Abstract
Beach aquifers, located in the subsurface of sandy beaches, are unique ecosystems with steep chemical and physical gradients resulting from the mixing of terrestrial fresh groundwater and saline groundwater from the sea. While work has rapidly progressed to understand the physics and chemistry in this environment, much less is known about the microorganisms present despite the fact that they are responsible for vital biogeochemical processes. This paper presents a review of the current state of knowledge of microbes within beach aquifers and the mechanisms that control the beach aquifer microbiome. We review literature describing the distribution and diversity of microorganisms in the freshwater-saltwater mixing zone of beach aquifers, and identify just 12 papers. We highlight knowledge gaps, as well as future research directions: The understanding of beach aquifer microorganisms is informed primarily by 16S ribosomal RNA gene sequences. Metagenomics and metatranscriptomics have not yet been applied but are promising approaches for elucidating key metabolic and ecological roles of microbes in this environment. Additionally, variability in field sampling and analytical methods restrict comparison of data across studies and geographic locations. Further, documented evidence on the migration of microbes within the beach aquifer is limited. Taking into account the physical transport of microbes through sand by flowing groundwater may be critical for understanding the structure and dynamics of microbial communities. Quantitative measurements of rates of elemental cycling in the context of microbial diversity need further investigation, in order to understand the roles of microbes in mediating biogeochemical fluxes from the beach aquifer to the coastal ocean. Lastly, understanding the current state of beach aquifers in regulating carbon stocks is critical to foster a better understanding of the contribution of the beach aquifer microbiome to global climate models.
Introduction
Oceanic sandy beaches are dynamic environments teeming with life. Sandy beaches represent 31% of the world’s unfrozen shoreline (Luijendijk et al., 2018) and are defined as shorelines consisting of permeable, sandy sediments (Charette et al., 2005). Serving as functional links between land and sea, sandy beaches provide numerous invaluable ecosystem services (Rocha, 2008), including coastal protection (Hanley et al., 2014), filtration and purification of water (Brown and McLachlan, 2002), nutrient mineralization (Schlacher et al., 2007), storage and discharge of submarine groundwater (Kim and Heiss, 2021), and the provision of nursery and nesting areas for fish and bird species (Brown and McLachlan, 2002; Defeo et al., 2009). Beaches are also economically important (Hanley et al., 2014; Domínguez-Tejo et al., 2018) because they provide a place for recreation and fishing. In California, a location globally renowned for its beaches, revenue from beach-related tourism is estimated at $17.6 billion per year (Eastern Research Group, 2015).
During the past decade, interest in studying sandy beaches has increased (Dugan et al., 2010). One of the most rapidly progressing research areas at sandy beaches involves the study of beach aquifers (located in the subsurface of beaches) and submarine groundwater discharge from the beach aquifer to the sea (Robinson et al., 2018).
Characteristics of a Beach Aquifer
The beach aquifer is defined as the region below the surface of the sandy beach saturated with groundwater (Figure 1). Groundwater in the beach aquifer consists of land-derived freshwater and saline water of marine origin, and can be described by three distinct regions (Abarca et al., 2013): an intertidal saltwater cell (where seawater is flushed through the beach forced by waves and tides) (Robinson et al., 2006; Bakhtyar et al., 2012); an area of terrestrial freshwater (originating from the upland watershed) (Heiss et al., 2017); and a deep saltwater wedge (Heiss and Michael, 2014). The interface between the three water types is termed the subterranean estuary and has steep physical and chemical gradients, similar to a surface water estuary (Moore, 1999). Water that is discharged to the coastal ocean from the beach aquifer is referred to as submarine groundwater discharge (Robinson et al., 2018). Depending in part on the geomorphology of the beach, the relative spatial extents of the different regions in Figure 1 will vary. For example, the presence of an impermeable layer in the beach aquifer could appreciably affect this conceptual model.
FIGURE 1
Beach Aquifers as Biogeochemical Reactors and Important Pathways to the Coastal Ocean
The mixing of freshwater and saltwater in beach aquifer was first described in detail in the landmark work by Lebbe (1981). The interplay between physical and chemical gradients set up by the mixing of seawater and freshwater influences the resultant precipitation of minerals in the beach aquifer (Robinson et al., 2018). Although early works have reviewed the physical characteristics of beach aquifers fairly extensively (Michael et al., 2005; Vandenbohede and Lebbe, 2006), in recent years, geochemical and microbial properties of beach aquifer systems are gaining attention (Kim and Heiss, 2021). Physical characteristics such as terrestrial freshwater hydraulic gradients, waves and tidal inputs of seawater, and general geometry of the region influence the mixing of seawater and freshwater within the beach aquifer (Robinson et al., 2007; Beck et al., 2011; Abarca et al., 2013; Kim and Heiss, 2021). Among chemical characteristics within the beach aquifer (Charette and Sholkovitz, 2002; Charette et al., 2005; McAllister et al., 2015), land-derived freshwater can contribute elevated concentrations of nitrate, phosphate, dissolved silica and organic carbon (Abarca et al., 2013; Kim and Heiss, 2021); on the other hand, infiltrating seawater contributes oxygen, salt, sulfate, and organic matter (Kim and Heiss, 2021). The existence of physical and chemical gradients has been documented in several beaches around the world (Santoro et al., 2008; Beck et al., 2011; Robinson et al., 2018).
Within the beach aquifer, microbially mediated biogeochemical processes break down organic matter (Ahrens et al., 2020) thereby changing the oxidation state and form of nutrients (Santoro et al., 2006; Bone et al., 2007; Kroeger and Charette, 2008; McAllister et al., 2015), trace metals (Charette and Sholkovitz, 2002; Charette et al., 2005), carbon (Dorsett et al., 2011) and oxygen (Heiss et al., 2017). In fact, a wide range of biogeochemical reactions has been documented within beach aquifers (Heiss et al., 2017), including but not limited to nitrification (Ullman et al., 2003), denitrification (Santoro, 2010; Wegner et al., 2018), anammox (Slomp and Van Cappellen, 2004; Kroeger and Charette, 2008; Sáenz et al., 2012), iron oxidation-reduction (Charette and Sholkovitz, 2002; Beck et al., 2011; McAllister et al., 2015), manganese oxidation-reduction (Ahrens et al., 2020), sulfate oxidation-reduction (McAllister et al., 2015), organic carbon degradation (Kroeger and Charette, 2008; Sirois et al., 2018), and methanogenesis (Adyasari et al., 2020). Therefore, the beach aquifer can essentially be viewed as a “biogeochemical reactor” (Anschutz et al., 2009) where mixing of freshwater and saltwater has dramatic effects on microbial elemental cycling (Kroeger and Charette, 2008).
Work to date suggests that the above mentioned biogeochemical processes may modulate the flux of chemicals to the coastal ocean (Boehm et al., 2006; Santos et al., 2009; Kim et al., 2012; Santos et al., 2014; Yang et al., 2015; Robinson et al., 2018; Welch et al., 2019; McKenzie et al., 2020) when coupled to the inflow and outflow of groundwater to and from the beach aquifer as well as oceanic forcing (Kim and Heiss, 2021). Indeed some field studies demonstrate elemental fluxes from submarine groundwater discharge of beach aquifers that are on par with those from large nearby river systems (Moore, 2010).
While it is understood that microbes play a role in these various processes in the beach aquifer, there is a paucity of studies describing the distribution, diversity, and function of microbes there. Further, recent work has highlighted the importance of the interplay of physical and chemical processes in controlling the distribution and transformation of chemicals in the beach aquifer, and their flux to the coastal ocean (Robinson et al., 2018), but similar work exploring the added (and critical) role of microbes is scarce. Therefore, the goal of this study is to review the literature on microbes in the beach aquifer to identify knowledge gaps and future research directions. While this paper was in review, a complementary review by Ruiz-González et al. (2021) on the microbial dimension of submarine groundwater discharge was published.
Literature Review
We conducted literature searches on 23rd December 2020 using the web-of-science search terms [(beach or subterranean estuary) and microorganisms] and [submarine groundwater and beach] to identify published studies that have investigated the beach microbiome, and we also identified seemingly relevant references within those papers. Papers were included if they described the microorganisms in the beach aquifer. We excluded papers that were not specifically carried out in the aquifers of sandy beaches. Papers describing research in tidal flats, intertidal beach sands, subtidal sediments, or impermeable sediments were excluded, as they were not within the scope of this study.
We identified only 12 studies that have investigated the distribution, diversity, or function of microorganisms in the beach aquifer (Figure 2; Table 1). However, we identified a number of papers that examined microbial diversity in subtidal sands (sands consistently bathed in seawater at the bottom of the water column, yet not within the beach aquifer) and in intertidal sands (sands exposed to air at low tide and underwater at high tide, also not within the beach aquifer); we briefly review those papers here, due to their potential relevance to the beach aquifer microbiome given the close proximities of these environments and similar physical characteristics.
