- 1Department of Oral Mucosal Diseases, Affiliated Stomatology Hospital of Guangzhou Medical University, Guangdong Engineering Research Center of Oral Restoration and Reconstruction, Guangzhou Key Laboratory of Basic and Applied Research of Oral Regenerative Medicine, Guangzhou, China
- 2Key Laboratory for Oral Biomedical Engineering of the Ministry of Education, Department of Oral Implantology, School and Hospital of Stomatology of Wuhan University, Wuhan, China
- 3Department of Periodontics, Affiliated Stomatology Hospital of Guangzhou Medical University, Guangdong Engineering Research Center of Oral Restoration and Reconstruction, Guangzhou Key Laboratory of Basic and Applied Research of Oral Regenerative Medicine, Guangzhou, China
- 4Department of Periodontics, Wuxi Stomatology Hospital, Wuxi, China
- 5Department of Basic Oral Medicine, Affiliated Stomatology Hospital of Guangzhou Medical University, Guangdong Engineering Research Center of Oral Restoration and Reconstruction, Guangzhou Key Laboratory of Basic and Applied Research of Oral Regenerative Medicine, Guangzhou, China
- 6Department of Pathology, School and Hospital of Stomatology of Wuhan University, Wuhan, China
Objectives: To investigate the role of hepatocyte growth factor (HGF)/c-Met signaling in oral malignant transformation.
Methods: We used immunohistochemistry to investigate HGF and c-Met expression in 53 oral squamous cell carcinoma (OSCC) specimens and 21 adjacent nontumor specimens and evaluated the associations between HGF and c-Met expression and clinicopathological parameters. Additionally, HGF-overexpression transgenic (HGF-Tg) and wild-type (Wt) mice were treated with 4-nitroquinoline-1-oxide (4NQO) to induce oral carcinogenesis for 16 weeks. At 16, 20, and 24 weeks, tongue lesions were collected for clinical observation; estimation of HGF, c-Met, and PCNA expression; apoptosis (TUNEL) assays; and RNA sequencing (RNA-seq).
Results: HGF and c-Met were positively expressed in 92.5% and 64% of OSCC samples, respectively. High HGF expression was significantly associated with smaller tumor size (p = 0.006) and inferior TNM stage (p = 0.032). No correlation between HGF and c-Met levels and other clinical parameters or prognosis was noted. In addition, HGF and c-Met expression was elevated in 4NQO-induced lesions of Wt mice. Compared with Wt mice, HGF-Tg mice have lower tumor incidence, number, volume, and lesion grade. In addition, the percentage of PCNA-positive cells in Wt mice was significantly higher than that in HGF-Tg mice at different time points. At 16 weeks, HGF-Tg mice exhibited less apoptotic cells compared with Wt mice (p < 0.000), and these levels gradually increased until the levels were greater than that of Wt mice at 24 weeks (p < 0.000). RNA-seq data revealed that 140 genes were upregulated and 137 genes were downregulated in HGF-Tg mice. KEGG enrichment analysis showed that upregulated differentially expressed genes (DEGs) are highly correlated with oxidative and metabolic signaling and that downregulated DEGs are related to MAPK and PI3K-AKT signaling.
Conclusions: HGF and c-Met expression is upregulated in OSCC tissues and is associated with the occurrence and development of OSCC. HGF overexpression in normal oral epithelial tissue can inhibit 4NQO-induced tumorigenesis potentially through inhibiting proliferation and accelerating apoptosis via MAPK and PI3K-AKT signaling.
Introduction
Oral squamous cell carcinoma (OSCC) is one of the most common malignant neoplasms with a high morbidity and mortality (1, 2). The development of OSCC is a multistep process that requires the accumulation of multiple genetic changes as a consequence of a patient’s genetic predisposition and long-term exposure to carcinogens, including tobacco, cigarette smoke, and alcohol (3). The features of aggressive invasion and high metastasis and recurrence rate lead to a poor prognosis for OSCC (4). Despite advances in current combination therapy, including surgery, radiotherapy, and chemotherapy, the 5-year survival rate remains less than 50%, and the patient’s quality of life is seriously affected (5). However, early detection and diagnosis followed by appropriate treatment can greatly improve survival. Over the past few decades, numerous studies have been devoted to the discovery of promising tumor molecular markers to help the diagnosis, classification, prognosis, and treatment of cancer.
Hepatocyte growth factor (HGF) is a multifunctional cytokine secreted primarily by mesenchymal cells that contributes to cell proliferation, motility, survival, and morphogenesis (6, 7). HGF binding to mesenchymal–epithelial transition factor (c-Met), known as its unique high-affinity receptor and expressed on epithelial cells, plays an important role in embryogenesis, organogenesis, tissue repair, and wound healing (8, 9). However, studies have confirmed that the HGF/c-Met signaling pathway is abnormally activated in various types of cancer, such as lung cancer, breast cancer, gastric cancer, and head and neck squamous cell carcinoma (HNSCC), and is involved in the oncogenesis, invasion, and angiogenesis of tumors (10, 11). Therefore, HGF/c-Met signaling has become one of the most popular targets for tumor treatment. A variety of targeted drugs targeting this signaling pathway have been developed, and most of them have entered the stage of preclinical research and clinical trials (12). Seiwert et al. showed that the proportion of HGF-positive expression was 59%, and Met overexpression was noted in greater than 80% of 97 HNSCC tissues (13). In addition, serum HGF levels in HNSCC patients were significantly greater than that in healthy people and declined after initial treatment (14). Several studies have suggested that HGF/c-Met signaling is related to OSCC progression (15, 16). In vitro, HGF promoted migration and invasion, whereas c-Met inhibitors inhibited migration and viability and promoted apoptosis in OSCC cell lines by activating AKT, ERK1/2, and NF-ĸB signaling (17).
However, recent studies have found that the HGF/c-Met pathway plays a multifunctional role in regulating physiological and pathological processes, including cancer. Finisguerra et al. demonstrated that HGF/c-Met-dependent nitric oxide released by neutrophils can kill tumor cells and protect the tissues around tumors, inhibiting tumor growth and metastasis. However, this signaling pathway can promote the development of Met-addicted tumors, indicating that the HGF/c-Met pathway plays a double-edged role in cancer (18). Moreover, Met deficiency in immune cells can promote tumor progression, and the therapeutic benefit of Met kinase inhibitors is partly weakened by the inhibition of antitumor neutrophils, which require HGF/c-Met activation. Additionally, Dong et al. recently discovered that pulp stem cells overexpressing HGF have dual effects in rheumatoid arthritis (RA). In the early stage before RA induction, HGF inhibited the progression of RA due to its immunosuppressive effect. However, in the late stage, HGF promoted synovitis by producing pathogenic interleukin-6 (IL-6), accelerated proliferation and induced apoptosis resistance (19). Therefore, the HGF/c-Met pathway may have different effects at different stages of disease development.
Previous studies explored the function and role of HGF/c-Met in OSCC mainly through in vitro cell experiments. Moreover, they observed the effects of HGF/c-Met signaling on the proliferation, metastasis, and angiogenesis of OSCC cell lines in vitro; however, only a few studies have explored the function and role of this pathway in carcinogenesis that refers to the transformation of a normal cell into a tumoral neoplastic cell. Abnormally activated HGF/c-Met signaling in OSCC can promote the development of tumor cells. However, according to the latest research hypothesis, its role in the stage of oral carcinogenesis may be different.
In this study, we aimed to explore the role of HGF/c-Met signaling in the malignant stage of the oral mucosal epithelium by establishing tongue cancer models of HGF transgenic mice (HGF-Tg) and wild-type (Wt) mice induced by chemical carcinogens (4NQO). In addition, we examined HGF and c-Met expression in 53 OSCC tissues and evaluated the correlation between HGF and c-Met expression and clinicopathological parameters.
Materials and Methods
Patients
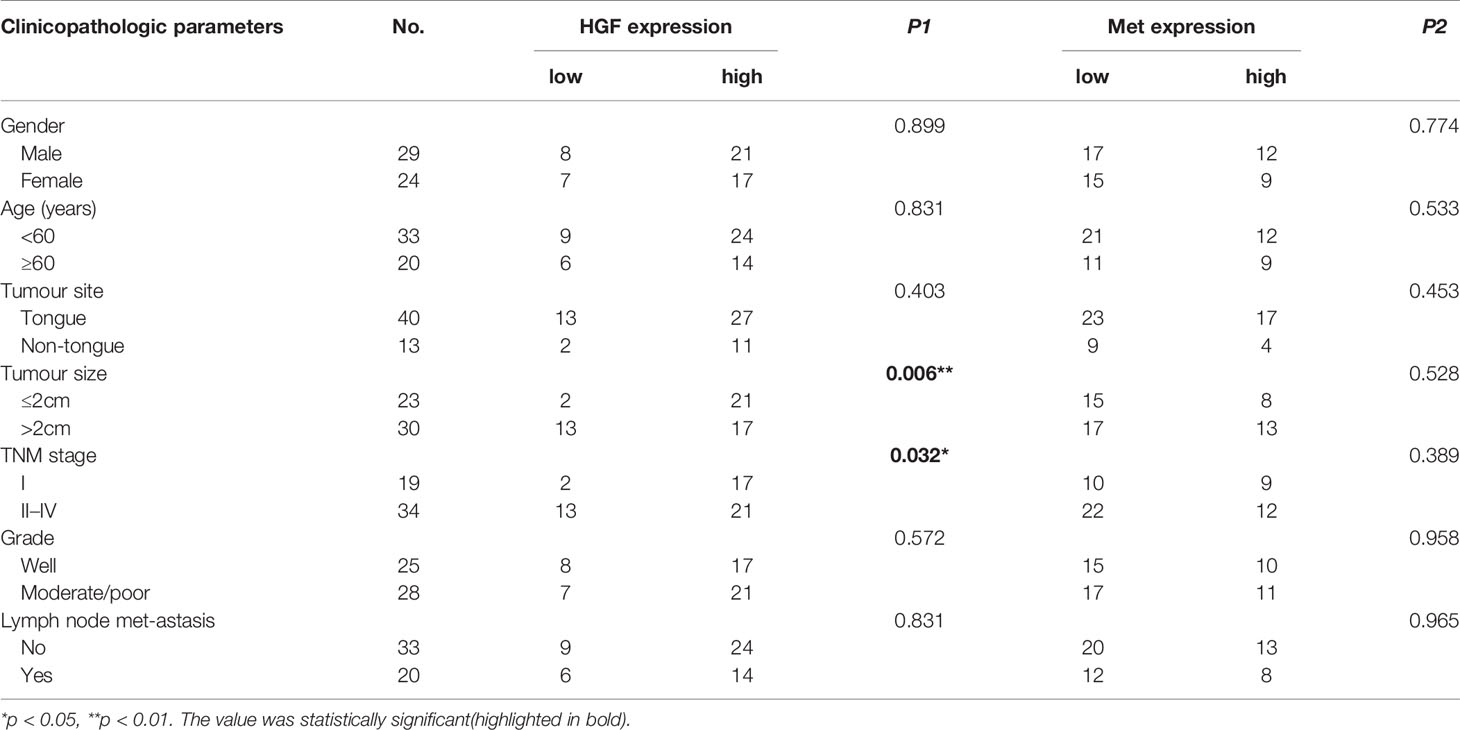
This study was approved by the human ethics research committee of Stomatological Hospital Affiliated to Guangzhou Medical University (KY2019025), and informed consent was obtained from all participants. In total, 74 paraffin-embedded tissue samples, including 53 OSCC tissues and 21 adjacent nontumor tissues from 2002 to 2006, with complete clinicopathological data of patients were provided by Stomatological Hospital Affiliated to Wuhan University. Follow-up ranged from 3 to 99 months. These patients had no other malignancies in the 5-year period before therapy, and none received treatment including chemotherapy and radiotherapy before surgery or biopsy. In addition, distant metastasis was not noted in any case. The recorded clinicopathologic parameters included sex, age, tumor size, tumor site, tumor grade, lymph node metastasis, and TNM stage. TNM staging of OSCC was determined by two pathologists according to the eighth edition of the AJCC Cancer Staging Manual. The overall survival (OS) value was from the date of first diagnosis (by biopsy or surgery) to the time of death or the last follow-up. The detailed clinical information of the patients is listed in Table 1.
