- 1Department of Biology, University of Fribourg, Fribourg, Switzerland
- 2Landscape Ecology, Institute of Terrestrial Ecosystems, ETH Zürich, Zürich, Switzerland
- 3Swiss Federal Institute for Forest, Snow and Landscape Research, Birmensdorf, Switzerland
The geography of speciation is one of the most contentious topics at the frontier between ecology and evolution. Here, building on previous hypotheses, I propose that ecological constraints on species co-existence mediate the likelihood of speciation, via a trade-off between competitive and dispersal abilities. Habitat stability, as found in the tropics, selects for the evolution of stronger competitive abilities. Since resource investment in competitive and dispersal abilities should trade off, high competition in stable habitats reduces species dispersal ability, decreasing effective population sizes. In smaller local populations, higher fixation rates of molecular substitutions increases the likelihood of speciation. Species diversity triggers more speciation by further increasing the spatial structuring of populations and decreasing effective population sizes. Resource specialization also trades-off with dispersal ability and could account for speciation at higher trophic levels. Biotic interactions may promote parapatric speciation and generate spatial patterns in diversity such as the latitudinal diversity gradient. I discuss evidence for this mechanism and emphasize the need for studies coupling ecology and speciation theory within landscapes across latitude.
Introduction
The extraordinary diversity of species on earth provides infinite motivation to unravel the underlying ecological and evolutionary rules. Evidences that biodiversity is linked to spatial features suggest that speciation mechanisms cannot be understood outside of their geographical context (Levin, 1993). The most famous example of spatial gradient in species diversity is the latitudinal biodiversity gradient that already puzzled Darwin (1859) and Wallace (1878). The current global distribution of biodiversity follows a strongly negative gradient from the tropics to the poles across multiple taxonomic groups (Mittelbach et al., 2007). The association between species spatial distribution and diversification is also reflected in the fossil record, where geographic range size is negatively related to speciation (Jablonski and Roy, 2003) and extinction rates (Jablonski, 2007). Classic examples of exceptional events of speciation, including Darwin's finches (Grant and Grant, 2011), Anolis lizards (Losos, 1998) or African cichlids (Schliewen et al., 1994; Barluenga et al., 2006), took place in confined spatial areas such as islands or lakes. Together, this suggests that the rate of species diversification is strongly linked to the spatial distribution and connectivity of meta-populations. However, the role played by this aspect of the speciation process in shaping diversity patterns has generally been neglected (Kisel and Barraclough, 2010).
Gavrilets (2014) highlighted the need to integrate models of community ecology with those developed in speciation theory in order to better understand the processes of speciation. Community ecology and speciation biology share the common interest of understanding what generates clines in species diversity and many theoretical similarities exists between the fields (e.g., the neutral biodiversity theory, Hubbell, 2001). In particular, the link between the latitudinal diversity gradient, the diversity of biotic interactions and the likelihood of speciation is a long-standing idea (Dobzhansky, 1950; MacArthur, 1969). Dobzhansky proposed that predictable tropical climate should select for greater specialization and species diversity. Since differences in either the intensity of speciation or extinction could ultimately account for the latitudinal cline in biodiversity (Weir and Schluter, 2007), a higher frequency of divergent selection along ecological axes in the tropics (Mittelbach et al., 2007), was proposed to trigger frequent events of ecological speciation (Schluter, 2009). Nevertheless, for speciation to occur, a geographically defined group of individuals must diverge from others and accumulate genetic and/or phenotypic differences that distinguish it as a new species (Levin, 1993).
In contrast to the view that speciation is the direct outcome of competition for resources, a trade-off between dispersal and competition/specialization may enhance speciation by affecting effective population size. Fedorov (1966) remarked that tropical forests are shaped by a multitude of contiguous populations of closely related species and postulated that the role of genetic drift prevails over that of natural selection in shaping new species. Here, building on Fedorov's observations, I propose the hypothesis of a possible direct link between biotic interactions, connectivity among populations, and speciation that may ultimately explain the higher diversity in the tropics (Mittelbach et al., 2007). I argue that under stable conditions, biotic interactions promote speciation via a trade-off between investments in competitive efficiency and dispersal abilities. While the metabolic theory of molecular evolution, which postulates a link between temperature-dependant metabolic rate and speciation, has received recent attention (Allen et al., 2006; Gillooly and Allen, 2007; Stegen et al., 2009; Dowle et al., 2013), the role of neutral parapatric speciation due to lower dispersal in the tropics has remained little explored. Five decades after the publication of Federov's observations and based on new molecular evidences (e.g., Lasso et al., 2011), I hypothesize that the ecological trade-off between resource use efficiency and dispersal abilities promotes higher speciation rates in the tropics.
Stability Selects for High Competitive and Low Dispersal Abilities
Brown and Gibson (1983) rightly stated that “ultimately, all general patterns of diversity must be attributed to physical causes, either historical perturbations or contemporary variation in the physical environment.” In particular, the shaping of species lineages should be linked to the long-term physical history of habitats (Pellissier et al., 2014). Greater stability over geological time periods has long been proposed to promote biodiversity, mostly because the risk of extinction diminishes (Darwin, 1859; Wallace, 1878). Reconstructions of ancient climate indeed indicate that the tropics remained climatically more stable than higher latitudes, and thus should have retained the most ancient lineages (Dynesius and Jansson, 2000). Accordingly, most extant clades have tropical origins, where basal clades in phylogenies are distributed at lower latitudes (Wiens and Donoghue, 2004). While stability obviously prevents species extinction, evidences indicate that speciation is also faster in the tropics (Weir and Schluter, 2007), but the mechanism is poorly described. A self-sustaining mechanism should provide an explanation for the origin of species diversity in stable habitats.
In ecology, stable habitats are traditionally associated with species possessing a particular life history (MacArthur and Wilson, 1967; Grime, 1977). In Grime's C-S-R triangle theory, stable habitats support C species, which optimize competitive abilities (Grime, 1977). Vegetation climax after ecological successions provide the best examples for the link between increasing stability and competition (Horn, 1974). As vegetation becomes well established in response to persistent climatic conditions, species compete for space, light and nutrients and only those that are most efficient can persist (Horn, 1974; Tilman, 1985). Stable habitats have also been associated with K species along the r-K gradient of MacArthur and Wilson (1967). In a stable environment, species densities are close to the carrying capacity (K) and species invest more in fewer offspring, increasing the probability of survival to a long adulthood (MacArthur and Wilson, 1967). Ecological successions have counterparts at an evolutionary scale. Examples of “evolutionary successions” have been described in island systems (Böhle et al., 1996; Panero et al., 1999). Darwin (1859) noted that most tree species make poor long-distance dispersers due to the large size of their seeds and, as a result, new islands are more often colonized by herbaceous species rather than trees. Under competition for resources, herbaceous clades colonizing islands rapidly evolve more competitive traits such as woodiness and tree-like morphologies (Böhle et al., 1996; Panero et al., 1999). Stable environments are associated with species evolutionary trajectory toward higher level of competitive abilities.
