- Jiangxi Key Laboratory of Power Batteries and Materials, Department of Energy and Environmental Materials, School of Materials Science and Engineering, Jiangxi University of Sciences and Technology, Ganzhou, China
In order to develop new type of carbon anode material with high performance, a novel organic carbon material polyacrylonitrile (PAN) hard carbon was prepared by calcination of polypropylene cyanide at 1,050°C. The obtained PAN hard carbon is used as the negative electrode material of lithium ion battery, showing an initial capacity of 343.5 mAh g−1 which is equal to that of graphite electrode (348.6 mAh g−1), and a higher initial coulomb efficiency of 87.9% than that of graphite electrode (84.4%). Moreover, the PAN hard carbon electrode shows superior cycle stability and rate performance at different current rates. The charge capacities are 320.1, 219.0, 212.9, and 123.5 mAh g−1 at current rates of 0.2, 1, 2, and 3 C, with coulombic efficiencies of 98.1, 100.6, 88.1, and 110.0%, respectively, which are higher than those of graphite electrode (83.8, 97.6, 98.8, and 94.1%). In addition, the charging capacity of PAN hard carbon electrode can still remain at 284.3 mAh g−1 (0.2 C after 200 cycles), 226.4 mAh g−1 (1 C after 300 cycles), 149.5 mAh g−1 (2 C after 400 cycles), and 120.0 mAh g−1 (3 C after 100 cycles), with capacity retention rates of 78.2, 111.9, 79.7, and 88.6%, respectively. The capacity and capacity retention rate after cycling are significantly higher than those of graphite electrode, indicating that the PAN hard carbon electrode shows superior rate performance, which would provide a new idea for the development of novel negative electrode material with high performance.
Introduction
The lithium ion battery has a wide application in the fields of mobile communication, information technology, consumer electronics, and mobile automobiles due to its excellent energy storage characteristics. With the progress and development of the human society, advanced lithium-ion batteries are required to have higher capacity, better rate performance, and longer service life (Armand and Tarascon, 2008; Xin et al., 2012; Song and Zhou, 2013; Huang et al., 2016; Wang et al., 2016; Li et al., 2018; Tang et al., 2018; Lu et al., 2019; Shin et al., 2019). Among all the components of lithium-ion battery, electrode materials are the key factors that restrict the performance of lithium-ion batteries. Among them, negative electrode material, as an important part of lithium ion battery, has an important influence on the electrochemical performance of lithium ion battery (Chen et al., 2016a,b; Wang et al., 2017; Maruyama et al., 2018; Xu et al., 2018, 2019; Zhu et al., 2018; Chen J. et al., 2019; Xiao et al., 2019). In lithium ion battery anode material, carbon material has the advantages of low electrode potential, high cycle efficiency, long cycle life, and good safety performance, makes it the preferred anode material for lithium-ion batteries. At present, graphite is a common carbon anode material, which has a good layered structure suitable for lithium ion embedding and deintercalation, which makes it show the advantages of high conductivity and high reversible specific capacity, and has become a widely used traditional commercial negative electrode material (Cameán and Garcia, 2011; Huang et al., 2012; Ni et al., 2014; Kim et al., 2017; Raccichini et al., 2017; Maruyama et al., 2018).
However, there are still some shortcomings in the traditional graphite anode materials, which seriously limit its application (Groult et al., 2005; Wu et al., 2013; Chen T. et al., 2019; Hashimoto et al., 2019; Kim et al., 2019). First of all, the theoretical capacity of graphite negative electrode is only 372 mAh·g−1, which is far from meeting the requirements of high performance lithium-ion batteries; secondly, the stability of layered structure is poor, which is easy to collapse after a long charge-discharge cycle, resulting in a serious decrease in specific capacity and a large reduction in energy storage life; thirdly, electrolyte decomposition will result in large irreversible capacity during the first charge-discharge process. These deficiencies largely limit the application of graphite anode materials in high-performance lithium-ion batteries. Therefore, a lot of work has been focused on finding new carbon anode materials with high performance of high capacity, stable layered structure, long cycle life, etc. (Chen et al., 2013; Kim et al., 2014; Hou et al., 2017; Huang et al., 2017; Yu et al., 2019). These research work provides a good idea for the development of high-performance new carbon anode materials. However, for the layered graphite anode materials, in addition to the above mentioned shortcomings, the interlayer spacing is small and Li+ can only be embedded from the end face of the material, which is bound to increase the diffusion resistance of lithium ion. As a result, the rate performance of layered graphite negative electrode is poor, which limits its application in high power battery (Kumagai et al., 2015; Libich et al., 2017; Dagger et al., 2018; Mazur et al., 2019). Therefore, improving the capacity and rate performance of carbon anode materials is an important issue to be solved urgently at present.
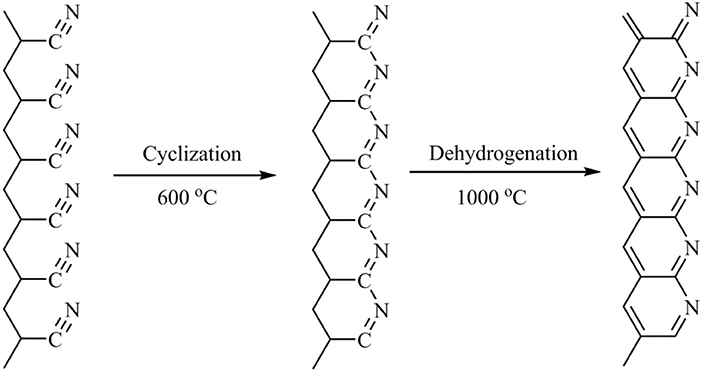
Herein, in this work, a new type of organic carbon material (Scheme 1) was developed to improve the rate performance of anode materials for lithium-ion batteries. The polyacrylonitrile (PAN) is cracked and carbonized at a high temperature of 1,050°C, and the PAN hard carbon is used as the negative electrode material of lithium ion battery, showing an initial capacity of 343.5 mAh g−1 which is equal to that of graphite electrode, and shows better cycle and rate performance compared with traditional graphite negative electrode, which will provide a new way for the development of novel carbon anode materials for lithium-ion batteries.
