- 1Skull Base Research Center, Loghman Hakim Hospital, Shahid Beheshti University of Medical Sciences, Tehran, Iran
- 2Department of Medical Genetics, Shahid Beheshti University of Medical Sciences, Tehran, Iran
- 3Urogenital Stem Cell Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran
Neuroblastoma is one of the utmost frequent neoplasms during the first year of life. This pediatric cancer is believed to be originated during the embryonic life from the neural crest cells. Previous studies have detected several types of chromosomal aberrations in this tumor. More recent studies have emphasized on expression profiling of neuroblastoma samples to identify the dysregulated genes in this type of cancer. Non-coding RNAs are among the mostly dysregulated genes in this type of cancer. Such dysregulation has been associated with a number of chromosomal aberrations that are frequently detected in neuroblastoma. In this study, we explain the role of non-coding transcripts in the malignant transformation in neuroblastoma and their role as biomarkers for this pediatric cancer.
Introduction
Neuroblastoma is a neoplasm originated from the neural crest of the sympathetic part of autonomic system (1) during the embryonic life (2). This malignancy is among the most common childhood cancers particularly during the first year of life (3). Neuroblastoma has a heterogeneous course in terms of both pathobiology and clinical manifestations. Several therapeutic options such as surgical removal of the tumor, chemotherapy, radiotherapy, and bone marrow transplantation are being applied for neuroblastoma (4). Spontaneous regression might also happen in the course of neuroblastoma (5). This tumor is associated with several genetic and chromosomal abnormalities that affect its clinical course and prognosis namely MYCN amplification, loss of distal portion of chromosome (chr) 1p and gain of 17q (6). Other chromosomal abnormalities detected in neuroblastoma are loss of 11q, 3p, 4p, 9p, 14q, and gain of 1q, 7q, 2p, and 11p (7–9). In addition to these chromosomal aberrations, dysregulation of several genes including non-coding RNAs (ncRNAs) are linked with this cancer (10). These kinds of transcripts have regulatory impact on other genes, hence constructing an epigenetic layer of gene regulation. They are classified based on their sizes to long non-coding RNAs (lncRNAs) and microRNAs (miRNAs) with the former having more than 200 nucleotides and the latter being about 22 nucleotides (11). Based on the speculation stated by the ENCODE consortium regarding the recognition of “biochemical functions for 80% of the genome” (12), ncRNAs have attained much attention during the recent decade particularly in the field of cancer research. In the current study, we explain the role of lncRNAs and miRNAs in the evolution of neuroblastoma and their role as biomarkers for this pediatric cancer.
Dysregulated miRNAs in Neuroblastoma
Chen and Stallings have measured expression of 157 miRNAs in neuroblastoma samples. They have displayed differential pattern of 32 miRNAs between tumor with favorable prognosis and those with poor prognosis. Notably, several of these miRNAs were down-regulated in neuroblastoma samples harboring MYCN amplification, which was associated with unfavorable outcome. Cell line studies have shown the role of retinoic acid in the modulation of expression of miRNAs in a MYCN-amplified cell line. Among the dysregulated miRNAs has been miR-184 which participates in the regulation of apoptosis. MYCN might exert its tumorigenic effects via modulating expression of miRNAs that participate in neural cell differentiation or apoptotic processes (13).
Among the firstly discovered tumor suppressor miRNAs in neuroblastoma was miR-34a (14), which is transcribed from a frequently deleted region in neuroblastoma i.e. 1p36.23. This miRNA was particularly down-regulated in neuroblastoma samples with 1p deletion (14). Since miR-34a inhibits expression of the E2F3 transcription factor, its down-regulation facilitates cell cycle progression (14). Subsequent studies have also verified the tumor suppressive impact of miR-34a in the neuroblastoma cells and its inhibitory effects on the expression of BCL2 and MYCN (15, 16). miR-34a also binds with the 3’ UTR of ATG5 and CD44 transcripts and decreases their expressions. Down-regulation of miR-34a in neuroblastoma cells results in the over-expression of ATG5 and CD44 (17, 18). CD44 is a cell surface receptor which can bind with hyaluronan and induce expression of genes that promote progression of cancer (19). ATG5 can dissociate V1V0-ATPase, increase pH in multivesicular bodies and enhance secretion of exosomes to facilitate cancer metastasis (20). Thus, miR-34a affects the progression of neuroblastoma through different mechanisms. Figure 1 shows some aspects of participation of miR-34a in the pathogenesis of neuroblastoma.
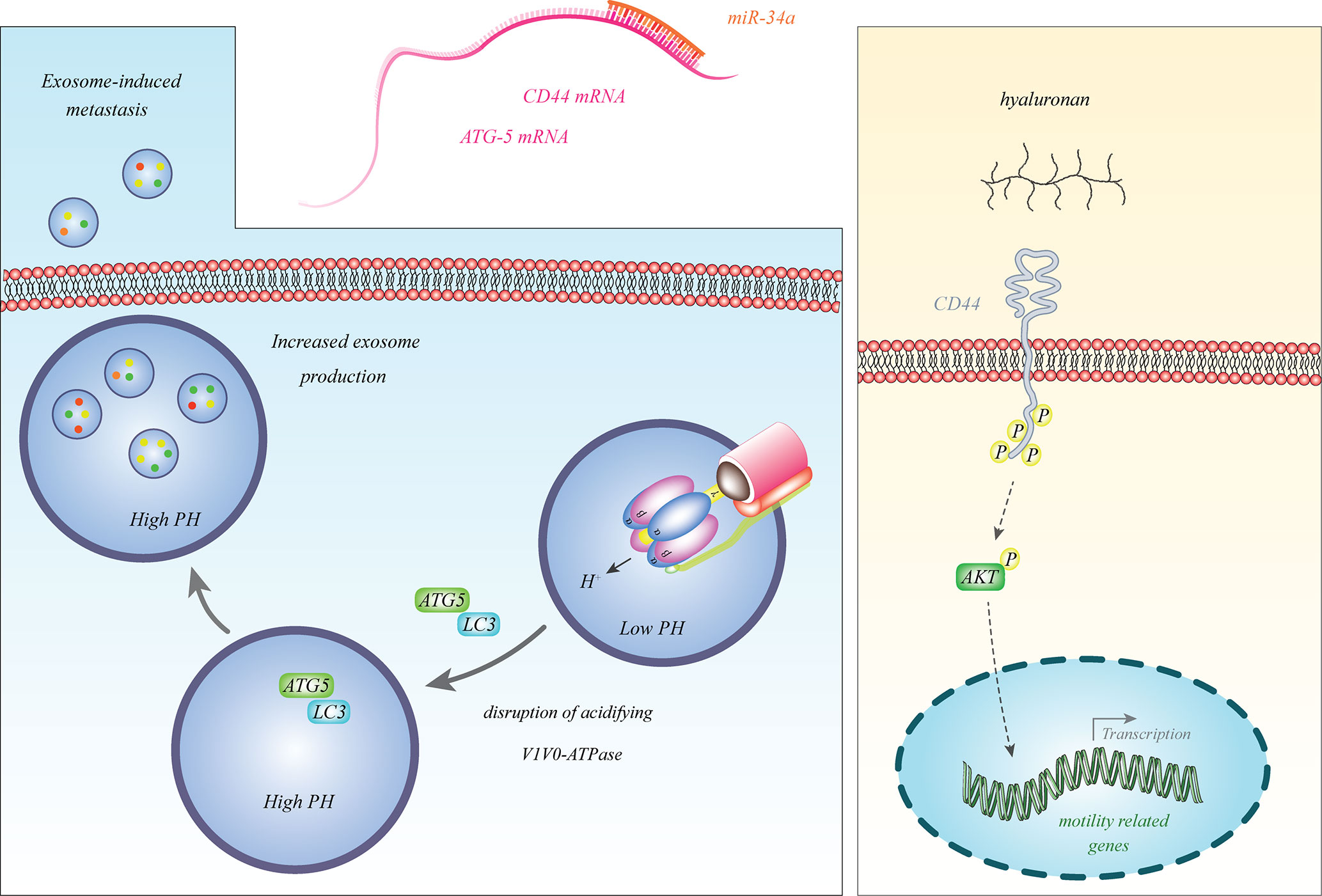
Figure 1 miR-34a binds with the 3’ UTR of ATG5 and CD44 transcripts to reduce their expressions. Decreased expression of miR-34a in neuroblastoma leads to over-expression of ATG5 and CD44 (17, 18). CD44 is a cell surface receptor which can bind with hyaluronan and induce expression of genes that promote progression of cancer (19). ATG5 can dissociate V1V0-ATPase, increase pH in multivesicular bodies and enhance secretion of exosomes to promote cancer metastasis (20).
miR-542-5p is another tumor suppressor miRNA whose down-regulation in neuroblastoma has conferred poor clinical outcome. Notably, forced up-regulation of miR-542-5p has resulted in attenuation of neuroblastoma invasive properties and tumor growth bot in vitro and in vivo (21). Moreover, expression of miR-490-5p has been diminished in neuroblastoma tissues and cells. Forced overexpression of miR-490-5p has diminished cell proliferation migration and invasiveness, prompted G0/G1 arrest in cells and induced cell apoptosis. MYEOV has been confirmed to be the target of miR-490-5p through which miR-490-5p blocks neuroblastoma progression (22). Table 1 recapitulates the results of studies which described down-regulation of miRNAs in neuroblastoma.
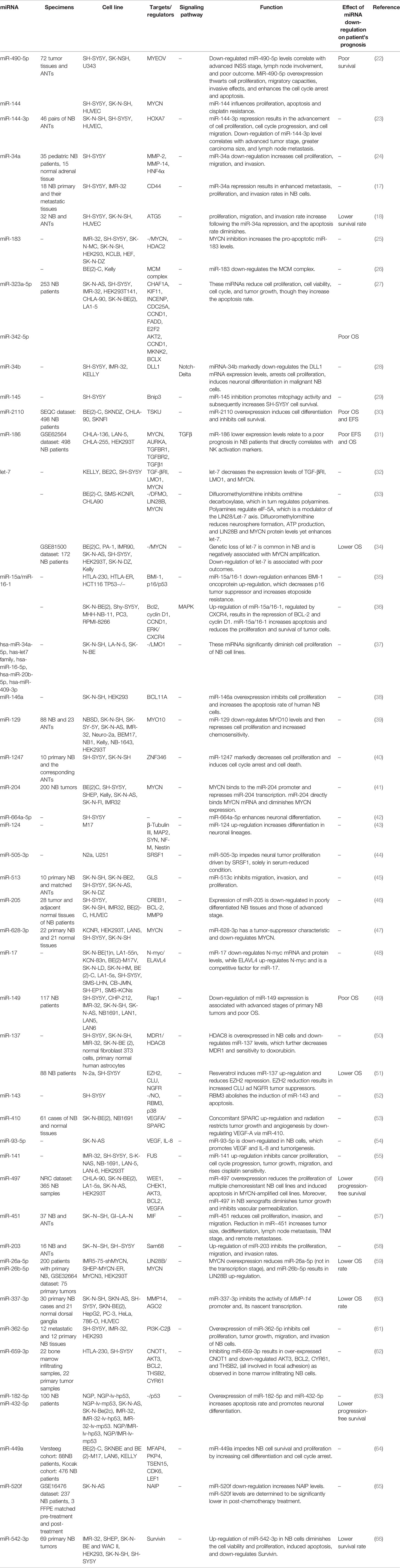
Table 1 Down-regulated miRNA in neuroblastoma (NB, Neuroblastoma; ANT, adjacent normal tissues; OS, overall survival; EFS, event-free survival).