Competitive abilities generally trade-off with dispersal abilities, referring to either dispersal distance or dispersal frequency (McPeek and Holt, 1992; Ehrlén and Groenendael, 1998; Turnbull et al., 1999; Cadotte et al., 2006). In general, plant species that are better competitors do not exhibit adaptations for long distance dispersal (Ehrlén and Groenendael, 1998). Accordingly, in habitats with stronger competition for light, species with poor dispersal capacities are more prevalent (Ozinga et al., 2004). The evolution of competitive traits over dispersal abilities under varying degrees of stability can be illustrated by simple metapopulation models adapted from Levins and Culver (1971). Given competition between two species where species 1 is a superior competitor to species 2 defined by the following equations,
where p is the proportion of occupied sites, d the dispersal ability to reach unoccupied sites, and e an external factor of extinction, which is inversely proportional to environmental stability. The globally stable equilibrium point is given by:
The relative proportion of sites occupied by each species depends on the degree of perturbation relative to the species dispersal ability. In conditions of intermediate stability, both species survive if d2 > d1. Ecologically, intermediate level of perturbation maintains the highest level of diversity (Roxburgh et al., 2004). In contrast, stability is expected to promote the dominance of the most competitive species, so that without perturbation (e = 0), only the most competitive species p1survives. Therefore, in stable conditions, only the best competitors are expected to co-exist. While a strategy based on dispersal is disadvantaged, stability selects for the evolution of competitive abilities (Calcagno et al., 2006).
Lower dispersal abilities impact the gene flow among individuals in a landscape. Wright (1946) pointed out the link between limited dispersal and reduced effective population size (Ne): “It is shown that in the absence of disturbing factors, short range dispersal […] leads to considerable differentiation not only among small subdivisions but also of large ones.” Limited dispersal leads to a less connected network of gene flow between populations and thus causes a decrease in effective population size. According to Wright's model with limited dispersal, isolation-by-distance reduces effective population size, even for species that are spread across a wide landscape. The resulting neighborhood effective population size is lower than the census size and related to species dispersal abilities by the following equation:
where D is the density of pairs contributing to the reproduction and σd the standard deviation of dispersal distance representing the dispersal kernel (Wright, 1946). This model linking effective population size and dispersal is in line with the positive correlation between the observed range sizes of species and their dispersal abilities (Gaston, 1998). Together, ecological and evolutionary demographic theories suggest a direct connection between stability, competition, lower dispersal, and decreased effective population size (Figure 1).
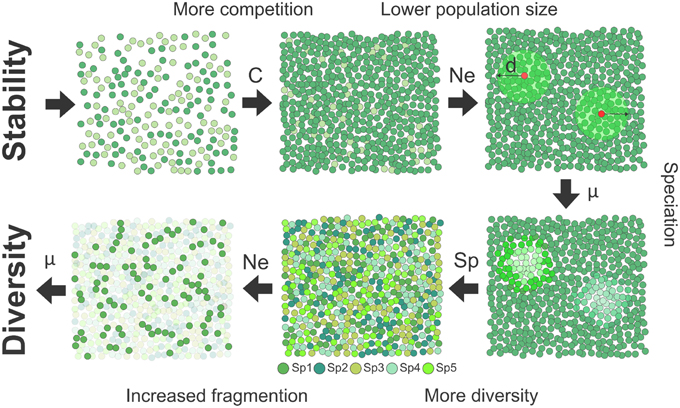
Figure 1. Schematic representation of the mechanism of speciation in a stable habitat. Habitat stability as found in the tropics selects for the evolution of stronger competitive abilities (C). Since resource investment in competition and dispersal (d) should trade off, inferior dispersal in competitive species implies a decrease in effective population size (Ne). In smaller populations, higher fixation rate of molecular substitutions (μ) increases the likelihood of speciation events. Higher diversity further promotes diversity since the presence of more species increases spatial structuring of populations, favoring genetic differentiation, and speciation.
Population Size, Nearly-neutral Substitutions and Speciation
A decrease in population size causes an increase in the rate of non-neutral substitutions (Ohta, 1992; Lanfear et al., 2014), and also in the rate of neutral substitution under specific geometric (e.g., non-circular meta-population distribution, Allen et al., 2015), or demographic conditions (e.g., overlapping generations, Balloux and Lehmann, 2012). As predicted by the nearly neutral theory of molecular evolution (Ohta, 1992), a decrease in effective population size (Ne) leads to easier fixation of nearly-neutral mutations (Ohta, 1992; Lanfear et al., 2014). The probability μf that the allele will be fixed in the population was first derived by Kimura (1957) and depends upon the effective Ne and census population sizes N and the strength of selection s as:
In the case of small effective population size, the coefficient of selection s is balanced by drift. As the effective population size decreases, the influence of drift increases, resulting in a higher ratio of non-synonymous to synonymous substitutions (dN/dS) (Kryazhimskiy and Plotkin, 2008). In particular, the probability of fixation of slightly deleterious mutations increases rapidly and approaches the neutral value (Ohta, 1992). Tachida and Iizuka (1991) further suggested that smaller population size might also increase the probability of fixation of slightly beneficial mutations, in comparison with the case of extensive dispersal within a large population. Selection can thus be stronger in small than in large populations (Ohta, 1992), but this depends on how the rate of migration scales with population size (Gavrilets and Gibson, 2002). As theoretically expected, accelerated rates of non-synonymous molecular substitution are typically found in geographically restricted populations, such as on islands (Johnson and Seger, 2001; Woolfit and Bromham, 2005).
In stable habitats, species should invest in competitive rather than dispersal traits, and reduced effective population size should increase intra-specific genetic structure (Wright, 1946; Ohta, 1992). The link between dispersal and genetic differentiation among lineages found in coral reef fishes (Riginos et al., 2014) or plants (Hardy et al., 2006; Duminil et al., 2007; Theim et al., 2014) suggests a direct link between dispersal, effective population size, and genetic structure. Furthermore, assuming a trade-off between competitive and dispersal abilities, intra-specific non-synonymous substitutions should correlate with traits representing syndromes of a K strategy. Accordingly, plant height showed a positive correlation with the ratio of non-synonymous to synonymous substitutions (dN/dS) across 138 families of flowering plants (Lanfear et al., 2013; Bromham et al., 2015), while longevity was a strong predictor of this ratio in animals (Romiguier et al., 2014). Similarly, large mammals and birds have a higher rate of amino acid substitutions in proteins (Popadin et al., 2007; Weber et al., 2014). In mammals, the subdivision of a species into sub-populations, such as in the case of competition for territories, promotes both high rates of speciation and chromosomal evolution consistent with an effect of small population size (Bush et al., 1977). Thus, as expected from a link between competitive abilities and effective population size, species displaying traits related to competition have a greater rate of non-neutral substitutions triggering protein evolution (Popadin et al., 2007; Lanfear et al., 2014; Romiguier et al., 2014; Weber et al., 2014).
