by Booth TC, Grzeda M, Chelliah A, Roman A, Al Busaidi A, Dragos C, Shuaib H, Luis A, Mirchandani A, Alparslan B, Mansoor N, Lavrador J, Vergani F, Ashkan K, Modat M and Ourselin S (2022) Front. Oncol. 12:799662. doi: 10.3389/fonc.2022.799662
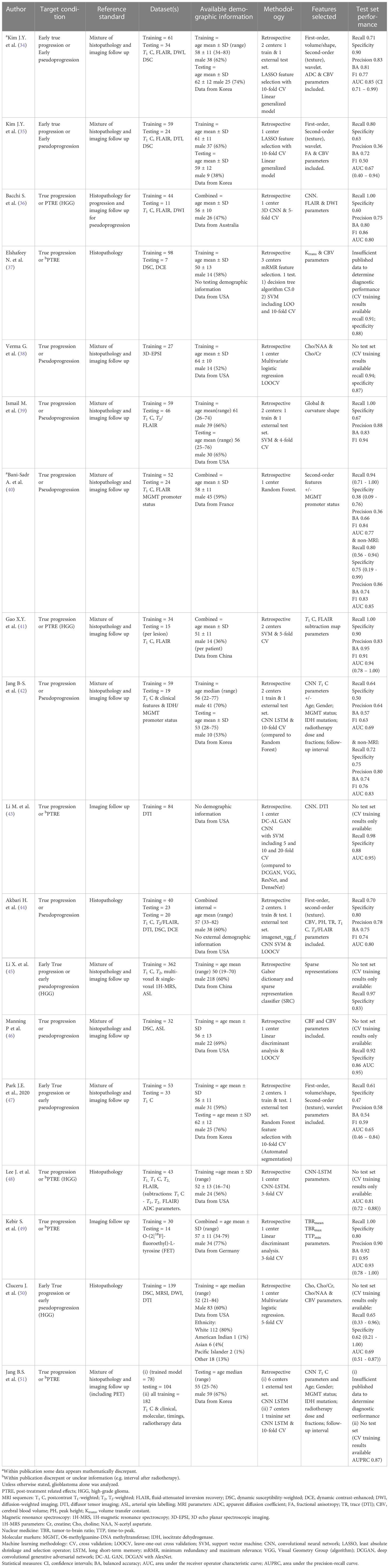
Due to a production error, there was an error in the published Table 1. The 4th row of the table started at the second column instead of the first column, causing the contents of the last column to move to the next row, resulting in a formatting error. The corrected Table 1 appears below.
The publisher apologizes for this error. The original version of this article has been updated.
Keywords: glioblastoma, machine learning, monitoring biomarkers, meta-analysis, artificial intelligence, treatment response, deep learning, glioma
Citation: Frontiers Production Office (2023) Erratum: Imaging biomarkers of glioblastoma treatment response: a systematic review and meta-analysis of recent machine learning studies. Front. Oncol. 13:1217461. doi: 10.3389/fonc.2023.1217461
Received: 05 May 2023; Accepted: 05 May 2023;
Published: 24 May 2023.
Approved by:
Frontiers Editorial Office, Frontiers Media SA, SwitzerlandCopyright © 2023 Frontiers Production Office. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Frontiers Production Office, cHJvZHVjdGlvbi5vZmZpY2VAZnJvbnRpZXJzaW4ub3Jn