- 1Department of Pharmacy, The University of Tokyo Hospital, Tokyo, Japan
- 2Department of Integrative Physiology and Bio-Nano Medicine, National Defense Medical College, Saitama, Japan
- 3Department of Preventive Medicine, Nagoya University Graduate School of Medicine, Aichi, Japan
- 4Program in Radiological and Medical Laboratory Sciences, Pathophysiological Laboratory Science, Nagoya University Graduate School of Medicine, Aichi, Japan
- 5Department of Pathophysiology, Tokyo University of Pharmacy and Life Sciences, Tokyo, Japan
- 6Department of Anatomy and Neurobiology, National Defense Medical College, Saitama, Japan
Dysfunctional missense variant of organic anion transporter 10 (OAT10/SLC22A13), rs117371763 (c.1129C>T; p.R377C), is associated with a lower susceptibility to gout. OAT10 is a urate transporter; however, its physiological role in urate handling remains unclear. We hypothesized that OAT10 could be a renal urate re-absorber that will be a new molecular target of urate-lowering therapy like urate transporter 1 (URAT1, a physiologically-important well-known renal urate re-absorber) and aimed to examine the effect of OAT10 dysfunction on renal urate handling. For this purpose, we conducted quantitative trait locus analyses of serum urate and fractional excretion of uric acid (FEUA) using samples obtained from 4,521 Japanese males. Moreover, we performed immunohistochemical and functional analyses to assess the molecular properties of OAT10 as a renal urate transporter and evaluated its potential interaction with urate-lowering drugs. Clinico-genetic analyses revealed that carriers with the dysfunctional OAT10 variant exhibited significantly lower serum urate levels and higher FEUA values than the non-carriers, indicating that dysfunction of OAT10 increases renal urate excretion. Given the results of functional assays and immunohistochemical analysis demonstrating the expression of human OAT10 in the apical side of renal proximal tubular cells, our data indicate that OAT10 is involved in the renal urate reabsorption in renal proximal tubules from urine. Additionally, we found that renal OAT10 inhibition might be involved in the urate-lowering effect of losartan and lesinurad which exhibit uricosuric effects; indeed, losartan, an approved drug, inhibits OAT10 more strongly than URAT1. Accordingly, OAT10 can be a novel potential molecular target for urate-lowering therapy.
Introduction
Elevated serum urate levels can cause gout, the most common form of inflammatory arthritis (Dalbeth et al., 2019). Under physiological conditions, uric acid—the terminal metabolite of purine metabolism in humans—primarily exists as urate, an anion form. Because urate cannot passively permeate across cellular membranes, urate transporters play a pivotal role in urate handling in our body. To date, urate transporter 1 (URAT1/SLC22A12) (Enomoto et al., 2002; Ichida et al., 2004), glucose transporter 9 (GLUT9/SLC2A9) (Matsuo et al., 2008; Dinour et al., 2010), and ATP-binding cassette transporter G2 (ABCG2/BCRP) (Matsuo et al., 2009; Woodward et al., 2009) have been identified as physiologically important urate transporters of which genetic dysfunction influences serum urate levels; therefore, these three transporters have been extensively studied in the context of urate regulation in our body. Nevertheless, such urate transporters do not provide a comprehensive picture of urate-handling systems which are also related to the risk of gout/hyperuricemia in humans. Besides, this point is supported by previous findings in animal studies. For instances, Abcg2 knockout mice retain approximately half levels of intestinal urate secretion activities compared with wild-type mice (Ichida et al., 2012); genetic disruption of Urat1 in uricase knockout mice results in the elevation of fractional excretion of uric acid (FEUA) (around to 50%, mean value) but not reaching to 100% (Hosoyamada et al., 2016), indicating the presence of other latent machineries that would be theoretically involved in the remaining portions of urate transport.
The kidney is an important organ for regulating serum urate levels by re-absorbing almost all of filtered urate and is responsible for two-thirds of the net urate secretion from the body. Regarding the renal urate re-absorption route, both URAT1 (as an importer) and GLUT9 (as an exporter) form a cascaded system of transcellular urate transport from urine to blood via renal proximal tubular cells as apical (urine side) and basal (blood side) machinery, respectively. It is noteworthy that this route remains functional to some extent even in subjects genetically lacking URAT1 (Ichida et al., 2004) as well as in animal models with Urat1 knockout genetic background (Hosoyamada et al., 2010; Hosoyamada et al., 2016), suggesting that there is an alternative to URAT1. Hitherto, based on urate transport activities determined by in vitro experiments and renal expression, several transporters have been proposed as the candidates including SLC22 family proteins (Koepsell, 2013; Halperin Kuhns and Woodward, 2021). Among them, organic anion transporter 10 (OAT10)/SLC22A13 is strongly expressed in the kidney and transport assays using Xenopus oocytes (Bahn et al., 2008) and mammalian culture cells (Higashino et al., 2020) demonstrated that urate is an OAT10 substrate. However, not only affinity of OAT10 protein to urate but also the effects of OAT10 dysfunction on the renal urate handling remain unclear.
In this study, we hypothesized that OAT10 could be a physiologically important machinery involved in the renal urate re-absorption based on our recent study (Higashino et al., 2020), in which clinico-genetic and functional studies revealed that a functionally null variant of OAT10 (c.1129C>T; p.R377C that diminishes the urate transport activity of OAT10 with little effect on its cellular protein level and plasma membrane localization) significantly decreases the risk of gout (odds ratio, OR = 0.67). This finding, together with the lower serum urate levels associated with this dysfunctional variant (Higashino et al., 2020), led us to reason that OAT10 could have a physiological impact on renal urate handling. To clarify the proposed involvement of OAT10 in renal urate re-absorption that remains speculation for a long time, we herein investigated the physiological role of OAT10 via a clinico-genetic approach. Moreover, to gain insight into the molecular properties of OAT10 as a urate transporter, including interaction with various urate-lowering drugs, we conducted functional analyses.
Materials and Methods
Ethics Approval
The studies involving human participants were reviewed and approved by the Institutional Ethics Committees of the National Defense Medical College and Nagoya University. All protocols were in accordance with the Declaration of Helsinki, and written informed consent was obtained from each participant in the present study.
Participants
A total of 4,521 Japanese men were recruited from health examination participants in Shizuoka and Daiko areas of the Japan Multi-Institutional Collaborative Cohort Study (Hamajima, 2007; Asai et al., 2009). The characteristics of the participants are shown in Table 1. Of note, under normal physiological conditions, uric acid (molecular weight of 168.1) exists in the blood mainly as urate (the anion form of uric acid, molecular weight of 167.1); on the other hand, in clinical testing method, this parameter is measured and converted to concentration of “uric acid (mg/dl)” in the serum, as described previously (Nakatochi et al., 2021). Nevertheless, based on the G-CAN (The Gout, Hyperuricemia, and Crystal-Associated Disease Network) nomenclature for gout (Bursill et al., 2019), we herein used the term of “serum urate,” instead of serum uric acid, for mentioning the circulating form.
Quantitative trait locus (QTL) analysis of serum urate was performed for all participants. Among them, for 1,518 men with available urinary data, we performed subsequent QTL analysis of FEUA. Based on the results of blood and urine tests, FEUA was calculated as follows [urinary urate (mg/dl) × serum creatinine (mg/dl)]/[serum urate (mg/dl) × urinary creatinine (mg/dl)] × 100 (%).
Genetic Analysis of Human OAT10/SLC22A13 Gene
Genomic DNA was extracted from whole peripheral blood cells of the 4,521 Japanese male participants according to methods described in our previous study (Sakiyama et al., 2021). To genotype rs117371763 (c.1129C>T: p.R377C) of the OAT10/SLC22A13 gene, we employed the TaqMan method (Thermo Fisher Scientific, Kanagawa, Japan) using a QuantStudio 5 real-time PCR system (Thermo Fisher Scientific), as described in our previous study (Higashino et al., 2020). Also, like in the previous study, to confirm the accuracy of the genotyping results, >100 samples were also subjected to direct sequencing with the following primers: 5′-TGGTGTGTGTTGGCAGAG-3′ and 5′-GGTCCCATCCACTGGAAC-3′. DNA sequence analysis was performed with a 3130xl Genetic Analyzer (Thermo Fisher Scientific). Genotype distributions were as follows: 1129C/C, 4,128 subjects; 1129C/T, 384 subjects; 1129T/T, 9 subjects. Among them, there were 1,369, 143, and 6 subjects with available urinary data, respectively.
