- 1Department of Urology, Kyushu University, Fukuoka, Japan
- 2Department of Urology, School of Medicine, University of Occupational and Environmental Health, Kitakyushu, Japan
- 3Department of Clinical Chemistry and Laboratory Medicine, Graduate School of Medical Sciences, Kyushu University, Fukuoka, Japan
Transforming growth factor-β1 (TGF-β1) plays a dual role in cancer, acting as a tumor suppressor in the early stage of cancer development and as a tumor promoter in the later stage of cancer progression in various cancers. In this study, we investigated the association between genetic polymorphisms in TGFB1 and clinicopathological characteristics or oncological outcome in prostate cancer cases treated with androgen-deprivation therapy (ADT) according to metastasis status. Japanese male patients with hormone-sensitive prostate cancer treated with ADT from 1993 to 2005 were included in this study. Genomic DNA was obtained from whole blood samples, and genotyping of TGFB1 (rs2241716 and rs4803455) was performed by PCR-based technique. No significant association between genetic polymorphisms in TGFB1 (rs2241716 and rs4803455) and clinicopathological parameters or prognosis was observed in patients with non-metastatic disease. In patients with metastatic disease, Gleason score in CT/TT carriers (rs2241716) and CA/AA carriers (rs4803455) was unfavorable compared with CC carriers. In addition, the CT/TT alleles in rs2241716 (hazard ratio, 1.82; 95% confidence interval, 1.12–2.94; P = 0.015) and the CA/AA alleles in rs4803455 (hazard ratio, 1.75; 95% confidence interval, 1.03–2.98; P = 0.040) were associated with a higher risk of progression during ADT compared with the CC allele in patients with metastatic disease. TGFB1 genetic variations were associated with adverse characteristics and progression risk in ADT among patients with metastatic disease, but not those with non-metastatic disease, supporting a distinct role of TGF-β signaling between non-metastatic and metastatic prostate cancer.
Introduction
Although most prostate cancer cases primarily respond to androgen-deprivation therapy (ADT), most of them eventually progress to castration-resistant prostate cancer (CRPC) (1). The aberrant activation of androgen receptor (AR) signaling, despite low levels of serum androgen, has been revealed to be critical in the progression to CRPC (2). Recently, intensive up-front therapies using docetaxel or novel AR-pathway inhibitors for metastatic hormone-sensitive prostate cancer have been proven to prolong survival and become standard therapy (3–5). However, although several risk models have been developed to estimate patient prognosis, it has been difficult to precisely predict the survival (3, 4, 6, 7).
Metastasis is the critical step for cancer progression, and the major cause of cancer-related mortality (8). Epithelial-mesenchymal transition (EMT) of cancer cells, which involves morphological and functional changes, is required for cells to metastasize to distant regions (9). Transforming growth factor-β1 (TGF-β1) is a pleiotropic polypeptide that forms multimeric complexes with two type I and two type II receptors and regulates various cellular functions such as differentiation, cellular proliferation, survival, apoptosis, migration, adhesion, angiogenesis, and immune surveillance (10). TGF-β1 has been shown to play a dual role in cancer, acting as a tumor suppressor in the early stage of cancer development and as a tumor promoter in the later stage of various cancers including prostate cancer (11). TGF-β signaling also interacts with EMT as well as AR signaling in prostate cancer, which may affect the therapeutic effect of ADT (12–15). Several studies have reported an association of genetic polymorphisms in TGFB1, which encodes TGF-β1, with cancer phenotypes in prostate cancer (16–19). Together, these findings suggest that genetic polymorphisms in TGFB1 may be associated with cancer phenotypes in the early and later stages.
In this study, we investigated the association between genetic polymorphisms in TGFB1 and clinicopathological characteristics or oncological outcomes in patients with prostate cancer during ADT by cancer stage.
Patients and Methods
Patients
This study included Japanese patients with non-metastatic prostate cancer treated with primary ADT or salvage ADT for prostate-specific antigen (PSA) recurrence after definitive therapy with radical prostatectomy or radiotherapy to prostate (non-metastatic disease) as well as patients with de novo metastatic prostate cancer to distant sites treated with primary ADT (metastatic disease) at the University of Occupational and Environmental Health (Kitakyushu, Japan) and Kyushu University Hospital (Fukuoka, Japan) from 1993 to 2005, as described previously (20–22).
Clinical TNM staging was determined in accordance with the unified TNM criteria based on the results of digital rectal examination, transrectal ultrasound, magnetic resonance imaging, computed tomography, and bone scan (23). ADT was performed with surgical castration or continuous medical castration using a gonadotropin-releasing hormone agonist (goserelin acetate or leuprorelin acetate) and/or an antiandrogen agent (bicalutamide, flutamide, or chlormadinone acetate). Progressive disease was defined as an increase in serum PSA levels >2 ng/mL and a 25% increase over the nadir, the appearance of a new lesion, or the progression of one or more known lesions classified according to the Response Evaluation Criteria in Solid Tumors (24).