Organisms prioritizing competitive over dispersal abilities can become geographically isolated more easily, which should enhance speciation (Mayr, 1963). For marine fishes the association between genetic structure, dispersal and species richness suggests that reduction in gene flow can promote speciation (Riginos et al., 2014). Kisel and Barraclough (2010) found that both dispersal and gene flow in terrestrial taxa were good predictors of speciation rates. Furthermore, small ranged species are over-represented in global biodiversity, which may indicate that speciation via dispersal limitation and small population size is an important mechanism in nature (Gaston, 1998). Under limited dispersal, speciation may arise from gradual separation of sub-populations as molecular substitutions become fixed locally (Figure 1). Prezygotic or postzygotic isolating barriers may further counter gene flow to avoid maladapted hybrids (Mayr, 1963; Ramsey et al., 2003; Lukhtanov et al., 2005; McBride and Singer, 2010). Greater isolation by distance (Martin and McKay, 2004), genetic divergence (Eo et al., 2008), but limited occurrence of hybrids (Hopkins, 2013; Surget-Groba and Kay, 2013) has been documented in the tropics and may represent on-going speciation events within local sub-populations.
Non-synonymous substitutions arising in small populations could promote evolution of novel ecological preferences. Since the rate of fixation of non-neutral mutations might be higher (Tachida and Iizuka, 1991; Ohta, 1992) and proteins evolve faster in small populations (Ohta, 2002), this may increase the overall rate of morphological evolution. For instance, in the fossil record small ranged and transient trilobite fossil species show increased morphological variation (Hopkins, 2011), while large ranged species are more likely to show morphological stasis (Gould and Eldredge, 1977). As formulated in the theory of punctuated equilibria (Gould and Eldredge, 1977; Eldredge et al., 2005), ecological innovations might arise at higher frequency in smaller populations. This would also explain the faster rate of morphological evolution in tropical islands (Millien, 2006) and lakes (Schliewen et al., 1994), where population size is smaller (Woolfit and Bromham, 2005) compared to larger continental or ocean surface. Stable habitats reunite the theoretical conditions expected to fuel speciation, including lower effective population size, local mating linked to dispersal limitation and high levels of local genetic variation (Gavrilets, 2004). Together this suggests that molecular evolution fuelling speciation is faster under stable conditions, high competition and limited dispersal, which is characteristic of the tropics. Population differentiation might be further fuelled by higher metabolic and mutation rates expected at lower latitudes (Wright et al., 2006; Stegen et al., 2009).
The current argument suggests a link between rate of molecular substitution and rate of speciation. Evidences of a link between the rates of molecular evolution and diversification have been reported (Eo and DeWoody, 2010; Lanfear et al., 2010; see Dowle et al., 2013). For instance, Lanfear et al. (2010) identified a correlation between clade rates of molecular evolution and net-diversification in birds. However, the demonstration that the correlation between molecular evolution and speciation hinges on population sizes would require a sampling at the scale of population subdivision. Moreover, as raised by Dowle et al. (2013), there is currently a lack of evidence of differential population size along latitudinal gradient. The absence of evidence primarily result the lack of studies investigating the genetic structure of population with a comparable sampling design across latitudes. Nevertheless, many studies highlight unexpected high genetic structure in low latitude species (Martin and McKay, 2004; Born et al., 2008; Eo et al., 2008; Lasso et al., 2011), which provide clues of a lower gene flow and smaller effective population sizes due to limited dispersal. Fedorov's ideas relied on the hypothesis of higher rate self-pollination in tropical trees, which was later contradicted (Bawa, 1974). Gene flow through pollen dispersal in tropical trees could potentially occur across long distances (White et al., 2002), but whether pollen transfer across large distance is common or not still remains to be documented across many species. Further studies are therefore required to quantify gene flow among populations across latitudes.
At the other extreme, in less stable environments larger range size resulting from higher dispersal may buffer species against extinction, as suggested by the positive relationship between range size and duration in the fossil record (Jablonski, 2008). This could explain the larger range size at higher latitudes (Rapoport's rule, Stevens, 1989), which may reduce the risk of extinction under less stable environmental conditions. Species with larger range size have greater dispersal ability (Gaston, 1998) and shorter generation time, like r species along the r-K gradient (MacArthur and Wilson, 1967). Species with shorter generation time accumulate synonymous substitutions faster and generally show a higher level of neutral polymorphism than species with greater longevity or offspring quality (Lanfear et al., 2014; Romiguier et al., 2014). Species with a wide distribution and high dispersal are stabilized in their genetic variation by their large population size and the process of gene flow (Gould and Eldredge, 1977), but which also limits morphological or ecological evolution. Yet, at higher latitude, in less stable environments, speciation may also happen neutrally related to species range dynamics. For instance, range dynamic in interaction with habitat heterogeneity can result in range fragmentation and high rate of speciation (Arenas et al., 2012, 2013; Mona et al., 2014). Populations in expansion are expected to fix mutations though genetic drift occurring in populations located on the edge of the expansion, which may promote speciation (Excoffier and Ray, 2008). However, pronounced range dynamic at higher latitude should also have increased extinction rates (Dynesius and Jansson, 2000). Better sampled phylogenies at the scale of ongoing population divisions (i.e., infra-species) within landscapes is necessary to compare diversification processes across latitudes.
Diversity Increases Landscape Fragmentation
Species diversity itself could be a driver of species diversification following the famous “diversity begets diversity” model, since speciation rates correlate with diversity (Emerson and Kolm, 2005). I propose that, when more than one species co-exist in a landscape, and assuming the same potential density of pairs in all species, the effective population size of a species i follow the formula:
where D is the density of pairs contributing to the reproduction σdi the standard deviation of dispersal distance of the species i and Sp the number of species. An increase in species diversity causes a decrease in species relative density, thus reducing effective population size. Increasing the number of species thus decreases the local population size, enhancing genetic drift, and the likelihood of divergence (Wright, 1946; Ohta, 1992).
A higher rate of neutral molecular evolution has been found in tropical clades (Wright et al., 2006; Gillman et al., 2009), and in clades with greater numbers of species (Duchene and Bromham, 2013), but small population size was dismissed as an explanation based on the assumption that neutral substitution is not influenced by population size (Charlesworth, 2009). However, the neutral substitution rate is higher for smaller populations in the presence of overlapping generations, as is largely the case in tropical species (Charlesworth, 2009; Balloux and Lehmann, 2012). In addition, limited dispersal in highly diverse landscape may shape more patchy distribution of populations with non-regular spatial structures (Figure 1). Allen et al. (2015) demonstrated that singular geometric spatial structures of individuals or populations that form in high diversity landscapes (Figure 1) can increase the rate of synonymous substitution. The higher rate of molecular evolution in tropical clades supports the central role of effective population size in tropical speciation (Wright et al., 2006; Gillman et al., 2009, 2012). Therefore, a higher rate of both neutral and non-neutral substitution could fuel speciation in stable habitats in interaction with population sizes.
Ecological Island Syndrome at Higher Trophic Levels
Specialisation of antagonistic and mutualistic interactions is another typical response of higher trophic levels to resource competition in stable habitats (Futuyma and Moreno, 1988). Specialisation allows an increase in the efficiency of the use of a given resource, to the detriment of a wider trophic regime (Futuyma and Moreno, 1988). Trophic specialization is central to models of adaptive radiation (Schluter, 2000), and may underlie much of the shaping of species diversity (Jocque et al., 2010; Forister et al., 2012). Like traits related to competitive abilities, trophic specialization is expected to be negatively related to dispersal, as the probability of finding suitable conditions elsewhere declines with specialization (Salisbury et al., 2012). Following Janzen (1968), who described plants as islands in space for the herbivorous insects that feed on them, a specialist will only be distributed in the area overlapping host or prey species, thus increasing the fragmentation of its populations (Figure 2). Hence, specialization results in a limited population size, which should increase the probability of speciation (Wright, 1946; Ohta, 1992).