Materials
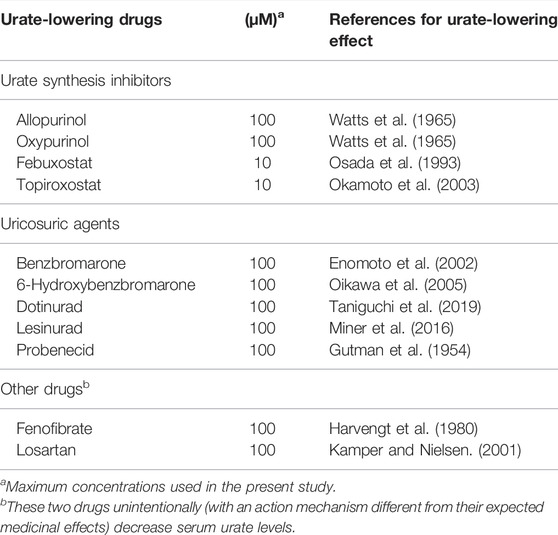
The critical materials and resources used in this study were summarized in Supplementary Table S1. Dotinurad was kindly provided by Fuji Yakuhin (Saitama, Japan) under a material transfer agreement. All other chemicals used were commercially available and of analytical grade. In this study, we evaluated the following urate-lowering drugs and their active metabolites: urate synthesis inhibitors (allopurinol, oxypurinol, febuxostat, and topiroxostat), uricosuric agents (benzbromarone, 6-hydroxybenzbromarone, dotinurad, lesinurad, and probenecid), and other drugs that decrease serum urate levels with an action mechanism different from their expected medicinal effects (fenofibrate and losartan). Detailed information is summarized in Table 2.
Generation of an Anti-OAT10 Polyclonal Antibody
An anti-human organic anion transporter 10 (OAT10) polyclonal antibody was raised in rabbits according to methods described in our previous study (Takada et al., 2015). In brief, the rabbit polyclonal antibody was generated against two KLH-coupled human OAT10 peptides as follows: amino acids #523–536 (KLH)-C + PKSVPSEKETEAKG; amino acids #293–306, (KLH)-C + KAASVNRRKLSPEL, which were also used to determine the titer of antiserum. On day 77, whole blood was collected from an adequately-immunized rabbit. The prepared antiserum was further purified before use using an epitopes-conjugated affinity column.
Immunohistochemistry
Immunohistochemical analysis using the anti-OAT10 antibody was conducted according to our previous study (Chiba et al., 2015). The analyzed tissue was the normal part of the human kidney obtained from a male patient with renal cell carcinoma. In brief, with 4-μm paraffin sections of the kidney, antigen retrieval was conducted by microwave oven-heating in a retrieval solution (0.01 M citrate buffer, pH 6.0) for 10 min; followed by inactivation of endogenous peroxidase activity with hydrogen peroxide (3% in methanol) treatment for 10 min, and further washing with Tris-buffered saline containing 0.05% Tween 20 (TBST). After blocking with 5% goat serum in TBS for 10 min, the sections were incubated with the anti-OAT10 antibody (1:50 dilution in TBS containing 1% bovine serum albumin, BSA) at 4°C overnight; subsequently the sections were treated with the Dako EnVision™ Kit/horseradish peroxidase (HRP) (K1491; Dako, Kyoto, Japan) as a secondary antibody, followed by 3,3′-diaminobenzidine (Dako) treatment for the detection of the immune reactions. Nuclei were counterstained with Mayer’s hematoxylin. For antigen absorption test, to block specific staining, the anti-OAT10 antibody was pre-incubated with the OAT10 peptides, which were used at a concentration of 100 μg/ml, before staining procedure. After visualization, the specimens were imaged using an upright microscope (Eclipse 50i; Nikon, Tokyo, Japan) equipped with a Nikon Digital Sight DS-5M camera (Nikon).
Plasmid Construction
The full-length human OAT10/SLC22A13 wild-type (WT) (NCBI accession; NM_004256) open reading frame (ORF) in pEGFP-N1 plasmid was constructed in our previous study (Higashino et al., 2020). The full-length human URAT1/SLC22A12 WT (NCBI accession; NM_144585.3) ORF in pEGFP-C1 was derived from our previous study (Toyoda et al., 2020a). Prior to further experiments described below, each plasmid containing the transporter and control empty vectors were obtained in the same lot using a PureLink™ HiPure Plasmid Filter Midiprep Kit (Thermo Fisher Scientific).
Cell Culture
Human embryonic kidney 293 (HEK293)-derived 293A cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (Nacalai Tesque, Kyoto, Japan) supplemented with 10% fetal bovine serum (Biowest, Nuaillé, France), 1% penicillin-streptomycin (Nacalai Tesque), 2 mM L-glutamine (Nacalai Tesque), and 1 × Non-Essential Amino Acid (Life Technologies, Tokyo, Japan) at 37°C in a humidified atmosphere of 5% (v/v) CO2 in air; we confirmed that 293A cells used were negative for mycoplasma contamination using MycoAlert™ mycoplasma detection kit (Lonza, Basel, Switzerland). Madin-Darby canine kidney II (MDCKII) cells (a well-used polarized renal cell line to investigate the cellular localization of transporter proteins—apical or basal membranes) were also maintained in a similar way.
For the in vitro transport assay, 24 h after the seeding of the cells (0.92 × 105 cells/cm2) onto 12-well cell culture plates, each vector plasmid was transfected into 293A cells with a forward transfection approach using polyethyleneimine “MAX” (PEI-MAX) (Polysciences, Warrington, PA, United States) as described previously (Toyoda et al., 2020a). All experiments were carried out with 293A cells at passages 12–20. Of note, such mammalian cell lines have been well-used to evaluate urate transport activity of target protein (Miner et al., 2016; Toyoda et al., 2020b; Higashino et al., 2020).
For confocal microscopy, each vector plasmid was transfected into MDCKII cells with a reverse transfection approach using PEI-MAX as described previously (Toyoda et al., 2016). In brief, MDCKII cells were collected after trypsinization, followed by centrifugation at 1,000 × g for 5 min. The cell pellet was re-suspended in fresh DMEM, and the resulting suspension was mixed with plasmid/PEI-MAX mixture (2 μg of plasmid/10 μl of PEI-MAX for 1 × 106 cells of MDCKII cells). Then, the MDCKII cells were re-seeded onto cell culture plates at a concentration of 1 × 105 cells/cm2. For whole cell lysate preparations, MDCKII cells were re-seeded onto 6-well cell culture plates at the same concentration. All experiments were carried out using MDCKII cells at passages 7–10.
Preparation of Protein Lysates and Immunoblotting
After preparation of whole cell lysates with cell lysis buffer A [50 mM Tris/HCl (pH 7.4), 1 mM dithiothreitol, 1% (v/v) Triton X-100, and cOmplete™, EDTA-free protease inhibitor cocktail (Roche, Basel, Switzerland)], to examine the N-linked glycosylation status of OAT10 and URAT1 proteins, whole cell lysate samples were treated with Peptide N-glycosidase F (PNGase F) (New England Biolabs, Ipswich, MA, United States) as described previously (Higashino et al., 2020). The protein concentration was determined using the Pierce™ BCA Protein Assay Kit (Thermo Fisher Scientific) with bovine serum albumin (BSA) as a standard according to the manufacturer’s protocol. The samples were separated using SDS-PAGE and transferred to an Immobilon-P PVDF membrane (Millipore, Bedford, MA, United States) by electroblotting at 15 V for 60 min. Blots were probed with appropriate antibodies (Supplementary Table S1) according to previous studies (Toyoda et al., 2020a; Higashino et al., 2020), and the signals were visualized by using chemiluminescence and detected using a multi-imaging analyzer Fusion Solo 4™ system (Vilber Lourmat, Eberhardzell, Germany).
Confocal Microscopic Observation
Confocal laser scanning microscopic observation was conducted as described previously, with some minor modifications (Toyoda et al., 2016). In brief, 72 h after the transfection, MDCKII cells were fixed with ice-cold methanol for 10 min at room temperature. After washing three times with PBS (−) containing 1% BSA (BSA-PBS), the cells were incubated with a mouse anti-gp135 monoclonal antibody (3F2/D8; the Developmental Studies Hybridoma Bank, Iowa City, IA, United States) diluted 100-fold in BSA-PBS for 1 h at room temperature, followed by incubation with a goat anti-mouse IgG-Alexa Fluor 546-conjugate (A-11003; Thermo Fisher Scientific), diluted 200-fold in BSA-PBS, for 1 h at room temperature in the dark. Subsequently, cells were subjected to TO-PRO-3 Iodide (Molecular Probes, Eugene, OR, United States) staining. After the visualization of the nuclei, the cells were mounted in VECTASHIELD Mounting Medium (Vector Laboratories, Burlingame, CA, United States). To analyze the localization of EGFP-fused OAT10 or EGFP-fused URAT1 proteins, fluorescence was detected using an FV10i Confocal Laser Scanning Microscope (Olympus, Tokyo, Japan).