Written informed consent was obtained from all patients. Patients who chose not to participate in this study were excluded. This study was performed in accordance with the principles described in the Declaration of Helsinki and the Ethical Guidelines for Epidemiological Research enacted by the Japanese Government and was approved by each institutional review board.
Genotyping
Genomic DNA was extracted from whole blood samples from patients as previously described (20–22). Rs2241716 and rs4803455 were selected as representative single nucleotide polymorphisms of the TGFB1 gene as described previously (20). Minimum minor allele frequency was set as 0.05 according to the HapMap database (http://hapmap.ncbi.nlm.nih.gov/index.html). Linkage disequilibrium analysis was performed with HaploView and the minimum r2 threshold was set as 0.8. Genotyping of TGFB1 (rs2241716 and rs4803455) was performed on a CFX Connect Real-Time System (Bio-Rad, Hercules, CA, USA) with pre-designed TaqMan SNP Genotyping Assays (C_15873887_10 and C_30031638_10, respectively; Life Technologies, Carlsbad, CA, USA) and TaqMan Gene Expression Master Mix (Life Technologies), according to the manufacturers’ protocols.
Statistical Analyses
All statistical analyses were performed using JMP14 software (SAS Institute, Cary, NC, USA). Categorical and continuous data were analyzed by Pearson’s chi-square and Wilcoxon rank sum tests, respectively. Survival analyses were conducted using the Kaplan–Meier method and the log-rank test. Univariate and multivariate analyses were performed using the Cox hazard proportional model to estimate hazard ratios (HRs). The differential prognostic value of TGFB1 genotype was investigated through interaction tests. All P-values were two-sided. P-values < 0.05 were considered significant.
Results
The clinical and pathological characteristics of the 101 prostate cancer patients with non-metastatic disease and 93 prostate cancer patients with metastatic disease included in this study are listed in Tables 1 and 2. In patients with non-metastatic disease, during the median follow-up for patients alive at the date of censor of 78 months (interquartile range [IQR], 44–114 months), 27 patients (26.7%) and 18 patients (17.8%) experienced progression and any-cause mortality, respectively. In patients with metastatic disease, during the median follow-up for patients alive at the date of censor of 70 months (IQR, 33–112 months), 78 patients (93.9%) and 55 patients (59.1%) experienced progression and any-cause mortality, respectively.
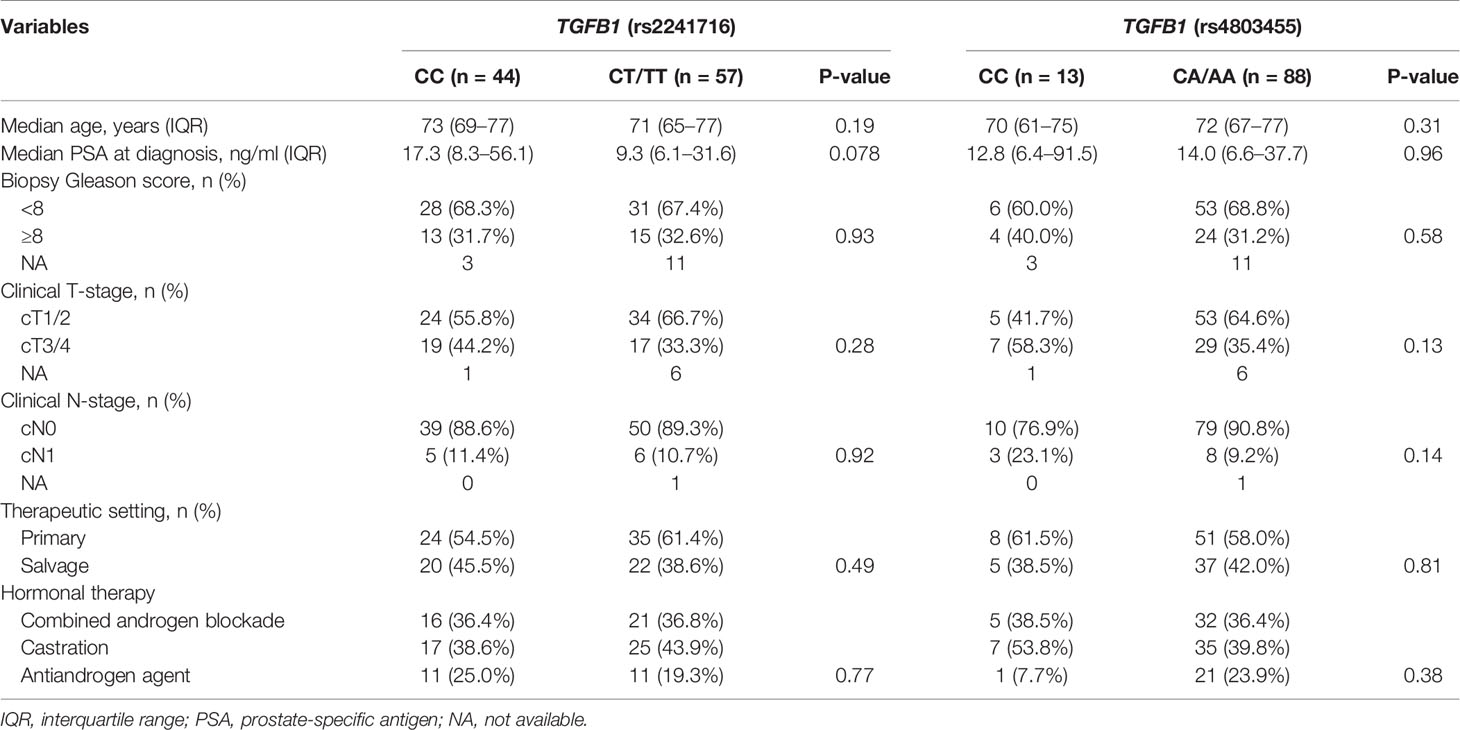
Table 1 Clinicopathological characteristics of patients with non-metastatic prostate cancer according to TGFB1 polymorphisms.