Animals
This research protocol was approved by the Ethics Committee for Animal Research (No. 2016-067) in accordance with the Ethical Principal in Animal Experimentation by Guangzhou Medical University. The generation of HGF high expression transgenic mice (HGF-Tg, n = 50, female) and genotyping followed previously used protocols (20). C57BL/6 mice (Wt, n = 54, female) were purchased from Guangdong Experimental Animal Center. The experiments were performed under controlled conditions, including a 24-h light/dark cycle, and the mice were maintained in a room with a relative humidity level of 30%–50% and a temperature of 18–25°C. The mice were fed standard mouse chow.
4NQO-Induced Tumorigenesis
The carcinogen 4NQO (Sigma-Aldrich, St. Louis, MO, USA) was prepared at a concentration of 5 mg/ml in propylene glycol and stored at 4°C. The 4NQO stock solution was diluted to a concentration of 50 µg/ml in drinking water, and the water was replaced once weekly. Six- to eight-week-old HGF-Tg mice and Wt mice were randomly divided into a control group (n = 18) and a 4NQO group (n = 86), respectively. The experimental group was drinking water containing 4NQO continually for 16 weeks, whereas the control group contained no 4NQO and the same volume of propylene glycol. Mice were sacrificed after 16, 20, or 24 weeks or when the body weight loss was ≥1/3 (21). The death time of naturally dead mice was recorded throughout the experiment, and the remaining surviving mice were sacrificed at 28 weeks. All mice were carefully inspected daily and weighed weekly. The experimental process is shown in Figure 1.
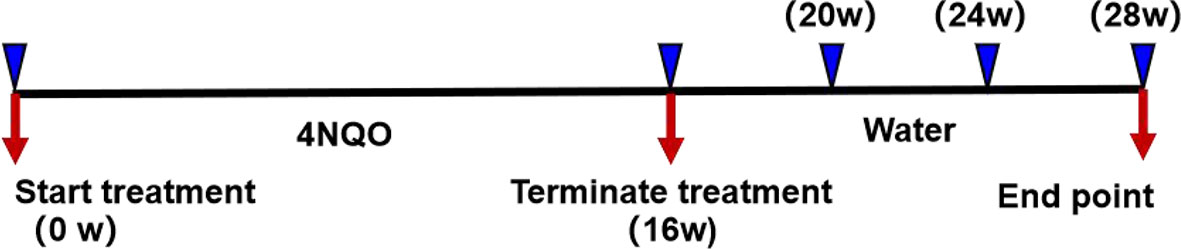
Figure 1 Representation of experimental design. After 4NQO treatment for 16 weeks, mice were sacrificed at 16, 20, 24, and 28 weeks.
Clinical Analysis
The mouse tongue was collected and photographed by stereo light microscopy. The number of tongue tumors larger than 0.5 mm was recorded. The volume of exophytic tumors was calculated using the formula 0.5 × (L × S2), where L is the longest diameter and S is the shortest diameter. The tumor volume of each mouse was the sum of all tumor volumes (22). To reduce the experimental error, photographing and recording the dimensions of the tumor were performed by Xiaoxi He.
Histopathological Analysis
The tongues were fixed in 10% formalin, embedded in paraffin, sectioned into 4-µm sections, and stained with hematoxylin and eosin (H&E). The sections were examined by two pathologists at our institution and classified as follows: (a) normal epithelia, (b) mild dysplasia, (c) moderate dysplasia, (d) severe dysplasia, and (e) oral cancer, including extraneous papillary tumors (23, 24).
Immunohistochemistry
IHC was performed using the streptavidin-peroxidase (SP) method as previously reported (25). In short, after fixation with 10% formalin for at least 24 h, the paraffin-embedded tissue was sliced into 4-µm sections. The sections were deparaffinized and rehydrated with xylene and ethanol. Antigen recovery was performed in a pressure cooker for 10 min in citrate buffer (pH = 6.0). Then, endogenous peroxidase activity was blocked with 3% H2O2 for 15 min. After blocking with 5% bovine serum albumin (BSA) for 30 min, sections was incubated overnight at 4℃ with the following antibodies: HGF for mouse paraffin sections (1:100, Bioss, #bs-1025R, Beijing, China), HGF for human paraffin sections (1:200, Proteintech, #26881-1-AP, Chicago, IL, United States), Met for mouse paraffin sections (1:100, Bioss, #bs-0668R, Beijing, China), Met for human paraffin sections (1:200, Proteintech, #25869-1-AP, Chicago, IL, United States), and PCNA (1:100, Proteintech, #24036-1-AP, Chicago, IL, United States). The tissue sections were then washed, incubated with the secondary antibody labeled with horseradish peroxidase (1:100, Beyotime, #A0208, Shanghai, China) for 1 h at room temperature and developed using DAB (SignalStain® DAB Substrate Kit, #8059, Cell Signaling, Boston, MA, USA). Finally, the nuclei were counterstained with hematoxylin. Negative controls by omitting the primary antibody and positive controls were done for all tissues.
Each section was independently evaluated by two pathologists. The average percentage of PCNA-positive cells was measured in ≥5 random fields at ×400 magnification in each section. Only distinct nuclear staining was considered positive. The images were obtained using a Nikon microscope, and Image-Pro Plus software was used to count the number of positive cells. The HGF and c-Met staining scores were multiplied by the staining intensity (0, no staining; 1, light yellow; 2, moderate yellow brown; 3, strong brown staining) and the proportion of positive cells (0, negative; 1, 1%–25%; 2, 26%–50%; 3, >50%) (17, 26). The scores ranged from 0 to 9.
TUNEL Assay
TUNEL staining was performed on 4-µm sections according to the manufacturer’s protocol (KeyGEN, #KGA704, Beijing, China). In brief, the sections were deparaffinized followed by antigen retrieval, labeled with biotin-dUTP for 30 min at room temperature, and incubated with streptavidin-HRP solution for 30 min in the dark at room temperature. The sections were then washed and developed using DAB. Finally, the nucleus was dyed with hematoxylin. A nucleus with positive expression appeared brownish yellow, and a nucleus with negative expression appeared blue. Apoptosis was evaluated by counting the proportion of TUNEL-positive cells in ≥5 images randomly selected at ×400 magnification.
RNA Sequencing
High-throughput sequencing was conducted by Origingene (Shanghai, China). In short, Trizol reagent (Invitrogen, Carlsbad, CA, USA) was used to extract total RNA from the tongue tumor samples of Wt and HGF-Tg mice at 24 weeks (n = 4). The RNA amount and purity were detected using a NanoDrop spectrophotometer (NanoDrop 2000, Wilmington, DE, United States), and the RNA concentration was determined using a Qubit. The RNA integrity was assessed by Agilent 2100 with RIN number >7.0. RNA-Seq libraries were prepared using a TruSeq™ RNA sample preparation Kit (Illumina) and sequenced using an Illumina HiSeq X-Ten (LC Bio, China) following the vendor’s recommended protocol. Cutadapt was used to remove the reads that contained adaptor contamination, low-quality bases, and undetermined bases. Then, sequence quality was verified using FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). RNA-Seq read data were mapped to the reference genome of mouse species (GRCm38) using HISAT2 software. The mapped reads of each sample were assembled using StringTie (27). Then, the transcriptomes from all samples were merged to reconstruct a comprehensive transcriptome using Perl scripts. After the final transcriptome was generated, StringTie and Ballgown were used to estimate the expression levels of all transcripts (27, 28). Differentially expressed genes (DEGs) were examined using the R/Bioconductor package edgeR and established based on a log2-fold change and p-value (|log2(FC)| ≥ 1.00; p < 0.05). Gene set pathway analysis was performed using DAVID bioinformatics tools. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was performed by setting all the KEGG pathway genes as background genes. The enrichment p-value calculation was performed with the Fisher’s exact test.