Urate Uptake Assay
With OAT10, urate uptake assay using OAT10-expressing 293A cells was conducted according to our previous study (Higashino et al., 2020). In brief, 48 h after the plasmid transfection, the cells were washed twice with a transport buffer (Ringer solution: 130 mM NaCl, 4 mM KCl, 1 mM Na2HPO4, 1 mM MgSO4, 1 mM CaCl2, 20 mM HEPES, 18 mM D-glucose, at pH 6.4 unless otherwise indicated) and pre-incubated in Ringer solution for 10 min at 37°C. The cells were further incubated for 1 min in pre-warmed fresh Ringer solution containing 10 μM [8–14C]-urate unless otherwise indicated. To examine the OAT10-inhibitory effects of the target compounds, Ringer solution either without (i.e., with only vehicle control, 0.1% dimethyl sulfoxide, DMSO) or with the individual compounds at the indicated concentrations was used. Subsequently, the cells were washed three times with ice-cold Ringer solution, lysed with 500 μl of 0.2 M NaOH, and neutralized with 100 μl of 1 M HCl. We then measured the radioactivity in the lysate using a liquid scintillator (Tri-Carb 3110TR; PerkinElmer, Waltham, MA, United States). Protein concentrations were determined using the Pierce™ BCA Protein Assay Kit, as described above. The urate transport activity was calculated as the incorporated clearance (μL/mg protein/min) [incorporated level of urate (DPM/mg protein/min)/urate level in the incubation mixture (DPM/μL)]. The OAT10-mediated urate transport activity was calculated by subtracting the urate transport activity of mock cells from that of OAT10-expressing cells. To examine the OAT10-inhibitory effects of the target compounds, their effects on the urate transport activity of mock cells were also examined.
With URAT1, urate uptake assay using URAT1-expressing 293A cells was conducted with a Cl−-free transport buffer [Cl−-free Hanks’ Balanced Salt Solution (HBSS): 125 mM Na-gluconate, 4.8 mM K-gluconate, 1.2 mM KH2PO4, 1.2 mM MgSO4, 1.3 mM Ca-gluconate, 25 mM HEPES, 5.6 mM D-glucose, at pH 6.4 unless otherwise indicated] in a similar way according to our previous studies (Miyata et al., 2016; Toyoda et al., 2020a). In this case, after pre-incubation in Cl−-free HBSS for 15 min at 37°C, the cells were further incubated for 20 s in pre-warmed fresh Cl−-free HBSS containing 10 μM [8–14C]-urate either without (i.e., with only vehicle control, 0.1% DMSO) or with the individual compounds at the indicated concentrations. Of note, imposing a Cl− gradient by removal of external Cl− results in the acceleration of URAT1-mediated urate uptake into the cell (Enomoto et al., 2002), bringing a relative reduction in the background levels for urate transport activities. Given this advantage for conducting in vitro assays, the Cl−-free condition has been used for functional investigations on URAT1 in various studies (Anzai et al., 2004; Miyata et al., 2016; Bao et al., 2019; Toyoda et al., 2020a). In this point, we had confirmed that the Cl−-free HBSS is a better choice for inhibitory tests against URAT1-mediated urate transport than Ringer solution (Supplementary Figure S1).
Calculation of the Half-Maximal Inhibitory Concentration Values
To calculate the half maximal inhibitory concentration (IC50) values of the target compounds against urate transport by OAT10 or URAT1, the urate transport activities were measured in the presence of each compound at several concentrations. The OAT10- or URAT1-mediated transport activities were expressed as a percentage of the control (100%). Based on the calculated values, fitting curves were obtained according to the following formula using the least-squares method with Excel 2019 (Microsoft, Redmond, WA, United States) as described previously (Toyoda et al., 2020a):
where, Emax is the maximum effect, EC50 is the half-maximal effective concentration, C is the concentration of the test compound, and n is the sigmoid-fit factor. Finally, based on these results, the IC50 was calculated.
Statistics
With clinical analyses, statistical analyses were performed using SPSS v.22.0J (IBM Japan, Tokyo, Japan). Univariate linear regression analysis was used for quantitative trait locus analyses. With functional analyses, all statistical analyses were performed using Excel 2019 with Statcel4 add-in software (OMS publishing, Saitama, Japan). To determine the kinetic parameters of the OAT10-mediated urate transport—Michaelis-Menten constant (Km) and maximal velocity (Vmax), the Michaelis–Menten model was fitted to the experimental values of the urate transport rates, and the urate concentrations by non-linear regression curve fitting using GraphPad Prism 8 (GraphPad Software, San Diego, CA, United States). Different statistical tests were used for different experiments, as described in the table captions of figure legends which include the number of biological replicates (n). In the case of a single pair of quantitative data, after comparing the variances of a set of data using an F-test, an unpaired Student’s t-test was performed. Statistical significance was defined in terms of p values less than 0.05 or 0.01.
Results
Increase of Net Renal Urate Secretion Associated With the Genetic Dysfunction of OAT10
To investigate the effect of a genetic dysfunction of OAT10 on renal urate secretion, we conducted clinico-genetic analyses. Regarding the functionally null variant (OAT10 c.1129C>T), only a few subjects were homozygotes (1129T/T); the others were heterozygotes (1129C/T) or wild-types (1129C/C). Thus, we compared the following parameters related to renal urate handling between the latter two groups (OAT10 1129C/T vs. 1129C/C) (Figure 1). Serum urate levels were significantly lower (Figure 1A; Supplementary Table S2) and the values of FEUA were significantly higher (Figure 1B; Supplementary Table S3) in heterozygous carriers of OAT10 c.1129T (C377) than those in non-carriers. This inverse association suggested that, like URAT1, OAT10 could be involved in urate re-uptake from urine. Additionally, the value of coefficient for the effect allele with the urate-lowering effect was −0.135 mg/dl/allele (Supplementary Table S2) which was not so large but could support the physiological importance of OAT10 as discussed later (see Discussion). Also, similar results were obtained when we combined the homozygotes with the heterozygotes (Supplementary Tables S4, S5). Moreover, immunohistochemistry revealed the luminal expression of OAT10 in the proximal tubules in the cortex of the human kidney (Figure 2), but the expression of OAT10 was hardly detected in the medulla, which is a similar pattern to that of URAT1 (Enomoto et al., 2002). The specificity of the anti-OAT10 antibody was verified by antigen absorption test (Supplementary Figure S2). Also, this luminal expression of OAT10 in the proximal tubules is consistent with the data in the Human Protein Atlas (Uhlén et al., 2015) (https://www.proteinatlas.org/ENSG00000172940-SLC22A13/tissue/kidney#img). Besides, when expressed in polarized MDCKII cells, OAT10 was matured as a glycoprotein (Figure 3A) and localized on the apical membrane, similar to URAT1 (Figure 3B). These results indicate that OAT10 prefers to be on the apical side of renal proximal tubule cells in the kidney, the position required for the urate re-uptake from urine into the cell. These findings are consistent with a result of immunoblotting addressing renal expression of rat Oat10 in a previous study (Bahn et al., 2008). Given these clinical and experimental data, OAT10 would be a renal urate re-absorber in humans that influences serum urate levels and net amount of renal urate excretion.
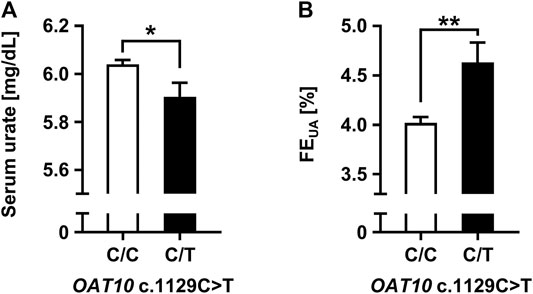
FIGURE 1. Increased renal urate excretion associated with genetic dysfunction of OAT10. Effects of genetic dysfunction of OAT10 on the renal urate handling in humans; serum urate levels (A), the values of fractional excretion of uric acid (FEUA) (B). Data are expressed as the mean ± SEM. *p < 0.05; **p < 0.01 (univariate linear regression analysis).
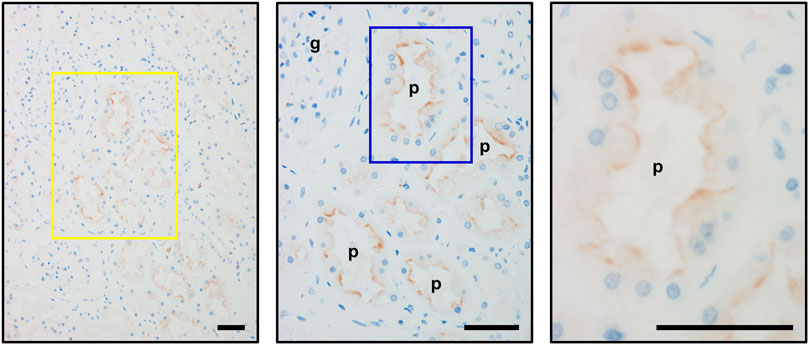
FIGURE 2. Immunohistochemical detection of OAT10 on the luminal membranes of the renal proximal tubules. Representative immunohistochemical micrographs of fixed sections of the human kidney, indicating the luminal expression of OAT10 in renal proximal tubular cells. Middle and right panels, the magnified image of the representative area outlined by a yellow and a blue rectangle, respectively. Bars, 50 μm; g, glomerulus; p, proximal tubule.