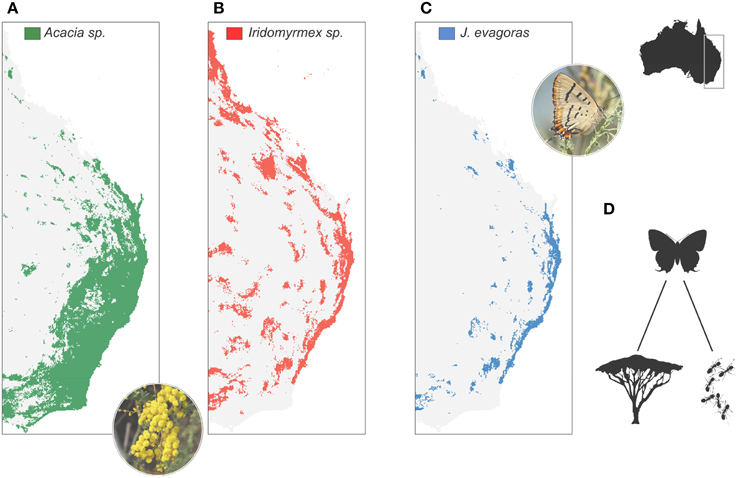
Figure 2. Example of biotic interactions impacting the spatial configuration of populations. The butterfly Jalmenus evagoras of the Lycaenidae feeds on ~16 Acacia plant species (A) (Pierce and Nash, 1999), and is in an obligatory mutualism with principally two Iridomyrmex ant species (Iridomyrmex anceps, I. rufoniger) (B). As a consequence, the distribution of J. evagoras (C) is not only constrained by the host plant, but also by the mutualist ant, causing population fragmentation and likely genetic differentiation (Eastwood et al., 2006). (D) Schematic view of the trophic link between the butterfly J. evagoras and its host plants and ant species. Distribution maps were obtained using species distribution models with a random forest statistical approach applied to occurrences from http://www.gbif.org/. Pseudo-absences were generated randomly across Australia. The distribution of J. evagoras was constrained by the forecasted range of the host plants and mutualistic ant species. Acacia spectabilis image by Melburnian [Own work (digital photograph by author)] [CC BY 3.0 (http://creativecommons.org/licenses/by/3.0)], via Wikimedia Commons. Jalmenus-evagoras-ventral image by Benjamint444 (Own work) [GFDL 1.2 (http://www.gnu.org/licenses/old-licenses/fdl-1.2.html)], via Wikimedia Commons.
Fragmentation of populations following the appearance of strong biotic interactions may trigger an increased rate of molecular substitution and speciation (Gavrilets et al., 2000). In both marine and terrestrial ecosystems, biotic specialization is associated with marked intra-specific spatial genetic structure. Fishes in close mutualism with corals, sea urchins, or anemones show exceptional spatial genetic structure (Hoffman et al., 2005) and a higher rate of diversification (Litsios et al., 2012). Zayed and Packer (2007) found that a species of bee with a specialized pollen diet had considerably higher spatial genetic variation among populations compared to a generalist counterpart. Strong spatial genetic differentiation has also been found among populations of mutualistic butterflies of the family Lycaenidae (Eastwood et al., 2006; Pellissier et al., 2012), which has led to even stronger differentiation in one of their specialized parasitoid wasps (Anton et al., 2007). Higher degrees of specialization or multiple biotic constraints (e.g., the required presence of a mutualist in addition to the trophic host, Figure 2) should reinforce the degree of fragmentation. The idea that biotic constraints play a major role in the process of tropical diversification is not novel (Dobzhansky, 1950), and has led to the hypothesis that the latitudinal diversity gradient is mainly due to latitudinal differences in biotic interactions (Wallace, 1878; Dobzhansky, 1950). Jocque et al. (2010) also argued that the trade-off between specialization and dispersal underlies the latitudinal diversity gradient, since specialized biotic interactions are more common in the tropics (Schemske et al., 2009). Accordingly, I propose that speciation associated with biotic interaction may not be necessarily due to filling novel niches, but results from spatial fragmentation of populations due to limited dispersal which promotes local divergence.
Conclusion
Although it is becoming increasingly clear that many tropical clades experience higher speciation rates, very little is known about the processes of divergence among populations (Surget-Groba and Kay, 2013). Increased evidences of unexpected high degree of genetic differentiation among populations of tropical species argue for pursuing Fedorov's idea (Lasso et al., 2011). Here, I discussed how biotic constraints may modulate population size, the rate of molecular evolution and speciation in stable habitats like those found in the tropics. I propose that under stable environmental conditions, biotic constraints promote speciation through a trade-off between competition and dispersal. In turn, this can be extended to specialization-dispersal trade-offs at higher trophic levels (Jocque et al., 2010). The current theory can be broken down into a series of elements which can be validated independently on empirical data and thus represent a testable framework (Table 1). The strength of the present concept is its ability to bridge theories in ecology (C-S-R, r-K theories, Rapoport's rule), paleontology (punctuated equilibria) and evolution (the nearly neutral theory of molecular evolution). Only by cutting across disciplines can we hope to unravel the mechanisms driving the origin of species diversity on earth. The continued reductions in DNA sequencing costs which allow sampling many populations across tropical and temperate landscape, and estimating both Ne and substitution rates, promise many advances in the years to come.
