- 1College of Life Science and Bioengineering, School of Science, Beijing Jiaotong University, Beijing, China
- 2Research Unit of Cell Death Mechanism, Institute of Biotechnology, Chinese Academy of Medical Science, Beijing, China
- 3School of Medicine, Nankai University, Tianjin, China
- 4Department of Thoracic Surgery, The Second Affiliated Hospital of Air Force Military Medical University, Xi’an, China
- 5Department of Thoracic Surgery, The Fifth Medical Center of Chinese PLA General Hospital, Beijing, China
- 6Institute of Acupuncture and Moxibustion, China Academy of Chinese Medical Sciences, Beijing, China
- 7Department of Pathology, The Fifth Medical Center of Chinese PLA General Hospital, Beijing, China
- 8Department of Radiotherapy, The Second Affiliated Hospital of Air Force Military Medical University, Xi’an, China
- 9Department of Oncology, Beijing Shijitan Hospital of Capital Medical University, Beijing, China
Cell-in-cell (CIC) structures are defined as the special structures with one or more cells enclosed inside another one. Increasing data indicated that CIC structures were functional surrogates of complicated cell behaviors and prognosis predictor in heterogeneous cancers. However, the CIC structure profiling and its prognostic value have not been reported in human esophageal squamous cell Carcinoma (ESCC). We conducted the analysis of subtyped CIC-based profiling in ESCC using “epithelium-macrophage-leukocyte” (EML) multiplex staining and examined the prognostic value of CIC structure profiling through Kaplan-Meier plotting and Cox regression model. Totally, five CIC structure subtypes were identified in ESCC tissue and the majority of them was homotypic CIC (hoCIC) with tumor cells inside tumor cells (TiT). By univariate and multivariate analyses, TiT was shown to be an independent prognostic factor for resectable ESCC, and patients with higher density of TiT tended to have longer post-operational survival time. Furthermore, in subpopulation analysis stratified by TNM stage, high TiT density was associated with longer overall survival (OS) in patients of TNM stages III and IV as compared with patients with low TiT density (mean OS: 51 vs 15 months, P = 0.04) and T3 stage (mean OS: 57 vs 17 months, P=0.024). Together, we reported the first CIC structure profiling in ESCC and explored the prognostic value of subtyped CIC structures, which supported the notion that functional pathology with CIC structure profiling is an emerging prognostic factor for human cancers, such as ESCC.
Introduction
Esophageal cancer ranks the top 10 among most deadly cancers. It has two major histological types: adenocarcinoma (EAC) and squamous cell carcinoma (ESCC) (1). In China, over 90% of the cases of esophageal cancer are ESCC, which is the fourth most prevalent cancer of the country. ESCC is a highly aggressive malignancy with poor prognosis. However, even the esophageal cancers that belong to same stage are variable in terms of recurrence, mortality rates, and disease prognosis. Therefore, further exploration of mechanism underlying heterogeneity and identifying novel prognostic factor is needed (2–6).
Cell-in-cell (CIC) structures are defined as the special structures with one or more cells enclosed inside another one and prevalent in a wide range of human cancers (7). In cancer tissues, CIC structures could form homotypically between tumor cells or heterotypically between different types of cells, such as tumor cell and immune cells (8). The profiling of CIC structures indicated the direct interaction between tumor cells, and immune cell in micro-environment could promote the formation of CIC structures that lead to the death of the internalized cells. An increasing number of studies have suggested that more CIC structures were associated with poor prognosis in cancers and reports on CIC structure profiling as functional pathological surrogate of complicated cell behaviors and prognostic predictor in specific cancers are being reported (9–14).
In this study, we explored the CIC structure profiling in ESCC tissues by using the EML method established previously (15), and the association between CIC subtypes and prognosis of esophageal cancer was analyzed.
Materials and Methods
Patients and Samples
We performed a retrospective analysis of patients with histologically confirmed ESCC after esophagectomy. This study consisted of 71 samples (whole slides) of ESCC patients, who were treated between February 2014 and March 2019 at the Fifth Medical Center of Chinese PLA General Hospital (Beijing, China) and the Second Affiliated Hospital of Air Force Military Medical University (Xian, China), and 70 samples of ESCC patients from a tumor microarray (TMA) purchased from Shanghai Outdo Biotech Co. Ltd (Heso-Squ172Sur-01, XT11-020). The tissue microarray spotted with 172 tissue cores with a diameter of 1.5 mm, consisting of 94 esophageal cancer samples and 78 samples from adjacent esophageal cancer tissue. Finally, 70 esophageal cancer samples in the tissue microarray were involved in the final analysis based on pathological characteristic selection. Informed written consent was obtained from the patients. The study was approved by the institutional ethical committee of the Fifth Medical Center and Second Affiliated Hospital of Air Force Military Medical University. Patients that exhibited other types of malignancy or had succumbed to illness within 1 month post the surgery were excluded from the study. Follow-up data were available for all 141 patients, ranging between 1 and 85 months post the surgery (mean, 29.96 months).
Immunostaining and Image Processing of Tissue Samples
Totally, 141 tissue samples of ESCC were collected for immunostaining. In addition, 10 paired para-carcinoma tissue and 80 normal tissues were used as control, respectively. The thickness of each tissue section is 4 μm. “EML method” was used to subtype CIC structure as previously reported (15), with antibodies against E-cadherin for epithelium, CD68 for macrophage, and CD45 for leukocyte. In brief, samples were first stained with antibody against CD45 (mouse mAb from Boster, BM0091) at dilution of 1:400 by Opal Multiplex tissue staining kit (Perkin Elmer, NEL791001KT) according to the standard protocol provided, which was eventually labeled with cyanine 3 fluorophore. Slides were then incubated with mixed antibodies against CD68 (rabbit pAb from Proteintech, 25747–1-AP) and E-cadherin (mouse mAb from BD Biosciences, 610181), followed by secondary antibodies of Alexa Fluor 647 anti-rabbit antibody (Invitrogen, A21245) and Alexa Fluor 488 anti-mouse antibody (Invitrogen, A11029), respectively. Fluorescence exited at the 488- and 647-nm wavelength was in spectrum similar to FITC and Cy5. Samples were also co-stained with antibodies against Caveolin-1 (rabbit mAb Cell Signaling Technology, 2267) and Ezrin (BD Biosciences, 610602), labeled with Cy5 and Cy3, respectively, as well as LAMP3 (Santa Cruz, sc-5275), labeled with Cy5. All slides were counterstained with DAPI to show nuclei and mounted with antifade reagent (Invitrogen, Carlsbad, CA). Multispectral images were taken with TMA modules of Vectra® Automated Imaging System (Akoya) by 20× objective lens. Nuance system (Akoya) was used to build libraries of each spectrum (DAPI, FITC, Cy5, and Cy3) and unmix multispectral images with autofluorescence subtracted in high contrast and accuracy. InForm automated image analysis software package (Akoya) was used for batch analysis of multispectral images based on specified algorithms.
CIC Structure Profiling and Quantification
Cellular structure where one or more cells morphologically fully enclosed by another cell with crescent nucleus was scored as CIC structures. Cell boundary of epithelial cells could be told by E-cadherin, which labels cell membrane. Additionally, CD45 and CD68 were applied to label cell body of leukocytes and macrophages, respectively. According to the expression feature of these three markers, we defined five different CIC structure subtypes in ESCC tissues, including (A) tumor cell in tumor cell (TiT, both cells expressed E-cadherin but negatively for CD45 and CD68), (B) leucocyte in tumor (LiT, inner cell is only positive for CD45, while outer cell only express E-cadherin), (C) tumor in macrophage (TiM, inner cell is only positive for E-cadherin, while outer cell only express CD68), (D) leukocytes in macrophages (LiM, inner cell is only positive for CD45, while outer cell only express CD68), and (E) macrophage in tumor (MIT, inner cell is only positive for CD68, while outer cell only express E-cadherin).