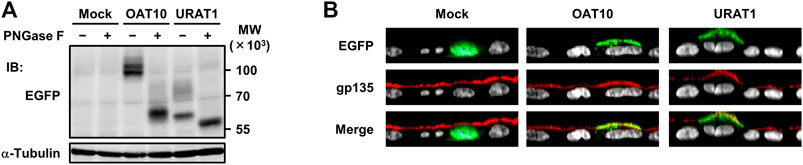
FIGURE 3. Expression and apical localization of OAT10 in polarized MDCKII cells. Immunoblot detection of OAT10 and URAT1 in whole cell lysate samples prepared from MDCKII cells 72 h after transfection (A). OAT10 or URAT1 fused with EGFP was detected using an anti-EGFP antibody. PNGase F sensitive signals indicate that the target proteins were maturated as glycoproteins; α-tubulin, loading control. Representative confocal microscopic Z-sectioning images (B). An endogenous apical membrane marker gp135 was immunostained using an anti-gp135 antibody (red). Nuclei were stained with TO-PRO-3 iodide (gray). Mock (EGFP), a control vector.
Molecular Properties of OAT10 as a Urate Transporter
To gain insight into the molecular properties of OAT10 as a urate transporter, we performed in vitro functional analyses using transiently OAT10-expressing HEK-derived 293A cells. The measured amount of radiolabeled urate incorporated into the cells revealed that OAT10 was more active at lower pH, mimicking urine conditions (Figure 4A). A similar result was obtained in URAT1 (Supplementary Figure S1). On the other hand, in contrast to the case in URAT1, OAT10-mediated urate transport was remarkably decreased by excluding Cl− from the Ringer solution (Figure 4B); the calculated OAT10-mediated urate transport activities in the Cl−-free condition were 6.1 ± 4.5% of those in the normal Ringer solution. This chloride-dependency is consistent with previous results obtained in an oocyte study (Mandal et al., 2017). Moreover, from time-course experiments we used uptake at 1 min to determine the initial rate of urate uptake by OAT10 (Figure 4C). Hence, uptake for 1 min at pH 6.4—a condition appropriate for the evaluation of OAT10-mediated urate transport—was examined in subsequent analyses. Under the experimentally maximum urate concentration (500 μM, which was due to the limited solubility of uric acid in the transport buffer), OAT10-mediated urate uptake was not completely saturated; thus, the estimated Km and Vmax for urate were 558 μM and 518 pmol/min per mg protein, respectively (Figure 4D). Considering that the Km value of OAT10 was the second lowest among the already characterized human urate transporters working as importers next to URAT1 (Km for urate, 371 μM) (Enomoto et al., 2002) (Table 3), OAT10 may have a role as a parallel backup of URAT1.
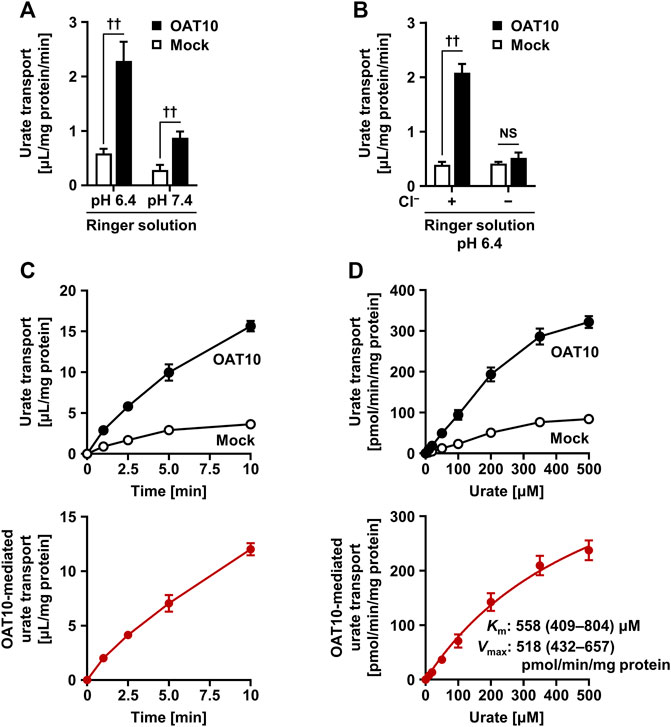
FIGURE 4. Characterization of molecular properties of OAT10 as a urate transporter. Transiently OAT10-expressing 293A cells were subjected to a cell-based urate transport assay, 48 h after transfection. Acidic pH (A), external Cl− (B), time (C), and concentration (D) dependent urate transport activities of OAT10 in Ringer solution (pH 6.4) containing 10 μM [8–14C]-urate (unless otherwise indicated). To prepare Cl−-free Ringer solution, all chloride salts in Ringer solution were replaced with each corresponding gluconate salt (B). OAT10-mediated urate transport [lower panels: (C,D)] was calculated by subtracting the urate transport activity in mock cells from that in OAT10-expressing cells [upper panels: (C,D)]. Data are expressed as the mean ± SD; n = 3 (A), 4 (B–D). ††p < 0.01; NS, not significantly different between groups (two-sided t-test). With the estimated Michaelis-Menten constant (Km) and maximal velocity (Vmax), the values of 95% confidence interval were in parentheses.
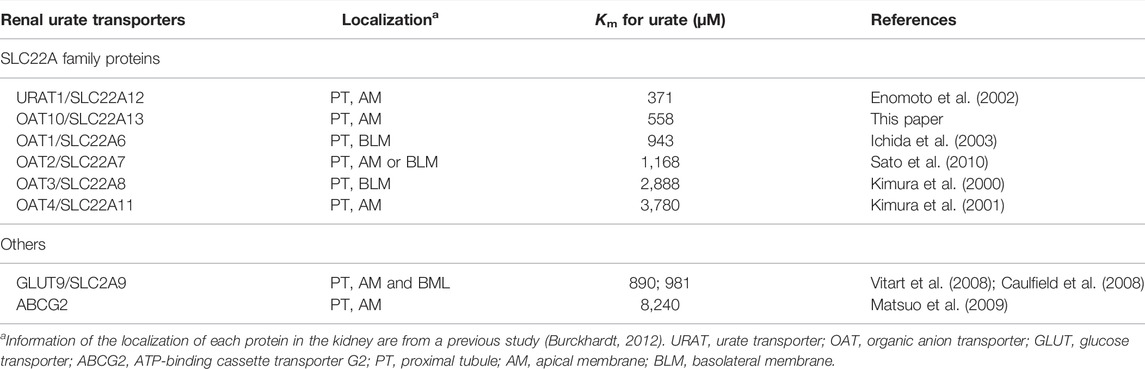
TABLE 3. Experimentally determined values of Michaelis-Menten constant (Km) for urate of human urate transporters expressed in the kidney.
Effects of Urate-Lowering Drugs and Their Reactive Metabolites on OAT10-Mediated Urate Transport
Although some drugs used for urate-lowering therapy inhibit URAT1 function, their effects on OAT10 remain unclear. Thus, we investigated the OAT10-inhibitory activities of urate-lowering drugs and their active metabolites (total 11), including urate synthesis inhibitors, uricosuric agents, fenofibrate, and losartan, five of which were compared with those against URAT1 (Figure 5).
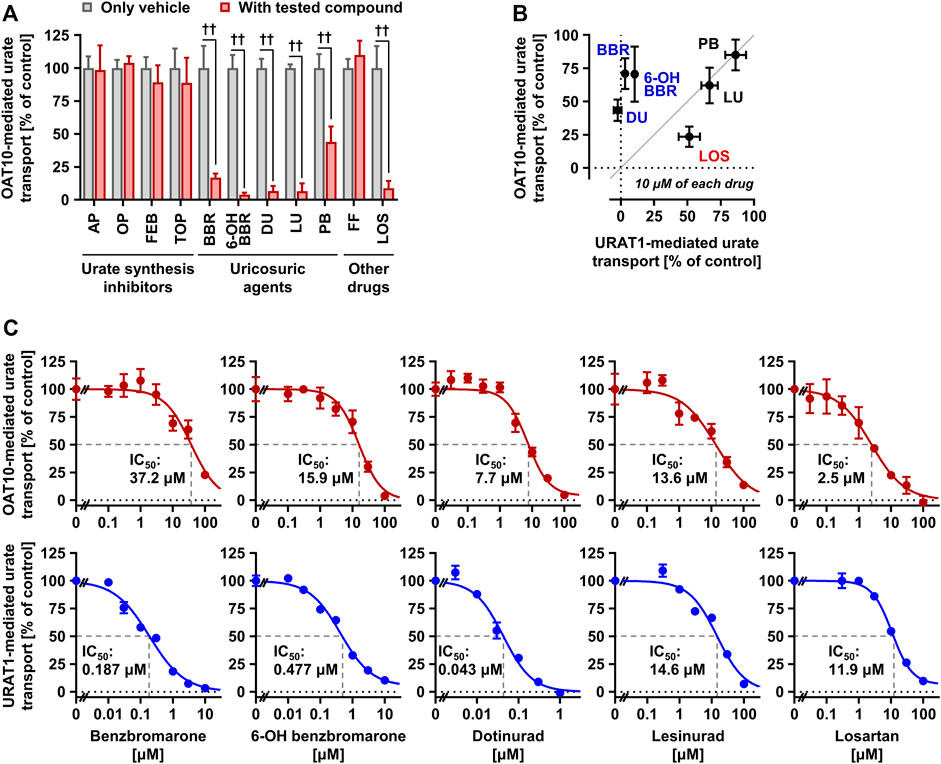
FIGURE 5. Effects of urate-lowering drugs and their reactive metabolites on the urate transport activity of OAT10. (A) OAT10-mediated urate transport activities in the presence or absence of urate-lowering drugs, including their reactive metabolites, at 100 μM or experimentally maximum concentrations (Table 2). AP, allopurinol; OP, oxypurinol; FEB, febuxostat; TOP, topiroxostat; BBR, benzbromarone; 6-OH BBR, 6-hydroxybenzbromarone; DU, dotinurad; LU, lesinurad; PB, probenecid; FF, fenofibrate; LOS, losartan. (B) Comparison of the effects of each compound (10 μM) on the urate transport activities of URAT1 and OAT10. Values are shown as % of vehicle control (only 0.1% dimethyl sulfoxide); data are expressed as the mean ± SD; n = 3–4 (A,B). ††p < 0.01 (two-sided t-test). (C) Concentration-dependent inhibition of OAT10-mediated urate transport by BBR, 6-OH BBR, DU, LU, and LOS. The urate transport activities were measured in the presence of each compound at indicated concentrations for 1 min. Values are shown as % of vehicle control; data are expressed as the mean ± SEM; n = 4.
