- 1Department of Molecular Pathology, The Affiliated Cancer Hospital of Zhengzhou University and Henan Cancer Hospital, Zhengzhou, China
- 2Henan Key Laboratory of Molecular Pathology, Zhengzhou, China
- 3Department of Gynecologic Oncology, The Affiliated Cancer Hospital of Zhengzhou University and Henan Cancer Hospital, Zhengzhou, China
- 4Department of Pathophysiology, School of Basic Medical Science, Zhengzhou University, Zhengzhou, China
- 5Department of Pathology, The Affiliated Cancer Hospital of Zhengzhou University and Henan Cancer Hospital, Zhengzhou, China
Predisposition of germline BRCA1/2 mutations (gBRCAMUT) increases the risk of breast and ovarian cancer in females, but the mutation prevalence and spectrum are highly ethnicity-specific with different recurrent mutations being reported in different populations. Hereby, we performed hybridization-based target sequencing of BRCA1/2 in 530 ovarian cancer patients from Henan, the central region of China, followed by haplotype analysis of six short tandem repeat (STR) markers in the patients with recurrent mutations to determine their founder effect. About 28.3% (150/530) of the OC patients in our cohort harbored gBRCAMUT; of the 151 mutations, 117 in BRCA1 and 34 in BRCA2, identified in this study, BRCA1:c.5470_5477del, c.981_982del, and c.4065_4068del are the top three mutants, recurrently detected in eight, seven, and six independent patients respectively. Haplotype analysis identified a region of 0.6 MB genomic length covering BRCA1 highly conserved across all eight carriers of BRCA1:c.5470_5477del, but not c.981_982del, suggesting a consequence of founder effect. Retrospective analysis in a subgroup of serous ovarian cancer patients revealed gBRCAMUT status was not associated with the progression-free survival (PFS); instead, an expression of Ki-67% ≥50% was associated with a shorter PFS (p = 0.041). In conclusion, patients with pathogenic or likely pathogenic gBRCAMUT account for 28.3% of the OC cases from Henan, and BRCA1:c.5470_5477del, the most frequently detected mutation in Henan patients, is a founder mutation in the population.
Introduction
Ovarian cancer (OC) is the seventh commonest cancer in women, with a global morbidity and mortality rate of 6.6 and 3.9 per 100,000 population respectively, and the incidence has been increasing moderately over the last decade (1, 2). OC usually doesn’t cause noticeable symptoms and there’s no effective screening method; therefore, many patients are not diagnosed until the late stage of the disease, resulting in a relatively high mortality rate (3, 4). Several factors have been associated with the risk of developing OC, e.g., endometriosis, family history, inherited genetic aberrations, etc.; particularly, 39–44% of women who inherited a pathogenic mutation in Breast Cancer 1 (BRCA1) and 11–17% of women with a pathogenic mutation in Breast Cancer 2 (BRCA2) will develop OC by 70–80 years of age (5, 6). BRCA1 and BRCA2, located on 17q21 and 13q13 respectively, are well-established tumor suppressor genes that their protein products are of paramount importance in maintaining the genomic stability and integrity by facilitating error-free homologous recombination repair (HRR) of DNA double strand breaks (DSBs). Multiple studies have revealed that mutations in BRCA1 and BRCA2 are prevalent throughout the whole coding region and the flanking splice sites without any hot spots (7, 8); meanwhile, some mutations have been reported in specific populations with relatively high frequencies as a consequence of founder effect, i.e., the 185delAG in Ashkenazi Jews (9), the 2804delAA in Dutch (10), and E881X in south Africans (11) etc. Khoo et al. first reported the BRCA1:c.1081del as a founder mutation in southern Chinese OC patients in 2002 (12). However, other groups to investigate the prevalence of germline BRCA1/2 mutations (gBRCA1/2MUT) in Chinese OC patients revealed different mutations, i.e., BRCA1:c.5470_5477del and c.981_982del etc. (7, 13) Previous dispute, probably caused by different places of origin of the patients, inspired us to identify the BRCA1/2 founder mutations in Henan OC patients; this is because Henan, located in the middle of China with over 100 million population mainly composed of Han Chinese, is the origin of Chinese civilization.
Material and Methods
Study Population
Patients diagnosed as ovarian cancer and referred to BRCA1/2 mutation test in Henan Cancer Hospital from March 2018 to May 2020 were included in this study. Clinical information of these patients was retrieved from their medical records, including disease on-set age, location of the primary lesion, family history, immunohistochemistry (IHC) staining, treatment strategy, and response. Family history was defined as at least one first- or second-degree relative has been diagnosed as breast, ovarian cancer, prostate cancer or pancreatic cancer. Progression of the disease was evaluated according to Response Evaluation Criteria in Solid Tumors (RECIST) 1.1 criteria. The study was approved by the Ethics Committee of Henan Cancer Hospital and written informed consent was obtained from all patients.
DNA Extraction and BRCA Mutation Detection
Genomic DNA was extracted from 500 µl peripheral-blood using QIAamp DNA isolation kit (QIAGEN, Germany) according to the manufacturer’s instructions. DNA concentration was determined by using the Qubit dsDNA HS assay (Life Technologies, the U.S.), and purity was evaluated with NanoDrop 2000 UV-Vis Spectrophotometer (Thermo Scientific, the U.S.) by measuring the ratio of absorbance at 260 and 280 nm. A total of 200 ng genomic DNA was used for library construction. Briefly, the harvested DNA was first sheared to size of approximately 300 bp by using the Bioruptor sonication device (Diagenode, U.S.A.); then the fragmented DNA was blunt-end-repaired and A-tailed to ligate with adapters, followed by PCR amplification and purification. Target-enriched library was prepared by using the cancer susceptible gene detection kit according to the instructions from the manufacturer (Novogene, China), covering a 0.26 Mb genomic region of 45 breast/ovarian cancer related genes. The enriched library was then processed for sequencing on a NextSeq550 sequencer (Illumina, the U.S.) generating paired-end reads of 150 bp to a targeted coverage of >500 unique reads.
