- 1Cancer Institute, Xinqiao Hospital, Army Medical University, Chongqing, China
- 2Chongqing Key Laboratory of Infectious Diseases and Parasitic Diseases, Department of Infectious Diseases, The First Affiliated Hospital of Chongqing Medical University, Chongqing, China
- 3Department of Ultrasound, The 941st Hospital of the PLA Joint Logistic Support Force, Xining, China
- 4Lung Cancer Center, Cancer Center and State Key Laboratory of Biotherapy, West China Hospital of Sichuan University, Chengdu, China
Objective: This study aimed to investigate the incidence of the pulmonary sarcomatoid carcinoma (PSC), to compare the clinical characteristics and overall survival (OS) of patients with PSC and those with other non-small-cell lung cancer (oNSCLC), so as to analyze the factors affecting the OS of patients with PSC and construct a nomogram prediction model.
Methods: Data of patients with PSC and those with oNSCLC diagnosed between 2004 and 2015 from the Surveillance, Epidemiology, and End Results database were collected. The age-adjusted incidence of PSC was calculated. The characteristics of patients with PSC and those with oNSCLC were compared, then the patients were matched 1:2 for further survival analysis. Patients with PSC were randomly divided into training set and testing set with a ratio of 7:3. The Cox proportional hazards model was used to identify the covariates associated with the OS. Significant covariates were used to construct the nomogram, and the C-index was calculated to measure the discrimination ability. The accuracy of the nomogram was compared with the tumor–node–metastasis (TNM) clinical stage, and the corresponding area under the curve was achieved.
Results: A total of 1049 patients with PSC were enrolled, the incidence of PSC was slowly decreased from 0.120/100,000 in 2004 to 0.092/100,000 in 2015. Before PSM, 793 PSC patients and 191356 oNSCLC patients were identified, the proportion of male, younger patients (<65 years), grade IV, TNM clinical stage IV was higher in the PSC. The patients with PSC had significantly poorer OS compared with those with oNSCLC. After PSM, PSC still had an extremely inferior prognosis. Age, sex, TNM clinical stage, chemotherapy, radiotherapy, and surgery were independent factors for OS. Next, a nomogram was established based on these factors, and the C-indexs were 0.775 and 0.790 for the training and testing set, respectively. Moreover, the nomogram model indicated a more comprehensive and accurate prediction than the TNM clinical stage.
Conclusions: The incidence of PSC was slowly decreased. PSC had a significantly poor prognosis compared with oNSCLC. The nomogram constructed in this study accurately predicted the prognosis of PSC, performed better than the TNM clinical stage.
Introduction
Pulmonary sarcomatoid carcinoma (PSC) is a rare subtype of non-small-cell lung cancer (NSCLC), accounting for only 0.1%–0.4% of lung cancers (1). PSC refers to poorly differentiated NSCLC containing sarcoma or sarcoma-like components or carcinomas consisting of spindle cells and giant cells (2). According to the 2015 World Health Organization (WHO) classification of lung tumors and International Classification of Disease for Oncology, 3rd Edition (ICD-O-3), PSC was classified into five categories: pleomorphic carcinoma, spindle cell carcinoma, giant cell carcinoma, carcinosarcoma, and pulmonary blastoma (3). Previous studies showed that PSC was more aggressive than other non-small-cell lung cancer (oNSCLC) and had a worse prognosis (4–6).
At present, the increasing number of studies on PSC are case reports or retrospective analyses focusing on the clinical-pathological characteristics and prognostic factors (6, 7). Despite numerous efforts to study the features of PSC, large population-based study never specifically investigated the incidence of PSC. The Surveillance, Epidemiology, and End Results (SEER) database is a systematic population-based cancer database. Therefore, this study was conducted to provide an overview of PSC incidence based on the data of SEER database.
Besides, only a few studies provided limited information about the detailed distinction between PSC and oNSCLC, and constructed a prediction model for PSC without external validation (8). Hence, this study also aimed to compare the clinicopathological characteristics and survival outcomes with oNSCLC, to explore the clinical features related to PSC overall survival (OS), and to construct and validate a nomogram prediction model.
Methods
Data Source
The patient data were obtained from the SEER 18 registries (www.seer.cancer.gov) of the National Cancer Institute using the SEER*Stat software (SEER*Stat 8.3.6). The database (1975–2016) covered about 27.8% of the American population and was released in April 2019 based on the November 2018 submission.
The inclusion criteria were as follows: patients with site record “lung and bronchus” and “one primary only”, diagnosed from 2004 to 2015. The diagnosis was confirmed by positive histology, and the type of reporting source was not autopsy or death certificate. Collectively, pleomorphic carcinoma (8022/3), giant cell carcinoma (8031/3), spindle cell carcinoma (8032/3), pulmonary blastoma (8972/3), and carcinosarcoma (8980/3) were grouped under the term PSC, and adenocarcinoma (8140/3, 8144/3, 8230/3, 8250/3, 8253/3, 8254/3, 8260/3, 8333/3, 8480/3, 8551/3), squamous cell carcinoma (8070/3–8072/3, 8083/3), adenosquamous carcinoma (8560/3), large cell carcinoma (8012/3), and others (8200/3, 8240/3, 8249/3, 8430/3, 8562/3) were grouped under the term oNSCLC. The patients lacking data about tumor, node, and metastases (TNM) clinical stage (American Joint Committee on Cancer Staging Manual, Sixth Edition) were excluded.
Study Design
The process and study design are presented in a flow-chart (Figure 1). The age-adjusted incidence of PSC was calculated with the patients with PSC. Then, the patients with PSC and the patients with oNSCLC were matched 1:2 between for further survival analysis using propensity score matching (PSM). Eligible patients with PSC were randomly divided into training set and testing set with a ratio of 7:3, and prognostic nomogram to predict 1-year survival for PSC was constructed based on training set and was validated using concordance index (C-index) and calibration curves in two sets. The total nomogram score of each patient was obtained, and the corresponding area under the curve (AUC) was achieved to compare the accuracy of the nomogram with the TNM clinical stage.
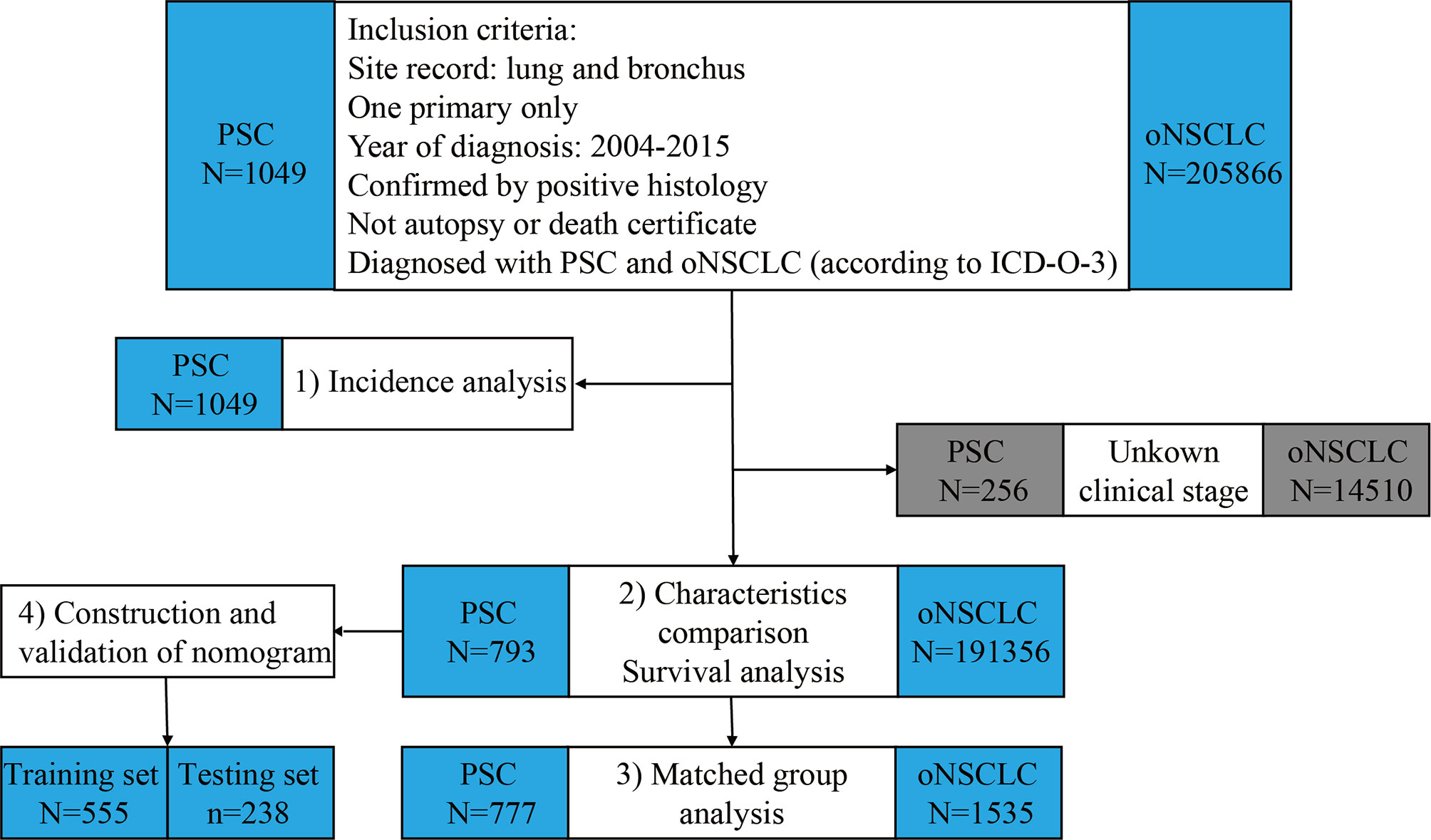
Figure 1 Flowchart of study design and patient selection. PSC, pulmonary sarcomatoid carcinoma; oNSCLC, other non-small-cell lung cancer.