First, we conducted inhibitory tests in the presence of each authentic compound at the maximum soluble levels in the transport buffer (Table 2). The results revealed that, contrary to all the urate synthesis inhibitors and fenofibrate, five compounds—benzbromarone, 6-hydroxybenzbromarone (an active metabolite of benzbromarone), dotinurad, lesinurad, and losartan—decreased the urate transport activity of OAT10 to < 20% that of the vehicle control (Figure 5A). Next, we compared their effects at 10 μM on the urate transport activity of OAT10 and URAT1 (Figure 5B), which demonstrated that i) benzbromarone, 6-hydroxybenzbromarone, and dotinurad were more specific to URAT1 than OAT10; ii) lesinurad inhibited OAT10 and URAT1 comparably; iii) losartan exhibited a stronger inhibitory effect on OAT10 than URAT1. These results were confirmed by further investigations of concentration-dependent inhibitory effects and calculated IC50 values (Figure 5C). Given the proposed mechanism, lesinurad (Miner et al., 2016) and losartan (Lee and Terkeltaub, 2006) do and can inhibit URAT1 in the kidney, respectively; hence, they may also act as renal OAT10 inhibitors in clinical situations (Figure 6). Of note, given that lesinurad reportedly inhibits not only URAT1 but also other transporters such as OATP1B1, OCT1, OAT1 and OAT3 in vitro (Garg et al., 2018), lesinurad is not necessarily highly-specific to URAT1. Thus, the OAT10-inhibitory effect of lesinurad we identified here is not inconsistent with previous findings.
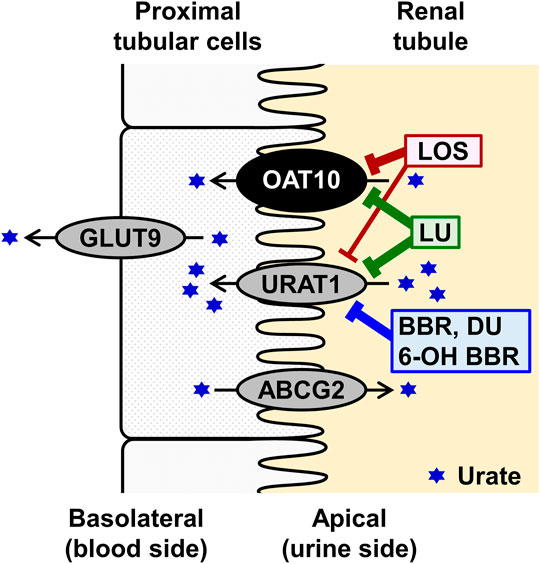
FIGURE 6. Schematic illustration of OAT10 function as a renal urate re-absorber. The results of interaction with uricosuric compounds and their abbreviations are shown in Figure 5 and its legend, respectively.
Discussion
In this study, we investigated the effect of OAT10 c.1129C>T on renal urate handling in a Japanese male population. The results, including immunohistochemical analyses, demonstrated that the dysfunction of OAT10 leads to an increase in the net activities of both renal urate secretion and a decrease in serum urate levels (Figure 1), which is consistent with our working hypothesis. With the urate-lowering effect of OAT10 c.1129C>T, β (coefficient for the effect allele, 1129T) was −0.135 mg/dl/allele (Supplementary Table S2). Regarding ABCG2 c.421C>A (p.Q141K) that is a well-established gout/hyperuricemia risk allele in the physiologically important urate transporter (Matsuo et al., 2009; Woodward et al., 2009), the calculated β for serum urate in the Japanese males (carriers with ABCG2 421C/A vs. 421C/C) was 0.20 mg/dl/allele in a previous report (Matsuo et al., 2009). Given these pieces of information, together with the protective effect of OAT10 c.1129C>T on gout susceptibility (Higashino et al., 2020), our data support the physiological importance of OAT10 as a urate transporter in the kidney, although the β of OAT10 was not so large compared with cases of causal genes for renal hypouricemia (which means that OAT10 has a smaller contribution to renal urate re-uptake than URAT1 and GLUT9). Moreover, functional analyses revealed that OAT10 has a high affinity for urate transport following that of URAT1 (Figure 4; Table 3); we identified lesinurad and losartan as clinically available potential inhibitors of OAT10 (Figure 5). These findings deepen our understanding of the physiological and pharmacological impact of OAT10.
While the function of OAT10 as a urate transporter was reported previously (Bahn et al., 2008; Higashino et al., 2020), the affinity of OAT10 to urate has been poorly studied. With this matter, our kinetic analyses revealed that OAT10 has relatively strong affinity for urate [Km (95% confidence interval), 558 (409–804) μM] among already-characterized urate transporters in humans (Table 3). A previous in vitro study using Xenopus oocytes showed that urate inhibited OAT10-mediated nicotinate transport in a concentration dependent manner with the IC50 value of 759 ± 501 μM (Bahn et al., 2008) which was the only published data evaluating the concentration-dependent effect of urate on the OAT10 function. Given these similar values, although little information is available, urate and nicotinate might share an internal tunnel for membrane transport in OAT10 protein. Additionally, based on reported Km values for nicotinate of OAT10 (<100 μM, lower than the Km value for urate we determined) (Bahn et al., 2008), OAT10 has a higher affinity for nicotinate than urate. Nicotinate (pyridine-3-carboxylate) is smaller than urate (a purine compound), which may make it easier to interact with OAT10.
Considering the protective effect of OAT10 dysfunction on gout susceptibility (OR = 0.67) (Higashino et al., 2020), in the context of the potentially simultaneous inhibition of renal URAT1 and OAT10 (Figure 6), it would be interesting to determine whether lesinurad, which had been an approved uricosuric agent that was recently withdrawn from the market unfortunately because of low market penetration, not because of therapeutic issues (Jansen et al., 2022), could have an anti-gout effect stronger than other uricosuric agents that inhibit only URAT1 in clinical situations. It is possible that the positive effects of OAT10 dysfunction may depend on OAT10 expressed not only in the kidney but also in other tissues; such a notion should be addressed in future clinical studies.
Regarding losartan, we found that its IC50 value against OAT10 (2.5 μM) is smaller than that against URAT1 (11.9 μM) (Figure 5). This relationship means that losartan would have a higher affinity to OAT10 than to URAT1; when losartan exists at concentrations enough for URAT1 inhibition, it also inhibits OAT10 in theory. In this context, losartan may have a stronger inhibitory activity towards OAT10 than URAT1 in clinical situations given the similarity of their renal expression as apical transporters (Figure 6), suggesting that the uricosuric effect of losartan may involve the inhibition of both OAT10 and URAT1. The beneficial effect of this angiotensin II receptor blocker, generally used as an antihypertensive drug, has been historically explained by the inhibition of URAT1, but not that of OAT10. Therefore, the stronger inhibitory effect of losartan on OAT10 than URAT1 extends our knowledge of the molecular basis underlying losartan-related uricosuric action.
In contrast to urate synthesis inhibitors, uricosuric agents tended to inhibit OAT10. Regarding the inhibitory potency of benzbromarone, our results are consistent with previous reports wherein high-dose benzbromarone inhibited OAT10 function using a Xenopus oocyte system (Bahn et al., 2008; Mandal et al., 2017). However, benzbromarone, 6-hydroxybenzbromarone, and dotinurad, considering already-characterized pharmacokinetic information including unbound plasma maximum concentrations (Supplementary Table S6) (Miyata et al., 2016; Taniguchi et al., 2019) and urine levels, would be selective inhibitors of URAT1 in clinical situations. Hence, combination therapy or improvement of chemical structures to enhance the OAT10 inhibitory activity of such urate-lowering drugs may be a potential strategy to provide a more effective anti-gout (urate-lowering) therapy.