Assessment of Variants’ Pathogenicity
Pathogenicity of the variants detected in BRCA1/2 of this study was evaluated according to the American College of Medical Genetics and Genomics (ACMG) guideline (14) and Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) criteria (v 2.5.1). Classification of the variants was performed by two clinical geneticists independently. Annotations of the variants followed the Human Genome Variant Society (HGVS) recommendations (15).
Genotype Haplotype Analysis
The individuals with recurrent mutations and from irrelevant families were genotyped at six different polymorphic short tandem repeats (STR) loci adjacent to BRCA1 on chromosome17, including D17S951, 17S1789, D17S846, D17S1818, D17S1327, and D17S1320. Sequences of the primers used in this study were obtained from the UCSC genome browser (http://www.genome.ucsc.edu/). PCR reaction was performed using fluorescently end-labeled primers with (Supplementary Table 1) the following program: 95°C for 3 min, followed by 10 cycles of 94°C for 30 s, 60°C for 30 s, and 72°C for 30 s, then 35 cycles of 94°C for 30 s, 55°C for 30 s and 72°C for 30 s, finally end with 72°C for 5 min. The amplicon was processed for size fractionation on a 3730xl Genetic Analyzer (Applied Biosystems, the U.S.) and analyzed using Genemapper™ software (Thermo Fisher Scientific, the U.S.) by Sangon Biotech. (Sangon, China).
Statistical Analysis
A chi-square test was used to determine the statistical significance for categorial variables. Disease on-set age distribution of the patients was compared by using a log-rank test stratified according to their mutation status. Median progression-free survival of the patients was calculated with the Kaplan–Meier method and compared using the log-rank test. A p-value of <0.05 was considered as significant.
Data Deposition
According to the Management of Human Genetic Resources in the People’s Republic of China, sequencing data related to this study is available from the corresponding author upon reasonable request.
Results
Patient Characteristics and the Association With BRCA Germline Mutation
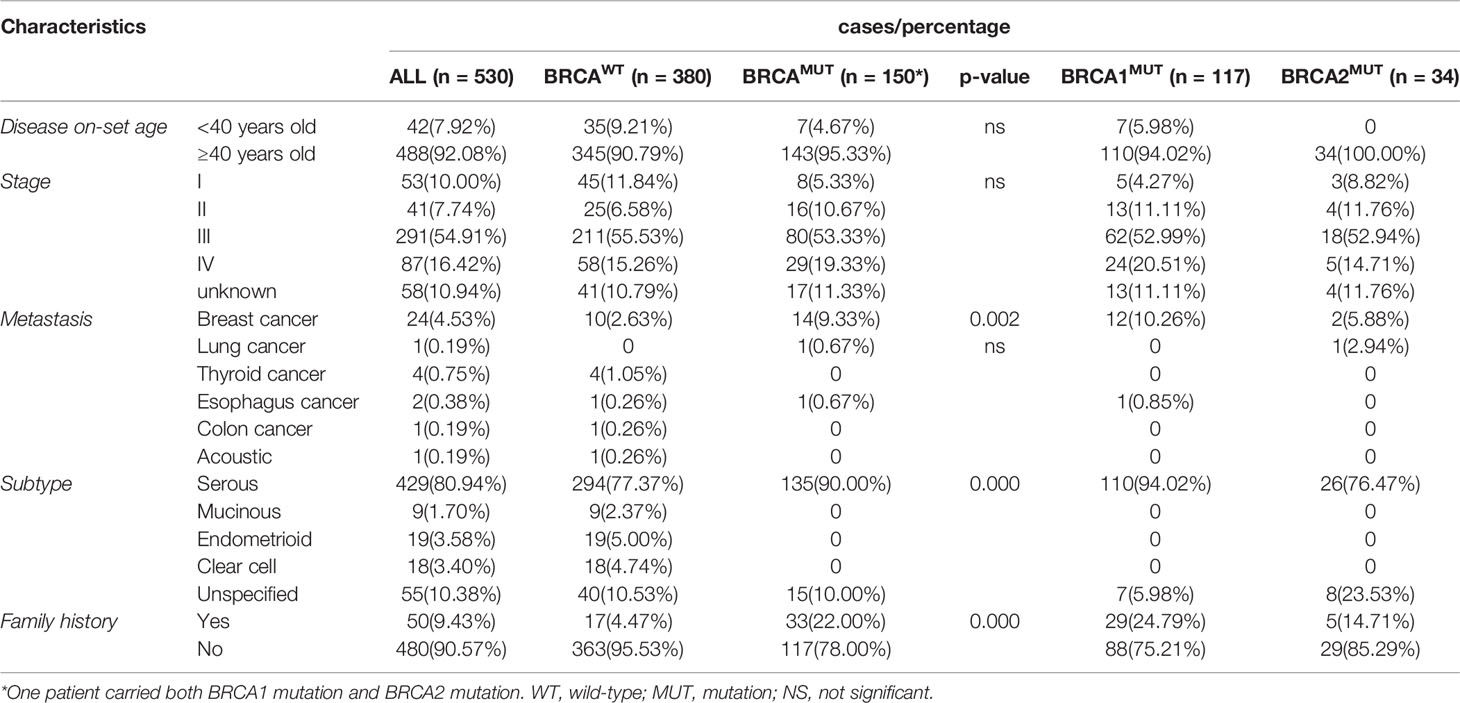
In total, 530 patients were included in this study. Individuals of <40 years old at diagnosis accounted for 7.92% (42/530) versus 92.08% (488/530) of the patients ≥40 years old. Only 17.74% (94/530) of the patients were diagnosed at earlier stages (53 in stage-1 and 41 in stage-2), and majority presented with advanced disease (54.91% in stage-3 and 16.42% in stage-4). Breast cancer was observed in 4.53% (24/530) of the patients, in which 14 were gBRCA1/2MUT and 10 were gBRCA1/2WT carriers; other concurrent cancers were rare in this cohort. Serous ovarian cancer was the dominant pathological subtype, accounting for 80.94% (429/530) of the patients, followed by clear cell, endometrioid, and mucinous carcinoma. There were 55 patients referred from other hospitals, and their diagnosis was only recorded as ovarian cancer without clear subtyping. About 9.43% (50/530) of the patients in our study were recorded with family history (Table 1).