Experimental Section
Preparation of PAN Organic Carbon
Fifteen grams of polyacrylonitrile powder were paved on a porcelain boat, placed in a tubular furnace, raised to 280°C in oxygen atmosphere at 1°C/min, and kept for 2 h (pre-oxidation for 2 h), and then naturally lowered to room temperature to obtain stabilized PAN powder. The PAN powder sample was then heated to about 600°C in an atmosphere of nitrogen for 30 min. finally, raising the temperature to about 1,050°C by 3°C/min, continuously carbonizing for 30 min at the temperature, then naturally cooling to room temperature to obtain a 4.5 g of PAN hard carbon-based material. The obtained PAN organic carbon was characterized and confirmed by IR, XRD, SEM, and TEM images, and the results were consistent with the target products.
Characterization and Electrochemical Tests
A ZEISS Crossbeam 340 scanning electron microscopy analyzer was used to capture the Scanning electron microscopy (SEM) images. The TEM images were obtained by a FEI Tecnai G2F20 transmission electron microscope. Infrared spectra were measured on a Fourier transform infrared (FTIR) spectrophotometer Bruker IFS66/S. The X-ray diffraction (XRD) spectra were collected on a Bruker powder X-ray diffractometer with a Cu Kα radiation.
Before the electrochemical performance was evaluated, all cells were pretreated by discharging them against lithium from an open-circuit voltage (OCV) to 0.001 V at a current of 0.1 mA and eventually charging them against lithium to 1.0 V at the same rate. The electrochemical performance was evaluated using an Arbin battery test station (BT2000, Arbin Instruments, College Station, TX, USA). Cyclic voltammograms were recorded at a scan rate of 0.1 mVs−1 between 0.001 and 2.5 V for half cells and between 2.5 and 4.75 V for full cells using a Solatron 1260/1287 Electrochemical Interface (Solatron Metrology, Bognor Regis, UK). Impedance analysis was also assessed using a Solatron 1260/1287 Electrochemical Interface kit.
Preparation of Electrode Sheet and Battery
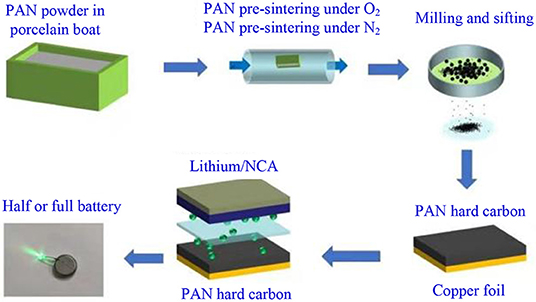
The preparation of electrode sheet and the assembly process of battery are similar with that of our published work (Chen et al., 2016b; Xu et al., 2018, 2019; Zhu et al., 2018; Chen J. et al., 2019; Xiao et al., 2019), as shown in Scheme 2. PAN organic carbon powder, super P (SP), and poly(vinylidene fluoride) (PVDF) binder were mixed in a ratio of 85:10:5 by weight, and 35 mL of N-methylpyrrolidone (NMP) solvent was added to 4 g of this mixture. After mixing evenly, the obtained slurry was coated onto the surface of copper foil, dried at 120°C for 2 h, and eventually dried for 24 h in a vacuum oven at 60°C. The film was cut into circular pieces with a diameter of 1.2 cm, and the area mass loading of the electrode was ~5.2 mg cm−2. Half-type Li-ion batteries were assembled with lithium metal as the counter electrode, and full-type Li-ion batteries were assembled with LiNi0.8Co0.15Al0.05O2 (NCA) as the counter electrode, with 1 M LiPF6 in a mixture of ethylene carbonate/diethyl carbonate (EC/DEC, 1:1 by volume) as the electrolyte, and Celgard®3501 (Celgard, Corp., LLC., Charlotte, NC USA) as the separator. In order to compare the performance of PAN organic carbon material with graphite negative electrode, half-battery, and full battery with graphite as electrode have been assembled as reference batteries.
