- 1Institute of Laboratory Medicine, Universities of Giessen and Marburg Lung Center (UGMLC), Philipps University Marburg, German Center for Lung Research (DZL) Marburg, Marburg, Germany
- 2Vienna Vaccine Safety Initiative - Pediatric Infectious Diseases and Vaccines, Berlin, Germany
- 3UMR Chrono-Environnement, Université Bourgogne Franche-Comté, Besançon, France
- 4ESCMID Study Group for Respiratory Viruses (ESGREV), Basel, Switzerland
- 54th Department of Internal Medicine, Attikon University Hospital, National and Kapodistrian University of Athens, Athens, Greece
The availability of pathogen-specific treatment options for respiratory tract infections (RTIs) increased the need for rapid diagnostic tests. Besides, retrospective studies, improved lab-based detection methods and the intensified search for new viruses since the beginning of the twenty-first century led to the discovery of several novel respiratory viruses. Among them are human bocavirus (HBoV), human coronaviruses (HCoV-HKU1, -NL63), human metapneumovirus (HMPV), rhinovirus type C (RV-C), and human polyomaviruses (KIPyV, WUPyV). Additionally, new viruses like SARS coronavirus (SARS-CoV), MERS coronavirus (MERS-CoV), novel strains of influenza virus A and B, and (most recently) SARS coronavirus 2 (SARS-CoV-2) have emerged. Although clinical presentation may be similar among different viruses, associated symptoms may range from a mild cold to a severe respiratory illness, and thus require a fast and reliable diagnosis. The increasing number of commercially available rapid point-of-care tests (POCTs) for respiratory viruses illustrates both the need for this kind of tests but also the problem, i.e., that the majority of such assays has significant limitations. In this review, we summarize recently published characteristics of POCTs and discuss their implications for the treatment of RTIs. The second key aspect of this work is a description of new and innovative diagnostic techniques, ranging from biosensors to novel portable and current lab-based nucleic acid amplification methods with the potential future use in point-of-care settings. While prototypes for some methods already exist, other ideas are still experimental, but all of them give an outlook of what can be expected as the next generation of POCTs.
Introduction
Respiratory viruses such as influenza A viruses (IAV) or human respiratory syncytial virus (RSV) are well-known, circulate worldwide, and are associated with significant morbidity and mortality (Iuliano et al., 2018; Shi et al., 2019). On the other hand, there are emerging infectious diseases which were, according to the definition by the WHO, hitherto unknown or rare but are now rapidly spreading either in number of cases or geographically (WHO, 2014a). In the last 20 years, in addition to the emergence of novel influenza and coronaviruses, advances in molecular detection methods have led to the discovery of new respiratory viruses already circulating worldwide (Jartti et al., 2012).
In 2001 human metapneumovirus (HMPV) was discovered (van den Hoogen et al., 2001). The outbreaks of severe acute respiratory syndrome coronavirus (SARS-CoV) in 2003, Middle East respiratory syndrome coronavirus (MERS-CoV) since April 2012, and SARS-CoV-2 since December 2019 highlight the danger of emerging (zoonotic) and highly pathogenic respiratory viruses (Fouchier et al., 2003; Zaki et al., 2012; WHO, 2020c). Two other coronaviruses, human coronaviruses (HCoV) NL63 and HKU1 were discovered in 2004 and 2005, respectively (Fouchier et al., 2004; van der Hoek et al., 2004; Woo et al., 2005). In 2005, Allander et al. described a new member of the family Parvoviridae, human bocavirus (HBoV) (Allander et al., 2005). The two new human polyomaviruses KI and WU (KIPyV, WUPyV) were first discovered in 2007 (Allander et al., 2007; Gaynor et al., 2007) and although often detected in samples, their role in causality of respiratory illness is still under discussion (reviewed in Jartti et al., 2012). In 2007, the human rhinovirus group C (RV-C) was introduced when newly sequenced strains differed significantly from the existing groups A and B (Lee et al., 2007). Finally, avian influenza viruses (AIV) such as IAV H5N1, H7N7, or H9N2 crossed the species barrier to infect humans several times in the last years (reviewed in Kim et al., 2016).
Lab-based techniques still dominate the field of virus diagnostics. Classical methods such as virus cultures, electron microscopy, and serology have been complemented by nucleic acid amplification tests (NAATs), sequencing (including next generation sequencing), and different antigen detection methods. Today, in most clinical settings, NAATs have replaced virus cultures as the gold standard, due to their high specificity, faster turnaround times, and absence of limitations posed by the need for susceptible cell lines (Fox, 2007).
In contrast to lab-based tests, point-of-care tests (POCTs) are performed at the site of sample collection (e.g., bedside, physician's office, or emergency department) and provide results usually in <2 h (Basile et al., 2018; Vos et al., 2019). Furthermore, they require only little hands-on time and no specific laboratory training as most critical steps are automated in a single device. The latter may range from handheld to benchtop size and is not designed for high-throughput sample processing. POCTs and other fast diagnostic tests performed in laboratories but provide results within 1–2 h may be called near-POCTs. Prompt identification of the causative pathogen may help the responsible healthcare professional choosing the appropriate treatment or take the right decisions in outbreak situations, regarding hospitalization and quarantine (Brendish et al., 2015).
In this review, we provide a brief overview of currently available POCTs for the diagnosis of emerging and new respiratory viruses along with their advantages and limitations and discuss recently published approaches and techniques with a potential use in future POCTs.
