- 1Department of Internal Medicine, Wake Forest University School of Medicine, Winston-Salem, NC, United States
- 2Department of Biostatistics and Data Science, Wake Forest University School of Medicine, Winston-Salem, NC, United States
- 3Department of Hematology and Oncology, Wake Forest University School of Medicine, Winston-Salem, NC, United States
Introduction: Metastatic colorectal cancer (mCRC) exhibits significant heterogeneity in molecular profiles, influencing treatment response and patient outcomes. Mutations in v-raf murine sarcoma viral oncogene homolog B1 (BRAF) and rat sarcoma (RAS) family genes are commonly observed in mCRC. Though originally thought to be mutually exclusive, recent data have shown that patients may present with concomitant RAS and BRAF mutations, posing unique challenges and implications for clinical management.
Methods: Below we present a retrospective study on 13 patients with concomitant BRAF and RAS (KRAS, NRAS) mutations in mCRC and describe their clinical features and treatment outcomes. We reviewed over 750 samples from a database of CRC patients from Guardant360 and FoundationOne kept by the Wake Forest Baptist Health Comprehensive Cancer Center. The study population included patients greater than the age of 18 who were diagnosed with mCRC harboring both BRAF and RAS mutations, as identified by next generation sequencing.
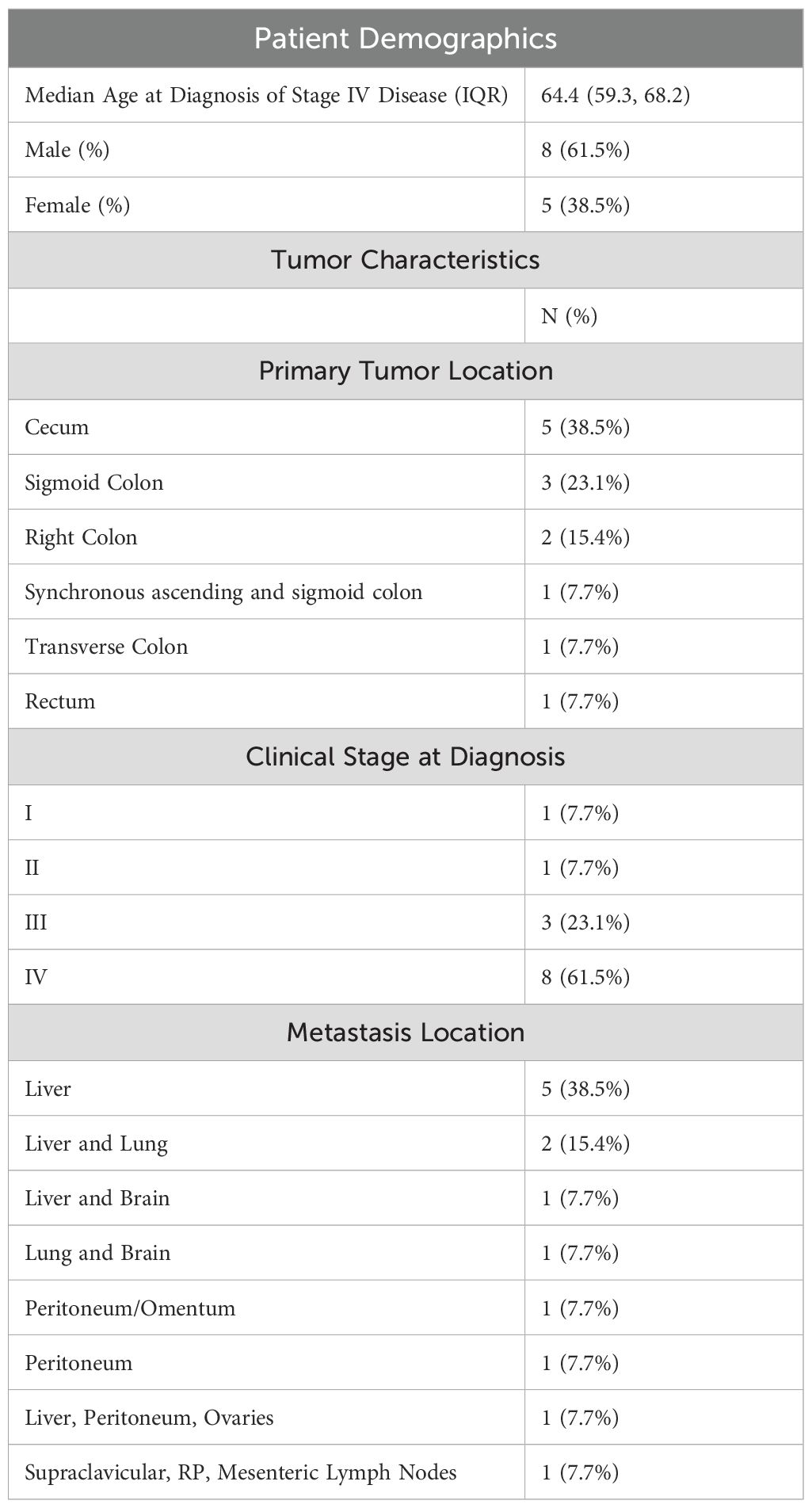
Results: Thirteen mCRC patients, 61.5% male, with a median age of diagnosis of 64.4 years had concomitant BRAF and RAS mutation. 61.5% of patients had right-sided primary disease. 61.5% patients had mutations in codon 12 of KRAS, 15.4% had BRAF G466V, and 15.4% had BRAF V600E mutations. 69.2% patients had liver metastasis, 23.1% had peritoneal metastases and 7.7% suffered metastasis to supraclavicular, retroperitoneal, and mesenteric lymph nodes. Median time from diagnosis of stage IV disease to progression was 25.3 months and median overall survival was 4.9 years.
Discussion: This study adds more insight to the limited existing data regarding rare mCRC cases with concomitant BRAF and RAS family mutations and exposes the need for future research on larger populations of this rare subset of patients.
Introduction