Results and Discussion
Figure 1A shows the IR spectra of PAN samples carbonized at different temperatures. It can be clearly seen that the PAN pristine sample shows obvious characteristic peaks of functional groups. The sharp and strong peak at 2,245 cm−1 is attributed to the characteristic stretching vibration of the -CN group. The 2,941 cm−1 bands is ascribed to the characteristic stretching vibration of the –CH2 bond. The bands at 1,451 and 1,356 cm−1 are attributed to the characteristic bending vibration of the –CH2 and –CH groups. The bands at 1,227 and 1,059 cm−1 are attributed to the skeleton vibration of the C-C and C-N groups. However, after carbonation of PAN at different temperatures, it is found that the characteristic vibration peaks of those CH functional groups almost disappear, the IR curve is close to a smooth straight line with no apparent functional characteristic vibration peaks, indicating that PAN polymers are almost completely carbonized at high temperature, in which the organic bonds decompose and break to form PAN hard carbon. After high temperature carbonization of polypropylene cyanide, the resulting hard carbon contains C=C and C=N double bonds. however, as shown in the Scheme 1 molecular structure, the six-membered ring containing the double bonds of C=C and C=N converges together, and the vibration of its six-membered ring is implicated in the polymerization, making the signal of the characteristic peak generated weak. However, the weak vibration peaks can still be seen, as shown in Figure 1A, the weak bands at 1,061, 1,379, and 1,652 cm−1 might be attributed to the characteristic bending vibration of the C=C and C=N groups. Similar to the characteristic peaks in the IR spectrum of the polynaphthalene, the signal of the vibration peak becomes weaker when the six-membered ring is limited by the surrounding ring. It can also be seen from Figure 1A that there is no obvious difference in IR spectra at different carbonation temperatures. However, after careful observation of IR spectra, it is found that with the increase of carbonation temperature, the IR curve is smoother, indicating a higher degree of carbonation. The EDS spectrum (Figure S1) of PAN hard carbon shows that the average contents of C, N and O elements were 74.41, 20.97, and 4.62%, respectively, which is close to the theoretical value of element content in PAN polymer, indicating the full carbonation of PAN polymer.
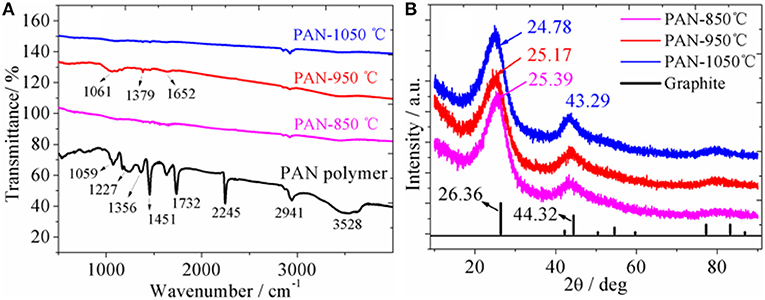
Figure 1. (A) IR spectra of PAN polymer carbonized at different temperatures; (B) XRD patterns of PAN polymer carbonized at different temperatures and standard patterns of graphite.
XRD patterns (Figure 1B) show that the graphite sample displays two main diffraction peaks (2θ = 26.36 and 44.32°), with lattice spacings of 3.38 and 2.04 Å, respectively. For the PAN hard carbon samples, two main broad prominent XRD diffraction peaks are also observed, indicating that PAN hard carbon has a amorphous structure with low crystallization degree. The XRD diffraction peaks are shifted to small angles with the increase of the carbonization temperature. For the PAN-1,050°C sample, the diffraction peaks at 24.78 and 43.29° with lattice spacings of 3.59 and 2.09 Å are observed. Compared with graphite, the diffraction angle of PAN hard carbon is smaller and the corresponding interlayer spacing is larger, which will be more beneficial to the transport of lithium ion.
Figure 2 shows the SEM and TEM images of PAN hard carbon and graphite samples. As can be seen from Figure 2a, the graphite samples are composed of relatively regular bulk particles of 10–15 μm, which is formed by stacking a plurality of graphite sheets. While for the PAN hard carbon sample, it is composed of extremely irregular shaped particles, which having a wide particle size distribution of between 5 and 30 μm (Figure 2b). This extremely irregular morphology is due to mechanical grinding of PAN hard carbon blocks after high temperature carbonization. At different carbonization temperatures, the surface morphology of PAN hard carbon is not very different (Figure S2), and with the increase of the carbonization temperature, the small particles adhered to the surface of the hard carbon particles produced are less, and the uniformity of the particles is better, which will be more helpful to electrochemical performance. Correspondingly, regular graphite particles show clear and obvious lattice fringes with lattice spacing of 0.335 nm (Figure 2c). However, as shown in Figure 2d, the irregular PAN hard carbon shows fuzzy and curved lattice stripes with a spacing of about 0.363 nm. The results suggest that graphite particles are layered structure with high crystallization degree, while PAN hard carbon is amorphous structure with low crystallization degree, and it is a carbon material with mixed crystallization and disordered structure, which is in good agreement with the analytical data of XRD diffraction peak as discussed in Figure 1B.
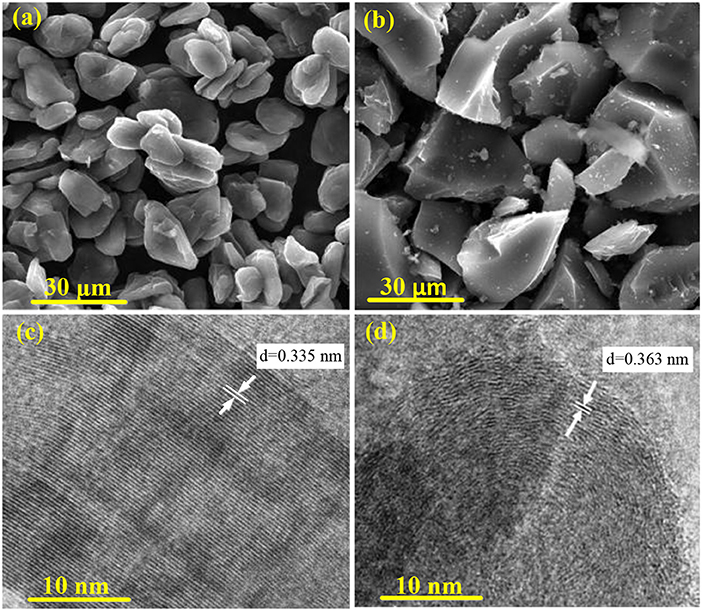

Figure 2. (a) SEM images of graphite sample; (b) SEM images of PAN hard carbon; (c) TEM images of graphite sample; (d) TEM images of PAN hard carbon.