FIGURE 2
TABLE 1
| Source | Study site | Main findings | Sample type | Method |
|---|---|---|---|---|
| Hong et al. (2019) | Gloucester point, VA, United States | 1) aerobic-anaerobic transition zone (AATZ) had the highest diversity of microbes | Sediment core (100 cm in length), groundwater samples (between 25 cm and 100 cm below surface of sand, adjacent to sediment core) | Target:V4 |
| 2) AATZ was a hotspot for biogeochemical processes in the subterranean estuary (STE) | 16S rRNA-based illumina MiSeq sequencing | |||
| Jiang et al. (2020) | Beach in sanggou bay national | 1) active carbon turnover in the STE | Sediment core (22 cm in length), porewater samples (between 0 cm and 20 cm below surface of sand, adjacent to sediment core) | Target: V4-V5 |
| 2) seasonality affects microbial activity that stimulates DOC production and removal | 16S rRNA-based illumina MiSeq sequencing | |||
| 3) porewater flow rate influences DOC production and removal by microbes | — | |||
| 4) heterotrophic species abundance accelerates carbon transformation rates (in higher temperatures) | — | |||
| Degenhardt et al. (2020) | Spiekeroog island, Germany | 1) No clear correlation of the microbial community composition with STE salinity gradients along a sandy high-energy beach | Sediment core (100 cm in length), porewater samples (at 0 cm, 10 cm, 30 cm, 50 cm and 100 cm below surface of sand, adjacent to sediment core) | Target: V4-V5 |
| 2) equilibrium state of microbial diversity down to a depth of 1 m defined as “core” community | 16S rRNA-based illumina MiSeq sequencing | |||
| 3) prevalence of generalist microorganisms, adapted to quickly changing environmental conditions | — | |||
| Chen et al. (2020) | Shengsi island, China | 1) diversity of dominant bacteria in coastal well water and porewater is different | Well water, pore water (100 cm from surface) | Target: V4-V5 |
| 2) main species present in sample types participate in the degradation of organic pollutants and utilization of nitrate in STEs | 16S rRNA-based illumina MiSeq sequencing | |||
| Adyasari et al. (2020) | Mobile bay, gulf of Mexico | 1) higher microbial diversity was found in the coastal pore water than the surface water samples | Sediment core (150 cm from surface of sand), river water (from surface), inland groundwater (300 cm from surface), coastal pore water (2500 cm from surface), bay water (from surface) | Target: V4-V5 |
| 2) distinct archaeal communities dominated the peat-dominated and sandy-dominated SGD samples | 16S rRNA-based illumina MiSeq sequencing | |||
| Adyasari et al. (2019) | Jepara, Indonesia | 1) microbial community composition varied with salinity | Dug wells, pore water, seepage meters, river water, seawater (pore water and seepage meter samples were categorized as submarine groundwater discharge samples). All samples were collected with vertical profiles between 0 cm to 20 cm below surface of sand | Target: V4-V5 |
| 2) fecal indicators and potential pathogens were identified | 16S rRNA-based illumina MiSeq sequencing | |||
| 3) microbial community composition varied with hydrology | — | |||
| Chen et al. (2019) | Qinzhou bay, China | 1) microbial community composition varied with size fraction | Well water (at 4 m depth below surface of sand) | Target: V4-V5 |
| 2) some key microbial groups from the two size fractions were utilized in the transformation of carbon, nutrients and iron | 16S rRNA-based illumina MiSeq sequencing | |||
| 3) microbial communities have important roles in carbon, nitrogen and iron cycling in STEs | — | |||
| Lee et al. (2017) | Gongcheonpo beach, jeju island, korea | 1) less diverse microbial groups were detected in seawater-dominated samples when compared with groundwater-dominated samples | Coastal water samples, and black sand samples (collected according to tidal stage) | Target: V1-V3 |
| 2) significant relationships between environmental factors and microbial communities were observed | 16S rRNA gene-based pyrosequencing | |||
| 3) community composition varied significantly with tidal fluctuation | — | |||
| Ye et al. (2016) | Yellow sea coast, China | 1) less diverse bacterial groups were detected in well water samples when compared with those detected in pore water samples | Pore water in unconsolidated deposits (15 m from surface of sand), bedrock fissure water (0.3–0.4 m from surface of sand), beach coarse sand pore water (0.5 m–1.5 m from surface of sand) | Target: V4-V5 |
| 2) well water samples were associated with bacterial communities that were involved in nitrate cycling, while brackish porewater samples were associated with microbes involved in ammonium cycling | 16S rRNA-based illumina MiSeq sequencing | |||
| 3) bacterial communities were correlated with the salinity gradient | — | |||
| 4) distinct bacterial communities were detected among different porewater samples | — | |||
| McAllister et al. (2015) | Cape shores, lewes, Delaware | 1) microbial communities significantly influenced Fe and S mineralization | Porewater samples from long-term multilevel sampling wells (390 cm–3600 cm from surface of sand) | Target: V1-V3 SSU; 454 pyrosequencing |
| 2) hydrological fluctuations affect the location of Fe & S mineralization | ||||
| Santoro et al. (2006) | Huntington beach, California, United States | 1) first study characterizing N-cycling functional genes and denitrifier community composition in the beach aquifer system | 5 cubic cm of sediment, pore water | Target: nirK, nirS |
| 2) geochemistry (nitrate and salinity) plays an important role in shaping denitrifying bacterial communities | ABI 3100 or ABI 3730 capillary sequencer (PE applied biosystems) | |||
| 3) nonequilibrium state of coastal aquifers is likely playing a vital role in shaping the microbial community | — | |||
| Santoro et al. (2008) | Huntington beach, California, United States | 1) pioneering study documenting the abundance and community composition of both AOA and AOB (and nitrification genes) in the beach aquifer | 5 cubic cm of sediment, pore water | Target: Bacterial amoA, archaeal amoA |
| 2) salinity appeared to be responsible for the shift from an AOA-dominated community at low-salinities to an AOB-dominated community at the most marine site | ABI 3730XL capillary sequencer (PE applied biosystems) | |||
| 3) AOA diversity was high at all sites and time points, while AOB was extremely low | — |
Summary of findings on microbial communities in beach aquifers.
In subtidal sands, researchers have identified a diverse group of bacteria capable of many potential metabolic functions (Urakawa et al., 2000; Rusch et al., 2003; Hunter et al., 2006; Sørensen et al., 2007; Mills et al., 2008; Böer et al., 2009; Gaidos et al., 2011; Gobet et al., 2012). They show that despite their low carbon content, subtidal sands could harbor a diverse and robust microbiome. Beck et al. (2011) documented microbial abundance and metabolic rates in sediment cores in a tidal flat to contrast paleo-environmental imprints and modern-day processes.
In intertidal sands, a few studies have characterized the diversity of microbes associated with several potential metabolic functions, as well as potential human pathogens (Yamahara et al., 2007; Cui et al., 2013; Staley and Sadowsky, 2016; Romão et al., 2017). In these studies, the intertidal sands sampled were from a location where one might place a beach towel; many of these studies sought primarily to understand whether human pathogens are present in beach sands.
Below we summarize the 12 papers on microbial distribution, diversity, and function in the beach aquifer. Our summary is guided by the following questions 1) what do we know about the beach aquifer microbiome? 2) what are the biggest challenges for characterizing and reporting microbial communities and associated metabolisms in beach aquifers? and 3) which future research directions warrant attention?
What do We Know About the Beach Aquifer Microbiome?
Microbial Community Characteristics in the Beach Aquifer
Nine of the 12 studies investigated the microbial community using 16S rRNA amplicon sequencing (Table 2) and these studies found that Proteobacteria dominated all samples. Chloroflexi and Bacteroidetes were the next most dominant phyla. Other documented phyla were Planctomycetes, Acidobacteria, Actinobacteria, Firmicutes, and Vernucomicrobia. Within Proteobacteria, genus-level identification revealed members belonging to Alphaproteobacteria, Betaproteobacteria and Gammaproteobacteria, which varied in proportion among saline groundwater and brackish/fresh well water samples. Less commonly observed were genera belonging to Epsilonproteobacteria and Zetaproteobacteria (only noted in McAllister et al. (2015)). More commonly observed genera were Sphingobium, Pseudoalternomonas, Flavobacterium, Limnohabitans, Rhodobacter, Aquiluna, Pseudomonas, Marinobacterium, and Nitrospira. Although some genera were consistently observed globally, at the species-level, data were not provided in all the publications.