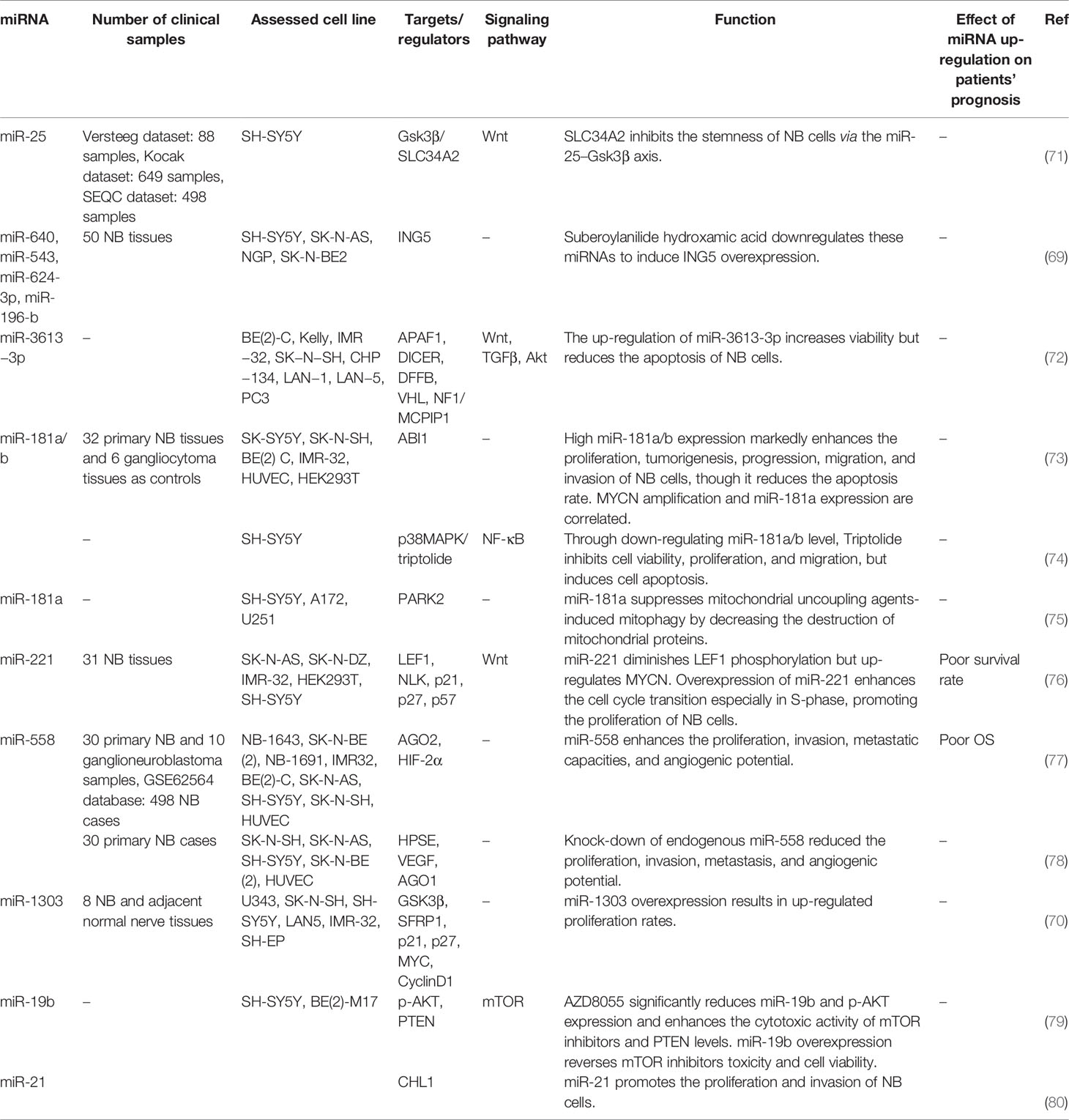
Schulte et al. have identified seven miRNAs whose expressions have been increased by MYCN in vitro and are over-expressed in primary neuroblastomas that harbor MYCN amplification. Notably, three of them were from the miR-106a and miR-17 clusters whose expressions are controlled by c-Myc. They also demonstrated up-regulation of miR-221 by MYCN in neuroblastoma (67). Montana et al. have shown transactivation of the miRNA 17-5p-92 cluster by MYCN. These miRNAs have further been demonstrated to suppress expression of p21 and BIM, thus influencing cell cycle transition and apoptosis, respectively. Notably, forced up-regulation of miRNA 17-5p-92 cluster in neuroblastoma cell lines that do not harbor MYCN amplification enhances their tumorigenic potential in animal models. On the other hand, suppression of miR-17-5p attenuates the proliferation of MYCN-amplified neuroblastoma cells via up-regulation of p21 and BIM. Over-expression of miR-17-5p has also been verified in primary neuroblastoma patients especially those with MYCN amplification and poor clinical outcome (68). miR‐640, miR‐543, miR‐624‐3p, and miR‐196‐b are among up-regulated miRNAs in neuroblastoma. Notably, these miRNAs target ING5 transcript. miRNA‐ING5‐histone acetylation axis has been recognized as the main route through which two anti-cancer drugs namely a histone deacetylase inhibitor and a proteasome inhibitor block progression of neuroblastoma (69). miR-1303 is another over-expressed miRNA in neuroblastoma. Up-regulation of this miRNA enhanced proliferation of neuroblastoma cells through targeting GSK3β and SFRP1. miR-1303 also increased levels of MYC and CyclinD1, and diminished p21 and p27 levels (70). Table 2 lists up-regulated miRNAs in neuroblastoma.
Aberrant expression of miRNAs in neuroblastoma samples can be used as biomarkers for prediction of the course of malignancy. For instance, down-regulation of miR-490-5p has been correlated with INSS stage, lymph node involvement, and poor clinical outcome of patients with neuroblastoma (22). Similarly, decreased expression of miR-186, let-7, miR-497 and miR-432-5p predicts lower survival rates (31, 34, 56, 63). Table 3 reviews the results of studies which evaluated this aspect of miRNAs.
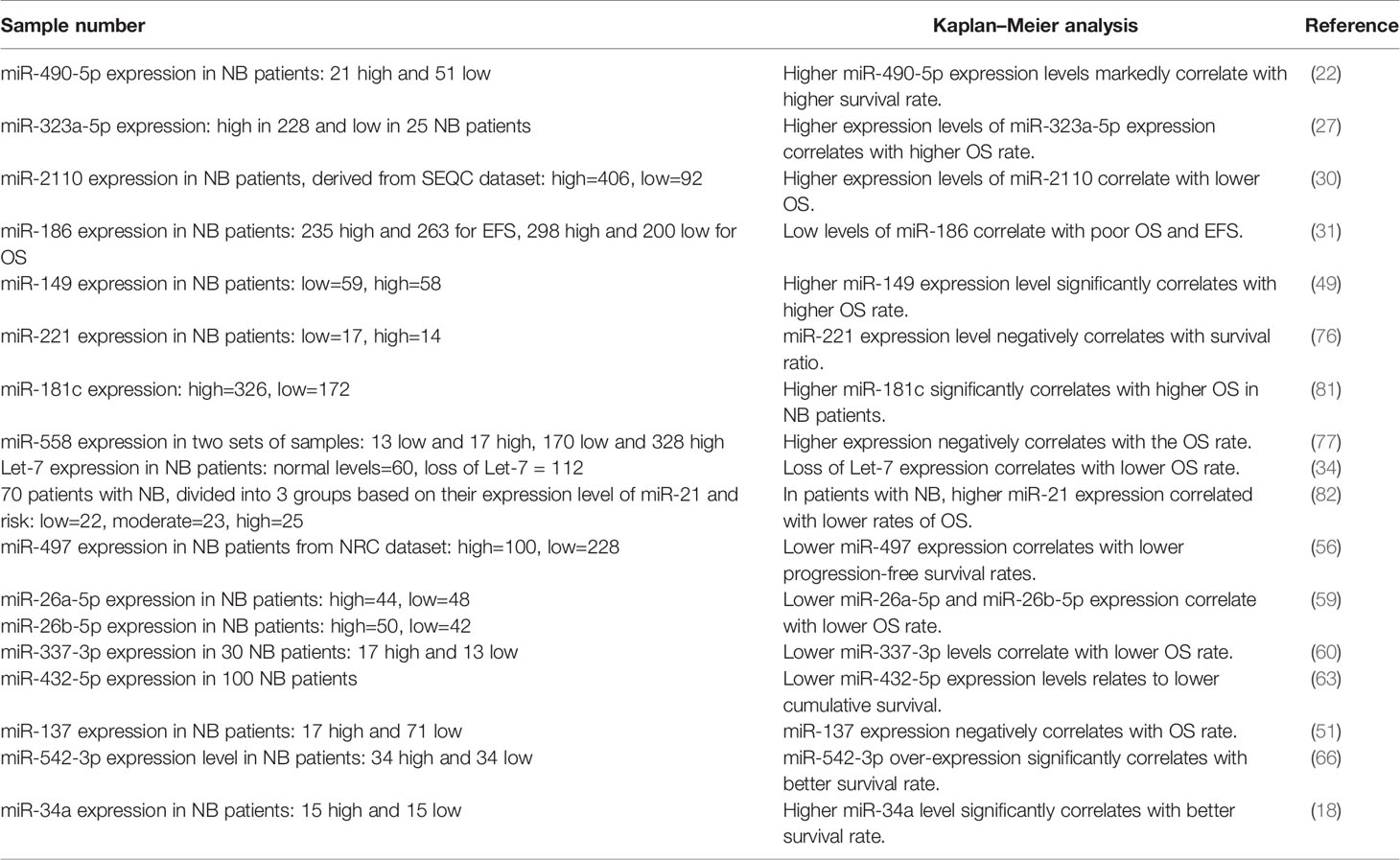
Table 3 Diagnostic importance of miRNAs in neuroblastoma (NB, neuroblastoma; OS, overall survival; EFS, event-free survival).