Cyclic voltammograms were recorded at a scan rate of 0.1 mVs−1 between 0.001 and 2.5 V as shown in Figure 3A. For graphite electrodes, there is a significant oxidation-reduction peak at 0.29 and 0.15 V. Compared with graphite, the PAN hard-carbon electrode has no significant oxidation-reduction peak (Figure 3B), its CV curve has a wider area distribution to the capacity, indicating that the PAN hard carbon electrode has a wider voltage platform than that of graphite electrode, and its voltage platform is not flat but oblique. Figure 3C shows the EIS spectra of PAN hard carbon and graphite electrodes, it can be seen that the semicircular area of PAN hard carbon electrode is larger than that of graphite negative electrode in high frequency region, which indicates that PAN hard carbon electrode has greater impedance than graphite negative electrode. The PAN hard carbon electrode has a larger impedance value, which is mainly due to the following reasons. First, PAN hard carbon has larger particle size, so that the transmission and the migration of the electric charge inside the electrode material are more slow. Secondly, the C content in PAN hard carbon material is only about 74%, which is obviously lower than the carbon content in graphite material, which is also the reason for its poor electrical conductivity. Although the conductivity of the PAN hard carbon electrode is not advantageous, it shows a good initial charge and discharge performance. As shown in Figure 3D, the initial charging/discharging curves of PAN hard carbon and graphite electrode investigated from 0.001 to 2.5 V at a current rate of 0.2 C were investigated. For the PAN hard carbon electrode and graphite electrode, the charging capacities are 343.5 and 348.6 mAh g−1, respectively, with discharging capacities of 390.6 and 413.1 mAh g−1. The Coulombic efficiencies are 87.9 and 84.4% for the PAN hard carbon electrode and graphite electrode, respectively. The PAN hard carbon electrode shows a higher initial coulomb efficiency, indicating that the first irreversible capacity of the PAN hard carbon electrode is smaller, as a negative electrode material, which is very advantageous for the performance of the full batteries.
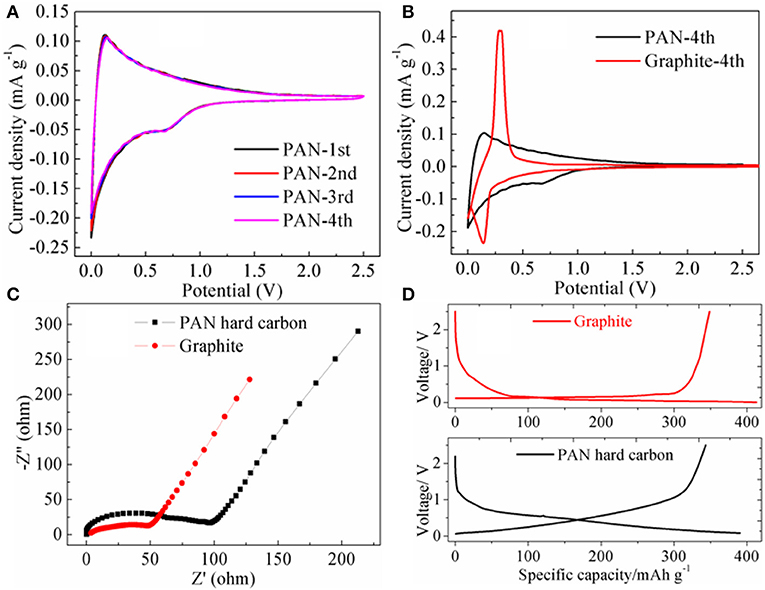
Figure 3. (A) Cyclic voltammetry curves of PAN hard carbon electrode; (B) the 4th CV curve comparison of PAN hard carbon and graphite electrodes; (C) EIS spectra of PAN hard carbon and graphite electrodes; (D) Initial charging/discharging curves of PAN hard carbon and graphite electrodes.
Rate charge and discharge performance of graphite and PAN hard carbon electrodes were investigated at different rate of 0.2, 1, 2, and 3 C, as shown in Figure 4. For the graphite electrode (Figure 4A), the discharge capacities are 348.6, 222.9, 135.9, and 92.0 mAh g−1 at current rates of 0.2, 1, 2, and 3 C, with charge capacities of 415.9, 230.5, 137.6, and 97.8 mAh g−1, respectively. The charge and discharge efficiencies at rates of 0.2, 1, 2, and 3 C are 83.8, 97.6, 98.8, and 94.1%, respectively. However, for the PAN hard carbon electrode (Figure 4B), the discharge capacities are 314.0, 220.3, 187.5, and 135.9 mAh g−1, and the charge capacities are 320.1, 219.0, 212.9, and 123.5 mAh g−1 at current rates of 0.2, 1, 2, and 3 C, with coulombic efficiencies of 98.1, 100.6, 88.1, and 110.0%, respectively. Although the charge-discharge capacity of PAN hard carbon electrode at 0.2 C is smaller than that of graphite electrode, the capacities of PAN hard carbon electrode is higher than that of graphite electrode at high rates, especially at 2C and 3C ratios. In addition, the charging and discharging coulomb efficiency of PAN hard carbon electrode is also higher than that of graphite electrode, which indicates that PAN hard carbon electrode has better rate performance.