Only those structures with inner cells morphologically fully enclosed were counted. CIC structure subtypes were defined based on the types of cell involved: TiT for E-cadherin+ cells in E-cadherin+ cells (Tumor cells in Tumor cells); TiM for E-cadherin+ cells in CD68+ cells (Tumor cells in Macrophages); MiT for CD68+ cells in E-cadherin+ cells (Macrophages in Tumor cells); LiT for CD45+ cells in E-cadherin+ cells (Leukocytes in Tumor cells); LiM for CD45+ cells in CD68+ cells (Leukocytes in Macrophages). Overall CIC structures (oCICs) indicated the total of all kinds of CIC structures. CIC structure density in tissue was calculated as CIC structure number per mm2. Double blind reviews were performed by three experienced investigators in the quantification of CIC structure subtypes.
Statistical Analysis
OS duration was defined as time from the date of surgery to death or to the most recent contact or visit. Associations between CIC structure counts and the clinicopathological characteristics of the patients were analyzed using the Spearman rank test. Proportional hazard assumption was checked by both graphically and hypothetically using a hypothesis test called Shoenfield residual test for the oCIC and TiT variables. Results showed that Shoenfield residuals were not associated with the time (p > 0.05), suggesting that this model satisfies the proportional hazards assumption (Figure S2). The continuous variables were dichotomized for OS using the “surv_cutpoint” function of the “survminer” R package, which determine the optimal cutpoints to separate high and low cell-in-cell groups at once based on maximally selected rank statistics (16, 17). The standardized log-rank statistics across all the cutoffs were shown to depict the selection of optimal cutpoints (Figure S3). Survival curves were plotted using the Kaplan-Meier method, and the differences in survival distributions between two groups were compared by the log-rank test. Cox univariate and multivariate regression analyses were conducted to determine the factors that were independently associated with patients’ OS. All statistical analyses were performed using SPSS 24.0 soft-ware (IBM, Armonk, NY, USA) and GraphPad Prism 5.0. For all these analyses, a P value less than 0.05 was considered statistically significant.
Results
Patient Characteristics
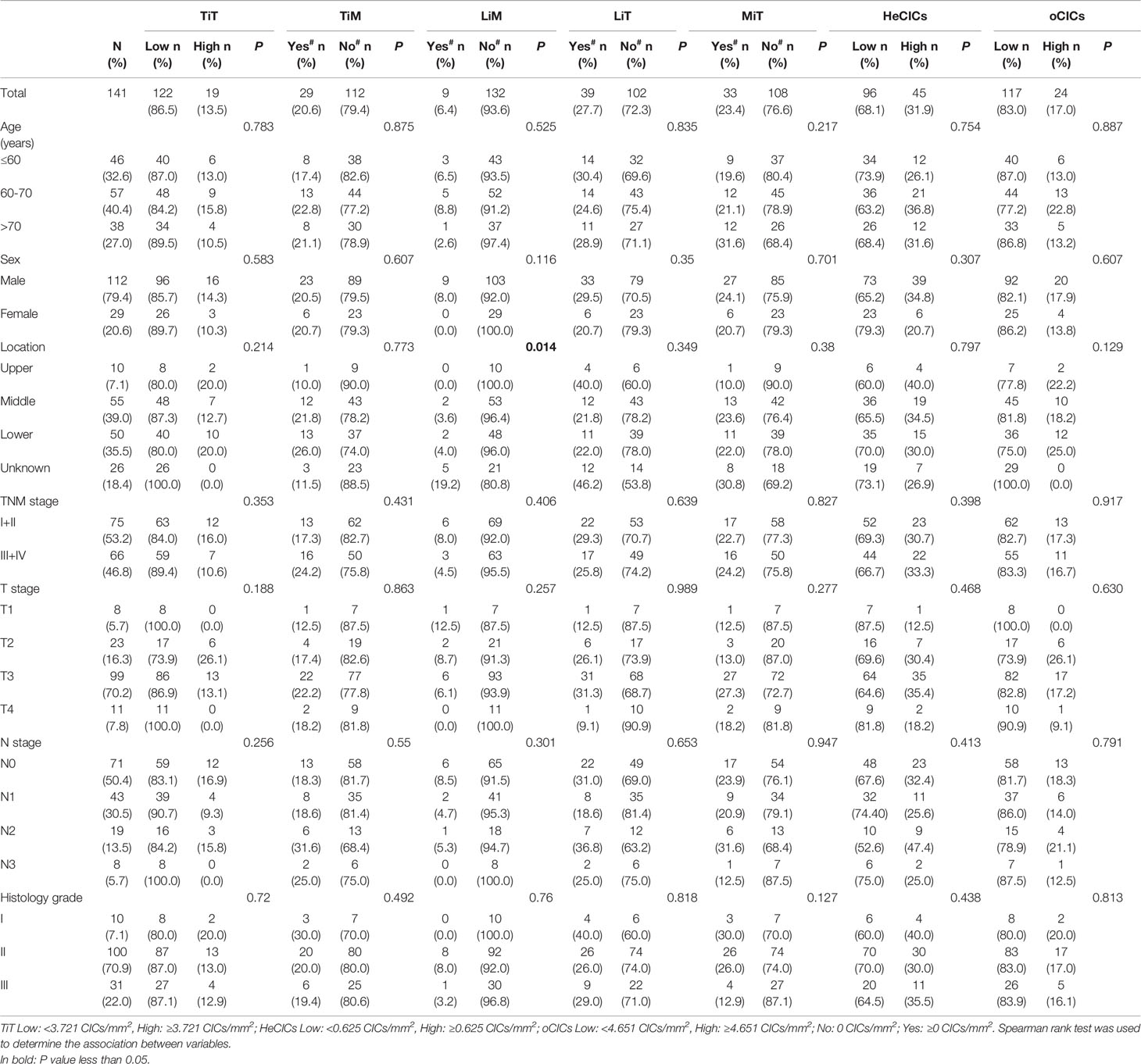
Totally, the data of 141 patients with ESCC were analyzed in this retrospective study. The patient characteristics were shown in Table 1 and Table S1. The median age was 64.0 years (range, 31–81 years), and most patients were male (79.4%). The most common locations of lesion were middle esophagus (39.0%) and lower esophagus (34.0%). None of patients had received preoperative chemotherapy and radiotherapy.
CIC Profiling in ESCC
Totally, 141 ESCC tissue samples, 10 paired para-carcinoma tissues and 80 paired normal tissues were stained by using the EML method. CIC structures were positive in 121 tumor tissue with the average density as 2.5 CIC/mm² (range, 0.5–24.0 CIC/mm²; Figure 1). However, there was no CIC detected in para-carcinoma or normal esophagus tissue.
