- 1Department of Otolaryngology, Wuwei People’s Hospital, Wuwei, China
- 2Department of Otolaryngology, The First Hospital of Lanzhou University, Lanzhou, China
Background: The involvement of microRNA-338-5p in modulating NPC pathogenesis is still largely unknown, and this study aimed to investigate this issue.
Methods: The expressions of cancer associated genes were determined by Real-Time qPCR and Western Blot, and cell apoptosis was determined by flow cytometer (FCM). CCK-8 assay and colony formation assay were respectively used to determine cell proliferation and colony formation abilities. Transwell assay was used to evaluate cell migration. The expression levels of Ki67 protein in mice tissues were measured by Immunohistochemistry (IHC) assay.
Results: The present study found that microRNA-338-5p suppressed NPC progression by degrading its downstream target, Wnt family member 2B (WNT2B). Specifically, microRNA-338-5p tended to be low-expressed in NPC tissues and cell lines, compared to the non-tumor nasopharyngeal mucosa tissues and normal nasopharyngeal cell line (NP69). Upregulation of microRNA-338-5p inhibited proliferation, mobility, and epithelial-mesenchymal transition (EMT) in NPC cells in vitro, while silencing of microRNA-338-5p had opposite effects. Consistently, microRNA-338-5p suppressed tumorigenesis of NPC cells in vivo. In addition, microRNA-338-5p targeted WNT2B for degradation and inhibition, and the inhibiting effects of microRNA-338-5p overexpression on NPC development were reversed by upregulating WNT2B.
Conclusions: Taken together, we concluded that microRNA-338-5p targeted WNT2B to hinder NPC development.
Background
Nasopharyngeal carcinoma (NPC) is a common head and neck malignancy worldwide (1, 2), that is characterized by highly malignant local invasion and distant metastasis (3, 4), and imposes a huge health burden on human beings. Unfortunately, clinical data indicated that the traditional therapies, such as radiotherapy and chemoresistance, were not efficacious to cure NPC as the result of radio-resistance (5, 6) and chemo-resistance (7, 8), resulting in worse prognosis in NPC patients. Therefore, development of alternative therapy strategies for NPC became necessary and meaningful (9, 10), and uncovering the underlying mechanisms of NPC pathogenesis might be the first step. Recently, researchers focused on screening out tumor suppressors and oncogenes in NPC, and those cancer associated genes included the genes with or without coding abilities (11–14). Especially, the non-coding RNAs always participated in the regulation of cellular functions via serving as post-transcriptional regulators, and our team concentrated on delving into the regulating mechanisms of non-coding RNAs, such as circular RNAs (circ-RNAs) (15, 16), long non-coding RNAs (LncRNAs) (13, 14), and microRNAs (miRNAs) (17, 18), in NPC.
Among all the non-coding RNAs, multiple miRNAs had been identified to regulate NPC development in previous publications (17, 18). For example, miR-99a inhibited cell proliferation and served as a prognostic factor in NPC (19), while other researchers noticed that miR-155 acted as an oncogene to facilitate NPC development (20). Interestingly, the role of microRNA-338-5p in regulating cancer progression varied according to different cancer types (21, 22). On the one hand, microRNA-338-5p hampered the development of esophageal squamous cancer (22), on the other, upregulation of microRNA-338-5p promoted cancer metastasis in colorectal cancer (21). Of note, Ying Shan et al. noticed that microRNA-338-5p inhibited migration and proliferation of NPC cells (23), however, the detailed molecular mechanisms still need to be elucidated. According to previous publications (21, 22, 24), miRNAs targeted the 3’ untranslated regions (3’UTR) of their downstream target genes, and the existing information suggested that microRNA-338-5p negatively regulated hypoxia-induced factor 1α (HIF-1α) in NPC cells (23). Our preliminary data suggested that there existed potential binding sites between microRNA-338-5p and 3’UTR of Wnt family member 2B (WNT2B), and WNT2B had been identified as an oncogene to promote NPC progression (25, 26). Nevertheless, up until now, no literature reported the regulating mechanisms of microRNA-338-5p and WNT2B in NPC cells.
Thus, we designed this study to investigate the role of the microRNA-338-5p/WNT2B axis in regulating NPC pathogenesis, which not only broadened our knowledge in this field, but provided novel biomarkers for NPC diagnosis and treatment.
Methods
Clinical Samples Collection and Preparation
The NPC tissues (N = 25) and non-tumor nasopharyngeal mucosa tissues (N = 20) were collected from 2014 to 2017 in Wuwei People’s Hospital. None of the patients accepted other therapies, including chemotherapy, radiotherapy, and adjuvant therapies, before surgical resection, and the NPC patients were histologically and clinically judged by two experienced pathologists, and the clinical tissues were frozen at −70°C conditions. The written informed consent forms had been obtained from all the participants, and our clinic-associated experiments were all allowed by the Ethics Committee in Wuwei People’s Hospital, and the approval number was No. 2017DS632913.
Cell Culture and Vectors Delivery
We purchased the NPC cell line SUNE1 and SUNE2, and normal nasopharyngeal cell line NP69 from the Cell Bank of the Chinese Academy of Sciences (China) and American Type Culture Collection (ATCC, USA). As previously documented (27), the cells were cultured in Dulbecco’s Modified Eagle’s medium (Gibco, USA) containing 10% fetal bovine serum (FBS, Gibco, USA) under the culture conditions at 37°C with 5% CO2 humidified atmosphere. The microRNA-338-5p mimic (50 nM) and inhibitor (100 nM) and WNT2B overexpression vectors (100 nM) were designed and synthesized by a commercial third-party company (GenePharma, Shanghai, China), which were all transfected into the cells by using the Lipofectamine 2000 reagent (Invitrogen, USA).
Real-Time qPCR
A commercial Trizol kit (Invitrogen, USA) was used for RNA extraction in NPC cells and tissues, and Real-Time qPCR was conducted to quantify microRNA-338-5p and WNT2B mRNA, which were respectively normalized by U6 and β-actin. The detailed experimental procedures could be found in the previous publication (28), and the primer sequences had been documented in Table 1.
Western Blot Analysis
Protein extraction from NPC cells and tissues were finished by using the RIPA lysis buffer reagent (Beyotime, Shanghai, China), and the protein quality was determined by BCA kit (Beyotime, Shanghai, China). Next, proteins were separated by SDS-PAGE, transferred onto the PVDF membranes (Millipore, USA), and incubated with primary antibodies against WNT2B, N-cadherin, and Vimentin, and subsequently with the secondary antibodies. Finally, an electrochemiluminescence (ECL) system was used for protein bands visualization, which were analyzed by using the Image J software. The antibodies’ information had been recorded in Table 2.
Cell-Counting Kit-8 (CCK-8) Assay
A commercial CCK-8 kit (YEASEN Biotechnology, Shanghai, China) was used to examine cell proliferation in NPC cells in keeping with the manufacturer’s protocol. Specifically, NPC cells were cultured for different time points, and were subsequently incubated with CCK-8 reaction solution for 2.5 h, which were fully vortexed, and a microplate reader (ThermoFisher Scientific, USA) was employed to determine the optical density (OD) values with 450 nm wavelength.