The limitations of this study include the lack of pharmacological evaluations in human bodies regarding the effects of OAT10 inhibition by potentially influential drugs that we proposed above. It will be interesting to examine whether existing drugs that unexpectedly affect serum urate levels can interact with OAT10 or not. Moreover, whereas this OAT10 variant (c.1129C>T; p.R377C) is common in the Japanese, it is very rare in other populations (Higashino et al., 2020); this should be a reason why previous single nucleotide polymorphism-based approaches (i.e., genome-wide association studies) including meta-analyses (Major et al., 2018; Tin et al., 2019) could not find OAT10 as a gene associated with serum urate levels. Therefore, when other common and/or rare dysfunctional variants of OAT10 are found in other populations, further replication studies on OAT10-mediated renal urate handling in humans will be required. Last, as this study is restricted in males, it will be a next issue to investigate whether there could be gender differences in the effects of genetic dysfunction of OAT10 on serum urate levels and renal urate handling. With this matter, a previous study proposed the presence of a gender difference in the renal expression of OAT10 based on immunoblotting data showing stronger signal in a female rat than in a male rat while the data lacked a loading control for the comparison of rat Oat10 protein levels (Bahn et al., 2008). Of note, there are reportedly intersexual differences in urate handling—age-fertile women show lower and higher levels of serum urate and renal urate excretion than men of similar ages, respectively (Mateos Antón et al., 1986; Stiburkova and Bleyer, 2012); however, post-menopausal women exhibit higher serum urate levels than pre-menopausal women (Wingrove et al., 1998; Hak and Choi, 2008). Based on these complicacies, the potential gender differences should be carefully evaluated in the near future.
Before closing, some future directions of this study are warrant mention. First, given the luminal expression of OAT10 in the renal proximal tubules (Figure 2), detailed histological studies investigating the segmental distribution of OAT10 will be a next step because renal proximal tubules are separated into S1–S3 segments that have reportedly different roles in renal urate handling (Maesaka and Fishbane, 1998). According to a recent transcriptome study that examined renal tubule segments microdissected from murine kidneys (Chen et al., 2021), Oat10 was highly expressed in the S2 and S3 segments than other renal tubule segments; a similar expression pattern was observed in Urat1. To deepen our understanding, confirmation of segmental distribution of each protein in the kidney of mice and humans will be required. Second, addressing how OAT10 and URAT1 can physiologically cooperate/interact with each other will be a future topic. Given their reverse chloride-dependency, when URAT1 function could be cis-inhibited by the (locally) elevated urine Cl−, OAT10-mediated urate transport might be enhanced, conversely. Additionally, in patients with renal hypouricemia, extremely high FEUA (≥100%) is typical in renal hypouricemia type 2 (characterized by homozygous of non-functional GLUT9) but not in renal hypouricemia type 1 (characterized by homozygous of non-functional URAT1; typical FEUA, tens of %) (Kawamura et al., 2021). This difference can support the presence of a latent alternative that contribute to renal urate re-uptake in the absence of URAT1 function. This context suggests that OAT10 may be a compensator of URAT1 in certain conditions.
In conclusion, we revealed that the dysfunction of OAT10 enhances renal urate excretion evidenced by both the increase in FEUA and the decrease in serum urate. This means that OAT10 acts as a renal urate re-absorber which could be a molecular target for urate-lowering therapy. Our findings provide a better understanding of renal urate handling systems, paving the way for more effective therapeutic strategies for urate-related diseases including gout and hyperuricemia as modulating renal functions is an important approach for the treatment of life-style related diseases.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding authors.
Ethics Statement
The studies involving human participants were reviewed and approved by the Institutional Ethics Committees of the National Defense Medical College and Nagoya University. The participants provided their written informed consent to participate in this study.
Author Contributions
Conceptualization, YTo, TT (14th Author), and HM; validation, YTo, YK (2nd Author), and AN; formal analysis, YTo, YK (2nd Author), and AN; investigation, YTo, YK (2nd Author), AN, SS, YTa, TT (7th Author), TK, YK (9th Author), YK (13th Author), and HM: i.e., YTo was responsible for biochemical and functional analyses, and analyzed the data; YK (2nd Author), AN, SS, YTa, TT (7th Author), TK, YK (9th Author), and HM collected and analyzed the participants’ clinical data and YTo, YK (2nd Author), AN, and HM are responsible for genetic and statistical analyses; YK (13th Author) and HM organized immunohistochemical analyses; resources, YTo, KM, TT (7th Author), TK, and YK (9th Author); data curation, YTo, YK (2nd Author), and AN; writing—original draft preparation, YTo; writing—review and editing, YTo, AN, TT (14th Author), and HM; visualization, YTo and YK (2nd Author); supervision, KI, HS, NS, and YK(13th Author); project administration, YTo, TT (14th Author), and HM; funding acquisition, YTo, AN, TT (14th Author), and HM; TT (14th Author) and HM organized this collaborative study as corresponding authors; all authors read and approved the final manuscript.
Funding
This study was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI (grant numbers 19K16441 and 21H03350 to YTo; 20K23152 to AN; 16H01808, 18KK0247, and 20H00568 to TT (14th Author); 17H04128, 19K22786, 20H00566, and 21KK0173 to HM; 17015018, 221S0001, 16H06277 (CoBiA), and 16H06279) as well as the Ministry of Defense of Japan. TT (14th Author) and HM have received research grants from “Gout and uric acid foundation of Japan”. TT (14th Author) has received research grants from “Takeda Science Foundation,” “Suzuken Memorial Foundation,” “Mochida Memorial Foundation for Medical and Pharmaceutical Research,” and “The Pharmacological Research Foundation, Tokyo.”
Conflict of Interest
Dotinurad was kindly provided from Fuji Yakuhin to TT (14th Author) under a material transfer agreement. KI and TT (14th Author) received fees for advisory boards and speaking fees from Mochida Pharmaceutical and Fuji Yakuhin. KI also received speaking fees from Teijin Pharma. HM received speaking fees from Mochida Pharmaceutical, Fuji Yakuhin, Teijin Pharma, and Sanwa Kagaku Kenkyusho. All the companies had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Acknowledgments
We would like to express our appreciation to all the participants for their generous involvement in this study. Our heartfelt gratitude goes to the members of the Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study) for their support. The authors also specially acknowledge N. Hamajima, H. Tanaka, and K. Wakai for sample collection and their continuous encouragement of our study. Moreover, the authors appreciate Fuji Yakuhin (Saitama, Japan) for providing dotinurad. We also thank K. Morichika, M. Miyazawa, and M. Seki (National Defense Medical College) for genetic analyses, and M. Watanabe (National Defense Medical College) for immunohistochemical analyses. YTo is an Excellent Young Researcher in the MEXT Leading Initiative for Excellent Young Researchers.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fphar.2022.842717/full#supplementary-material
Abbreviations
ABCG2, ATP-binding cassette transporter G2; BSA, bovine serum albumin; DMSO, dimethyl sulfoxide; FEUA, fractional excretion of uric acid; GLUT9, glucose transporter 9; HBSS, Hanks’ Balanced Salt Solution; IC50, half maximal inhibitory concentration; MDCKII, Madin-Darby canine kidney II; Vmax, maximal velocity; Km, Michaelis-Menten constant; OR, odds ratio; OAT10, organic anion transporter 10; QTL, quantitative trait locus; URAT1, urate transporter 1.
References
Anzai, N., Miyazaki, H., Noshiro, R., Khamdang, S., Chairoungdua, A., Shin, H. J., et al. (2004). The Multivalent PDZ Domain-Containing Protein PDZK1 Regulates Transport Activity of Renal Urate-Anion Exchanger URAT1 via its C Terminus. J. Biol. Chem. 279 (44), 45942–45950. doi:10.1074/jbc.M406724200
Asai, Y., Naito, M., Suzuki, M., Tomoda, A., Kuwabara, M., Fukada, Y., et al. (2009). Baseline Data of Shizuoka Area in the Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study). Nagoya J. Med. Sci. 71 (3-4), 137–144.