Commercially Available Test Systems
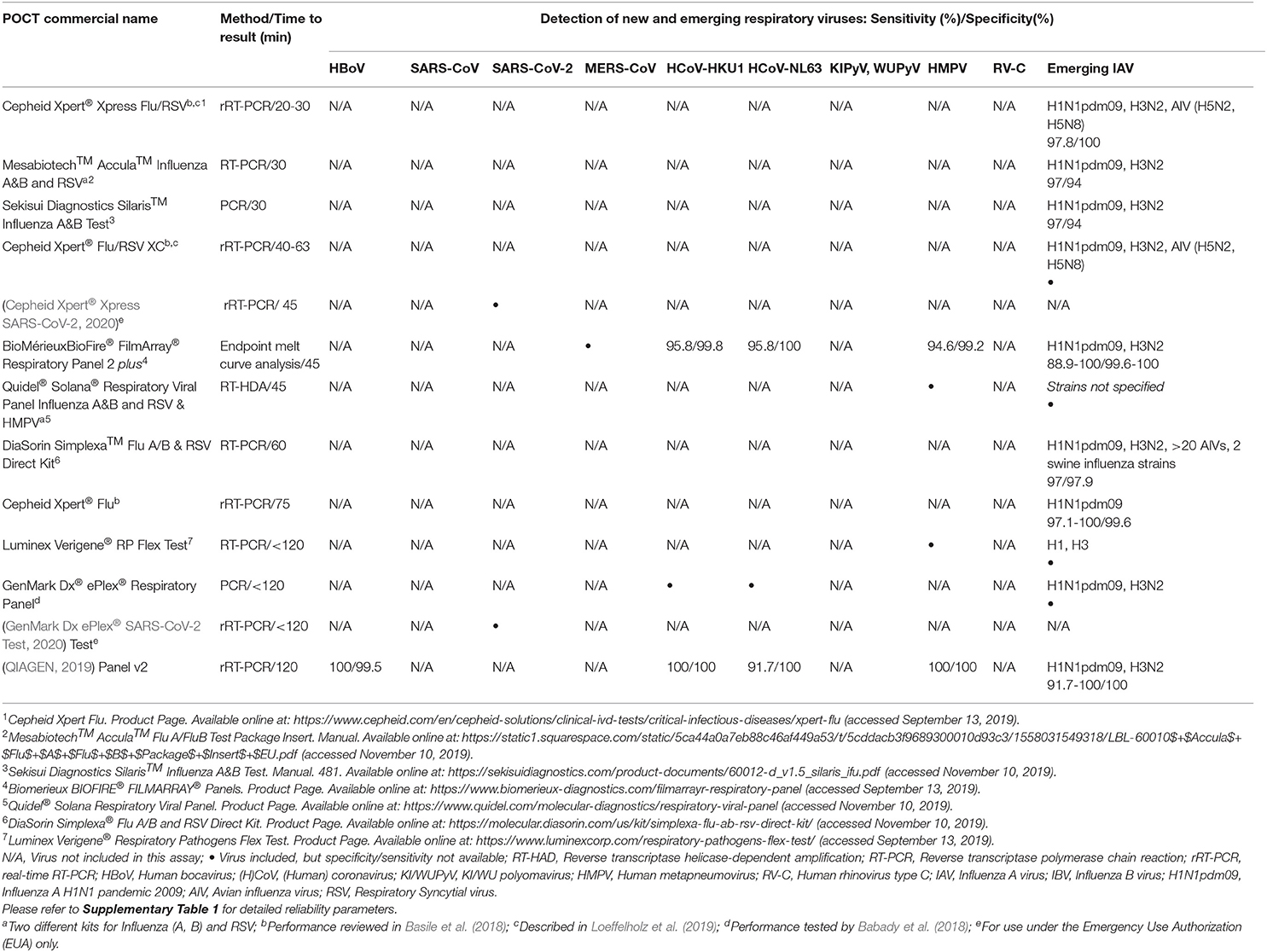
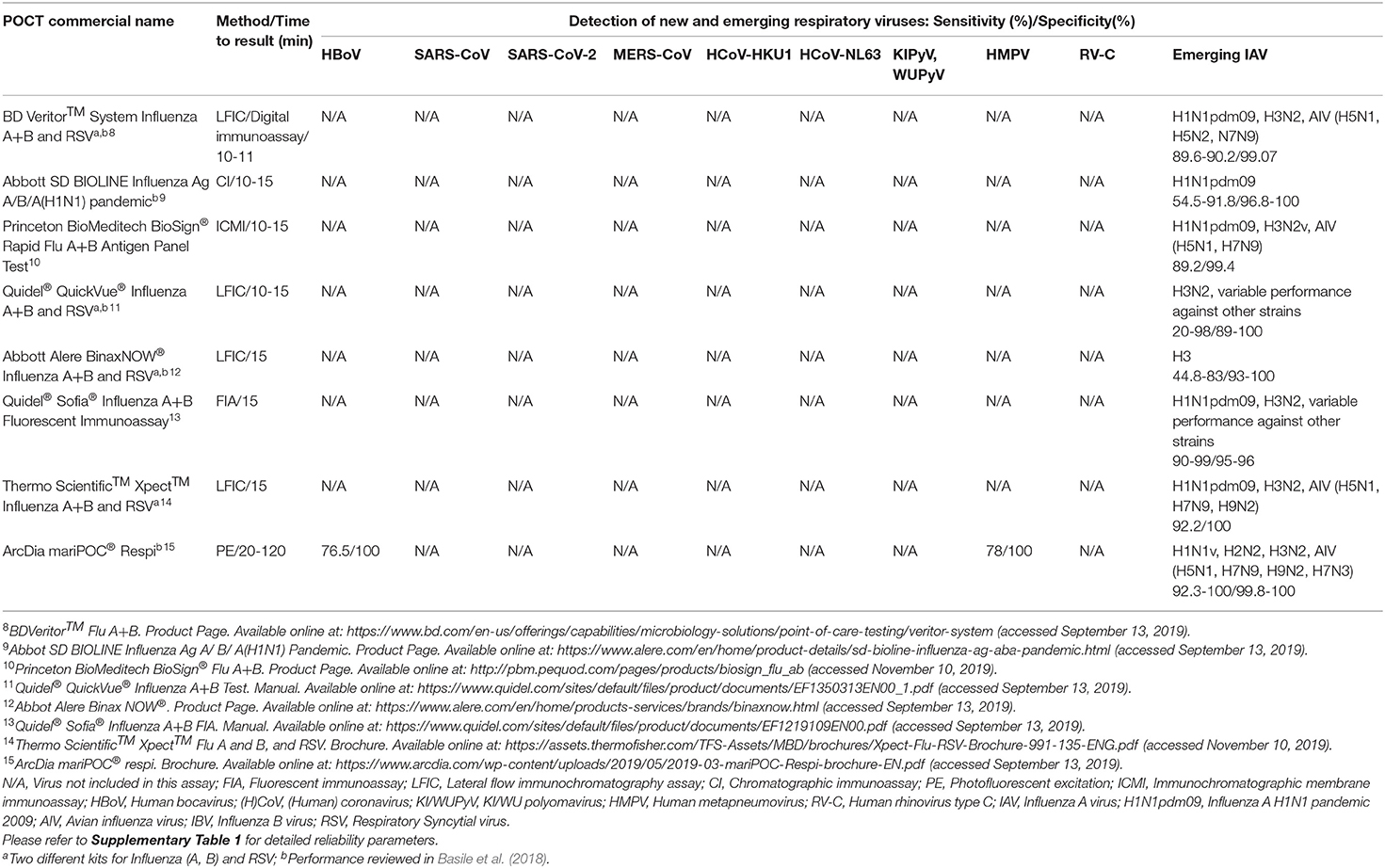
For the diagnosis of commonly encountered respiratory viruses such as IAV, influenza B virus (IBV), and RSV many commercially available POCTs and near-POCTs with different sensitivities and specificities for each virus are available (Supplementary Table 1). However, a recent meta-analysis has demonstrated that three newer generation rapid multiplex polymerase chain reaction systems (mPCRs) (bioMérieuxBioFire® FilmArray® RP, Nanosphere Verigene® RV+ test, and Hologic Gen-Probe Prodesse assays) are highly accurate, though usually more expensive, and may provide important diagnostic information for early identification of IAV, IBV, and RSV (Huang H. S. et al., 2018; Rabold and Waggoner, 2019).