Real-Time Quantitative PCR Analysis
Eleven DEGs involved in the PI3K-AKT and MAPK pathways were verified by Q-PCR. Total RNA was extracted using an RNA Extraction Kit (19221ES50, YEASEN, China). The quantity and quality of extracted RNA were analyzed with 1.8 to 2.0 of the OD260/280 using a spectrophotometer (Nanodrop 2000, Thermo Fisher Scientific, USA). Double-stranded cDNA was synthesized using a PrimeScript RT kit (Cat No. AG11706, ACCURATE BIOLOGY, China) according to the manufacturer’s instructions. Q-PCR was performed using SYBR Premix ExTaq™ II (Cat No. AG11718, ACCURATE BIOLOGY, China). The 2−δδCT method was used to quantify the relative expression of each mRNA, and GAPDH served as an internal reference gene. All experiments were conducted in triplicate. The primer sequences are shown in Supplementary Table 1.
Statistical Analysis
The data were analyzed using SPSS 17.0 software and GraphPad Prism 6.0. According to the IHC score of HGF and c-Met expression, OSCC patients were divided into a low group (0–4) and a high group (>4). The chi-square test was used to evaluate the significance of the association between HGF expression and clinicopathological parameters. Kaplan–Meier analysis and log-rank tests were performed to estimate cumulative survival. Statistical significance among two or more groups was tested by unpaired Student’s t-test and Mann–Whitney test. p < 0.05 was considered a significant difference.
Results
HGF and c-Met expression in OSCC Specimens
Tissue samples from 53 OSCC patients were immunohis-tochemically stained for HGF and c-Met expression. HGF protein was expressed in cytoplasm, cell membrane, and nucleus, and Met protein was mainly expressed in cytoplasm and membrane. Immunostaining for HGF and c-Met was negative or extremely weak in the normal oral mucosa but positive in 92.5% (49/53) and 64% (34/53) of OSCC samples, respectively. Compared with adjacent nontumor tissues, HGF and c-Met expression were relatively higher in tumor tissues (Figure 2A).
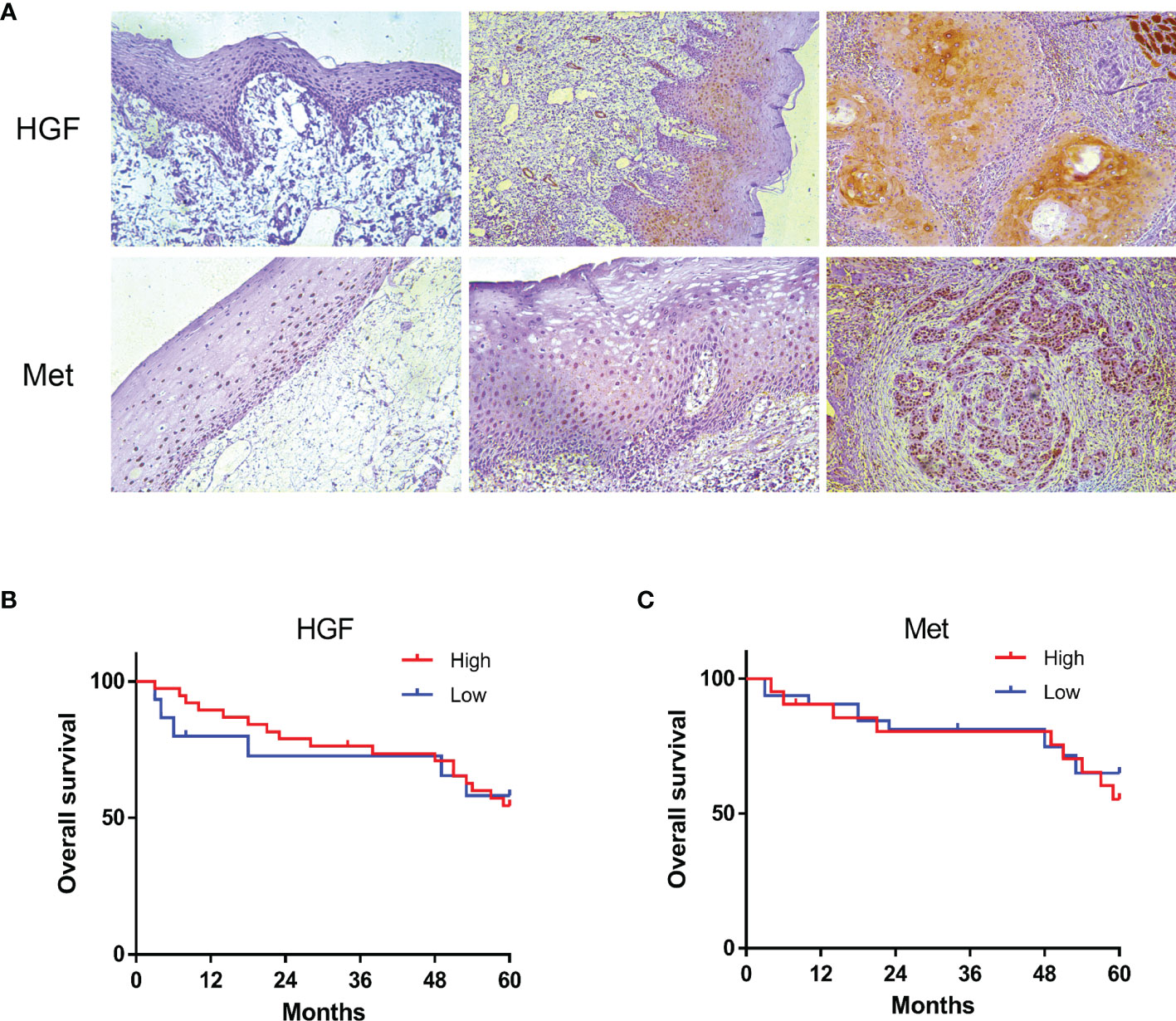
Figure 2 HGF and c-Met immunostaining in human normal oral mucosa, paracancerous tissue, and OSCC tissue. ×200 (A). The 5-year overall survival rate of OSCC patients (p > 0.05). Kaplan–Meier analysis (B, C).
To further understand the role of HGF and c-Met in OSCC progression, we analyzed the associations between tissue HGF and c-Met levels and clinicopathological features. As shown in Table 1, high HGF expression was correlated with smaller tumor size (p < 0.006) and inferior TNM stage (p < 0.032), and no significant relationship was found between HGF expression and sex, age, grade, lymph node metastasis, or any other clinicopathological parameters. However, no significant correlation was noted between c-Met expression levels and clinical parameters. In addition, HGF and c-Met expression was not significantly associated with overall survival (OS) in patients with OSCC (Figures 2B, C).
Expression of HGF and Its Receptor c-Met in Tumors Induced by 4NQO
Similar to human OSCC tissues, HGF was not expressed or was weakly expressed in normal oral mucosa of the Wt-control group but was upregulated in epithelial dysplasia and tumor tissues in the Wt-4NQO group (Figure 3A, left panels). As a unique receptor of HGF, we evaluated c-Met staining by immunohistochemistry in different groups. We found that c-Met expression was also upregulated during carcinogenesis but was slightly positively expressed in the normal tongue epithelium (Figure 3B, left panels). From 16 to 24 weeks, the epithelial lesions of 4NQO-treated mice gradually deteriorated from mild dysplasia to invasive carcinoma. In this process, we found that HGF expression was gradually weakened in the Wt-4NQO group, but the result was not statistically significant (data not shown). In addition, we did not observe a difference in c-Met expression between dysplasia and tumor tissue in Wt mice (Figure 3B, left panels).
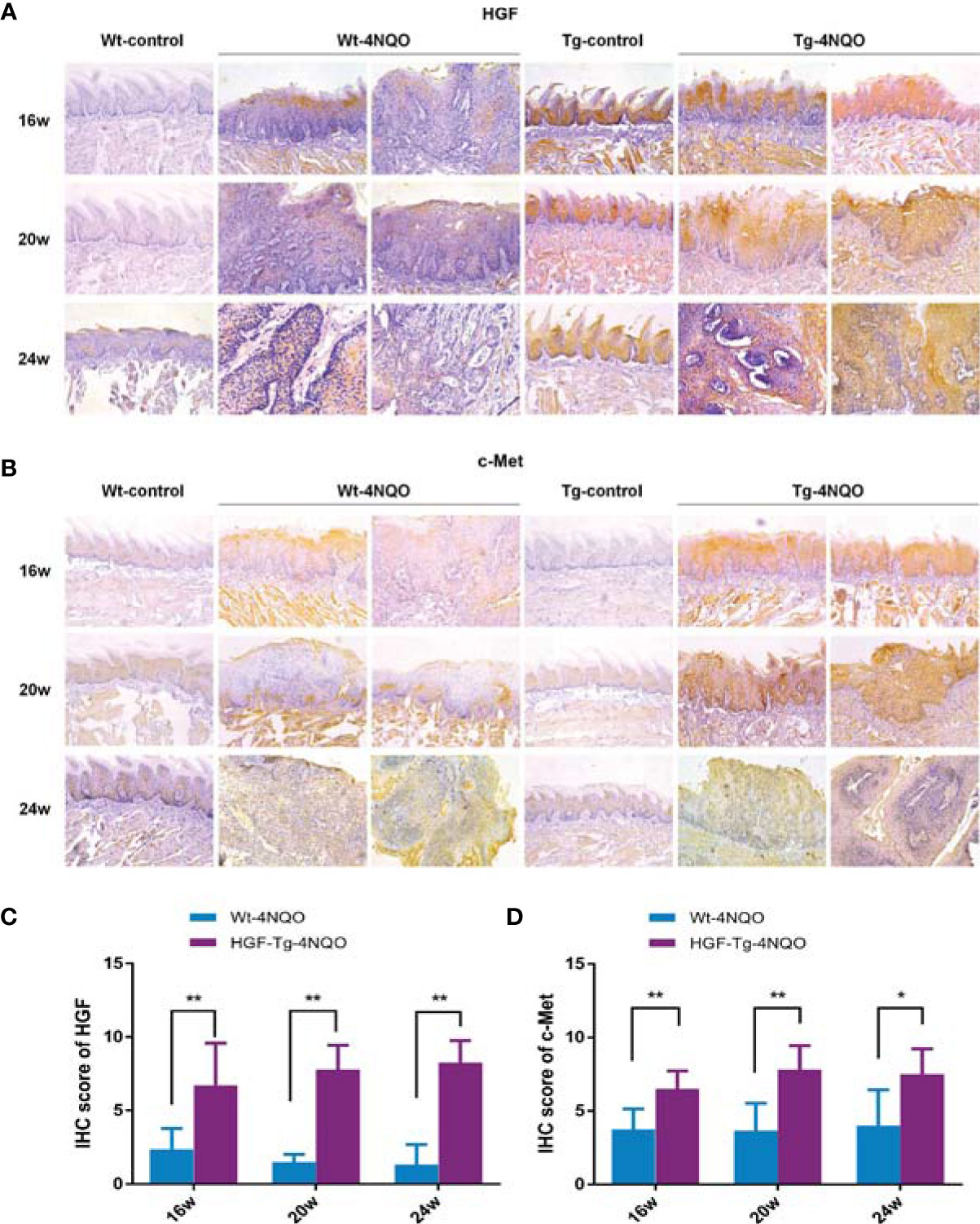
Figure 3 HGF and c-Met immunostaining in mouse tongue lesions of different groups. Representative IHC images of HGF (A) and c-Met (B).×200. IHC scores of HGF (C) and c-Met (D). 4NQO group, 16 w: n = 8; 20 w: n = 7 (Tg) and n = 8 (Wt); 24 w: n = 6 (Tg) and n = 9 (Wt). Mean ± SEM. Mann–Whitney test. *p < 0.05, **p < 0.01.