Bahn, A., Hagos, Y., Reuter, S., Balen, D., Brzica, H., Krick, W., et al. (2008). Identification of a New Urate and High Affinity Nicotinate Transporter, hOAT10 (SLC22A13). J. Biol. Chem. 283 (24), 16332–16341. doi:10.1074/jbc.M800737200
Bao, R., Liu, M., Wang, D., Wen, S., Yu, H., Zhong, Y., et al. (2019). Effect of Eurycoma Longifolia Stem Extract on Uric Acid Excretion in Hyperuricemia Mice. Front. Pharmacol. 10, 1464. doi:10.3389/fphar.2019.01464
Burckhardt, G. (2012). Drug Transport by Organic Anion Transporters (OATs). Pharmacol. Ther. 136 (1), 106–130. doi:10.1016/j.pharmthera.2012.07.010
Bursill, D., Taylor, W. J., Terkeltaub, R., Abhishek, A., So, A. K., Vargas-Santos, A. B., et al. (2019). Gout, Hyperuricaemia and Crystal-Associated Disease Network (G-CAN) Consensus Statement Regarding Labels and Definitions of Disease States of Gout. Ann. Rheum. Dis. 78 (11), 1592–1600. doi:10.1136/annrheumdis-2019-215933
Caulfield, M. J., Munroe, P. B., O'Neill, D., Witkowska, K., Charchar, F. J., Doblado, M., et al. (2008). SLC2A9 Is a High-Capacity Urate Transporter in Humans. Plos Med. 5 (10), e197. doi:10.1371/journal.pmed.0050197
Chen, L., Chou, C.-L., and Knepper, M. A. (2021). A Comprehensive Map of mRNAs and Their Isoforms across All 14 Renal Tubule Segments of Mouse. J. Am. Soc. Nephrol. 32 (4), 897–912. doi:10.1681/ASN.2020101406
Chiba, T., Matsuo, H., Kawamura, Y., Nagamori, S., Nishiyama, T., Wei, L., et al. (2015). NPT1/SLC17A1 Is a Renal Urate Exporter in Humans and its Common Gain-of-Function Variant Decreases the Risk of Renal Underexcretion Gout. Arthritis Rheumatol. 67 (1), 281–287. doi:10.1002/art.38884
Dalbeth, N., Choi, H. K., Joosten, L. A. B., Khanna, P. P., Matsuo, H., Perez-Ruiz, F., et al. (2019). Gout. Nat. Rev. Dis. Primers 5 (1), 69. doi:10.1038/s41572-019-0115-y
Dinour, D., Gray, N. K., Campbell, S., Shu, X., Sawyer, L., Richardson, W., et al. (2010). Homozygous SLC2A9 Mutations Cause Severe Renal Hypouricemia. J. Am. Soc. Nephrol. 21 (1), 64–72. doi:10.1681/ASN.2009040406
Enomoto, A., Kimura, H., Chairoungdua, A., Shigeta, Y., Jutabha, P., Cha, S. H., et al. (2002). Molecular Identification of a Renal Urate Anion Exchanger that Regulates Blood Urate Levels. Nature 417 (6887), 447–452. doi:10.1038/nature742
Garg, Y., Gawri, V., Gore, R., Garg, R. S., and Kumar, A. (2018). Lesinurad, a Novel Uricosuric Drug for Allopurinol-Refractory Gout Patients. J. Clin. Diagn. Res. 12 (2), FE01–FE05. doi:10.7860/jcdr/2018/27997.11149
Gutman, A. B., Yu, T. F., and Randolph, V. (1954). Further Observation on the Uricosuric Effects of Probenecid (Benemid) in Tophaceous Gout. Trans. Assoc. Am. Physicians 67, 250–260.
Hak, A. E., and Choi, H. K. (2008). Menopause, Postmenopausal Hormone Use and Serum Uric Acid Levels in US Women-Tthe Third National Health and Nutrition Examination Survey. Arthritis Res. Ther. 10 (5), R116. doi:10.1186/ar2519
Halperin Kuhns, V. L., and Woodward, O. M. (2021). Urate Transport in Health and Disease. Best Pract. Res. Clin. Rheumatol. 35 (4), 101717. doi:10.1016/j.berh.2021.101717
Hamajima, N. and J-MICC Study Group (2007). The Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study) to Detect Gene-Environment Interactions for Cancer. Asian Pac. J. Cancer Prev. 8 (2), 317–323.
Harvengt, C., Heller, F., and Desager, J. P. (1980). Hypolipidemic and Hypouricemic Action of Fenofibrate in Various Types of Hyperlipoproteinemias. Artery 7 (1), 73–82.
Higashino, T., Morimoto, K., Nakaoka, H., Toyoda, Y., Kawamura, Y., Shimizu, S., et al. (2020). Dysfunctional Missense Variant of OAT10/SLC22A13 Decreases Gout Risk and Serum Uric Acid Levels. Ann. Rheum. Dis. 79 (1), 164–166. doi:10.1136/annrheumdis-2019-216044
Hosoyamada, M., Takiue, Y., Morisaki, H., Cheng, J., Ikawa, M., Okabe, M., et al. (2010). Establishment and Analysis of SLC22A12 (URAT1) Knockout Mouse. Nucleosides Nucleotides Nucleic Acids 29 (4-6), 314–320. doi:10.1080/15257771003738634
Hosoyamada, M., Tsurumi, Y., Hirano, H., Tomioka, N. H., Sekine, Y., Morisaki, T., et al. (2016). Urat1-Uox Double Knockout Mice Are Experimental Animal Models of Renal Hypouricemia and Exercise-Induced Acute Kidney Injury. Nucleosides Nucleotides Nucleic Acids 35 (10-12), 543–549. doi:10.1080/15257770.2016.1143559
Ichida, K., Hosoyamada, M., Hisatome, I., Enomoto, A., Hikita, M., Endou, H., et al. (2004). Clinical and Molecular Analysis of Patients with Renal Hypouricemia in Japan-influence of URAT1 Gene on Urinary Urate Excretion. J. Am. Soc. Nephrol. 15 (1), 164–173. doi:10.1097/01.asn.0000105320.04395.d0
Ichida, K., Hosoyamada, M., Kimura, H., Takeda, M., Utsunomiya, Y., Hosoya, T., et al. (2003). Urate Transport via Human PAH Transporter hOAT1 and its Gene Structure. Kidney Int. 63 (1), 143–155. doi:10.1046/j.1523-1755.2003.00710.x
Ichida, K., Matsuo, H., Takada, T., Nakayama, A., Murakami, K., Shimizu, T., et al. (2012). Decreased Extra-renal Urate Excretion Is a Common Cause of Hyperuricemia. Nat. Commun. 3, 764. doi:10.1038/ncomms1756
Jansen, T. L., Tanja, G., and Matthijs, J. (2022). A Historical Journey of Searching for Uricosuric Drugs. Clin. Rheumatol. 41 (1), 297–305. doi:10.1007/s10067-021-05930-1
Kamper, A., and Nielsen, A. H. (2001). Uricosuric Effect of Losartan in Renal Transplanted Patients. Transpl. Proc. 33 (1-2), 1201. doi:10.1016/s0041-1345(00)02385-x
Kawamura, Y., Nakayama, A., Shimizu, S., Toyoda, Y., Nishida, Y., Hishida, A., et al. (2021). A Proposal for Practical Diagnosis of Renal Hypouricemia: Evidenced from Genetic Studies of Nonfunctional Variants of URAT1/SLC22A12 Among 30,685 Japanese Individuals. Biomedicines 9 (8), 1012. doi:10.3390/biomedicines9081012
Kimura, H., Ichida, K., Hosoyamada, M., Oono, I., Endou, H., and Hosoya, T. (2000). Analysis of Uric Acid Transport via hOAT3. Gout Nucleic Acid Metab. 24 (2), 115–121. doi:10.6032/gnam1999.24.2_115
Kimura, H., Ichida, K., Hosoyamada, M., Oono, I., Endou, H., and Hosoya, T. (2001). Urate Transport via hOAT4. Gout Nucleic Acid Metab. 25 (2), 113–120. doi:10.6032/gnam1999.25.2_113
Koepsell, H. (2013). The SLC22 Family with Transporters of Organic Cations, Anions and Zwitterions. Mol. Aspects Med. 34 (2-3), 413–435. doi:10.1016/j.mam.2012.10.010
Lee, S. J., and Terkeltaub, R. A. (2006). New Developments in Clinically Relevant Mechanisms and Treatment of Hyperuricemia. Curr. Rheumatol. Rep. 8 (3), 224–230. doi:10.1007/s11926-996-0029-z
Maesaka, J. K., and Fishbane, S. (1998). Regulation of Renal Urate Excretion: a Critical Review. Am. J. Kidney Dis. 32 (6), 917–933. doi:10.1016/s0272-6386(98)70067-8
Major, T. J., Dalbeth, N., Stahl, E. A., and Merriman, T. R. (2018). An Update on the Genetics of Hyperuricaemia and Gout. Nat. Rev. Rheumatol. 14 (6), 341–353. doi:10.1038/s41584-018-0004-x
Mandal, A. K., Mercado, A., Foster, A., Zandi-Nejad, K., and Mount, D. B. (2017). Uricosuric Targets of Tranilast. Pharmacol. Res. Perspect. 5 (2), e00291. doi:10.1002/prp2.291
Mateos Antón, F., García Puig, J., Ramos, T., González, P., and Ordás, J. (1986). Sex Differences in Uric Acid Metabolism in Adults: Evidence for a Lack of Influence of Estradiol-17β (E2) on the Renal Handling of Urate. Metabolism 35 (4), 343–348. doi:10.1016/0026-0495(86)90152-6
Matsuo, H., Chiba, T., Nagamori, S., Nakayama, A., Domoto, H., Phetdee, K., et al. (2008). Mutations in Glucose Transporter 9 Gene SLC2A9 Cause Renal Hypouricemia. Am. J. Hum. Genet. 83 (6), 744–751. doi:10.1016/j.ajhg.2008.11.001
Matsuo, H., Takada, T., Ichida, K., Nakamura, T., Nakayama, A., Ikebuchi, Y., et al. (2009). Common Defects of ABCG2, a High-Capacity Urate Exporter, Cause Gout: a Function-Based Genetic Analysis in a Japanese Population. Sci. Transl. Med. 1 (5), 5ra11. doi:10.1126/scitranslmed.3000237
Miner, J. N., Tan, P. K., Hyndman, D., Liu, S., Iverson, C., Nanavati, P., et al. (2016). Lesinurad, a Novel, Oral Compound for Gout, Acts to Decrease Serum Uric Acid through Inhibition of Urate Transporters in the Kidney. Arthritis Res. Ther. 18 (1), 214. doi:10.1186/s13075-016-1107-x
Miyata, H., Takada, T., Toyoda, Y., Matsuo, H., Ichida, K., and Suzuki, H. (2016). Identification of Febuxostat as a New Strong ABCG2 Inhibitor: Potential Applications and Risks in Clinical Situations. Front. Pharmacol. 7, 518. doi:10.3389/fphar.2016.00518
Nakatochi, M., Toyoda, Y., Kanai, M., Nakayama, A., Kawamura, Y., Hishida, A., et al. (2021). An X Chromosome-wide Meta-Analysis Based on Japanese Cohorts Revealed that Non-autosomal Variations Are Associated with Serum Urate. Rheumatology (Oxford) 60 (9), 4430–4432. doi:10.1093/rheumatology/keab404
Nakayama, A., Matsuo, H., Ohtahara, A., Ogino, K., Hakoda, M., Hamada, T., et al. (2019). Clinical Practice Guideline for Renal Hypouricemia (1st Edition). Hum. Cell 32 (2), 83–87. doi:10.1007/s13577-019-00239-3
Oikawa, T., Kunishima, C., Matsumoto, K., Ishikawa, S., and Endou, H. (2005). The Inhibitory Effect of Benzbromarone and 6-hydroxybenzbromarone on Urate Transporter (URAT1). J. New Rem. Clin. 54 (6), 15–20.