Covariates
The study covariates included age at diagnosis, sex, race, year of diagnosis, grade, laterality, TNM clinical stage, chemotherapy, radiotherapy, surgery, survival months, and vital status.
Statistical Analysis
Covariates were presented as frequency and percentages and compared using Pearson’s chi-square test. The age-adjusted incidence was calculated with Rate Session in the SEER*STAT software. PSM method was used to balance baseline covariates between PSC and oNSCLC. Survival analysis was accomplished by the Kaplan–Meier method and the log-rank test. In the PSC group, the Cox proportional hazards model was used to identify the covariates associated with OS and calculate the hazard ratio (HR) with 95% confidence interval (CI). The results were displayed using the forest plot. Based on the Cox model, significant covariates (P < 0.05) were used to construct the nomogram and the C-index was calculated to measure the discrimination ability. Statistical analyses were performed using the IBM SPSS Statistics, version 23.0, and the “survival,” “MatchIt,” “createtableone,” “love.plot,” “rms,” “nomogramEx,” “nomogramFormula,” and “survivalROC” packages in the R version 4.0.0 (http://www.r-project.org/).
Results
Annual Incidence
A total of 1049 patients with PSC were enrolled to analysis the rate of incidence. The age-adjusted incidence of PSC was calculated based on the SEER 18 registries. Overall, the incidence of PSC was slowly decreased from 0.120/100,000 in 2004 to 0.092/100,000 in 2015 (Figure 2A). The incidence of male was obviously higher than female (Figure 2B). Among pathological subgroups, the incidence of carcinosarcoma increased from 0.015/100,000 in 2004 to 0.028/100,000 in 2015, and the incidence of giant cell carcinoma and spindle cell carcinoma decreased from 0.038/100,000 in 2004 to 0.014/100,000 in 2015 and 0.040/100,000 in 2004 to 0.020/100,000 in 2015, respectively. The incidence of pulmonary blastoma was stable (Figure 2C). In addition, the incidence decreased significantly in TNM clinical stage III from 0.028/100,000 in 2004 to 0.009/100,000 in 2015 (Figure 2D). Supplementary Table 1 shows the detailed incidence data of PSC.
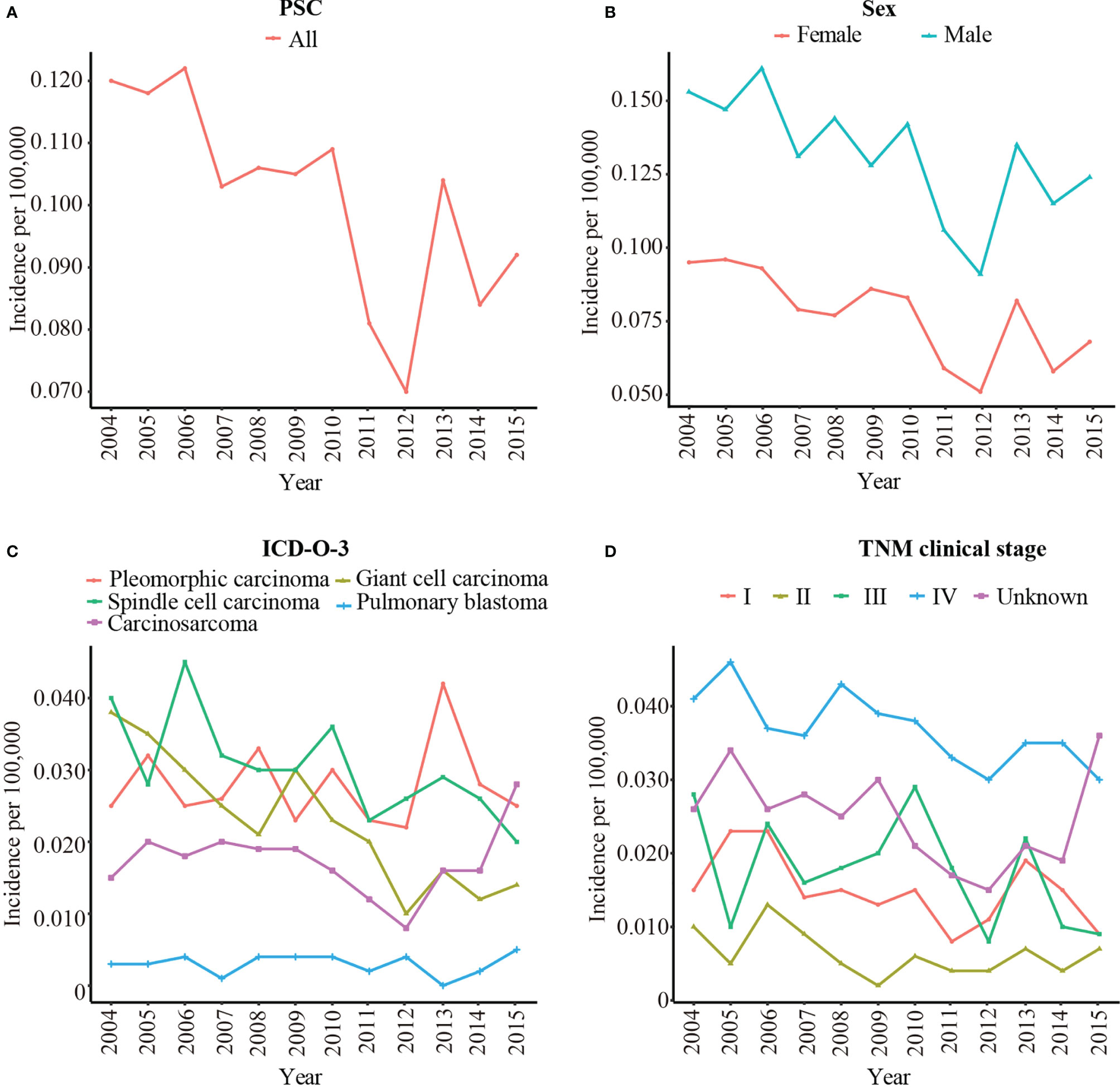
Figure 2 The age-adjusted incidence of PSC from 2004 to 2015 by sex, pathological type, and TNM clinical stage. (A) The age-adjusted incidence of all patients with PSC; (B) The age-adjusted incidence of PSC by sex; (C) The age-adjusted incidence of PSC by pathological type; (D) The age-adjusted incidence of PSC by TNM clinical stage. PSC, pulmonary sarcomatoid carcinoma; TNM clinical stage, tumor–node–metastasis clinical stage; ICD-O-3, International Classification of Disease for Oncology, 3rd Edition.