Colorectal cancer (CRC) is the third most common cause of cancer-related death in both men and women and ranks second in cancer-related deaths overall (1). Approximately 65% of patients diagnosed with CRC will either present with metastatic cancer at diagnosis or develop distant metastasis later in the course of the disease. The treatment of metastatic CRC (mCRC) has been individualized due to biomarkers that can be both prognostic and predictive, aiding in treatment decisions (2, 3).
Rat sarcoma (RAS) family genes (KRAS, NRAS) and v-raf murine sarcoma viral oncogene homolog B1 (BRAF) play a key role in the epithelial growth factor receptor (EGFR) pathway, an essential pathway for survival of cancer cells. Mutations in RAS family and BRAF genes lead to loss of cell cycle regulation and are the primary driver mutations for colorectal carcinogenesis (4). Specifically, RAS mutations are activating mutations which increase the expression and activity of RAS, driving an increase in cellular mitosis resulting in continuous proliferation and cell growth leading to carcinogenesis (5, 6). RAS mutations are present in about 40% of mCRC cases (5). KRAS is the most common of the RAS family genes to be involved in CRC and majority have missense gain of function mutation including codon 12, 13, and 61, most commonly G12D, G12V, G13D. Other codons that can be involved are 59, 117, 146. NRAS mutations in CRC are usually found in codons 12, 13, 61, most commonly G12A, G12D, G13S, Q61Q. The presence of a RAS mutation has been found to be a negative prognostic factor (5).
Conversely, BRAF mutations are only present in about 8-12% of mCRC cases (5). These mutations drive MAP-kinase pathway activation which leads to cellular growth, proliferation, and differentiation (5). BRAF mutations have been divided into three categories based upon their RAS dependency, kinase activity, and susceptibility to inhibitors. Class I are RAS-independent, kinase-activating V600 mutations, which signal as monomers and are characteristically responsive to BRAF inhibitors. Class II are RAS-independent, kinase-activating non-V600E mutations, which signal as dimers. Class III are RAS-dependent, BRAF variants characterized by the activation of tyrosine kinase receptors and enhancement of the ERK pathway. Class III mutations are associated with a better overall survival (OS), left-sided tumors, and absence of lymph node and peritoneal metastases (7).
