SM, Francis H. Disruption of epithelial cell-matrix interactions induces apoptosis. J Cell Biol (1994) 124(4):619–26. doi: 10.1083/jcb.124.4.619
4. Vivanco I, Sawyers CL. The phosphatidylinositol 3-Kinase AKT pathway in human cancer. Nat Rev Cancer (2002) 2(7):489–501. doi: 10.1038/nrc839
5. Miranda-Saavedra D, Gabaldón T, Barton GJ, Langsley G, Doerig C. The kinomes of apicomplexan parasites. Microbes Infect (2012) 14(10):796–810. doi: 10.1016/j.micinf.2012.04.007
6. Miranda-Saavedra D, Stark MJ, Packer JC, Vivares CP, Doerig C, Barton GJ. The complement of protein kinases of the microsporidium Encephalitozoon cuniculi in relation to those of Saccharomyces cerevisiae and Schizosaccharomyces pombe. BMC Genomics (2007) 8:309. doi: 10.1186/1471-2164-8-309
7. Lin SC, Hardie DG. AMPK: Sensing Glucose as well as Cellular Energy Status. Cell Metab (2018) 27(2):299–313. doi: 10.1016/j.cmet.2017.10.009
8. Viollet B, Horman S, Leclerc J, Lantier L, Foretz M, Billaud M, et al. AMPK inhibition in health and disease. Crit Rev Biochem Mol Biol (2010) 45(4):276–95. doi: 10.3109/10409238.2010.488215
9. Jeon SM, Hay N. The double-edged sword of AMPK signaling in cancer and its therapeutic implications. Arch Pharm Res (2015) 38(3):346–57. doi: 10.1007/s12272-015-0549-z
10. Zadra G, Batista JL, Loda M. Dissecting the Dual Role of AMPK in Cancer: From Experimental to Human Studies. Mol Cancer Res (2015) 13(7):1059–72. doi: 10.1158/1541-7786.MCR-15-0068
11. Jeon SM, Chandel NS, Hay N. AMPK regulates NADPH homeostasis to promote tumour cell survival during energy stress. Nature (2012) 485(7400):661–5. doi: 10.1038/nature11066
12. Park HU, Suy S, Danner M, Dailey V, Zhang Y, Li H, et al. AMP-activated protein kinase promotes human prostate cancer cell growth and survival. Mol Cancer Ther (2009) 8(4):733–41. doi: 10.1158/1535-7163.MCT-08-0631
13. Xu ZX, Liang J, Haridas V, Gaikwad A, Connolly FP, Mills GB, et al. A plant triterpenoid, avicin D, induces autophagy by activation of AMP-activated protein kinase. Cell Death Differ (2007) 14(11):1948–57. doi: 10.1038/sj.cdd.4402207
14. Labuschagne CF, Cheung EC, Blagih J, Domart MC, Vousden KH. Cell Clustering Promotes a Metabolic Switch that Supports Metastatic Colonization. Cell Metab (2019) 30(4):720–34.e5. doi: 10.1016/j.cmet.2019.07.014
15. AlGhamdi AA, Mohammed MRS, Zamzami MA, Al-Malki AL, Qari MH, Khan MI, et al. Untargeted Metabolomics Identifies Key Metabolic Pathways Altered by Thymoquinone in Leukemic Cancer Cells. Nutrients (2020) 12(6):1792. doi: 10.3390/nu12061792
16. Hassan MA, Al-Sakkaf K, Shait Mohammed MR, Dallol A, Al-Maghrabi J, Aldahlawi A, et al. Integration of Transcriptome and Metabolome Provides Unique Insights to Pathways Associated With Obese Breast Cancer Patients. Front Oncol (2020) 10:804. doi: 10.3389/fonc.2020.00804
17. Gaboon NEA, Banaganapalli B, Nasser K, Razeeth M, Alsaedi MS, Rashidi OM, et al. Exome sequencing and metabolomic analysis of a chronic kidney disease and hearing loss patient family revealed RMND1 mutation induced sphingolipid metabolism defects. Saudi J Biol Sci (2020) 27(1):324–34. doi: 10.1016/j.sjbs.2019.10.001
18. Hardie DG. Molecular Pathways: Is AMPK a Friend or a Foe in Cancer? Clin Cancer Res (2015) 21(17):3836–40. doi: 10.1158/1078-0432.CCR-14-3300
19. Shait Mohammed MR, Krishnan S, Amrathlal RS, Jayapal JM, Namperumalsamy VP, Prajna L, et al. Local Activation of the Alternative Pathway of Complement System in Mycotic Keratitis Patient Tear. Front Cell Infect Microbiol (2020) 10:205. doi: 10.3389/fcimb.2020.00205
20. Intlekofer AM, Dematteo RG, Venneti S, Finley LW, Lu C, Judkins AR, et al. Hypoxia Induces Production of L-2-Hydroxyglutarate. Cell Metab (2015) 22(2):304–11. doi: 10.1016/j.cmet.2015.06.023
21. Jiang L, Shestov A, Swain P, Yang C, Parker SJ, Wang QA, et al. Reductive carboxylation supports redox homeostasis during anchorage-independent growth. Nature (2016) 532:255–8. doi: 10.1038/nature17393
22. Riganti C, Gazzano E, Polimeni M, Aldieri E, Ghigo D. The pentose phosphate pathway: an antioxidant defense and a crossroad in tumor cell fate. Free Radic Biol Med (2012) 53(3):421–36. doi: 10.1016/j.freeradbiomed.2012.05.006
23. Nathan C, Ding A. SnapShot: Reactive Oxygen Intermediates (ROI). Cell (2010) 140(6):951–951.e2. doi: 10.1016/j.cell.2010.03.008
24. Kalhan SC, Hanson RW. Resurgence of serine: an often neglected but indispensable amino Acid. J Biol Chem (2012) 287(24):19786–91. doi: 10.1074/jbc.R112.357194
25. Dizdaroglu M. Chemistry of free radical damage to DNA andnucleoproteins. DNA and Free Radicals. Halliwell B, Arouma OI, editors. Chichester: Ellis Horwood (1993) p. 19–39.