Compared with Wt mice, HGF staining was significantly stronger in normal epithelial tissue of Tg mice (Figure 3A). HGF was mainly overexpressed in normal oral epithelium of HGF-Tg mice, while it was upregulated in both epithelium and connective tissues of 4NQO-treated Tg and Wt mice (Figure 3A). However, c-Met was slightly positively expressed in the normal epithelium of both Wt and HGF-Tg mice (Figure 3B). In an oral carcinogenesis model induced by 4NQO, HGF and c-Met proteins maintained high levels compared to Wt mice from 16 to 24 weeks (Figures 3C, D).
HGF-Tg Mice Were Less Susceptible to Oral Cancer Induced by 4NQO
As shown in Figure 4A, the surface of normal tongue mucosa was pink and soft, the lingual papillae were evenly distributed, and there were no white patches or tumor formation. After drinking 4NQO water for 16 weeks, the lingual papilla disappeared regionally, and white patches of different sizes emerged. Over time, the tongue surface showed several leukoplakia and developed into papillary tumors.
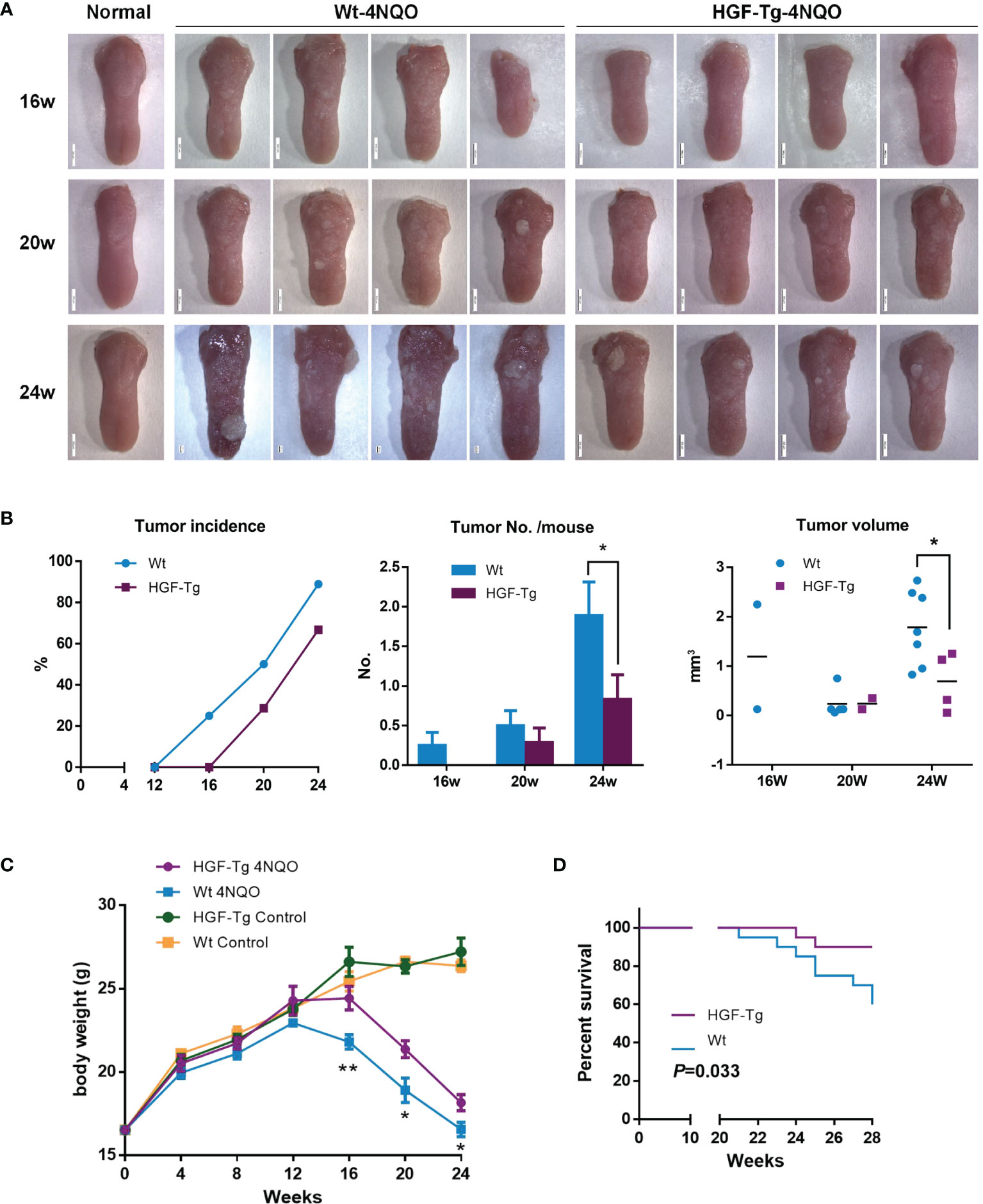
Figure 4 Induction of mouse tongue tumorigenesis by 4NQO. Representative images of mouse tongue (A). Tumor incidence, numbers, and volume in mice. Student’s t-test (B). The body weights of control and 4NQO-treated mice. Wt mice exhibited significantly reduced body weight compared with HGF-Tg mice treated with 4NQO. Student’s t-test (D). Kaplan–Meier survival analysis (E). 4NQO group, 16 w: n = 8; 20 w: n = 7 (Tg) and n = 8 (Wt); 24 w: n = 6 (Tg) and n = 9 (Wt); 28 w: n = 20. Control group, n = 3 (16/20/24 w). Mean ± SEM. *p < 0.05, **p < 0.01.
Wt mice began to develop tumors 12–16 weeks after 4NQO induction, whereas HGF-Tg mice started to develop tumors at 16–20 weeks. The incidence of tumors in Tg mice was always lower than that in Wt mice (Figure 4B, left panels). The tumor number on the tongue surface of HGF-Tg mice was significantly less than that in Wt mice (Figure 4B, middle). At week 24, most 4NQO-treated mice showed typical pathological features of tongue tumors, with the tumor volume of the HGF-Tg group being significantly smaller (Figure 4B, right panels).
Throughout the 28-week experimental period, we noticed that both HGF-Tg mice and Wt mice exposed to 4NQO began to lose weight at 12 weeks, but the weight of Wt mice decreased more seriously than HGF-Tg mice (Figure 4C). In addition, the survival rate of HGF-Tg mice was significantly greater than that of Wt mice (p < 0.033, Figure 4D).
H&E staining directly revealed dynamic histopathological changes in carcinogenesis induced by 4NQO compared with normal epithelial tissue (Figure 5A). At the 16th week, the tongue epithelium of Tg mice showed mild and moderate dysplasia, while the Wt mice had papillary tumors. At 24 weeks, invasive tumors were noted in the Wt group, but the basement membrane of the tumor in transgenic mice remained intact. The grade of dysplasia lesions and neoplasm incidence in HGF-Tg mice treated with 4NQO were significantly lower than those in Wt mice (p < 0.05, Figure 5B). Histopathological analysis further confirmed that HGF-Tg mice were less sensitive to 4NQO-induced oral tumorigenesis.