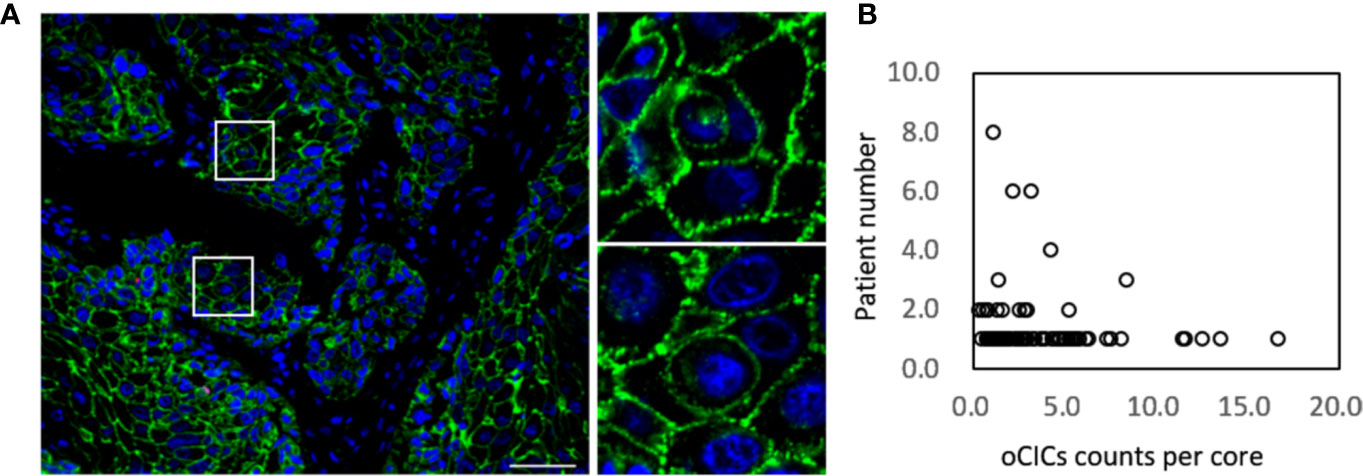
Figure 1 Cell-in-cell structures in esophagus carcinoma. (A) Representative image for CICs in human esophagus carcinoma co-stained with antibodies for E-cadherin, CD45, CD68 and DAPI. Right panels show zoomed images for boxed regions in the left image. Scale bar: 100 μm. (B) Distribution of overall CICs (oCICs) across esophagus carcinoma tissues from different patients.
We exploited three molecules, E-cadherin for epithelial cell membrane, CD45 for the leukocytes, and CD68 for the macrophages to identify CIC subtypes. Based on the “epithelium-macrophage-leukocyte” (EML) multiplex staining method, we defined five different CIC structure subtypes in ESCC tissues including (A) tumor cell in tumor cell (TiT), (B) leucocyte in tumor (LiT), (C) tumor in macrophage (TiM), (D) leukocytes in macrophages (LiM), and (E) macrophage in tumor (MIT) (Figures 2A–E). TiT was also named with homotypic CIC (HoCIC) structures, and the other four subtypes of CIC structures were summarized as heterotypic CIC (HeCIC) structures. Besides, molecules, like Caveolin-1, Ezrin, and LAMP3, were also checked in esophageal carcinoma tissues. The result showed that the positive rate of Ezrin in HoCIC structure was higher than that of caveolin-1, indicating a more active role of Ezrin in CIC structures of ESCC tissues. Interestingly, though LAMP3 expressed well in most of the ESCC tumor cells, only a small portion of CIC structures was positive in LAMP3, probably suggesting a low level of lysosomal activity in these CIC structures (Figure S1).
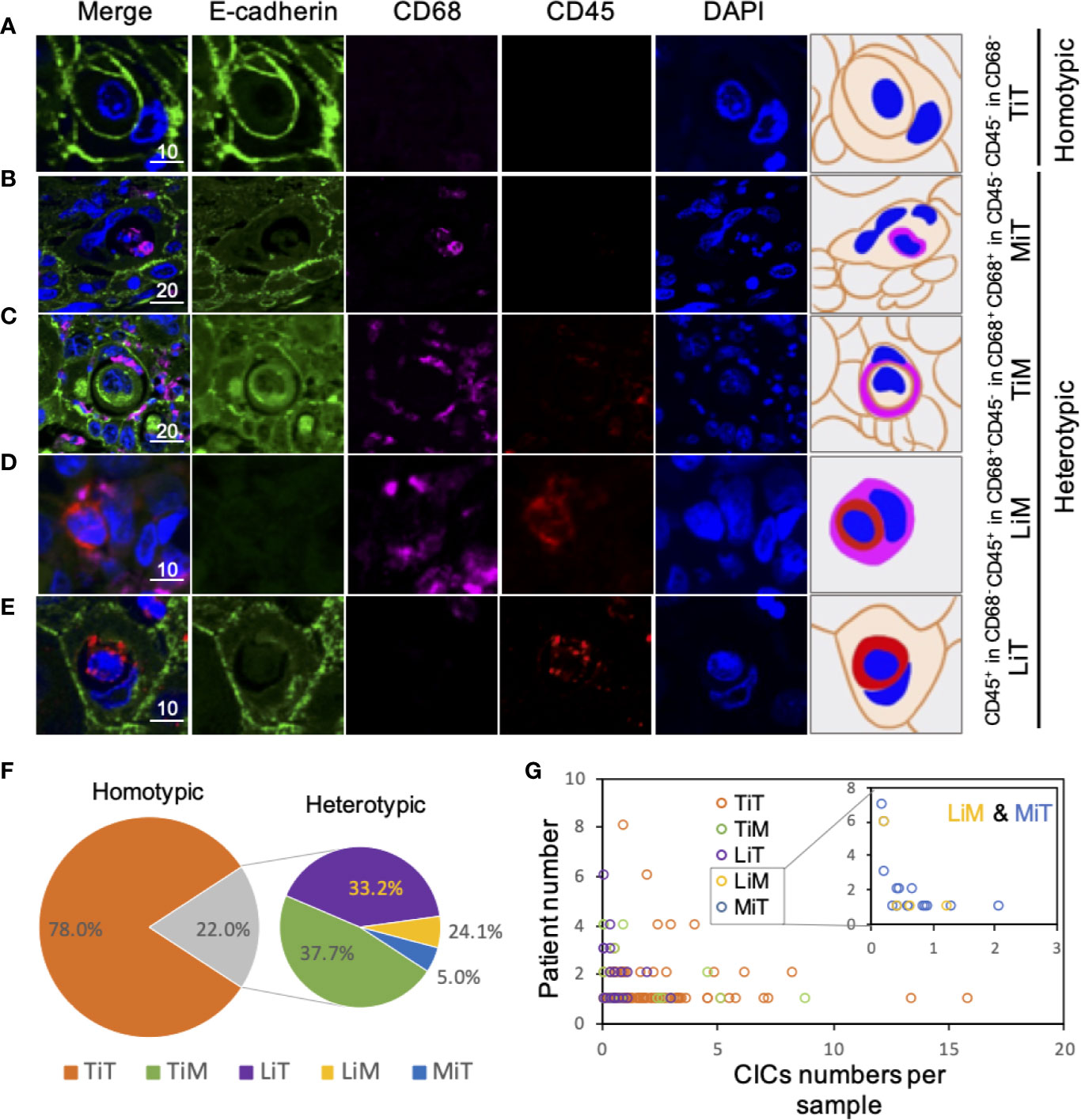
Figure 2 Subtype profiling of cell-in-cell structures in esophagus cancer. (A–E) Representative images for five CIC subtypes as indicated. Right panels of pictures demonstrate the schematic structure for each CIC subtype. Scale bar: 10 or 20 μm. (F) Distribution of five CIC subtypes across esophagus cancer tissues in all patients. (G) Distribution of five CIC subtypes in esophagus cancer tissues from different patients.