TABLE 2
| Study | Bacteria | Archaea | Taxonomic classification level |
|---|---|---|---|
| Hong et al. (2019) | Proteobacteria, Chloroflexi, Planctomycetes, Bacteroidetes, Acidobacteria, Cyanobacteria, unclassified phyla | Crenarchaeota, Euryarchaeota, Parvarchaeota, and unclassified phyla | Phylum |
| Jiang et al. (2020) | Proteobacteria, Actinobacteria, Bacteroidetes, and Firmicutes | — | Phylum |
| Pseudoalteromona, Marinobacter, Colwellia, Sphingobium, Thioprofundum, Gp10, and Ilumatobacter | Nitrosopumilus | Genus | |
| Degenhardt et al. (2020) | Acidobacteria, | — | Phylum |
| Actinobacteria, Chloroflexi, Planctomycetes, Proteobacteria, | |||
| Verrucomicrobia, Bacteroidetes, | |||
| Latescibacteria, and Firmicutes | |||
| Gillisia, Lewinella, Aquibacter, Sediminicola, Winogradskyella, Nitrospira, Fuerstia, Sulfurifustis, Persicirhabdus, Planococcus, Bacillus and Planomicrobium, and several unclassified genera | — | Genus | |
| Chen et al. (2020) | Gemmatimonadetes, Armatimonadia, Elusimicrobia, Cyanobacteria, SAR202_clade, Nitrospira, Acidobacteria, Planctomycetacia, Sphingobacteriia, deltaproteobacteria, cytophagia, Gammaproteobacteria, Flavobacteriia, Actinobacteria, Alphaproteobacteria, Betaproteobacteria | — | Class |
| Marivivens donghaensis, Pseudoalteromonas hodoensis, Pseudomonas pseudoalcaligenes, Acinetobacter populi, Pelagibacterium nitratireducens, Enterobacter tabaci, Flavobacterium jejuense, Erythrobacter citreus, Halomonas zhaodongensis, Vibrio toranzoniae, Acidovorax soil strain BL21, Rhodobacter ovatus, Novosphingobium naphthae | — | Species (OTUs with >98% similarity) | |
| Adyasari et al. (2020) | Acidimicrobiia, Actinobacteria, Alphaproteobacteria, Anaerolineae, Bacteroidia, Dehaloccoidia, Deltaproteobacteria, Gammaproteobacteria, Holophagae, Methanomicrobia, Oxyphotobacteria, Parcubacteria, Planctomycetacia, Thermodesulfovibrionia, Thermoleophilia, Verrucomicrobiae, unclassified | — | Class |
| Acinetobacter, Catenococcus, Cutibacterium, Cyanobium, Flavobacterium, Gaiellales, Massilia, Mesorhizobium, Methylobacter, Nocardioides, Pirellulaceae, Polynucleobacter, Rheinheimera, Sphingomonas, Spirosoma, Turicella, Verrucomicrobium, Vogesella, unclassified | — | Genus | |
| Adyasari et al. (2019) | Gammaproteobacteria, Alphaproteobacteria, Bacteroidia, Oxyphotobacteria, Actinobacteria | — | Class |
| Burkholderiaceae, Novosphingobium, Limnohabitans, Pseudarcicella, Sediminibacterium, Methylomonas, Methylomonaceae, Flavobacterium, Marivivens, Acidovorax, Synechococcus, Marinobacterium, Halomonas | — | Genus | |
| Chen et al. (2019) | Limnohabitans, Gallionella, Terrimonas, Perlucidibaca, Ferruginibacter, Sphingobium, Polynucleobacter, Pseudomonas, Sphingopyxis, Simiduia, Flavobacterium, Novosphingobium | Methanobacterium, Methanosaeta, unclassified Parvarchaeota, unclassified Bathyarchaeota, unclassified Woesearchaeota | Genus |
| Limnohabitans australis, Limnohabitans parvus, Novosphingobium pentaromativorans, Sphingopyxis soli, Flavobacterium chungnamense, Flavobacterium columnare, Polynucleobacter necessarius, Pseudomonas extremorientalis, Perlucidibaca piscinae, Acinetobacter vivianii, Caulobacter virbioides, Sphingobium cupriresistens, Ferruginibacter profundus, Terrimonas lutea, Methanococcus mairipludis | Methanosaeta concilli, Methanobacterium lacus, Nitrosopumilus maritimus, several other OTUs with > 98% similarity with clones found in groundwater from studies listed in Chen et al. (2019) | Species (OTUs with >98% similarity) | |
| Lee et al. (2017) | Alphaproteobacteria, Gammaproteobacteria, Flavobacteria, Betaproteobacteria, Sphingobacteria, Planctomycetacia, Deltaproteobacteria, Chloracidobacterium, Chroobacteria, Anaerolineae, Acidimicrobiia, Rhotdothermus, Bacteroidia, Coscinodiscophyce, Actinobacteria, Unclassified | — | Class |
| Loktanella, Pelagibacter, Roseovirus, Polaribacter, Shwanella, Pseudomonas, Flavobacterium, Sterolibacterium, Blastocella, Dechloromonas | — | Genus | |
| Ye et al. (2016) | Proteobacteria, Actinobacteria, Bacteroidetes, Planctomycetes, Acidobacteria, Cyanobacteria | — | Phylum |
| Comamonas, Limnohabitans, Flavobacterium, Flectobacillus, Novosphingobium, Rhodobacter, Sediminibacterium, Perlucidibaca, Aquabacterium, Hydrogenophaga, Simplicispira, Fluviicola, Pseudarcicella, Polynucleobacter, Winogradskyella, Chryseolinea, Blastopirellula, Acinetobacter, Prochlorococcus, Rhodopirellula, Nitrospira, Lentibacter, Erythrobacter, Leisingera, Loktanella, Glaceicola | — | Genus | |
| Rhodoluna lacicola, Rhodoluna limnophila, Aquiluna rubra, Polynucleobacter acidophobus, Calothrix desertica, Karenia brevis, Prochlorococcus marinu, Vibrio splendidus, Pseudoalteromonas marina, Rhodopirellula baltica | — | Species (OTUs with >98% similarity) | |
| McAllister et al. (2015) | Deltaproteobacteria, Epsilonproteobacteria, Gammaproteobacteria, Betaproteobacteria, Zetaproteobacteria | — | Class |
| Desulfovibrio, Desulfopila, Desulfurispora, Geopsychobacter, Geobacter, Sulfurimonas, Acidiferrobacter, Ferritrophicum, Thioclava, Mariprofundus, Ferritrophicum | — | Genus | |
| Santoro et al. (2008) | Betaproteobacteria (Nitrosospira-like) | Thaumarchaeota (Nitrosopumilus- and Nitrosarchaeum-like) | amoA-based OTUs with >95% identity |
| Santoro et al. (2006) | Unknown denitrifiying bacteria. Sequences were more closely related to other environmental sequences than known cultivated denitrifiers | — | nirK or nirS-based OTUs with >95% identity |
Summary of microbial communities in beach aquifers. The organisms listed were extracted from figures and tables provided in the original study, the lists include all the organisms in the study.
Four of the 12 studies characterized archaea in the beach aquifer. Santoro et al. (2008) characterized the archaeal community composition in a beach aquifer, focusing specifically on ammonia-oxidizing Thaumarchaeota. Other studies (Chen et al., 2019; Hong et al., 2019; Jiang et al., 2020) characterized the archaeal community composition in addition to the bacterial community composition using amplicon sequencing. Hong et al. (2019) documented Crenarchaeota (60%), Euryarchaeota (30%) as the dominant archaeal phyla and Parvarchaeota as a minor phylum at the study site. Chen et al. (2019) also noted Euryarchaeota as a dominant archaeal phylum, in addition to Bathyarchaeota, Woesearchaeota and Parvarchaeota. Jiang et al. (2020) noted a change in the abundance of Thaumarchaetoa to changing seasons in comparison with the major phylum–Proteobacteria.
Adyasari et al. (2019) documented the first evidence of the distribution of potential pathogens (1%-10% of total microbial community composition recorded) in beach aquifers, with potential implications on the water quality of the coastal ocean. The dominant genera were Vibrio, Prevotella, Staphylococcus, Leptospira, and members of the family Enterobacteriaceae.
To date, there is no evidence for the presence of endemic microbes in beach aquifer systems, although previous work on terrestrial aquifers suggests this may be a strong possibility (Anantharaman et al., 2016).
Functional Characteristics of the Beach Aquifer Microbiome
Five of the twelve studies (Santoro et al., 2006; Santoro et al., 2008; McAllister et al., 2015; Chen et al., 2019; Hong et al., 2019) aimed to provide substantive insights into the functional diversity of microbes in the beach aquifer. Two of those (Santoro et al., 2006; Santoro et al., 2008) investigated microorganisms involved in the nitrogen cycling using quantification and/or sequencing of functional genes involved in denitrification and nitrification, and three (McAllister et al., 2015; Chen et al., 2019; Hong et al., 2019) inferred microbial function from 16S rRNA amplicon sequencing.
Nitrogen cycling was investigated in the beach aquifer extensively by Santoro et al. (2006), Santoro et al. (2008). Santoro et al. (2006) characterized the diversity of denitrifying bacteria by sequencing nirS and nirK nitrite reductase genes (using Sanger sequencing). Little community compositional overlap between sampling sites was observed, suggesting high denitrifier diversity along small spatial scales (<40 m). Santoro et al. (Santoro et al., 2008) investigated the relative diversity and abundance of betaproteobacterial ammonia-oxidizing bacteria (AOB) and ammonia-oxidizing archaea (AOA) in the same beach aquifer using sequencing and quantitative polymerase chain reaction (qPCR) analysis of the functional gene encoding ammonia monooxygenase subunit A (amoA). Interestingly, a clear transition from an AOB-dominated community to an AOA-dominated community was observed as the location of the brackish mixing zone in the beach aquifer shifted seaward with a change in the season. This study also revealed a striking difference in relative bacterial and archaeal amoA gene diversity at all stations and time points. The study demonstrated an intimate link between microbial community composition, physicochemical gradients (e.g., especially salinity), and groundwater hydrology.