Transwell Assay
The NPC cells were cultured in the serum-free medium of the upper chamber of the Invasion Chamber (BD Bioscience, CA, USA) with Matrigel-coated membranes, and the same volume of the medium with 10% FBS was added in the lower chamber to serve as the chemotactic content. The above cells were cultured at 37°C in the incubator for 24 h, and the cells in the upper surface of the membranes were removed, and cells in the lower chamber were fixed with methanol and stained with 0.1% crystal violet for visualization. The stained cells were counted under an invert light microscope.
Colony Formation Assay
As previously described (29), the colonies, formation abilities in NPC cells were determined by performing colony formation assay. Specifically, NPC cells were cultured in 96-well plates with 1,000 cells/well at 37°C for 14 days, and were subsequently stained with 0.1% crystal violet (Beyotime Biotechnology, Shanghai, China). Then, a light microscope (ThermoFisher Scientific, USA) was used to count the colonies above 15 cells.
Flow Cytometry (FCM) for Cell Apoptosis Analysis
The NPC cells were respectively stained with Annexin V-FITC and propidium iodide (PI) at room temperature in darkness, and Flow Cytometer (Partec, Germany) was employed to examine cell apoptosis ratio. The cells double stained with Annexin V-FITC and PI were regarded as late apoptosis, with Annexin V-FITC alone being early apoptosis, and with PI alone being necroptosis.
Bioinformatics Analysis
The targeting sites microRNA-338-5p and 3’UTR of WNT2B mRNA were predicted by the online starBase software (http://starbase.sysu.edu.cn/), and the association of microRNA-338-5p with neck squamous cell carcinoma (HNSC) was analyzed by using the Pan-cancer analysis (http://starbase.sysu.edu.cn/panCancer.php).
Dual-Luciferase Reporter Gene System Assay
The binding sites in the 3’UTR of WNT2B mRNA were mutated, and the wild-type and mutant WNT2B sequences were cloned into the luciferase reporter plasmids, and named as Wt-WNT2B and Mut-WNT2B, respectively. The above vectors were co-transfected with miR-NC, microRNA-338-5p mimic, and inhibitor into the NPC cells by using the commercial Lipofectamine 2000 reagent (Invitrogen, USA) according to the manufacturer’s protocol. After 48 h post-transfection, the Dual-luciferase reporter gene system (Promega, USA) was employed to determine both firefly luciferase and renilla luciferase activities, and the Luminometer (Promega, USA) was performed to estimate the relative luciferase activities.
Establishment of Mice Models
The nude female mice (5 weeks old) were purchased from Lanzhou University, and the mice were fed under specific pathogen-free circumstances. The NPC cells were subcutaneously injected into the right flanks of mice at the density of 2 × 106 cells per mouse, and each group contained at least five mice. The caliper was used to measure tumor volumes every 5 days, and at day 25, mice were anesthetized by intravenously injecting Barbiturate (100 mg/kg), and sacrificed by using the cervical dislocation method. The tumors were obtained for further analysis, and the animal experiments were approved by Wuwei People’s Hospital (No. 2017DS632823).
Immunohistochemistry (IHC) for Ki67 Protein Examination
The expression patterns, including localization and expression levels, were examined by using the IHC assay in mice tumor tissues. The detailed experimental procedures could be found in previous publications (30–32). Briefly, the mice tumor tissues were fixed by paraffin and embedded by wax, and were spliced into about 4-μm thick sections. Next, the IHC was performed by using a Benchmark XT automated staining machine (Ventana, USA), and the primary antibody against Ki67 (Ventana, USA) was diluted into 1:100 to incubate with the tumor tissues. After that, the sections were incubated with the secondary antibody labeled with horseradish peroxidase (HRP, Ventana, USA), and the 3, 3’ diaminobenzidine (DAB) was used to visualize the Ki-67 positive cells. A light microscope was employed to photograph the images to evaluate the expression status of Ki67 protein in mice tumor tissues, and the cells stained in yellow were regarded as Ki67-positive cells.
Analysis of the Data
Data were presented as Means ± Standard Deviation (SD) and analyzed by SPSS 18.0 software (IBM, USA) and GraphPad Prism 8 (GraphPad Software, USA). Means from two groups were compared by Student’s t-test, and means in multiple groups were analyzed by one-way analysis of variance (ANOVA). Besides, genes’ correlations were analyzed by Pearson Correlation analysis. *P < 0.05 meant statistical significance.
Results
MicroRNA-338-5p Was Aberrantly Downregulated in NPC Tissues and Cells
The NPC tissues (N = 25) and non-tumor nasopharyngeal mucosa tissues (N = 20) were obtained from NPC patients, and Real-Time qPCR was conducted to examine the expression levels of microRNA-338-5p in the above clinical tissues (Figure 1A). As shown in Figure 1A, we validated that microRNA-338-5p was downregulated in NPC tissues, in contrast with the normal tissues (P < 0.05). Consistently, by performing the Pan-cancer analysis, we found that microRNA-338-5p also tended to be low-expressed in the cancer tissues collected from 497 patients with head and neck squamous cell carcinoma (HNSC), instead of the 44 normal samples (P < 0.05, Figure 1B). In addition, the HNSC patients with low-expressed microRNA-338-5p tended to have a worse prognosis although without statistical significance (P = 0.1, Figure 1C). Also, the above results were validated by our cellular experiments, which showed that lower levels of microRNA-338-5p were observed in NPC cell lines (SUNE1 and SUNE2), compared to the normal nasopharyngeal cell line (NP69) (P < 0.05, Figure 1D).