CRC with a RAS mutation (8) or class I or II BRAF mutations (9) may be resistant to anti-EGFR therapies (5, 10). BRAF mutations in mCRC can be targeted with anti-BRAF agents and anti-EGFR agents in combination (11). Moreover, patients with BRAF V600E mutations may benefit from first-line therapy with encorafenib, cetuximab, and mFOLFOX 6 based on the BREAKWATER phase III trial (12). Until recently, mutations in BRAF and RAS were originally considered to be mutually exclusive in CRC and as such, very limited data have been published on their co-occurrence. One study showed that concurrent KRAS/BRAF mutations are reported in about 0.05% of metastatic CRC cases (5). We present a retrospective study that examines 13 patients with concomitant RAS and BRAF mutations in mCRC and its association with clinicopathological features and outcomes.
Materials and methods
We reviewed 763 samples from Guardant360 and FoundationOne databases kept by the Wake Forest Baptist Health Comprehensive Cancer Center. Descriptions of these databases can be found in Supplementary Tables 1 and 2. The study population included patients greater than age 18 with mCRC who were identified to have both a BRAF and RAS mutation by next generation sequencing (NGS). From these databases, we identified patients that had concomitant BRAF and RAS mutations in the CRC population. Further, we excluded patients who did not have mCRC and/or who had very limited data on chart review. We also excluded patients who had ARF mutations or RAF mutations. Ultimately, we identified 13 patients with mCRC and concomitant BRAF and RAS family gene mutations. NGS data was obtained on all 13 patients. The NGS sample was obtained from primary tumor (colon) in four patients, metastatic site in seven patients, and from circulating tumor DNA (ctDNA) in five patients. Three patients had samples obtained from both metastatic site and ctDNA. A retrospective record review was conducted in January 2024 on this sample to examine specific demographics and clinical features of the select population including sex, age at initial diagnosis and at diagnosis of stage IV disease, location of CRC, stage of the CRC at initial diagnosis, type of RAS mutation, type