Diagnosis of emerging and novel viruses, including HBoV, RV-C, coronaviruses (e.g., HCoV-HKU1, HCoV-NL63, SARS-CoV-2) as well as specific subtypes of AIV (H5N1, H7N9, H10N8) and reassortant IAV strains remains challenging (Schuster and Williams, 2018; WHO, 2020a). The diagnosis of most of these viruses is based on molecular techniques that can only be performed at specialized referral centers. Recently, there has been increased interest in using POCTs in other settings, such as emergency departments, although implementation might be hampered by the need for specific training (Bouzid et al., 2020). NAATs have higher sensitivity than immunochromatographic assays, but generally, they require a higher degree of technical skills and training (Drancourt et al., 2016). Polymerase chain reaction (PCR) remains the gold standard technique for the diagnosis of AIV subtypes while WHO recommends against the utilization of rapid tests in avian flu diagnosis (WHO, 2005; Monne et al., 2008; Schuster and Williams, 2018). RV-C, is typically detected from nasopharyngeal specimens using reverse transcriptase PCR (RT-PCR) and specific species and serotypes can be further identified by semi-nested PCRs or sequencing (Bochkov et al., 2011; Jartti et al., 2012; Schuster and Williams, 2018).
Diagnosis of HBoV was based on the detection of the specific IgM antibody along with a 4-fold increase of the IgG titer or low IgG avidity indicative of seroconversion, but adaptive immune response requires several hours or even days to develop, thus the utility of such tests in acute settings is limited (Soderlund-Venermo et al., 2009). As HBoV DNA persists in airway secretions for months after an acute infection, quantitative PCR along with serology are currently the preferred diagnostic methods (Christensen et al., 2019). Viral DNAemia, mRNA detection via RT-PCR, and antigen immunodetection assays have shown some promising results but further studies to define their sensitivity, specificity, and applicability in clinical practice are needed (Soderlund-Venermo et al., 2009; Christensen et al., 2010, 2013, 2019; Proenca-Modena et al., 2011; Xu et al., 2017).
The detection of KIPyV and WUPyV, found in secretion from both symptomatic and asymptomatic patients, is based on PCR and serology testing (Neske et al., 2010; Touze et al., 2010; Jartti et al., 2012).
SARS-CoV-2, HCoV-NL63, and -HKU1 as well as HMPV diagnosis is based primarily on RT-PCR methods (van den Hoogen et al., 2004; Nichols et al., 2008; Jartti et al., 2012; WHO, 2020a). No data is available for fluorescent antibody, rapid cultures and enzyme immunoassay (EIA) diagnostic tests for coronaviruses NL63 and HKU1, whereas fluorescent antibody assays may be of some utility for HMPV (Nichols et al., 2008; Jartti et al., 2012). For HMPV specifically, multiplex ligation-dependent probe amplification (MLPA) on a nasopharyngeal swab has high sensitivity and specificity rates (100 and 96%, respectively) (Reijans et al., 2008; Panda et al., 2014; Hoppe et al., 2016).
Available tests for the now extinct SARS-CoV include antibody testing using an EIA and RT-PCR tests in respiratory, blood, and stool specimens (CDC, 2004). For the detection of MERS-CoV, that has its epicenter in the Arabian peninsula, the United States Centers for Disease Control and Prevention (CDC) and the WHO recommend sampling from the lower respiratory tract and real-time RT-PCR (rRT-PCR) testing with specific primers since this appears to be more sensitive than testing of upper respiratory tract specimens (WHO, 2014b,c; CDC, 2015).
NAATs have been proven to be highly accurate and easily scalable tests during large outbreaks of novel or emerging viruses, like the SARS-CoV-2 pandemic. Rapid genome sequencing analysis accommodates the fast development of reliable in-house and commercially available NAATs reagents shortly after an outbreak onset (CDC, 2020; WHO, 2020b).
Innovative Approaches for Future POCTs
Biosensors
Biosensors can be a reliable and cost-effective way to detect specific pathogens in point-of-care settings. Different types of sensors for rapid identification of respiratory viruses have been developed recently. By using a gold-coated array of carbon electrodes, the authors were able to detect MERS-CoV spike protein in the picogram range within 20 min (Layqah and Eissa, 2019). This electrochemical assay is based on the competitive binding of a MERS-CoV antibody either to the virus in the sample or to the immobilized antigen on the electrode, which can be measured by a reduced peak current through the chip. In theory, this technique can be easily expanded to simultaneously detect multiple viruses, however, its diagnostic performance needs to be validated with the use of patient samples.
Different types of biosensors for AIV detection use nanobio hybrid materials (reviewed in Lee et al., 2018). In one of those approaches, a DNA probe coupled to a field-effect transistor enabled detection of target DNA down to 1 fM (Lin et al., 2009). Another sensitive technique is based on surface plasmon resonance (SPR), in which biomolecules bound to a metal surface lead to the reduction of the reflection of an incident light beam (Tang et al., 2010; Chang et al., 2018). With a new antibody against a recombinant AIV H7N9, the authors were able to reach a detection limit of a few hundred copies per mL nasal fluid within 10 min of processing time. Although still experimental, the characteristics of this approach render it a promising candidate for a future rapid POCT.
New Techniques and Prototypes
In a capillary convective PCR (CCPCR), the reagents circulate across a temperature gradient in a simple capillary tube, which allows run times shorter than 30 min (Chou et al., 2011). Together with a self-made dipstick detection method, this principle was already used to test for non-respiratory viruses like hepatitis C virus (Zhang et al., 2013, 2014). Zhou et al. integrated this method into a 1.5 kg device for the automation of the detection of different IAV strains (Zhuo et al., 2018). Although fast and sensitive, manual RNA extraction limits the use as POCT so far. Hardick et al. presented another portable NAAT device not only for the detection of IAV, but also for IBV, RSV, and MERS-CoV. It is based on a RT-PCR in microfluidic cards but likewise lacks automated nucleic acid extraction (Hardick et al., 2018).
Alternatively to nucleic acid-based techniques, giant magnetoresistive (GMR) biosensors function comparable to an enzyme-linked immunosorbent assay but use magnetic labels instead of enzymes or fluorophores coupled to detection antibodies (Hall et al., 2010). With such a sensor, Wu et al. constructed a handheld device which, in connection with a computer or smartphone, was able to detect IAV H3N2 in purified and disrupted virus solutions (Wu et al., 2017). Although H3N2 strains are circulating already since 1968 in the human population (Smith et al., 2004), this method is likely adaptable to emerging AIVs with the use of appropriate capture antibodies.