In total, pathogenic or likely pathogenic gBRCA1/2MUT were identified in 28.3% (150/530) of the OC patients in our cohort; 117 patients carried BRCA1MUT and 33 carried BRCA2MUT and one patient carried both (Figure 1A). Although the disease incidence in both BRCAWT and BRCAMUT patients peak at 50–54 years old, BRCAMUT carriers had a significantly higher likelihood of developing OC between the age of 40 and 44 as compared to the BRCAWT carriers (p = 0.019, Figure 1B). Generally, the disease on-set age was slightly earlier in BRCAMUT carriers as compared to BRCAWT carriers (median: 52 vs 54 years old, p = 0.0252, Figure 1C), and in BRCA1MUT carriers versus BRCA2MUT carriers (median: 51 vs 55 years old, p = 0.0055, Figure 1D). Among the 42 patients presenting the disease before 40, seven patients harbored BRCA1MUT and the other 35 were BRCAWT, suggesting other unclarified risk factors contributing to their early disease on-set. Not surprisingly, BRCAMUT carriers more often developed breast cancers than other cancers (p = 0.002); BRCAMUT predominantly present in serous ovarian cancers rather than other subtypes (p = 0.000), and patients with a family showed a higher likelihood to carry BRCAMUT than the ones without (p = 0.000; Table 1).
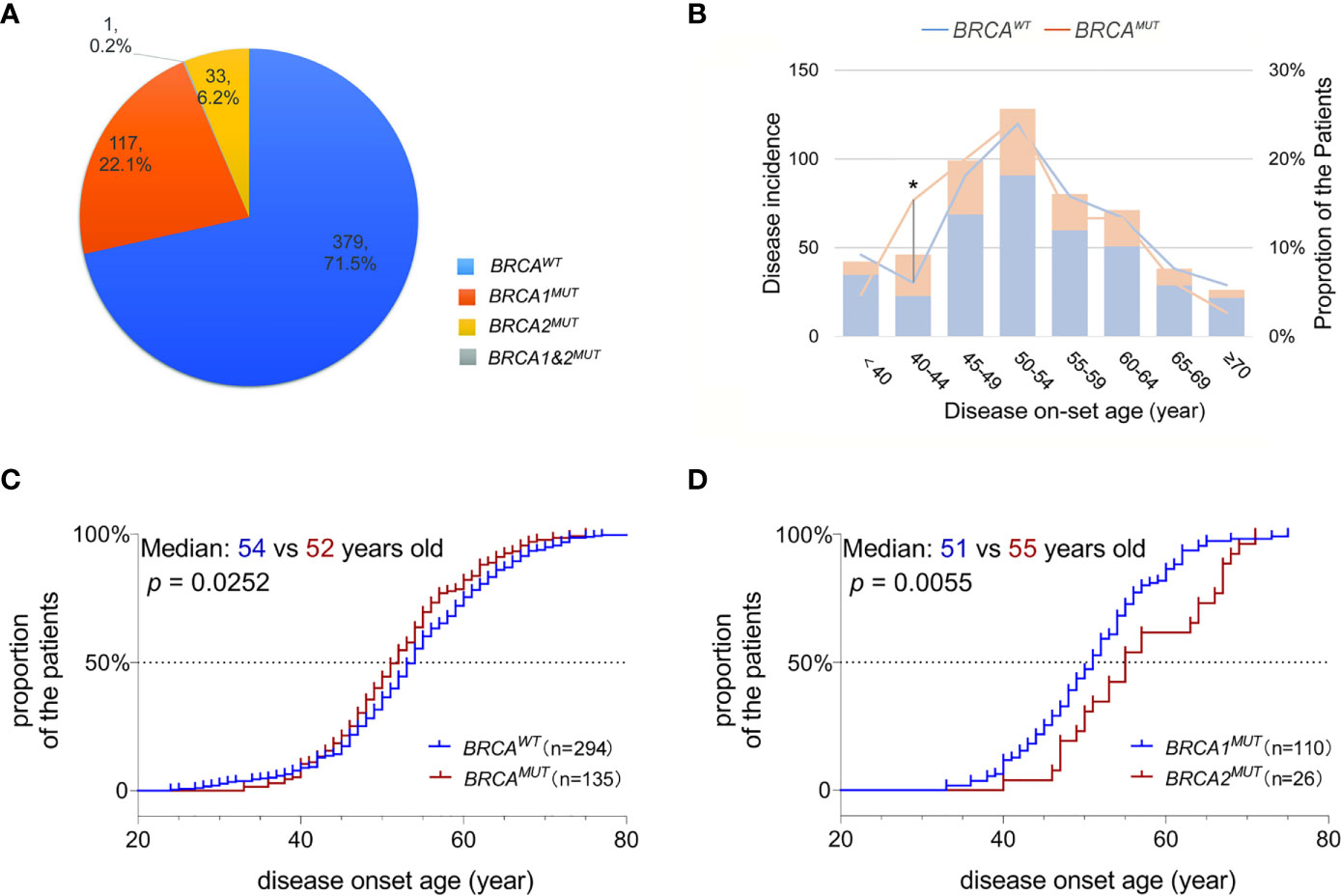
Figure 1 BRCA mutations associated with disease on-set age of OC. (A) A pie plot shows the number and proportion of BRCA1 and BRCA2 germline mutation carriers in 530 Henan OC patients. WT, wild-type; MUT, mutation. (B) Incidence (left Y-axis) and proportion of the patients developed serous OC (right Y-axis) at defined age (X-axis) in BRCAMUT (orange) and BRCAWT (blue) carriers. A chi-square test was used to determine whether the frequency of OC incidence is different between BRCAMUT and BRCAWT carriers of 40–44 years old. *p < 0.05. (C) Comparison of the disease on-set age of serous OC between BRCAMUT versus BRCAWT carriers, and BRCA1MUT versus BRCA2MUT carriers (D) by plotting the cumulative incidence curve. A log-ranked test was used to compare the difference in disease on-set age between different groups, and a p-value of < 0.05 was considered as significant.