of BRAF mutation, location of metastasis, first-line treatment of metastatic cancer, second and further lines of treatment of the patients CRC, response to therapy, duration of response, time to progression, and survival length. We also reviewed tumor mutation burden (TMB) and microsatellite instability (MSI) status for each patient. MSI status was determined through NGS for eight patients. The others were determined through an alternative method. We used primarily descriptive statistics (means or percentages), and estimated time to progression and overall survival using Kaplan-Meier methods.
Results
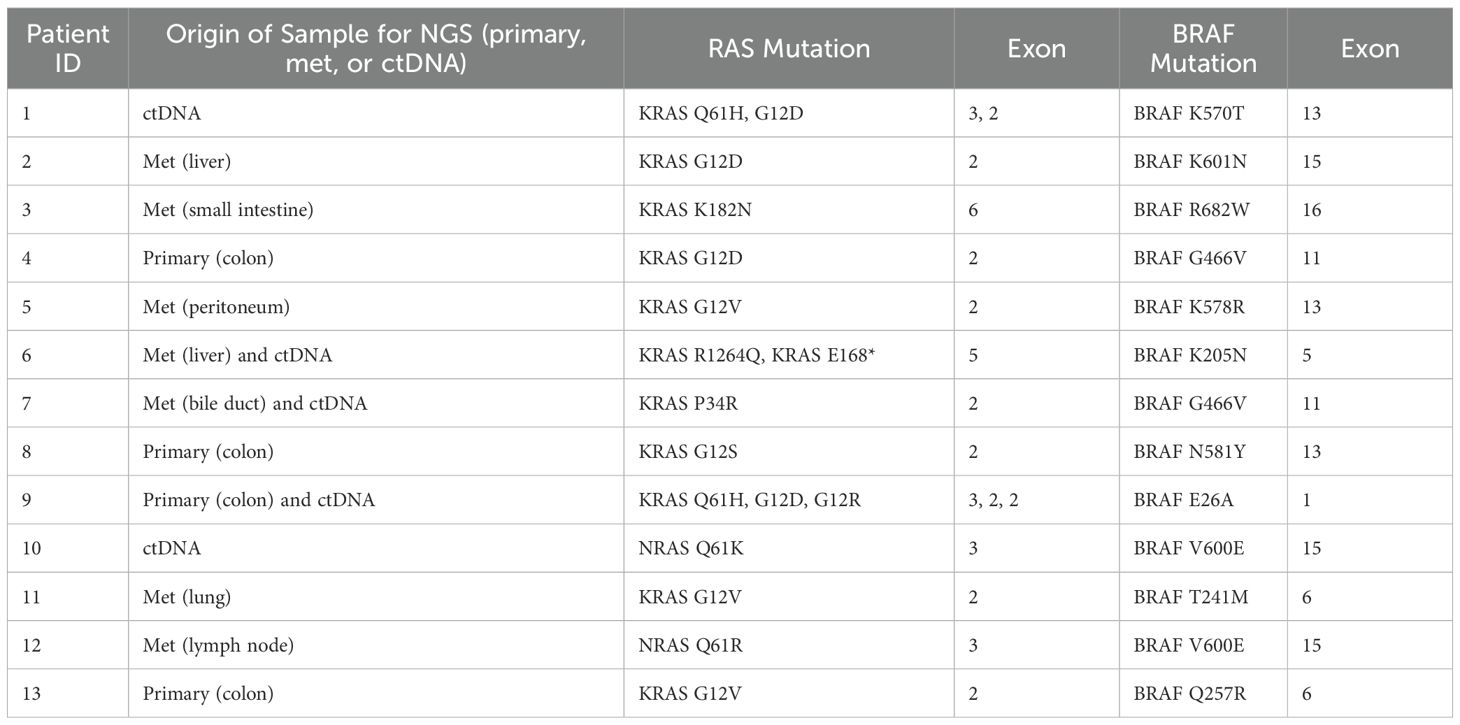
Eight patients (61.5%) were newly diagnosed with stage IV disease, the other five were diagnosed at stages I-III and later progressed to stage IV disease (Table 1). The following analyses were conducted on patients from the time of stage IV presentation. Patients were between 32 and 92 years old (median age 64.4). Eight (61.5%) were male and five (38.5%) were female. Eight patients had right-sided primary disease, four had left-sided, and one had synchronous primaries of the ascending and sigmoid colon. Table 2 shows each patient’s unique combination of mutations, the origin of the NGS sample, and the exon of each gene. KRAS mutation in codon 12 was the most prevalent RAS family mutation. Only two patients with a RAS family mutation harbored a BRAF V600E mutation.
Efficacy results include a median time to progression (from stage IV diagnosis) of 25.3 months (95% CI 8.3, 39.5). Nine patients (69.2%) were progression-free at 12 months and three patients (27.7%) were progression free at 36 months. Nine patients (69.2%) have died. The median time to death (from diagnosis of stage IV disease) was 4.9 years (95% CI 3.5, 5.5). 71.8% of patients survived four years, 13.7% survived six years. Three patients (23.1%) did not have progression of their disease at 30, 40, and 60 months. Two of these patients possessed KRAS G12D mutation. Table 3 shows additional information regarding the patients who did not have progression, including MSI and TMB status. MSI status is categorized into microsatellite instability high (MSI-H) and microsatellite stable (MSS).