By designing a prototype for a lateral flow assay for approximately 5 US$, Huang et al. proved that the simultaneous detection of IAV and IBV in swab samples is possible at very low costs (Huang et al., 2017). However, the sensitivity of this prototype needs significant improvements before it can be used under clinical conditions. Instead of constructing a completely new apparatus, Cui et al. used commercially available glucose test strips together with specifically designed glucose-bearing substrates to test for the cleavage activity of IAV neuraminidase in spiked samples (Cui et al., 2017).
Although the majority of the presented approaches and prototypes focuses on the detection of influenza viruses, most of them can theoretically be applied to emerging or new viruses with only minor changes.
Lab-Based NAATs With Potential Use as Point-of-Care Applications
In comparison to PCRs, isothermal NAATs do not require complex devices when working with extracted nucleic acids. Reverse transcription strand invasion-based amplification (RT-SIBA) and reverse transcription loop-mediated isothermal amplification (RT-LAMP) are two examples which have been used to detect e.g., HMPV (Song et al., 2014), IAV (Eboigbodin et al., 2016), and MERS-CoV (Huang P. et al., 2018). Wang et al. went on to integrate seven RT-LAMP assays into a microfluidic chip for the multiplex detection of different respiratory viruses in a device weighing <3 kg (Wang et al., 2018). Another chip-based system, named iROAD, uses reverse transcription-based recombinase polymerase amplification (RT-RPA) and was able to rapidly identify IAV, different HCoVs, and other respiratory viruses in extracted nucleic acids (Koo et al., 2017).
Discussion
Clinical Performance of POCTs
The development of new laboratory and point-of-care diagnostic tests for influenza, RSV, and emerging respiratory viruses has taken up pace in recent years. Healthcare professionals, hospital managers, and laboratory directors will need to update and re-evaluate best practices regularly.
The advancement of diagnostic capabilities may change the way we identify, document, and communicate respiratory viral infections in the future. Along with these developments, the expectations of patients and healthcare professionals may change as well, i.e., patients will want to know the responsible pathogen and their clinical prognosis. With the development of specific antiviral therapies and vaccines, new diagnostic algorithms will be needed to ensure the highest quality of care while containing costs.
From a viewpoint of quality of care and clinical management, timely infection control, and the ability to act upon results, the future will likely belong to portable, CLIA-waived rapid diagnostic tests that take 10–20 min.
A key criterion for the evaluation of diagnostic tests will be their clinical utility, i.e., their ability to identify the current culprit for the patient's symptoms, and to distinguish relevant pathogen(s) from bystander pathogens. More research is needed to correlate clinical outcomes and laboratory data.
A second concern will be the correct timing of diagnostic testing with regards to a patient's course of illness. The sensitivity and positive predictive value of a diagnostic test depend on specimen quality and virus load (which is usually higher in children and early in the course of illness), duration of viral shedding, and patient's immune status. Future diagnostic algorithms will need to consider these factors in addition to the epidemiology of viruses in the respective season or region.
Advantages and Limitations of POCTs
The initial criticism of POCTs was directed toward a lack of sensitivity and a high degree of variability in test results. Early studies during the 2009/10 flu pandemic reported sensitivities for influenza POCTs ranging from 20 to 70% for the same test kit (Rath et al., 2012). In published evaluation studies, it was not entirely clear whether POCTs were performed at the bedside or whether samples were sent to the laboratory, which would constitute a near-POCT. These methodological inconsistencies also impaired the value of meta-analyses comparing the point-of-care performance of different commercial tests (Chartrand et al., 2015). The reported performance differences also hint to the fact that the training and experience of the staff performing the test have an impact. Procedural concerns were most pronounced with early-stage lateral flow (“strip”) tests, where the result was read manually. Second-generation antigen POCTs and modern molecular POCTs achieve significantly higher reproducibility and sensitivity through automation of key steps in the process. In 2018, the FDA implemented new performance regulations setting new sensitivity thresholds required for POCTs to maintain approval (FDA, 2017). As a result, only several influenza POCTs were taken off the market. The role of regulators in setting quality standards for POCTs cannot be underestimated (Zhang et al., 2016; Azar and Landry, 2018).
In addition to the reliability of the POCT result itself, the hands-on time and the time-to-result are still critical for user-acceptance at the bedside. It is expected that future POCTs will be more robust and easier to use.
As the majority of acute respiratory infections (ARI) is of viral origin, these infections are common reasons for inappropriate antibiotic use (Harris et al., 2016; Tief et al., 2016). Studies have raised the expectation that expanded use of virus diagnostics at the point-of-care may help to limit the use of antibiotics (Bonner et al., 2003). The European Health Action Plan on AMR (European Parliament, 2018) and the O'Neill Report on “Tackling Drug-resistant infections globally” (O'Neill, 2016) both point to rapid diagnostics as a key instrument in tackling AMR. Including POCTs in antimicrobial stewardship programs might increase their acceptance by physicians.
The ability to direct “the right treatment to the right patient at the right time” facilitates precision medicine. In the future, advanced virus diagnostics may be combined with biomarker POCTs for the prediction of individual-level host responses to further target treatment to those who are most likely to benefit from it.
A major obstacle to expanding the use of POCTs is cost. In fact, POCTs would be most impactful in settings where the majority of early treatment decisions are made. The current pricing schemes, however, seem too high for broad-scale testing in the community and emergency departments. In addition to pricing, reimbursement strategies need to be reconsidered. Even in hospital emergency rooms, per-capita standard reimbursements disincentivize the use of virus diagnostics in patients with ARI. Up-to-date economic models are needed to clarify the cost-effectiveness of different types of POCTs.
The steepest increase in antimicrobial resistance is being observed in low-middle income countries (LMIC) (WHO-PAHO, 2018). For LMIC, portable POCTs with multi-modality for known and emerging pathogens, or simple lab-based instrumentation with no/minimal need for cold chain or refrigeration of reagents, may be the most likely to succeed.
Priorities for the Development of New POCTs
None of the currently commercially available POCTs covers all viruses discussed here (Tables 1, 2, Supplementary Table 1). To date, the same holds true for new technologies as well. However, it is conceivable that these techniques may be adapted to other respiratory viruses after having shown their usefulness in practice. Especially biosensors have the potential for a wider spectrum of applications. They also do not encounter the major limitation of NAATs as POCTs: the extraction of nucleic acids (Ali et al., 2017). For these tests, even less obvious complications, like the stability of the plastic materials against required chemicals, have to be overcome for future highly specific nucleic acid-based POCTs.
Any ideal POCT should fulfill the ASSURED criteria of the World Health Organization to be applicable in resource-limited settings (Mabey et al., 2004). Currently, this is most likely true for tests based on lateral flow immunochromatography but coming improvements e.g., in miniaturization and battery capacity may facilitate the use of other test principles (Basile et al., 2018).
While economic benefits of POCTs and better outcomes for the patients are still discussed, it is likely that these tests will gain further importance with decreasing processing costs and improved robustness.