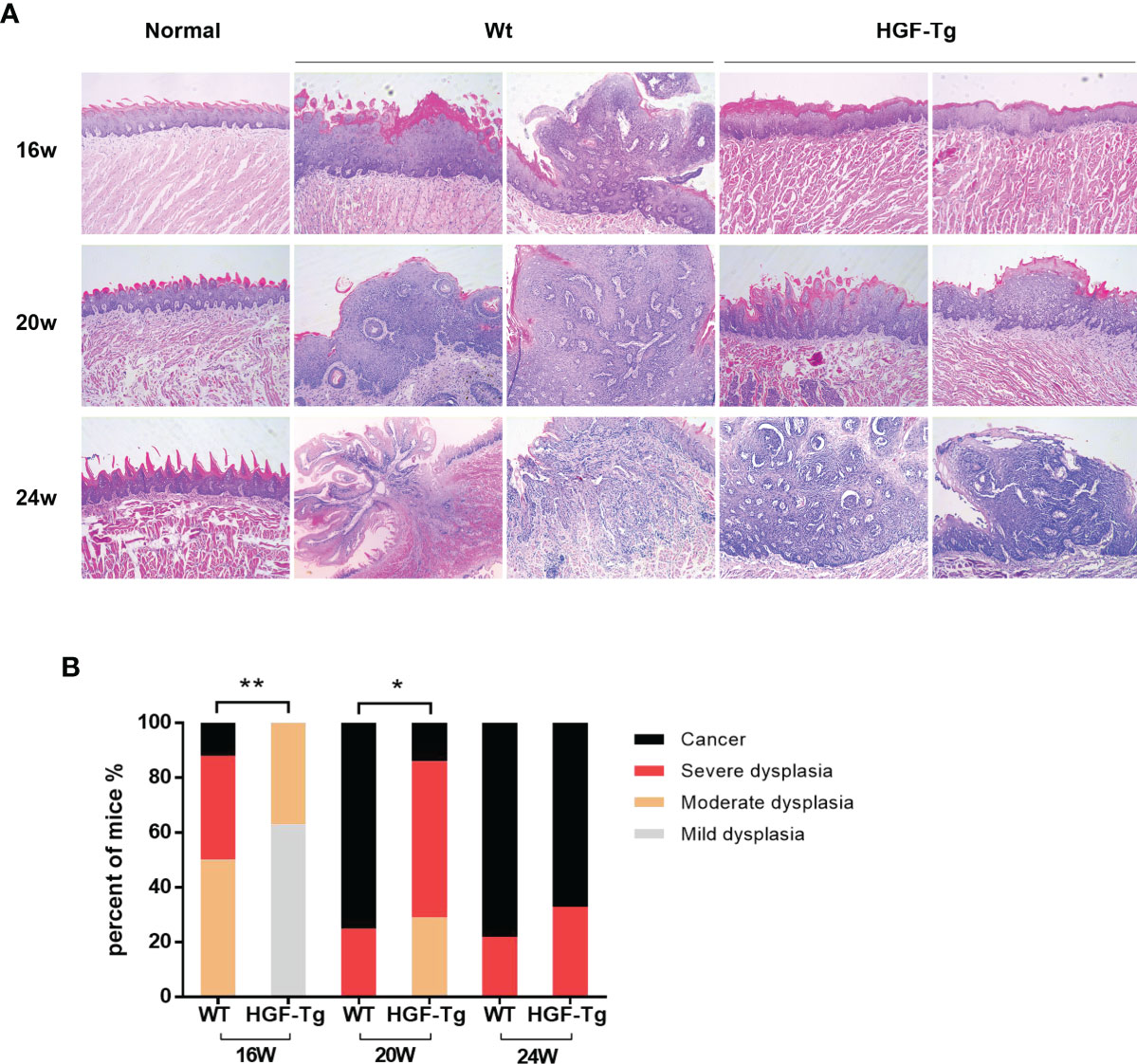
Figure 5 Representative HE sections of mice tongue, including dysplastic tissues and carcinoma. With the exception of the left image of Wt mice at 24 weeks, which was magnified at ×50, the remaining images were magnified at ×100 (A). Quantification of histopathological grading of tongue. Chi-square test (B). 4NQO group, 16 w: n = 8; 20 w: n = 7 (Tg) and n = 8 (Wt); 24 w: n = 6 (Tg) and n = 9 (Wt). *p < 0.05, **p < 0.01.
Overexpression of HGF Affects Cellular Proliferation and Apoptosis in Tumor Tissue
To further observe the pathological changes of tongue lesions of Tg mice and Wt mice, we used IHC to detect PCNA and TUNEL analysis for apoptosis (Figure 6). During 4NQO-induced carcinogenesis, PCNA-positive cells increased continuously in the process of transforming from mild dysplasia to cancerous tissue. Moreover, the proportion of proliferating cells in Wt mice was relatively greater than that in Tg mice treated with 4NQO from 16 to 24 weeks (p < 0.05) (Figures 6A, C), suggesting that HGF may inhibit cell proliferation during tumorigenesis.
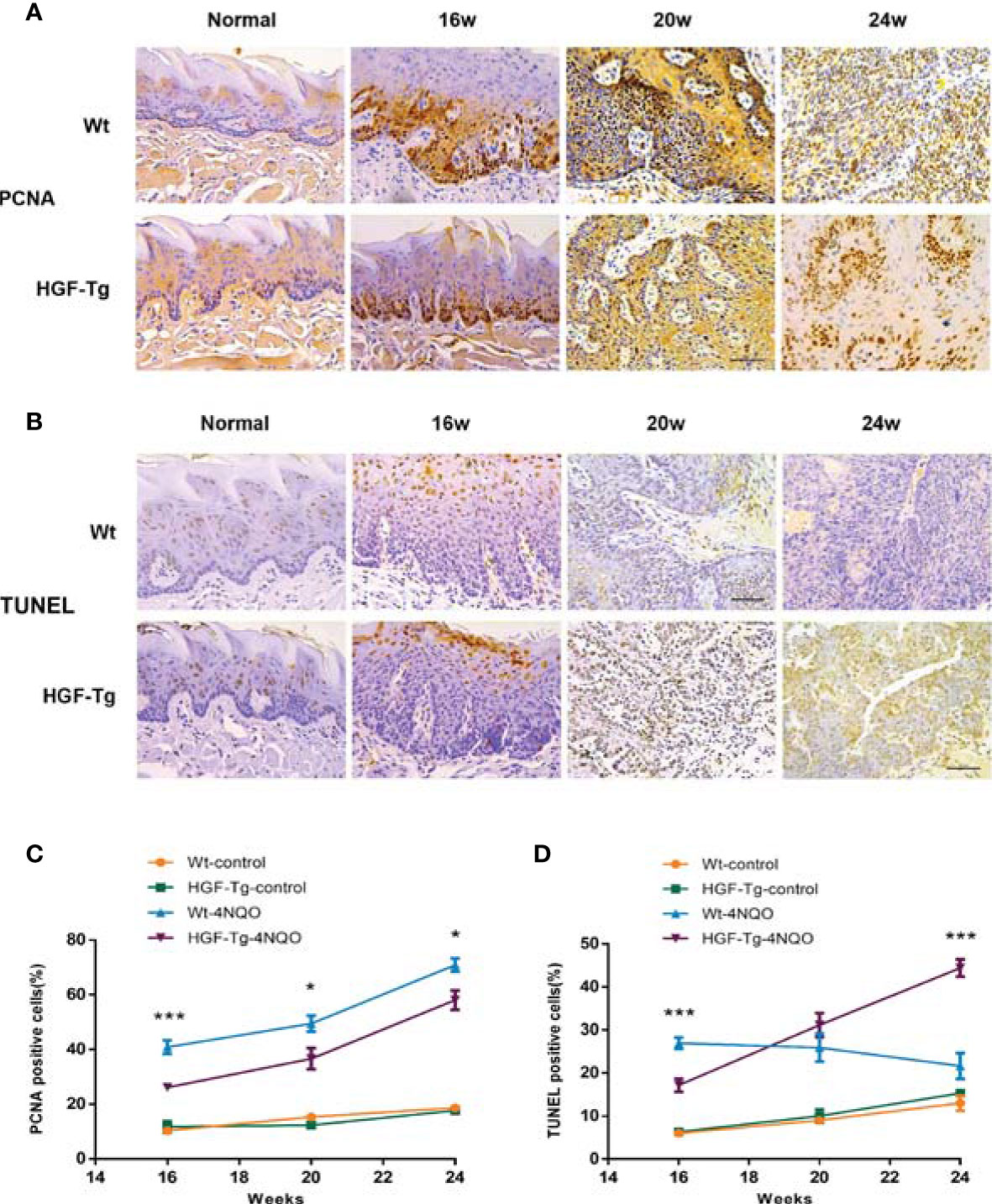
Figure 6 Proliferation and apoptosis in tongue lesions. Representative IHC PCNA staining images (A) and percentage of PCNA-positive cells (C). Representative TUNEL staining images (B) and percentage of TUNEL-positive cells (D). All pictures were obtained at ×400 magnification. 4NQO group, 16 w: n = 8; 20 w: n = 7 (Tg) and n = 8 (Wt); 24 w: n = 6 (Tg) and n = 9 (Wt). Control group, n = 3 (16/20/24 w). Mean ± SEM. Student’s t-test. *p < 0.05, ***p < 0.001.
Additionally, the number of apoptotic cells in lesions of 4NQO-treated Wt mice gradually declined from 16 to 24 weeks, whereas that in Tg mice increased during the experiment. At 16 weeks, the number of TUNEL-positive cells in the Tg-4NQO group was significantly lower than that in the Wt-4NQO group. However, at 24 weeks, the number was significantly greater than that in the Wt-4NQO group (Figures 6B, D).
Downregulation of MAPK and PI3K-AKT Signaling Potentially Affects Tumorigenesis in HGF-Tg Mice
To further explore the downstream molecular mechanism of HGF overexpression, we performed RNA-seq using tongue tumor tissues harvested from Wt and HGF-Tg mice. As indicated in the heatmap generated for DE genes (Figure 7A), 140 upregulated DEGs and 137 downregulated DEGs were identified from our RNA-seq data. To determine pathways regulated by HGF overexpression, KEGG analysis was performed, and the data are presented as a bubble plot. KEGG analysis showed that the upregulated DEGs were generally enriched in signaling pathways, such as oxidative phosphorylation (p = 0.00064); metabolic pathways (p = 0.00084); starch and sucrose metabolism (p = 0.001); and the citrate cycle (TCA cycle) (p = 0.019) (Figure 7B). The downregulated DEGs were significantly enriched in MAPK signaling (p = 0.0055) and the PI3K-AKT pathway (p = 0.018) (Figure 7C). We further analyzed the expression pattern of DEGs associated with MAPK and PI3K-Akt signaling that are most related to proliferation and apoptosis. Q-PCR results showed that the mRNA expression of DEGs (Atf2, Cacna1d, Flna, Pdgfrb, Rapgef2, Bcl2L1, and Ptk2) in HGF-Tg mice was significantly decreased compared with that in Wt mice, which was consistent with the RNA-seq data (Figure 7D). Therefore, our data suggested that inhibition of the MAPK and PI3K-AKT pathways may play a role in the progression of tongue tumor of HGF-Tg mice.