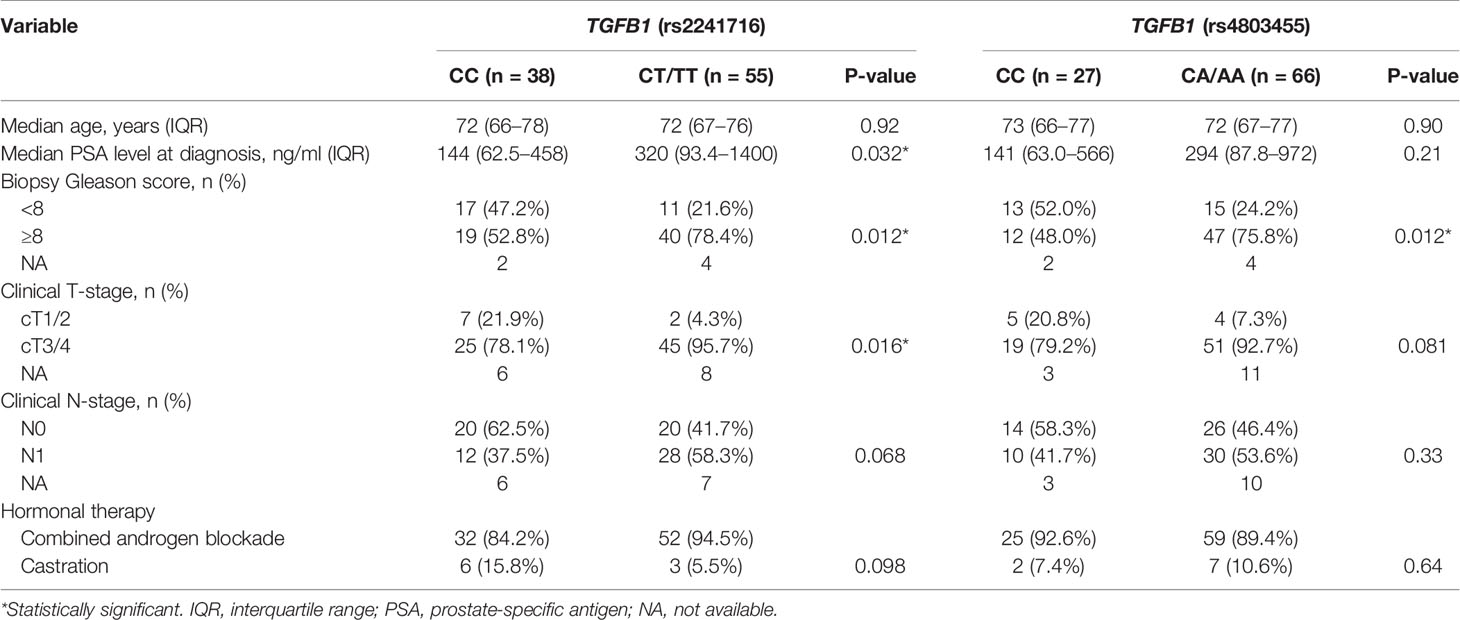
Table 2 Clinicopathological characteristics of patients with metastatic prostate cancer according to TGFB1 polymorphisms.
We analyzed the association of genetic polymorphisms in TGFB1 with clinicopathological characteristics and prognosis in patients with non-metastatic disease. Patient backgrounds were comparable in the two subgroups of TGFB1 genotypes (rs2241716 and rs4803455) in patients with non-metastatic prostate cancer (Table 1). No significant association between genetic polymorphisms in TGFB1 (rs2241716 and rs4803455) and prognosis including progression-free survival (PFS) and overall survival (OS) in patients with non-metastatic disease was observed (Table 3, Supplementary Table 1 and Figures 1A, B).