Dysregulated lncRNAs in Neuroblastoma
LncRNAs can regulate expression of genes via different mechanisms including alterations in chromatin configuration, modulation of transcription, splicing, mRNA stability and bioavailability as well as post-translational modifications (83). Therefore, they contribute in the pathogenesis of human cancers. Prajapati et al. have analyzed RNA-seq data of a number of neuroblastoma samples to recognize their differential expression in among primary neuroblastoma, relapsed ones and metastasized tumors. They reported up-regulation of RFPL1S, PPP1R26-AS1, RP11-439E19.3, CASC15, AC004540.5, and CTD-2881E23.2 while down-regulation of USP3-AS1, CHRM3-AS2 and RP6-99M1.2 in tumor cells compared with the corresponding non-tumor mononuclear cells isolated from bone marrow (MNCs). Moreover, expression of theses up-regulated lncRNAs along with ZRANB2-AS2 and LINC00511 were increased in the disseminated tumor cells (DTCs) compared with the corresponding MNCs. They suggested CASC15, PPP1R26-AS1, and USP3-AS1 lncRNAs as putative markers in clinical investigations in this type of pediatric cancer (84). Pandey et al. have assessed transcript signature of low-risk and high-risk neuroblastoma samples. They have reported association between a certain lncRNA namely neuroblastoma associated transcript-1 (NBAT-1) and prognosis of neuroblastoma. Altered expression of this lncRNA between the mentioned groups of neuroblastoma has been attributed to CpG methylation and the presence of a certain functional polymorphism on chr 6p22. Mechanistically, NBAT-1 down-regulation enhances proliferation and invasion of neuroblastoma cells through suppression of expression of target genes as well as induction of expression of neuronal-specific transcription factor NRSF/REST (85). Liu et al. have reported co-amplification of the lncUSMycN with MYCN in a portion of human neuroblastoma samples. This lncRNA has been shown to bind with the RNA-binding protein NonO, resulting in N-Myc up-regulation (86). Barnhill et al. have revealed that low levels of CAI2 expression in normal tissues in spite of its over-expression in the majority of tumor cell lines with a normal 9p21 locus. This lncRNA has been suggested to modulate expression of p16 and/or ARF. CAI2 expression has been higher in advanced-stage neuroblastomas in an independent manner from MYCN amplification (87). Watters et al. have shown modulation of expression of several transcribed Ultra-conserved regions (T-UCRs) in response to all-trans-retinoic acid (ATRA). Among these transcripts has been the lncRNA T-UC.300A which has imperative impacts in the regulation of cell proliferation, invasion and the suppression of differentiation of neuroblastoma cells before exposure to ATRA (88). Yu et al. have identified a transcript which has been over-expressed in neuroblastoma and named it the non-coding RNA expressed in aggressive neuroblastoma (ncRAN). Over-expression of this transcript has been associated with poor survival of patients. This lncRNA has been mapped to the region of 17q which is amplified in neuroblastoma and exerts oncogenic effects in this type of cancer (89). Tables 4 and 5 enlist over-expression and decreased expression lncRNAs in neuroblastoma, respectively.
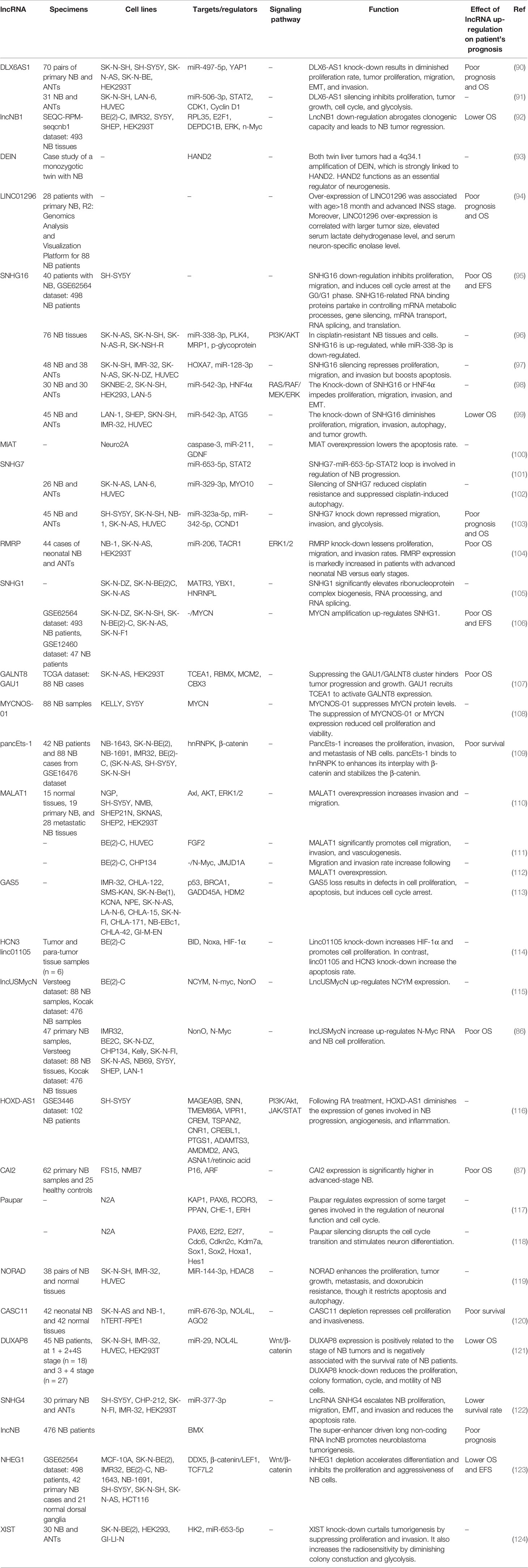
Table 4 Up-regulated lncRNAs in neuroblastoma (ANT, adjacent normal tissue; NB, Neuroblastoma; EMT, epithelial-mesenchymal transition; OS, overall survival; EFS, event-free survival).
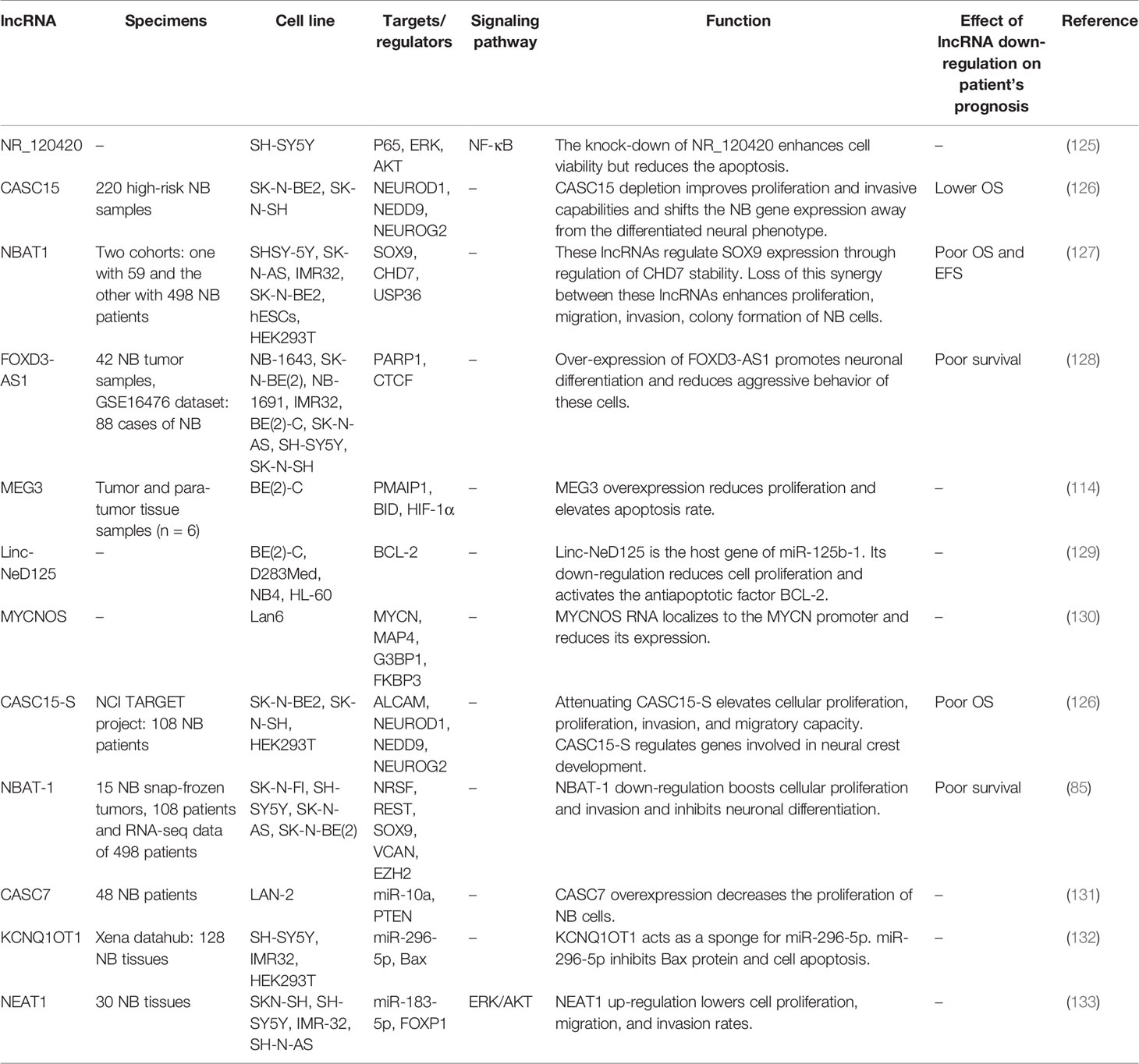
Table 5 Down-regulated lncRNAs in neuroblastoma (NB, Neuroblastoma; OS, overall survival; EFS, event-free survival).
Dysregulation of several lncRNAs in neuroblastoma samples has been correlated with survival of patients. For instance, high levels of DLX6-AS1, lncNB1, LINC01296, SNHG16 and RMRP expression have been linked with poor prognosis and lower survival (90, 92, 94, 95, 104). Table 6 summarizes the results of studies which assessed correlation between expression levels of lncRNAs and survival of patients with neuroblastoma.

Table 6 Prognostic value of lncRNAs in neuroblastoma (NB, neuroblastoma; OS, overall survival; EFS, event-free survival).