The Founder Effect of BRCA1:c.5470_5477del in Henan OC Patients
Of the 151 mutations identified in our study, 117 were in BRCA1 and 34 in BRCA2; frameshift is the dominant subtype (66.9%,101/151), followed by missense (33.1%, 50/151); six mutations were located on the canonical splice sites, and an intronic conversion of A>G at BRCA1:c.213_12, creating a novel 3′ acceptor splicing sites, was also identified (Figures 2A, B). About 21.9% (33/151) of the detected mutations have not been reported in either Clinvar or BRCA exchange database (accessed in Jan. 2020), suggesting a different BRCA mutation spectrum in Chinese as compared to other populations. BRCA1:c.5470_5477del was the most recurrently detected mutation in our cohort, presenting in eight independent individuals; followed by BRCA1:c.981_982del and BRCA1:c.4065_4068del. In total, the top three mutants accounted for 13.9% (21/151) of all the BRCA mutations detected in this study.
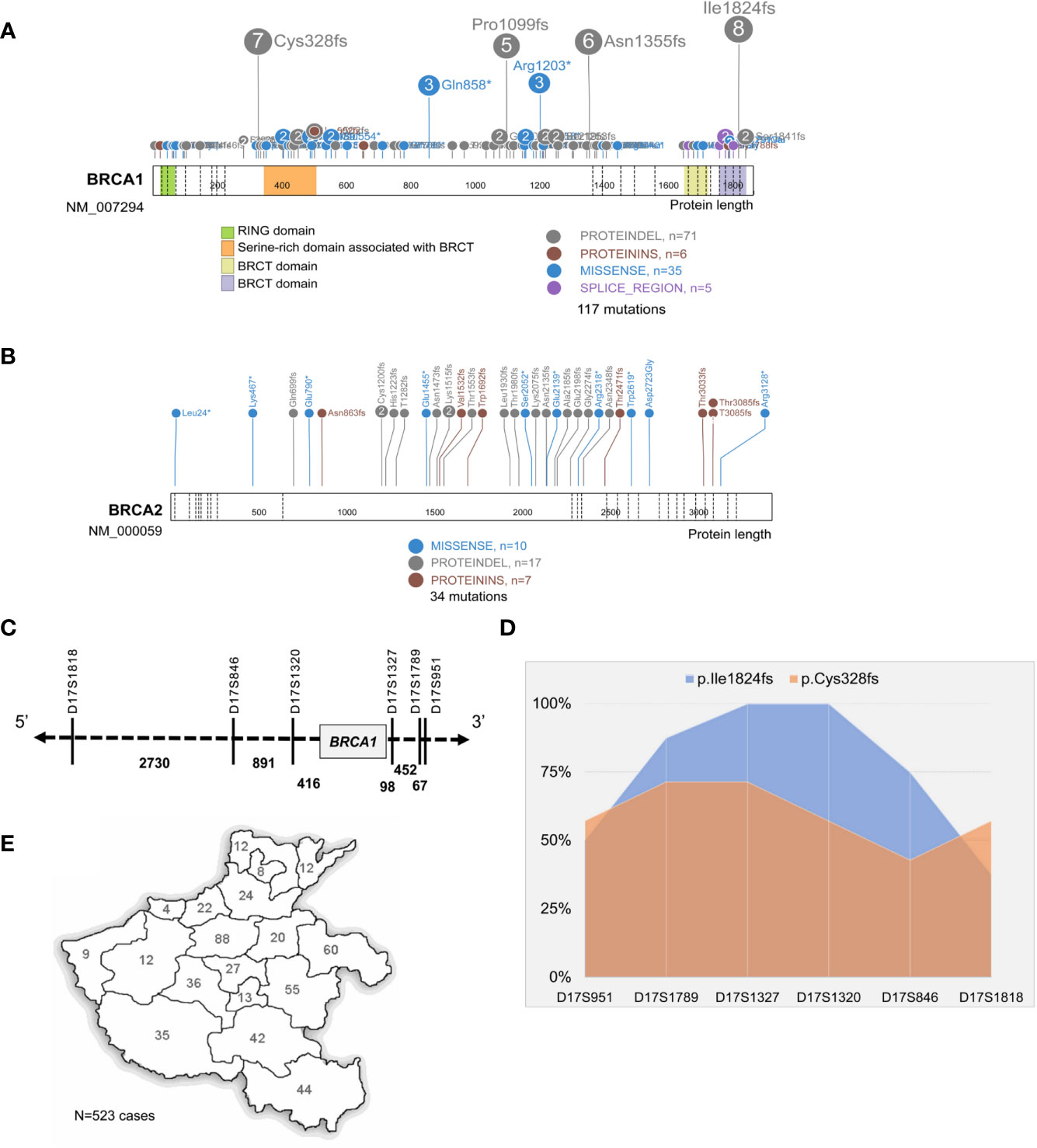
Figure 2 Identification of BRCA1:c.5470_5477del as the founder mutation in Henan OC patients. Protein paint shows the pathogenic and likely pathogenic mutations detected in BRCA1 (A) and BRCA2 (B) from 530 Henan OC patients. (C) A schematic diagram illustrates the genomic location of BRCA1 and its flanking STR markers selected for haplotype analysis. (D) Haplotype analysis of six STR markers across 8 BRCA1:c.5470_5477del (p.lle1824fs) carriers (blue) and seven BRCA1:c.981_982del (p.Cys328fs) carriers (orange). STR loci are indicated on x-axis and the proportion of patients sharing the same allele is indicated on Y-axis. (E) A map shows the places of origin of the patients in this OC cohort.