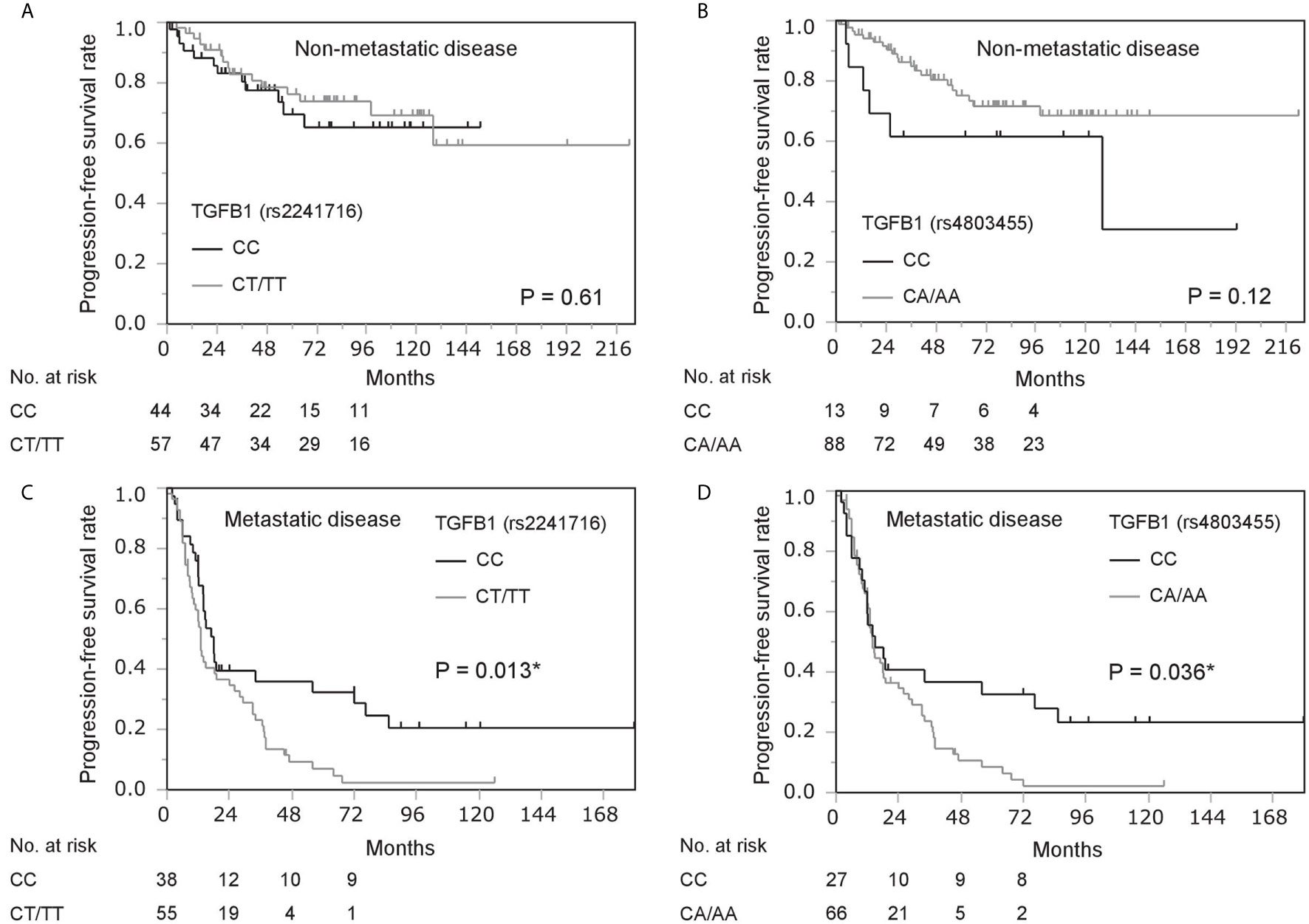
Figure 1 Kaplan–Meier survival analysis of progression-free survival in prostate cancer patients stratified by gene polymorphisms in TGFB1 (rs2241716 and rs4803455). (A, B) Progression-free survival in patients with non-metastatic disease by gene polymorphisms in TGFB1 [rs2241716 (A) and rs4803455 (B)]. (C, D) Progression-free survival in patients with metastatic disease by gene polymorphisms in TGFB1 [rs2241716 (C) and rs4803455 (D)]. *Statistically significant.