Among the subtypes of CIC structures, TiT was most prevalent subtype, which accounted for 78.0% of the total number of oCIC structures, and 97.5% (118) of oCIC-positive tumor tissue was TiT positive. As for HeCIC structures, TiM had the largest number, accounting for 37.7% of the total HeCIC structures, and 8.3% of oCIC structures; LiT and MiT ranked the second and third, accounting for 33.1% and 24.1% of the total HeCIC, 7.3% and 5.3% of oCIC structures, respectively; LiM was rare and accounted for 5.1% of the total HeCICs (Figure 2F).
Association Between CIC Structure Subtypes and Clinicopathological Characteristics
To quantify the CIC structures more accurately, CIC density, for CIC structure counts per mm2, was introduced in this study. The CIC density in the 141 samples was shown in Figure 2G. Considering that majority of the CIC structures was HoCIC (or TiT) and HeCIC quantity was rather low, only oCIC structures, HeCIC structures, and TiTs were evaluated by density in subsequent analysis. The patients were divided into two groups according to optional cutpoint of CIC structures density based on the maximally selected rank statistics. Take oCIC structures as an example, the patients with density < 4.651 CIC structures/mm² were categorized as low oCIC structure group, while those with density ≥ 4.651 CIC structures/mm² were categorized as high CIC structure group. As such, the patients were dichotomize based on the optional cutpoints of HeCIC structures (0.625 CIC structures/mm²) and TiT (3.721 CIC structures/mm²) (Figure S3). For TiM, LiM, LiT, and MiT, the dichotomy was applied based on the presence and absence of corresponding CIC structure owing to the scarce of these CIC structure subtypes.
The association between CIC structure subtypes and the clinical information of patients was further analyzed, as shown in Table 1. The result indicated that, compared with the samples of upper ESCC, there were more LiM detected in the samples of lower ESCC (0% vs 50%, P = 0.014). CIC structure subtypes demonstrated no association with other traditional clinicopathological factors, including TNM stage, age, sex, and histology grade.
Association Between CIC Structure Subtypes and Patients’ Survival
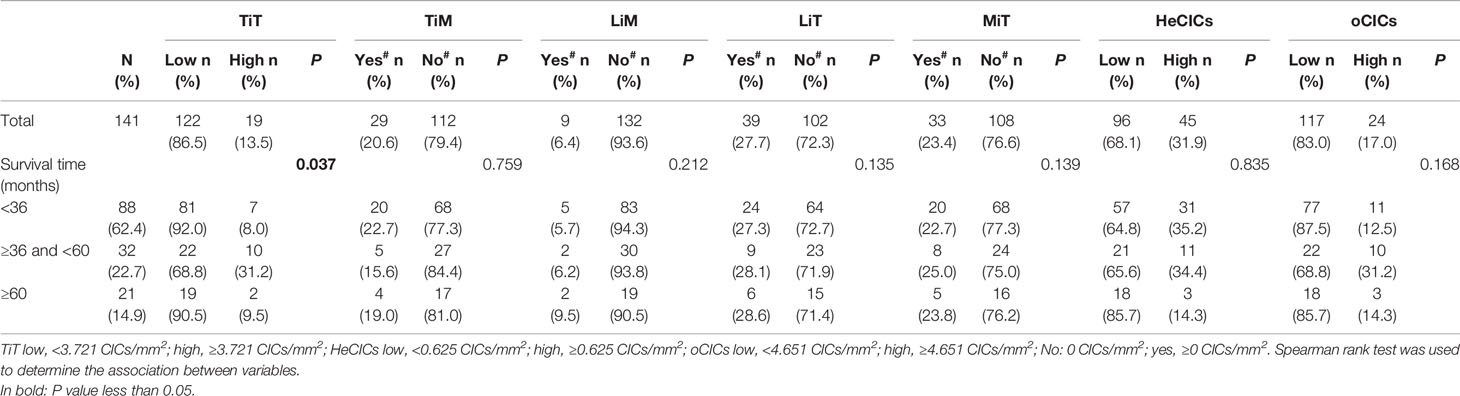
With the patients’ return visit, the survival information of 141 patients was collected, and the correlation between the levels of various CIC structure subtypes and survival time of the patients was analyzed. The results in Table 2 showed that in esophagus cancer, the distribution of TiT was significantly different in patients with distinct survival duration. More narrowly, the proportion of more TiT was much higher in patients with survival duration 36 to 60 months than that in patients with survival duration less than 36 months or longer than 60 months.
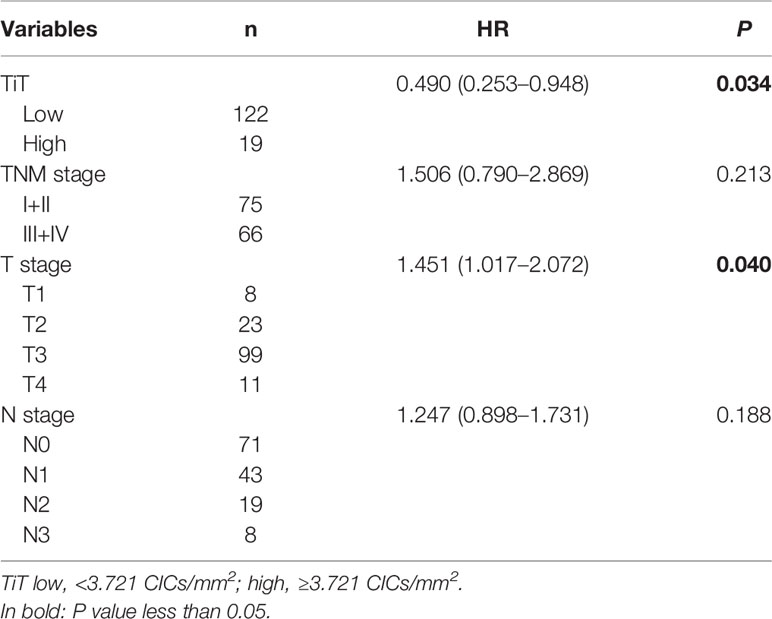
The relation between CIC structures and patients’ survival was then assessed by Cox regression model. In univariate analysis, TiT density, TNM stage, T stage, and N stage was significantly correlated with the OS of ESCC patients, respectively, as shown in Table 3 (HR: 0.48249882, P = 0.028905; HR: 2.45798584, P = 0.000008; HR: 1.76727404, P = 0.000513; HR: 1.57737502, P = 0.000022).