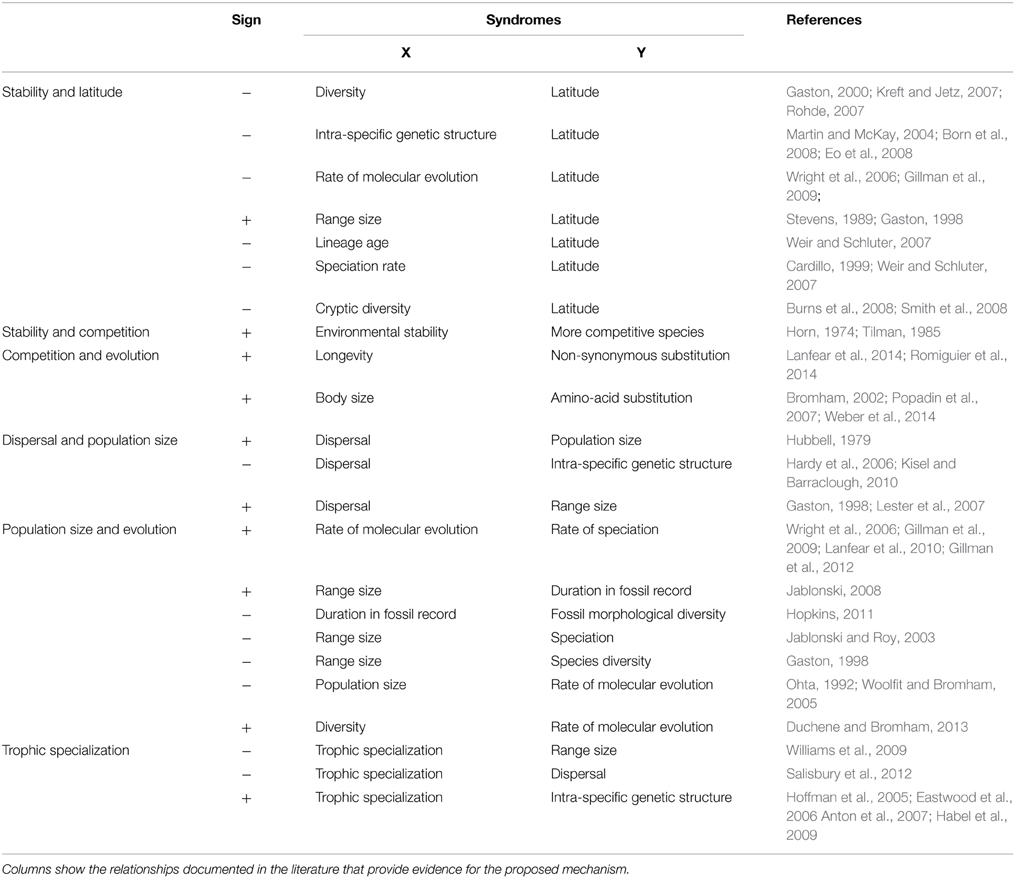
Table 1. List of syndromes that provide indication of the links between competitive abilities, dispersal abilities, population sizes, and species diversification along a latitudinal gradient.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Jens-Christian Svenning, Mary S. Wisz, Doyle McKay, Nadir Alvarez, Rudolf Rohr, Russell Naisbit, Nadine Sandau as well as the reviewers are thanked for their comments on a previous version of the manuscript.
References
Allen, A. P., Gillooly, J. F., Savage, V. M., and Brown, J. H. (2006). Kinetic effects of temperature on rates of genetic divergence and speciation. Proc. Natl. Acad. Sci. U.S.A. 103, 9130–9135. doi: 10.1073/pnas.0603587103
Allen, B., Sample, C., Dementieva, Y., Medeiros, R. C., Paoletti, C., and Nowak, M. A. (2015). The molecular clock of neutral evolution can be accelerated or slowed by asymmetric spatial structure. PLoS Comput. Biol. 11:e1004108. doi: 10.1371/journal.pcbi.1004108
Anton, C., Zeisset, I., Musche, M., Durka, W., Boomsma, J. J., and Settele, J. (2007). Population structure of a large blue butterfly and its specialist parasitoid in a fragmented landscape. Mol. Ecol. 16, 3828–3838. doi: 10.1111/j.1365-294X.2007.03441.x
Arenas, M., François, O., Currat, M., Ray, N., and Excoffier, L. (2013). Influence of admixture and Paleolithic range contractions on current European diversity gradients. Mol. Biol. Evol. 30, 57–61. doi: 10.1093/molbev/mss203
Arenas, M., Ray, N., Currat, M., and Excoffier, L. (2012). Consequences of range contractions and range shifts on molecular diversity. Mol. Biol. Evol. 29, 207–218. doi: 10.1093/molbev/msr187
Balloux, F., and Lehmann, L. (2012). Substitution rates at neutral genes depend on population size under fluctuating demography and overlapping generations. Evolution 66, 605–611. doi: 10.1111/j.1558-5646.2011.01458.x
Barluenga, M., Stölting, K. N., Salzburger, W., Muschick, M., and Meyer, A. (2006). Sympatric speciation in Nicaraguan crater lake cichlid fish. Nature 439, 719–723. doi: 10.1038/nature04325
Bawa, K. S. (1974). Breeding systems of tree species of a lowland tropical community. Evolution 28, 85–92. doi: 10.2307/2407241
Böhle, U. R., Hilger, H. H., and Martin, W. F. (1996). Island colonization and evolution of the insular woody habit in Echium L. (Boraginaceae). Proc. Natl. Acad. Sci. U.S.A. 93, 11740–11745. doi: 10.1073/pnas.93.21.11740
Born, C., Hardy, O. J., Chevallier, M. H., Ossari, S., Atteke, C., Wickings, E., et al. (2008). Small−scale spatial genetic structure in the Central African rainforest tree species Aucoumea klaineana: a stepwise approach to infer the impact of limited gene dispersal, population history and habitat fragmentation. Mol. Ecol. 17, 2041–2050. doi: 10.1111/j.1365-294X.2007.03685.x
Bromham, L. (2002). Molecular clocks in reptiles: life history influences rate of molecular evolution. Mol. Biol. Evol. 19, 302–309. doi: 10.1093/oxfordjournals.molbev.a004083
Bromham, L., Hua, X., Lanfear, R., and Cowman, P. F. (2015). Exploring the relationships between mutation rates, life history, genome size, environment, and species richness in flowering plants. Am. Nat. 185, 507–524. doi: 10.1086/680052
Burns, J. M., Janzen, D. H., Hajibabaei, M., Hallwachs, W., and Hebert, P. D. N. (2008). DNA barcodes and cryptic species of skipper butterflies in the genus Perichares in Area de Conservacion Guanacaste, Costa Rica. Proc. Natl. Acad. Sci. U.S.A. 105, 6350–6355. doi: 10.1073/pnas.0712181105
Bush, G. L., Case, S. M., Wilson, A. C., and Patton, J. L. (1977). Rapid speciation and chromosomal evolution in mammals. Proc. Natl. Acad. Sci. U.S.A. 74, 3942–3946. doi: 10.1073/pnas.74.9.3942
Cadotte, M. W., Mai, D. V., Jantz, S., Collins, M. D., Keele, M., and Drake, J. A. (2006). On testing the competition-colonization trade-off in a multispecies assemblage. Am. Nat. 168, 704–709. doi: 10.1086/508296
Calcagno, V., Mouquet, N., Jarne, P., and David, P. (2006). Coexistence in a metacommunity: the competition–colonization trade−off is not dead. Ecol. Lett. 9, 897–907. doi: 10.1111/j.1461-0248.2006.00930.x
Cardillo, M. (1999). Latitude and rates of diversification in birds and butterflies. Proc. R. Soc. B Biol. Sci. 266, 1221. doi: 10.1098/rspb.1999.0766
Charlesworth, B. (2009). Fundamental concepts in genetics: effective population size and patterns of molecular evolution and variation. Nat. Rev. Genet. 10, 195–205. doi: 10.1038/nrg2526
Dowle, E. J., Morgan-Richards, M., and Trewick, S. A. (2013). Molecular evolution and the latitudinal biodiversity gradient. Heredity 110, 501–510. doi: 10.1038/hdy.2013.4
Duchene, D., and Bromham, L. (2013). Rates of molecular evolution and diversification in plants: chloroplast substitution rates correlated with species-richness in the Proteaceae. BMC Evol. Biol. 13:65. doi: 10.1186/1471-2148-13-65
Duminil, J., Fineschi, S., Hampe, A., Jordano, P., Salvini, D., Vendramin, G. G., et al. (2007). Can population genetic structure be predicted from life−history traits? Am. Nat. 169, 662–672. doi: 10.1086/513490