Patient Characteristics
Of the 1049 patients with PSC and 205866 patients with oNSCLC in the primary SEER database, patients who had unknown TNM clinical stage (PSC: N = 256; oNSCLC: N = 14510) were excluded. Thus, 793 patients with PSC and 191356 patients with oNSCLC were identified (Figure 1).
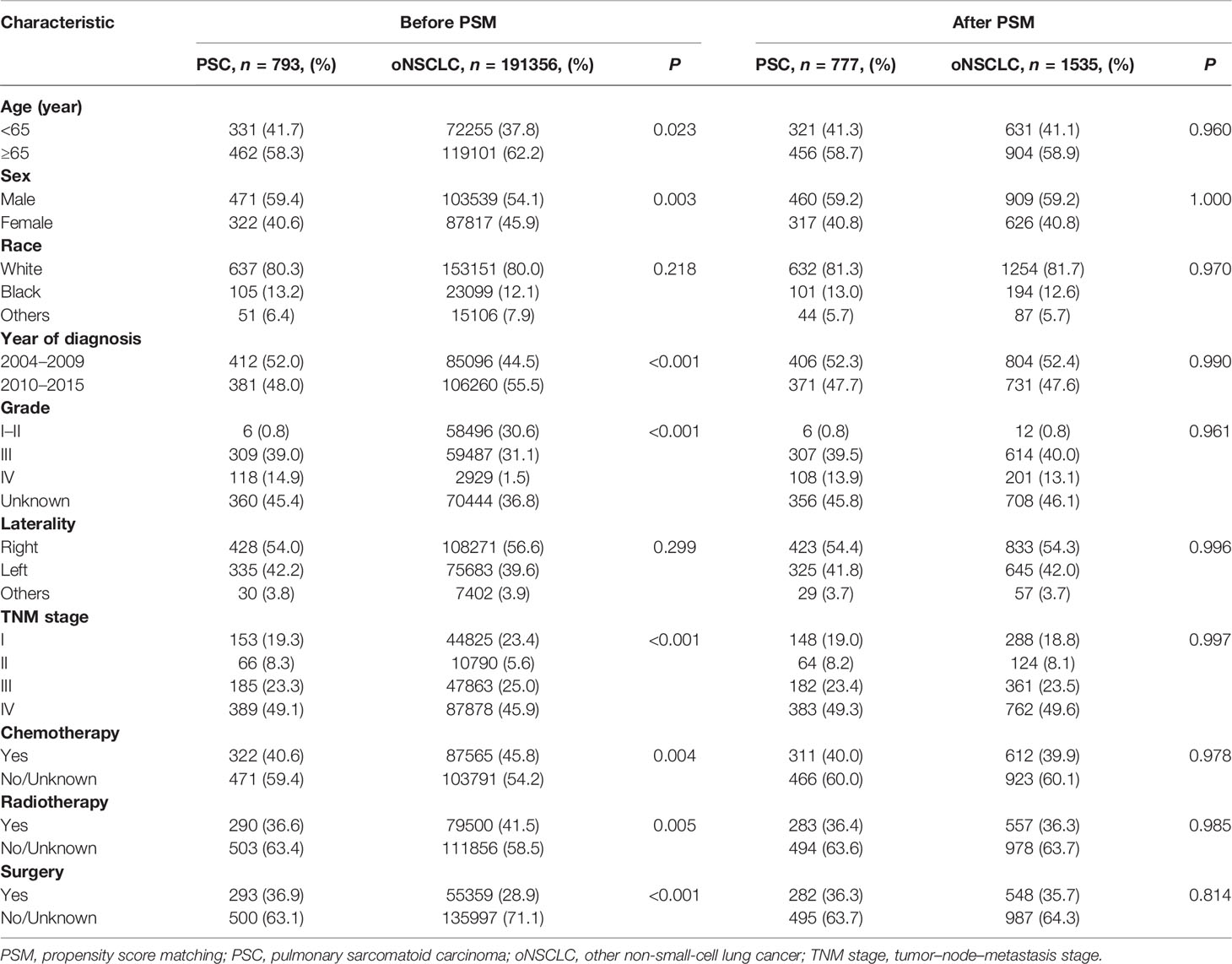
The clinicopathological characteristics between two groups were shown as follows. A majority were men, in both PSC (59.4%) and oNSCLC (54.1%, P = 0.003), and most of them were white (80.3% vs 80.0%, P = 0.218). In total, 58.3% of the patients with PSC versus 62.2% of the patients with oNSCLC were older than 65 years (P = 0.023). PSCs had a significantly lower proportion during 2010–2015 (48.0% vs 55.5%, P < 0.001). Compared with oNSCLC, significantly more patients with PSC had grade IV tumors (14.9% vs 1.5%, P < 0.001) as well as TNM clinical stage IV (49.1% vs 45.9%, P < 0.001). More patients with oNSCLC received chemotherapy (45.8% vs 40.6%, P = 0.004), and radiotherapy (41.5% vs 36.6%, P = 0.005). More patients with PSC underwent surgery (36.9% vs 28.9%, P < 0.001). No significant difference was observed in laterality (Table 1).
PSM for PSC and oNSCLC
Thus, the PSM method was used to balance all characteristics, including age, sex, race, year of diagnosis, grade, laterality, TNM clinical stage, chemotherapy, radiotherapy, and surgery between the two groups.
After PSM, the clinicopathological characteristics between 777 patients with PSC and 1535 patients with oNSCLC were shown as follows. Most of the patients were men, in both PSC (59.2%) and oNSCLC (59.2%, P = 1.000). Approximately 59% patients were older than 65 years in two groups (P = 0.960). Grade IV tumors (13.9% vs 13.1%, P = 0.961) as well as TNM clinical stage IV (49.3% vs 49.6%, P = 0.997) were balanced in two groups, and other covariates, including race, year of diagnosis, laterality, chemotherapy, radiotherapy, and surgery also showed no significantly difference (Table 1 and Figure 3).