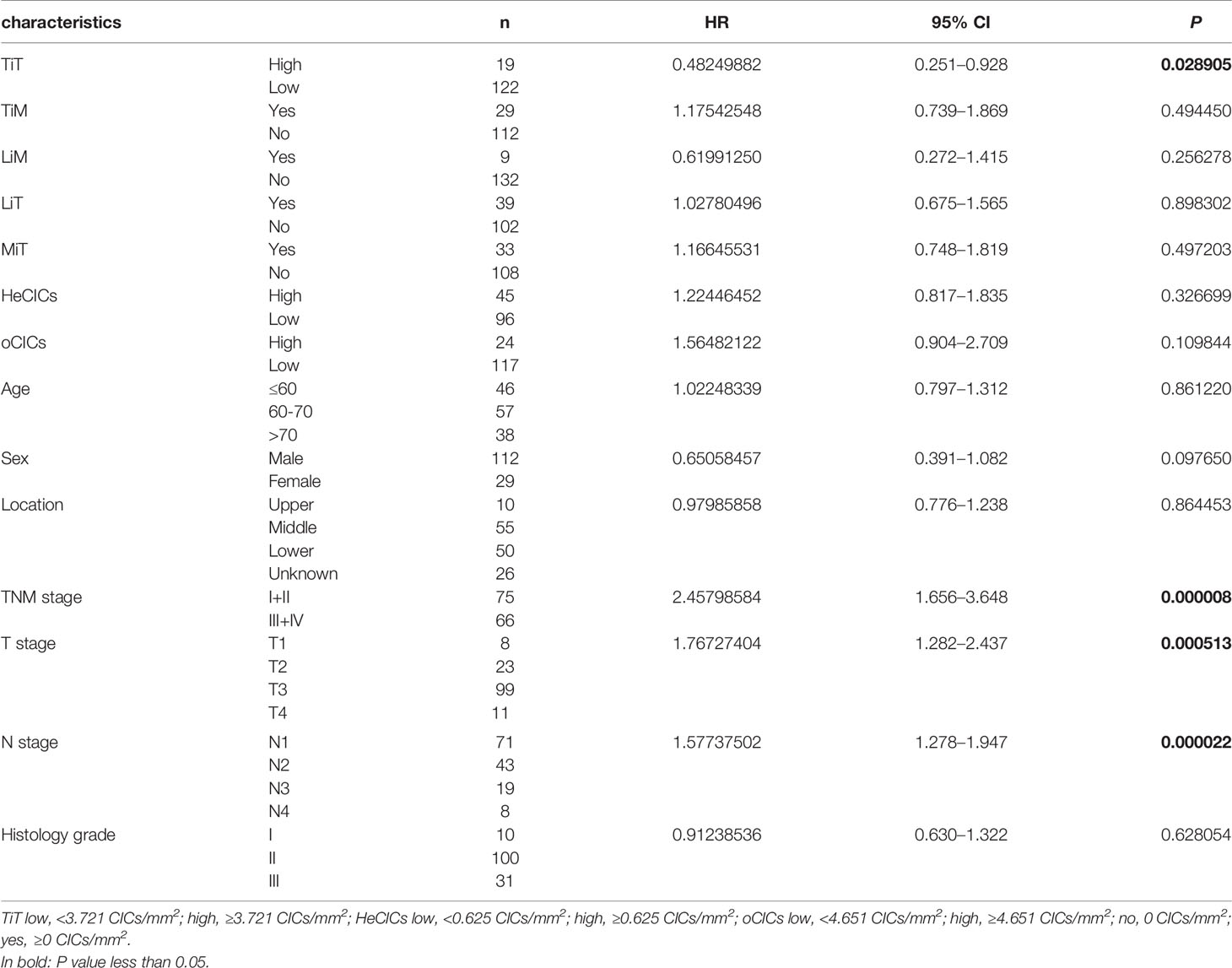
Table 3 Association of overall survival with clinicopathological parameters and CICs by univariate Cox regression analysis.
Survival curve was plotted by Kaplan-Meier method, and the difference was assessed by log-rank test. The results indicated that patients with high TiT density showed a favorable prognosis and longer median OS (mOS: 57 vs 23 months, P=0.025) (Figure 3B) compared with patients with low TiT density. However, oCIC structures failed to show any association with survival time (Figures 3A, B).
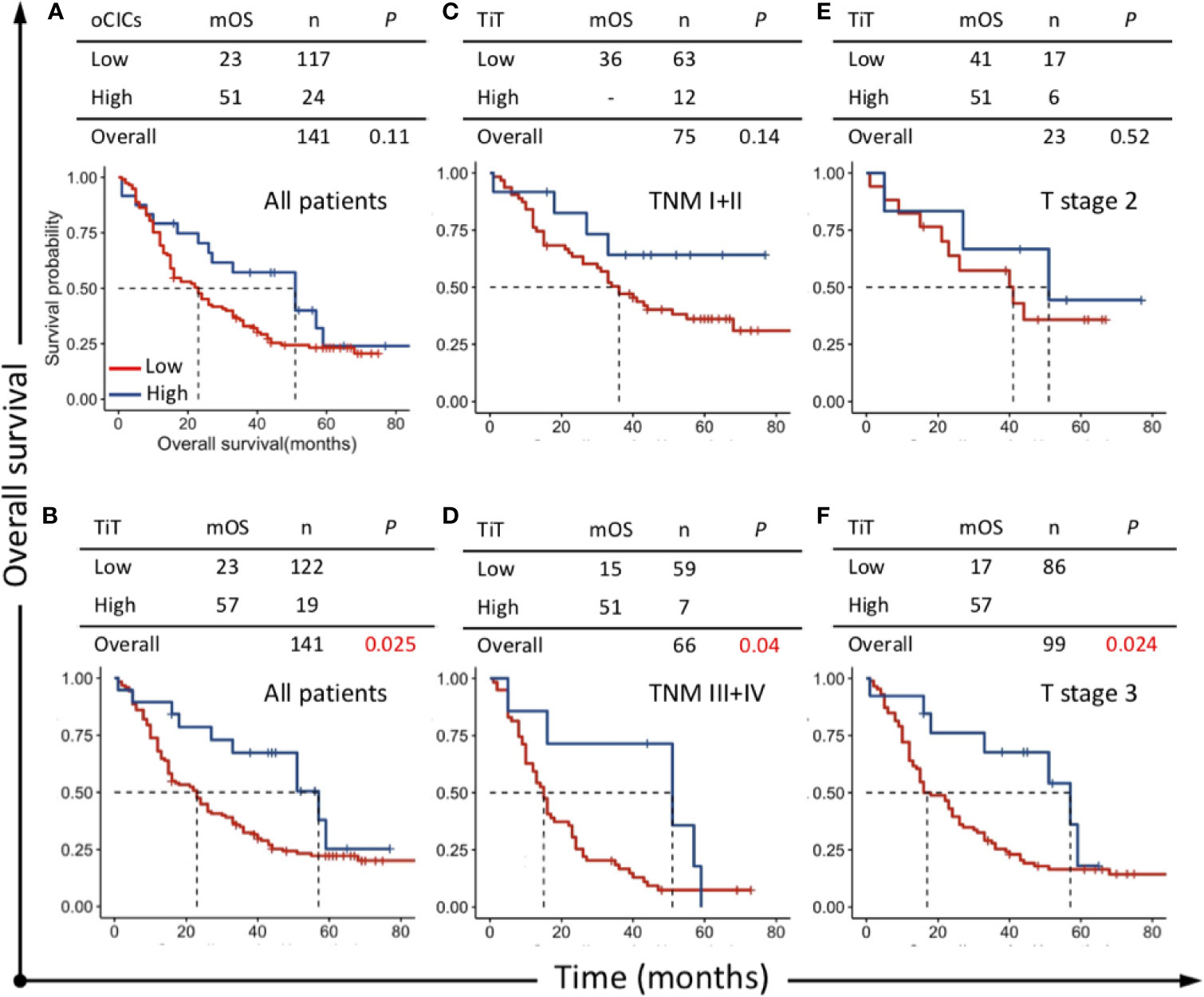
Figure 3 Survival impacts of CICs on overall survival (OS) of esophagus cancer patients. Kaplan–Meier plotting for OS curves of (A) oCICs, (B) TiT, and (C–F) TiT in stratified patients with different TNM and T stages. TiT Low: <3.721 CICs/mm2, High: ≥3.721 CICs/mm2; oCICs Low: <4.651 CICs/mm2, High: ≥4.651 CICs/mm2.