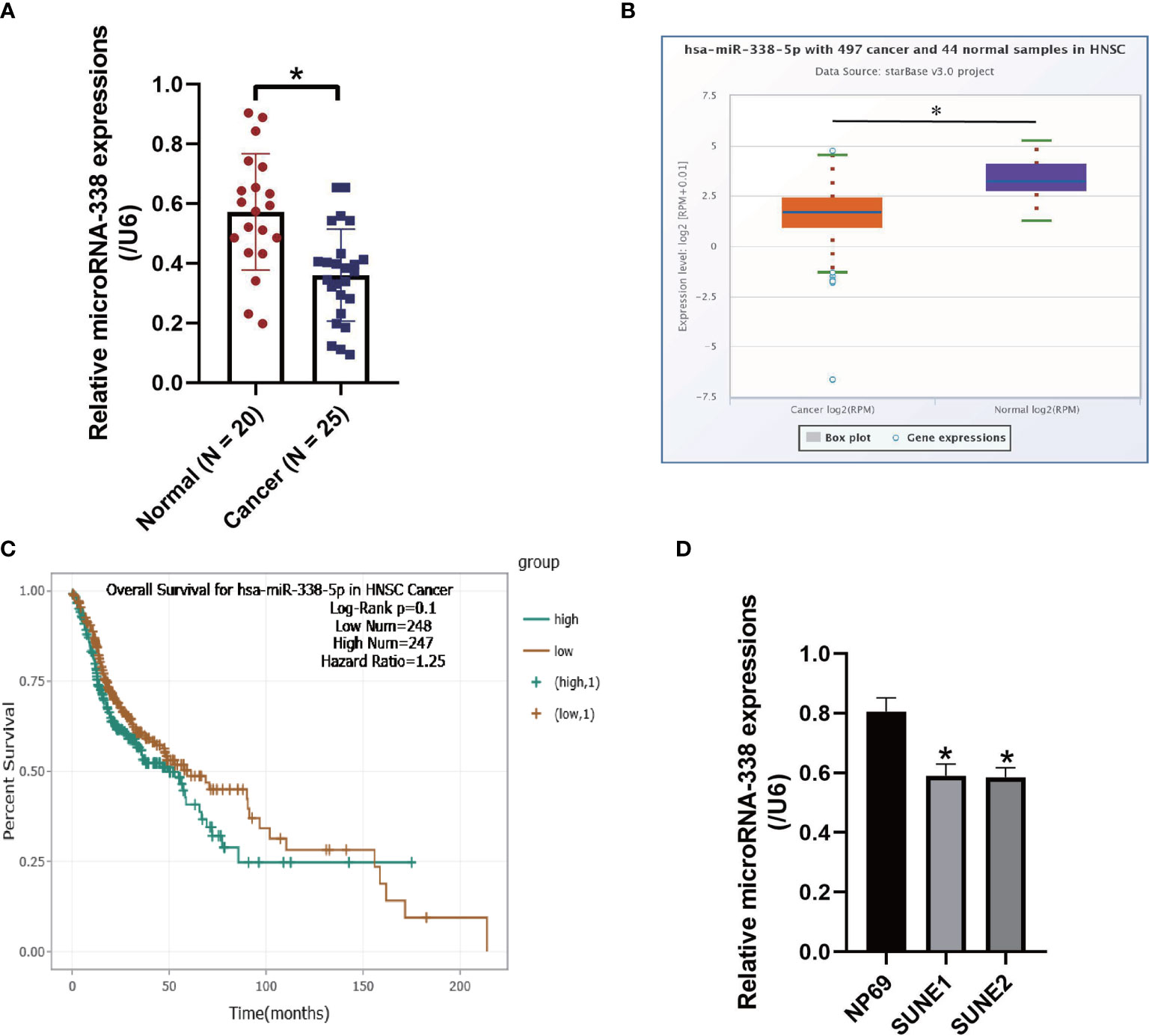
Figure 1 The expression status of microRNA-338-5p were examined in the NPC clinical tissues and cell lines. (A) The NPC tissues (N = 25) and non-tumor nasopharyngeal mucosa tissues (N = 20) were obtained, and Real-Time qPCR was conducted to examine the expression levels of microRNA-338-5p. (B) The expression levels of microRNA-338-5p in the clinical tissues collected from neck squamous cell carcinoma (HNSC) were analyzed by using the online Pan-cancer analysis. (C) Pan-cancer analysis was performed to analyze the correlations of microRNA-338-5p levels and patients’ prognosis in HNSC. (D) Real-Time qPCR was used to examine the levels of microRNA-338-5p in NPC cells. Each experiment repeated at least three times, and *P < 0.05.
MicroRNA-338-5p Negatively Regulated Cell Growth and Viability in NPC In Vitro and In Vivo
Previous publications reported that the role of microRNA-338-5p in regulating cancer progression is controversial according to cancer types (21, 22), and our data suggested that microRNA-338-5p acted as a tumor suppressor to hinder the development of NPC (Figure 2). Specifically, the microRNA-338-5p mimic and inhibitor were constructed and delivered into NPC cells (SUNE1 and SUNE2) to overexpress and downregulate microRNA-338-5p (Figure S1), and the CCK-8 assay results showed that microRNA-338-5p negatively regulated cell proliferation abilities in NPC cells (P < 0.05, Figures 2A, B). Consistently, the colony formation assay evidenced that microRNA-338-5p inhibited colonies formation abilities in NPC cells (P < 0.05, Figure 2C). Next, by performing the Annexin V-FITC/PI double staining assay, we found that overexpression of microRNA-338-5p triggered apoptotic cell death in NPC cells (P < 0.05, Figure 2D). In addition, the SUNE1 and SUNE2 cells with differential vectors transfection were used to establish xenograft tumor-bearing mice models, and the results suggested that microRNA-338-5p overexpression slowed down tumor growth (P < 0.05, Figures 2E–G) and decreased the expression levels of Ki67 protein (Figure 2H) to inhibit tumorigenesis of NPC cells in vivo, while silencing of microRNA-338-5p had opposite effects (Figures 2E–H).
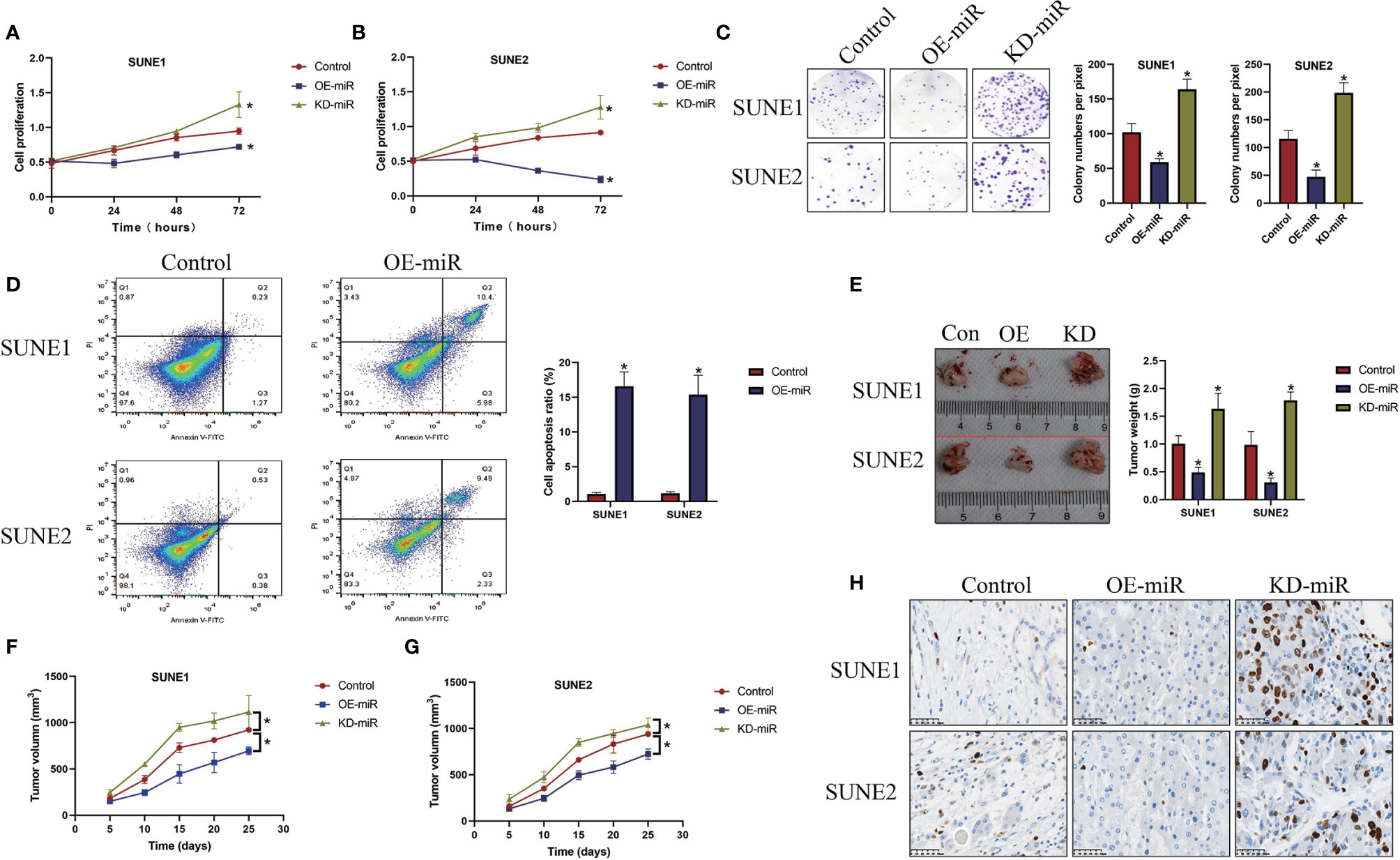
Figure 2 MicroRNA-338-5p inhibited NPC cell proliferation and viability in vitro and in vivo. (A, B) The NPC cells (SUNE1 and SUNE2) were pre-transfected with different vectors, and were cultured for 0, 24, 48, and 72 h, respectively. The CCK-8 assay was performed to evaluate cell proliferation abilities. (C) Colony formation assay was used to determine colonies formation abilities in NPC. (D) The NPC cells were stained with Annexin V-FITC and PI, and cell apoptosis ratio was measured by flow cytometer (FCM). The xenograft tumor-bearing mice models were established, and (E) tumor weight and (F, G) volume were measured and documented. (H) Immunohistochemistry (IHC) assay was performed to examine the localization and expression levels of Ki67 protein in mice tumor tissues. Each experiment repeated at least three times, and *P < 0.05.