Then we performed haplotype analysis in the patients harboring BRCA1: c.5470_5477del by investigating six highly polymorphic STR markers flanking BRCA1 on chromosome17 (Figure 2C); seven other patients, sharing the BRCA1:c.981_982del mutation, were included for comparison. The results showed there was a highly conservative region, with a genomic length of 0.6 MB covering BRCA1, shared by all eight unrelated BRCA1:c.5470_5477del carriers, and the conservation gradually descend toward both sides. Meanwhile, this region was only partially shared by the BRCA1:c.981_982del carriers (Figure 2D and S Figure 1). Collectively, our data support BRCA1:c.981_982del as the suspected founder mutation in Henan OC patients since 98.7% (523/530) of the patients in this cohort are from different regions of Henan province (Figure 2E).
Ki-67 Expression, but Not gBRCAMUT Status, Associated With the PFS of Serous OC Patients
To prevent the confounding influence from patients with other subtypes of ovarian cancer, we focused our survival analysis on serous cancer only. In total, 165 serous OC patients at stage-3 or -4 disease with continuous follow-up data were investigated for their progression-free survival (PFS), and 103 patients were subjected to surgery resection followed by chemotherapy and the other 62 received adjuvant chemotherapy in advance to surgery. The analysis revealed neither the order of chemotherapy and surgery nor the gBRCAMUT status influenced the PFS of patients (Figures 3A, B); instead, the patients with a high Ki-67 expression of ≥50% nuclear staining in the FFPE sections showed a significantly shorter PFS (Figure 3C, p = 0.041), suggesting as a valuable prognostic predictor (Hazard ratio = 1.557 with 95% CI of 1.018–2.379). We didn’t observe any evident difference in the PFS between the patients with BRCA1:c.5470_5477del founder mutation and the ones with other gBRCAMUT (data not shown).
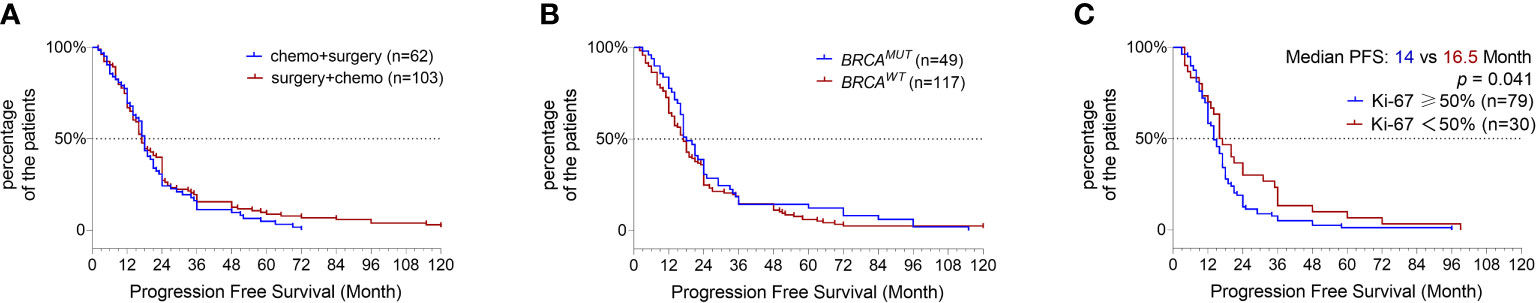
Figure 3 Progression-free survival (PFS) analysis. Kaplan–Meier plot shows the comparison of PFS in serous OC patients primarily treated by surgery versus adjuvant chemotherapy (A), BRCAMUT carriers versus BRCAWT carriers (B), and the patients with high ki-67 expression of ≥50% versus <50% (C).