Dynesius, M., and Jansson, R. (2000). Evolutionary consequences of changes in species' geographical distributions driven by Milankovitch climate oscillations. Proc. Natl. Acad. Sci. U.S.A. 97, 9115–9120. doi: 10.1073/pnas.97.16.9115
Eastwood, R., Pierce, N. E., Kitching, R. L., and Hughes, J. M. (2006). Do ants enhance diversification in lycaenid butterflies? Phylogeographic evidence from a model myrmecophile, Jalmenus evagoras. Evolution 60, 315–327. doi: 10.1111/j.0014-3820.2006.tb01109.x
Ehrlén, J., and Groenendael, J. M. (1998). The trade-off between dispersability and longevity-an important aspect of plant species diversity. Appl. Veg. Sci. 1, 29–36. doi: 10.2307/1479083
Eldredge, N., Thompson, J. N., Brakefield, P. M., Gavrilets, S., Jablonski, D., Jackson, J. B. C., et al. (2005). The dynamics of evolutionary stasis. Paleobiology 31, 133–145. doi: 10.1666/0094-8373(2005)031[0133:TDOES]2.0.CO;2
Emerson, B. C., and Kolm, N. (2005). Species diversity can drive speciation. Nature 434, 1015–1017. doi: 10.1038/nature03450
Eo, S. H., and DeWoody, J. A. (2010). Evolutionary rates of mitochondrial genomes correspond to diversification rates and to contemporary species richness in birds and reptiles. Proc. Biol. Sci. 277, 3587–3592. doi: 10.1098/rspb.2010.0965
Eo, S. H., Wares, J. P., and Carroll, J. P. (2008). Population divergence in plant species reflects latitudinal biodiversity gradients. Biol. Lett. 4, 382–384. doi: 10.1098/rsbl.2008.0109
Excoffier, L., and Ray, N. (2008). Surfing during population expansions promotes genetic revolutions and structuration. Trends Ecol. Evol. 23, 347–351. doi: 10.1016/j.tree.2008.04.004
Fedorov, A. A. (1966). The structure of the tropical rain forest and speciation in the humid tropics. J. Ecol. 1, 1–11. doi: 10.2307/2257656
Forister, M. L., Dyer, L. A., Singer, M. S., Stireman, J. O., and Lill, J. T. (2012). Revisiting the evolution of ecological specialization, with emphasis on insect-plant interactions. Ecology 93, 981–991. doi: 10.1890/11-0650.1
Futuyma, D. J., and Moreno, G. (1988). The evolution of ecological specialization. Annu. Rev. Ecol. Syst. 19, 207–233. doi: 10.1146/annurev.es.19.110188.001231
Gaston, K. J. (1998). Species-range size distributions: products of speciation, extinction and transformation. Philos. Trans. R. Soc. B Biol. Sci. 353, 219–230. doi: 10.1098/rstb.1998.0204
Gavrilets, S. (2004). Fitness Landscapes and the Origin of Species. Princeton, NJ: Princeton university press.
Gavrilets, S. (2014). Models of speciation: where are we now? J. Hered. 105, 743–755. doi: 10.1093/jhered/esu045
Gavrilets, S., and Gibson, N. (2002). Fixation probabilities in a spatially heterogeneous environment. Popul. Ecol. 44, 51–58. doi: 10.1007/s101440200007
Gavrilets, S., Li, H., and Vose, M. D. (2000). Patterns of parapatric speciation. Evolution 54, 1126–1134. doi: 10.1111/j.0014-3820.2000.tb00548.x
Gillman, L. N., Keeling, D. J., Ross, H. A., and Wright, S. D. (2009). Latitude, elevation and the tempo of molecular evolution in mammals. Proc. Biol. Sci. 276, 3353–3359. doi: 10.1098/rspb.2009.0674
Gillman, L. N., McCowan, L. S. C., and Wright, S. D. (2012). The tempo of genetic evolution in birds: body mass and climate effects. J. Biogeogr. 39, 1567–1572. doi: 10.1111/j.1365-2699.2012.02730.x
Gillooly, J. F., and Allen, A. P. (2007). Linking global patterns in biodiversity to evolutionary dynamics using metabolic theory. Ecology 88, 1890–1894. doi: 10.1890/06-1935.1
Gould, S. J., and Eldredge, N. (1977). Punctuated equilibria: the tempo and mode of evolution reconsidered. Paleobiology 3, 115–151.
Grant, P. R., and Grant, B. R. (2011). How and Why Species Multiply: the Radiation of Darwin's Finches. New Jersey: Princeton University Press.
Grime, J. P. (1977). Evidence for the existence of three primary strategies in plants and its relevance to ecological and evolutionary theory. Am. Nat. 111, 1169. doi: 10.1086/283244
Habel, J. C., Meyer, M., and Schmitt, T. (2009). The genetic consequence of differing ecological demands of a generalist and a specialist butterfly species. Biodivers. Conserv. 18, 1895–1908. doi: 10.1007/s10531-008-9563-5
Hardy, O. J., Maggia, L., Bandou, E., Breyne, P., Caron, H., Chevallier, M. H., et al. (2006). Fine-scale genetic structure and gene dispersal inferences in 10 Neotropical tree species. Mol. Ecol. 15, 559–571. doi: 10.1111/j.1365-294X.2005.02785.x
Hoffman, E. A., Kolm, N., Berglund, A., Arguello, J. R., and Jones, A. G. (2005). Genetic structure in the coral-reef-associated Banggai cardinalfish, Pterapogon kauderni. Mol. Ecol. 14, 1367–1375. doi: 10.1111/j.1365-294X.2005.02538.x
Hopkins, M. J. (2011). How species longevity, intraspecific morphological variation, and geographic range size are related: a comparison using Late Cambrian trilobites. Evolution 65, 3253–3273. doi: 10.1111/j.1558-5646.2011.01379.x
Horn, H. S. (1974). The ecology of secondary succession. Annu. Rev. Ecol. Syst. 5, 25–37. doi: 10.1146/annurev.es.05.110174.000325
Hubbell, S. P. (1979). Tree dispersion, abundance, and diversity in a tropical dry forest. Science 203, 1299–1309. doi: 10.1126/science.203.4387.1299
Hubbell, S. P. (2001). The Unified Neutral Theory of Biodiversity and Biogeography. New Jersey: Princeton University Press.