Upregulation of MicroRNA-338-5p Inhibited NPC Cell Mobility In Vitro
Next, we examined the effects of microRNA-338-5p on cell mobility in NPC cells in vitro (Figure 3). As shown in Figures 3A, B, the transwell assay results showed that upregulation of microRNA-338-5p inhibited cell migration abilities in both SUNE1 and SUNE2 cells (P < 0.05), which were significantly inhibited by silencing miR-338-5p (P < 0.05). Also, microRNA-338-5p negatively regulated epithelial-mesenchymal transition (EMT) in NPC cells (Figures 3C–F). Specifically, the Western Blot analysis results indicated that overexpression of microRNA-338-5p decreased the expression levels of N-cadherin and Vimentin to hamper EMT in NPC cells, and microRNA-338-5p knock-down had opposite effects (P < 0.05, Figures 3C–F).
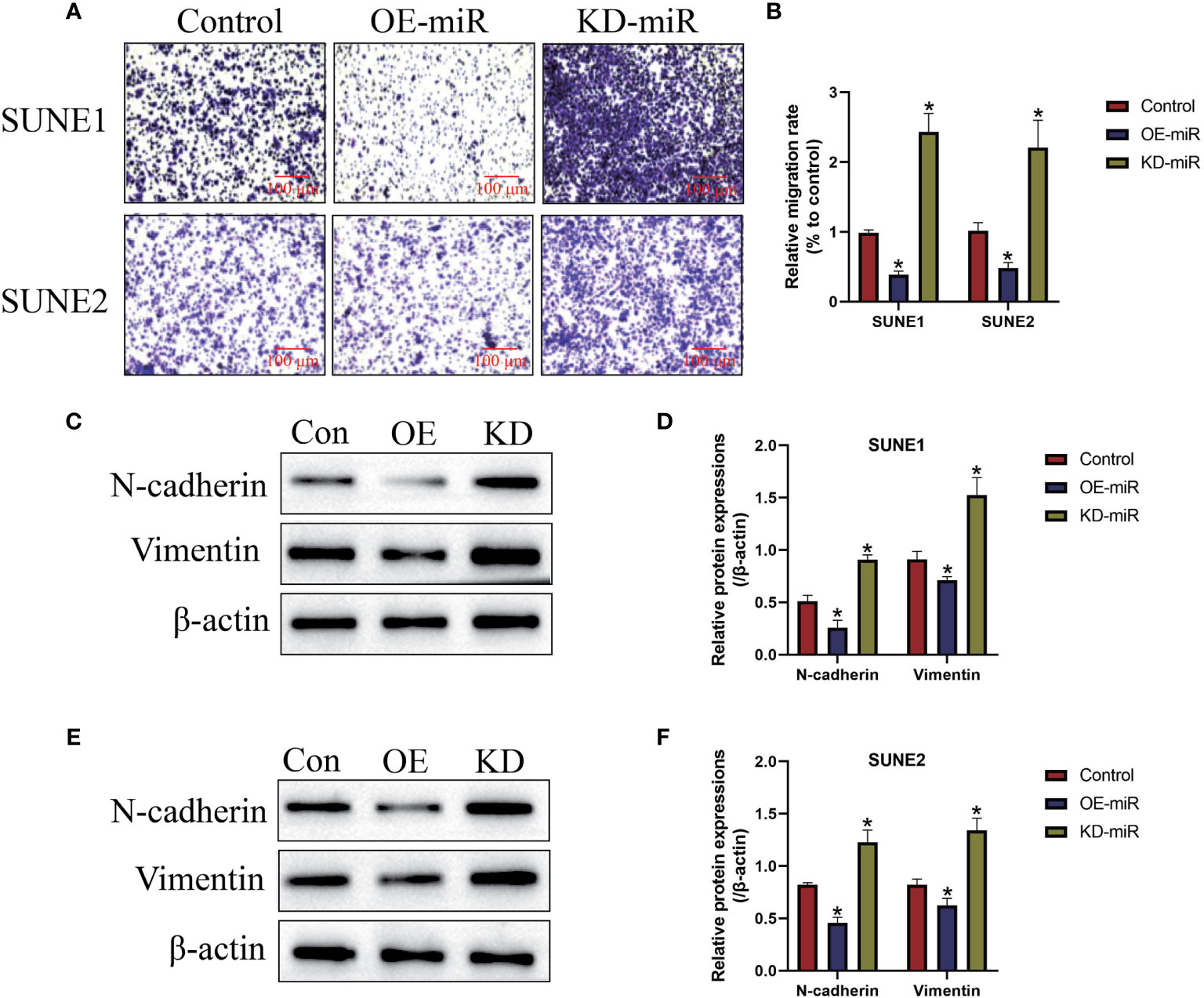
Figure 3 The regulating effects of microRNA-338-5p on cell migration and epithelial-mesenchymal transition (EMT). (A, B) Transwell assay was employed to detect cell migration abilities in NPC cells. (C–F) Western Blot analysis was conducted to examine the expression levels of EMT associated proteins (N-cadherin and Vimentin) in NPC cells. Each experiment repeated at least three times, and *P < 0.05.