Expression and Function of circRNAs in Neuroblastoma
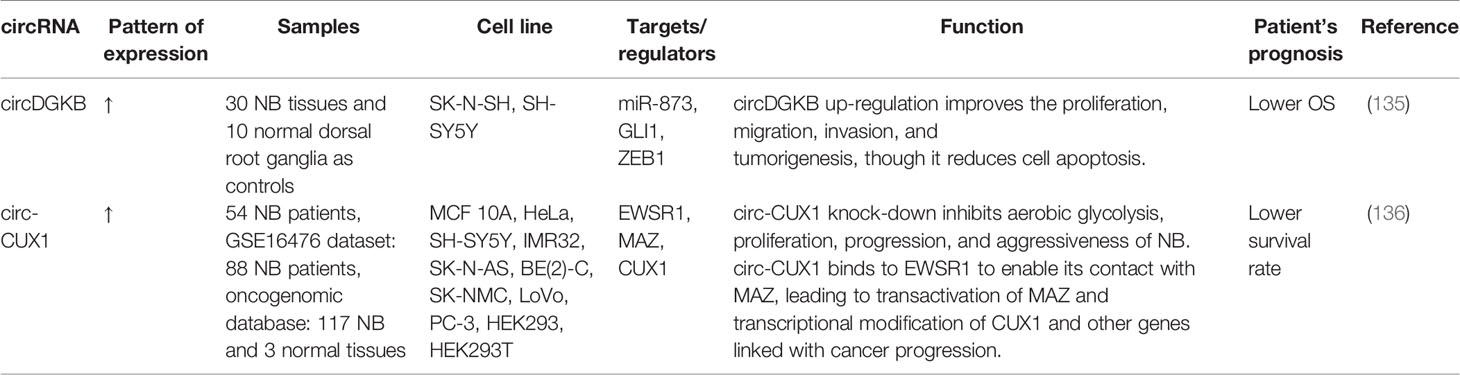
Circular RNAs (circRNAs) constitute a group of ncRNAs which are produced from exons or introns through construction of covalently-closed circles (134). Recent studies have shown dysregulation of this type of ncRNAs in cancers. For instance, circDGKB has been shown to be over-expressed in neuroblastoma tissues versus normal dorsal root ganglia. Notably, over-expression of this circRNA has been an indicator of poor survival of these patients. Mechanistically, circDGKB enhances cell proliferation, migration and invasion of neuroblastoma cells while inhibiting cell apoptosis. Moreover, up-regulation of circDGKB reduced expression level of miR-873 and increased GLI1 expression (135). Table 7 recapitulates the results of studies which assessed function of circRNAs in neuroblastoma.
Polymorphisms Within ncRNAs and Risk of Neuroblastoma
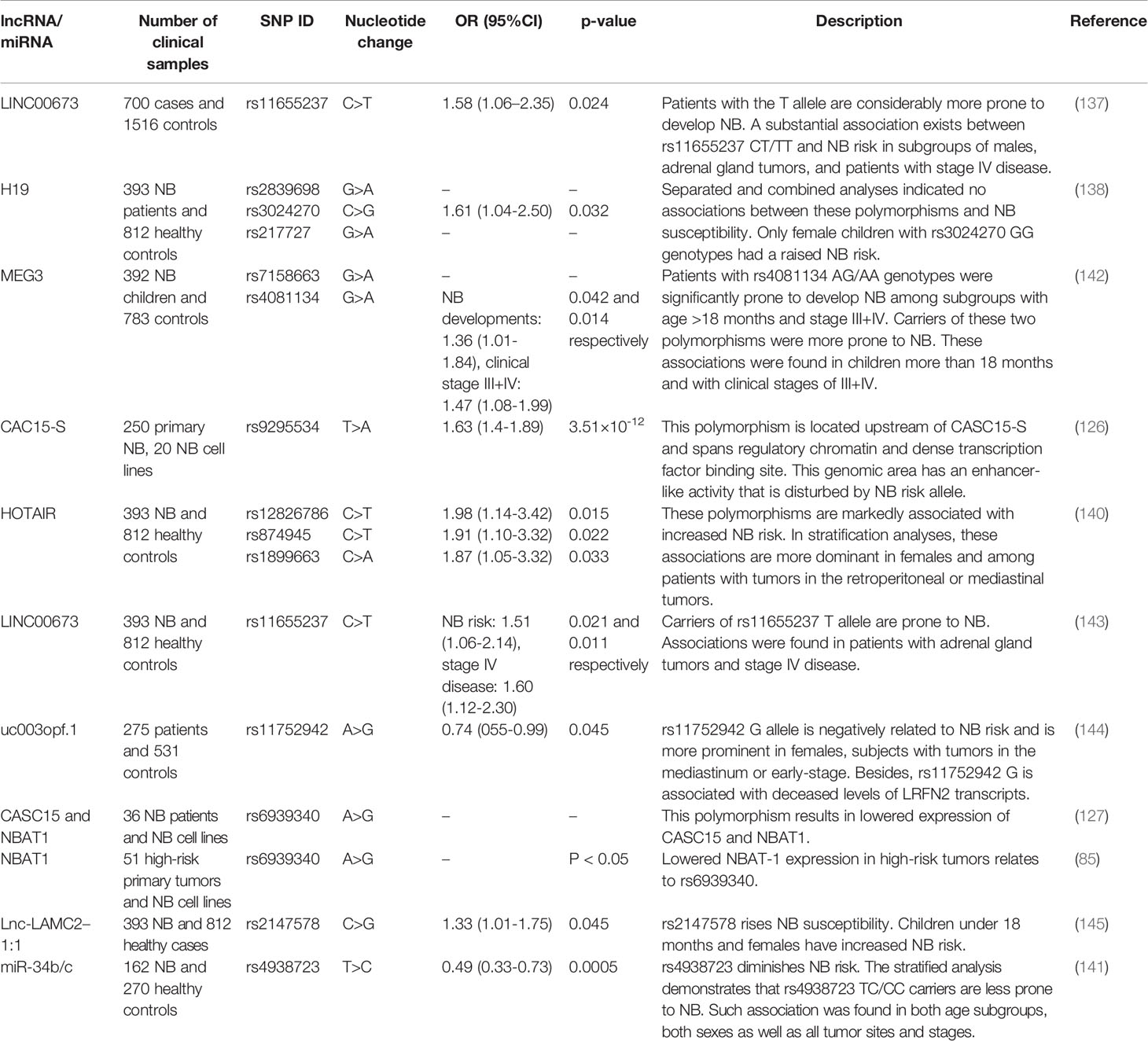
Single nucleotide polymorphisms (SNPs) within lncRNAs or miRNAs can modulate expression or activity of these transcripts, thus being implicated in the development of neuroblastoma. The role of a number of SNPs within lncRNAs such as LINC00673, H19, MEG3 and HOTAIR has been evaluated in this regard (137–140). Moreover, the rs4938723 within miR-34b/c has been associated with risk of this kind of cancer (141). Notably, some studies have appraised these associations in certain subgroups of patients. For instance, the association between rs4938723 TC and CC genotypes is prominent in all age-based subgroups, both sexes, retroperitoneal tumors as well as tumors originated from other sites, and all clinical stages (141). Such detailed analyses have not been done for all assessed SNPs. Table 8 summarizes the results of studies which assessed contribution of SNPs within ncRNAs in conferring the risk of neuroblastoma.
Discussion
Recent studies have demonstrated abnormal expression of lncRNAs, miRNAs and circRNAs in neuroblastoma. Besides, some SNPs within lncRNAs and miRNAs confer risk of neuroblastoma. In vitro studies have shown the functional interactions between a number of these ncRNAs and MYCN, the oncogene that has essential roles in the pathogenesis of this type of cancer. Moreover, certain miRNAs have been shown to target tyrosine kinase receptors. For instance, hsa-miR-376c is predicted to target ALK tyrosine kinase receptor. Notably, this miRNA has been up-regulated in neuroblastoma samples of long-survivors (146). Expressions of a number of other ncRNAs have been shown to stratify neuroblastoma patients based on their risk of recurrence and clinical outcome.
The observed dysregulation of ncRNAs in neuroblastoma can be explained by their association with the frequent chromosomal abnormalities in this kind of cancer. Amplification of genomic loci corresponding to these transcripts is a possible route for their up-regulation (86). Moreover, epigenetic factors participate in the regulation of ncRNAs expression in neuroblastoma, as several lines of evidence points to the role of retinoic acid and its derivatives in the reversal of such dysregulation. Consistent with these observations, ATRA has been lately shown to induce differentiation of a number of neuroblastoma cell lines or activate apoptosis in these cells (147).
As a number of ncRNAs regulate tumorigenic process downstream of MYCN, dysregulation of these transcripts might represent an alternative mechanism of MYCN up-regulation/amplification in neuroblastoma. In vivo studies have demonstrated the efficacy of miRNA antagonism in suppression of proliferation of MYCN-amplified neuroblastoma cells in animal models (68). However, these results have not been replicated in clinical settings. Administration of miRNA mimics in clinical settings has encountered some problems most of the being related with the distribution of these transcripts in the body and enrichment in the target organs. Encapsulation of these small transcripts in nanoparticle vesicles is expected to enhance their stability and their presence in the circulation, permitting further time for their amassment in tumor tissues (148).
Multidrug resistance is a problem in the treatment of patients with neuroblastoma. Such phenotype has been associated with a number of genetic abnormalities such as over-expression of MYCN oncogene, hyper-activation of tyrosine kinase receptors (BDNF-TrkB) or reduced expression and activity of tumor suppressor genes including p53 (148). Therefore, ncRNAs that modulate expression of these elements or function in the downstream of these molecules can also be involved in the multidrug resistance of these cells. Therefore, modulation of expression of these transcripts represents a novel modality to combat multidrug resistance in neuroblastoma.
Expression profile of ncRNAs has been correlated with patients’ survival. The underlying mechanism of this observation has been clarified in some cases. For instance, hsa-miR-383, hsa-miR-548d-5p, hsa-miR-939 and hsa-miR-877* miRNAs which have been down-regulated in neuroblastoma samples from long-survivors (146) target a number of genes being involved in the neuronal differentiation (149).
Taken together, the above-mentioned evidence suggests the crucial roles of ncRNAs in the regulation of important aspects of cell survival, proliferation and differentiation and their participation in the pathogenesis of neuroblastoma. Their potential as therapeutic targets for this type of cancer should be more explored in the future studies. The main limitation of studies which assessed expression of ncRNAs in neuroblastoma is lack of longitudinal assessment of expression of these transcripts to unravel temporal changes during the course of disease. Conduction of this type of studies would facilitate approval of the diagnostic and prognostic power of ncRNAs.