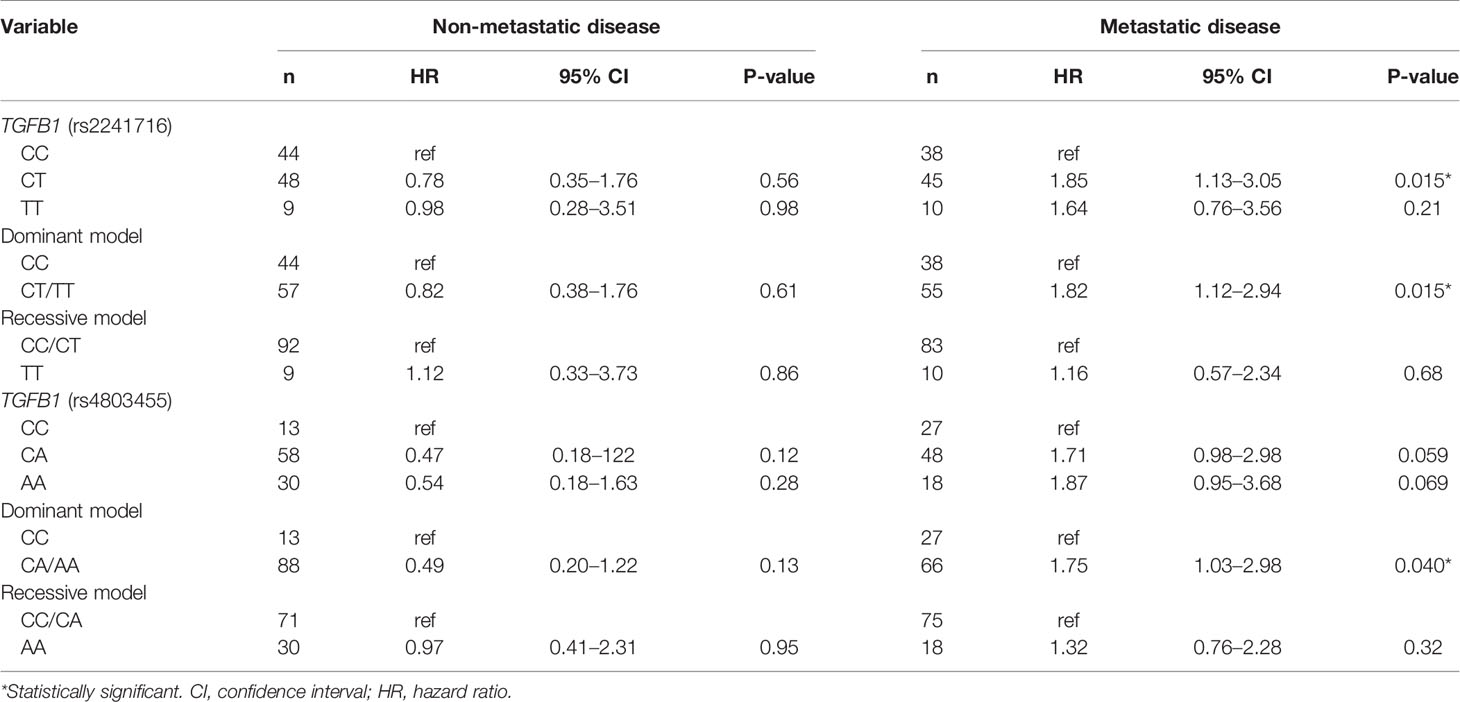
We next analyzed the significance of TGFB1 genotype among patients with metastatic prostate cancer in the same manner. Analysis of patient backgrounds revealed that PSA value at diagnosis in CT/TT carriers (rs2241716) was higher than that of CC carriers in patients with metastatic disease (Table 2). In addition, Gleason score in CT/TT carriers (rs2241716) and CA/AA carriers (rs4803455) was unfavorable compared with that in CC carriers in patients with metastatic disease (Table 2). Moreover, clinical T-stage in CT/TT carriers (rs2241716) was more advanced than that of CC carriers in patients with metastatic disease (Table 2). Consistent with these findings, the CT/TT alleles in rs2241716 (HR, 1.82; 95% confidence interval [CI], 1.12–2.94; P = 0.015) and the CA/AA alleles in rs4803455 (HR, 1.75; 95% CI, 1.03–2.98; P = 0.040) were associated with a higher risk of progression during ADT compared with that of the CC allele in patients with metastatic disease (Table 3). Similarly, Kaplan–Meier curve showed worse PFS among patients carrying the CT/TT alleles in rs2241716 and the CA/AA alleles in rs4803455 compared with patients carrying the CC allele (Figures 1C, D). However, when multivariate analyses incorporating PSA value, Gleason score, clinical T-stage for rs2241716, and Gleason score for rs4803455 were performed, the significance of the CT/TT alleles in rs2241716 (HR, 1.79; 95% CI, 0.98–3.27; P = 0.057) and the CA/AA alleles in rs2241716 (HR, 1.34; 95% CI, 0.76–2.35; P = 0.31) on PFS diminished. With regard to OS, there was no significant association between the genetic polymorphisms in TGFB1 and mortality risk in patients with metastatic disease (Supplementary Table 1).