A broad suite of microbially mediated elemental cycling in the beach aquifer was investigated by McAllister et al. (2015), Hong et al. (2019), and Chen et al. (2019). McAllister et al. (2015) identified putative iron-cycling microbes (2.5% of the groundwater community) and sulfur-cycling microbes (29% of the groundwater community) based on the metabolic characteristics of the closely related (>98% OTU similarity) microbes documented in literature in other habitats. Hong et al. (2019) inferred metabolic functions (nitrogen, methane, and sulfur metabolism) from classified 16S rRNA OTUs using the bioinformatics program, PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States), along a redox gradient in the beach aquifer. Their results indicated the presence of an erobic-anaerobic transition zone where microbial diversity was the highest. Chen et al. (2019) also noted the dominance of the archaeal phylum Bathyarchaeota, which may play a significant role in anaerobic carbon metabolism in groundwater.
Linkages Between Environmental Factors and Microbial Community Composition in the Beach Aquifer
Among the 12 papers, authors investigate associations between the microbial community and various physical (temperature, size fraction, sediment lithology and sample retrieval depth) and chemical properties (pH, salinity, concentrations of dissolved oxygen, nitrogen, phosphorous, iron, silica, sulfur, and carbon) of the sampling location (summarized in Table 3).
TABLE 3
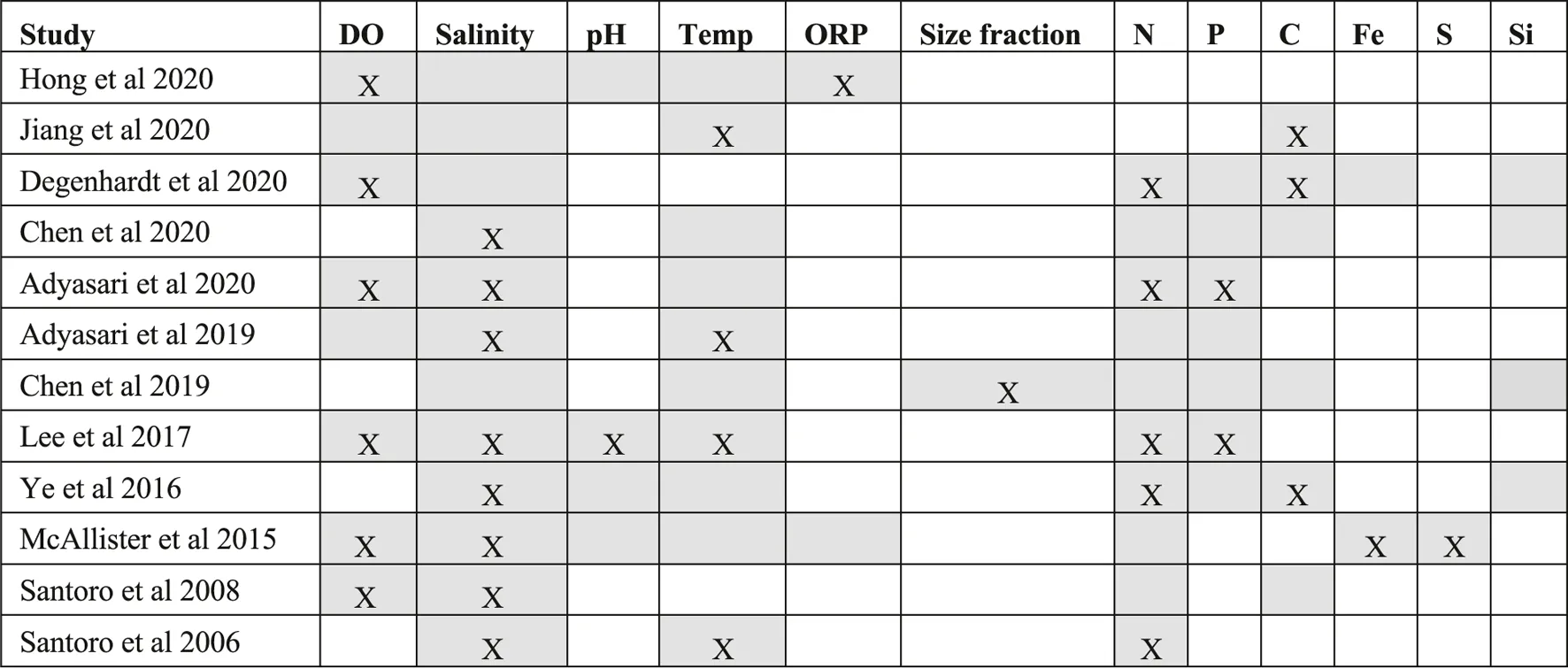 |
Relationships between beach aquifer microbiota and environmental parameters (Empty grey cells refer to measured environmental data, Grey cells with “X” indicates authors reported, statistically significant relationship between microbiome community composition and the environmental parameter; Empty white cells indicate parameters that were not measured by a given study. DO = dissolved oxygen, Temp = Temperature, ORP = Oxidation-Reduction Potential, Size fraction = Chen et al. (2019) was the only study that used cellulose acetate filters to generate two size fractions: 0.2–0.45 µm and >0.45 µm and study its relationship with the environment, N = nitrogen concentration, P = phosphorous concentration, C = carbon concentration, Fe = iron concentration, S = sulfide concentration, Si = silicate concentration).
Among the physical properties, authors presented evidence of links between the microbial community and sample retrieval depth from surface (Hong et al., 2019), size fraction (Chen et al., 2019), sediment lithology (Adyasari et al., 2020) and temperature (Santoro et al., 2006; Lee et al., 2017; Adyasari et al., 2019; Jiang et al., 2020). Temperature was the most commonly investigated physical parameter. Adyasari et al. (2019) found that temperature explained the patterns of microbial diversity. Lee et al. (2017) also observed a significant positive correlation between microbial community composition and temperature measurements, but only for groundwater samples collected at low tide. Although Santoro et al. (2006) measured temperature in the groundwater samples, selection for microbial communities and nitrogen cycling genes based on this variable was not apparent. Temperature might also be linked to other environmental parameters such as subsurface salinity gradients (Geng et al., 2016). Complementarily, Jiang et al. (2020) observed a link between sediment microbial activity and carbon transformations in groundwater samples with higher temperatures.
Among the chemical properties, which were more extensively investigated compared to the physical properties, two of the most commonly reported factors that affected the composition of the microbiome were salinity and oxygen. Eight of the 12 studies (Table 3) reported a significant relationship between salinity and microbial community composition. Contrastingly, Degenhardt et al. (2020) did not observe a correlation between microbial community composition and salinity gradients in the aquifer of a sandy high-energy beach in Spiekeroog Island. Hong et al. (2019) also did not observe clear patterns of bacterial or archaeal community shifts with variation in salinity. Chen et al. (2019) and Jiang et al. (2020) measured salinity, but neither study reported any significant relationships with microbes within the beach aquifer systems. In this case, salinity could be tightly linked to other environmental parameters such as concentration of organic matter, temperature, and pH (Rath et al., 2019). For example, when salinity decreases, it promotes the adsorption of dissolved organic carbon to sediment and may cause a reduction in groundwater dissolved organic carbon concentration (Setia et al., 2013; Jiang et al., 2020). Contrastingly, when salinity increases, groundwater dissolved organic carbon concentration may increase (Setia et al., 2013; Jiang et al., 2020).
Six of the 12 studies (Table 3) concluded that dissolved oxygen plays a vital role in regulating microbial distribution and function. Three studies highlighted the importance of dissolved oxygen in governing the distribution of bacterial communities (Santoro et al., 2008; McAllister et al., 2015; Degenhardt et al., 2020). One study (Hong et al., 2019) showed a dissolved oxygen profile drove the distribution of distinct erobic microbial groups (Gammaproteobacteria, Betaproteobacteria, Acidobacteria, Flavobacteria, Thaumarchaeta) and anaerobic microbial groups (sulfate-reducing bacteria, methanogenic archaea). Adyasari et al. (2020) also observed changes in microbial community composition with oxygen distribution. Lastly, Lee et al. (2017) showed that microbial community composition was significantly correlated to dissolved oxygen concentration, but only in samples collected at high tide.
What Are the Biggest Knowledge Gaps and Data Needs for Characterizing and Reporting Microbial Communities and Associated Metabolisms in Beach Aquifers?
Lawton (1999) stated “although details of single organisms matter and are of great interest, ecologists would profit most from uncovering underlying patterns, rules and laws”. Beach aquifers are vulnerable global hotspots for microbe-mediated biogeochemical cycling. Yet, there are limited studies to-date that characterize the microbial community (“who is there?”) in the beach aquifer. Questions that remain unanswered are “what are they doing there?” and “how did they get there?” Further, biogeochemical cycling estimates in beach aquifers remain fragmented. Therefore, the biggest challenge is the lack of data on the controls of the beach aquifer microbiome, or the “rules of life” of this environment. For the purpose of this study, we have identified knowledge gaps informed by our review and highlight corresponding data needs that warrant attention for future research.