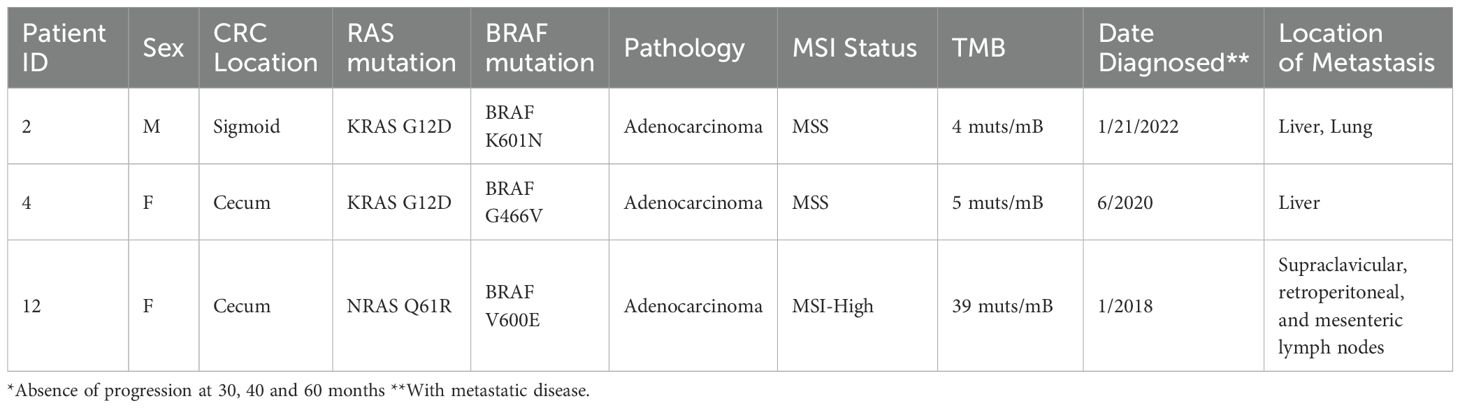
Table 3. Characteristics of the patients without progression* and were still living at the time of data collection.
Discussion
The presence of concomitant BRAF and RAS mutations has historically been thought to be mutually exclusive in mCRC. One study showed 0.064% in a population of 6,251 (13). A review of 11 papers identified a total of only 30 cases of concomitant RAS and BRAF mutations in mCRC (5). The present study describes 13 patients with concomitant RAS and BRAF mutations in mCRC.
In our cohort of patients, we identified many different combinations of concomitant BRAF and RAS mutations (Table 2). Primary tumor location and sites of metastasis, as listed in Table 1, show that the majority of the patients had liver metastasis, 69.2%. Three patients (23.1%) had peritoneal metastases and one (7.7%) with metastasis to supraclavicular, retroperitoneal (RP), and mesenteric lymph nodes. This distribution appears similar to what has been reported with BRAF-mutant mCRC patients except for the percentage of RP nodes (14, 15). Our cohort shares some features with BRAF-mutant-only mCRC, but the lower incidence of RP node involvement and higher liver-predominant disease suggests that concomitant RAS mutation may modify the metastatic pattern.
Individually, BRAF and RAS mutations in mCRC are associated with adverse prognosis. Prior studies have reported a median overall survival (OS) of 18.9 months in patients with BRAF-mutated mCRC, compared to 33.2 months in those with wild-type BRAF tumors (16) and median OS of 27.5 months for RAS wild-type and 17.3 months for RAS-mutated tumors (17). However, the presence of RAS and RAF mutations may reflect a more heterogeneous biology than previously appreciated since in our cohort, the median OS from the time of diagnosis of metastatic disease is 4.9 years (95% CI 3.5, 5.5) and three patients were still alive at the time of data collection.
Several factors may contribute to this prolonged survival, including access to sequential lines of effective systemic therapy, the use of multimodal treatment approaches, and the possibility of distinct tumor biology within this dual-mutant subset. Furthermore, these outcomes may reflect careful patient selection, as well as improvements in supportive care and molecularly-guided treatment strategies over time. This observation indicates the need to re-examine the prognostic assumptions in molecularly defined subgroups and highlights the importance of continued molecular characterization and individualized treatment planning.