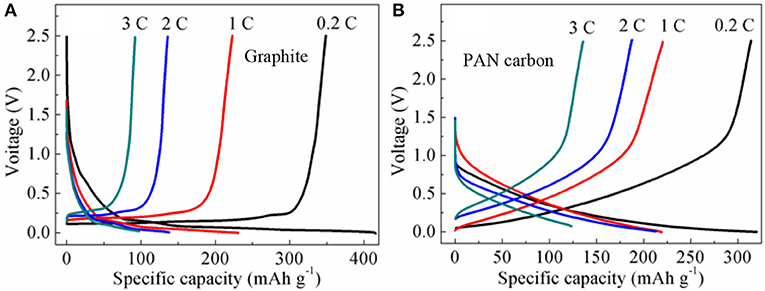
Figure 4. (A) Rate charge and discharge performance of graphite electrode; (B) Rate charge and discharge performance of PAN hard carbon electrode.
The advantage of rate performance for the PAN hard carbon electrode is also fully reflected in the rate cycle performance. As shown in Figure 5, rate cycle performance of PAN hard carbon and graphite electrodes at different current rates of 0.2, 1, 2, and 3 C were investigated. At 0.2 C, the initial charge capacities are 363.4 and 340.6 mAh g−1 for PAN hard carbon electrode and graphite electrode, respectively (Figure 5A). After 200 cycles, the capacities decreases to 284.3 and 267.9 mAh g−1, with capacity retention ratio of 78.2 and 78.6%, the results showed that the initial cycle performance of graphite was better at a low rate of 0.2 C, but the PAN hard carbon showed better cycle performance after more than 180 cycles. This performance trend is also well-reflected in 1 C current rate cycle performance. As shown in Figure 5B, although PAN hard carbon electrode shows smaller initial charging capacity, its cycle stability is better. After 300 cycles, the capacity increases, and the capacity retention rate reaches 111.9%, which is much higher than that of graphite electrode (73.4%). However, under 2 and 3 C rate currents, it can be clearly seen that the initial capacity of PAN hard carbon electrode is much higher than that of graphite electrode. Similarly, the PAN hard carbon electrode also shows more stable cycle performance than the graphite electrode. The charge capacity at 2 C are 187.5 and 142.3 mAh g−1 for the PAN hard carbon electrode and graphite electrode (Figure 5C). After 400 cycles, the capacity retention rate are 79.7 and 73.2% for the PAN hard carbon electrode and graphite electrode, respectively. The 3 C rate cycle performance also shows the same rule, as shown in Figure 5D, for the PAN hard carbon electrode and graphite electrode, the charge capacity at 3 C are 135.4 and 106.0 mAh g−1 with capacity retention rate of 88.6 and 44.8%, respectively. Compared with graphite electrode, PAN hard carbon electrode exhibits superior rate cycling performance. This is mainly due to the irregular structure of hard carbon materials is more beneficial to the intercalation and removal of lithium ions than the layered structure of graphite materials. As discussed in Figures 1, 2, compared with graphite material, the PAN hard carbon material is more irregular, long-range disorder and short-range order. Moreover, the interlayer spacing of the PAN hard carbon material is larger than that of the graphite layer, and the insertion and removal channels of the lithium ion are wider, which are more beneficial to the intercalation and the removal of the lithium ions. In addition. This is the reasons why PAN hard carbon material has better rate cycle performance than graphite material.
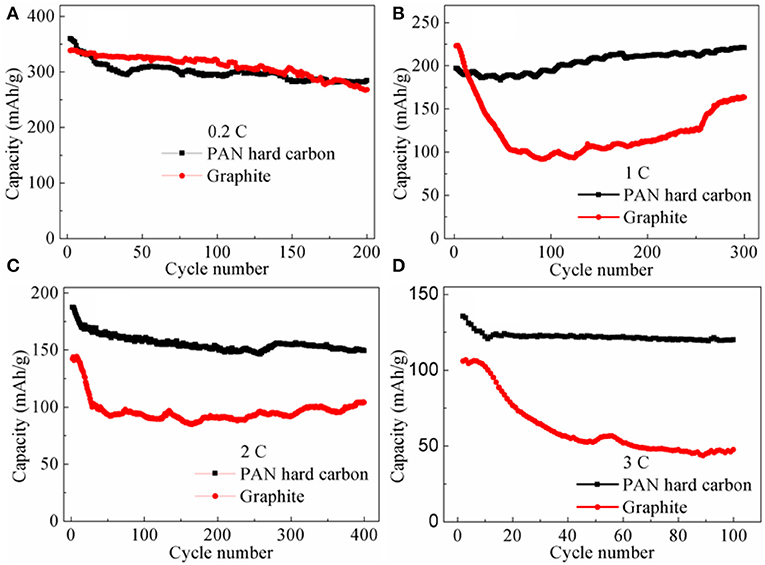
Figure 5. Rate cycle performance of PAN hard carbon electrode and graphite electrode at different current rates (A) 0.2 C; (B) 1 C; (C) 2 C; (D) 3 C.