Discussion
In this study, we identified 28.3% (150/530) of the OC patients in this Henan cohort as gBRCAMUT carriers, including one with concurrent mutations in both BRCA1 and BRCA2. Haplotype analysis revealed a region of 0.6 MB genomic length spanning BRCA1 highly conserved across all the independent carriers of BRCA1:c.5470_5477del, supporting it as a founder mutation in Henan population. Survival analysis showed the gBRCAMUT status of the serous OC patients was not associated with their PFS; instead, a nuclear expression of Ki-67% over 50% of the malignant cells appeared to be an independent predictor for a shorter PFS.
Mounting studies have shown that the prevalence of gBRCAMUT in ovarian cancer patients varies across different populations, ranging from 13.8% in Americans to 40% in Ashkenazi Jews (16–18). A nationwide multi-center study carried out by Wu et al. revealed the prevalence of gBRCAMUT in Chinese OC patients was 28.5% (7), which is comparable to our result. Interestingly, the top recurrently detected mutations varies between different studies, including the BRCA1:c.1081del, c.964del, c.3770_3771del, c.2371_2372del and c.5470_5477del etc. (12, 19–21); this is probably caused by the different ethnic origins and geographic locations of the included patients. Several studies have reported the BRCA1:c.5470_5477del as the most frequently detected mutation in Chinese OC patients; of note, the patients enrolled in these studies were mainly from northern China (13, 20, 22), and studies on the southern Chinese revealed different top mutations (12, 19, 21). In this study, almost all the patients are from Henan, the central region of China and the origin of Chinese civilization.
Predisposition of gBRCAMUT is known to promote the early-onset of breast and ovarian cancer (23). In our cohort, this phenomenon is more evident in the gBRCA1MUT carriers but not the gBRCA2MUT carriers of serous OC patients (median age: 51 vs 55; p = 0.0055) and to a much less extent in the BRCAMUT carriers in general as compared to the BRCAWT patients (median age: 52 vs 54; p = 0.0252). To our surprise, about 6.6% (35/530) of BRCAWT patients developed the disease before 40, in which only one presents with a clear family history, suggesting other unclarified risk factors contributing to their early disease on-set. In addition, the BRCAMUT carriers appeared to have the first peak of incidence from 40 to 44 years old, suggesting the BRCAMUT carriers should carry out preventative screening from 40.
STR analysis supports BRCA1:c.5470_5477del as the founder mutation in our OC cohort, but a validation of this finding in a larger cohort, maybe a nation-wide multi-centered study, could make the conclusion more solid. The conserved region from chr17:42627978 to 43223568 was only partially shared by BRCA1:c.981_982del carriers, which is probably caused by genomic rearrangement or other aberrations that occurred during the course of heredity. Recently, BRCA1:c.5470_5477del was also revealed as a founder mutation in a cohort of 9505 Han breast cancer patients, which made our conclusion more solid (24). Of note, studies carried out in the southern part of China identified different founder mutations, i.e., BRCA1:1081del (12). A non-negligible limitation in all these studies, including this one, could be that only the coding regions of BRCA1/2 and the flanking splice sites were sequenced, which missed the pathogenic aberrations in introns. For example, BRCA1:c.442-22_442-13del has been identified as an ancient founder mutation in the patients from the southern part of China (25).
Currently the first-line therapy against advanced ovarian cancer is maximal cytoreductive surgical debulking followed by chemotherapy with carboplatin–paclitaxel regimen (26). In this study, no difference in PFS was observed if the patients received adjuvant chemotherapy before surgery. Two independent studies showed neoadjuvant chemotherapy followed by surgery was not inferior to surgery followed by chemotherapy and with less postoperative adverse events (27, 28); however, a national cancer database study showed a superior overall survival in patients with primary surgery versus neoadjuvant chemotherapy (29). Surgery first or chemo first? There’s still not a conclusive answer at this point.
The prognosis of BRCAMUT carriers was shown better than BRCAWT carriers (30, 31), but we didn’t observe significant PFS benefit in gBRCAMUT carriers. This result is consistent with a recent study performed by You et al. that the benefit in PFS was only observed when the somatic BRCAMUT was also included into the analysis (22). Ki-67, a well-established biomarker of cellular proliferation, is frequently used in routine clinical workflow (32). Previous study showed Ki-67 expression was related to the overall survival of OC patients (33); here, our results confirmed that an expression of Ki-67 ≥50% indicated a shorter PFS. The efficiency of Ki-67 expression in predicting the prognostics of OC patients still needs improvement, and this could be addressed as the implication of high throughput sequencing based clinical tests are becoming increasingly prevalent.
In conclusion, our study reveals 28.3% of Henan OC patients are gBRCAMUT carriers, and BRCA1:c.5470_5477del is a founder mutation in Henan population. A nation-wide large cohort study may help us to understand the founder effect of BRCA mutations in Chinese and design a cost-effective screening test for the high-risk population.
Data Availability Statement
The data presented in the study are deposited in the Genome Sequence Archive repository (https://bigd.big.ac.cn/gsa/), accession number PRJCA004762.
Ethics Statement
The studies involving human participants were reviewed and approved by the ethics committee of Henan Cancer Hospital. The patients provided their written informed consent to participate in this study.