Going Beyond Community Characteristics
Although the 12 studies identified in our review are extremely valuable, there is plenty of room for further investigation into the “potential function” of the microbiome using metagenomics (where the total community DNA is sequenced) and the “actual function” (or “expressional activity”) of the microbiome using metatranscriptomics (where the total community mRNA is sequenced). Recently, Anantharaman et al. (2016) applied metagenomic sequencing, assembly, and binning approaches to examine the genetic and metabolic potential of sediment and groundwater bacteria and archaea within a shallow aquifer near the Colorado River. Results demonstrating tremendous novelty in the aquifer system highlight the potential of this approach for biological discovery in subsurface beach aquifers as well. Moran et al. (2013) argued that instantaneous inventories of mRNA can be highly informative about ecologically relevant processes. When metagenomics is coupled with metatranscriptomics, these approaches can advance beach aquifer microbial diversity/function research.
Complexities in Sampling
Among the studies identified in our review, there was considerable variability in the sample type, sampling method, frequency, depth, location, and storage. First, the studies reviewed undertook sampling at different points along the shoreline (defined as mean sea level). Most samples were collected within ±50 m from the shoreline. Only a few studies sampled beyond ±100 m from the shoreline (Ye et al., 2016; Chen et al., 2019; Degenhardt et al., 2020). Second, several types of samples were collected for microbial community analysis with varying terminologies–e.g., well water, groundwater, pore water, sediment. Well water, groundwater, and pore water represent water collected from within the porous media of the beach aquifer which we refer to as “groundwater” here. Third, samples were collected at varying depths beneath the surface of the sand. Often, when sediment cores were used to sample sediment, water adjacent to the sediment core was also sampled at varying depths. Sediment cores were usually 100 cm in length, except in Jiang et al. (2020) where they were 22 cm in length. Groundwater was extracted from anywhere between 0 cm and 3600 cm below the sand surface and were obtained using push-point piezometers and peristaltic pumps or pre-rinsed polyethylene syringes (Charette and Allen, 2006), except one study that used an in-situ profiler (0–20 cm) (Ibánhez et al., 2011). Fourth, the frequency of sampling varied–from hourly to weekly and monthly. Finally, once collected, samples were stored at −20°C (Ye et al., 2016; Adyasari et al., 2019; Chen et al., 2020; Degenhardt et al., 2020; Jiang et al., 2020), or −80°C (Santoro et al., 2006; Santoro et al., 2008; McAllister et al., 2015; Chen et al., 2019; Hong et al., 2019) or −70°C (Lee et al., 2017) for downstream analysis.
Free-Living Versus Attached Microbes in Beach Aquifer Systems
Among the papers we reviewed, the microbial ecology of the beach aquifers targeted prokaryotes in both groundwater (free-living) and sediments (attached). However, to date there are no direct comparisons between the free-living and attached microbial communities documented in coastal aquifer systems (Ruiz-Gonzalez et al., 2021). Specifically, no study has directly compared groundwater and sediment microbial communities in beach aquifer systems. Some works did report differences in the microbial communities with changes in particle size fraction (for example: Chen et al., 2019). Therefore, addressing the difference in the microbial diversity of suspended versus attached communities will form an important part of future microbiome research in beach aquifer systems.
Variability in Analytical Methods
The papers we reviewed relied heavily on 16S rRNA gene amplicon sequencing. Downstream analysis for microbial community composition and functional characteristics consistently targeted the hypervariable V4-V5 region of the 16S rRNA gene on the MiSeq Illumina sequencing platform, except McAllister et al. (2015) and Lee et al. (2017), which adopted tagged pyrosequencing of the V1-V3 region. However, the fusion-primers targeting the regions were different among the papers reviewed. We observed consistency in the usage of forward primers (515F: (Ye et al., 2016; Adyasari et al., 2019; Chen et al., 2019; Hong et al., 2019; Chen et al., 2020). However, the same studies used different reverse primers (926R–(Adyasari et al., 2019, 2020); 907R–(Ye et al., 2016; Chen et al., 2019, 2020); 541R–(Lee et al., 2017); 806R–(Hong et al., 2019)). Degenhardt et al. (Degenhardt et al., 2020) used a unique set of forward and reverse primers as per Klindworth et al. (2013). Overall, this variability in the primers and variable regions used for 16S rRNA amplicon sequencing makes comparisons across studies (and field sites) challenging. To date, no published studies have used metagenomics or metatranscriptomics to characterize the beach aquifer microbiome. Such studies have the potential to yield extremely valuable and unprecedented information on the metabolic functions of microbes in the beach aquifer, rather than relying solely on inferences made using 16S rRNA gene amplicon sequencing.
Metadata Reporting
Metadata reporting, including site description, access to the sediment, and relevant properties of the beach aquifer system, were fragmented or inconsistent across many of the studies identified in our review. For instance, not all studies report groundwater flow rate and/or velocity, and properties of the sand (such as texture, grain size) were often not mentioned. In cases where sediments are not homogenous and contain iron-containing minerals, it might not be possible to isolate the effect of groundwater chemistry on the microbiome, since sediment mineral composition co-varies with groundwater chemistry (Luo et al., 2018). Detailed sediment and site characterization can help document potential confounding effects.
Taxonomic Levels of Classification of Microbial Data
Some studies have chosen to report microbial community data only at the phylum or class level. While this provides high-level details regarding microbiome composition of beach aquifers, in the future, whenever possible it will be important to document evidence at the genus and/or species level to provide more insight into the function of microbiome. For example, there is increasing evidence to suggest the presence of aquifer microbes that are distinct from other microbes found in sediments or surface waters (e.g., iron-cycling, sulfur-cycling microbes; (Anantharaman et al., 2016; Probst et al., 2017; Probst et al., 2018; Kadnikov et al., 2020). Yet, it continues to remain unclear as to whether these differences are apparent at the genus and/or species level, as studies often report data at higher taxonomic levels of classification. Moreover, some phyla (e.g., Proteobacteria) are known to be extremely diverse with numerous members when compared to other phyla. Therefore, phylum-level classification may not provide a fine enough description of the community composition.
Microbial Transport Within the Beach Aquifer
It is likely that the microbiome is readily transported with water as it moves over wave-, tidal- and seasonal time scales through the beach aquifer. To date, experiments on microbial transport in the subsurface have focused primarily on the movement of pathogens in relation to the protection of water resources (de Sieyes et al., 2016). Boehm et al. (2014) tested whether the microbiome can be transported from seawater through unsaturated, intertidal sands and the vadose or unsaturated zone to the aquifer. While the groundwater surrounding the sediment may contain the same microorganisms as the sediment (suggesting a constant exchange of microbes between the sediment and groundwater), this does not mean that microorganisms in the groundwater can be readily transported through the subsurface. Surface-surface interactions caused by electrostatic, hydrophobic, and steric forces between sediments and microorganisms, or straining, may impede their transport through the porous media and cause their transport to be retarded (Bradford et al., 2002; Bradford et al., 2003; Bradford et al., 2013). Therefore, even if there is exchange of microbes between sediment and groundwater, this does not imply microbial transport can occur. Indeed, investigation into the potential for immigration of microbes between different regions (via transport through sands and adaptations to changes in groundwater chemistry) in the beach aquifer is required to understand the mechanisms controlling microbial structure.
Quantifying Rates of Elemental Cycling Within Beach Aquifer Systems
Data on the rates of key microbially mediated biogeochemical processes (e.g., nitrification/denitrification) in beach aquifer systems is lacking. For example, denitrification (the transformation of nitrate to nitrogen gas) in the beach aquifer is vital to reduce the nitrogen load in groundwater that gets discharged to the coastal ocean (Galloway et al., 2003). Organic carbon mineralization is tightly linked to nitrate-rich freshwater denitrification (Anschutz et al., 2009) and quantifying the reduction of sulfate to sulfide is also important, as sulfur cycling is closely linked to nitrogen cycling (Heiss et al., 2017). Therefore, nuances of the reactive and dynamic nature of the beach aquifer (quantitative measurements of rates of elemental cycling) in the context of its microbial diversity needs further investigation.
Beach Aquifers and the Global Carbon Budget
One of the emergent issues in beach aquifers is the behavior and lability of terrestrially derived dissolved organic matter, as well as the source-sink dynamics of carbon (Waska et al., 2021). In a thorough review on groundwater, Griebler and Lueders (2009) highlighted the important association between microbial communities and dissolved organic carbon as a driver of free-living versus attached communities in addition to other variables such as the availability of nutrients, sediment grain size and minerology. Further, Dittmar et al. (2001) noted that subsurface fluxes of organic matter are significant, yet quantitative estimates of transformations of organic matter through microbially mediated cycling need further investigation. This highlights the importance of beach aquifers as potential sources of organic and inorganic carbon and their need to be incorporated into global climate system models. It is possible that investigating carbon cycling in beach aquifers can potentially alter the global estimates of CO2 outgassing to the atmosphere, as current budgets underestimate the flux of CO2 from groundwater to the atmosphere (Ward et al., 2017).