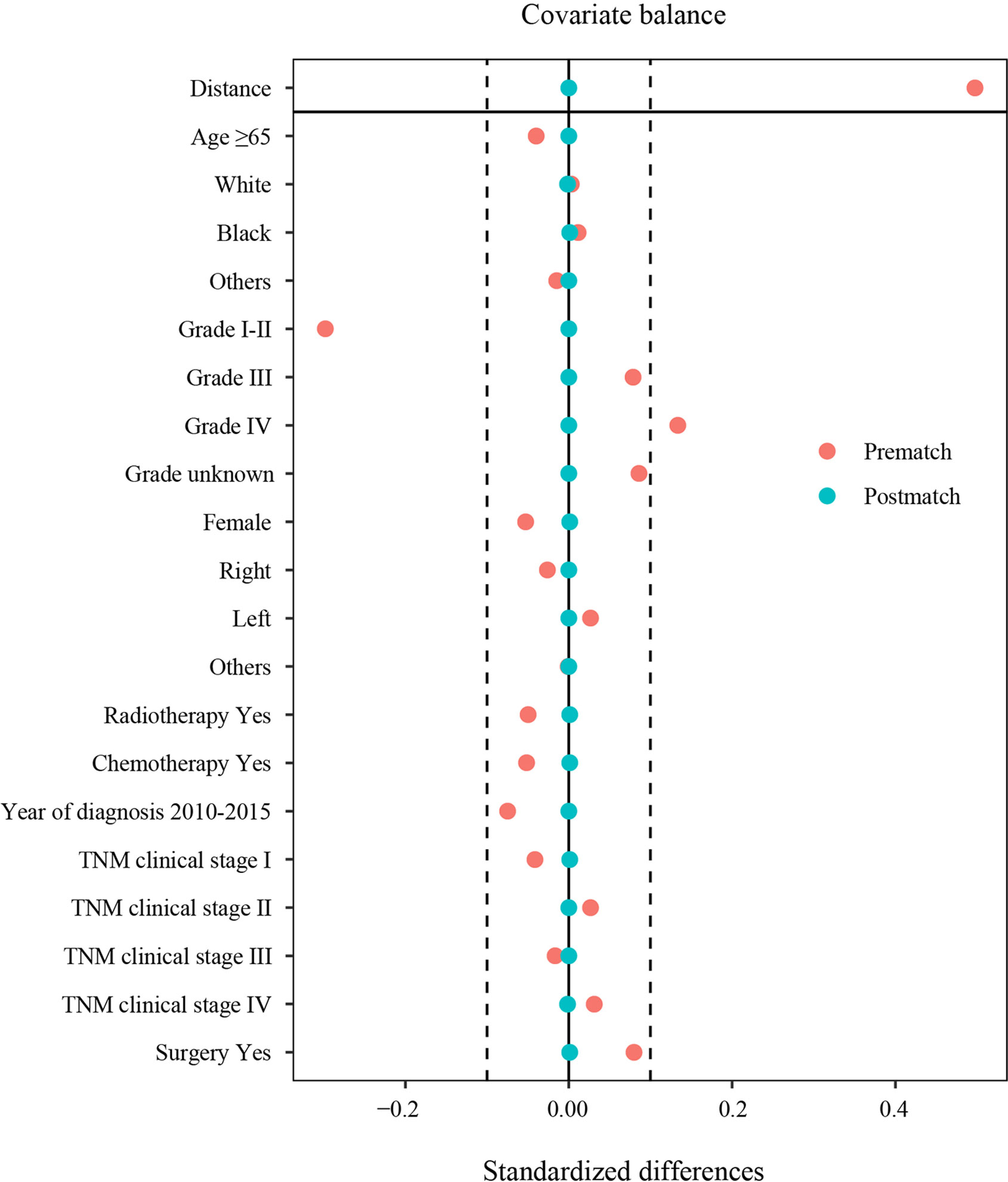
Figure 3 Standardized differences in characteristics before and after PSM. TNM clinical stage, tumor–node–metastasis clinical stage; PSM, propensity score matching.
Survival Analysis of PSC and oNSCLC
Before PSM, patients with PSC had significantly poorer OS compared with patients with oNSCLC (mOS: 5.0 months vs 12.0 months, P < 0.001) (Figure 4A). Patients with PSC who had stage I (mOS: 46.0 months vs 61.0 months, P = 0.012), stage III (mOS: 7.0 months vs 12.0 months, P = 0.003) and stage IV (mOS: 2.0 months vs 5.0 months, P < 0.001) revealed an inferior survival compared with oNSCLC, respectively. But stage II showed no difference (mOS: 14.0 months vs 27.0 months, P = 0.160; Figures 4B–E).
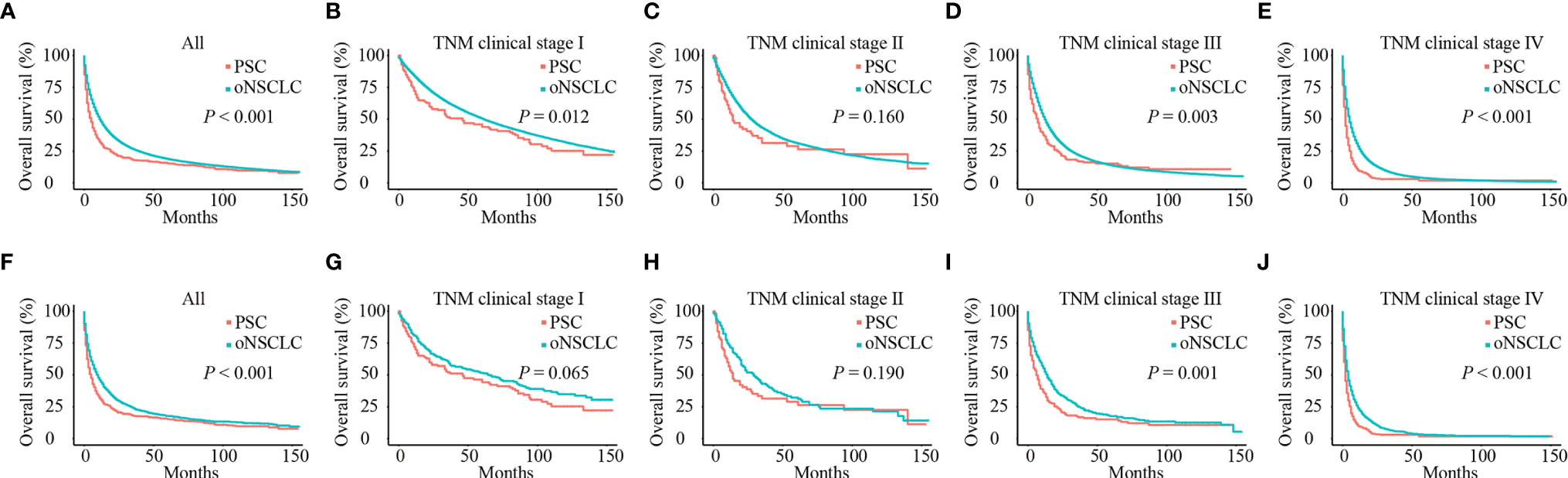
Figure 4 Survival curves of patients with PSC and oNSCLC. (A) Before PSM, OS in patients with PSC and oNSCLC; (B–E) Before PSM, OS in patients with PSC and oNSCLC at different clinical stage; (F) After PSM, OS in patients with PSC and oNSCLC; (G–J) After PSM, OS in patients with PSC and oNSCLC at different clinical stage. PSC, pulmonary sarcomatoid carcinoma; oNSCLC, other non-small-cell lung cancer; PSM, propensity score matching.
After adjusting for covariates (age, sex, year of diagnosis, grade, TNM clinical stage, chemotherapy, radiotherapy and surgery), PSC still performed worse survival (mOS: 5.0 months vs 10.0 months, P < 0.001; Figure 4F). Also, patients with PSC revealed significantly inferior survival compared with patients with oNSCLC at stage III (mOS: 6.0 months vs 14.0 months, P = 0.001) and stage IV (mOS: 2.0 months vs 4.0 months, P < 0.001; Figures 4I, J), respectively. But patients with PSC and patients with oNSCLC had stage I (mOS: 46.0 months vs 65.0 months, P = 0.065) and stage II (mOS: 14.0 months vs 29.0 months, P = 0.190) showed no difference, respectively (Figures 4G, H).
Multivariate Analysis of PSC
A total of 793 patients were included in the PSC group. With a ratio of 7:3, patients with PSC were randomly assigned to the training set (N=555) and testing set (N=238). Supplementary Table 2 lists the baseline characteristics, no significant difference was found between the two sets.
As shown (Figure 5), in the training set, age (≥65 years, HR 1.36, P = 0.003) had a correlation with worse prognosis while female (HR 0.73, P = 0.002) was associated with better OS. The TNM clinical stage (II vs I, HR 2.39, P < 0.001; III vs I, HR 2.70, P < 0.001; IV vs I, HR 5.02, P < 0.001) were also covariates having effect on OS. Patients who received chemotherapy (HR 0.46, P < 0.001) or radiotherapy (HR 0.75, P = 0.005) or underwent surgery (HR 0.40, P < 0.001) experienced superior survival compared with those who did not. Other characteristics (race, year of diagnosis, pathological type, grade, and laterality) had no statistically significant difference (P > 0.05) in the model.