To further demonstrate the advantages of the PAN hard carbon material, the full cell performance of an NCA/PAN hard carbon battery was studied using LiNi0.8Co0.15Al0.05O2 cathode material as a counter electrode. Figure 6A shows the initial charge and discharge performance of NCA/PAN hard carbon full cell, the charge and discharge capacities are 181.7 and 163.6 mAh g−1, respectively, with initial coulomb efficiency of 90.1%. The charge and discharge platform of the NCA/ PAN hard carbon full cell is not obvious, which is mainly related to the charge and discharge platform of the PAN hard carbon half-cell, which has been discussed in the above Figure 3D, the charge and discharge platform of the half-cell is a diagonal line and is not a significant horizontal platform, which leads to the discharging and charging platform of the full battery also showing oblique line, and there is no obvious platform. This can also be well-reflected in cyclic voltammetry curves, as shown in Figure 6B, the NCA/PAN hard carbon full cell exhibited a significant redox peak at voltages of 4.04 and 4.15 V, with no significant redox peaks below 4 V. Figure 6C shows the Cycle performance of NCA/PAN hard carbon full cell, an initial discharge capacity of 163.2 mAh g−1 is observed, and after 80 cycles, the discharge capacity decrease to 104.2 mAh g−1, the capacity retention rate is 64%. It can be seen that the early cycle capacity decay is fast, indicating that the irreversible capacity of the full cell is large.
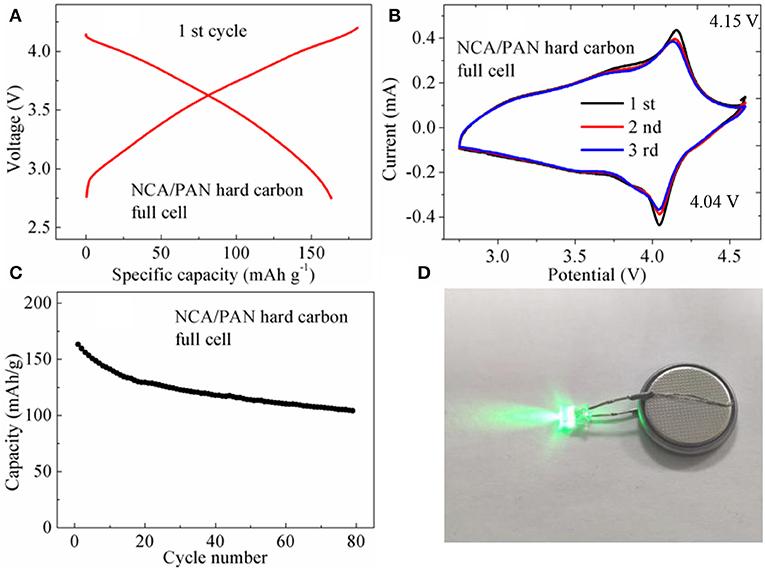
Figure 6. (A) Initial charge and discharge performance of NCA/PAN hard carbon full cell; (B) Cyclic voltammetry curves of NCA/PAN hard carbon full cell; (C) Cycle performance of NCA/PAN hard carbon full cell; (D) Physical diagram of full battery driving LED lamp.
As a new negative electrode material, when assembled with the positive electrode material into a full cell (Figure 6D), the formulation design of the electrode material, the capacity matching and the matching between the electrodes, is a process that takes a long time to explore. The PAN hard carbon electrode material in this work shows excellent cycle performance and rate performance. After further optimizing the formula and matching of the full cell, it is believed that the PAN hard carbon electrode material has a potential application prospect and will provide a new idea for the design of high performance new carbon negative electrode material.
Conclusion
After the PAN polymer is sintered at a high temperature of 1,050°C and then cracked and carbonized to obtain a novel organic hard carbon material (PAN hard carbon). The resulting hard-carbon material exhibits a disordered, amorphous structure with a wider layer spacing than graphite, providing a spacious channel for the embedding and removal of lithium ions. The obtained PAN hard carbon is used as the negative electrode material of lithium ion battery, showing an initial capacity which is equal to that of graphite electrode, and a higher initial coulomb efficiency than that of graphite electrode. Moreover, due to irregular amorphous structure, porous morphology and wider interlayer spacing, the PAN hard carbon electrode shows superior cycle stability and rate performance at different current rates compared with graphite electrode with regular crystalline layered structure. This new type of PAN hard carbon material with superior cycle and rate properties, which would provide a new idea for the development of novel negative electrode material with high performance.
Data Availability Statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation, to any qualified researcher.
Author Contributions
XR, YL, and HL synthesized PAN hard carbon, did test analysis experiment, data processing, and so on. JC and SZ designed the experimental, analyzed and discussed data, and wrote the article. BC, WW, HF, and HL were involved in some experiments, data processing, and so on.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The handling Editor declared a past co-authorship with one of the authors JC.