Of the three patients without progression at 30, 40, and 60 months and were still alive at the time of data collection, one had MSI-H disease with a high TMB of 39 mutations per megabase (muts/mB). This patient responded favorably to the immune checkpoint inhibitor pembrolizumab and remained progression-free at the time of data collection. Pembrolizumab is a standard first-line therapy for mCRC in patients with MSI-H or mismatch repair–deficient (dMMR) tumors (18, 19). Emerging data suggest that immunotherapy may be equally effective in MSI-H mCRC regardless of BRAF mutation status (20, 21). Ongoing trials, such as SEAMARK, are investigating whether adding targeted therapy offers additional benefit in this population, or whether immunotherapy alone is sufficient (22). Further, the CheckMate 8HW trial showed that patients with MSI-H/dMMR mCRC treated with nivolumab plus ipilimumab had significantly longer progression-free survival (PFS) when compared to treatment with chemotherapy alone, including patients with baseline RAS or BRAF mutations (23). However, it remains unclear whether this included patients with co-occurring RAS and BRAF mutations and how that might impact outcomes.
The remaining two long-term survivors both harbored the KRAS G12D mutation. One patient was receiving fourth-line treatment with TAS-102 and bevacizumab as of August 2023, while the other underwent right hemicolectomy and liver metastasectomy following conversion therapy with FOLFOX, followed by FOLFIRI and bevacizumab. The KRAS G12D mutation is among the most commonly identified KRAS alterations in colorectal and other gastrointestinal carcinomas (6, 24). Interestingly, this mutation has been associated with a more favorable OS compared to other KRAS variants in colorectal cancer (6). These observations raise the hypothesis that KRAS G12D may confer a prognostic advantage. Further investigation is warranted to explore the biological and clinical implications of this specific mutational profile.
The V600E mutation normally comprises approximately 90% of BRAF mutations present in CRC (23). Surprisingly, of the 13 patients in this series, only two possessed BRAF V600E mutation (15.4%). One of the two patients is still living as described above. The other, also female, had cecal primary with metastasis to liver and lung, concomitant NRAS Q61K mutation, unknown microsatellite status, and 9 muts/mB. She only lived four years from diagnosis despite multiple lines of treatment. This raises the question of whether non-V600E BRAF mutations may be more common in the presence of concomitant RAS expression.
Though still limited, there have been increasing amounts of data and studies describing concomitant RAS and BRAF mutations in mCRC. Despite this, it remains challenging to draw clear conclusions regarding the prognostic role of these concomitant mutations, their impact on PFS, OS, or how current chemotherapy regimens can be modified or combined to gain better control of the disease. The limitations of the current study include its small sample size, its retrospective nature, its lack of comparison group, and selection bias in that it represents a sample of patients from one small geographic area. Further, we are unable to draw conclusions about whether concomitant mutations effects OS as we did not compare our sample to those without concomitant mutations.
Our cohort suggests that double-mutated mCRC may have a metastatic pattern more similar to RAS-mutant CRC than solitary BRAF-mutated disease. It posits that KRAS G12D may contribute to improved prognosis even in the presence of BRAF mutation. And it questions the existing literature that the group I and II BRAF mutations are associated with worse prognosis. It also raises the question of whether other factors, like the presence of concomitant RAS family mutation can alter the trajectory of patients’ overall disease course. Ultimately, this study elucidates the need for more research on larger populations of patients who possess concomitant RAS and BRAF mutations in mCRC.
Data availability statement
The data analyzed in this study is subject to the following licenses/restrictions: Guardant360 and FoundationOne are Next-Generation Sequencing (NGS) Vendors. Wake Forest has established data use agreements with both Guardant360 and FoundationOne, allowing them to use the data for research purposes. Requests to access these datasets should be directed to https://guardanthealth.com/contact/; ZGF0YWNvbGxhYm9yYXRpb25zQGZvdW5kYXRpb25tZWRpY2luZS5jb20=.
Ethics statement
This study involving human participants was reviewed and approved by the Advocate Health - Wake Forest University School of Medicine Institutional Review Board. Written informed consent from the patients was not required to participate in this study in accordance with the national legislation and the institutional requirements.
Author contributions
JS: Conceptualization, Data curation, Investigation, Visualization, Writing – original draft, Writing – review & editing. ML: Data curation, Investigation, Visualization, Writing – original draft, Writing – review & editing. JE: Formal analysis, Investigation, Writing – review & editing. RP: Supervision, Validation, Writing – review & editing. CR: Conceptualization, Methodology, Supervision, Validation, Writing – review & editing.