Author Contributions
MT and SG-F wrote the draft and revised it. OR, KHT, and MH performed the data collection, designed the tables and figures. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Grau E, Oltra S, Orellana C, Hernández-Martí M, Castel V. There is no evidence that the SDHB gene is involved in neuroblastoma development. Oncol Res Featuring Preclinical Clin Cancer Ther (2005) 15(7-8):393–8. doi: 10.3727/096504005776449671
2. Kushner BH, Cheung N-KV. Neuroblastoma—from genetic profiles to clinical challenge. New Engl J Med (2005) 353(21):2215–7. doi: 10.1056/NEJMp058251
3. Heck JE, Ritz B, Hung RJ, Hashibe M, Boffetta P. The epidemiology of neuroblastoma: a review. Paediatr Perinat Epidemiol (2009) 23(2):125–43. doi: 10.1111/j.1365-3016.2008.00983.x
4. Forouzani-Moghaddam MJ, Nabian P, Gholami A, Dehghanbaghi N, Azizipanah M, Jokar K, et al. A review of neuroblastoma: prevalence, diagnosis, related genetic factors, and treatment. Iran J Pediatr Hematol Oncol (2018) 8(4):237–46 doi: 10.1038/nature11247
5. Brodeur GM. Spontaneous regression of neuroblastoma. Cell Tissue Res (2018) 372(2):277–86. doi: 10.1007/s00441-017-2761-2
6. Brinkschmidt C, Christiansen H, Terpe HJ, Simon R, Lampert F, Boecker W, et al. Distal chromosome 17 gains in neuroblastomas detected by comparative genomic hybridization (CGH) are associated with a poor clinical outcome. Med Pediatr Oncol (2001) 36(1):11–3. doi: 10.1002/1096-911X(20010101)36:1<11::AID-MPO1004>3.0.CO;2-M
7. Vandesompele J, Baudis M, De Preter K, Van Roy N, Ambros P, Bown N, et al. Unequivocal delineation of clinicogenetic subgroups and development of a new model for improved outcome prediction in neuroblastoma. J Clin Oncol Off J Am Soc Clin Oncol (2005) 23(10):2280–99. doi: 10.1200/JCO.2005.06.104
8. Plantaz D, Vandesompele J, Van Roy N, Lastowska M, Bown N, Combaret V, et al. Comparative genomic hybridization (CGH) analysis of stage 4 neuroblastoma reveals high frequency of 11q deletion in tumors lacking MYCN amplification. Int J Cancer (2001) 91(5):680–6. doi: 10.1002/1097-0215(200002)9999:9999<::AID-IJC1114>3.0.CO;2-R
9. Breen CJ, O’Meara A, McDermott M, Mullarkey M, Stallings RL. Coordinate deletion of chromosome 3p and 11q in neuroblastoma detected by comparative genomic hybridization. Cancer Genet Cytogenet (2000) 120(1):44–9. doi: 10.1016/S0165-4608(99)00252-6
10. Stallings RL. MicroRNA involvement in the pathogenesis of neuroblastoma: potential for microRNA mediated therapeutics. Curr Pharm Des (2009) 15(4):456–62. doi: 10.2174/138161209787315837
11. Palazzo AF, Lee ES. Non-coding RNA: what is functional and what is junk? Front Genet (2015) 6:2. doi: 10.3389/fgene.2015.00002
12. Dunham I, Birney E, Lajoie BR, Sanyal A, Dong X, Greven M, et al. An integrated encyclopedia of DNA elements in the human genome2012. Nature (2012) 498(7414):57–74. doi: 10.1038/nature11247
13. Chen Y, Stallings RL. Differential patterns of microRNA expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res (2007) 67(3):976–83. doi: 10.1158/0008-5472.CAN-06-3667
14. Welch C, Chen Y, Stallings RL. MicroRNA-34a functions as a potential tumor suppressor by inducing apoptosis in neuroblastoma cells. Oncogene (2007) 26(34):5017–22. doi: 10.1038/sj.onc.1210293
15. Cole KA, Attiyeh EF, Mosse YP, Laquaglia MJ, Diskin SJ, Brodeur GM, et al. A functional screen identifies miR-34a as a candidate neuroblastoma tumor suppressor gene. Mol Cancer Res (2008) 6(5):735–42. doi: 10.1158/1541-7786.MCR-07-2102
16. Wei JS, Song YK, Durinck S, Chen QR, Cheuk AT, Tsang P, et al. The MYCN oncogene is a direct target of miR-34a. Oncogene (2008) 27(39):5204–13. doi: 10.1038/onc.2008.154
17. Chen J, Hongting L, Shaoping L, Xin C, Qian D. MiR-34-a acts as a suppressor in neuroblastoma progression by targeting CD44. J Pak Med Assoc (2017) 67(10):1524–31.
18. Cheng X, Xu Q, Zhang Y, Shen M, Zhang S, Mao F, et al. miR-34a inhibits progression of neuroblastoma by targeting autophagy-related gene 5. Eur J Pharmacol (2019) 850:53–63. doi: 10.1016/j.ejphar.2019.01.071
19. Misra S, Hascall VC, Markwald RR, Ghatak S. Interactions between Hyaluronan and Its Receptors (CD44, RHAMM) Regulate the Activities of Inflammation and Cancer. Front Immunol (2015) 6:201–1. doi: 10.3389/fimmu.2015.00201
20. Guo H, Chitiprolu M, Roncevic L, Javalet C, Hemming FJ, Trung MT, et al. Atg5 disassociates the V1V0-ATPase to promote exosome production and tumor metastasis independent of canonical macroautophagy. Dev Cell (2017) 43(6):716–730. e7. doi: 10.1016/j.devcel.2017.11.018
21. Bray I, Tivnan A, Bryan K, Foley NH, Watters KM, Tracey L, et al. MicroRNA-542-5p as a novel tumor suppressor in neuroblastoma. Cancer Lett (2011) 303(1):56–64. doi: 10.1016/j.canlet.2011.01.016
22. Wang J, Zhang X, Yao H, Le Y, Zhou W, Li J, et al. MiR-490-5p functions as tumor suppressor in childhood neuroblastoma by targeting MYEOV. Hum Cell (2020) 33(1):261–71. doi: 10.1007/s13577-019-00302-z
23. Cao X, Sun Z, Zhang L, Chen M, Yuan B. microRNA-144-3p suppresses human neuroblastoma cell proliferation by targeting HOXA7. Eur Rev Med Pharmacol Sci (2019) 23(2):716–23. doi: 10.26355/eurrev_201901_16885
24. Li Z, Chen H. miR-34a inhibits proliferation, migration and invasion of paediatric neuroblastoma cells via targeting HNF4α. Artif Cells Nanomed Biotechnol (2019) 47(1):3072–8. doi: 10.1080/21691401.2019.1637886
25. Sadra K.B.K.H.A., Huh S-O. Targeting the Difficult-to-Drug CD71 and MYCN with Gambogic Acid and Vorinostat in a Class of Neuroblastomas. Cell Physiol Biochem (2019) 53:258–80. doi: 10.33594/000000134
26. Lodrini M, Poschmann G, Schmidt V, Ẅnschel J, Dreidax D, Witt O, et al. Minichromosome maintenance complex is a critical node in the miR-183 signaling network of MYCN-amplified neuroblastoma cells. J Proteome Res (2016) 15(7):2178–86. doi: 10.1021/acs.jproteome.6b00134
27. Soriano A, Masanas M, Boloix A, Masiá N, París-Coderch L, Piskareva O, et al. Functional high-throughput screening reveals miR-323a-5p and miR-342-5p as new tumor-suppressive microRNA for neuroblastoma. Cell Mol Life Sci (2019) 76(11):2231–43. doi: 10.1007/s00018-019-03041-4
28. Bettinsoli P, Ferrari-Toninelli G, Bonini S, Prandelli C, Memo M. Notch ligand Delta-like 1 as a novel molecular target in childhood neuroblastoma. BMC Cancer (2017) 17(1):1–12. doi: 10.1186/s12885-017-3340-3
29. Liu H, Huang H, Li R, Bi W, Feng L, Lingling E, et al. Mitophagy protects SH-SY5Y neuroblastoma cells against the TNFα-induced inflammatory injury: involvement of microRNA-145 and Bnip3. Biomed Pharmacother (2019) 109:957–68. doi: 10.1016/j.biopha.2018.10.123
30. Zhao Z, Partridge V, Sousares M, Shelton SD, Holland CL, Pertsemlidis A, et al. microRNA-2110 functions as an onco-suppressor in neuroblastoma by directly targeting Tsukushi. PloS One (2018) 13(12):e0208777. doi: 10.1371/journal.pone.0208777
31. Neviani P, Wise PM, Murtadha M, Liu CW, Wu C-H, Jong AY, et al. Natural killer–derived exosomal miR-186 inhibits neuroblastoma growth and immune escape mechanisms. Cancer Res (2019) 79(6):1151–64. doi: 10.1158/0008-5472.CAN-18-0779
32. Wang X-H, Wu H-Y, Gao J, Wang X-H, Gao T-H, Zhang S-F. FGF represses metastasis of neuroblastoma regulated by MYCN and TGF-β1 induced LMO1 via control of let-7 expression. Brain Res (2019) 1704:219–28. doi: 10.1016/j.brainres.2018.10.015
33. Lozier AM, Rich ME, Grawe AP, Peck AS, Zhao P, Chang AT-T, et al. Targeting ornithine decarboxylase reverses the LIN28/Let-7 axis and inhibits glycolytic metabolism in neuroblastoma. Oncotarget (2015) 6(1):196. doi: 10.18632/oncotarget.2768
34. Powers JT, Tsanov KM, Pearson DS, Roels F, Spina CS, Ebright R, et al. Multiple mechanisms disrupt the let-7 microRNA family in neuroblastoma. Nature (2016) 535(7611):246–51. doi: 10.1038/nature18632
35. Marengo B, Monti P, Miele M, Menichini P, Ottaggio L, Foggetti G, et al. Etoposide-resistance in a neuroblastoma model cell line is associated with 13q14. 3 mono-allelic deletion and miRNA-15a/16-1 down-regulation. Sci Rep (2018) 8(1):1–15. doi: 10.1038/s41598-018-32195-7
36. Klein S, Abraham M, Bulvik B, Dery E, Weiss ID, Barashi N, et al. CXCR4 Promotes Neuroblastoma Growth and Therapeutic Resistance through miR-15a/16-1–Mediated ERK and BCL2/Cyclin D1 Pathways. Cancer Res (2018) 78(6):1471–83. doi: 10.1158/0008-5472.CAN-17-0454
37. Saeki N, Saito A, Sugaya Y, Amemiya M, Sasaki H. Indirect Down-regulation of Tumor-suppressive let-7 Family MicroRNAs by LMO1 in Neuroblastoma. Cancer Genomics-Proteomics (2018) 15(5):413–20. doi: 10.21873/cgp.20100
38. Li S-H, Li J-P, Chen L, Liu J-L. miR-146a induces apoptosis in neuroblastoma cells by targeting BCL11A. Med Hypotheses (2018) 117:21–7. doi: 10.1016/j.mehy.2018.05.019
39. Wang X, Li J, Xu X, Zheng J, Li Q. miR-129 inhibits tumor growth and potentiates chemosensitivity of neuroblastoma by targeting MYO10. Biomed Pharmacother (2018) 103:1312–8. doi: 10.1016/j.biopha.2018.04.153
40. Wu T, Lin Y, Xie Z. MicroRNA-1247 inhibits cell proliferation by directly targeting ZNF346 in childhood neuroblastoma. Biol Res (2018) 51(1):1–10. doi: 10.1186/s40659-018-0162-y