Acknowledgments
The authors would like to express their sincere thanks to the National Natural Science Foundation of China (21762019 and 51874151), the Science and Technology Project of Jiangxi Province (20161BAB213082 and 20171BAB206017), and the Program for Excellent Young Talents of JXUST.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fenrg.2020.00003/full#supplementary-material
References
Armand, M., and Tarascon, J. M. (2008). Building better batteries. Nature 451, 652–657. doi: 10.1038/451652a
Cameán, I., and Garcia, A. B. (2011). Graphite materials prepared by HTT of unburned carbon from coal combustion fly ashes: performance as anodes in lithium-ion batteries. J. Power Sources 196, 4816–4820. doi: 10.1016/j.jpowsour.2011.01.041
Chen, J., Guo, J. K., Zhang, T., Wang, C. X., Ding, N. W., Zhang, Q., et al. (2016a). Electrochemical properties of carbonyl substituted phthalocyanines as electrode materials for lithiumion batteries. RSC Adv. 6, 52850–52853. doi: 10.1039/C6RA09826F
Chen, J., Xu, Y., Cao, M. H., Zhu, C. J., Liu, X. L., Li, Y. T., et al. (2019). A stable 2D nano-columnar sandwich layered phthalocyanine negative electrode for lithium-ion batteries. J. Power Sources 426, 169–177. doi: 10.1016/j.jpowsour.2019.04.027
Chen, J., Zhang, Q., Zeng, M., Ding, N. W., Li, Z. F., and Zhong, S. W. (2016b). Carboxyl conjugated phthalocyanines used as novel electrode materials with high specific capacity for lithium-ion batteries. J. Solid State Electrochem. 20,1285–1294. doi: 10.1007/s10008-016-3126-6
Chen, T., Jia, W. S., Yao, Z. Y., Liu, Y. C., and Li, J. Z. (2019). Partly lithiated graphitic carbon foam as 3D porous current collectors for dendrite-free lithium metal anodes. Electrochem. Commun. 107:106535. doi: 10.1016/j.elecom.2019.106535
Chen, Y. M., Li, X. Y., Park, K., Song, J., Hong, J. H., Zhou, L. M., et al. (2013). Goodenough, hollow carbon-nanotube/carbon-nanofiber hybrid anodes for Li-Ion batteries. J. Am. Chem. Soc. 135, 16280–16283. doi: 10.1021/ja408421n
Dagger, T., Kasnatscheew, J., Vortmann-Westhoven, B., Schwieters, T., and Schappacher, F. M. (2018). Performance tuning of lithium ion battery cells with area-oversized graphite based negative electrodes. J. Power Sources 396, 519–526. doi: 10.1016/j.jpowsour.2018.06.043
Groult, H., Nakajima, T., Perrigaud, L., Ohzawa, Y., and Kumagai, N. (2005). Surface-fluorinated graphite anode materials for Li-ion batteries. J. Fluor. Chem. 126, 1111–1116. doi: 10.1016/j.jfluchem.2005.03.014
Hashimoto, H., Muramatsu, Y., Nishina, Y., and Asoh, H. (2019). Bipolar anodic electrochemical exfoliation of graphite powders. Electrochem. Commun. 104:106475. doi: 10.1016/j.elecom.2019.06.001
Hou, H. S., Qiu, X. Q., Wei, W. F., Zhang, Y., and Ji, X. B. (2017). Carbon anode materials for advanced sodium-ion batteries. Adv. Energy Mater. 7:201602898. doi: 10.1002/aenm.201602898
Huang, B., Xu, B. Y., Li, Y. T., Zhou, W. D., You, Y., Zhong, S. W., et al. (2016). Li-Ion Conduction and Stability of Perovskite Li3/8Sr7/16Hf1/4Ta3/4O3. ACS Appl. Mater. Interfaces 8, 14552–14557. doi: 10.1021/acsami.6b03070
Huang, J. T., Lin, Y. M., Ji, M. W., Cong, G. T., and Xu, J. (2017). Nitrogen-doped porous carbon derived from foam polystyrene as an anode material for lithium-ion batteries. Appl. Surf. Sci. 2019:144398. doi: 10.1016/j.apsusc.2019.144398
Huang, X., Zeng, Z. Y., Fan, Z. X., Liu, J. Q., and Zhang, H. (2012). Graphene-based electrodes. Adv. Mater. 24, 5979–6004. doi: 10.1002/adma.201201587
Kim, D. S., Chung, D. J., Bae, J., Jeong, G., and Kim, H. (2017). Surface engineering of graphite anode material with black TiO2-x for fast chargeable lithium ion battery. Electrochim. Acta 258, 336–342. doi: 10.1016/j.electacta.2017.11.056
Kim, J., Park, K., Woo, H., Gil, B., and Park, B. (2019). Selective removal of nanopores by triphenylphosphine treatment on the natural graphite anode. Electrochim. Acta 326:134993. doi: 10.1016/j.electacta.2019.134993
Kim, K.-J., Lee, T. S., Kim, G. S., Lim, H., and Lee, S. M. (2014). A hard carbon/microcrystalline graphite/carbon composite with a core-shell structure as novel anode materials for lithium-ion batteries. Electrochim. Acta 135, 27–34. doi: 10.1016/j.electacta.2014.04.171
Kumagai, S., Ishikawa, T., and Sawa, N. (2015). Cycle performance of lithium-ion capacitors using graphite negative electrodes at different pre-lithiation levels. J. Energy Storage 2, 1–7. doi: 10.1016/j.est.2015.05.005
Li, Y. T., Chen, X., Dolocan, A., Cui, Z. M., Xin, S., Xue, L. G., et al. (2018). Garnet electrolyte with an ultralow interfacial resistance for Li-Metal batteries. J. Am. Chem. Soc. 140, 6448–6455 doi: 10.1021/jacs.8b03106
Libich, J., Máca, J., Vondrák, J., Cech, O., and Sedlaríková, M. (2017). Irreversible capacity and rate-capability properties of lithium-ion negative electrode based on natural graphite. J. Energy Storage 14, 383–390. doi: 10.1016/j.est.2017.03.017