Okamoto, K., Eger, B. T., Nishino, T., Kondo, S., Pai, E. F., and Nishino, T. (2003). An Extremely Potent Inhibitor of Xanthine Oxidoreductase. Crystal Structure of the Enzyme-Inhibitor Complex and Mechanism of Inhibition. J. Biol. Chem. 278 (3), 1848–1855. doi:10.1074/jbc.M208307200
Osada, Y., Tsuchimoto, M., Fukushima, H., Takahashi, K., Kondo, S., Hasegawa, M., et al. (1993). Hypouricemic Effect of the Novel Xanthine Oxidase Inhibitor, TEI-6720, in Rodents. Eur. J. Pharmacol. 241 (2-3), 183–188. doi:10.1016/0014-2999(93)90201-r
Sakiyama, M., Matsuo, H., Toyoda, Y., Yonekura, Y., Ishikawa, T., Nakayama, A., et al. (2021). Porphyrin Accumulation in Humans with Common Dysfunctional Variants of ABCG2, a Porphyrin Transporter: Potential Association with Acquired Photosensitivity. Hum. Cell 34 (4), 1082–1086. doi:10.1007/s13577-021-00534-y
Sato, M., Mamada, H., Anzai, N., Shirasaka, Y., Nakanishi, T., and Tamai, I. (2010). Renal Secretion of Uric Acid by Organic Anion Transporter 2 (OAT2/SLC22A7) in Human. Biol. Pharm. Bull. 33 (3), 498–503. doi:10.1248/bpb.33.498
Stiburkova, B., and Bleyer, A. J. (2012). Changes in Serum Urate and Urate Excretion with Age. Adv. Chronic Kidney Dis. 19 (6), 372–376. doi:10.1053/j.ackd.2012.07.010
Takada, T., Yamanashi, Y., Konishi, K., Yamamoto, T., Toyoda, Y., Masuo, Y., et al. (2015). NPC1L1 Is a Key Regulator of Intestinal Vitamin K Absorption and a Modulator of Warfarin Therapy. Sci. Transl. Med. 7 (275), 275ra23. doi:10.1126/scitranslmed.3010329
Taniguchi, T., Ashizawa, N., Matsumoto, K., Saito, R., Motoki, K., Sakai, M., et al. (2019). Pharmacological Evaluation of Dotinurad, a Selective Urate Reabsorption Inhibitor. J. Pharmacol. Exp. Ther. 371 (1), 162–170. doi:10.1124/jpet.119.259341
Tin, A., Marten, J., Halperin Kuhns, V. L., Li, Y., Wuttke, M., Kirsten, H., et al. (2019). Target Genes, Variants, Tissues and Transcriptional Pathways Influencing Human Serum Urate Levels. Nat. Genet. 51 (10), 1459–1474. doi:10.1038/s41588-019-0504-x
Toyoda, Y., Takada, T., Miyata, H., Ishikawa, T., and Suzuki, H. (2016). Regulation of the Axillary Osmidrosis-Associated ABCC11 Protein Stability by N-Linked Glycosylation: Effect of Glucose Condition. PLoS One 11 (6), e0157172. doi:10.1371/journal.pone.0157172
Toyoda, Y., Takada, T., Miyata, H., Matsuo, H., Kassai, H., Nakao, K., et al. (2020). Identification of GLUT12/SLC2A12 as a Urate Transporter that Regulates the Blood Urate Level in Hyperuricemia Model Mice. Proc. Natl. Acad. Sci. U S A. 117 (31), 18175–18177. doi:10.1073/pnas.2006958117
Toyoda, Y., Takada, T., Saito, H., Hirata, H., Ota-Kontani, A., Kobayashi, N., et al. (2020). Inhibitory Effect of Citrus Flavonoids on the In Vitro Transport Activity of Human Urate Transporter 1 (URAT1/SLC22A12), a Renal Re-absorber of Urate. NPJ Sci. Food 4, 3. doi:10.1038/s41538-020-0063-7
Uhlén, M., Fagerberg, L., Hallström, B. M., Lindskog, C., Oksvold, P., Mardinoglu, A., et al. (2015). Proteomics. Tissue-Based Map of the Human Proteome. Science 347 (6220), 1260419. doi:10.1126/science.1260419
Vitart, V., Rudan, I., Hayward, C., Gray, N. K., Floyd, J., Palmer, C. N., et al. (2008). SLC2A9 Is a Newly Identified Urate Transporter Influencing Serum Urate Concentration, Urate Excretion and Gout. Nat. Genet. 40 (4), 437–442. doi:10.1038/ng.106
Watts, R. W., Watts, J. E., and Seegmiller, J. E. (1965). Xanthine Oxidase Activity in Human Tissues and its Inhibition by Allopurinol (4-Hydroxypyrazolo[3,4-D] Pyrimidine). J. Lab. Clin. Med. 66 (4), 688–697. doi:10.5555/uri:pii:0022214365900533
Wingrove, C. S., Walton, C., and Stevenson, J. C. (1998). The Effect of Menopause on Serum Uric Acid Levels in Non-obese Healthy Women. Metabolism 47 (4), 435–438. doi:10.1016/s0026-0495(98)90056-7
Keywords: uricosuric agent, urate reabsorption inhibitor, urate-lowering therapy, losartan, renal urate handling, lesinurad, urate transporter
Citation: Toyoda Y, Kawamura Y, Nakayama A, Morimoto K, Shimizu S, Tanahashi Y, Tamura T, Kondo T, Kato Y, Ichida K, Suzuki H, Shinomiya N, Kobayashi Y, Takada T and Matsuo H (2022) OAT10/SLC22A13 Acts as a Renal Urate Re-Absorber: Clinico-Genetic and Functional Analyses With Pharmacological Impacts. Front. Pharmacol. 13:842717. doi: 10.3389/fphar.2022.842717
Received: 24 December 2021; Accepted: 09 February 2022;
Published: 06 April 2022.
Edited by:
Francisca Sivera, Hospital General de Elda, SpainReviewed by:
David Mount, Brigham and Women’s Hospital and Harvard Medical School, United StatesMariano Andres, Miguel Hernández University of Elche, Spain
Copyright © 2022 Toyoda, Kawamura, Nakayama, Morimoto, Shimizu, Tanahashi, Tamura, Kondo, Kato, Ichida, Suzuki, Shinomiya, Kobayashi, Takada and Matsuo. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Tappei Takada, dGFwcGVpLXRreUB1bWluLmFjLmpw; Hirotaka Matsuo, aG1hdHN1by5uZG1jQGdtYWlsLmNvbQ==
†These authors have contributed equally to this work
 Yu Toyoda
Yu Toyoda Yusuke Kawamura
Yusuke Kawamura Akiyoshi Nakayama
Akiyoshi Nakayama Keito Morimoto1
Keito Morimoto1 Kimiyoshi Ichida
Kimiyoshi Ichida Nariyoshi Shinomiya
Nariyoshi Shinomiya Yasushi Kobayashi
Yasushi Kobayashi Tappei Takada
Tappei Takada Hirotaka Matsuo
Hirotaka Matsuo