41. Ooi CY, Carter DR, Liu B, Mayoh C, Beckers A, Lalwani A, et al. Network modeling of microRNA–mRNA interactions in neuroblastoma tumorigenesis identifies miR-204 as a direct inhibitor of MYCN. Cancer Res (2018) 78(12):3122–34. doi: 10.1158/0008-5472.CAN-17-3034
42. Watanabe K, Yamaji R, Ohtsuki T. Micro RNA-664a-5p promotes neuronal differentiation of SH-SY 5Y cells. Genes Cells (2018) 23(3):225–33. doi: 10.1111/gtc.12559
43. Sharif S, Ghahremani MH, Soleimani M. Induction of morphological and functional differentiation of human neuroblastoma cells by miR-124. J Biosci (2017) 42(4):555–63. doi: 10.1007/s12038-017-9714-5
44. Yang K, Tong L, Li K, Zhou Y, Xiao J. A SRSF1 self-binding mechanism restrains Mir505-3p from inhibiting proliferation of neural tumor cell lines. Anti-cancer Drugs (2018) 29(1):40–9. doi: 10.1097/CAD.0000000000000564
45. Xia H-L, Lv Y, Xu C-W, Fu M-C, Zhang T, Yan X-M, et al. MiR-513c suppresses neuroblastoma cell migration, invasion, and proliferation through direct targeting glutaminase (GLS). Cancer Biomarkers (2017) 20(4):589–96. doi: 10.3233/CBM-170577
46. Chen S, Jin L, Nie S, Han L, Lu N, Zhou Y. miR-205 inhibits neuroblastoma growth by targeting cAMP-responsive element-binding protein 1. Oncol Res Featuring Preclinical Clin Cancer Ther (2018) 26(3):445–55. doi: 10.3727/096504017X14974834436195
47. Megiorni F, Colaiacovo M, Cialfi S, McDowell HP, Guffanti A, Camero S, et al. A sketch of known and novel MYCN-associated miRNA networks in neuroblastoma. Oncol Rep (2017) 38(1):3–20. doi: 10.3892/or.2017.5701
48. Samaraweera L, Spengler BA, Ross RA. Reciprocal antagonistic regulation of N-myc mRNA by miR−17 and the neuronal-specific RNA-binding protein HuD. Oncol Rep (2017) 38(1):545–50. doi: 10.3892/or.2017.5664
49. Xu Y, Chen X, Lin L, Chen H, Yu S, Li D. MicroRNA-149 is associated with clinical outcome in human neuroblastoma and modulates cancer cell proliferation through Rap1 independent of MYCN amplification. Biochimie (2017) 139:1–8. doi: 10.1016/j.biochi.2017.04.011
50. Zhao G, Wang G, Bai H, Li T, Gong F, Yang H, et al. Targeted inhibition of HDAC8 increases the doxorubicin sensitivity of neuroblastoma cells via up regulation of miR-137. Eur J Pharmacol (2017) 802:20–6. doi: 10.1016/j.ejphar.2017.02.035
51. Ren X, Bai X, Zhang X, Li Z, Tang L, Zhao X, et al. Quantitative nuclear proteomics identifies that miR-137-mediated EZH2 reduction regulates resveratrol-induced apoptosis of neuroblastoma cells. Mol Cell Proteomics (2015) 14(2):316–28. doi: 10.1074/mcp.M114.041905
52. Yang H-J, Ju F, Guo X-X, Ma S-P, Wang L, Cheng B-F, et al. RNA-binding protein RBM3 prevents NO-induced apoptosis in human neuroblastoma cells by modulating p38 signaling and miR-143. Sci Rep (2017) 7(1):1–11. doi: 10.1038/srep41738
53. Boyineni J, Tanpure S, Gnanamony M, Antony R, Fernández KS, Lin J, et al. SPARC overexpression combined with radiation retards angiogenesis by suppressing VEGF-A via miR−410 in human neuroblastoma cells. Int J Oncol (2016) 49(4):1394–406. doi: 10.3892/ijo.2016.3646
54. Fabbri E, Montagner G, Bianchi N, Finotti A, Borgatti M, Lampronti I, et al. MicroRNA miR-93-5p regulates expression of IL-8 and VEGF in neuroblastoma SK-N-AS cells. Oncol Rep (2016) 35(5):2866–72. doi: 10.3892/or.2016.4676
55. Wang Z, Lei H, Sun Q. MicroRNA-141 and its associated gene FUS modulate proliferation, migration and cisplatin chemosensitivity in neuroblastoma cell lines. Oncol Rep (2016) 35(5):2943–51. doi: 10.3892/or.2016.4640
56. Soriano A, París-Coderch L, Jubierre L, Martínez A, Zhou X, Piskareva O, et al. MicroRNA-497 impairs the growth of chemoresistant neuroblastoma cells by targeting cell cycle, survival and vascular permeability genes. Oncotarget (2016) 7(8):9271. doi: 10.18632/oncotarget.7005
57. Liu G, Xu Z, Hao D. MicroRNA−451 inhibits neuroblastoma proliferation, invasion and migration by targeting macrophage migration inhibitory factor. Mol Med Rep (2016) 13(3):2253–60. doi: 10.3892/mmr.2016.4770
58. Zhao D, Tian Y, Li P, Wang L, Xiao A, Zhang M, et al. MicroRNA-203 inhibits the malignant progression of neuroblastoma by targeting Sam68. Mol Med Rep (2015) 12(4):5554–60. doi: 10.3892/mmr.2015.4013
59. Beckers A, Van Peer G, Carter DR, Gartlgruber M, Herrmann C, Agarwal S, et al. MYCN-driven regulatory mechanisms controlling LIN28B in neuroblastoma. Cancer Lett (2015) 366(1):123–32. doi: 10.1016/j.canlet.2015.06.015
60. Xiang X, Mei H, Zhao X, Pu J, Li D, Qu H, et al. miRNA-337-3p suppresses neuroblastoma progression by repressing the transcription of matrix metalloproteinase 14. Oncotarget (2015) 6(26):22452. doi: 10.18632/oncotarget.4311
61. Wu K, Yang L, Chen J, Zhao H, Wang J, Xu S, et al. miR-362-5p inhibits proliferation and migration of neuroblastoma cells by targeting phosphatidylinositol 3-kinase-C2β. FEBS Lett (2015) 589(15):1911–9. doi: 10.1016/j.febslet.2015.05.056
62. Stigliani S, Scaruffi P, Lagazio C, Persico L, Carlini B, Varesio L, et al. Deregulation of focal adhesion pathway mediated by miR-659-3p is implicated in bone marrow infiltration of stage M neuroblastoma patients. Oncotarget (2015) 6(15):13295. doi: 10.18632/oncotarget.3745
63. Rihani A, Van Goethem A, Ongenaert M, De Brouwer S, Volders P-J, Agarwal S, et al. Genome wide expression profiling of p53 regulated miRNAs in neuroblastoma. Sci Rep (2015) 5:9027. doi: 10.1038/srep09027
64. Zhao Z, Ma X, Sung D, Li M, Kosti A, Lin G, et al. microRNA-449a functions as a tumor suppressor in neuroblastoma through inducing cell differentiation and cell cycle arrest. RNA Biol (2015) 12(5):538–54. doi: 10.1080/15476286.2015.1023495
65. Harvey H, Piskareva O, Creevey L, Alcock LC, Buckley PG, O’Sullivan MJ, et al. Modulation of chemotherapeutic drug resistance in neuroblastoma SK-N-AS cells by the neural apoptosis inhibitory protein and mi R-520f. Int J Cancer (2015) 136(7):1579–88. doi: 10.1002/ijc.29144
66. Althoff K, Lindner S, Odersky A, Mestdagh P, Beckers A, Karczewski S, et al. miR-542-3p exerts tumor suppressive functions in neuroblastoma by downregulating S urvivin. Int J Cancer (2015) 136(6):1308–20. doi: 10.1002/ijc.29091
67. Schulte JH, Horn S, Otto T, Samans B, Heukamp LC, Eilers UC, et al. MYCN regulates oncogenic MicroRNAs in neuroblastoma. Int J Cancer (2008) 122(3):699–704. doi: 10.1002/ijc.23153
68. Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M, et al. Antagomir-17-5p abolishes the growth of therapy-resistant neuroblastoma through p21 and BIM. PloS One (2008) 3(5):e2236. doi: 10.1371/journal.pone.0002236
69. Wu JC, Jiang HM, Yang XH, Zheng HC. ING5-mediated antineuroblastoma effects of suberoylanilide hydroxamic acid. Cancer Med (2018) 7(9):4554–69. doi: 10.1002/cam4.1634
70. Li Z, Xu Z, Xie Q, Gao W, Xie J, Zhou L. miR-1303 promotes the proliferation of neuroblastoma cell SH-SY5Y by targeting GSK3β and SFRP1. Biomed Pharmacother (2016) 83:508–13. doi: 10.1016/j.biopha.2016.07.010
71. Chen J, Wang P, Cai R, Peng H, Zhang C, Zhang M. SLC 34A2 promotes neuroblastoma cell stemness via enhancement of miR-25/Gsk3β-mediated activation of Wnt/β-catenin signaling. FEBS Open Bio (2019) 9(3):527–37. doi: 10.1002/2211-5463.12594
72. Nowak I, Boratyn E, Durbas M, Horwacik I, Rokita H. Exogenous expression of miRNA-3613-3p causes APAF1 downregulation and affects several proteins involved in apoptosis in BE (2)-C human neuroblastoma cells. Int J Oncol (2018) 53(4):1787–99. doi: 10.3892/ijo.2018.4509
73. Liu X, Peng H, Liao W, Luo A, Cai M, He J, et al. MiR-181a/b induce the growth, invasion, and metastasis of neuroblastoma cells through targeting ABI1. Mol Carcinogenesis (2018) 57(9):1237–50. doi: 10.1002/mc.22839
74. Jiang J, Song X, Yang J, Lei K, Ni Y, Zhou F, et al. Triptolide inhibits proliferation and migration of human neuroblastoma SH-SY5Y cells by upregulating microRNA-181a. Oncol Res Featuring Preclinical Clin Cancer Ther (2018) 26(8):1235–43. doi: 10.3727/096504018X15179661552702
75. Cheng M, Liu L, Lao Y, Liao W, Liao M, Luo X, et al. MicroRNA-181a suppresses parkin-mediated mitophagy and sensitizes neuroblastoma cells to mitochondrial uncoupler-induced apoptosis. Oncotarget (2016) 7(27):42274. doi: 10.18632/oncotarget.9786
76. He X-y, Tan Z-l, Mou Q, Liu F-j, Liu S, Yu C-w, et al. MicroRNA-221 enhances MYCN via targeting nemo-like kinase and functions as an oncogene related to poor prognosis in neuroblastoma. Clin Cancer Res (2017) 23(11):2905–18. doi: 10.1158/1078-0432.CCR-16-1591
77. Qu H, Zheng L, Song H, Jiao W, Li D, Fang E, et al. microRNA-558 facilitates the expression of hypoxia-inducible factor 2 alpha through binding to 5′-untranslated region in neuroblastoma. Oncotarget (2016) 7(26):40657. doi: 10.18632/oncotarget.9813
78. Qu H, Zheng L, Pu J, Mei H, Xiang X, Zhao X, et al. miRNA-558 promotes tumorigenesis and aggressiveness of neuroblastoma cells through activating the transcription of heparanase. Hum Mol Genet (2015) 24(9):2539–51. doi: 10.1093/hmg/ddv018
79. Chen Y, Tsai Y-H, Tseng B-J, Pan H-Y, Tseng S-H. Suppression of miR-19b enhanced the cytotoxic effects of mTOR inhibitors in human neuroblastoma cells. J Pediatr Surg (2016) 51(11):1818–25. doi: 10.1016/j.jpedsurg.2016.07.003
80. Li Y, Shang YM, Wang QW. MicroRNA-21 promotes the proliferation and invasion of neuroblastoma cells through targeting CHL1. Minerva Med (2016) 107(5):287–93.