Lu, Y., Hou, X. S., Miao, L. C., Li, L., Shi, R. J., Liu, L. J., et al. (2019). Cyclohexanehexone with ultrahigh capacity as cathode materials for lithium-ion batteries. Angew. Chem. Int. Ed. 58, 7020–7024. doi: 10.1002/anie.201902185
Maruyama, S., Fukutsuka, T., Miyazaki, K., and Abe, T. (2018). Observation of the intercalation of dimethyl sulfoxide-solvated lithium ion into graphite and decomposition of the ternary graphite intercalation compound using in situ Raman spectroscopy. Electrochim. Acta 265, 41–46. doi: 10.1016/j.electacta.2018.01.035
Mazur, P., Mrlik, J., Pocedic, J., Vrana, J., and Bystron, T. (2019). Effect of graphite felt properties on the long-term durability of negative electrode in vanadium redox flow battery. J. Power Sources 414, 354–365. doi: 10.1016/j.jpowsour.2019.01.019
Ni, S. B., Lv, X. H., Zhang, J. C., Ma, J. J., Yang, X. L., and Zhang, L. L. (2014). The electrochemical performance of lithium vanadate/natural graphite composite material as anode for lithium ion batteries. Electrochim. Acta 145, 327–334. doi: 10.1016/j.electacta.2014.09.018
Raccichini, R., Varzi, A., Wei, D., and Passerini, S. (2017). You have full text access to this onlineopen articlecritical insight into the relentless progression toward graphene and graphene-containing materials for lithium-ion battery anodes. Adv. Mater. 29:160321. doi: 10.1002/adma.201603421
Shin, D.-Y., Park, D.-H., and Ahn, H.-J. (2019). Interface modification of an Al current collector for ultrafast lithium-ion batteries. Appl. Surf. Sci. 475, 519–523. doi: 10.1016/j.apsusc.2019.01.016
Song, Z. P., and Zhou, H. S. (2013). Towords sustainable and versatile energy storge devices: an overview of organic electrode materials. Energy Environ. Sci. 6:2280. doi: 10.1039/c3ee40709h
Tang, M., Zhu, S., Liu, Z., Jiang, C., Wu, Y., Li, H., et al. (2018). Tailoring pi-conjugated systems: from pi-pi stacking to high-rate-performance organic cathodes. Chem 4, 2600–2614. doi: 10.1016/j.chempr.2018.08.014
Wang, C., Jiang, C., Xu, Y., Liang, L., Zhou, M., Jiang, J., et al. (2016). A selectively permeable membrane for enhancing cyclability of organic sodium-ion batteries. Adv. Mater. 28, 9182–9187. doi: 10.1002/adma.201603240
Wang, Y. K., Chen, J., Jiang, C. C., Ding, N. W., Wang, C. X., Li, D., et al. (2017). Tetra-β-nitro-substituted phthalocyanines: a new organic electrode material for lithium batteries. J. Solid State Electrochem. 21, 947–954. doi: 10.1007/s10008-016-3419-9
Wu, M. H., Chen, J., Wang, C., Wang, F. Q., and Yi, B. L. (2013). Non-graphitic PPy-based carbon nanotubes anode materials for lithium-ion batteries. Electrochim. Acta 105, 462–467. doi: 10.1016/j.electacta.2013.05.019
Xiao, Z. E., Chen, J., Liu, J., Liang, T. X., Xu, Y., Zhu, C. J., et al. (2019). Microcrystalline copper foil as a high performance collector for lithium-ion batteries. J. Power Sources 438:226973. doi: 10.1016/j.jpowsour.2019.226973
Xin, S., Guo, Y. G., and Wan, L. J. (2012). Nanocarbon Networks for advanced rechargeable lithium batteries. Acc. Chem. Res. 45,1759–1769. doi: 10.1021/ar300094m
Xu, Y., Chen, J., Xiao, Z. E., Ou, C. X., Lv, W. X., Tao, L. H., et al. (2019). Porous diatomite-mixed 1,4,5,8-NTCDA nanowires as high-performance electrode materials for lithium-ion batteries. Nanoscale 11, 15881–15891. doi: 10.1039/C9NR06186J
Xu, Y., Chen, J., Zhu, C. J., Zhang, P. W., Jiang, G. X., Wang, C. X., et al. (2018). High-performance of sodium carboxylate-derived materials for electrochemical energy storage. Sci. China Mater. 61, 707–718. doi: 10.1007/s40843-017-9210-1
Yu, K. F., Wang, Y., Wang, X. F., Liu, W. P., and Liang, C. (2019). Preparation of porous carbon anode materials for lithium-ion battery from rice husk. Mater. Lett. 253, 405–408. doi: 10.1016/j.matlet.2019.07.126
Keywords: lithium-ion battery, polyphenylamine, organic carbon, anode, rate performance
Citation: Rao X, Lou Y, Chen J, Lu H, Cheng B, Wang W, Fang H, Li H and Zhong S (2020) Polyacrylonitrile Hard Carbon as Anode of High Rate Capability for Lithium Ion Batteries. Front. Energy Res. 8:3. doi: 10.3389/fenrg.2020.00003
Received: 27 November 2019; Accepted: 09 January 2020;
Published: 28 January 2020.
Edited by:
Yutao Li, University of Texas at Austin, United StatesReviewed by:
Biyi Xu, University of Texas at Austin, United StatesNan Wu, Beijing Institute of Technology, China
Chengliang Wang, Huazhong University of Science and Technology, China
Copyright © 2020 Rao, Lou, Chen, Lu, Cheng, Wang, Fang, Li and Zhong. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jun Chen, Y2hlbmp1bkBqeHVzdC5lZHUuY24=; Shengwen Zhong, emhvbmdzaHdAMTI2LmNvbQ==
 Xianfa Rao
Xianfa Rao Jun Chen
Jun Chen Weiting Wang
Weiting Wang