Conclusion
Although there has been advancement in the knowledge and understanding of beach aquifer systems, there is presently a paucity of data on the beach aquifer microbiome. The mechanisms that control the beach aquifer microbiome or the “rules of life” for this environment need attention. Specifically, the extent to which the environment (groundwater chemistry, sediment characteristics) controls the beach aquifer system and the ability of the microbiome present naturally in the beach sediments to be transported to other locations in the aquifer remains largely unknown. Beach aquifer microbiome research will directly benefit society by furthering the understanding of the ecosystem services provided by beaches in terms of biogeochemical cycling. As beaches are threatened by changing environmental conditions such as sea-level rise, this information is essential to anticipate the overall contribution of beaches to biogeochemical cycling on the planet.
Statements
Author contributions
AB conceptualized the framework for the review. AA wrote the manuscript. AB, CF and AA reviewed the manuscript and analysis. All authors contributed to the article and approved the submitted version.
Funding
We are grateful to the National Science Foundation (NSF Emerging Frontiers Award Number:2024504) for funding the project “The sandy beach microbiome: physical, chemical and biological controls on diversity and function”.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1
AbarcaE.KaramH.HemondH. F.HarveyC. F. (2013). Transient Groundwater Dynamics in a Coastal Aquifer: The Effects of Tides, the Lunar Cycle, and the Beach Profile. Water Resour. Res.49, 2473–2488. 10.1002/wrcr.20075
2
AdyasariD.HassenrückC.MontielD.DimovaN. (2020). Microbial Community Composition across a Coastal Hydrological System Affected by Submarine Groundwater Discharge. PLoS One15, e0235235. 10.1371/journal.pone.0235235
3
AdyasariD.HassenrückC.OehlerT.SabdaningsihA.MoosdorfN. (2019). Microbial Community Structure Associated with Submarine Groundwater Discharge in Northern Java (Indonesia). Sci. Total Environ.689, 590–601. 10.1016/j.scitotenv.2019.06.193
4
AhrensJ.BeckM.MarchantH. K.AhmerkampS.SchnetgerB.BrumsackH. J. (2020). Seasonality of Organic Matter Degradation Regulates Nutrient and Metal Net Fluxes in a High Energy Sandy Beach. J. Geophys. Res. Biogeosci.125. 10.1029/2019JG005399
5
AnantharamanK.BrownC. T.HugL. A.SharonI.CastelleC. J.ProbstA. J.et al (2016). Thousands of Microbial Genomes Shed Light on Interconnected Biogeochemical Processes in an Aquifer System. Nat. Commun.7, 13219. 10.1038/ncomms13219
6
AnschutzP.SmithT.MouretA.DebordeJ.BujanS.PoirierD.et al (2009). Tidal Sands as Biogeochemical Reactors. Estuarine, Coastal Shelf Sci.84, 84–90. 10.1016/j.ecss.2009.06.015
7
BakhtyarR.BarryD. A.BrovelliA. (2012). Numerical Experiments on Interactions between Wave Motion and Variable-Density Coastal Aquifers. Coastal Eng.60, 95–108. 10.1016/j.coastaleng.2011.09.001
8
BeckM.RiedelT.GraueJ.KösterJ.KowalskiN.WuC. S.et al (2011). Imprint of Past and Present Environmental Conditions on Microbiology and Biogeochemistry of Coastal Quaternary Sediments. Biogeosciences8, 55–68. 10.5194/bg-8-55-2011
9
BoehmA. B.PaytanA.ShellenbargerG. G.DavisK. A. (2006). Composition and Flux of Groundwater from a California Beach Aquifer: Implications for Nutrient Supply to the Surf Zone. Continental Shelf Res.26, 269–282. 10.1016/j.csr.2005.11.008
10
BoehmA. B.YamaharaK. M.SassoubreL. M. (2014). Diversity and Transport of Microorganisms in Intertidal Sands of the California Coast. Appl. Environ. Microbiol.80, 3943–3951. 10.1128/AEM.00513-14
11
BöerS. I.HedtkampS. I. C.van BeusekomJ. E. E.FuhrmanJ. A.BoetiusA.RametteA. (2009). Time- and Sediment Depth-Related Variations in Bacterial Diversity and Community Structure in Subtidal Sands. Isme J.3, 780–791. 10.1038/ismej.2009.29
12
BoneS. E.CharetteM. A.LamborgC. H.GonneeaM. E. (2007). Has Submarine Groundwater Discharge Been Overlooked as a Source of Mercury to Coastal Waters?. Environ. Sci. Technol.41, 3090–3095. 10.1021/es0622453
13
BradfordS. A.MoralesV. L.ZhangW.HarveyR. W.PackmanA. I.MohanramA.et al (2013). Transport and Fate of Microbial Pathogens in Agricultural Settings. Crit. Rev. Environ. Sci. Tech.43, 775–893. 10.1080/10643389.2012.710449
14
BradfordS. A.SimunekJ.BettaharM.van GenuchtenM. T.YatesS. R. (2003). Modeling Colloid Attachment, Straining, and Exclusion in Saturated Porous Media. Environ. Sci. Technol.37, 2242–2250. 10.1021/es025899u
15
BradfordS. A.YatesS. R.BettaharM.SimunekJ. (2002). Physical Factors Affecting the Transport and Fate of Colloids in Saturated Porous Media. Water Resour. Res.38, 63–71. 10.1029/2002WR001340
16
BrownA. C.McLachlanA. (2002). Sandy Shore Ecosystems and the Threats Facing Them: Some Predictions for the Year 2025. Envir. Conserv.29, 62–77. 10.1017/S037689290200005X
17
CharetteM. A.AllenM. C. (2006). Precision Ground Water Sampling in Coastal Aquifers Using a Direct-Push, Shielded-Screen Well-Point System. Groundwater Monit. Remediation26, 87–93. 10.1111/j.1745-6592.2006.00076.x
18
CharetteM. A.SholkovitzE. R.HanselC. M. (2005). Trace Element Cycling in a Subterranean Estuary: Part 1. Geochemistry of the Permeable Sediments. Geochimica et Cosmochimica Acta69, 2095–2109. 10.1016/j.gca.2004.10.024
19
CharetteM. A.SholkovitzE. R. (2002). Oxidative Precipitation of Groundwater-Derived Ferrous Iron in the Subterranean Estuary of a Coastal Bay. Geophys. Res. Lett.29, 85–91. 10.1029/2001GL014512
20
ChenX.YeQ.DuJ.ZhangJ. (2019). Bacterial and Archaeal Assemblages from Two Size Fractions in Submarine Groundwater Near an Industrial Zone. Water11, 1261. 10.3390/w11061261
21
ChenX.YeQ.SandersC. J.DuJ.ZhangJ. (2020). Bacterial-derived Nutrient and Carbon Source-Sink Behaviors in a Sandy Beach Subterranean Estuary. Mar. Pollut. Bull.160, 111570. 10.1016/j.marpolbul.2020.111570
22
CuiH.YangK.PagalingE.YanT. (2013). Spatial and Temporal Variation in Enterococcal Abundance and its Relationship to the Microbial Community in Hawaii Beach Sand and Water. Appl. Environ. Microbiol.79, 3601–3609. 10.1128/aem.00135-13
23
de SieyesN. R.RussellT. L.BrownK. I.MohantyS. K.BoehmA. B. (2016). Transport of Enterococci and F+ Coliphage through the Saturated Zone of the Beach Aquifer. J. Water Health14, 26–38. 10.2166/wh.2015.290
24
DefeoO.McLachlanA.SchoemanD. S.SchlacherT. A.DuganJ.JonesA.et al (2009). Threats to Sandy Beach Ecosystems: A Review. Estuarine, Coastal Shelf Sci.81, 1–12. 10.1016/j.ecss.2008.09.022
25
DegenhardtJ.DlugoschL.AhrensJ.BeckM.WaskaH.EngelenB. (2020). Seasonal Dynamics of Microbial Diversity at a Sandy High Energy Beach Reveal a Resilient Core Community. Front. Mar. Sci.7, 573570. 10.3389/fmars.2020.573570
26
DittmarT.LaraR. J.KattnerG. (2001). River or Mangrove? Tracing Major Organic Matter Sources in Tropical Brazilian Coastal Waters. Mar. Chem.73, 253–271. 10.1016/S0304-4203(00)00110-9
27
Domínguez-TejoE.MetternichtG.JohnstonE. L.HedgeL. (2018). Exploring the Social Dimension of Sandy Beaches through Predictive Modelling. J. Environ. Manage.214, 379–407. 10.1016/j.jenvman.2018.03.006
28
DorsettA.CherrierJ.MartinJ. B.CableJ. E. (2011). Assessing Hydrologic and Biogeochemical Controls on Pore-Water Dissolved Inorganic Carbon Cycling in a Subterranean Estuary: A 14C and 13C Mass Balance Approach. Mar. Chem.127, 76–89. 10.1016/j.marchem.2011.07.007
29
DuganJ. E.DefeoO.JaramilloE.JonesA. R.LastraM.NelR.et al (2010). Give Beach Ecosystems Their Day in the Sun. Science329, 1146. 10.1126/science.329.5996.1146-a
30
Eastern Research Group (2015). “The National Significance of California’s Ocean Economy.” 2015. https://coast.noaa.gov/data/digitalcoast/pdf/california-ocean-economy.pdf (Accessed January 11, 2021).