81. Maugeri M, Barbagallo D, Barbagallo C, Banelli B, Di Mauro S, Purrello F, et al. Altered expression of miRNAs and methylation of their promoters are correlated in neuroblastoma. Oncotarget (2016) 7(50):83330. doi: 10.18632/oncotarget.13090
82. Zhou Y, Sheng B. Association of microRNA 21 with biological features and prognosis of neuroblastoma. Cancer Control (2016) 23(1):78–84. doi: 10.1177/107327481602300113
83. Fernandes JCR, Acuña SM, Aoki JI, Floeter-Winter LM, Muxel SM. Long Non-Coding RNAs in the Regulation of Gene Expression: Physiology and Disease. Noncoding RNA (2019) 5(1):17. doi: 10.3390/ncrna5010017
84. Prajapati B, Fatma M, Fatima M, Khan MT, Sinha S, Seth PK. Identification of lncRNAs Associated With Neuroblastoma in Cross-Sectional Databases: Potential Biomarkers. Front Mol Neurosci (2019) 12:293–3. doi: 10.3389/fnmol.2019.00293
85. Pandey GK, Mitra S, Subhash S, Hertwig F, Kanduri M, Mishra K, et al. The risk-associated long noncoding RNA NBAT-1 controls neuroblastoma progression by regulating cell proliferation and neuronal differentiation. Cancer Cell (2014) 26(5):722–37. doi: 10.1016/j.ccell.2014.09.014
86. Liu PY, Erriquez D, Marshall GM, Tee AE, Polly P, Wong M, et al. Effects of a novel long noncoding RNA, lncUSMycN, on N-Myc expression and neuroblastoma progression. JNCI: J Natl Cancer Inst (2014) 106(7):dju113. doi: 10.1093/jnci/dju113
87. Barnhill LM, Williams RT, Cohen O, Kim Y, Batova A, Mielke JA, et al. High expression of CAI2, a 9p21-embedded long noncoding RNA, contributes to advanced-stage neuroblastoma. Cancer Res (2014) 74(14):3753–63. doi: 10.1158/0008-5472.CAN-13-3447
88. Watters KM, Bryan K, Foley NH, Meehan M, Stallings RL. Expressional alterations in functional ultra-conserved non-coding RNAs in response to all-trans retinoic acid–induced differentiation in neuroblastoma cells. BMC Cancer (2013) 13:184. doi: 10.1186/1471-2407-13-184
89. Yu M, Ohira M, Li Y, Niizuma H, Oo ML, Zhu Y, et al. High expression of ncRAN, a novel non-coding RNA mapped to chromosome 17q25.1, is associated with poor prognosis in neuroblastoma. Int J Oncol (2009) 34(4):931–8. doi: 10.3892/ijo_00000219
90. Li C, Wang S, Yang C. Long non-coding RNA DLX6-AS1 regulates neuroblastoma progression by targeting YAP1 via miR-497-5p. Life Sci (2020), 252:117657. doi: 10.1016/j.lfs.2020.117657
91. Hu Y, Sun H, Hu J, Zhang X. LncRNA DLX6-AS1 Promotes the Progression of Neuroblastoma by Activating STAT2 via Targeting miR-506-3p. Cancer Manage Res (2020) 12:7451–63. doi: 10.2147/CMAR.S252521
92. Liu PY, Tee AE, Milazzo G, Hannan KM, Maag J, Mondal S, et al. The long noncoding RNA lncNB1 promotes tumorigenesis by interacting with ribosomal protein RPL35. Nat Commun (2019) 10(1):1–17. doi: 10.1038/s41467-019-12971-3
93. Shatara M, Xavier AC, Dombkowski A, Cukovic D, Poulik JM, Altinok D, et al. Monozygotic twins with neuroblastoma MS have a similar molecular profile: a case of twin-to-twin metastasis. Br J Cancer (2019) 121(10):890–3. doi: 10.1038/s41416-019-0594-3
94. Wang J, Wang Z, Yao W, Dong K, Zheng S, Li K. The association between lncRNA LINC01296 and the clinical characteristics in neuroblastoma. J Pediatr Surg (2019) 54(12):2589–94. doi: 10.1016/j.jpedsurg.2019.08.032
95. Yu Y, Chen F, Yang Y, Jin Y, Shi J, Han S, et al. lncRNA SNHG16 is associated with proliferation and poor prognosis of pediatric neuroblastoma. Int J Oncol (2019) 55(1):93–102. doi: 10.3892/ijo.2019.4813
96. Xu Z, Sun Y, Wang D, Sun H, Liu X. SNHG16 promotes tumorigenesis and cisplatin resistance by regulating miR-338-3p/PLK4 pathway in neuroblastoma cells. Cancer Cell Int (2020) 20(1):1–13. doi: 10.1186/s12935-020-01291-y
97. Bao J, Zhang S, Meng Q, Qin T. SNHG16 Silencing Inhibits Neuroblastoma Progression by Downregulating HOXA7 via Sponging miR-128-3p. Neurochem Res (2020), 1–12. doi: 10.1007/s11064-020-02955-x
98. Deng D, Yang S, Wang X. Long non-coding RNA SNHG16 regulates cell behaviors through miR-542-3p/HNF4α axis via RAS/RAF/MEK/ERK signaling pathway in pediatric neuroblastoma cells. Biosci Rep (2020) 40(5). doi: 10.1042/BSR20200723
99. Wen Y, Gong X, Dong Y, Tang C. Long Non Coding RNA SNHG16 Facilitates Proliferation, Migration, Invasion and Autophagy of Neuroblastoma Cells via Sponging miR-542-3p and Upregulating ATG5 Expression. OncoTargets Ther (2020) 13:263. doi: 10.2147/OTT.S226915
100. Li E-y, Zhao P-j, Jian J, Yin B-q, Sun Z-y, Xu C-x, et al. LncRNA MIAT overexpression reduced neuron apoptosis in a neonatal rat model of hypoxic-ischemic injury through miR-211/GDNF. Cell Cycle (2019) 18(2):156–66. doi: 10.1080/15384101.2018.1560202
101. Chi R, Chen X, Liu M, Zhang H, Li F, Fan X, et al. Role of SNHG7-miR-653-5p-STAT2 feedback loop in regulating neuroblastoma progression. J Cell Physiol (2019) 234(8):13403–12. doi: 10.1002/jcp.28017
102. Wang S, Wang X, Zhang C. LncRNA SNHG7 enhances chemoresistance in neuroblastoma through cisplatin-induced autophagy by regulating miR-329-3p/MYO10 axis. Eur Rev Med Pharmacol Sci (2020) 24(7):3805–17. doi: 10.26355/eurrev_202004_20847
103. Jia J, Zhang D, Zhang J, Yang L, Zhao G, Yang H, et al. Long non-coding RNA SNHG7 promotes neuroblastoma progression through sponging miR-323a-5p and miR-342-5p. Biomed Pharmacother (2020) 128:110293. doi: 10.1016/j.biopha.2020.110293
104. Pan J, Zhang D, Zhang J, Qin P, Wang J. LncRNA RMRP silence curbs neonatal neuroblastoma progression by regulating microRNA-206/tachykinin-1 receptor axis via inactivating extracellular signal-regulated kinases. Cancer Biol Ther (2019) 20(5):653–65. doi: 10.1080/15384047.2018.1550568
105. Yang T-W, Sahu D, Chang Y-W, Hsu C-L, Hsieh C-H, Huang H-C, et al. RNA-binding proteomics reveals MATR3 interacting with lncRNA SNHG1 to enhance neuroblastoma progression. J Proteome Res (2018) 18(1):406–16. doi: 10.1021/acs.jproteome.8b00693
106. Sahu D, Hsu C-L, Lin C-C, Yang T-W, Hsu W-M, Ho S-Y, et al. Co-expression analysis identifies long noncoding RNA SNHG1 as a novel predictor for event-free survival in neuroblastoma. Oncotarget (2016) 7(36):58022. doi: 10.18632/oncotarget.11158
107. Chai P, Jia R, Jia R, Pan H, Wang S, Ni H, et al. Dynamic chromosomal tuning of a novel GAU1 lncing driver at chr12p13. 32 accelerates tumorigenesis. Nucleic Acids Res (2018) 46(12):6041–56. doi: 10.1093/nar/gky366
108. O’Brien EM, Selfe JL, Martins AS, Walters ZS, Shipley JM. The long non-coding RNA MYCNOS-01 regulates MYCN protein levels and affects growth of MYCN-amplified rhabdomyosarcoma and neuroblastoma cells. BMC Cancer (2018) 18(1):1–13. doi: 10.1186/s12885-018-4129-8
109. Li D, Wang X, Mei H, Fang E, Ye L, Song H, et al. Long noncoding RNA pancEts-1 promotes neuroblastoma progression through hnRNPK-mediated β-catenin stabilization. Cancer Res (2018) 78(5):1169–83. doi: 10.1158/0008-5472.CAN-17-2295
110. Bi S, Wang C, Li Y, Zhang W, Zhang J, Lv Z, et al. LncRNA-MALAT1-mediated Axl promotes cell invasion and migration in human neuroblastoma. Tumor Biol (2017) 39(5). 1010428317699796. doi: 10.1177/1010428317699796
111. Tee AE, Liu B, Song R, Li J, Pasquier E, Cheung BB, et al. The long noncoding RNA MALAT1 promotes tumor-driven angiogenesis by up-regulating pro-angiogenic gene expression. Oncotarget (2016) 7(8):8663. doi: 10.18632/oncotarget.6675
112. Tee AE, Ling D, Nelson C, Atmadibrata B, Dinger ME, Xu N, et al. The histone demethylase JMJD1A induces cell migration and invasion by up-regulating the expression of the long noncoding RNA MALAT1. Oncotarget (2014) 5(7):1793. doi: 10.18632/oncotarget.1785
113. Mazar J, Rosado A, Shelley J, Marchica J, Westmoreland TJ. The long non-coding RNA GAS5 differentially regulates cell cycle arrest and apoptosis through activation of BRCA1 and p53 in human neuroblastoma. Oncotarget (2017) 8(4):6589. doi: 10.18632/oncotarget.14244
114. Tang W, Dong K, Li K, Dong R, Zheng S. MEG3, HCN3 and linc01105 influence the proliferation and apoptosis of neuroblastoma cells via the HIF-1α and p53 pathways. Sci Rep (2016) 6(1):1–9. doi: 10.1038/srep36268
115. Liu PY, Atmadibrata B, Mondal S, Tee AE, Liu T. NCYM is upregulated by lncUSMycN and modulates N-Myc expression. Int J Oncol (2016) 49(6):2464–70. doi: 10.3892/ijo.2016.3730
116. Yarmishyn AA, Batagov AO, Tan JZ, Sundaram GM, Sampath P, Kuznetsov VA, et al. HOXD-AS1 is a novel lncRNA encoded in HOXD cluster and a marker of neuroblastoma progression revealed via integrative analysis of noncoding transcriptome. BMC Genomics (2014) 15(S9):S7. doi: 10.1186/1471-2164-15-S9-S7