Author Contributions
JL and YG conceived and designed the study. SH and JL analyzed the data and drafted the manuscript with the support from YG and HL. XW and CZ performed NGS test and bioinformatic analysis. YL and LW evaluated the clinical outcome of the patients. JM, BW, YW, and QX analyzed the IHC staining and were responsible for the pathological diagnosis. YG and HL supervised the project. All authors read and approved the submitted manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was financially supported by the funding from Major public welfare projects in Henan Province (grant number: 201300310400) and Henan science and technology project (grant number: 212102310675). JL was also supported by the Henan provincial young researcher program. National Natural Science Foundation of China, 81802779 to JL; Henan Provincial Health Commission, SBGJ202002020 to JL; and Henan Science and Technology Project, 212102310675 to JL.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would also like to thank the other colleagues in the department of molecular pathology for helpful suggestions.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.655709/full#supplementary-material
References
1. Momenimovahed Z, Tiznobaik A, Taheri S, Salehiniya H. Ovarian Cancer in the World: Epidemiology and Risk Factors. Int J Womens Health (2019) 11:287–99. doi: 10.2147/IJWH.S197604
2. Gaona-Luviano P, Medina-Gaona LA, Magana-Perez K. Epidemiology of Ovarian Cancer. Chin Clin Oncol (2020) 9(4):47. doi: 10.21037/cco-20-34
3. Patni R. Screening for Ovarian Cancer: An Update. J Midlife Health (2019) 10(1):3–5. doi: 10.4103/jmh.JMH_46_19
4. Stewart C, Ralyea C, Lockwood S. Ovarian Cancer: An Integrated Review. Semin Oncol Nurs (2019) 35(2):151–6. doi: 10.1016/j.soncn.2019.02.001
5. Chen S, Parmigiani G. Meta-Analysis of BRCA1 and BRCA2 Penetrance. J Clin Oncol (2007) 25(11):1329–33. doi: 10.1200/JCO.2006.09.1066
6. Kuchenbaecker KB, Hopper JL, Barnes DR, Phillips KA, Mooij TM, Roos-Blom MJ, et al. Risks of Breast, Ovarian, and Contralateral Breast Cancer for BRCA1 and BRCA2 Mutation Carriers. JAMA (2017) 317(23):2402–16. doi: 10.1001/jama.2017.7112
7. Wu X, Wu L, Kong B, Liu J, Yin R, Wen H, et al. The First Nationwide Multicenter Prevalence Study of Germline BRCA1 and BRCA2 Mutations in Chinese Ovarian Cancer Patients. Int J Gynecol Cancer (2017) 27(8):1650–7. doi: 10.1097/IGC.0000000000001065
8. Heramb C, Wangensteen T, Grindedal EM, Ariansen SL, Lothe S, Heimdal KR, et al. BRCA1 and BRCA2 Mutation Spectrum - an Update on Mutation Distribution in a Large Cancer Genetics Clinic in Norway. Hered Cancer Clin Pract (2018) 16:3. doi: 10.1186/s13053-017-0085-6
9. Struewing JP, Abeliovich D, Peretz T, Avishai N, Kaback MM, Collins FS, et al. The Carrier Frequency of the BRCA1 185delAG Mutation is Approximately 1 Percent in Ashkenazi Jewish Individuals. Nat Genet (1995) 11(2):198–200. doi: 10.1038/ng1095-198
10. Zeegers MP, van Poppel F, Vlietinck R, Spruijt L, Ostrer H. Founder Mutations Among the Dutch. Eur J Hum Genet (2004) 12(7):591–600. doi: 10.1038/sj.ejhg.5201151
11. Reeves MD, Yawitch TM, van der Merwe NC, van den Berg HJ, Dreyer G, van Rensburg EJ. BRCA1 Mutations in South African Breast and/or Ovarian Cancer Families: Evidence of a Novel Founder Mutation in Afrikaner Families. Int J Cancer (2004) 110(5):677–82. doi: 10.1002/ijc.20186
12. Khoo US, Chan KY, Cheung AN, Xue WC, Shen DH, Fung KY, et al. Recurrent BRCA1 and BRCA2 Germline Mutations in Ovarian Cancer: A Founder Mutation of BRCA1 Identified in the Chinese Population. Hum Mutat (2002) 19(3):307–8. doi: 10.1002/humu.9015
13. Li A, Xie R, Zhi Q, Deng Y, Wu Y, Li W, et al. BRCA Germline Mutations in an Unselected Nationwide Cohort of Chinese Patients With Ovarian Cancer and Healthy Controls. Gynecol Oncol (2018) 151(1):145–52. doi: 10.1016/j.ygyno.2018.07.024
14. Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and Guidelines for the Interpretation of Sequence Variants: A Joint Consensus Recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med (2015) 17(5):405–24. doi: 10.1038/gim.2015.30
15. den Dunnen JT, Dalgleish R, Maglott DR, Hart RK, Greenblatt MS, McGowan-Jordan J, et al. Hgvs Recommendations for the Description of Sequence Variants: 2016 Update. Hum Mutat (2016) 37(6):564–9. doi: 10.1002/humu.22981
16. Robles-Diaz L, Goldfrank DJ, Kauff ND, Robson M, Offit K. Hereditary Ovarian Cancer in Ashkenazi Jews. Fam Cancer (2004) 3(3-4):259–64. doi: 10.1007/s10689-004-9552-0
17. Pal T, Permuth-Wey J, Betts JA, Krischer JP, Fiorica J, Arango H, et al. BRCA1 and BRCA2 Mutations Account for a Large Proportion of Ovarian Carcinoma Cases. Cancer (2005) 104(12):2807–16. doi: 10.1002/cncr.21536
18. Kwon BS, Byun JM, Lee HJ, Jeong DH, Lee TH, Shin KH, et al. Clinical and Genetic Characteristics of BRCA1/2 Mutation in Korean Ovarian Cancer Patients: A Multicenter Study and Literature Review. Cancer Res Treat (2019) 51(3):941–50. doi: 10.4143/crt.2018.312
19. Kwong A, Shin VY, Au CH, Law FB, Ho DN, Ip BK, et al. Detection of Germline Mutation in Hereditary Breast and/or Ovarian Cancers by Next-Generation Sequencing on a Four-Gene Panel. J Mol Diagn (2016) 18:580–94.