Finally, we analyzed the impact of TGFB1 genotype on survival between patients with non-metastatic and metastatic diseases. Intriguingly, the dominant model of rs4803455 (CC vs. CA/AA; interaction test, P = 0.016) but not the dominant model of rs2241716 (CC vs. CT/TT; Interaction test, P = 0.091) was differentially associated with PFS between patients with non-metastatic and metastatic diseases. However, the significance of TGFB1 genotypes (rs2241716 and rs4803455) on patient backgrounds and OS did not differ between patients with non-metastatic and metastatic diseases (data not shown).
Discussion
This study showed that genetic polymorphisms in TGFB1 were associated with unfavorable clinicopathological parameters including PSA value, Gleason score, and clinical T-stage patients with metastatic prostate cancer. Consistent with these associations between TGFB1 variations and clinicopathological characteristics, the progression risk during ADT was associated with TGFB1 genotypes, suggesting that TGFB1 genotypes were associated with PFS through unfavorable tumor characteristics. In addition, TGFB1 variations were not associated with clinicopathological characteristics and prognosis in patients with non-metastatic disease, and a differential impact of TGFB1 variation (rs4803455) on PFS between non-metastatic and metastatic disease was observed. Since TGF-β1 has been suggested to play a dual role in the early and later stages of cancer development (11), the differential impact of TGFB1 genotype on non-metastatic and metastatic diseases may be explained by the distinct biological role of TGF-β signaling according to tumor stage.
A previous study showed that genetic variation in TGFB1 (509C>T, rs1800469) was associated with Gleason score and tumor stage in prostate cancer (17, 18). Similarly, another genetic polymorphism (TGFB1+869T>C, rs1982073) combined with a genetic polymorphism in epidermal growth factor was reported to be associated with time to CRPC (19). Similarly, it has been reported that genetic polymorphism in the promoter region of TGFBR2 gene coding TGF-βRII was associated with Gleason score and risk of early relapse after ADT among patients with both non-metastatic and metastatic prostate cancer (25). In addition, this study showed that other polymorphisms in TGFB1 (rs2241716 and rs4803455) were associated with adverse characteristics and progression risk during ADT. These results support the robustness of the association between TGFB1 genotype and tumor aggressiveness in metastatic prostate cancer, which indicates altered progression risk according to TGFB1 genotype.
The interactions of TGF-β signaling with EMT and AR signaling may be a possible molecular basis underlying the findings in this study. We previously showed that TGF-β induces AR expression including AR variants through the Twist1 transcription factor, which results in increased EMT phenotype and augmented castration resistance, which is reversed by TGF-β1 inhibitor (13, 14). Therefore, TGFB1 genotyping may be helpful to identify promising candidates for therapeutics using TGF-β inhibitors, which are under clinical trials (26, 27). As well, TGFB1 genotype could predict durable responders to primary ADT as shown by Kaplan-Meier curve on PFS (Figures 1C, D). Although the reason why durable responders carried CC genotype in TGFB1 (rs2241716 and rs4803455), it was suggested that EMT regulated by TGF signaling may play an important role in long-lasting response to ADT.
This study had several limitations. First, this study had a retrospective design. In addition, the study population was limited to Japanese patients, and intensive up-front therapies using docetaxel and novel AR pathway inhibitors were not used at the time of the study. Thus, the significance of TGFB1 variation in up-front therapies for metastatic hormone-sensitive prostate cancer should be investigated in the future. In addition, the functional effects of the genetic polymorphisms investigated in this study remain unclear. Finally, the correlation between TGFB1 variation and genetic polymorphism in TGFBR or the expression of TGF-β receptor in prostate cancer has not been investigated. Comprehensive investigation on the relationship between TGF-β signaling and ADT would be required in the future.
In conclusion, this study showed that TGFB1 genetic variations were associated with adverse characteristics and risk of progression during ADT among patients with metastatic disease, but not those with non-metastatic disease. This finding supports a distinct functional role of TGF-β signaling in non-metastatic and metastatic prostate cancer. In addition, TGFB1 genotyping may be useful to identify candidates for TGF-β signaling–targeting therapies.
Data Availability Statement
All data relevant to the study are included in the article or uploaded as supplemental information. Deidentified participant data are available upon request.
Ethics Statement
The studies involving human participants were reviewed and approved by Kyushu University Hospital and University of Occupational and Environmental Health review boards. The patients/participants provided their written informed consent to participate in this study.