26. Sies H. Oxidative Stress and Antioxidants. Exp Physiol. Sies H, editor. London: Academic (1991) p. 99–116.
27. Grollman AP, Moriya M. Mutagenesis by 8-oxoguanine: an enemy within. Trends Genet (1993) 9:246–9. doi: 10.1016/0168-9525(93)90089-z
28. Davison CA, Durbin SM, Thau MR, Zellmer VR, Chapman SE, Diener J, et al. Antioxidant enzymes mediate survival of breast cancer cells deprived of extracellular matrix. Cancer Res (2013) 73(12):3704–15. doi: 10.1158/0008-5472.CAN-12-2482
29. Bissell MJ, Radisky D. Putting tumours in context. Nat Rev Cancer (2001) 1(1):46–54. doi: 10.1038/35094059
30. Wang YN, Lu YX, Liu J, Jin Y, Bi HC, Zhao Q, et al. AMPKα1 confers survival advantage of colorectal cancer cells under metabolic stress by promoting redox balance through the regulation of glutathione reductase phosphorylation. Oncogene (2020) 39(3):637–50. doi: 10.1038/s41388-019-1004-2
31. Davison CA, Chapman SE, Sasser TA, Wathen C, Diener J, Schafer ZT, et al. Multimodal optical, X-ray CT, and SPECT imaging of a mouse model of breast cancer lung metastasis. Curr Mol Med (2013) 13(3):368–76. doi: 10.2174/1566524011313030006
32. Anastasiou D, Poulogiannis G, Asara JM, Boxer MB, Jiang JK, Shen M, et al. Inhibition of pyruvate kinase M2 by reactive oxygen species contributes to cellular antioxidant responses. Science (2011) 334(6060):1278–83. doi: 10.1126/science.1211485
33. Stincone A, Prigione A, Cramer T, Wamelink MM, Campbell K, Cheung E, et al. The return of metabolism: biochemistry and physiology of the pentose phosphate pathway. Biol Rev Camb Philos Soc (2015) 90(3):927–63. doi: 10.1111/brv.12140
34. Kowalik MA, Columbano A, Perra A. Emerging Role of the Pentose Phosphate Pathway in Hepatocellular Carcinoma. Front Oncol (2017) 7:87. doi: 10.3389/fonc.2017.00087
35. Alfarouk KO, Ahmed S, Elliott RL, Benoit A, Alqahtani SS, Ibrahim ME, et al. The Pentose Phosphate Pathway Dynamics in Cancer and Its Dependency on Intracellular pH. Metabolites (2020) 10(7):285. doi: 10.3390/metabo10070285
36. Kulis M, Esteller M. DNA methylation and cancer. Adv Genet (2010) 70:27–56. doi: 10.1016/B978-0-12-380866-0.60002-2
Keywords: AMP-activated protein kinase, metabolomic analysis, extra cellular matrix detachment, compound C, oxidative phosphorylation
Citation: Shait Mohammed MR, Alghamdi RA, Alzahrani AM, Zamzami MA, Choudhry H and Khan MI (2021) Compound C, a Broad Kinase Inhibitor Alters Metabolic Fingerprinting of Extra Cellular Matrix Detached Cancer Cells. Front. Oncol. 11:612778. doi: 10.3389/fonc.2021.612778
Received: 30 September 2020; Accepted: 19 January 2021;
Published: 25 February 2021.
Edited by:
Federica Sotgia, University of Salford, United KingdomReviewed by:
Jianrong Lu, University of Florida, United StatesKhalid Omer Alfarouk, Alfarouk Biomedical Research LLC, United States
Copyright © 2021 Shait Mohammed, Alghamdi, Alzahrani, Zamzami, Choudhry and Khan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mohammad Imran Khan, bWlraGFuQGthdS5lZHUuc2E=
†These authors have contributed equally to this work
 Mohammed Razeeth Shait Mohammed
Mohammed Razeeth Shait Mohammed Raed Ahmed Alghamdi
Raed Ahmed Alghamdi Abdulaziz Musa Alzahrani
Abdulaziz Musa Alzahrani Mazin A. Zamzami
Mazin A. Zamzami Hani Choudhry
Hani Choudhry Mohammad Imran Khan
Mohammad Imran Khan