CIC Structures Are Independent Prognostic Factor for Esophagus Cancer
To analyze whether CIC structures and their subtypes are independent prognostic factors for postoperative survival of ESCC patients, all the variables identified in univariate analysis (TiT density, TNM stage, T stage and N stage) were then included in further multivariate analysis using Cox proportional hazards model. The result indicated that TiT density (HR = 0.490, P = 0.034) and T stage (HR = 1.451, P = 0.040) were independent prognostic factors for ESCC patients (Table 4). In detail, more TiT in tumor tissue and early T stage could decrease the death risk of ESCC patients by nearly 50%.
Homotypic CIC Structures Preferentially Impact the Survival of Patients of Late TNM Stage
To clarify the prognostic value of TiT in different subpopulation of ESCC patients, the correlation of TiT density and OS was analyzed when the patients were stratified by different TNM stage or T stage. The result indicated that high TiT density was associated with longer OS in patients of TNM stage III and IV as compared with patients with low TiT density (mOS: 51 vs 15 months, P = 0.04) and T3 stage (mOS: 57 vs 17 months, P=0.024). Meanwhile, in patients with early stage such as TNM stage I and II, there was no difference in prognosis between high and low TiT density groups, as well as T2 stage (Figures 3C–F). These results suggested that TiT density may potentially have better prognostic value in patients of later stage or higher risks; however, it should be validated in further study with amplified patient size and in a prospective way.
Discussion
In this study, we examined the CIC profiles of ESCC and paired adjacent normal tissues and identified five CIC subtypes in ESCC tissues. Among the five subtypes, TiT served as an independent prognostic factor. On top of that, TiT tended to demonstrate better prognostic performance in patients of rather later TNM stage or T stage (P = 0.04 or 0.024), which warrants further validation in future studies.
CIC structure is a special way of cell death with high environmental specificity that kills the internalized cells generally in an acidified lysosome-dependent way (8, 18). Cell death caused by CICs is mediated by three core elements including adherens junction, contractile actomyosin and mechanical ring as well (19–21). Besides, CIC structure formation is also regulated by a set of factors, such as CDKN2a (22), PCDH7 (23), IL8 (24) and membrane lipids (25). Functionally, CIC structure-mediated non-autonomous inner cell death is conducive to cell competition in mammals (7, 26–28). Therefore, CIC structures are also regarded as a way for tissues to maintain homeostasis (29, 30). Disorder in this process may lead to tumors or diseases such as immune-related disorders (8, 31–33). At present, literatures have reported CIC structures in tumor tissues, including urothelial carcinoma (34), buccal mucosa squamous cell carcinoma (10), pancreatic ductal adenocarcinoma (9), metastatic adenocarcinoma (34), head and neck squamous cell carcinoma (35), renal cell carcinoma (36), gastric cancer (37), breast carcinoma (11, 38), small cell carcinoma of lung (39), benign tendon sheath giant cell carcinoma specimen (40), malignant mesothelioma (41), leiomyolipoma (42), etc., but there is no study on the CIC structures in the tissues of esophageal cancer.
Current determination of CIC structures was mainly based on the cellular morphology that was readout by ways of tissue staining. Hematoxylin-Eosin (H&E) staining and May-Grunwald-Giemsa (MGG) staining were two usual methods to distinguish cell nucleus and membrane. Papanicolaou staining was used for polychromatic staining, of which the section transparency is better, and the cytoplasm in a variety of cells present different colors, which is convenient for determining cell types. Similarly, EML staining can also help to distinguish cell types, and more precisely, in which epithelial cells, macrophages and leukocytes were marked by epithelial cadherin, CD68, and CD45, respectively. The results showed that the 5 subtypes reported in other cancers were also detected in human ECSS tissues, of which TiT was much higher than the other 4 subtypes. In addition, the density of subtyped CIC structures are correlated with location and prognosis of ESCC, suggesting that CIC structures may be a candidate marker for clinical diagnosis of esophagus cancer. There may also be other types of cell cannibalism, which requires screening with more cellular markers.
As for the level of CIC structures in the previous studies, other quantitative methods were used. In urothelial carcinoma and metastatic adenocarcinoma, where the number of CIC structures in 100 tumor cells was set as CIC structure index (34); Except for this, in the experiment of breast ductal carcinoma, the number of CIC structures in 1000 tumor cells was also served as CIC structure index (43). To calculate CIC structure index, the total number of tumor cells, other than those in CIC structures, in a defined field shall be counted, which tended to produce more systemic errors. While in this paper, we used CIC structure density as the readout, which was calculated as CIC structure number in a defined area, so it is more suitable to clinical practices.
Previous experiments about melanoma cells showed that cell ingestion in cancer arises owing to the need for metabolism of individual cells, which increases intracellular nutrient pools in order to support cancer cell survival and proliferation (44). Similarly, the internalization of tumor cells may increase the instability of genome in target cells, thereby promoting the development of tumors in the long term. On the other hand, CIC formation as a mechanism of cell cannibalism that is induced by the establishment of epithelial adhesion could inhibit transformed growth, suggesting that CIC structure-mediated cell killing might also be a potential tumor suppressive pathway (7, 20). Cannibalistic behavior, as one of the mechanisms involving CIC formation, may increase nutrient intake by feeding upon other cells, and escape from the specific immune response by engulfing lymphocytes (33). In this scenario, upregulation of phagocytosis in human tumor cells may be similar to that of some unicellular microorganisms, of which the goal is to survive and propagate in a hostile microenvironment (8, 45). Moreover, while this study dealt with resectable cancer, increased CIC formation was also identified in metastatic cancer cells. For example, Lugini et al. found that the metastatic, but not the primary, melanoma cells displayed strong cannibalistic activity against live lymphocytes (44). This activity seems to be associated with the expression of TM9SF4, an important protein associated with phagocytic activity (45, 46), as well as v-ATPase, a master controller of vacuolar pH that is in complex with TM9SF4 to create a unique milieu favoring tumor metastasis and chemo/immune-resistance traits of solid tumors (47). Future investigations on the expressions and roles of TM9SF4 and v-ATPase in CIC structures of metastatic cancers would shed novel lights on the CIC structure formation by tumor cell cannibalism.
Together with classic clinicopathologic factors such as TNM stage, CIC subtype may provide more accurate information of prognosis prediction, especially in the situation where traditional prognostic factors have reached the ceiling roof. For ESCC patients with late TNM stage, low TiT could be helpful to identify patients with poorer outcomes who should be given more intensive treatment.
There were still several limitations in this study. First, the retrospective nature of this study may lead to bias inevitably, and well-designed prospective study will be needed. Second, there was no validation cohort in this study which should be considered in the future. Third, some samples came from the commercial TMA and the related treatment information including chemotherapy and radiotherapy were unavailable which may decrease the credibility of result to certain extents. Additionally, the quantification of CIC structures in tissues relays on multiple experienced investigators, which calls for an algorithm-based program for more standard and efficient quantification similar to that achieved recently on cytospins (48).
To sum up, we reported the CIC profiling in ESCC for the first time and preliminarily explored the prognostic value of CIC subtype in this study. TiT was identified as a potent prognostic marker in ESCC and showed more prognostic value in patients with late TNM stage or high risks. Our work also supports the notion that function pathology with CIC profiling is an emerging prognostic factor for human cancers.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding authors.
Ethics Statement
The studies involving human participants were reviewed and approved by The Outdo Biotech Co. Ltd., National Human Genetic Resources Sharing Service Platform. The patients/participants provided their written informed consent to participate in this study.