The Regulating Mechanisms of MicroRNA-338-5p and WNT2B in NPC Cells
The online starBase software (http://starbase.sysu.edu.cn/) predicted that miR-338-5p potentially bound to the 3’ UTR of WNT2B mRNA (Figure 4A), and previous data suggested that microRNAs served as post-transcriptional regulators for their downstream target genes. Therefore, we conjectured that there might exist potential regulating mechanisms between miR-338-5p and WNT2B in NPC cells. To investigate this issue, the targeting sites in WNT2B were mutated, and were co-transfected with miR-338-5p mimic and inhibitor into SUNE1 and SUNE2 cells, respectively. The dual-luciferase reporter gene system results indicated that miR-338-5p overexpression decreased the luciferase activity in NPC cells co-transfected with wild-type WNT2B (wt-WNT2B), instead of the mutant WNT2B (Mut-WNT2B), while silencing of miR-338-5p had opposite effects (P < 0.05, Figures 4B, C). Next, by performing Real-Time qPCR (Figure 4D) and Western Blot analysis (Figure 4E), we proved that miR-338-5p negatively regulated WNT2B expressions in NPC cells at both transcriptional and translated levels. Also, WNT2B mRNA was upregulated in NPC clinical tissues, in contrast with the normal tissues (P < 0.05, Figure 4F). Interestingly, miR-338-5p negatively correlated with WNT2B mRNA in NPC tissues (P < 0.05, Figure 4G).
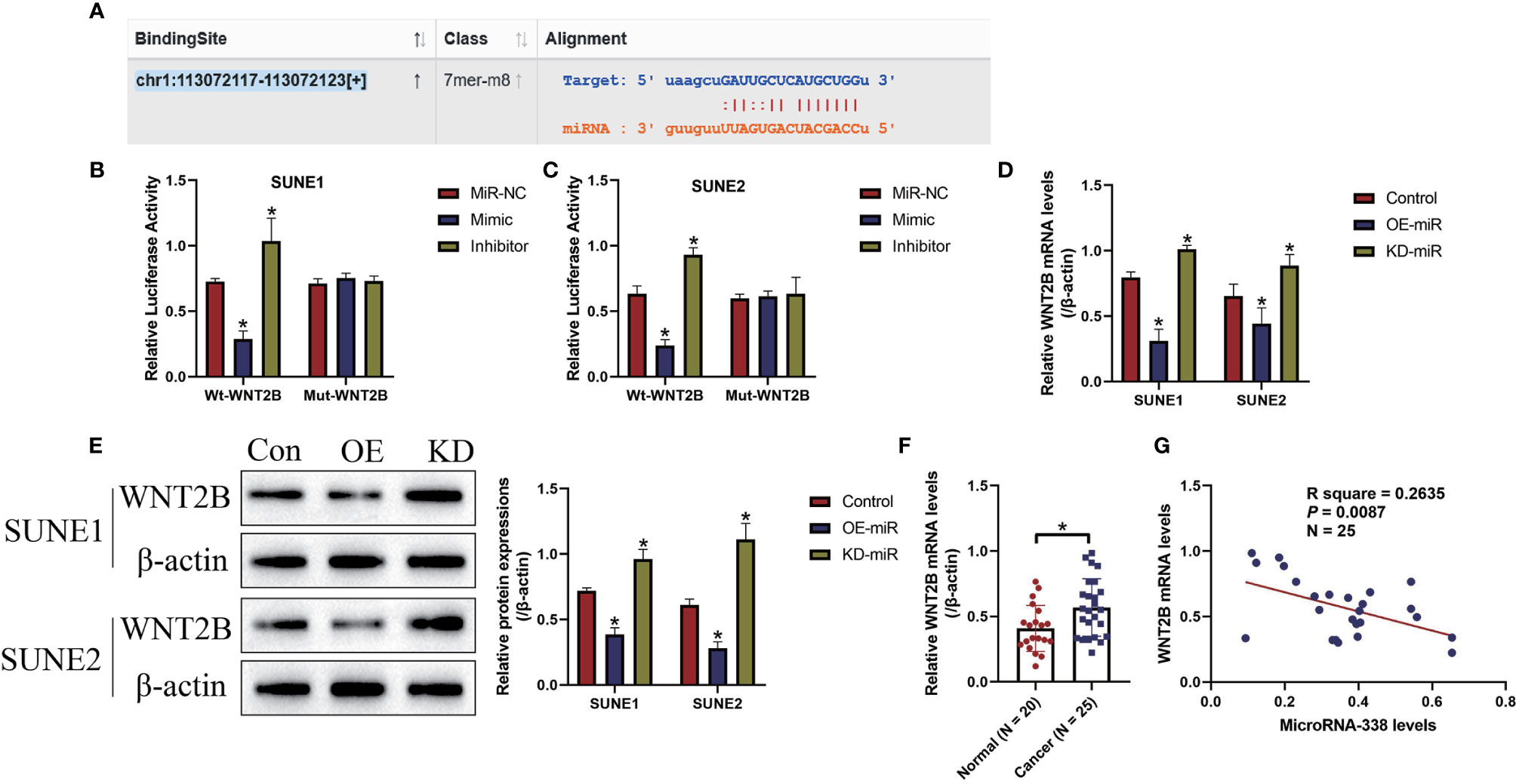
Figure 4 MicroRNA-338-5p targeted 3’ untranslated region (3’UTR) of WNT2B mRNA for its degradation and inhibition. (A) The binding sites in microRNA-338-5p and 3’UTR of WNT2B mRNA were predicted by using the online starBase software (http://starbase.sysu.edu.cn/). (B, C) The above targeting sites were validated by the dual-luciferase reporter gene system assay in NPC cells. MicroRNA-338-5p negatively regulated the expression levels of WNT2B at both (D) transcriptional and (E) translated levels, determined by Real-Time qPCR and Western Blot analysis, respectively. (F) WNT2B mRNA was upregulated in NPC tissues, compared to the normal tissues. (G) Pearson correlation analysis was performed to analyze the correlations between microRNA-338-5p and WNT2B mRNA in NPC tissues. Each experiment repeated at least three times, and *P < 0.05.
Overexpression of MicroRNA-338-5p Hindered NPC Progression by Targeting WNT2B
Given that miR-338-5p inhibited WNT2B expressions in NPC cells, we next investigated whether miR-338-5p inhibited NPC development by targeting WNT2B. To achieve this, the miR-NC, microRNA-338-5p mimic (OE-miR), and WNT2B overexpression vectors (OE-WNT2B) were delivered into NPC cells, respectively, and the results in Figure S2 suggested that the OE-WNT2B vectors had been successfully delivered into NPC cells (P < 0.05). As expected, the CCK-8 assay (Figures 5A, B) and colony formation assay (Figure 5C) results showed that WNT2B overexpression abrogated the inhibiting effects of upregulated microRNA-338-5p on both cell proliferation and colonies, formation abilities (P < 0.05). Consistently, as shown in Figures 5D, E, the Annexin V-FITC/PI double staining assay results showed that overexpression of microRNA-338-5p induced cell apoptosis in NPC cells, which were reversed by upregulating WNT2B (P < 0.05). Furthermore, the transwell assay was performed to evaluate cell migration, and we validated that microRNA-338-5p inhibited NPC cell migration through targeting WNT2B (P < 0.05, Figures 5F, G).