31
GaidosE.RuschA.IlardoM. (2011). Ribosomal Tag Pyrosequencing of DNA and RNA from Benthic Coral Reef Microbiota: Community Spatial Structure, Rare Members and Nitrogen-Cycling Guilds. Environ. Microbiol.13, 1138–1152. 10.1111/j.1462-2920.2010.02392.x
32
GallowayJ. N.AberJ. D.ErismanJ. W.SeitzingerS. P.HowarthR. W.CowlingE. B.et al (2003). The Nitrogen Cascade. BioScience53, 341. 10.1641/0006-3568(2003)053[0341:TNC]2.0.CO;2
33
GengX.BoufadelM. C.JacksonN. L. (2016). Evidence of Salt Accumulation in Beach Intertidal Zone Due to Evaporation. Sci. Rep.6, 31486. 10.1038/srep31486
34
GobetA.BöerS. I.HuseS. M.van BeusekomJ. E. E.QuinceC.SoginM. L.et al (2012). Diversity and Dynamics of Rare and of Resident Bacterial Populations in Coastal Sands. Isme J.6, 542–553. 10.1038/ismej.2011.132
35
GrieblerC.LuedersT. (2009). Microbial Biodiversity in Groundwater Ecosystems. Freshw. Biol.54, 649–677. 10.1111/j.1365-2427.2008.02013.x
36
HanleyM. E.HoggartS. P. G.SimmondsD. J.BichotA.ColangeloM. A.BozzedaF.et al (2014). Shifting Sands? Coastal Protection by Sand Banks, Beaches and Dunes. Coastal Eng.87, 136–146. 10.1016/j.coastaleng.2013.10.020
37
HeissJ. W.MichaelH. A. (2014). Saltwater-freshwater Mixing Dynamics in a Sandy Beach Aquifer over Tidal, Spring-Neap, and Seasonal Cycles. Water Resour. Res.50, 6747–6766. 10.1002/2014WR015574
38
HeissJ. W.PostV. E. A.LaattoeT.RussonielloC. J.MichaelH. A. (2017). Physical Controls on Biogeochemical Processes in Intertidal Zones of Beach Aquifers. Water Resour. Res.53, 9225–9244. 10.1002/2017WR021110
39
HongY.WuJ.WilsonS.SongB. (2019). Vertical Stratification of Sediment Microbial Communities along Geochemical Gradients of a Subterranean Estuary Located at the Gloucester Beach of Virginia, United States. Front. Microbiol.9, 3343. 10.3389/fmicb.2018.03343
40
HunterE. M.MillsH. J.KostkaJ. E. (2006). Microbial Community Diversity Associated with Carbon and Nitrogen Cycling in Permeable Shelf Sediments. Appl. Environ. Microbiol.72, 5689–5701. 10.1128/aem.03007-05
41
IbánhezJ. S. P.LeoteC.RochaC. (2011). Porewater Nitrate Profiles in Sandy Sediments Hosting Submarine Groundwater Discharge Described by an Advection-Dispersion-Reaction Model. Biogeochemistry103, 159–180. 10.1007/s10533-010-9454-1
42
JiangS.ZhangY.JinJ.WuY.WeiY.WangX.et al (2020). Organic Carbon in a Seepage Face of a Subterranean Estuary: Turnover and Microbial Interrelations. Sci. Total Environ.725, 138220. 10.1016/j.scitotenv.2020.138220
43
KadnikovV. V.MardanovA. V.BeletskyA. V.KarnachukO. V.RavinN. V. (2020). Microbial Life in the Deep Subsurface Aquifer Illuminated by Metagenomics. Front. Microbiol.11, 572252. 10.3389/fmicb.2020.572252
44
KimK. H.HeissJ. W. (2021). Methods in Capturing the Spatiotemporal Dynamics of Flow and Biogeochemical Reactivity in Sandy Beach Aquifers: A Review. Water13, 782. 10.3390/w13060782
45
KimT.-H.WaskaH.KwonE.SuryaputraI. G. N.KimG. (2012). Production, Degradation, and Flux of Dissolved Organic Matter in the Subterranean Estuary of a Large Tidal Flat. Mar. Chem.142-144 (144), 1–10. 10.1016/j.marchem.2012.08.002
46
KlindworthA.PruesseE.SchweerT.PepliesJ.QuastC.HornM.et al (2013). Evaluation of General 16S Ribosomal RNA Gene PCR Primers for Classical and Next-Generation Sequencing-Based Diversity Studies. Nucleic Acids Res.41, e1. 10.1093/nar/gks808
47
KroegerK. D.CharetteM. A. (2008). Nitrogen Biogeochemistry of Submarine Groundwater Discharge. Limnol. Oceanogr.53, 1025–1039. 10.4319/lo.2008.53.3.1025
48
LawtonJ. H. (1999). Are There General Laws in Ecology?. Oikos84, 177. 10.2307/3546712
49
LebbeL. (1981). The Subterranean Flow of Fresh and Salt Water Underneath the Western Belgian Beach. 7th Salt Water Intrusion Meeting, Uppsala. Sveriges Geologiska Underso kning Rapporter och Meddelanden, 27, 193–219.
50
LeeE.ShinD.HyunS. P.KoK.-S.MoonH. S.KohD.-C.et al (2017). Periodic Change in Coastal Microbial Community Structure Associated with Submarine Groundwater Discharge and Tidal Fluctuation. Limnol. Oceanogr.62, 437–451. 10.1002/lno.10433
51
LuijendijkA.HagenaarsG.RanasingheR.BaartF.DonchytsG.AarninkhofS. (2018). The State of the World’s Beaches. Sci. Rep.8, 6641. 10.1038/s41598-018-24630-6
52
LuoW.GaoX.ZhangX. (2018). Geochemical Processes Controlling the Groundwater Chemistry and Fluoride Contamination in the Yuncheng Basin, China-An Area with Complex Hydrogeochemical Conditions. PLoS One13, e0199082. 10.1371/journal.pone.0199082
53
McAllisterS. M.BarnettJ. M.HeissJ. W.FindlayA. J.MacDonaldD. J.DowC. L.et al (2015). Dynamic Hydrologic and Biogeochemical Processes Drive Microbially Enhanced Iron and Sulfur Cycling within the Intertidal Mixing Zone of a Beach Aquifer. Limnol. Oceanogr.60, 329–345. 10.1002/lno.10029
54
McKenzieT.HollowayC.DulaiH.TuckerJ. P.SugimotoR.NakajimaT.et al (2020). Submarine Groundwater Discharge: A Previously Undocumented Source of Contaminants of Emerging Concern to the Coastal Ocean (Sydney, Australia). Mar. Pollut. Bull.160, 111519. 10.1016/j.marpolbul.2020.111519
55
MichaelH. A.MulliganA. E.HarveyC. F. (2005). Seasonal Oscillations in Water Exchange between Aquifers and the Coastal Ocean. Nature436, 1145–1148. 10.1038/nature03935
56
MillsH. J.HunterE.HumphrysM.KerkhofL.McGuinnessL.HuettelM.et al (2008). Characterization of Nitrifying, Denitrifying, and Overall Bacterial Communities in Permeable Marine Sediments of the Northeastern Gulf of Mexico. Appl. Environ. Microbiol.74, 4440–53. 10.1128/aem.02692-07
57
MooreW. S. (2010). A Reevaluation of Submarine Groundwater Discharge along the Southeastern Coast of North America. Glob. Biogeochem. Cycles24, a–n. 10.1029/2009GB003747
58
MooreW. S. (1999). The Subterranean Estuary: a Reaction Zone of Ground Water and Sea Water. Mar. Chem.65, 111–125. 10.1016/S0304-4203(99)00014-6
59
MoranM. A.SatinskyB.GiffordS. M.LuoH.RiversA.ChanL.-K.et al (2013). Sizing up Metatranscriptomics. ISME J.7, 237–243. 10.1038/ismej.2012.94
60
ProbstA. J.CastelleC. J.SinghA.BrownC. T.AnantharamanK.SharonI.et al (2017). Genomic Resolution of a Cold Subsurface Aquifer Community Provides Metabolic Insights for Novel Microbes Adapted to High CO2concentrations. Environ. Microbiol.19, 459–474. 10.1111/1462-2920.13362
61
ProbstA. J.LaddB.JarettJ. K.Geller-McGrathD. E.SieberC. M. K.EmersonJ. B.et al (2018). Differential Depth Distribution of Microbial Function and Putative Symbionts through Sediment-Hosted Aquifers in the Deep Terrestrial Subsurface. Nat. Microbiol.3, 328–336. 10.1038/s41564-017-0098-y
62
RathK. M.FiererN.MurphyD. V.RouskJ. (2019). Linking Bacterial Community Composition to Soil Salinity along Environmental Gradients. ISME J.13, 836–846. 10.1038/s41396-018-0313-8
63
RobinsonC. E.XinP.SantosI. R.CharetteM. A.LiL.BarryD. A. (2018). Groundwater Dynamics in Subterranean Estuaries of Coastal Unconfined Aquifers: Controls on Submarine Groundwater Discharge and Chemical Inputs to the Ocean. Adv. Water Resour.115, 315–331. 10.1016/j.advwatres.2017.10.041