Jablonski, D. (2007). Scale and hierarchy in macroevolution. Palaeontology 50, 87–109. doi: 10.1111/j.1475-4983.2006.00615.x
Jablonski, D. (2008). Extinction and the spatial dynamics of biodiversity. Proc. Natl. Acad. Sci. U.S.A. 105, 11528–11535. doi: 10.1073/pnas.0801919105
Jablonski, D., and Roy, K. (2003). Geographical range and speciation in fossil and living molluscs. Proc. Biol. Sci. 270, 401–406. doi: 10.1098/rspb.2002.2243
Janzen, D. H. (1968). Host plants as islands in evolutionary and contemporary time. Am. Nat. 102, 592–595. doi: 10.1086/282574
Jocque, M., Field, R., Brendonck, L., and De Meester, L. (2010). Climatic control of dispersal-ecological specialization trade-offs: a metacommunity process at the heart of the latitudinal diversity gradient? Glob. Ecol. Biogeogr. 19, 244–252. doi: 10.1111/j.1466-8238.2009.00510.x
Johnson, K. P., and Seger, J. (2001). Elevated rates of nonsynonymous substitution in island birds. Mol. Biol. Evol. 18, 874–881. doi: 10.1093/oxfordjournals.molbev.a003869
Kimura, M. (1957). Some problems of stochastic processes in genetics. Ann. Math. Stat. 4, 882–901. doi: 10.1214/aoms/1177706791
Kisel, Y., and Barraclough, T. G. (2010). Speciation has a spatial scale that depends on levels of gene flow. Am. Nat. 175, 316–334. doi: 10.1086/650369
Kreft, H., and Jetz, W. (2007). Global patterns and determinants of vascular plant diversity. Proc. Natl. Acad. Sci. U.S.A. 104, 5925–5930. doi: 10.1073/pnas.0608361104
Kryazhimskiy, S., and Plotkin, J. B. (2008). The population genetics of dN/dS. PLoS Genet. 4:e1000304. doi: 10.1371/journal.pgen.1000304
Lanfear, R., Ho, S. Y., Love, D., and Bromham, L. (2010). Mutation rate is linked to diversification in birds. Proc. Natl. Acad. Sci. U.S.A. 107, 20423–20428. doi: 10.1073/pnas.1007888107
Lanfear, R., Ho, S. Y. W., Jonathan Davies, T., Moles, A. T., Aarssen, L., Swenson, N. G., et al. (2013). Taller plants have lower rates of molecular evolution. Nat. Commun. 4:1879. doi: 10.1038/ncomms2836
Lanfear, R., Kokko, H., and Eyre-Walker, A. (2014). Population size and the rate of evolution. Trends Ecol. Evol. 29, 33–41. doi: 10.1016/j.tree.2013.09.009
Lasso, E., Dalling, J. W., and Bermingham, E. (2011). Strong spatial genetic structure in five tropical Piper species: should the Baker–Fedorov hypothesis be revived for tropical shrubs? Ecol. Evol. 1, 502–516. doi: 10.1002/ece3.40
Lester, S. E., Ruttenberg, B. I., Gaines, S. D., and Kinlan, B. P. (2007). The relationship between dispersal ability and geographic range size. Ecol. Lett. 10, 745–758. doi: 10.1111/j.1461-0248.2007.01070.x
Levins, R., and Culver, D. (1971). Regional coexistence of species and competition between rare species. Proc. Natl. Acad. Sci. U.S.A. 68, 1246–1248. doi: 10.1073/pnas.68.6.1246
Litsios, G., Sims, C. A., Wüest, R. O., Pearman, P. B., Zimmermann, N. E., and Salamin, N. (2012). Mutualism with sea anemones triggered the adaptive radiation of clownfishes. BMC Evol. Biol. 12:212. doi: 10.1186/1471-2148-12-212
Losos, J. B. (1998). Contingency and determinism in replicated adaptive radiations of island lizards. Science 279, 2115–2118. doi: 10.1126/science.279.5359.2115
Lukhtanov, V. A., Kandul, N. P., Plotkin, J. B., Dantchenko, A. V., Haig, D., and Pierce, N. E. (2005). Reinforcement of pre-zygotic isolation and karyotype evolution in Agrodiaetus butterflies. Nature 436, 385–389. doi: 10.1038/nature03704
MacArthur, R. H. (1969). Patterns of communities in the tropics. Biol. J. Linnean Soc. 1, 19–30. doi: 10.1111/j.1095-8312.1969.tb01809.x
MacArthur, R. H., and Wilson, E. O. (1967). The Theory of Island Biogeography. Monographs in Population Biology. New Jersey: Princeton University Press.
Martin, P. R., and McKay, J. K. (2004). Latitudinal variation in genetic divergence of populations and the potential for future speciation. Evolution 58, 938–945. doi: 10.1111/j.0014-3820.2004.tb00428.x
McBride, C. S., and Singer, M. C. (2010). Field studies reveal strong postmating isolation between ecologically divergent butterfly populations. PLoS Biol. 8:e1000529. doi: 10.1371/journal.pbio.1000529
McPeek, M. A., and Holt, R. D. (1992). The Evolution of dispersal in spatially and temporally varying environments. Am. Nat. 140, 1010. doi: 10.1086/285453
Millien, V. (2006). Morphological evolution is accelerated among island mammals. PLoS Biol. 4:e321. doi: 10.1371/journal.pbio.0040321
Mittelbach, G. G., Schemske, D. W., Cornell, H. V., Allen, A. P., Brown, J. M., Bush, M. B., et al. (2007). Evolution and the latitudinal diversity gradient: speciation, extinction and biogeography. Ecol. Lett. 10, 315–331. doi: 10.1111/j.1461-0248.2007.01020.x
Mona, S., Ray, N., Arenas, M., and Excoffier, L. (2014). Genetic consequences of habitat fragmentation during a range expansion. Heredity 112, 291–299. doi: 10.1038/hdy.2013.105
Ohta, T. (1992). The nearly neutral theory of molecular evolution. Annu. Rev. Ecol. Evol. Syst. 23, 263–286. doi: 10.1146/annurev.es.23.110192.001403
Ohta, T. (2002). Near-neutrality in evolution of genes and gene regulation. Proc. Natl. Acad. Sci. U.S.A. 99, 16134–16137. doi: 10.1073/pnas.252626899
Ozinga, W. A., Bekker, R. M., Schaminée, J. H. J., and van Groenendael, J. M. (2004). Dispersal potential in plant communities depends on environmental conditions. J. Ecol. 92, 767–777. doi: 10.1111/j.0022-0477.2004.00916.x
Panero, J. L., Francisco-Ortega, J., Jansen, R. K., and Santos-Guerra, A. (1999). Molecular evidence for multiple origins of woodiness and a new world biogeographic connection of the Macaronesian island endemic Pericallis (Asteraceae: senecioneae). Proc. Natl. Acad. Sci. U.S.A. 96, 13886–13891. doi: 10.1073/pnas.96.24.13886
Pellissier, L., Leprieur, F., Parravicini, V., Cowman, P. F., Kulbicki, M., Litsios, G., et al. (2014). Quaternary coral reef refugia preserved fish diversity. Science 344, 1016–1019. doi: 10.1126/science.1249853
Pellissier, L., Litsios, G., Guisan, A., and Alvarez, N. (2012). Molecular substitution rate increases in myrmecophilous lycaenid butterflies (Lepidoptera). Zool. Scr. 41, 651–658. doi: 10.1111/j.1463-6409.2012.00556.x
Pierce, N. E., and Nash, D. R. (1999). The imperial blue, Jalmenus evagoras (Lycaenidae). Monogr. Aust. Lepidoptera 6, 279–315.