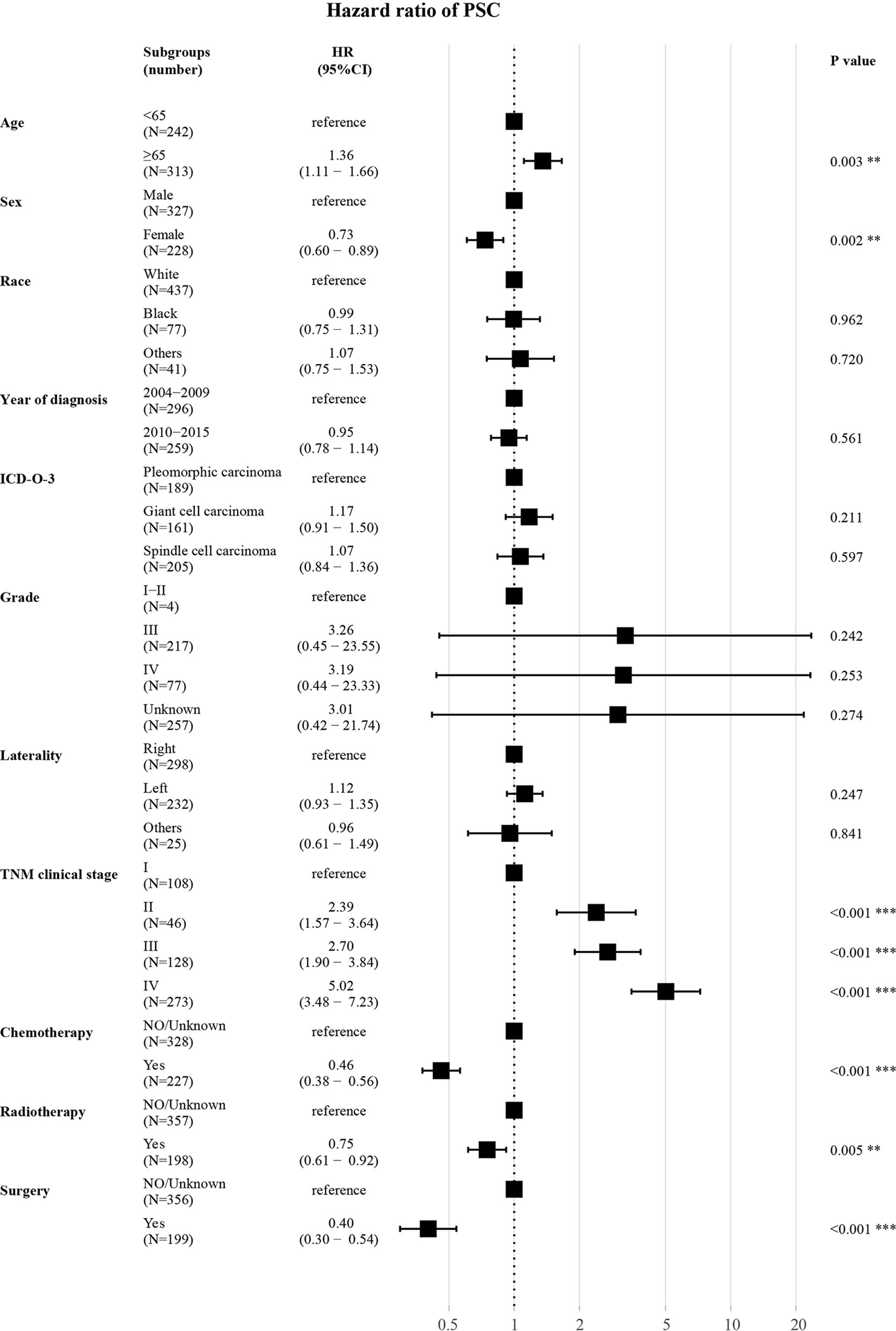
Figure 5 Forest plot for multivariate analysis of the effect of different variables. HR, hazard ratio; CI, confidence interval; ICD-O-3, International Classification of Disease for Oncology, 3rd Edition; TNM clinical stage, tumor–node–metastasis clinical stage. **P < 0.01; ***P < 0.001.
Construction and Validation of the Nomogram
The Cox regression analysis demonstrated that age, sex, TNM clinical stage, chemotherapy, radiotherapy, and surgery were independent prognostic factors for OS. So a nomogram was established to predict 1-year survival based on the results (Figure 6A). The TNM clinical stage was the largest contributing covariate to prognosis, followed by surgery and chemotherapy. In addition, age, sex and radiotherapy also presented an impact on OS. Each subtype of significant characteristics corresponded to a unique point. The total points of every patient were calculated. It was convenient to estimate the probability of 1-year survival by locating it on the scale. Supplementary Table 3 shows the detailed score for each characteristic.
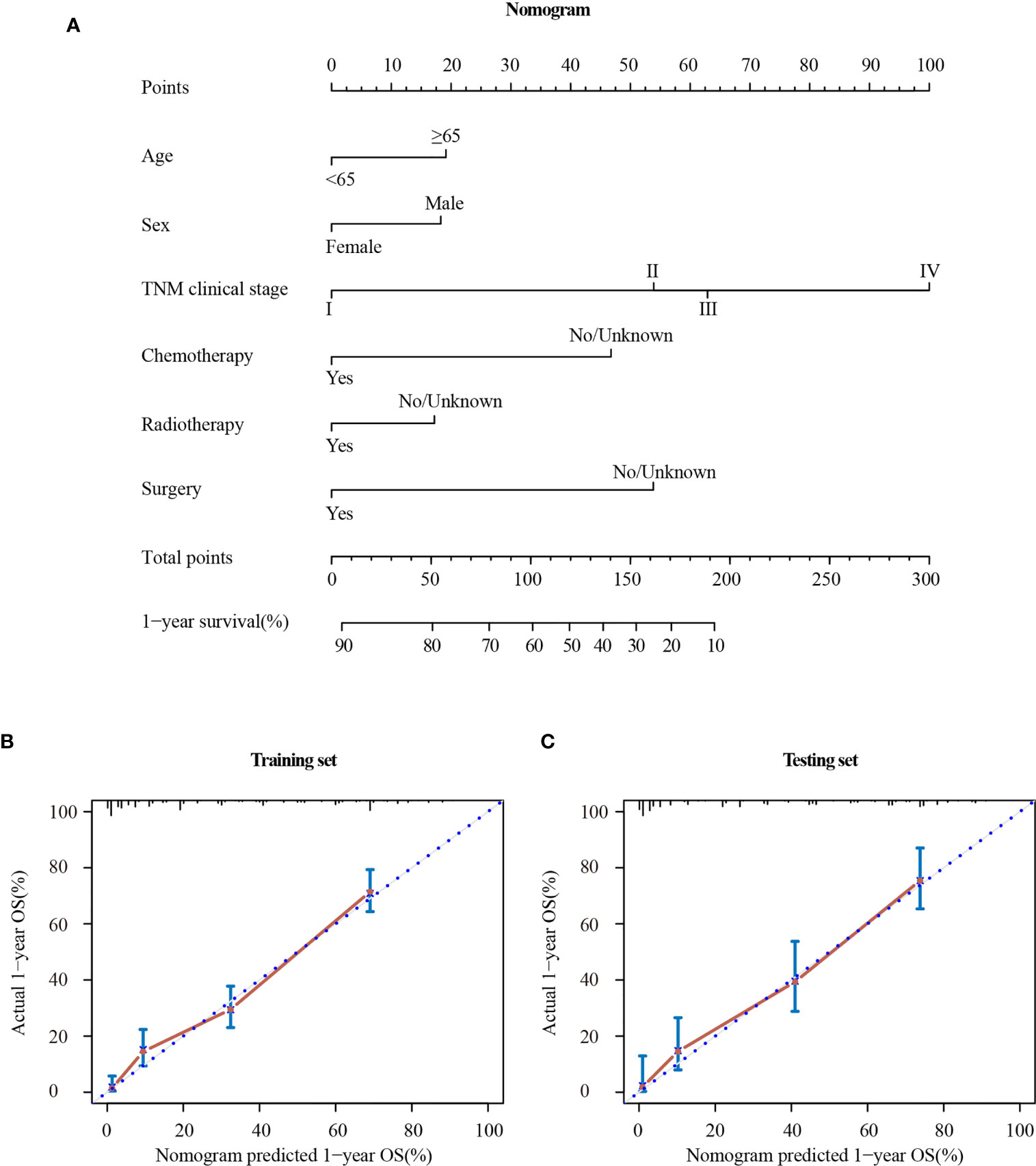
Figure 6 Nomogram (A) and calibration curve in the training set (B) and testing set (C). TNM clinical stage, tumor–node–metastasis clinical stage.