Author Contributions
CS and PN planned, structured, and edited the manuscript. PN searched the literature and integrated all contributions. PN, BR, PF, EA, and ST participated in the writing of the first draft of the manuscript and subsequent revisions. BR and PF contributed equally to this work. All authors critically read, reviewed, and approved the submitted final version of the manuscript.
Funding
CS was supported by Universities Giessen and Marburg Lung Center (UGMLC), the German Center for Lung Research (DZL), University Hospital Giessen and Marburg (UKGM) research funding according to article 2, section 3 cooperation agreement, and the Deutsche Forschungsgemeinschaft (DFG)-funded-SFB 1021 (C04), -KFO 309 (P10), and SK 317/1-1 (project number 428518790). PF was supported by Doctorate scholarship by the State Scholarships Foundation (IKY), Partnership Agreement (PA) 2014-2020, co-financed by Greece and the European Union (European Social Fund - ESF) through the Operational Programme Human Resources Development, Education and Lifelong Learning 2014-2020.
Conflict of Interest
CS received consultancy fees and research funding from Hycor Biomedical and Thermo Fisher Scientific, research funding from Mead Johnson Nutrition (MJN), and consultancy fees from Bencard Allergie.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Authors would like to thank the ESCMID Study Group on Respiratory Viruses (ESGREV) for providing a platform for interaction between authors.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2020.00181/full#supplementary-material
References
Ali, N., Rampazzo, R. D. C. P., Costa, A. D. T., and Krieger, M. A. (2017). Current nucleic acid extraction methods and their implications to point-of-care diagnostics. Biomed. Res. Int. 2017:9306564. doi: 10.1155/2017/9306564
Allander, T., Andreasson, K., Gupta, S., Bjerkner, A., Bogdanovic, G., Persson, M. A. A., et al. (2007). Identification of a third human polyomavirus. J. Virol. 81, 4130–4136. doi: 10.1128/JVI.00028-07
Allander, T., Tammi, M. T., Eriksson, M., Bjerkner, A., Tiveljung-Lindell, A., and Andersson, B. (2005). Cloning of a human parvovirus by molecular screening of respiratory tract samples. Proc. Natl. Acad. Sci. U.S.A. 102, 12891–12896. doi: 10.1073/pnas.0504666102
Azar, M. M., and Landry, M. L. (2018). Detection of influenza A and B viruses and respiratory syncytial virus by use of clinical laboratory improvement amendments of 1988 (CLIA)-waived point-of-care assays: a paradigm shift to molecular tests. J. Clin. Microbiol. 56:e00367-18. doi: 10.1128/JCM.00367-18
Babady, N. E., England, M. R., Jurcic Smith, K. L., He, T., Wijetunge, D. S., Tang, Y.-W., et al. (2018). Multicenter evaluation of the eplex respiratory pathogen panel for the detection of viral and bacterial respiratory tract pathogens in nasopharyngeal swabs. J. Clin. Microbiol. 56:e01658-17. doi: 10.1128/JCM.01658-17
Basile, K., Kok, J., and Dwyer, D. E. (2018). Point-of-care diagnostics for respiratory viral infections. Expert. Rev. Mol. Diagn. 18, 75–83. doi: 10.1080/14737159.2018.1419065
Bochkov, Y. A., Palmenberg, A. C., Lee, W. M., Rathe, J. A., Amineva, S. P., Sun, X., et al. (2011). Molecular modeling, organ culture and reverse genetics for a newly identified human rhinovirus C. Nat. Med. 17, 627–632. doi: 10.1038/nm.2358
Bonner, A. B., Monroe, K. W., Talley, L. I., Klasner, A. E., and Kimberlin, D. W. (2003). Impact of the rapid diagnosis of influenza on physician decision-making and patient management in the pediatric emergency department: results of a randomized, prospective, controlled trial. Pediatrics 112, 363–367. doi: 10.1542/peds.112.2.363
Bouzid, D., Zanella, M.-C., Kerneis, S., Visseaux, B., May, L., Schrenzel, J., et al. (2020). Rapid diagnostic tests for infectious diseases in the emergency department. Clin. Microbiol. Infect. doi: 10.1016/j.cmi.2020.02.024. [Epub ahead of print].
Brendish, N. J., Schiff, H. F., and Clark, T. W. (2015). Point-of-care testing for respiratory viruses in adults: the current landscape and future potential. J. Infect. 71, 501–510. doi: 10.1016/j.jinf.2015.07.008
CDC (2004). Severe Acute Respiratory Syndrome. Available online at: https://www.cdc.gov/sars/surveillance/absence.pdf (accessed September 13, 2019).
CDC (2015). Interim Guidelines for Collecting, Handling, and Testing Clinical Specimens from Patients Under Investigation (PUIs) for Middle East Respiratory Syndrome Coronavirus (MERS-CoV) – Version 2.1. Available online at: https://www.cdc.gov/coronavirus/mers/downloads/Guidelines-Clinical-Specimens.pdf (accessed March 01, 2019).
CDC (2020). Frequently Asked Questions on COVID-19 Testing at Laboratories. Available online at: https://www.cdc.gov/coronavirus/2019-ncov/lab/testing-laboratories.html (accessed March 14, 2020).
Cepheid Xpert® Xpress SARS-CoV-2 (2020). Assay Manual. Available online at: https://www.fda.gov/media/136314/download (accessed March 25, 2020).
Chang, Y.-F., Wang, W.-H., Hong, Y.-W., Yuan, R.-Y., Chen, K.-H., Huang, Y.-W., et al. (2018). Simple strategy for rapid and sensitive detection of avian influenza a H7N9 virus based on intensity-modulated SPR biosensor and new generated antibody. Anal. Chem. 90, 1861–1869. doi: 10.1021/acs.analchem.7b03934
Chartrand, C., Tremblay, N., Renaud, C., and Papenburg, J. (2015). Diagnostic accuracy of rapid antigen detection tests for respiratory syncytial virus infection: systematic review and meta-analysis. J. Clin. Microbiol. 53, 3738–3749. doi: 10.1128/JCM.01816-15
Chou, W. P., Chen, P. H., Miao, M., Kuo, L. S., Yeh, S. H., and Chen, P. J. (2011). Rapid DNA amplification in a capillary tube by natural convection with a single isothermal heater. BioTechniques 50, 52–57. doi: 10.2144/000113589
Christensen, A., Dollner, H., Skanke, L. H., Krokstad, S., Moe, N., and Nordbo, S. A. (2013). Detection of spliced mRNA from human bocavirus 1 in clinical samples from children with respiratory tract infections. Emerg. Infect. Dis. 19, 574–580. doi: 10.3201/eid1904.121775
Christensen, A., Kesti, O., Elenius, V., Eskola, A. L., Døllner, H., Altunbulakli, C., et al. (2019). Human bocaviruses and paediatric infections. Lancet 3, 418–426. doi: 10.1016/S2352-4642(19)30057-4
Christensen, A., Nordbo, S. A., Krokstad, S., Rognlien, A. G., and Dollner, H. (2010). Human bocavirus in children: mono-detection, high viral load and viraemia are associated with respiratory tract infection. J. Clin. Virol. 49, 158–162. doi: 10.1016/j.jcv.2010.07.016
Cui, X., Das, A., Dhawane, A. N., Sweeney, J., Zhang, X., Chivukula, V., et al. (2017). Highly specific and rapid glycan based amperometric detection of influenza viruses. Chem. Sci. 8, 3628–3634. doi: 10.1039/C6SC03720H
Drancourt, M., Michel-Lepage, A., Boyer, S., and Raoult, D. (2016). The point-of-care laboratory in clinical microbiology. Clin. Microbiol. Rev. 29, 429–447. doi: 10.1128/CMR.00090-15
Eboigbodin, K., Filén, S., Ojalehto, T., Brummer, M., Elf, S., Pousi, K., et al. (2016). Reverse transcription strand invasion based amplification (RT-SIBA): a method for rapid detection of influenza A and B. Appl. Microbiol. Biotechnol. 100, 5559–5567. doi: 10.1007/s00253-016-7491-y
European Parliament (2018). European One Health Action Plan Against Antimicrobial Resistance (AMR). Available online at: https://oeil.secure.europarl.europa.eu/oeil/popups/ficheprocedure.do?lang=en&reference=2017/2254(INI) (accessed April 02, 2019).