Author Contributions
QS, HJ, and HH: concept and design. YW and ZN: staining and imaging. YW, ZN, LZ, YAZ, QM, YCZ, ML, YS, JG LG, and JS: data acquisition. QS, HH, and YW: manuscript drafting. YAZ, QM, QJS, and YT: pathological judgment and confirmation. YW and LZ: statistical analysis. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Professor Zhaolie Chen for experimental assistance and guidance. This study was supported by Beijing Municipal Natural Science Foundation (KZ202110025029), the National Natural Science Foundation of China (31970685), and Beijing Municipal Administration of Hospitals Incubating Program (PX2021033).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.670051/full#supplementary-material
References
1. Siegel RL, Miller KD and Jemal A. Cancer Statistics, 2018. CA Cancer J Clin (2018) 68(1):7–30. doi: 10.3322/caac.21442
2. Yang CS, Chen X and Tu S. Etiology and Prevention of Esophageal Cancer. Gastrointest Tumors (2016) 3(1):3–16. doi: 10.1159/000443155
3. Shaheen O, Ghibour A and Alsaid B. Esophageal Cancer Metastases to Unexpected Sites: A Systematic Review. Gastroenterol Res Pract (2017) 2017:1657310. doi: 10.1155/2017/1657310
4. Castro C, Peleteiro B and Lunet N. Modifiable Factors and Esophageal Cancer: A Systematic Review of Published Meta-Analyses. J Gastroenterol (2018) 53(1):37–51. doi: 10.1007/s00535-017-1375-5
5. Arnal MJD. Esophageal Cancer: Risk Factors, Screening and Endoscopic Treatment in Western and Eastern Countries. World J Gastroenterology (2015) 21(26):7933–43. doi: 10.3748/wjg.v21.i26.7933
6. Torre LA, Bray F, Siegel RL, Ferlay J. Lortet-Tieulent J and Jemal a. Global Cancer Statistics, 2012. CA: A Cancer J Clin (2015) 65(2):87–108. doi: 10.3322/caac.21262
7. Sun Q, Huang H and Overholtzer M. Cell-in-Cell Structures are Involved in the Competition Between Cells in Human Tumors. Mol Cell Oncol (2015) 2(4):e1002707. doi: 10.1080/23723556.2014.1002707
8. Fais S, Overholtzer M. Cell-in-Cell Phenomena in Cancer. Nat Rev Cancer (2018) 18(12):758–66. doi: 10.1038/s41568-018-0073-9
9. Huang H, He M, Zhang Y, Zhang B, Niu Z, Zheng Y, et al. Identification and Validation of Heterotypic Cell-in-Cell Structure as an Adverse Prognostic Predictor for Young Patients of Resectable Pancreatic Ductal Adenocarcinoma. Signal Transduct Target Ther (2020) 5(1):246–8. doi: 10.1038/s41392-020-00346-w
10. Fan J, Fang Q, Yang Y, Cui M, Zhao M, Qi J, et al. Role of Heterotypic Neutrophil-in-Tumor Structure in the Prognosis of Patients With Buccal Mucosa Squamous Cell Carcinoma. Front Oncol (2020) 10:541878(2284). doi: 10.3389/fonc.2020.541878
11. Zhang X, Niu Z, Qin H, Fan J, Wang M, Zhang B, et al. Subtype-Based Prognostic Analysis of Cell-in-Cell Structures in Early Breast Cancer. Front Oncol (2019) 9:895. doi: 10.3389/fonc.2019.00895
12. Ruan B, Niu Z, Jiang X, Li Z, Tai Y, Huang H, et al. High Frequency of Cell-in-Cell Formation in Heterogeneous Human Breast Cancer Tissue in a Patient With Poor Prognosis: A Case Report and Literature Review. Front Oncol (2019) 9:1444. doi: 10.3389/fonc.2019.01444
13. Hayashi A, Yavas A, McIntyre CA, Ho YJ, Erakky A, Wong W, et al. Genetic and Clinical Correlates of Entosis in Pancreatic Ductal Adenocarcinoma. Mod Pathol (2020) 33(9):1822–31. doi: 10.1038/s41379-020-0549-5
14. Schwegler M, Wirsing AM, Schenker HM, Ott L, Ries JM, Buttner-Herold M, et al. Prognostic Value of Homotypic Cell Internalization by Nonprofessional Phagocytic Cancer Cells. BioMed Res Int (2015) 2015:359392. doi: 10.1155/2015/359392
15. Huang H, Chen A, Wang T, Wang M, Ning X, He M, et al. Detecting Cell-in-Cell Structures in Human Tumor Samples by E-cadherin/CD68/CD45 Triple Staining. Oncotarget (2015) 6(24):20278–87. doi: 10.18632/oncotarget.4275
16. Zhang B, Wu Q, Li B, Wang D, Wang L and Zhou YL. Ma Regulator-Mediated Methylation Modification Patterns and Tumor Microenvironment Infiltration Characterization in Gastric Cancer. Mol Cancer (2020) 19(1):53. doi: 10.1186/s12943-020-01170-0
17. Alcala N, Leblay N, Gabriel AAG, Mangiante L, Hervas D, Giffon T, et al. Integrative and Comparative Genomic Analyses Identify Clinically Relevant Pulmonary Carcinoid Groups and Unveil the Supra-Carcinoids. Nat Commun (2019) 10(1):3407. doi: 10.1038/s41467-019-11276-9
18. Su Y, Ren H, Tang M, Zheng Y, Zhang B, Wang C, et al. Role and Dynamics of Vacuolar Ph During Cell-in-Cell Mediated Death. Cell Death Dis (2021) 12(1):119. doi: 10.1038/s41419-021-03396-2
19. Wang M, Niu Z, Qin H, Ruan B, Zheng Y, Ning X, et al. Mechanical Ring Interfaces Between Adherens Junction and Contractile Actomyosin to Coordinate Entotic Cell-in-Cell Formation. Cell Rep (2020) 32(8):108071. doi: 10.1016/j.celrep.2020.108071
20. Sun Q, Cibas ES, Huang H. Hodgson L and Overholtzer M. Induction of Entosis by Epithelial Cadherin Expression. Cell Res (2014) 24:1288–98. doi: 10.1038/cr.2014.137
21. Niu Z, He M and Sun Q. Molecular Mechanisms Underlying Cell-in-Cell Formation: Core Machineries and Beyond. J Mol Cell Biol (2021) 24:1288–98. doi: 10.1093/jmcb/mjab015
22. Liang J, Fan J, Wang M, Niu Z, Zhang Z, Yuan L, et al. Cdkn2a Inhibits Formation of Homotypic Cell-in-Cell Structures. Oncogenesis (2018) 7(6):1–8. doi: 10.1038/s41389-018-0056-4
23. Wang C, Chen A, Ruan B, Niu Z, Su Y, Qin H, et al. Pcdh7 Inhibits the Formation of Homotypic Cell-in-Cell Structure. Front Cell Dev Biol (2020) 8:329(329). doi: 10.3389/fcell.2020.00329
24. Ruan B, Wang C, Chen A, Liang J, Niu Z, Zheng Y, et al. Expression Profiling Identified IL-8 as a Regulator of Homotypic Cell-in-Cell Formation. BMB Rep (2018) 51(8):412–7. doi: 10.5483/BMBRep.2018.51.8.089