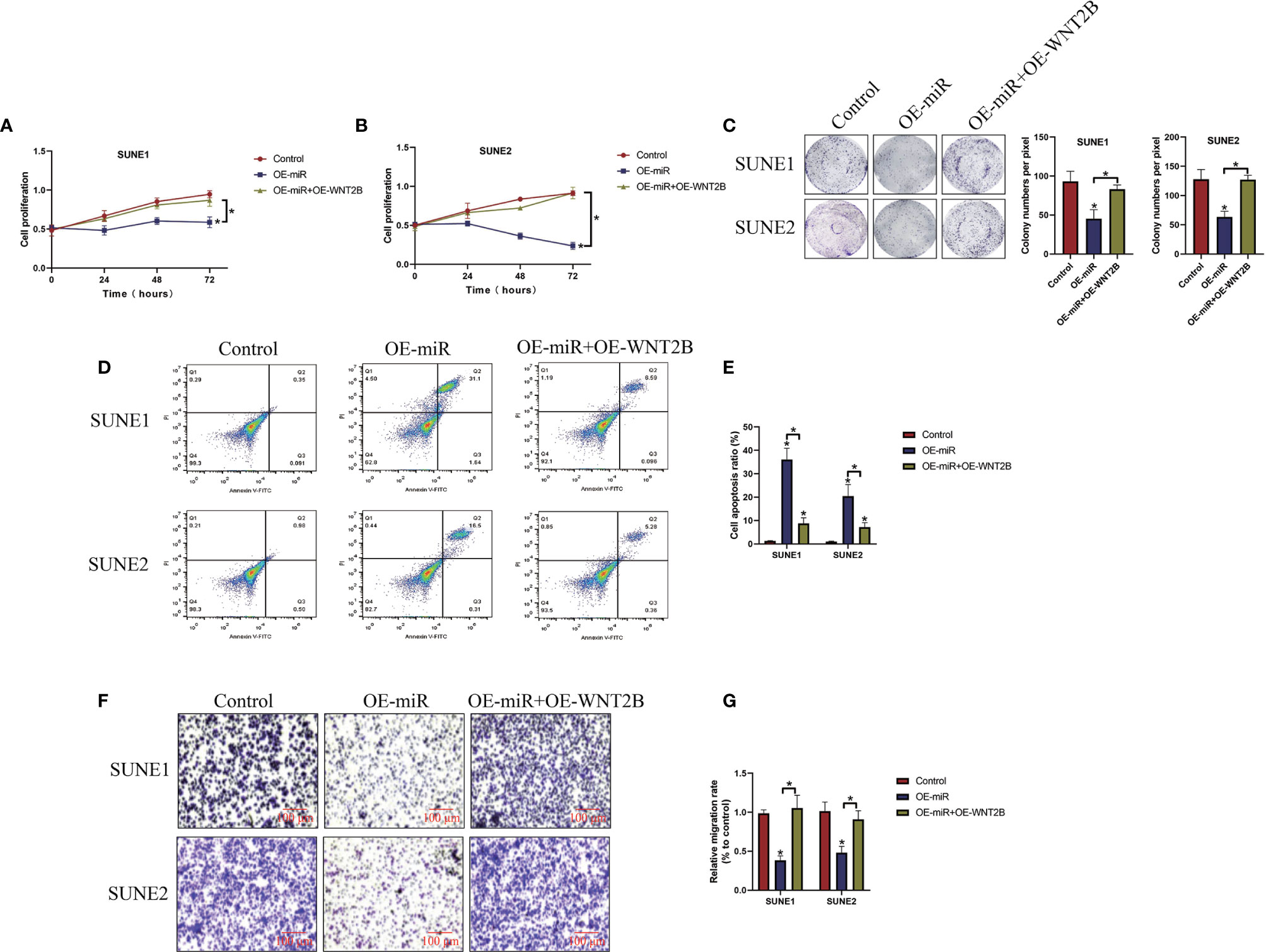
Figure 5 Overexpression of microRNA-338-5p inhibited cancer progression in NPC by inhibiting WNT2B. The NPC cells were transfected with microRNA-338-5p mimic and WNTB overexpression vectors, respectively. (A, B) CCK-8 assay was used to examine cell proliferation abilities. (C) Colony formation assay was used to determine colonies formation abilities in NPC cells. (D, E) Annexin V-FITC/PI double staining assay was performed to measure cell apoptosis ratio. (F, G) Transwell assay was performed to examine cell migration abilities. Each experiment repeated at least three times, and *P < 0.05.
Discussion
Emerging evidence suggested that miRNAs played important roles in regulating NPC pathogenesis, and targeting the cancer associated miRNAs had been proven as novel strategies to hinder the development of NPC (17, 18). Among all the miRNAs, microRNA-338-5p was identified as a tumor suppressor in colorectal cancer (21), but it served as an oncogene to promote esophageal squamous cancer progression (22), suggesting that the role of microRNA-338-5p in regulating cancer development was controversial based on different cancer types. According to the data from Ying Shan et al., microRNA-338-5p inhibited migration and proliferation of NPC cells (23), but the detailed mechanisms were still not fully delineated. In line with the previous work (23), we validated that microRNA-338-5p functioned as a tumor suppressor to hamper the development of NPC in vitro and in vivo by targeting its downstream target Wnt family member 2B (WNT2B). Mechanistically, overexpression of microRNA-338-5p inhibited cell proliferation, colonies, formation abilities, migration, epithelial-mesenchymal transition (EMT), and tumorigenesis, while promoted cell apoptosis in NPC cells. Consistently, knock-down of microRNA-338-5p had opposite effects on the above malignant phenotypes.
Based on the existing information from the previous publications (21, 22, 24), miRNAs often targeted the 3’ untranslated regions (3’UTRs) of their downstream target genes for degradation and inhibition, to regulate biological functions in cancer cells. In addition, multiple cancer associated genes, such as hypoxia-induced factor 1α (HIF-1α) (23), Sox 4 (33), E26 transformation specific-1 (ETS1) (34), and sphingosine kinase 2 (SphK2) (35), could be targeted by microRNA-338-5p. Therefore, we conjectured that microRNA-338-5p might regulate NPC progression by inhibiting its corresponding downstream cancer associated genes in a similar manner. To validate this hypothesis, the online starBase software (http://starbase.sysu.edu.cn/) was used, and we predicted that microRNA-338-5p potentially bound to the 3’ UTR of WNT2B mRNA. Furthermore, we validated that microRNA-338-5p negatively regulated WNT2B expressions in NPC cells at both transcriptional and translated levels. Interestingly, data from previous literature indicated that WNT2B promoted the development of multiple cancers, including NPC (25, 26), cervical cancer (36), ovarian cancer (37), and so on, indicating that WNT2B acted as an oncogene and exerted opposite effects with microRNA-338-5p in regulating cancer progression. Finally, our data showed that the inhibiting effects of microRNA-338-5p overexpression on NPC development were abrogated by upregulating WNT2B, suggesting that microRNA-338-5p targeted WNT2B to suppress cancer progression in NPC.
Conclusions
Collectively, the present study first identified a novel microRNA-338-5p/WNT2B axis that regulated the development of NPC. Mechanistically, microRNA-338-5p targeted the 3’UTR of WNT2B for degradation, resulting in the inhibiting effects on NPC progression in vitro and in vivo.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
Ethics Statement
The studies involving human participants were reviewed and approved by the Ethics Committee of Wuwei People’s Hospital. The patients/participants provided their written informed consent to participate in this study. The animal study was reviewed and approved by the Ethics Committee of Wuwei People’s Hospital.