FDA (2017). Microbiology Devices; Reclassification of Influenza Virus Antigen Detection Test Systems Intended for Use Directly With Clinical Specimens. Docket No. FDA-2014-N-0440. Available online at: https://www.govinfo.gov/content/pkg/FR-2017-01-12/pdf/2017-00199.pdf (accessed November 14, 2019).
Fouchier, R. A. M., Hartwig, N. G., Bestebroer, T. M., Niemeyer, B., Jong, J. C., de Simon, J. H., et al. (2004). A previously undescribed coronavirus associated with respiratory disease in humans. Proc. Natl. Acad. Sci. U.S.A. 101, 6212–6216. doi: 10.1073/pnas.0400762101
Fouchier, R. A. M., Kuiken, T., Schutten, M., van Amerongen, G., van Doornum, G. J. J., van den Hoogen, B. G., et al. (2003). Aetiology: koch's postulates fulfilled for SARS virus. Nature 423:240. doi: 10.1038/423240a
Fox, J. D. (2007). Nucleic acid amplification tests for detection of respiratory viruses. J. Clin. Virol. 40, S15–S23. doi: 10.1016/S1386-6532(07)70005-7
Gaynor, A. M., Nissen, M. D., Whiley, D. M., Mackay, I. M., Lambert, S. B., Wu, G., et al. (2007). Identification of a novel polyomavirus from patients with acute respiratory tract infections. PLoS Pathog. 3:e64. doi: 10.1371/journal.ppat.0030064
GenMark Dx ePlex® SARS-CoV-2 Test (2020). Assay Manual. Available online at: https://www.fda.gov/media/136282/download (accessed March 25, 2019).
Hall, D. A., Gaster, R. S., Lin, T., Osterfeld, S. J., Han, S., Murmann, B., et al. (2010). GMR biosensor arrays: a system perspective. Biosens. Bioelectron. 25, 2051–2057. doi: 10.1016/j.bios.2010.01.038
Hardick, J., Metzgar, D., Risen, L., Myers, C., Balansay, M., Malcom, T., et al. (2018). Initial performance evaluation of a spotted array mobile analysis platform (MAP) for the detection of influenza A/B, RSV, and MERS coronavirus. Diagn. Microbiol. Infect. Dis. 91, 245–247. doi: 10.1016/j.diagmicrobio.2018.02.011
Harris, A. M., Hicks, L. A., and Qaseem, A. (2016). Appropriate antibiotic use for acute respiratory tract infection in adults: advice for high-value care from the american college of physicians and the centers for disease control and prevention. Ann. Intern. Med. 164, 425–434. doi: 10.7326/M15-1840
Hoppe, B. P. C., Jongh, E., de Griffioen-Keijzer, A., Zijlstra-Baalbergen, J. M., IJzerman, E. P. F., and Baboe, F. (2016). Human metapneumovirus in haematopoietic stem cell transplantation recipients: a case series and review of the diagnostic and therapeutic approach. Neth. J. Med. 74, 336–341. Available online at: http://www.njmonline.nl/article.php?a=1762&d=1172&i=199
Huang, H. S., Tsai, C. L., Chang, J., Hsu, T. C., Lin, S., and Lee, C. C. (2018). Multiplex PCR system for the rapid diagnosis of respiratory virus infection: systematic review and meta-analysis. Clin. Microbiol. Infect. 24, 1055–1063. doi: 10.1016/j.cmi.2017.11.018
Huang, P., Wang, H., Cao, Z., Jin, H., Chi, H., Zhao, J., et al. (2018). A rapid and specific assay for the detection of MERS-CoV. Front. Microbiol. 9:1101. doi: 10.3389/fmicb.2018.01101
Huang, S., Abe, K., Bennett, S., Liang, T., Ladd, P. D., Yokobe, L., et al. (2017). Disposable autonomous device for swab-to-result diagnosis of influenza. Anal. Chem. 89, 5776–5783. doi: 10.1021/acs.analchem.6b04801
Iuliano, A. D., Roguski, K. M., Chang, H. H., Muscatello, D. J., Palekar, R., Tempia, S., et al. (2018). Estimates of global seasonal influenza-associated respiratory mortality: a modelling study. Lancet 391, 1285–1300. doi: 10.1016/S0140-6736(17)33293-2
Jartti, T., Jartti, L., Ruuskanen, O., and Söderlund-Venermo, M. (2012). New respiratory viral infections. Curr. Opin. Pulm. Med. 18, 271–278. doi: 10.1097/MCP.0b013e328351f8d4
Kim, S. M., Kim, Y.-I., Pascua, P. N. Q., and Choi, Y. K. (2016). Avian influenza a viruses: evolution and zoonotic infection. Semin. Respir. Crit. Care Med. 37, 501–511. doi: 10.1055/s-0036-1584953
Koo, B., Jin, C. E., Lee, T. Y., Lee, J. H., Park, M. K., Sung, H., et al. (2017). An isothermal, label-free, and rapid one-step RNA amplification/detection assay for diagnosis of respiratory viral infections. Biosens. Bioelectron. 90, 187–194. doi: 10.1016/j.bios.2016.11.051
Layqah, L. A., and Eissa, S. (2019). An electrochemical immunosensor for the corona virus associated with the middle east respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Mikrochim. Acta 186:224. doi: 10.1007/s00604-019-3345-5
Lee, T., Ahn, J.-H., Park, S. Y., Kim, G.-H., Kim, J., Kim, T.-H., et al. (2018). Recent advances in AIV biosensors composed of nanobio hybrid material. Micromachines 9:651. doi: 10.3390/mi9120651
Lee, W.-M., Kiesner, C., Pappas, T., Lee, I., Grindle, K., Jartti, T., et al. (2007). A diverse group of previously unrecognized human rhinoviruses are common causes of respiratory illnesses in infants. PLoS ONE 2:e966. doi: 10.1371/journal.pone.0000966
Lin, C.-H., Hung, C.-H., Hsiao, C.-Y., Lin, H.-C., Ko, F.-H., and Yang, Y.-S. (2009). Poly-silicon nanowire field-effect transistor for ultrasensitive and label-free detection of pathogenic avian influenza DNA. Biosens. Bioelectron. 24, 3019–3024. doi: 10.1016/j.bios.2009.03.014
Loeffelholz, M., Jones, R., Uy, C., Lo, D., Kele, A., and Baron, E. J. (2019). Cepheid assays Detect a Broad Range of Contemporary Human and Avian Influenza strains. Available online at: https://www.cepheid.com/administrator/components/com_productcatalog/library-files/13c07133b4e35667b67e9158de356e05-1353—Monograph-on-Human-Influenza-and-Avian-Strains-Final-08082019-08082021.pdf (accessed September 13, 2019).