25. Ruan B, Zhang B, Chen A, Yuan L, Liang J, Wang M, et al. Cholesterol Inhibits Entotic Cell-in-Cell Formation and Actomyosin Contraction. Biochem Biophys Res Commun (2018) 495(1):1440–6. doi: 10.1016/j.bbrc.2017.11.197
26. Sun Q, Luo T, Ren Y, Florey O, Shirasawa S, Sasazuki T, et al. Competition Between Human Cells by Entosis. Cell Res (2014) 24:1299–310. doi: 10.1038/cr.2014.138
27. Kroemer G, Perfettini J-L. Entosis, a Key Player in Cancer Cell Competition. Cell Res (2014) 24(11):1280–1. doi: 10.1038/cr.2014.133
28. Huang H, Chen Z, Sun Q. Mammalian Cell Competitions, Cell-in-Cell Phenomena and Their Biomedical Implications. Curr Mol Med (2015) 15(9):852–60. doi: 10.2174/1566524015666151026101101
29. Sedwick C, Overholtzer M. Answering Existential Questions on Entosis. J Cell Biol (2011) 15(9):852–60. doi: 10.1083/jcb.1956pi
30. Liang J, Niu Z, Zhang B, Yu X, Zheng Y, Wang C, et al. P53-Dependent Elimination of Aneuploid Mitotic Offspring by Entosis. Cell Death Differ (2021) 28(2):799–813. doi: 10.1038/s41418-020-00645-3
31. Ni C, Huang L, Chen Y, He M, Hu Y, Liu S, et al. Implication of Cell-in-Cell Structures in the Transmission of HIV to Epithelial Cells. Cell Res (2015) 25(11):1265–8. doi: 10.1038/cr.2015.119
32. Ni C, Chen Y, Zeng M, Pei R, Du Y, Tang L, et al. In-Cell Infection: A Novel Pathway for Epstein-Barr Virus Infection Mediated by Cell-in-Cell Structures. Cell Res (2015) 25(7):785–800. doi: 10.1038/cr.2015.50
33. Zhang Z, Zheng Y, Niu Z, Zhang B, Wang C, Yao X, et al. Sars-Cov-2 Spike Protein Dictates Syncytium-Mediated Lymphocyte Elimination. Cell Death Differ (2021) 25(7):785–800. doi: 10.1038/s41418-021-00782-3
34. Gupta K, Dey P. Cell Cannibalism: Diagnostic Marker of Malignancy. Diagn Cytopathol (2003) 28(2):86–7. doi: 10.1002/dc.10234
35. Schenker H, Buttner-Herold M, Fietkau R and Distel LV. Cell-in-Cell Structures are More Potent Predictors of Outcome Than Senescence or Apoptosis in Head and Neck Squamous Cell Carcinomas. Radiat Oncol (2017) 12(1):21. doi: 10.1186/s13014-016-0746-z
36. Kong Y, Liang Y and Wang J. Foci of Entotic Nuclei in Different Grades of Noninherited Renal Cell Cancers. IUBMB Life (2015) 67(2):139–44. doi: 10.1002/iub.1354
37. Barresi V. Phagocytosis (Cannibalism) of Apoptotic Neutrophils by Tumor Cells in Gastric Micropapillary Carcinomas. World J Gastroenterol (2015) 21(18):139–44. doi: 10.3748/wjg.v21.i18.5548
38. Routray S, Shankar AA and Swain N. Does Entosis Curb the Detached Cancer Cells Better? Oral Oncol (2014) 50(3):e9–e11. doi: 10.1016/j.oraloncology.2013.11.010
39. Sharma N, Dey P. Cell Cannibalism and Cancer. Diagn Cytopathol (2011) 39(3):229–33. doi: 10.1002/dc.21402
40. Fernandez-Flores A. Cannibalism in a Benign Soft Tissue Tumor (Giant-Cell Tumor of the Tendon Sheath, Localized Type): A Study of 66 Cases. Rom J Morphol Embryol (2012) 53(1):15–22. doi: 10.1016/j.oraloncology.2013.11.010
41. Kimura N, Dota K, Araya Y, Ishidate T and Ishizaka M. Scoring System for Differential Diagnosis of Malignant Mesothelioma and Reactive Mesothelial Cells on Cytology Specimens. Diagn Cytopathol (2009) 37(12):885–90. doi: 10.1002/dc.21128
42. Yokoo H, Isoda K, Nakazato Y, Nakayama Y, Suzuki Y, Nakamura T, et al. Retroperitoneal Epithelioid Angiomyolipoma Leading to Fatal Outcome. Pathol Int (2000) 50(8):649–54. doi: 10.1046/j.1440-1827.2000.01096.x
43. Abodief WT, Dey P and Al-Hattab O. Cell Cannibalism in Ductal Carcinoma of Breast. Cytopathology (2006) 17(5):304–5. doi: 10.1111/j.1365-2303.2006.00326.x
44. Lugini L, Matarrese P, Tinari A, Lozupone F, Federici C, Iessi E, et al. Cannibalism of Live Lymphocytes by Human Metastatic But Not Primary Melanoma Cells. Cancer Res (2006) 66(7):3629–38. doi: 10.1158/0008-5472.CAN-05-3204
45. Fais S, Fauvarque M-O. TM9 and Cannibalism: How to Learn More About Cancer by Studying Amoebae and Invertebrates. Trends Mol Med (2012) 18(1):4–5. doi: 10.1016/j.molmed.2011.09.001
46. Lozupone F, Perdicchio M, Brambilla D, Borghi M, Meschini S, Barca S, et al. The Human Homologue of Dictyostelium Discoideum phg1A is Expressed by Human Metastatic Melanoma Cells. EMBO Rep (2009) 10(12):1348–54. doi: 10.1038/embor.2009.236
47. Lozupone F, Borghi M, Marzoli F, Azzarito T, Matarrese P, Iessi E, et al. TM9SF4 is a Novel V-Atpase-Interacting Protein That Modulates Tumor Ph Alterations Associated With Drug Resistance and Invasiveness of Colon Cancer Cells. Oncogene (2015) 34(40):5163–74. doi: 10.1038/onc.2014.437
Keywords: subtype, entosis, EML staining, multiplex staining, prognosis, cell-in-cell structures, esophageal squamous cell carcinoma, functional pathology
Citation: Wang Y, Niu Z, Zhou L, Zhou Y, Ma Q, Zhu Y, Liu M, Shi Y, Tai Y, Shao Q, Ge J, Hua J, Gao L, Huang H, Jiang H and Sun Q (2021) Subtype-Based Analysis of Cell-in-Cell Structures in Esophageal Squamous Cell Carcinoma. Front. Oncol. 11:670051. doi: 10.3389/fonc.2021.670051
Received: 20 February 2021; Accepted: 19 May 2021;
Published: 11 June 2021.
Edited by:
Paolo Nuciforo, Vall d’Hebron Institute of Oncology (VHIO), SpainReviewed by:
Stefano Fais, National Institute of Health (ISS), ItalyArtur Mezheyeuski, Uppsala University, Sweden
Copyright © 2021 Wang, Niu, Zhou, Zhou, Ma, Zhu, Liu, Shi, Tai, Shao, Ge, Hua, Gao, Huang, Jiang and Sun. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hongyan Huang, aGhvbmd5MTk5OUAxMjYuY29t; Hong Jiang, amhvbmdAYmp0dS5lZHUuY24=; Qiang Sun, c3VucUBibWkuYWMuY24=
†These authors have contributed equally to this work
 Yuqi Wang
Yuqi Wang Zubiao Niu
Zubiao Niu Lulin Zhou2,3†
Lulin Zhou2,3† Hongyan Huang
Hongyan Huang Qiang Sun
Qiang Sun