The stability of the nomogram was validated using the calibration plot in the training set (Figure 6B) and testing set (Figure 6C), respectively. The calibration curves displayed high internal and external consistency with the actual observation for 1-year survival, and achieving C-indexs of 0.775 and 0.790 for the training and testing set, respectively.
Comparison of the Predictive Accuracy for OS Between Nomogram and TNM Clinical Stage
Compared with the TNM clinical stage, in the training test, the AUCs of nomogram and TNM clinical stage were 0.867 and 0.813, respectively; and in the testing set, the AUCs were 0.871 and 0.762, respectively. As shown in Figure 7, the nomogram had better prediction accuracy for the 1-year survival probability compared to TNM clinical stage.
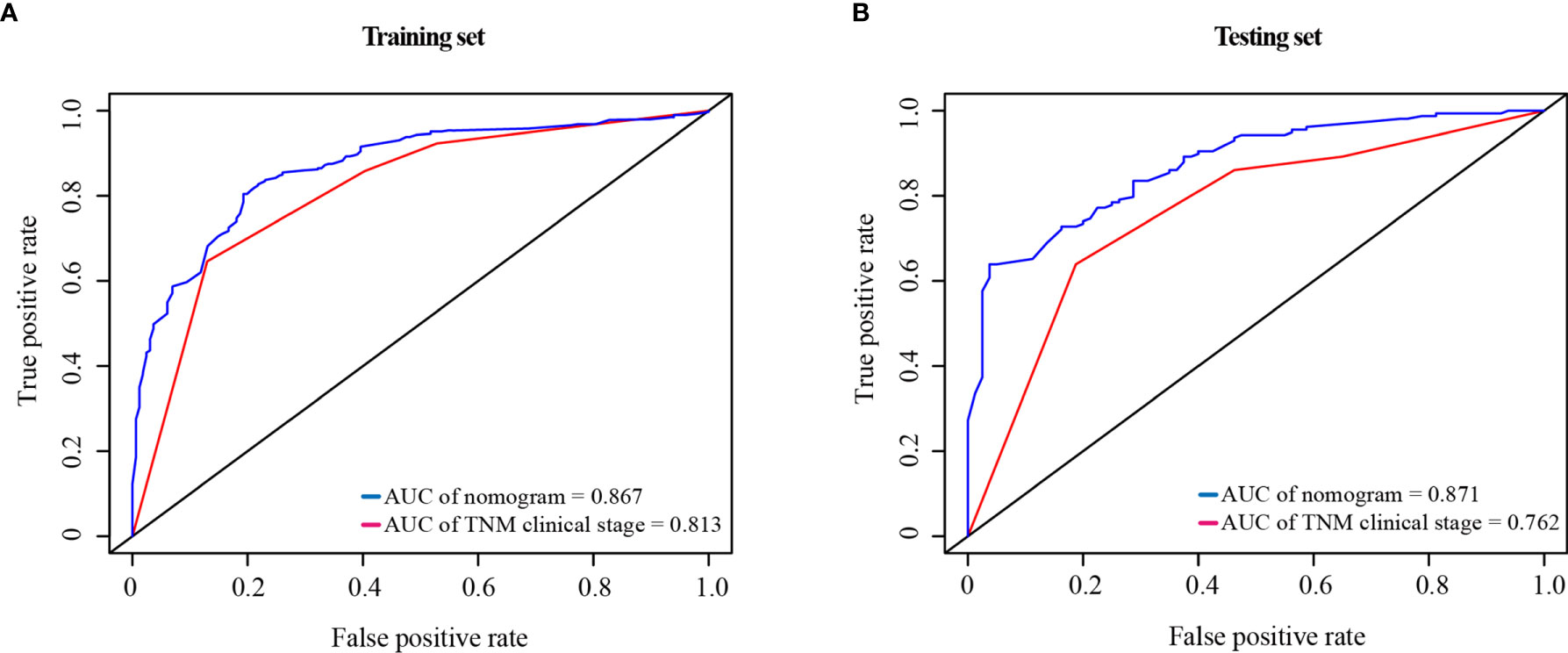
Figure 7 ROC curve of the nomogram and the TNM clinical stage in the training set (A) and testing set (B). AUC, area under the curve; ROC, receiver operating characteristic; TNM clinical stage, tumor–node–metastasis clinical stage.
Discussion
This large retrospective study showed that PSC was a rare cancer, accounting for less than 0.5% of oNSCLC, as described in previous studies (4, 5). It is remarkable that the incidence of PSC was slowly decreased. As for pathological, the incidence of carcinosarcoma was significant increased. PSC occurred more frequently in the elder (≥65 years) and male individuals (9). In the current study, PSC had a highly aggressive behavior, with a significantly higher proportion of poorly differentiated tumors (grade IV: 14.9% vs 1.5%, P <0.001) compared with oNSCLC, and up to 49.1% of patients were in stage IV when diagnosed.
PSC is difficult to diagnose, leading to a poor prognosis (1, 4). In small-sample studies on PSC, mOS was 5–8.5 months (10–14). In previous SEER-based studies, mOS was 6.0 months for patients with all stages and 3.0 months for patients with advanced disease (4, 6). In this study, mOS was 5.0 months for PSC and 12.0 months for oNSCLC (P < 0.001) before PSM. After equalizing significantly different characteristics in PSC and oNSCLC, PSCs were still found to have a significantly poorer clinical outcome compared with oNSCLC (mOS: 5.0 m vs 10.0 m, P < 0.001).
The multivariate Cox proportional hazards model revealed that elderly, male patients with advanced clinical stage had a worse prognosis (2, 15), whereas receiving chemotherapy, radiotherapy, or surgery could prolong OS. At present, the standard treatment of PSC is controversial (5, 9). Surgery in the early-stage PSC has been demonstrated to provide optimal OS benefit, but a high risk of recurrence and adjuvant chemotherapy should be considered (2, 5, 6, 15). Previous studies indicated that adjuvant chemotherapy after surgery was effective in improving survival outcomes (15, 16), and large population-based studies also revealed the benefit of chemotherapy (7). On the contrary, in a study on 69 patients with PSC, Liang et al. reported that adjuvant chemotherapy could not improve OS (5). PSC was a chemorefractory cancer in previous studies (9, 11, 17). Patients with PSC who received first-line platinum-based chemotherapy did not experience significant benefits (mOS: 7.0 months vs 5.3 months, P = 0.096) (18). Few studies on the effect of radiotherapy on the prognosis of PSC and a small-sample study reported that patients who received radiotherapy had a worse mOS (5.0 m vs 6.0 m, P < 0.001) (6, 19). However, a retrospective study based on SEER showed that radiotherapy improved the survival in stage I-III patients with PSC (8). In this multivariate analysis, both chemotherapy (HR 0.46, P < 0.001) and radiotherapy (HR 0.75, P = 0.005) were protective factors and improved the survival. By far, data showing that chemotherapy or radiotherapy prolongs survival is insufficient, and requires further prospective research.
In recent studies, MET exon 14 skipping mutations were found to be enriched in PSC, and the incidence rate was as high as 31.8% (20), which could be a potential target for therapy using MET inhibitors, such as volitinib, capmatinib, and tepotinib (21–23). In addition, patients with PSC had an increased tendency for the high expression of tumor mutation burden (>20 mutations per Mb) compared with those with oNSCLC (20% vs 14%, P = 0.056) (22). At the same time, PSC also had a high expression of PD-L1, suggesting that targeting PD-1/PD-L1 might be a potential treatment regimen for PSC. Patients with PSC who received PD-1/PD-L1 inhibitors exhibited high response rates and prolonged overall survival (24–31). Also some novel biomarkers were investigated to predict the survival outcome of PSC, such as CD8+ tumor-infiltrating lymphocytes, the epithelial−mesenchymal transition transcription factors (Twist1), the change of neutrophil-to-lymphocyte ratio, KRAS mutations and c-MET overexpression (32–36).