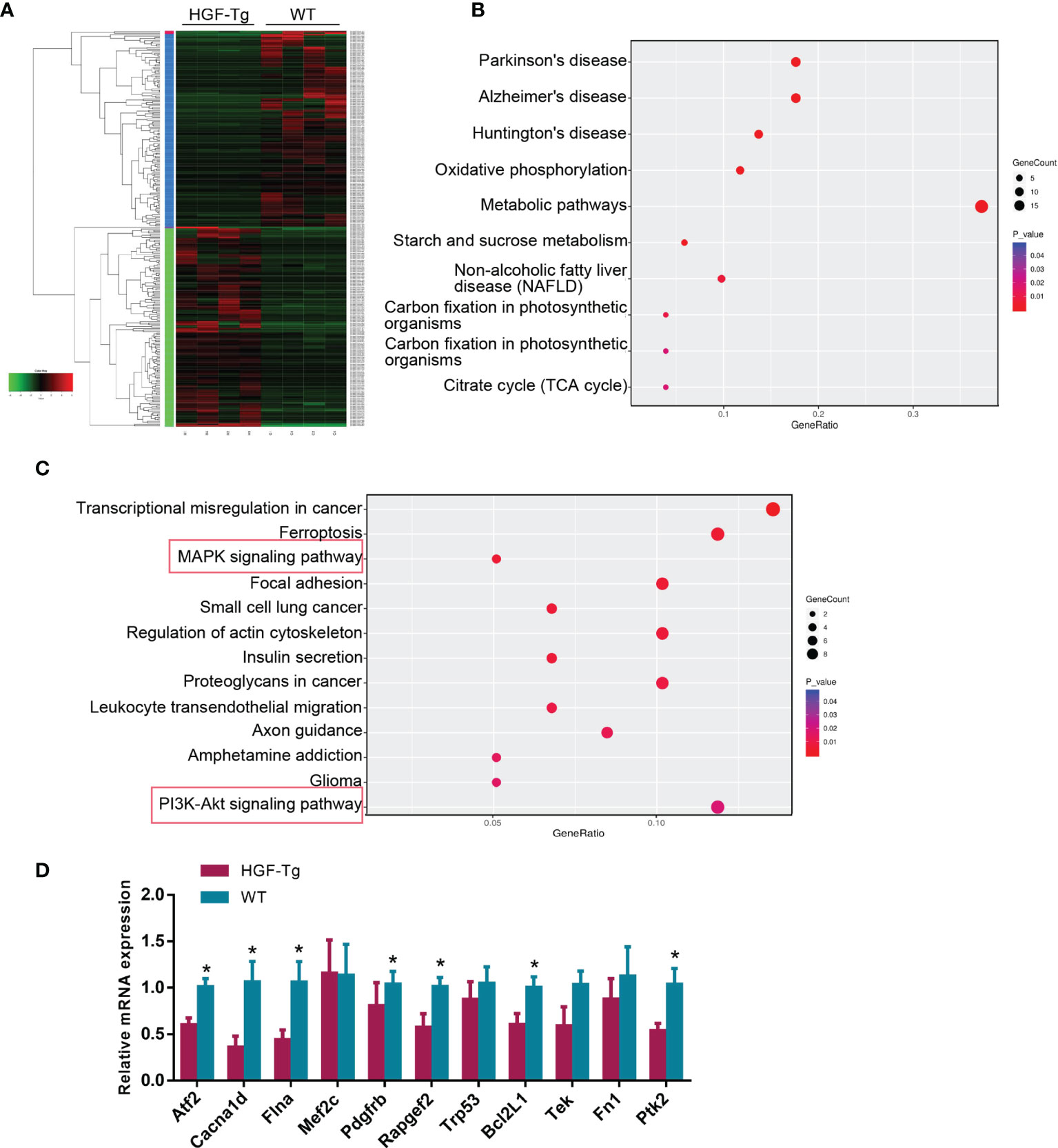
Figure 7 RNA sequencing of tongue tumor tissues of Wt and Tg mice at 24 weeks. Heatmap of 277 differentially expressed genes (A). Bubble plot represents the top 10 KEGG pathways with 140 upregulated DEGs (B) and top 13 KEGG pathways with 137 downregulated DEGs (C). Q-PCR data of DEGs enriched in the MAPK and PI3K-AKT pathway (D). Mean ± SEM. n = 4. Student’s t-test. *p < 0.05.
Discussion
It has been extensively reported that HGF and Met overexpression, gene amplification, or mutation is found in numerous tumors (12, 29). Similarly, in this study, we showed that HGF and c-Met were upregulated in OSCC and adjacent nontumor tissues. In addition, compared with normal epithelium, HGF and its receptor c-Met were positively expressed in dysplastic and tumor tissues in an oral cancer model of Wt mice. Moreover, we investigated the correlation between HGF/c-Met signaling and HNSCC, and its association with survival of HNSCC patients using data obtained from The Cancer Genome Atlas (TCGA) (Supplementary Materials). It showed that HGF expression increased in HNSCC patients with no statistical significance (Supplementary Figure A,a), but c-Met was significantly overexpressed compared with normal counterpart (Supplementary Figure B,a). The data suggested that HGF/c-Met signaling is closely correlated with the progression of OSCC.
In addition, we found that high HGF expression was associated with smaller tumor size and inferior TNM stage but was not related to other clinical parameters or OS in OSCC patients. TCGA data revealed that HGF expression was not correlated with prognosis of HNSCC patients (Supplementary Figure A,b). However, Uchida et al. reported no association between HGF expression in serum and oral cancer tissues and tumor size, lymph node status, metastasis and prognosis, although the serum levels of HGF in OSCC patients were significantly increased compared with those in healthy people (30). Chen et al. reported that only HGF expression in the tumor invasion front, not the tumor center, was correlated with lymph node status and the clinical stage of OSCC in Taiwan (31). Thus, a definite conclusion about the correlation between HGF expression level and clinical parameters and prognosis is lacking, and more clinical samples are needed.
A meta-analysis showed that c-Met overexpression in HNSCC was significantly correlated with poor OS and regional lymph node metastases (32). Another systematic review suggested that c-Met overexpression as assessed by immunohistochemistry could represent a promising prognostic biomarker of HNSCC, but the best scoring method remains to be determined (33). However, consistent with the findings of Sun et al., our study showed no significant correlation between c-Met expression in OSCC and clinicopathological variables. Moreover, according to TCGA data, there was no association between c-Met expression and OS of HNSCC patients (Supplementary Figure B,b).
In oral cancer models, HGF-Tg mice had a smaller tumor number and tumor volume and a higher survival rate than Wt mice. We observed greater HGF expression in dysplastic tissue compared with tumor tissue from the Wt-4NQO group, but the data were not statistically significant. At 16, 20, and 24 weeks, the pathological changes in the tongue in Wt mice were more serious than those in HGF-Tg mice. Based on our clinical samples and in vivo evidence, our research supports the view that HGF expression is associated with tumor size and the stage of oral cancer, and HGF overexpression can slow tongue tumor induction by 4NQO.
Intriguingly, we discovered that activation of the HGF/c-Met signaling pathway can inhibit the occurrence of oral cancer, although aberrant HGF and c-Met expression in tumor tissues is considered to be a promoting factor for the development of various types of cancer (34). In addition, Liu et al. reported that HGF can suppress hepatocarcinogenesis induced by diethylnitrosamine, indicating that HGF has different effects on the growth of normal hepatocytes and tumor hepatocytes in vivo (35). Therefore, combined with the latest relevant research results, we believe that the HGF/c-Met signaling pathway plays a two-way regulatory role in different cell types or under different conditions. Abnormal expression of the HGF/c-Met pathway in tumor tissues can promote tumor progression, whereas HGF overexpression in normal tissues can inhibit tumorigenesis.
As a multifunctional growth factor, HGF plays an important role in cell proliferation, survival, motility, and morphogenesis (36). In this study, PCNA immunostaining was used to assess the proliferation status of the lesions. The higher the cell proliferation rate, the higher the risk of cell mutation and malignant transformation (37). Findings showed that the HGF-Tg mice had fewer PCNA-positive cells than the Wt mice treated with 4NQO. Similarly, Tajima et al. demonstrated that HGF has a strong antiproliferative effect in various tumor cell lines, including HepG2 hepatocellular carcinoma cells, KB squamous carcinoma cells, and B6/F1 melanoma cells, in vitro and in vivo (38). In contrast, Knowles et al. showed that HGF promotes cell growth in HNSCC cell lines (39). Thus, we hypothesized that HGF may have multifunctional effects on cell growth in different cell types. Although the expression of PCNA is considered a useful marker in evaluating the malignant degree and progression of tumors (40), due to its long half-life (approximately 20 h), involvement in other cellular processes such as DNA repair, epitopic differences and other factors, and higher values of PCNA positivity might often be detected (41, 42); thus, there is a need to add other cell proliferation markers in subsequent studies.
During 4NQO-induced oral carcinogenesis, HGF overexpression promotes cell apoptosis based on TUNEL results. Conner et al. discovered that HGF inhibits hepatocarcinogenesis in transgenic mice and induces apoptosis in transformed rat liver epithelial cells (43). Furthermore, it has been reported that JNK1 induction, protein kinase C, and matrix metalloproteinase induction are involved in the apoptotic process induced by HGF (44). Consistent with our findings, Tulasne et al. revealed that the HGF/c-Met axis has proapoptotic and antiapoptotic properties in different cell types or stress conditions, which is important for maintaining the cell survival/apoptosis balance (45). They concluded that although HGF/c-Met signaling is involved in invasive growth and the antiapoptotic response in most tumor cell types, it can induce apoptosis and prevent malignant transformation in some cases, such as in some transformed cell lines.
4NQO is a water-soluble chemical carcinogen that causes DNA mutations and DNA double-strand breaks while increasing reactive oxygen species (ROS) production and forming large DNA adducts (46). An increase in intracellular ROS can affect cell apoptosis, survival, and differentiation via MAPK signaling consisting of ERK, JNK, and p38 (47, 48). Choi et al. demonstrated that HGF overexpression can reduce the level of intracellular ROS and inhibit H2O2-induced apoptosis (3). Similarly, previous studies have reported that HGF can protect vascular endothelial cells from oxidative damage induced by hypoxia/reoxygenation (49, 50). In addition, HGF can reduce H2O2-induced apoptosis of neural progenitor cells derived from human embryonic stem cells (51). These studies indicated that HGF regulates oxidative stress. In our study, RNA-Seq data showed that upregulated DEGs are highly correlated with oxidative and metabolic signaling.