Funding
The author(s) declare that no financial support was received for the research and/or publication of this article.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fonc.2025.1621412/full#supplementary-material
References
1. Siegel RL, Wagle NS, Cercek A, Smith RA, and Jemal A. Colorectal cancer statistics, 2023. CA Cancer J Clin. (2023) 73:233–54. doi: 10.3322/caac.21772
2. Crutcher M and Waldman S. Biomarkers in the development of individualized treatment regimens for colorectal cancer. Front Med (Lausanne). (2022) 9:1062423. doi: 10.3389/fmed.2022.1062423
3. Ruiz-Bañobre J, Kandimalla R, and Goel A. Predictive biomarkers in metastatic colorectal cancer: A systematic review. JCO Precis Oncol. (2019) 3:1–17. doi: 10.1200/PO.18.00260
4. Midthun L, Shaheen S, Deisch J, Senthil M, Tsai J, and Hsueh CT. Concomitant KRAS and BRAF mutations in colorectal cancer. J Gastrointest Oncol. (2019) 10:577–81. doi: 10.21037/jgo.2019.01.10
5. Afrăsânie VA, Gafton B, Marinca MV, Alexa-Stratulat T, Miron L, Rusu C, et al. The coexistence of RAS and BRAF mutations in metastatic colorectal cancer: A case report and systematic literature review. J Gastrointestin Liver Dis. (2020) 29:251–6. doi: 10.15403/jgld-1003
6. Koulouridi A, Karagianni M, Messaritakis I, Sfakianaki M, Voutsina A, Trypaki M, et al. Prognostic value of KRAS mutations in colorectal cancer patients. Cancers. (2022) 14:3320. doi: 10.3390/cancers14143320
7. Srivastava P, Mishra S, Shukla S, Sharma P, and Husain N. Concomitant non-V600E BRAF and KRAS mutations in colorectal carcinoma by next-generation sequencing: A distinct subtype. Int J Surg Pathology. (2023) 32:1186–90. doi: 10.1177/10668969231215425
8. Douillard JY, Oliner KS, Siena S, Tabernero J, Burkes R, Barugel M, et al. Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med. (2013) 369:1023–34. doi: 10.1056/NEJMoa1305275
9. Xu T, Li J, Wang Z, Zhang X, Zhou J, Lu Z, et al. Real-world treatment and outcomes of patients with metastatic BRAF mutant colorectal cancer. Cancer Med. (2023) 12:10473–84. doi: 10.1002/cam4.5783
10. Therkildsen C, Bergmann TK, Henrichsen-Schnack T, Ladelund S, and Nilbert M. The predictive value of KRAS, NRAS, BRAF, PIK3CA and PTEN for anti-EGFR treatment in metastatic colorectal cancer: A systematic review and meta-analysis. Acta Oncol. (2014) 53:852–64. doi: 10.3109/0284186X.2014.895036
11. Yaeger R, Cercek A, O’Reilly EM, Reidy DL, Kemeny N, Wolinsky T, et al. Pilot trial of combined BRAF and EGFR inhibition in BRAF-mutant metastatic colorectal cancer patients. Clin Cancer Res 15 March. (2015) 21:1313–20. doi: 10.1158/1078-0432.CCR-14-2779
12. Elez E, Yoshino T, Shen L, Lonardi S, Van Cutsem E, Eng C, et al. Encorafenib, cetuximab, and mFOLFOX6 in BRAF-mutated colorectal cancer. N Engl J Med. (2025) 392:2425–37. doi: 10.1056/NEJMoa2501912
13. Kopetz S, Tie J, Gibbs P, Jiang Z-Q, Lieu CH, Agarwal A, et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer. (2011) 117:4623–32. doi: 10.1002/cncr.26086
14. Lipsyc M and Yaeger R. Impact of somatic mutations on patterns of metastasis in colorectal cancer. J Gastrointest Oncol. (2015) 6:645–9. doi: 10.3978/j.issn.2078-6891.2015.045