In this study, six features, including age, sex, TNM clinical stage, chemotherapy, radiotherapy, and surgery, were used to construct a nomogram prognostic model. The total score was calculated using the quantitative score of each feature, and the 1-year survival rate was predicted scientifically and accurately. The C-indexs of this model were 0.775 and 0.790 in internal and external validation, respectively, indicating good agreement between predicted and actual 1-year survival. Next, the nomogram model was compared with the conventional TNM clinical stage and found to be superior in the 1-year survival of AUC both in the training set and testing set, indicating a more comprehensive and accurate prediction. This model could be used to individualize prognostic assessment and might serve an effective diagnostic tool for making treatment-related decisions (37, 38).
This study had several limitations. First, the variables enrolled were restricted; some important characteristics related to prognosis were not included in this study, such as smoking status, performance status score, gene mutation detection by next generation sequencing, MET expression detection, and PD-L1 immunohistochemistry assay. Second, treatment information was limited, without target therapy, immunotherapy, etc. The database only contained the status of surgery, chemotherapy, and radiotherapy, but some of which were not known. Third, the construction of this model was based on retrospective data and requires further confirmation.
In conclusion, the incidence of PSC was slowly decreased, and for pathological subgroups, the incidence of carcinosarcoma was increased. PSC had a significantly poor prognosis compared with oNSCLC. The nomogram constructed in this study accurately predicted the prognosis of PSC, and performed better than the TNM clinical stage. This model is expected to help pathologist and oncologist in designing clinical strategies.
Data Availability Statement
Publicly available datasets were analyzed in this study. This data can be found here: The SEER 18 registries (www.seer.cancer.gov) of the National Cancer Institute using the SEER*Stat software (SEER*Stat 8.3.6).
Author Contributions
JS, MC, and QY designed the study. MC, QY, and ZX contributed to data selection and assembly. BL, FL, and YY analysed data. MC and QY were involved in drafting the manuscript. JS critically revised the manuscript. All authors contributed to the article and approved the submitted version.
Funding
The study was supported by the National Natural Science Foundation of China (81773245, 81972858), the Clinical Research Project of Army Medical University (2018XLC1010), and the Chongqing Science and Technology Leading Talents Program (cstccxljrc201910).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors are grateful to all the staff in the National Cancer Institute (USA) for their contribution to the SEER program.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.630885/full#supplementary-material
References
1. Ung M, Rouquette I, Filleron T, Taillandy K, Brouchet L, Bennouna J, et al. Characteristics and Clinical Outcomes of Sarcomatoid Carcinoma of the Lung. Clin Lung Cancer (2016) 17(5):391–7. doi: 10.1016/j.cllc.2016.03.001
2. Roesel C, Terjung S, Weinreich G, Hager T, Chalvatzoulis E, Metzenmacher M, et al. Sarcomatoid Carcinoma of the Lung: A Rare Histological Subtype of Non-Small Cell Lung Cancer With a Poor Prognosis Even At Earlier Tumour Stages. Interact Cardiov Th (2016) 24(3):407–13. doi: 10.1093/icvts/ivw392
3. Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM, Beasley MB, et al. The 2015 World Health Organization Classification of Lung Tumors. J Thorac Oncol (2015) 10(9):1243–60. doi: 10.1097/JTO.0000000000000630
4. Yendamuri S, Caty L, Pine M, Adem S, Bogner P, Miller A, et al. Outcomes of Sarcomatoid Carcinoma of the Lung: A Surveillance, Epidemiology, and End Results Database Analysis. Surgery (2012) 152(3):397–402. doi: 10.1016/j.surg.2012.05.007
5. Lin Y, Yang H, Cai Q, Wang D, Rao H, Lin S, et al. Characteristics and Prognostic Analysis of 69 Patients With Pulmonary Sarcomatoid Carcinoma. Am J Clin Oncol (2016) 39(3):215–22. doi: 10.1097/COC.0000000000000101
6. Rahouma M, Kamel M, Narula N, Nasar A, Harrison S, Lee B, et al. Pulmonary Sarcomatoid Carcinoma: An Analysis of a Rare Cancer From the Surveillance, Epidemiology, and End Results Database. Eur J Cardiothorac Surg (2018) 53(4):828–34. doi: 10.1093/ejcts/ezx417
7. Sun L, Dai J, Chen Y, Duan L, He W, Chen Q, et al. Pulmonary Sarcomatoid Carcinoma: Experience From SEER Database and Shanghai Pulmonary Hospital. Ann Thorac Surg (2020) 110(2):406–13. doi: 10.1016/j.athoracsur.2020.02.071
8. Liang X, Cheng Y, Yuan Z, Yan Z, Li Q, Huang Y, et al. Clinical, Pathological and Treatment Factors Associated With the Survival of Patients With Pulmonary Sarcomatoid Carcinoma. Oncol Lett (2020) 19(6):4031–9. doi: 10.3892/ol.2020.11472
9. Ouziane I, Boutayeb S, Mrabti H, Lalya I, Rimani M, Errihani H. Sarcomatoid Carcinoma of the Lung: A Model of Resistance of Chemotherapy. N Am J Med Sci (2014) 6(7):342–5. doi: 10.4103/1947-2714.136920
10. Giroux Leprieur E, Antoine M, Vieira T, Duruisseaux M, Poulot V, Rabbe N, et al. Clinical and Molecular Features in Patients With Advanced Non-Small-Cell Lung Carcinoma Refractory to First-Line Platinum-Based Chemotherapy. Lung Cancer (2013) 79(2):167–72. doi: 10.1016/j.lungcan.2012.10.010
11. Bae H, Min HS, Lee S, Kim D, Chung DH, Lee J, et al. Palliative Chemotherapy for Pulmonary Pleomorphic Carcinoma. Lung Cancer (2007) 58(1):112–5. doi: 10.1016/j.lungcan.2007.05.006
12. Hong JY, Choi MK, Uhm JE, Park MJ, Lee J, Park YH, et al. The Role of Palliative Chemotherapy for Advanced Pulmonary Pleomorphic Carcinoma. Med Oncol (2009) 26(3):287–91. doi: 10.1007/s12032-008-9117-4
13. Ito K, Oizumi S, Fukumoto S, Harada M, Ishida T, Fujita Y, et al. Clinical Characteristics of Pleomorphic Carcinoma of the Lung. Lung Cancer (2010) 68(2):204–10. doi: 10.1016/j.lungcan.2009.06.002
14. Kaira K, Horie Y, Ayabe E, Murakami H, Takahashi T, Tsuya A, et al. Pulmonary Pleomorphic Carcinoma: A Clinicopathological Study Including EGFR Mutation Analysis. J Thorac Oncol (2010) 5(4):460–5. doi: 10.1097/JTO.0b013e3181ce3e3c
15. Huang S, Shen S, Li X. Pulmonary Sarcomatoid Carcinoma: A Clinicopathologic Study and Prognostic Analysis of 51 Cases. World J Surg Oncol (2013) 11(1):252. doi: 10.1186/1477-7819-11-252