64
RobinsonC.GibbesB.CareyH.LiL. (2007). Salt-freshwater Dynamics in a Subterranean Estuary over a Spring-Neap Tidal Cycle. J. Geophys. Res.112, C09007. 10.1029/2006JC003888
65
RobinsonC.GibbesB.LiL. (2006). Driving Mechanisms for Groundwater Flow and Salt Transport in a Subterranean Estuary. Geophys. Res. Lett.33, L03402. 10.1029/2005GL025247
66
RochaC. (2008). Sandy Sediments as Active Biogeochemical Reactors: Compound Cycling in the Fast Lane. Aquat. Microb. Ecol.53, 119–127. 10.3354/ame01221
67
RomãoD.StaleyC.FerreiraF.RodriguesR.SabinoR.VeríssimoC.et al (2017). Next-generation Sequencing and Culture-Based Techniques Offer Complementary Insights into Fungi and Prokaryotes in Beach Sands. Mar. Pollut. Bull.119, 351–358. 10.1016/j.marpolbul.2017.04.036
68
Ruiz-GonzálezC.RodellasV.Garcia-OrellanaJ. (2021). The Microbial Dimension of Submarine Groundwater Discharge: Current Challenges and Future Directions. FEMS Microbiol. Rev.10.1093/femsre/fuab010
69
RuschA.HuettelM.ReimersC. E.TaghonG. L.FullerC. M. (2003). Activity and Distribution of Bacterial Populations in Middle Atlantic Bight Shelf Sands. Fems Microbiol. Ecol.44, 89–100. 10.1111/j.1574-6941.2003.tb01093.x
70
SáenzJ. P.HopmansE. C.RogersD.HendersonP. B.CharetteM. A.SchoutenS.et al (2012). Distribution of Anaerobic Ammonia-Oxidizing Bacteria in a Subterranean Estuary. Mar. Chem.136-137 (137), 7–13. 10.1016/j.marchem.2012.04.004
71
SantoroA. E.BoehmA. B.FrancisC. A. (2006). Denitrifier Community Composition along a Nitrate and Salinity Gradient in a Coastal Aquifer. Aem72, 2102–2109. 10.1128/aem.72.3.2102-2109.2006
72
SantoroA. E.FrancisC. A.de SieyesN. R.BoehmA. B. (2008). Shifts in the Relative Abundance of Ammonia-Oxidizing Bacteria and Archaea across Physicochemical Gradients in a Subterranean Estuary. Environ. Microbiol.10, 1068–1079. 10.1111/j.1462-2920.2007.01547.x
73
SantoroA. E. (2010). Microbial Nitrogen Cycling at the Saltwater-Freshwater Interface. Hydrogeol J.18, 187–202. 10.1007/s10040-009-0526-z
74
SantosI. R.BryanK. R.PilditchC. A.TaitD. R. (2014). Influence of Porewater Exchange on Nutrient Dynamics in Two New Zealand Estuarine Intertidal Flats. Mar. Chem.167, 57–70. 10.1016/j.marchem.2014.04.006
75
SantosI. R.BurnettW. C.DittmarT.SuryaputraI. G. N. A.ChantonJ. (2009). Tidal Pumping Drives Nutrient and Dissolved Organic Matter Dynamics in a Gulf of Mexico Subterranean Estuary. Geochim. Cosmochim. Acta73, 1325–1339. 10.1016/j.gca.2008.11.029
76
SchlacherT. A.DuganJ.SchoemanD. S.LastraM.JonesA.ScapiniF.et al (2007). Sandy Beaches at the Brink. Divers. Distributions13, 556–560. 10.1111/j.1472-4642.2007.00363.x
77
SetiaR.RengasamyP.MarschnerP. (2013). Effect of Exchangeable Cation Concentration on Sorption and Desorption of Dissolved Organic Carbon in Saline Soils. Sci. Total Environ.465, 226–232. 10.1016/j.scitotenv.2013.01.010
78
SiroisM.CouturierM.BarberA.GélinasY.ChaillouG. (2018). Interactions between Iron and Organic Carbon in a Sandy Beach Subterranean Estuary. Mar. Chem.202, 86–96. 10.1016/j.marchem.2018.02.004
79
SlompC. P.Van CappellenP. (2004). Nutrient Inputs to the Coastal Ocean through Submarine Groundwater Discharge: Controls and Potential Impact. J. Hydrol.295, 64–86. 10.1016/j.jhydrol.2004.02.018
80
SørensenK. B.GlazerB.HannidesA.GaidosE. (2007). Spatial Structure of the Microbial Community in Sandy Carbonate Sediment. Mar. Ecol. Prog. Ser.346, 61–74.
81
StaleyC.SadowskyM. J. (2016). Regional Similarities and Consistent Patterns of Local Variation in Beach Sand Bacterial Communities throughout the Northern Hemisphere. Appl. Environ. Microbiol.82, 2751–2762. 10.1128/aem.00247-16
82
UllmanW. J.ChangB.MillerD. C.MadsenJ. A. (2003). Groundwater Mixing, Nutrient Diagenesis, and Discharges across a Sandy Beachface, Cape Henlopen, Delaware (USA). Estuarine, Coastal Shelf Sci.57, 539–552. 10.1016/S0272-7714(02)00398-0
83
UrakawaH.YoshidaT.NishimuraM.OhwadaK. (2000). Characterization of Depth-Related Population Variation in Microbial Communities of a Coastal Marine Sediment Using 16S rDNA-Based Approaches and Quinone Profiling. Environ. Microbiol.2, 542–554. 10.1046/j.1462-2920.2000.00137.x
84
VandenbohedeA.LebbeL. (2006). Occurrence of Salt Water above Fresh Water in Dynamic Equilibrium in a Coastal Groundwater Flow System Near De Panne, Belgium. Hydrogeol J.14, 462–472. 10.1007/s10040-005-0446-5
85
WardN. D.BianchiT. S.MedeirosP. M.SeidelM.RicheyJ. E.KeilR. G.et al (2017). Where Carbon Goes when Water Flows: Carbon Cycling across the Aquatic Continuum. Front. Mar. Sci.4. 10.3389/fmars.2017.00007
86
WaskaH.SimonH.AhmerkampS.GreskowiakJ.AhrensJ.SeibertS. L.et al (2021). Molecular Traits of Dissolved Organic Matter in the Subterranean Estuary of a High-Energy Beach: Indications of Sources and Sinks. Front. Mar. Sci.8, 607083. 10.3389/fmars.2021.607083
87
WegnerC.-E.GasparM.GeesinkP.HerrmannM.MarzM.KüselK. (2018). Biogeochemical Regimes in Shallow Aquifers Reflect the Metabolic Coupling of the Elements Nitrogen, Sulfur, and Carbon. Appl. Environ. Microbiol.85, e02346–18. 10.1128/AEM.02346-18
88
WelchE. M.DulaiH.El-KadiA.ShulerC. K. (2019). Submarine Groundwater Discharge and Stream Baseflow Sustain Pesticide and Nutrient Fluxes in Faga’alu Bay, American Samoa. Front. Environ. Sci.7, 162. 10.3389/fenvs.2019.00162
89
YamaharaK. M.LaytonB. A.SantoroA. E.BoehmA. B. (2007). Beach Sands along the California Coast Are Diffuse Sources of Fecal Bacteria to Coastal Waters. Environ. Sci. Technol.41, 4515–4521. 10.1021/es062822n
90
YangL.ChenC.-T. A.HongH.ChangY.-C.LuiH.-K. (2015). Mixing Behavior and Bioavailability of Dissolved Organic Matter in Two Contrasting Subterranean Estuaries as Revealed by Fluorescence Spectroscopy and Parallel Factor Analysis. Estuarine, Coastal Shelf Sci.166, 161–169. 10.1016/j.ecss.2014.10.018
91
YeQ.LiuJ.DuJ.ZhangJ. (2016). Bacterial Diversity in Submarine Groundwater along the Coasts of the Yellow Sea. Front. Microbiol.6, 1519. 10.3389/fmicb.2015.01519
Summary
Keywords
subterranean estuary, coastal aquifers, microbiome, sandy beach, groundwater, biogeochemistry
Citation
Archana A, Francis CA and Boehm AB (2021) The Beach Aquifer Microbiome: Research Gaps and Data Needs. Front. Environ. Sci. 9:653568. doi: 10.3389/fenvs.2021.653568
Received
14 January 2021
Accepted
08 April 2021
Published
26 April 2021
Volume
9 - 2021
Edited by
Carlos Rocha, Trinity College Dublin, Ireland
Reviewed by
Gwénaëlle Chaillou, Université du Québecà Rimouski, Canada
Wenzhi Liu, Chinese Academy of Sciences, China
Updates
Copyright
© 2021 Archana, Francis and Boehm.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Anand Archana, archanaa@stanford.edu
This article was submitted to Biogeochemical Dynamics, a section of the journal Frontiers in Environmental Science
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.