Mabey, D., Peeling, R. W., Ustianowski, A., and Perkins, M. D. (2004). Diagnostics for the developing world. Nat. Rev. Microbiol. 2, 231–240. doi: 10.1038/nrmicro841
Monne, I., Ormelli, S., Salviato, A., Battisti, C., de Bettini, F., Salomoni, A., et al. (2008). Development and validation of a one-step real-time PCR assay for simultaneous detection of subtype H5, H7, and H9 avian influenza viruses. J. Clin. Microbiol. 46, 1769–1773. doi: 10.1128/JCM.02204-07
Neske, F., Prifert, C., Scheiner, B., Ewald, M., Schubert, J., Opitz, A., et al. (2010). High prevalence of antibodies against polyomavirus WU, polyomavirus KI, and human bocavirus in German blood donors. BMC Infect. Dis. 10:215. doi: 10.1186/1471-2334-10-215
Nichols, W. G., Peck Campbell, A. J., and Boeckh, M. (2008). Respiratory viruses other than influenza virus: impact and therapeutic advances. Clin. Microbiol. Rev. 21, 274–90. doi: 10.1128/CMR.00045-07
O'Neill, J. (2016). Tackling Drug-Resistant Infections Globally, Final Report and Recommendations. Available online at: https://amr-review.org/sites/default/files/160518_Final%20paper_with%20cover.pdf (accessed March 20, 2020).
Panda, S., Mohakud, N. K., Pena, L., and Kumar, S. (2014). Human metapneumovirus: review of an important respiratory pathogen. Int. J. Infect. Dis. 25, 45–52. doi: 10.1016/j.ijid.2014.03.1394
Proenca-Modena, J. L., Gagliardi, T. B., Paula, F. E., Iwamoto, M. A., Criado, M. F., Camara, A. A., et al. (2011). Detection of human bocavirus mRNA in respiratory secretions correlates with high viral load and concurrent diarrhea. PLoS ONE 6:e21083. doi: 10.1371/journal.pone.0021083
QIAGEN (2019). QIAstat-Dx® Respiratory Panel Instructions for Use: Version 2. Manual. Available online at: https://www.qiagen.com/ie/resources/resourcedetail?id=371ac902-c220-4e88-9d22-9012c33f268b&lang=en (accessed November 10, 2019).
Rabold, E., and Waggoner, J. (2019). Rapid Diagnostic Tests for Infectious Diseases. Available online at: https://wwwnc.cdc.gov/travel/yellowbook/2020/posttravel-evaluation/rapid-diagnostic-tests-for-infectious-diseases (accessed March 08, 2020).
Rath, B., Tief, F., Obermeier, P., Tuerk, E., Karsch, K., Muehlhans, S., et al. (2012). Early detection of influenza A and B infection in infants and children using conventional and fluorescence-based rapid testing. J. Clin. Virol. 55, 329–333. doi: 10.1016/j.jcv.2012.08.002
Reijans, M., Dingemans, G., Klaassen, C. H., Meis, J. F., Keijdener, J., Mulders, B., et al. (2008). RespiFinder: a new multiparameter test to differentially identify fifteen respiratory viruses. J. Clin. Microbiol. 46, 1232–1240. doi: 10.1128/JCM.02294-07
Schuster, J. E., and Williams, J. V. (2018). Emerging respiratory viruses in children. Infect. Dis. Clin. North. Am. 32, 65–74. doi: 10.1016/j.idc.2017.10.001
Shi, T., Denouel, A., Tietjen, A. K., Campbell, I., Moran, E., Li, X., et al. (2019). Global disease burden estimates of respiratory syncytial virus-associated acute respiratory infection in older adults in 2015: a systematic review and meta-analysis. J. Infect. Dis. jiz059. doi: 10.1093/infdis/jiz059
Smith, D. J., Lapedes, A. S., Jong, J. C., de Bestebroer, T. M., Rimmelzwaan, G. F., Osterhaus, A. D. M. E., et al. (2004). Mapping the antigenic and genetic evolution of influenza virus. Science 305, 371–376. doi: 10.1126/science.1097211
Soderlund-Venermo, M., Lahtinen, A., Jartti, T., Hedman, L., Kemppainen, K., Lehtinen, P., et al. (2009). Clinical assessment and improved diagnosis of bocavirus-induced wheezing in children, Finland. Emerg. Infect. Dis. 15, 1423–1430. doi: 10.3201/eid1509.090204
Song, Q., Zhu, R., Sun, Y., Zhao, L., Wang, F., Deng, J., et al. (2014). Identification of human metapneumovirus genotypes A and B from clinical specimens by reverse transcription loop-mediated isothermal amplification. J. Virol. Methods 196, 133–138. doi: 10.1016/j.jviromet.2013.10.037
Tang, Y., Zeng, X., and Liang, J. (2010). Surface plasmon resonance: an introduction to a surface spectroscopy technique. J. Chem. Educ. 87, 742–746. doi: 10.1021/ed100186y
Tief, F., Hoppe, C., Seeber, L., Obermeier, P., Chen, X., Karsch, K., et al. (2016). An inception cohort study assessing the role of pneumococcal and other bacterial pathogens in children with influenza and ILI and a clinical decision model for stringent antibiotic use. Antivir. Ther. 21, 413–424. doi: 10.3851/IMP3034
Touze, A., Gaitan, J., Arnold, F., Cazal, R., Fleury, M. J., Combelas, N., et al. (2010). Generation of merkel cell polyomavirus (MCV)-like particles and their application to detection of MCV antibodies. J. Clin. Microbiol. 48, 1767–1770. doi: 10.1128/JCM.01691-09
van den Hoogen, B. G., Jong, J. C., de Groen, J., Kuiken, T., Groot, R., de Fouchier, R. A. M., et al. (2001). A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat. Med. 7, 719–724. doi: 10.1038/89098
van den Hoogen, B. G., Osterhaus, D. M. E., and Fouchier, R. A. M. (2004). Clinical impact and diagnosis of human metapneumovirus infection. Pediatr. Infect. Dis. J. 23, S25–32. doi: 10.1097/01.inf.0000108190.09824.e8
van der Hoek, L., Pyrc, K., Jebbink, M. F., Vermeulen-Oost, W., Berkhout, R. J. M., Wolthers, K. C., et al. (2004). Identification of a new human coronavirus. Nat. Med. 10, 368–373. doi: 10.1038/nm1024
Vos, L. M., Bruning, A. H. L., Reitsma, J. B., Schuurman, R., Riezebos-Brilman, A., Hoepelman, A. I. M., et al. (2019). Rapid molecular tests for influenza, respiratory syncytial virus, and other respiratory viruses: a systematic review of diagnostic accuracy and clinical impact studies. Clin. Infect. Dis. 69, 1243–1253. doi: 10.1093/cid/ciz056
Wang, R., Zhao, R., Li, Y., Kong, W., Guo, X., Yang, Y., et al. (2018). Rapid detection of multiple respiratory viruses based on microfluidic isothermal amplification and a real-time colorimetric method. Lab. Chip. 18, 3507–3515. doi: 10.1039/C8LC00841H
WHO (2005). Recommendations on the Use of Rapid Testing for Influenza Diagnosis. Available online at: https://www.who.int/influenza/resources/documents/RapidTestInfluenza_WebVersion.pdf (accessed September 13, 2019).