16. Karim NA, Schuster J, Eldessouki I, Gaber O, Namad T, Wang J, et al. Pulmonary Sarcomatoid Carcinoma: University of Cincinnati Experience. Oncotarget (2018) 9(3):4102–8. doi: 10.18632/oncotarget.23468
17. Raveglia F, Mezzetti M, Panigalli T, Furia S, Giuliani L, Conforti S, et al. Personal Experience in Surgical Management of Pulmonary Pleomorphic Carcinoma. Ann Thorac Surg (2004) 78(5):1742–7. doi: 10.1016/j.athoracsur.2004.04.084
18. Vieira T, Girard N, Ung M, Monnet I, Cazes A, Bonnette P, et al. Efficacy of First-Line Chemotherapy in Patients With Advanced Lung Sarcomatoid Carcinoma. J Thorac Oncol (2013) 8(12):1574–7. doi: 10.1097/01.JTO.0000437008.00554.90
19. Nakajima M, Kasai T, Hashimoto H, Iwata Y, Manabe H. Sarcomatoid Carcinoma of the Lung: A Clinicopathologic Study of 37 Cases. Cancer Am Cancer Soc (1999) 86(4):608–16. doi: 10.1002/(SICI)1097-0142(19990815)86:4<608::AID-CNCR9>3.0.CO;2-1
20. Tong JH, Yeung SF, Chan AWH, Chung LY, Chau SL, Lung RWM, et al. Met Amplification and Exon 14 Splice Site Mutation Define Unique Molecular Subgroups of Non–Small Cell Lung Carcinoma With Poor Prognosis. Clin Cancer Res (2016) 22(12):3048–56. doi: 10.1158/1078-0432.CCR-15-2061
21. Lee C, Usenko D, Frampton GM, McMahon C, Ali SM, Weiss J. Met 14 Deletion in Sarcomatoid Non-Small-Cell Lung Cancer Detected by Next-Generation Sequencing and Successfully Treated With a MET Inhibitor. J Thorac Oncol (2015) 10(12):e113–4. doi: 10.1097/JTO.0000000000000645
22. Schrock AB, Li SD, Frampton GM, Suh J, Braun E, Mehra R, et al. Pulmonary Sarcomatoid Carcinomas Commonly Harbor Either Potentially Targetable Genomic Alterations or High Tumor Mutational Burden as Observed by Comprehensive Genomic Profiling. J Thorac Oncol (2017) 12(6):932–42. doi: 10.1016/j.jtho.2017.03.005
23. Alì G, Bruno R, Poma AM, Affinito O, Monticelli A, Piaggi P, et al. Whole Transcriptome Targeted Gene Quantification Provides New Insights on Pulmonary Sarcomatoid Carcinomas. Sci Rep Uk (2019) 9(1):3536. doi: 10.1038/s41598-019-40016-8
24. Qin J, Chen B, Li C, Yan J, Lu H. Genetic Heterogeneity and Predictive Biomarker for Pulmonary Sarcomatoid Carcinomas. Cancer Genet Ny (2021) 250-251:12–9. doi: 10.1016/j.cancergen.2020.11.004
25. Sukrithan V, Sandler J, Gucalp R, Gralla R, Halmos B. Immune Checkpoint Blockade Is Associated With Durable Responses in Pulmonary Sarcomatoid Carcinoma. Clin Lung Cancer (2019) 20(3):e242–6. doi: 10.1016/j.cllc.2018.12.013
26. Babacan NA, Pina IB, Signorelli D, Prelaj A, Garassino MC, Tanvetyanon T. Relationship Between Programmed Death Receptor-Ligand 1 Expression and Response to Checkpoint Inhibitor Immunotherapy in Pulmonary Sarcomatoid Carcinoma: A Pooled Analysis. Clin Lung Cancer (2020) 21(5):e456–63. doi: 10.1016/j.cllc.2020.02.022
27. Velcheti V, Rimm DL, Schalper KA. Sarcomatoid Lung Carcinomas Show High Levels of Programmed Death Ligand-1 (PD-L1). J Thorac Oncol (2013) 8(6):803–5. doi: 10.1097/JTO.0b013e318292be18
28. Vieira T, Antoine M, Hamard C, Fallet V, Duruisseaux M, Rabbe N, et al. Sarcomatoid Lung Carcinomas Show High Levels of Programmed Death Ligand-1 (PD-L1) and Strong Immune-Cell Infiltration by TCD3 Cells and Macrophages. Lung Cancer (2016) 98:51–8. doi: 10.1016/j.lungcan.2016.05.013
29. Kotlowska MP, Rueda AG, Olmedo ME, Benito A, Roldán AS, Fernandez Méndez MA, et al. Efficacy of Immunotherapy in Sarcomatoid Lung Cancer, A Case Report and Literature Review. Respir Med Case Rep (2019) 26:310–4. doi: 10.1016/j.rmcr.2019.02.017
30. Roesel C, Kambartel K, Kopeika U, Berzins A, Voshaar T, Krbek T. Lazarus-Type Tumour Response to Therapy With Nivolumab for Sarcomatoid Carcinomas of the Lung. Curr Oncol (2019) 26(2):270–3. doi: 10.3747/co.26.4377
31. Domblides C, Leroy K, Monnet I, Mazieres J, Barlesi F, Gounant V, et al. Efficacy of Immune Checkpoint Inhibitors in Lung Sarcomatoid Carcinoma. J Thorac Oncol (2020) 15(5):860–6. doi: 10.1016/j.jtho.2020.01.014
32. Mignard X, Ruppert A, Antoine M, Vasseur J, Girard N, Mazières J, et al. C-MET Overexpression as a Poor Predictor of MET Amplifications or Exon 14 Mutations in Lung Sarcomatoid Carcinomas. J Thorac Oncol (2018) 13(12):1962–7. doi: 10.1016/j.jtho.2018.08.008
33. Chen J, He Q, Liu J, Xiao Y, Xiao C, Chen K, et al. CD8+ Tumor-Infiltrating Lymphocytes as a Novel Prognostic Biomarker in Lung Sarcomatoid Carcinoma, A Rare Subtype of Lung Cancer. Cancer Manag Res (2018) 10:3505–11. doi: 10.2147/CMAR.S169074
34. Liu T, Zhao X, Zheng X, Zheng Y, Dong X, Zhao N, et al. The EMT Transcription Factor, Twist1, as a Novel Therapeutic Target for Pulmonary Sarcomatoid Carcinomas. Int J Oncol (2020) 56(3):750–60. doi: 10.3892/ijo.2020.4972
35. Ding Y, Shao Y, Na C, Yin JC, Hua H, Tao R, et al. Genetic Characterisation of Sarcomatoid Carcinomas Reveals Multiple Novel Actionable Mutations and Identifies KRAS Mutation as a Biomarker of Poor Prognosis. J Med Genet (2020) 2020:107083. doi: 10.1136/jmedgenet-2020-107083
36. Seong YW, Han SJ, Jung W, Jeon JH, Cho S, Jheon S, et al. Perioperative Change in Neutrophil-to-Lymphocyte Ratio (NLR) Is a Prognostic Factor in Patients With Completely Resected Primary Pulmonary Sarcomatoid Carcinoma. J Thorac Dis (2019) 11(3):819–26. doi: 10.21037/jtd.2019.02.02
37. Wang Y, Li J, Xia Y, Gong R, Wang K, Yan Z, et al. Prognostic Nomogram for Intrahepatic Cholangiocarcinoma After Partial Hepatectomy. J Clin Oncol (2013) 31(9):1188–95. doi: 10.1200/JCO.2012.41.5984
Keywords: clinical characteristics, prediction model, nomogram, pulmonary sarcomatoid carcinoma, SEER database
Citation: Chen M, Yang Q, Xu Z, Luo B, Li F, Yu Y and Sun J (2021) Survival Analysis and Prediction Model for Pulmonary Sarcomatoid Carcinoma Based on SEER Database. Front. Oncol. 11:630885. doi: 10.3389/fonc.2021.630885
Received: 18 November 2020; Accepted: 11 May 2021;
Published: 31 May 2021.
Edited by:
Giulia Veronesi, Humanitas Research Hospital, ItalyReviewed by:
Alessandro Russo, A. O. Papardo, ItalyConghua Xie, Zhongnan Hospital of Wuhan University, China
Copyright © 2021 Chen, Yang, Xu, Luo, Li, Yu and Sun. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jianguo Sun, c3VuamcwOUBhbGl5dW4uY29t
†These authors have contributed equally to this work
 Mingjing Chen
Mingjing Chen Qiao Yang
Qiao Yang Zihan Xu1,4
Zihan Xu1,4 Jianguo Sun
Jianguo Sun