15. Tan E, Whiting J, Xie H, Imanirad I, Carballido E, Felder S, et al. BRAF mutations are associated with poor survival outcomes in advanced-stage mismatch repair-deficient/microsatellite high colorectal cancer. Oncologist. (2022) 27:191–7. doi: 10.1093/oncolo/oyab055
16. Ottaiano A, Normanno N, Facchini S, Cassata A, Nappi A, Romano C, et al. Study of ras mutations’ Prognostic value in metastatic colorectal cancer: STORIA analysis. Cancers. (2020) 12:1919. doi: 10.3390/cancers12071919
17. André T, Shiu KK, Kim TW, Jensen BV, Jensen LH, Punt C, et al. Pembrolizumab in microsatellite-instability-high advanced colorectal cancer. N Engl J Med. (2020) 383:2207–18. doi: 10.1056/NEJMoa2017699
18. Morris VK, Kennedy EB, Baxter NN, Benson III Al B, Cercek A, Cho MVan K. Morris, Erin B. Kennedy, Nancy N. Baxter, Al B. Benson III, Andrea Cercek, May Cho, et al. Treatment of metastatic colorectal cancer: ASCO guideline. JCO. (2023) 41:678–700. doi: 10.1200/JCO.22.01690
19. Gbolahan O and O’Neil B. Update on systemic therapy for colorectal cancer: biologics take sides. Transl Gastroenterol Hepatol. (2019) 4:9. doi: 10.21037/tgh.2019.01.12
20. Colle R, Lonardi S, Cachanado M, Overman MJ, Elez E, Fakih M, et al. BRAF V600E/RAS mutations and lynch syndrome in patients with MSI-H/dMMR metastatic colorectal cancer treated with immune checkpoint inhibitors. Oncologist. (2023) 28:771–9. doi: 10.1093/oncolo/oyad082
21. Zeissig MN, Ashwood LM, Kondrashova O, and Sutherland KD. Next batter up! Targeting cancers with KRAS-G12D mutations. Trends Cancer. (2023) 9:955–67. doi: 10.1016/j.trecan.2023.07.010
22. Elez E, Kopetz S, Tabernero J, Bekaii-Saab T, Taieb J, Yoshino T, et al. SEAMARK: phase II study of first-line encorafenib and cetuximab plus pembrolizumab for MSI-H/dMMR BRAFV600E-mutant mCRC. Future Oncol. (2024) 20:653–63. doi: 10.2217/fon-2022-1249
23. Andre T, Elez E, Van Cutsem E, Jensen LH, Bennouna J, Mendez G, et al. Nivolumab plus ipilimumab in microsatellite-instability-high metastatic colorectal cancer. N Engl J Med. (2024) 391:2014–26. doi: 10.1056/NEJMoa2402141
Keywords: colorectal cancer, BRAF mutation, KRAS mutation, NRAS mutation, RAS family mutation, concurrent RAS/BRAF variants, clinical-pathological features
Citation: Srivastav J, Lehman ME, Evans JK, Paluri R and Rocha Lima CMSP (2025) A compilation of 13 patients with metastatic colorectal cancer and concomitant BRAF and RAS family mutations. Front. Oncol. 15:1621412. doi: 10.3389/fonc.2025.1621412
Received: 01 May 2025; Accepted: 06 August 2025;
Published: 25 August 2025.
Edited by:
Caiyue Li, Louisiana State University, United StatesReviewed by:
Nadia Saoudi Gonzalez, Vall d’Hebron Institute of Oncology (VHIO), SpainGabriela Remonatto, Hospital de Clínicas de Porto Alegre, Brazil
Irem Bilgetekin, Memorial Ankara Hospital, Türkiye
Copyright © 2025 Srivastav, Lehman, Evans, Paluri and Rocha Lima. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Morgan E. Lehman, bW9yZ2FuLmxlaG1hbkBhZHZvY2F0ZWhlYWx0aC5vcmc=
 Jigisha Srivastav
Jigisha Srivastav Morgan E. Lehman
Morgan E. Lehman Joni K. Evans2
Joni K. Evans2 Ravi Paluri
Ravi Paluri