WHO (2014a). A Brief Guide to Emerging Infectious Diseases and Zoonoses. World Health Organization, Regional Office for South-East Asia. Available online at: https://apps.who.int/iris/handle/10665/204722 (accessed March 27, 2020).
WHO (2014b). Interim Surveillance Recommendations for Human Infection with Middle East Respiratory Syndrome Coronavirus. Available online at: https://www.who.int/csr/disease/coronavirus_infections/InterimRevisedSurveillanceRecommendations_nCoVinfection_14July2014.pdf?ua=1 (accessed September 13, 2019).
WHO (2014c). Laboratory Testing for Middle East Respiratory Syndrome Coronavirus: Interim Recommendations (revised). Available online at: https://www.who.int/csr/disease/coronavirus_infections/WHO_interim_recommendations_lab_detection_MERSCoV_092014.pdf?ua,=1 (accessed September, 13, 2019).
WHO (2020a). Coronavirus Disease (COVID-19) Technical Guidance: Laboratory Testing For 2019-nCoV in Humans. Available online at: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/laboratory-guidance (accessed March 08, 2020).
WHO (2020b). Laboratory Testing For 2019 Novel Coronavirus (2019-nCoV) in Suspected Human Cases: Interim Guidance. Available online at: https://www.who.int/publications-detail/laboratory-testing-for-2019-novel-coronavirus-in-suspected-human-cases-20200117 (accessed March 14, 2020).
WHO (2020c). Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19). Available online at: https://www.who.int/docs/default-source/coronaviruse/who-china-joint-mission-on-covid-19-final-report.pdf (accessed March 01, 2020).
WHO-PAHO (2018). Recommendations for Implementing Antimicrobial Stewardship Programs in Latin America and the Caribbean: Manual for Public Health Decision-Makers. Available online at: http://iris.paho.org/xmlui/handle/123456789/49645 (accessed November 13, 2020).
Woo, P. C. Y., Lau, S. K. P., Chu, C.-M., Chan, K.-H., Tsoi, H.-W., Huang, Y., et al. (2005). Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J. Virol. 79, 884–895. doi: 10.1128/JVI.79.2.884-895.2005
Wu, K., Klein, T., Krishna, V. D., Su, D., Perez, A. M., and Wang, J.-P. (2017). Portable GMR handheld platform for the detection of influenza a virus. ACS Sens. 2, 1594–1601. doi: 10.1021/acssensors.7b00432
Xu, M., Arku, B., Jartti, T., Koskinen, J., Peltola, V., Hedman, K., et al. (2017). Comparative diagnosis of human bocavirus 1 respiratory infection with messenger RNA reverse-transcription polymerase chain reaction (PCR), DNA quantitative PCR, and serology. J. Infect. Dis. 215, 1551–1557. doi: 10.1093/infdis/jix169
Zaki, A. M., van Boheemen, S., Bestebroer, T. M., Osterhaus, A. D. M. E., and Fouchier, R. A. M. (2012). Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 367, 1814–1820. doi: 10.1056/NEJMoa1211721
Zhang, L., Hao, M., Zhang, K., Zhang, R., Lin, G., Jia, T., et al. (2016). External quality assessment for the molecular detection of MERS-CoV in China. J. Clin. Virol. 75, 5–9. doi: 10.1016/j.jcv.2015.12.001
Zhang, S., Lin, Y., Wang, J., Wang, P., Chen, J., Xue, M., et al. (2014). A convenient nucleic acid test on the basis of the capillary convective PCR for the on-site detection of enterovirus 71. J. Mol. Diagn. 16, 452–458. doi: 10.1016/j.jmoldx.2014.04.002
Zhang, S., Xue, M., Zhang, J., Chen, Q., Chen, J., Wang, Z., et al. (2013). A one-step dipstick assay for the on-site detection of nucleic acid. Clin. Biochem. 46, 1852–1856. doi: 10.1016/j.clinbiochem.2013.10.013
Keywords: virus diagnostics, innovative approaches, biosensors, bedside testing, POCT, commercial point-of-care tests
Citation: Nelson PP, Rath BA, Fragkou PC, Antalis E, Tsiodras S and Skevaki C (2020) Current and Future Point-of-Care Tests for Emerging and New Respiratory Viruses and Future Perspectives. Front. Cell. Infect. Microbiol. 10:181. doi: 10.3389/fcimb.2020.00181
Received: 22 November 2019; Accepted: 06 April 2020;
Published: 29 April 2020.
Edited by:
David Ong, Franciscus Gasthuis & Vlietland, NetherlandsReviewed by:
Kai Huang, The University of Texas Medical Branch at Galveston, United StatesValentijn Schweitzer, University Medical Center Utrecht, Netherlands
Copyright © 2020 Nelson, Rath, Fragkou, Antalis, Tsiodras and Skevaki. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Chrysanthi Skevaki, Y2hyeXNhbnRoaS5za2V2YWtpQHVrLWdtLmRl
†These authors have contributed equally to this work
 Philipp P. Nelson
Philipp P. Nelson Barbara A. Rath
Barbara A. Rath Paraskevi C. Fragkou
Paraskevi C. Fragkou Emmanouil Antalis
Emmanouil Antalis Sotirios Tsiodras
Sotirios Tsiodras Chrysanthi Skevaki
Chrysanthi Skevaki