20. Bu H, Chen J, Li Q, Hou J, Wei Y, Yang X, et al. BRCA Mutation Frequency and Clinical Features of Ovarian Cancer Patients: A Report From a Chinese Study Group. J Obstet Gynaecol Res (2019) 45(11):2267–74. doi: 10.1111/jog.14090
21. Deng H, Chen M, Guo X, Heng J, Xu X, Peng L, et al. Comprehensive Analysis of Serum Tumor Markers and BRCA1/2 Germline Mutations in Chinese Ovarian Cancer Patients. Mol Genet Genomic Med (2019) 7(6):e672. doi: 10.1002/mgg3.672
22. You Y, Li L, Lu J, Wu H, Wang J, Gao J, et al. Germline and Somatic Brca1/2 Mutations in 172 Chinese Women With Epithelial Ovarian Cancer. Front Oncol (2020) 10:295. doi: 10.3389/fonc.2020.00295
23. Antoniou A, Pharoah PD, Narod S, Risch HA, Eyfjord JE, Hopper JL, et al. Average Risks of Breast and Ovarian Cancer Associated With BRCA1 or BRCA2 Mutations Detected in Case Series Unselected for Family History: A Combined Analysis of 22 Studies. Am J Hum Genet (2003) 72(5):1117–30. doi: 10.1086/375033
24. Meng H, Yao L, Yuan H, Xu Y, Ouyang T, Li J, et al. Brca1 c.5470_5477del, a Founder Mutation in Chinese Han Breast Cancer Patients. Int J Cancer (2020) 146(11):3044–52. doi: 10.1002/ijc.32877
25. Shaw T, Chan SH, Teo JX, Chong ST, Li ST, Courtney E, et al. Investigation Into the Origins of an Ancient BRCA1 Founder Mutation Identified Among Chinese Families in Singapore. Int J Cancer (2021) 148(3):637–45. doi: 10.1002/ijc.33241
26. Cortez AJ, Tudrej P, Kujawa KA, Lisowska KM. Advances in Ovarian Cancer Therapy. Cancer Chemother Pharmacol (2018) 81(1):17–38. doi: 10.1007/s00280-017-3501-8
27. Vergote I, Trope CG, Amant F, Kristensen GB, Ehlen T, Johnson N, et al. Neoadjuvant Chemotherapy or Primary Surgery in Stage IIIC or IV Ovarian Cancer. N Engl J Med (2010) 363(10):943–53. doi: 10.1056/NEJMoa0908806
28. Kehoe S, Hook J, Nankivell M, Jayson GC, Kitchener H, Lopes T, et al. Primary Chemotherapy Versus Primary Surgery for Newly Diagnosed Advanced Ovarian Cancer (CHORUS): An Open-Label, Randomised, Controlled, non-Inferiority Trial. Lancet (9990) 2015) 386:249–57. doi: 10.1016/S0140-6736(14)62223-6
29. May T, Comeau R, Sun P, Kotsopoulos J, Narod SA, Rosen B, et al. A Comparison of Survival Outcomes in Advanced Serous Ovarian Cancer Patients Treated With Primary Debulking Surgery Versus Neoadjuvant Chemotherapy. Int J Gynecol Cancer (2017) 27:668–74.
30. Safra T, Lai WC, Borgato L, Nicoletto MO, Berman T, Reich E, et al. Monte-Carlo (Principality of Monaco), March 17-20, 1988. Abstracts. Chronobiologia (1988) 15(3):264–71.
31. Zhong Q, Peng HL, Zhao X, Zhang L, Hwang WT. Effects of BRCA1- and BRCA2-related Mutations on Ovarian and Breast Cancer Survival: A Meta-Analysis. Clin Cancer Res (2015) 21(1):211–20. doi: 10.1158/1078-0432.CCR-14-1816
32. Ross W, Hall PA. Ki67: From Antibody to Molecule to Understanding? Clin Mol Pathol (1995) 48(3):M113–7. doi: 10.1136/mp.48.3.m113
Keywords: BRCA1/2, haplotype analysis, ovarian cancer, founder mutation, Chinese
Citation: Li J, Han S, Zhang C, Luo Y, Wang L, Wang P, Wang Y, Xia Q, Wang X, Wei B, Ma J, Li H and Guo Y (2021) Identification of BRCA1:c.5470_5477del as a Founder Mutation in Chinese Ovarian Cancer Patients. Front. Oncol. 11:655709. doi: 10.3389/fonc.2021.655709
Received: 12 February 2021; Accepted: 06 April 2021;
Published: 11 May 2021.
Edited by:
Jian-Jun Wei, Northwestern University, United StatesReviewed by:
Deyin Xing, Johns Hopkins University, United StatesPatrizia Pasanisi, Istituto Nazionale dei Tumori (IRCCS), Italy
Copyright © 2021 Li, Han, Zhang, Luo, Wang, Wang, Wang, Xia, Wang, Wei, Ma, Li and Guo. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yongjun Guo, Z3VveW9uZ2p1bkB6enUuZWR1LmNu; Hongle Li, bGxobDczQDE2My5jb20=
†These authors have contributed equally to this work and share first authorship
 Jun Li1,2†
Jun Li1,2† Sile Han
Sile Han Cuiyun Zhang
Cuiyun Zhang Yongjun Guo
Yongjun Guo