117. Pavlaki I, Alammari F, Sun B, Clark N, Sirey T, Lee S, et al. The long non-coding RNA Paupar promotes KAP 1-dependent chromatin changes and regulates olfactory bulb neurogenesis. EMBO J (2018) 37(10):e98219. doi: 10.15252/embj.201798219
118. Vance KW, Sansom SN, Lee S, Chalei V, Kong L, Cooper SE, et al. The long non-coding RNA P aupar regulates the expression of both local and distal genes. EMBO J (2014) 33(4):296–311. doi: 10.1002/embj.201386225
119. Wang B, Xu L, Zhang J, Cheng X, Xu Q, Wang J, et al. LncRNA NORAD accelerates the progression and doxorubicin resistance of neuroblastoma through up-regulating HDAC8 via sponging miR-144-3p. Biomed Pharmacother (2020) 129:110268. doi: 10.1016/j.biopha.2020.110268
120. Yu Z, Zhang J, Han J. Silencing CASC11 curbs neonatal neuroblastoma progression through modulating microRNA-676-3p/nucleolar protein 4 like (NOL4L) axis. Pediatr Res (2020) 87(4):662–8. doi: 10.1038/s41390-019-0625-z
121. Nie L, Li C, Zhao T, Wang Y, Liu J. LncRNA double homeobox A pseudogene 8 (DUXAP8) facilitates the progression of neuroblastoma and activates Wnt/β-catenin pathway via microRNA-29/nucleolar protein 4 like (NOL4L) axis. Brain Res (2020) 1746:146947. doi: 10.1016/j.brainres.2020.146947
122. Yang H, Guo J, Zhang M, Li A. LncRNA SNHG4 promotes neuroblastoma proliferation, migration, and invasion by sponging miR-377-3p. Neoplasma (2020) 67:1054–62. doi: 10.4149/neo_2020_191023N1081
123. Zhao X, Li D, Yang F, Lian H, Wang J, Wang X, et al. Long Noncoding RNA NHEG1 Drives β-Catenin Transactivation and Neuroblastoma Progression through Interacting with DDX5. Mol Ther (2020) 28(3):946–62. doi: 10.1016/j.ymthe.2019.12.013
124. Mu L, Wang L, Zhang S, Wang Q. Long noncoding RNA XIST suppresses tumorigenesis and enhances radiosensitivity in neuroblastoma cells through regulating miR-653-5p/HK2 axis. (2020). doi: 10.2147/OTT.S170439
125. Tian C, Li Z, Zhang L, Dai D, Huang Q, Liu J, et al. lncRNA NR_120420 promotes SH-SY5Y cells apoptosis by regulating NF-κB after oxygen and glucose deprivation. Gene (2020) 728:144285. doi: 10.1016/j.gene.2019.144285
126. Russell MR, Penikis A, Oldridge DA, Alvarez-Dominguez JR, McDaniel L, Diamond M, et al. CASC15-S is a tumor suppressor lncRNA at the 6p22 neuroblastoma susceptibility locus. Cancer Res (2015) 75(15):3155–66. doi: 10.1158/0008-5472.CAN-14-3613
127. Mondal T, Juvvuna PK, Kirkeby A, Mitra S, Kosalai ST, Traxler L, et al. Sense-antisense lncRNA pair encoded by locus 6p22. 3 determines neuroblastoma susceptibility via the USP36-CHD7-SOX9 regulatory axis. Cancer Cell (2018) 33(3):417–434. e7. doi: 10.1016/j.ccell.2018.01.020
128. Zhao X, Li D, Huang D, Song H, Mei H, Fang E, et al. Risk-associated long noncoding RNA FOXD3-AS1 inhibits neuroblastoma progression by repressing PARP1-mediated activation of CTCF. Mol Ther (2018) 26(3):755–73. doi: 10.1016/j.ymthe.2017.12.017
129. Bevilacqua V, Gioia U, Di Carlo V, Tortorelli AF, Colombo T, Bozzoni I, et al. Identification of linc-NeD125, a novel long non coding RNA that hosts miR-125b-1 and negatively controls proliferation of human neuroblastoma cells. RNA Biol (2015) 12(12):1323–37. doi: 10.1080/15476286.2015.1096488
130. Vadie N, Saayman S, Lenox A, Ackley A, Clemson M, Burdach J, et al. MYCNOS functions as an antisense RNA regulating MYCN. RNA Biol (2015) 12(8):893–9. doi: 10.1080/15476286.2015.1063773
131. Zhou X, Lu H, Li F, Han L, Zhang H, Jiang Z, et al. LncRNA cancer susceptibility candidate (CASC7) upregulates phosphatase and tensin homolog by downregulating miR-10a to inhibit neuroblastoma cell proliferation. Neuroreport (2020) 31(5):381–6. doi: 10.1097/WNR.0000000000001411
132. Li MM, Liu XH, Zhao YC, Ma XY, Zhou YC, Zhao YX, et al. Long noncoding RNA KCNQ1OT1 promotes apoptosis in neuroblastoma cells by regulating miR-296-5p/Bax axis. FEBS J (2020) 287(3):561–77. doi: 10.1111/febs.15047
133. Pan W, Wu A, Yu H, Yu Q, Zheng B, Yang W, et al. NEAT1 Negatively Regulates Cell Proliferation and Migration of Neuroblastoma Cells by miR-183-5p/FOXP1 Via the ERK/AKT Pathway. Cell Transplant (2020) 29:0963689720943608. doi: 10.1177/0963689720943608
134. Lu W-Y. Roles of the circular RNA circ-Foxo3 in breast cancer progression. Cell Cycle (2017) 16(7):589. doi: 10.1080/15384101.2017.1278935
135. Yang J, Yu L, Yan J, Xiao Y, Li W, Xiao J, et al. Circular RNA DGKB Promotes the Progression of Neuroblastoma by Targeting miR-873/GLI1 Axis. Front Oncol (2020) 10:1104. doi: 10.3389/fonc.2020.01104
136. Li H, Yang F, Hu A, Wang X, Fang E, Chen Y, et al. Therapeutic targeting of circ-CUX 1/EWSR 1/MAZ axis inhibits glycolysis and neuroblastoma progression. EMBO Mol Med (2019) 11(12):e10835. doi: 10.15252/emmm.201910835
137. Li Y, Zhuo Z-J, Zhou H, Liu J, Liu Z, Zhang J, et al. Additional data support the role of LINC00673 rs11655237 C> T in the development of neuroblastoma. Aging (Albany NY) (2019) 11(8):2369. doi: 10.18632/aging.101920
138. Hu C, Yang T, Pan J, Zhang J, Yang J, He J, et al. Associations between H19 polymorphisms and neuroblastoma risk in Chinese children. Biosci Rep (2019) 39(4). doi: 10.1042/BSR20181582
139. Bayarmaa B, Wu Z, Peng J, Wang Y, Xu S, Yan T, et al. Association of LncRNA MEG3 polymorphisms with efficacy of neoadjuvant chemotherapy in breast cancer. BMC Cancer (2019) 19(1):877. doi: 10.1186/s12885-019-6077-3
140. Yang X, He J, Chang Y, Luo A, Luo A, Zhang J, et al. HOTAIR gene polymorphisms contribute to increased neuroblastoma susceptibility in Chinese children. Cancer (2018) 124(12):2599–606. doi: 10.1002/cncr.31353
141. Li Y, Zhuo Z-J, Zhou H, Liu J, Xiao Z, Xiao Y, et al. miR-34b/c rs4938723 T> C Decreases Neuroblastoma Risk: A Replication Study in the Hunan Children. Dis Markers (2019) 2019). doi: 10.1155/2019/6514608
142. Zhuo Z-J, Zhang R, Zhang J, Zhu J, Yang T, Zou Y, et al. Associations between lncRNA MEG3 polymorphisms and neuroblastoma risk in Chinese children. Aging (Albany NY) (2018) 10(3):481. doi: 10.18632/aging.101406
143. Zhang Z, Chang Y, Jia W, Zhang J, Zhang R, Zhu J, et al. LINC00673 rs11655237 C> T confers neuroblastoma susceptibility in Chinese population. Biosci Rep (2018) 38(1). doi: 10.1042/BSR20171667
144. Pan J, Lin H, Yang T, Yang J, Hu C, Zhu J, et al. lncRNA-uc003opf. 1 rs11752942 A> G polymorphism decreases neuroblastoma risk in Chinese children. Cell Cycle (2020) 19:1–6. doi: 10.1080/15384101.2020.1808382
145. Yang T, Zhang Z, Zhang J, Tan T, Yang J, Pan J, et al. The rs2147578 C> G polymorphism in the Inc-LAMC2–1: 1 gene is associated with increased neuroblastoma risk in the Henan children. BMC Cancer (2018) 18(1):948. doi: 10.1186/s12885-018-4847-y
146. Scaruffi P, Stigliani S, Moretti S, Coco S, De Vecchi C, Valdora F, et al. Transcribed-Ultra Conserved Region expression is associated with outcome in high-risk neuroblastoma. BMC Cancer (2009) 9:441–1. doi: 10.1186/1471-2407-9-441
147. Thiele CJ, Reynolds CP, Israel MA. Decreased expression of N-myc precedes retinoic acid-induced morphological differentiation of human neuroblastoma. Nature (1985) 313(6001):404–6. doi: 10.1038/313404a0
148. Boloix A, París-Coderch L, Soriano A, Roma J, Gallego S, de Toledo JS, et al. Novel micro RNA-based therapies for the treatment of neuroblastoma. Anales Pediatr (English Ed) (2016) 85(2):109. e1–109. e6. doi: 10.1016/j.anpede.2015.07.032
Keywords: miRNA, lncRNA, neuroblastoma, expression, polymorphism
Citation: Rezaei O, Honarmand Tamizkar K, Hajiesmaeili M, Taheri M and Ghafouri-Fard S (2021) Non-Coding RNAs Participate in the Pathogenesis of Neuroblastoma. Front. Oncol. 11:617362. doi: 10.3389/fonc.2021.617362
Received: 26 October 2020; Accepted: 11 January 2021;
Published: 24 February 2021.
Edited by:
Takaomi Sanda, National University of Singapore, SingaporeReviewed by:
Andrea Di Cataldo, University of Catania, ItalyJoanna Kitlinska, Georgetown University, United States
Copyright © 2021 Rezaei, Honarmand Tamizkar, Hajiesmaeili, Taheri and Ghafouri-Fard. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mohammad Taheri, bW9oYW1tYWRfODIzQHlhaG9vLmNvbQ==; Soudeh Ghafouri-Fard, cy5naGFmb3VyaWZhcmRAc2JtdS5hYy5pcg==
 Omidvar Rezaei1
Omidvar Rezaei1 Kasra Honarmand Tamizkar
Kasra Honarmand Tamizkar Mohammad Taheri
Mohammad Taheri