Popadin, K., Polishchuk, L. V., Mamirova, L., Knorre, D., and Gunbin, K. (2007). Accumulation of slightly deleterious mutations in mitochondrial protein-coding genes of large versus small mammals. Proc. Natl. Acad. Sci. U.S.A. 104, 13390–13395. doi: 10.1073/pnas.0701256104
Ramsey, J., Bradshaw, H. D., and Schemske, D. W. (2003). Components of reproductive isolation between the monkeyflowers Mimulus lewisii and M. cardinalis (Phrymaceae). Evolution 57, 1520–1534. doi: 10.1111/j.0014-3820.2003.tb00360.x
Riginos, C., Buckley, Y., Blomber, S. P., and Treml, E. A. (2014). Dispersal capacity predicts both population genetic structure and species richness in reef fishes. Am. Nat. 184, 52–64 doi: 10.1086/676505
Rohde, K. (2007). Latitudinal gradients in species diversity?: the search for the primary cause. Oikos 65, 514–527. doi: 10.2307/3545569
Romiguier, J., Gayral, P., Ballenghien, M., Bernard, A., Cahais, V., Chenuil, A., et al. (2014). Comparative population genomics in animals uncovers the determinants of genetic diversity. Nature 515, 261–263. doi: 10.1038/nature13685
Roxburgh, S. H., Shea, K., and Wilson, J. B. (2004). The intermediate disturbance hypothesis: patch dynamics and mechanisms of species coexistence. Ecology 85, 359–371. doi: 10.1890/03-0266
Salisbury, C. L., Seddon, N., Cooney, C. R., and Tobias, J. A. (2012). The latitudinal gradient in dispersal constraints: ecological specialisation drives diversification in tropical birds. Ecol. Lett. 15, 847–855. doi: 10.1111/j.1461-0248.2012.01806.x
Schemske, D. W., Mittelbach, G. G., Cornell, H. V., Sobel, J. M., and Roy, K. (2009). Is there a latitudinal gradient in the importance of biotic interactions? Annu. Rev. Ecol. Evol. Syst. 40, 245–269. doi: 10.1146/annurev.ecolsys.39.110707.173430
Schliewen, U. K., Tautz, D., and Pääbo, S. (1994). Sympatric speciation suggested by monophyly of crater lake cichlids. Nature 368, 629–632. doi: 10.1038/368629a0
Schluter, D. (2000). Ecological character displacement in adaptive radiation. Am. Nat. 156, S4–S16. doi: 10.1086/303412
Schluter, D. (2009). Evidence for ecological speciation and its alternative. Science 323, 737–741. doi: 10.1126/science.1160006
Smith, M. A., Rodriguez, J. J., Whitfield, J. B., Deans, A. R., Janzen, D. H., Hallwachs, W., et al. (2008). Extreme diversity of tropical parasitoid wasps exposed by iterative integration of natural history, DNA barcoding, morphology, and collections. Proc. Natl. Acad. Sci. U.S.A. 105, 12359–12364. doi: 10.1073/pnas.0805319105
Stegen, J. C., Enquist, B. J., and Ferriere, R. (2009). Advancing the metabolic theory of biodiversity. Ecol. Lett. 12, 1001–1015. doi: 10.1111/j.1461-0248.2009.01358.x
Stevens, G. C. (1989). The latitudinal gradient in geographical range: how so many species coexist in the tropics. Am. Nat. 133, 240–256. doi: 10.1086/284913
Surget-Groba, Y., and Kay, K. M. (2013). Restricted gene flow within and between rapidly diverging Neotropical plant species. Mol. Ecol. 22, 4931–4942. doi: 10.1111/mec.12442
Tachida, H., and Iizuka, M. (1991). Fixation probability in spatially changing environments. Genet. Res. 58, 243–251. doi: 10.1017/S0016672300029992
Theim, T. J., Shirk, R. Y., and Givnish, T. J. (2014). Spatial genetic structure in four understory Psychotria species (Rubiaceae) and implications for tropical forest diversity. Am. J. Bot. 101, 1189–1199. doi: 10.3732/ajb.1300460
Tilman, D. (1985). The resource-ratio hypothesis of plant succession. Am. Nat. 125, 827. doi: 10.1086/284382
Turnbull, L., Turnbull, L., Rees, M., Rees, M., Crawley, M. J., and Crawley, M. J. (1999). Seed mass and the competition/colonization trade-of: a sowing experiment. J. Ecol. 87, 899–912. doi: 10.1046/j.1365-2745.1999.00405.x
Weber, C. C., Nabholz, B., Romiguier, J., and Ellegren, H. (2014). Kr/Kc but not dN/dS correlates positively with body mass in birds, raising implications for inferring lineage-specific selection. Genome Biol. 15:542. doi: 10.1186/s13059-014-0542-8
Weir, J. T., and Schluter, D. (2007). The latitudinal gradient in recent speciation and extinction rates of birds and mammals. Science 315, 1574–1576. doi: 10.1126/science.1135590
White, G. M., Boshier, D. H., and Powell, W. (2002). Increased pollen flow counteracts fragmentation in a tropical dry forest: an example from Swietenia humilis Zuccarini. Proc. Natl. Acad. Sci. U.S.A. 99, 2038–2042. doi: 10.1073/pnas.042649999
Wiens, J. J., and Donoghue, M. J. (2004). Historical biogeography, ecology and species richness. Trends Ecol. Evol. 19, 639–644. doi: 10.1016/j.tree.2004.09.011
Williams, S. E., Williams, Y. M., VanDerWal, J., Isaac, J. L., Shoo, L. P., and Johnson, C. N. (2009). Ecological specialization and population size in a biodiversity hotspot: how rare species avoid extinction. Proc. Natl. Acad. Sci. U.S.A. 106(Suppl. 2), 19737–19741. doi: 10.1073/pnas.0901640106
Woolfit, M., and Bromham, L. (2005). Population size and molecular evolution on islands. Proc. Biol. Sci. 272, 2277–2282. doi: 10.1098/rspb.2005.3217
Wright, S., Keeling, J., and Gillman, L. (2006). The road from Santa Rosalia: a faster tempo of evolution in tropical climates. Proc. Natl. Acad. Sci. U.S.A. 103, 7718–7722. doi: 10.1073/pnas.0510383103
Keywords: molecular evolution, latitude diversity gradient, biotic interactions, competition, dispersal, population size
Citation: Pellissier L (2015) Stability and the competition-dispersal trade-off as drivers of speciation and biodiversity gradients. Front. Ecol. Evol. 3:52. doi: 10.3389/fevo.2015.00052
Received: 22 March 2015; Accepted: 05 May 2015;
Published: 03 June 2015.
Edited by:
Hao Zhu, Southern Medical University, ChinaReviewed by:
Miguel Arenas, Consejo Superior de Investigaciones Científicas, SpainNusha Keyghobadi, Western University, Canada
Copyright © 2015 Pellissier. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Loïc Pellissier, Department of Biology, University of Fribourg, Chemin du Musée 10, 1700 Fribourg, Switzerland,bG9pYy5wZWxsaXNzaWVyQHVuaWZyLmNo
 Loïc Pellissier
Loïc Pellissier