Moreover, HGF overexpression in normal tissues can inhibit cell proliferation and accelerate apoptosis in the progression of oral cancer potentially by downregulating MAPK and PI3K-AKT according to KEGG enrichment analysis. As important intracellular signal transduction pathways, the two signaling pathways have highly similar physiological functions and jointly regulate cell growth, development, differentiation, and apoptosis (52). Although studies have shown that the HGF/c-Met pathway is abnormally activated in cancer and can promote tumor proliferation, invasion, metastasis, and angiogenesis by stimulating the PI3K/AKT, Ras/MAPK, JAK/STAT, SRC, Wnt/β-catenin, and other signaling pathways (10, 34), our study suggested that HGF/c-Met signaling may inhibit the development of tumor cells transformed from normal cells with high expression of HGF through the regulation of these two pathways, and thus provides a direction for the mechanism of our future research.
Nowadays, HGF/c-Met signaling has been recognized as a promising cancer therapeutic target, and many types of inhibitors have been developed to eliminate the activation of this pathway (53). However, the therapeutic effect of a large number of HGF/c-Met targeted drugs is not satisfactory and faces great challenges at present (54). This might be due to the signal pathway having an anticancer effect in the malignant stage of normal cells, which requires the attention of researchers.
In this study, we used HGF-overexpression transgenic mice (globally expressed) and tongue tumor models induced by 4NQO to explore the role of HGF/c-Met signaling in oral tumorigenesis. A limitation of the research is that we mainly focused on the HGF protein secreted by oral epithelial tissues, but HGF was upregulated in both epithelium and stromal tissue of HGF-Tg and Wt mice with 4NQO treatment. Since HGF is expressed in multiple cells, multiple tissues, and multiple organs in Tg mice, the function and role of HGF produced by other cells or tissues could not be confirmed, which might also affect the malignant transformation of epithelial tissues. A previous study found that HGF is commonly secreted by stromal cells and then activates the Met receptors in tumor cells by paracrine in the tumor microenvironment, while HGF autocrine by tumor cells rarely occurs (10). Therefore, the next plan was to adopt the Cre-loxP methodology to generate transgenic mice with overexpression of specific cell or tissue expression, and separately culture epithelial cells and stromal cells, such as fibroblasts, in order to study its function and mechanism more accurately.
Conclusion
In conclusion, our study found that HGF/c-Met signaling was closely associated with the occurrence and development of OSCC, and the overexpression of HGF in normal oral epithelial tissue can inhibit 4NQO-induced tumorigenesis probably by reducing cell proliferation and accelerating apoptosis via MAPK and PI3K-AKT signaling; however, its function and mechanism of action require further experiments.
Data Availability Statement
All raw RNA-sequencing data can be accessed from NCBI SRA database (SRA accession: PRJNA779172).
Ethics Statement
The studies involving human participants were reviewed and approved by the Human Ethics Research Committee of Stomatological Hospital Affiliated to Guangzhou Medical University (KY2019025). The patients/participants provided their written informed consent to participate in this study. The animal study was reviewed and approved by the Ethics Committee for Animal Research (No.2016-067) in accordance with the Ethical Principal in Animal Experimentation by Guangzhou Medical University.
Author Contributions
XW, XH, and SC contributed to the conception and design of the study. SC and XC collected OSCC specimens and patient’s clinical data. XH, YT, XZ, ZW, and WL performed the experiments. XH, YT, LY, and SC performed the statistical analysis. XH wrote the first draft of the manuscript. XH, SC, LW, and XW critically revised the manuscript. All authors contributed to the manuscript revision and read and approved the submitted version.
Funding
This work was supported by the Medical Scientific Research Foundation of Guangdong Province, China (grant numbers A2016223) and by the National Natural Science Foundation of China (grant numbers 31801152).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2021.756479/full#supplementary-material
References
1. Johnson DE, Burtness B, Leemans CR, Lui VWY, Bauman JE, Grandis JR. Head and Neck Squamous Cell Carcinoma. Nat Rev Dis Primers (2020) 6:92. doi: 10.1038/s41572-020-00224-3
2. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2018. CA A Cancer J Clin (2018) 68:7–30. doi: 10.3322/caac.21442
3. Choi S, Myers JN. Molecular Pathogenesis of Oral Squamous Cell Carcinoma: Implications for Therapy. J Dent Res (2008) 87:14. doi: 10.1177/154405910808700104
4. Vigneswaran N, Williams MD. Epidemiologic Trends in Head and Neck Cancer and Aids in Diagnosis. Oral Maxillofac Surg Clin North Am (2014) 26:123–41. doi: 10.1016/j.coms.2014.01.001
5. Thariat J, Vignot S, Lapierre A, Falk A, Guigay J, Van Obberghen-Schilling E, et al. Integrating Genomics in Head and Neck Cancer Treatment: Promises and Pitfalls. Crit Rev Oncol Hematol (2015) 95:397–406. doi: 10.1126/science.1846706
6. Bottaro D, Rubin J, Faletto D, Chan A, Kmiecik T, Vande Woude G, et al. Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Sci (New York NY) (1991) 251:802–4. doi: 10.1126/science.1846706
7. Matsumoto K, Nakamura T. Hepatocyte Growth Factor: Molecular Structure, Roles in Liver Regeneration, and Other Biological Functions. Crit Rev Oncog (1992) 3:27–54.
8. Birchmeier C, Gherardi E. Developmental Roles of HGF/SF and its Receptor, the C-Met Tyrosine Kinase. Trends Cell Biol (1998) 4:404–10. doi: 10.1016/s0962-8924(98)01359-2
9. Molnarfi N, Benkhoucha M, Funakoshi H, Nakamura T, Lalive P. Hepatocyte Growth Factor: A Regulator of Inflammation and Autoimmunity. Autoimmun Rev (2015) 14:293–303. doi: 10.1016/j.autrev.2014.11.013
10. Owusu B, Galemmo R, Janetka J, Klampfer L. Hepatocyte Growth Factor, a Key Tumor-Promoting Factor in the Tumor Microenvironment. Cancers (Basel) (2017) 9(4):35. doi: 10.3390/cancers9040035
11. Sierra J, Tsao M. C-MET as a Potential Therapeutic Target and Biomarker in Cancer. Ther Adv Med Oncol (2011) 3:S21–35. doi: 10.1177/1758834011422557
12. De Silva D, Roy A, Kato T, Cecchi F, Lee Y, Matsumoto K, et al. Targeting the Hepatocyte Growth Factor/Met Pathway in Cancer. Biochem Soc Trans (2017) 45:855–70. doi: 10.1042/bst20160132
13. Seiwert T, Jagadeeswaran R, Faoro L, Janamanchi V, Nallasura V, El Dinali M, et al. The MET Receptor Tyrosine Kinase Is a Potential Novel Therapeutic Target for Head and Neck Squamous Cell Carcinoma. Cancer Res (2009) 69:3021–31. doi: 10.1158/0008-5472.Can-08-2881
14. Kim C, Lee J, Kang S, Bae J, Hong S, Kahng H. Serum Hepatocyte Growth Factor as a Marker of Tumor Activity in Head and Neck Squamous Cell Carcinoma. Oral Oncol (2007) 43:1021–5. doi: 10.1016/j.oraloncology.2006.11.016
15. Yasui H, Ohnishi Y, Nakajima M, Nozaki M. Migration of Oral Squamous Cell Carcinoma Cells are Induced by HGF/c-Met Signalling via Lamellipodia and Filopodia Formation. Oncol Rep (2017) 37:3674–80. doi: 10.3892/or.2017.5587
16. Dong G, Lee T, Yeh N, Geoghegan J, Van Waes C, Chen Z. Metastatic Squamous Cell Carcinoma Cells That Overexpress C-Met Exhibit Enhanced Angiogenesis Factor Expression, Scattering and Metastasis in Response to Hepatocyte Growth Factor. Oncogene (2004) 23:6199–208. doi: 10.1038/sj.onc.1207851
17. Sun Z, Liu Q, Ye D, Ye K, Yang Z, Li D. Role of C-Met in the Progression of Human Oral Squamous Cell Carcinoma and Its Potential as a Therapeutic Target. Oncol Rep (2018) 39:209–16. doi: 10.3892/or.2017.6073
18. Finisguerra V, Di Conza G, Di Matteo M, Serneels J, Costa S, Thompson A, et al. MET Is Required for the Recruitment of Anti-Tumoural Neutrophils. Nature (2015) 522:349–53. doi: 10.1038/nature14407
19. Dong X, Kong F, Liu C, Dai S, Zhang Y, Xiao F, et al. Pulp Stem Cells With Hepatocyte Growth Factor Overexpression Exhibit Dual Effects in Rheumatoid Arthritis. Stem Cell Res Ther (2020) 11:229. doi: 10.1186/s13287-020-01747-y