Author Contributions
Conception and design: MS. Acquisition of data: MS, NF, TM, ST, SN, SU, MU, EK, AT, JI, and TU. Analysis and interpretation of data (e.g., statistical analysis): MS and NF. Writing, review, and/or revision of the manuscript: MS and NF. Study supervision: ME. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by a grant from Takeda Science Foundation.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to thank Ms. Noriko Hakoda and Ms. Eriko Gunshima for technical assistance. We thank Gabrielle White Wolf, PhD, from Edanz Group (https://en-author-services.edanzgroup.com/ac) for editing a draft of this manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.697955/full#supplementary-material
References
1. Shiota M, Eto M. Current Status of Primary Pharmacotherapy and Future Perspectives Toward Upfront Therapy for Metastatic Hormone-Sensitive Prostate Cancer. Int J Urol (2016) 23:360–9. doi: 10.1111/iju.13091
2. Shiota M, Yokomizo A, Naito S. Pro-Survival and Anti-Apoptotic Properties of Androgen Receptor Signaling by Oxidative Stress Promote Treatment Resistance in Prostate Cancer. Endocr Relat Cancer (2012) 19:R243–53. doi: 10.1530/ERC-12-0232
3. Sweeney CJ, Chen YH, Carducci M, Liu G, Jarrard DF, Eisenberger M, et al. Chemohormonal Therapy in Metastatic Hormone-Sensitive Prostate Cancer. N Engl J Med (2015) 373:737–46. doi: 10.1056/NEJMoa1503747
4. Fizazi K, Tran N, Fein L, Matsubara N, Rodriguez-Antolin A, Alekseev BY, et al. Abiraterone Plus Prednisone in Metastatic, Castration-Sensitive Prostate Cancer. N Engl J Med (2017) 377:352–60. doi: 10.1056/NEJMoa1704174
5. Sathianathen NJ, Koschel S, Thangasamy IA, Teh J, Alghazo O, Butcher G, et al. Indirect Comparisons of Efficacy Between Combination Approaches in Metastatic Hormone-Sensitive Prostate Cancer: A Systematic Review and Network Meta-Analysis. Eur Urol (2020) 77:365–72. doi: 10.1016/j.eururo.2019.09.004
6. Cooperberg MR, Hinotsu S, Namiki M, Ito K, Broering J, Carroll PR, et al. Risk Assessment Among Prostate Cancer Patients Receiving Primary Androgen Deprivation Therapy. J Clin Oncol (2009) 27:4306–13. doi: 10.1200/JCO.2008.21.5228
7. Akamatsu S, Kubota M, Uozumi R, Narita S, Takahashi M, Mitsuzuka K, et al. Development and Validation of a Novel Prognostic Model for Predicting Overall Survival in Treatment-Naïve Castration-Sensitive Metastatic Prostate Cancer. Eur Urol Oncol (2019) 2:320–8. doi: 10.1016/j.euo.2019.01.013
8. Dongre A, Weinberg RA. New Insights Into the Mechanisms of Epithelial-Mesenchymal Transition and Implications for Cancer. Nat Rev Mol Cell Biol (2019) 20:69–84. doi: 10.1038/s41580-018-0080-4
9. Thiery JP, Sleeman JP. Complex Networks Orchestrate Epithelial-Mesenchymal Transitions. Nat Rev Mol Cell Biol (2006) 7:131–42. doi: 10.1038/nrm1835
10. Elliott RL, Blobe GC. Role of Transforming Growth Factor Beta in Human Cancer. J Clin Oncol (2005) 23:2078–93. doi: 10.1200/JCO.2005.02.047
11. Morikawa M, Derynck R, Miyazono K. Tgf-β and the TGF-β Family: Context-Dependent Roles in Cell and Tissue Physiology. Cold Spring Harb Perspect Biol (2016) 8:a021873. doi: 10.1101/cshperspect.a021873
12. Fuzio P, Ditonno P, Rutigliano M, Battaglia M, Bettocchi C, Loverre A, et al. Regulation of TGF-β1 Expression by Androgen Deprivation Therapy of Prostate Cancer. Cancer Lett (2012) 318:135–44. doi: 10.1016/j.canlet.2011.08.034
13. Shiota M, Zardan A, Takeuchi A, Kumano M, Beraldi E, Naito S, et al. Clusterin Mediates TGF-β-Induced Epithelial-Mesenchymal Transition and Metastasis Via Twist1 in Prostate Cancer Cells. Cancer Res (2012) 72:5261–72. doi: 10.1158/0008-5472.CAN-12-0254
14. Shiota M, Itsumi M, Takeuchi A, Imada K, Yokomizo A, Kuruma H, et al. Crosstalk Between Epithelial-Mesenchymal Transition and Castration Resistance Mediated by Twist1/AR Signaling in Prostate Cancer. Endocr Relat Cancer (2015) 22:889–900. doi: 10.1530/ERC-15-0225
15. Pu H, Begemann DE, Kyprianou N. Aberrant TGF-β Signaling Drives Castration-Resistant Prostate Cancer in a Male Mouse Model of Prostate Tumorigenesis. Endocrinology (2017) 158:1612–22. doi: 10.1210/en.2017-00086
16. Li Z, Habuchi T, Tsuchiya N, Mitsumori K, Wang L, Ohyama C, et al. Increased Risk of Prostate Cancer and Benign Prostatic Hyperplasia Associated With Transforming Growth Factor-Beta 1 Gene Polymorphism At Codon10. Carcinogenesis (2004) 25:237–40. doi: 10.1093/carcin/bgg197
17. Ewart-Toland A, Chan JM, Yuan J, Balmain A, Ma J. A Gain of Function TGFB1 Polymorphism may be Associated With Late Stage Prostate Cancer. Cancer Epidemiol Biomarkers Prev (2004) 13:759–64.