Author Contributions
SW: Conception, investigations, resources, manuscript drafting, data collection, and analysis. TY: Technical support, data visualization, and manuscript proofreading. ZH: Conception, guidance, funding acquisition, and manuscript review and submission. All authors contributed to the article and approved the submitted version.
Funding
This work was financially supported by the Basic Science Foundation of Gansu Province (Grant No. 2016DE56398210), and the funding body supported the present study in terms of resources, data collection/analysis, and manuscript drafting and publication.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.684462/full#supplementary-material
Abbreviations
3’UTRs, 3’ untranslated regions; WNT2B, Wnt family member 2B; NPC, Nasopharyngeal carcinoma; miRNAs, MicroRNAs; circRNAs, Circular RNAs; LncRNAs, Long non-coding RNAs; HIF-1α, Hypoxia-induced factor 1α; ATCC, American Type Culture Collection; FBS, Fetal bovine serum; ECL, Electrochemiluminescence; CCK-8, Cell-counting kit-8; OD, Optical density; FCM, Flow cytometry; PI, Propidium iodide; HNSC, Head and neck squamous cell carcinoma; IHC, Immunohistochemistry.
References
1. Liu G, Zeng X, Wu B, Zhao J, Pan Y. Rna-Seq Analysis of Peripheral Blood Mononuclear Cells Reveals Unique Transcriptional Signatures Associated With Radiotherapy Response of Nasopharyngeal Carcinoma and Prognosis of Head and Neck Cancer. Cancer Biol Ther (2020) 21(2):139–46. doi: 10.1080/15384047.2019.1670521
2. Luo WJ, Feng YF, Guo R, Tang LL, Chen L, Zhou GQ, et al. Patterns of EBV-Positive Cervical Lymph Node Involvement in Head and Neck Cancer and Implications for the Management of Nasopharyngeal Carcinoma T0 Classification. Oral Oncol (2019) 91:7–12. doi: 10.1016/j.oraloncology.2019.01.012
3. Yang JH, Lin LK, Zhang S. Effects of DACT1 Methylation Status on Invasion and Metastasis of Nasopharyngeal Carcinoma. Biol Res (2019) 52(1):31. doi: 10.1186/s40659-019-0238-3
4. Yang JH, Lin LK, Zhang S. Epigenetic Silencing of microRNA-335 Contributes to Nasopharyngeal Carcinoma Metastasis. Am J Otolaryngol (2020) 41(1):102302. doi: 10.1016/j.amjoto.2019.102302
5. Li LN, Xiao T, Yi HM, Zheng Z, Qu JQ, Huang W, et al. Retraction: MiR-125b Increases Nasopharyngeal Carcinoma Radioresistance by Targeting A20/Nf-κb Signaling Pathway. Mol Cancer Ther (2018) 17(11):2490. doi: 10.1158/1535-7163.MCT-18-0938
6. Liao L, Yan WJ, Tian CM, Li MY, Tian YQ, Zeng GQ. Knockdown of Annexin A1 Enhances Radioresistance and Inhibits Apoptosis in Nasopharyngeal Carcinoma. Technol Cancer Res Treat (2018) 17:1533034617750309. doi: 10.1177/1533034617750309
7. Zhang J, Xie T, Zhong X, Jiang HL, Li R, Wang BY, et al. Melatonin Reverses Nasopharyngeal Carcinoma Cisplatin Chemoresistance by Inhibiting the Wnt/β-Catenin Signaling Pathway. Aging (Albany NY) (2020) 12(6):5423–38. doi: 10.18632/aging.102968
8. Zhu Y, Hu Q, Li H. Isoprenylcysteine Carboxylmethyltransferase Is Associated With Nasopharyngeal Carcinoma Chemoresistance and Ras Activation. Biochem Biophys Res Commun (2019) 516(3):784–9. doi: 10.1016/j.bbrc.2019.06.074
9. Mo Y, Wang Y, Xiong F, Ge X, Li Z, Li X, et al. Proteomic Analysis of the Molecular Mechanism of Lovastatin Inhibiting the Growth of Nasopharyngeal Carcinoma Cells. J Cancer (2019) 10(10):2342–9. doi: 10.7150/jca.30454
10. Ouyang Y, Jin YB, Chen XP, Zhang GY, Mao SL, Ling F, et al. STIL Is Upregulated in Nasopharyngeal Carcinoma Tissues and Promotes Nasopharyngeal Carcinoma Proliferation, Migration and Invasion. Neoplasma (2020) 67(1):37–45. doi: 10.4149/neo_2019_190306N192
11. Gao Q, Tang L, Wu L, Li K, Wang H, Li W, et al. LASP1 Promotes Nasopharyngeal Carcinoma Progression Through Negatively Regulation of the Tumor Suppressor PTEN. Cell Death Dis (2018) 9(3):393. doi: 10.1038/s41419-018-0443-y
12. He Y, Wang Z, Liu C, Gong Z, Li Y, Lu T, et al. Protocadherin 17 Is a Tumor Suppressor and Is Frequently Methylated in Nasopharyngeal Carcinoma. Cancer Manag Res (2019) 11:1601–13. doi: 10.2147/CMAR.S191102
13. Qian W, Ren Z, Lu X. Knockdown of Long Non-Coding RNA TUG1 Suppresses Nasopharyngeal Carcinoma Progression by Inhibiting Epithelial-Mesenchymal Transition (EMT) Via the Promotion of Mir-384. Biochem Biophys Res Commun (2019) 509(1):56–63. doi: 10.1016/j.bbrc.2018.12.011
14. Zhang W, Du M, Wang T, Chen W, Wu J, Li Q, et al. Long Non-Coding RNA LINC01133 Mediates Nasopharyngeal Carcinoma Tumorigenesis by Binding to YBX1. Am J Cancer Res (2019) 9(4):779–90.