20. Wang X, Tang Y, Shen R, Chen J, Chen G, Luo G, et al. Hepatocyte Growth Factor (HGF) Optimizes Oral Traumatic Ulcer Healing of Mice by Reducing Inflammation. Cytokine (2017) 99:275–80. doi: 10.1016/j.cyto.2017.08.006
21. Tang X, Knudsen B, Bemis D, Tickoo S, Gudas L. Oral Cavity and Esophageal Carcinogenesis Modeled in Carcinogen-Treated Mice. Clin Cancer Res an Off J Am Assoc Cancer Res (2004) 10:301–13. doi: 10.1158/1078-0432.ccr-0999-3
22. Tseng S, Yang C, Yu E, Chang C, Lee Y, Liu C, et al. K14-EGFP-miR-31 Transgenic Mice Have High Susceptibility to Chemical-Induced Squamous Cell Tumorigenesis That Is Associating With Ku80 Repression. Int J Cancer (2015) 136:1263–75. doi: 10.1002/ijc.29106
23. Al-Afifi N, Alabsi A, Kaid F, Bakri M, Ramanathan A. Prevention of Oral Carcinogenesis in Rats by Dracaena Cinnabari Resin Extracts. Clin Oral Investig (2019) 23:2287–301. doi: 10.1007/s00784-018-2685-6
24. Thompson L. World Health Organization Classification of Tumours: Pathology and Genetics of Head and Neck Tumours. Ear Nose Throat J (2006) 85:74. doi: 10.1177/014556130608500201
25. Da Silva J, Moreira Dos Santos T, Sobral L, Queiroz-Junior C, Rachid M, Proudfoot A, et al. Relevance of CCL3/CCR5 Axis in Oral Carcinogenesis. Oncotarget (2017) 8:51024–36. doi: 10.18632/oncotarget.16882
26. Xu Z, Wang S, Wu M, Zeng W, Wang X, Dong Z. Tgfβ1 and HGF Protein Secretion by Esophageal Squamous Epithelial Cells and Stromal Fibroblasts in Oesophageal Carcinogenesis. Oncol Lett (2013) 6:401–06. doi: 10.3892/ol.2013.1409
27. Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL. StringTie Enables Improved Reconstruction of a Transcriptome From RNA-Seq Reads. Nat Biotechnol (2015) 33(3):290–95. doi: 10.1038/nbt.3122
28. Frazee AC, Pertea G, Jaffe AE, Langmead B, Salzberg SL, Leek JT. Ballgown Bridges the Gap Between Transcriptome Assembly and Expression Analysis. Nat Biotechnol (2015) 33(3):243–46. doi: 10.1038/nbt.3172
29. Hartmann S, Bhola N, Grandis J. HGF/Met Signaling in Head and Neck Cancer: Impact on the Tumor Microenvironment. Clin Cancer Res an Off J Am Assoc Cancer Res (2016) 22:4005–13. doi: 10.1158/1078-0432.Ccr-16-0951
30. Uchida D, Kawamata H, Omotehara F, Nakashiro Ki, Kimura-Yanagawa T, Hino S, et al. Role of HGF/c-Met System in Invasion and Metastasis of Oral Squamous Cell Carcinoma Cells In Vitro and its Clinical Significance. Int J Cancer (2001) 93:489–96. doi: 10.1002/ijc.1368
31. Chen Y, Wang J, Chang Y, Liu B, Wang Y, Sun A, et al. Expression of Hepatocyte Growth Factor and C-Met Protein is Significantly Associated With the Progression of Oral Squamous Cell Carcinoma in Taiwan. J Oral Pathol Med Off Publ Int Assoc Oral Pathologists Am Acad Oral Pathol (2004) 33:209–17. doi: 10.1111/j.0904-2512.2004.00118.x
32. Vsiansky V, Gumulec J, Raudenska M, Masarik M. Prognostic Role of C-Met in Head and Neck Squamous Cell Cancer Tissues: A Meta-Analysis. Sci Rep (2018) 8:10370. doi: 10.1038/s41598-018-28672-8
33. Szturz P, Budíková M, Vermorken J, Horová I, Gál B, Raymond E, et al. Prognostic Value of C-MET in Head and Neck Cancer: A Systematic Review and Meta-Analysis of Aggregate Data. Oral Oncol (2017) 74:68–76. doi: 10.1016/j.oraloncology.2017.09.009
34. Zhang Y, Xia M, Jin K, Wang S, Wei H, Fan C, et al. Function of the C-Met Receptor Tyrosine Kinase in Carcinogenesis and Associated Therapeutic Opportunities. Mol Cancer (2018) 17:45. doi: 10.1186/s12943-018-0796-y
35. Liu ML, Mars WM, Michalopoulos GK. Hepatocyte Growth Factor Inhibits Cell Proliferation In Vivo of Rat Hepatocellular Carcinomas Induced by Diethylnitrosamine. Carcinogenesis (1995) 16:841–3. doi: 10.1093/carcin/16.4.841
36. Fajardo-Puerta A, Mato Prado M, Frampton A, Jiao L. Gene of the Month: HGF. J Clin Pathol (2016) 69:575–9. doi: 10.1136/jclinpath-2015-203575
37. Redder CP, Pandit S, Desai D, Kandagal VS, Ingaleshwar PS, Shetty SJ, et al. Comparative Analysis of Cell Proliferation Ratio in Plaque and Erosive Oral Lichen Planus: An Immunohistochemical Study. Dent Res J (2014) 11(3):316–20.
38. Tajima H, Matsumoto K, Nakamura T. Hepatocyte Growth Factor has Potent Anti-Proliferative Activity in Various Tumor Cell Lines. FEBS Lett (1991) 291:229–32. doi: 10.1016/0014-5793(91)81291-f
39. Knowles L, Stabile L, Egloff A, Rothstein M, Thomas S, Gubish C, et al. HGF and C-Met Participate in Paracrine Tumorigenic Pathways in Head and Neck Squamous Cell Cancer. Clin Cancer Res an Off J Am Assoc Cancer Res (2009) 15:3740–50. doi: 10.1158/1078-0432.Ccr-08-3252
40. Chiang CP, Lang MJ, Liu BY, Wang JT, Leu JS, Hahn LJ, et al. Expression of Proliferating Cell Nuclear Antigen (PCNA) in Oral Submucous Fibrosis, Oral Epithelial Hyperkeratosis and Oral Epithelial Dysplasia in Taiwan. Oral Oncol (2000) 36(4):353–59. doi: 10.1016/s1368-8375(00)00014-2
41. Steck K, el-Naggar AK. Comparative Flow Cytometric Analysis of Ki-67 and Proliferating Cell Nuclear Antigen (PCNA) in Solid Neoplasms. Cytometry (1994) 17(3):258–65. doi: 10.1002/cyto.990170309
42. Madan M, Chandra S, Raj V, Madan R. Evaluation of Cell Proliferation in Malignant and Potentially Malignant Oral Lesions. J Oral Maxillofac Pathol (2015) 19(3):297–305. doi: 10.4103/0973-029X.174613
43. Conner E, Teramoto T, Wirth P, Kiss A, Garfield S, Thorgeirsson S. HGF-Mediated Apoptosis via P53/Bax-Independent Pathway Activating JNK1. Carcinogenesis (1999) 20:583–90. doi: 10.1093/carcin/20.4.583
44. Matteucci E, Modora S, Simone M, Desiderio M. Hepatocyte Growth Factor Induces Apoptosis Through the Extrinsic Pathway in Hepatoma Cells: Favouring Role of Hypoxia-Inducible Factor-1 Deficiency. Oncogene (2003) 22:4062–73. doi: 10.1038/sj.onc.1206519
45. Tulasne D, Foveau B. The Shadow of Death on the MET Tyrosine Kinase Receptor. Cell Death Differ (2008) 15:427–34. doi: 10.1038/sj.cdd.4402229
46. Yaeno A, Chikako N, Toru T, Shigenori O, Kanehisa M, Atsushi U, et al. 4-Nitroquinoline 1-Oxide Forms 8-Hydroxydeoxyguanosine in Human Fibroblasts Through Reactive Oxygen Species. Toxicol Sci (2006) 91(2):382–92. doi: 10.1093/toxsci/kfj161
47. Bedard K, Krause K. The NOX Family of ROS-Generating NADPH Oxidases: Physiology and Pathophysiology. Physiol Rev (2007) 87:245–313. doi: 10.1152/physrev.00044.2005
48. Coso S, Harrison I, Harrison C, Vinh A, Sobey C, Drummond G, et al. NADPH Oxidases as Regulators of Tumor Angiogenesis: Current and Emerging Concepts. Antioxidants Redox Signaling (2012) 16:1229–47. doi: 10.1089/ars.2011.4489
49. Zhang Y, Hu S, Chen Y. Hepatocyte Growth Factor Suppresses Hypoxia/Reoxygenation-Induced XO Activation in Cardiac Microvascular Endothelial Cells. Heart Vessels (2015) 30:534–44. doi: 10.1007/s00380-014-0547-y
50. Zhong W, Zhao Y, Tian Y, Chen M, Lai X. The Protective Effects of HGF Against Apoptosis in Vascular Endothelial Cells Caused by Peripheral Vascular Injury. Acta Biochim Biophys Sinica (2018) 50:701–08. doi: 10.1093/abbs/gmy048
51. Hu Z, Geng J, Liang D, Luo M, Li M. Hepatocyte Growth Factor Protects Human Embryonic Stem Cell Derived-Neural Progenitors From Hydrogen Peroxide-Induced Apoptosis. Eur J Pharmacol (2010) 645:23–31. doi: 10.1016/j.ejphar.2010.07.011
52. Lu S, Jang H, Gu S, Zhang J, Nussinov R. Drugging Ras GTPase: A Comprehensive Mechanistic and Signaling Structural View. Chem Soc Rev (2016) 45:4929–52. doi: 10.1039/c5cs00911a
53. Koch J, Aebersold D, Zimmer Y, Medová M. MET Targeting: Time for a Rematch. Oncogene (2020) 39:2845–62. doi: 10.1038/s41388-020-1193-8
Keywords: OSCC, HGF, c-Met, 4NQO, transgenic mouse, tumorigenesis
Citation: He X, Chen S, Tang Y, Zhao X, Yan L, Wu L, Wu Z, Liu W, Chen X and Wang X (2021) Hepatocyte Growth Factor Overexpression Slows the Progression of 4NQO-Induced Oral Tumorigenesis. Front. Oncol. 11:756479. doi: 10.3389/fonc.2021.756479
Received: 10 August 2021; Accepted: 15 November 2021;
Published: 14 December 2021.
Edited by:
Andre Luiz Sena Guimaraes, Unimontes, BrazilReviewed by:
Tarcilia A. Silva, Federal University of Minas Gerais, BrazilSutapa Mukherjee, Chittaranjan National Cancer Institute, India
Copyright © 2021 He, Chen, Tang, Zhao, Yan, Wu, Wu, Liu, Chen and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xinhong Wang, eGluaG9uZ3dhbmcyMDIwQDE2My5jb20=
†These authors have contributed equally to this work
 Xiaoxi He1†
Xiaoxi He1† Lihong Wu
Lihong Wu Xinhong Wang
Xinhong Wang