18. Brand TC, Bermejo C, Canby-Hagino E, Troyer DA, Baillargeon J, Thompson IM, et al. Association of Polymorphisms in TGFB1 and Prostate Cancer Prognosis. J Urol (2008) 179:754–8. doi: 10.1016/j.juro.2007.09.020
19. Teixeira AL, Ribeiro R, Morais A, Lobo F, Fraga A, Pina F, et al. Combined Analysis of EGF+61G>A and TGFB1+869T>C Functional Polymorphisms in the Time to Androgen Independence and Prostate Cancer Susceptibility. Pharmacogenom J (2009) 9:341–6. doi: 10.1038/tpj.2009.20
20. Shiota M, Fujimoto N, Imada K, Yokomizo A, Itsumi M, Takeuchi A, et al. Potential Role for YB-1 in Castration-Resistant Prostate Cancer and Resistance to Enzalutamide Through the Androgen Receptor V7. J Natl Cancer Inst (2016) 108:djw005. doi: 10.1093/jnci/djw005
21. Shiota M, Fujimoto N, Itsumi M, Takeuchi A, Inokuchi J, Tatsugami K, et al. Gene Polymorphisms in Antioxidant Enzymes Correlate With the Efficacy of Androgen-Deprivation Therapy for Prostate Cancer With Implications of Oxidative Stress. Ann Oncol (2017) 28:569–75. doi: 10.1093/annonc/mdw646
22. Shiota M, Narita S, Akamatsu S, Fujimoto N, Sumiyoshi T, Fujiwara M, et al. Association of Missense Polymorphism in HSD3B1 With Outcomes Among Men With Prostate Cancer Treated With Androgen-Deprivation Therapy or Abiraterone. JAMA Netw Open (2019) 2:e190115. doi: 10.1001/jamanetworkopen.2019.0115
23. International Union Against Cancer: Urologic Tumors. In: Sobin LH, Wittekind CH, editors. Prostate Tnm Classification of Malignant Tumors. 5th. New York: John Wiley & Sons (1997) p. 170–3.
24. Scher HI, Halabi S, Tannock I, Morris M, Sternberg CN, Carducci MA, et al. Design and End Points of Clinical Trials for Patients With Progressive Prostate Cancer and Castrate Levels of Testosterone: Recommendations of the Prostate Cancer Clinical Trials Working Group. J Clin Oncol (2008) 26:1148–59. doi: 10.1200/JCO.2007.12.4487
25. Teixeira AL, Gomes M, Nogueira A, Azevedo AS, Assis J, Dias F, et al. Improvement of a Predictive Model of Castration-Resistant Prostate Cancer: Functional Genetic Variants in Tgfβ1 Signaling Pathway Modulation. PLoS One (2013) 8:e72419. doi: 10.1371/journal.pone.0072419
26. Teixeira AF, Ten Dijke P, Zhu HJ. On-Target Anti-TGF-β Therapies Are Not Succeeding in Clinical Cancer Treatments: What Are Remaining Challenges? Front Cell Dev Biol (2020) 8:605. doi: 10.3389/fcell.2020.00605
Keywords: androgen-deprivation therapy, metastasis, prostate cancer, SNP, TGFB1
Citation: Shiota M, Fujimoto N, Matsumoto T, Tsukahara S, Nagakawa S, Ueda S, Ushijima M, Kashiwagi E, Takeuchi A, Inokuchi J, Uchiumi T and Eto M (2021) Differential Impact of TGFB1 Variation by Metastatic Status in Androgen-Deprivation Therapy for Prostate Cancer. Front. Oncol. 11:697955. doi: 10.3389/fonc.2021.697955
Received: 20 April 2021; Accepted: 28 April 2021;
Published: 25 May 2021.
Edited by:
Kouji Izumi, Kanazawa University, JapanCopyright © 2021 Shiota, Fujimoto, Matsumoto, Tsukahara, Nagakawa, Ueda, Ushijima, Kashiwagi, Takeuchi, Inokuchi, Uchiumi and Eto. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Masaki Shiota, c2hpb3RhLm1hc2FraS4xMDFAbS5reXVzaHUtdS5hYy5qcA==
 Masaki Shiota
Masaki Shiota Naohiro Fujimoto2
Naohiro Fujimoto2 Shigehiro Tsukahara
Shigehiro Tsukahara Eiji Kashiwagi
Eiji Kashiwagi Ario Takeuchi
Ario Takeuchi