15. Wei H, Liu D, Sun J, Mao Y, Zhao L, Zhu W, et al. Circular RNA Circ_0008450 Upregulates CXCL9 Expression by Targeting miR-577 to Regulate Cell Proliferation and Invasion in Nasopharyngeal Carcinoma. Exp Mol Pathol (2019) 110:104288. doi: 10.1016/j.yexmp.2019.104288
16. Zhu L, Liu Y, Yang Y, Mao XM, Yin ZD. Circrna ZNF609 Promotes Growth and Metastasis of Nasopharyngeal Carcinoma by Competing With microRNA-150-5p. Eur Rev Med Pharmacol Sci (2019) 23(7):2817–26. doi: 10.26355/eurrev_201904_17558
17. He H, Liao X, Yang Q, Liu Y, Peng Y, Zhong H, et al. Microrna-494-3p Promotes Cell Growth, Migration, and Invasion of Nasopharyngeal Carcinoma by Targeting Sox7. Technol Cancer Res Treat (2018) 17:1533033818809993. doi: 10.1177/1533033818809993
18. Zhang Y, Zhao Y, Liu L, Su H, Dong D, Wang J, et al. Microrna-19b Promotes Nasopharyngeal Carcinoma More Sensitive to Cisplatin by Suppressing Kras. Technol Cancer Res Treat (2018) 17:1533033818793652. doi: 10.1177/1533033818793652
19. Wu SH, Han L, Lu BC, Wang HY, Zheng CP. MiR-99a Inhibits Cell Proliferation of Nasopharyngeal Carcinoma by Targeting mTOR and Serves as a Prognostic Factor. Eur Rev Med Pharmacol Sci (2019) 23(5):2053–61. doi: 10.26355/eurrev_201903_17246
20. Wu S, Xie DL, Dai XY. Down-Regulation of miR-155 Promotes Apoptosis of Nasopharyngeal Carcinoma CNE-1 Cells by Targeting PI3K/AKT-FOXO3a Signaling. Eur Rev Med Pharmacol Sci (2019) 23(17):7391–8. doi: 10.26355/eurrev_201909_18847
21. Chu CA, Lee CT, Lee JC, Wang YW, Huang CT, Lan SH, et al. MiR-338-5p Promotes Metastasis of Colorectal Cancer by Inhibition of Phosphatidylinositol 3-Kinase, Catalytic Subunit Type 3-Mediated Autophagy Pathway. EBioMedicine (2019) 43:270–81. doi: 10.1016/j.ebiom.2019.04.010
22. Lin WC, Chen LH, Hsieh YC, Yang PW, Lai LC, Chuang EY, et al. miR-338-5p Inhibits Cell Proliferation, Colony Formation, Migration and Cisplatin Resistance in Esophageal Squamous Cancer Cells by Targeting FERMT2. Carcinogenesis (2019) 40(7):883–92. doi: 10.1093/carcin/bgy189
23. Shan Y, Li X, You B, Shi S, Zhang Q, You Y. MicroRNA-338 Inhibits Migration and Proliferation by Targeting Hypoxia-Induced Factor 1α in Nasopharyngeal Carcinoma. Oncol Rep (2015) 34(4):1943–52. doi: 10.3892/or.2015.4195
24. Yu N, Yong S, Kim HK, Choi YL, Jung Y, Kim D, et al. Identification of Tumor Suppressor miRNAs by Integrative miRNA and mRNA Sequencing of Matched Tumor-Normal Samples in Lung Adenocarcinoma. Mol Oncol (2019) 13(6):1356–68. doi: 10.1002/1878-0261.12478
25. Liu C, Li G, Ren S, Su Z, Wang Y, Tian Y, et al. miR-185-3p Regulates the Invasion and Metastasis of Nasopharyngeal Carcinoma by Targeting WNT2B In Vitro. Oncol Lett (2017) 13(4):2631–6. doi: 10.3892/ol.2017.5778
26. Liu C, Li G, Yang N, Su Z, Zhang S, Deng T, et al. miR-324-3p Suppresses Migration and Invasion by Targeting WNT2B in Nasopharyngeal Carcinoma. Cancer Cell Int (2017) 17:2. doi: 10.1186/s12935-016-0372-8
27. Ke Z, Xie F, Zheng C, Chen D. CircHIPK3 Promotes Proliferation and Invasion in Nasopharyngeal Carcinoma by Abrogating Mir-4288-Induced ELF3 Inhibition. J Cell Physiol (2019) 234(2):1699–706. doi: 10.1002/jcp.27041
28. Wang B, Sun L, Li J, Jiang R. miR-577 Suppresses Cell Proliferation and Epithelial-Mesenchymal Transition by Regulating the WNT2B Mediated Wnt/β-Catenin Pathway in Non-Small Cell Lung Cancer. Mol Med Rep (2018) 18(3):2753–61. doi: 10.3892/mmr.2018.9279
29. Hong W, Xue M, Jiang J, Zhang Y, Gao X. Circular RNA circ-CPA4/let-7 miRNA/PD-L1 Axis Regulates Cell Growth, Stemness, Drug Resistance and Immune Evasion in Non-Small Cell Lung Cancer (NSCLC). J Exp Clin Cancer Res (2020) 39(1):149. doi: 10.1186/s13046-020-01648-1
30. Baker E, Whiteoak N, Hall L, France J, Wilson D, Bhaskar P. Mammaglobin-a, VEGFR3, and Ki67 in Human Breast Cancer Pathology and Five Year Survival. Breast Cancer (Auckl) (2019) 13:1178223419858957. doi: 10.1177/1178223419858957
31. Leung SCY, Nielsen TO, Zabaglo LA, Arun I, Badve SS, Bane AL, et al. Analytical Validation of a Standardised Scoring Protocol for Ki67 Immunohistochemistry on Breast Cancer Excision Whole Sections: An International Multicentre Collaboration. Histopathology (2019) 75(2):225–35. doi: 10.1111/his.13880
32. Rimm DL, Leung SCY, McShane LM, Bai Y, Bane AL, Bartlett JMS, et al. An International Multicenter Study to Evaluate Reproducibility of Automated Scoring for Assessment of Ki67 in Breast Cancer. Mod Pathol (2019) 32(1):59–69. doi: 10.1038/s41379-018-0109-4
33. Li Y, Chen P, Zu L, Liu B, Wang M, Zhou Q. MicroRNA-338-3p Suppresses Metastasis of Lung Cancer Cells by Targeting the EMT Regulator Sox4. Am J Cancer Res (2016) 6(2):127–40.
34. Zhang L, Yan R, Zhang SN, Zhang HZ, Ruan XJ, Cao Z, et al. MicroRNA-338-3p Inhibits the Progression of Bladder Cancer Through Regulating ETS1 Expression. Eur Rev Med Pharmacol Sci (2019) 23(5):1986–95. doi: 10.26355/eurrev_201903_17237
35. Xiao G, Wang Q, Li B, Wu X, Liao H, Ren Y, et al. Microrna-338-3p Suppresses Proliferation of Human Liver Cancer Cells by Targeting Sphk2. Oncol Res (2018) 26(8):1183–9. doi: 10.3727/096504018X15151495109394
36. Li Q, Yu X, Yang L. MiR-145 Inhibits Cervical Cancer Progression and Metastasis by Targeting WNT2B by Wnt/β-Catenin Pathway. Int J Clin Exp Pathol (2019) 12(10):3740–51.
Keywords: microRNAs, nasopharyngeal carcinoma, microRNA-338-5p, malignant phenotypes, Wnt family member 2B
Citation: Wang S, Yang T and He Z (2021) Investigations on the Role of the MicroRNA-338-5p/Wnt Family Member 2B (WNT2B) Axis in Regulating the Pathogenesis of Nasopharyngeal Carcinoma (NPC). Front. Oncol. 11:684462. doi: 10.3389/fonc.2021.684462
Received: 23 March 2021; Accepted: 03 June 2021;
Published: 29 June 2021.
Edited by:
Alessandro Rimessi, University of Ferrara, ItalyReviewed by:
Lu Feng, Zhengzhou University, ChinaQingtao Ni, Jiangsu Taizhou People’s Hospital, China
Copyright © 2021 Wang, Yang and He. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhengxiang He, YmFjaGVsaGVAMTYzLmNvbQ==
†These authors share first authorship
 Suzhen Wang
Suzhen Wang Tianning Yang
Tianning Yang Zhengxiang He
Zhengxiang He
