- 1Department of Biology, SBA School of Science and Engineering, Lahore University of Management Sciences, Lahore, Pakistan
- 2Department of Neuroscience, The Scripps Research Institute, Jupiter, FL, United States
- 3Center on Aging, The Scripps Research Institute, Jupiter, FL, United States
Drosophila species have successfully spread and adapted to diverse climates across the globe. For Drosophila melanogaster, rotting vegetative matter provides the primary substrate for mating and oviposition, and also acts as a nutritional resource for developing larvae and adult flies. The transitory nature of decaying vegetation exposes D. melanogaster to rapidly changing nutrient availability. As evidenced by their successful global spread, flies are capable of dealing with fluctuating nutritional reserves within their respective ecological niches. Therefore, D. melanogaster populations might contain standing genetic variation to support survival during periods of nutrient scarcity. The natural history and genetic tractability of D. melanogaster make the fly an ideal model for studies on the genetic basis of resistance to nutritional stress. We review artificial selection studies on nutritionally-deprived D. melanogaster and summarize the phenotypic outcomes of selected animals. Many of the reported evolved traits phenocopy mutants of the nutrient-sensing PI3K/Akt pathway. Given that the PI3K/Akt pathway is also responsive to acute nutritional stress, the PI3K/Akt pathway might underlie traits evolved under chronic nutritional deprivation. Future studies that directly test for the genetic mechanisms driving evolutionary responses to nutritional stress will take advantage of the ease in manipulating fly nutrient availability in the laboratory.
Introduction
The colonization of diverse ecological niches exposes organisms to varied environmental stresses. These stresses exert selection pressures which might lead to changes in physiology and behavior. Changes that enhance organismal fitness can result in adaptive evolution, a process by which selection acts on heritable changes in the genetic architecture of a population. Thus, studies on how organisms respond to chronic stressors provide valuable insights on adaptation.
Recent advances have uncovered varied responses to stress in different organisms (Badyaev, 2005). Among the limited number of model systems that have been established for these studies, the vinegar fly, Drosophila melanogaster, remains one of the most tractable. Adaptation to diverse environmental conditions underlies the successful global spread of Drosophila (Stephan and Li, 2007), facilitating its dissemination across multifarious ecological habitats (Keller, 2007). Physiological and behavioral responses have helped the fly adapt to and survive in conditions encompassing a wide range of humidity (Matzkin et al., 2007; Ahuja et al., 2012; Griffin et al., 2016), oxygen concentration (Romero et al., 2007; Zhou et al., 2007), temperature (Bochdanovits and de Jong, 2003; Trotta et al., 2006), and nutrient availability (Kolss et al., 2009). Reproducing diverse environments in the laboratory has revealed how various parameters influence different fly traits, but the molecular mechanisms underlying adaptation to varied habitats are still unclear.
Due to the ease in which the fly diet can be manipulated in the laboratory, nutrition has been a major focus in studies of environmental factors that affect stress-induced acquired and heritable traits (Rion and Kawecki, 2007). In the wild, Drosophila larvae develop in ephemeral habitats such as rotting vegetative matter, resulting in rapid changes to food quantity and quality. Nutritional stress can include starvation due to food deprivation and malnourishment due to nutrient imbalance or depletion. Nutritional scarcity is further exacerbated by competition as greater larval density depletes scant resources. Moreover, other species share habitats with Drosophila, resulting in interspecies competition that further contributes to nutritional stress.
Drosophila larval populations selected on nutrient-poor food exhibit various behavioral and physiological changes (Kolss and Kawecki, 2008; Kolss et al., 2009; Vijendravarma et al., 2011, 2012a,b, 2013; Vijendravarma and Kawecki, 2013). These changes include altered foraging behavior, accelerated development, reduced body weight, and smaller body size (Kolss and Kawecki, 2008; Kolss et al., 2009; Vijendravarma et al., 2011, 2012b, 2013). These phenotypes are maintained when selected populations are subsequently raised on a richer diet, suggesting a genetic basis for these traits (Kolss and Kawecki, 2008; Kolss et al., 2009; Vijendravarma et al., 2012b, 2013). Interestingly, morphological changes in response to nutritional stress phenocopy mutants of PI3-kinase (PI3K)/Akt signaling, a pathway known to be responsive to nutritional status (Chen et al., 1996; Montagne et al., 1999; Oldham et al., 2000; Figure 1). Hence, under prolonged malnutrition, we hypothesize that fly nutrient-sensing pathways might be under selective pressure.
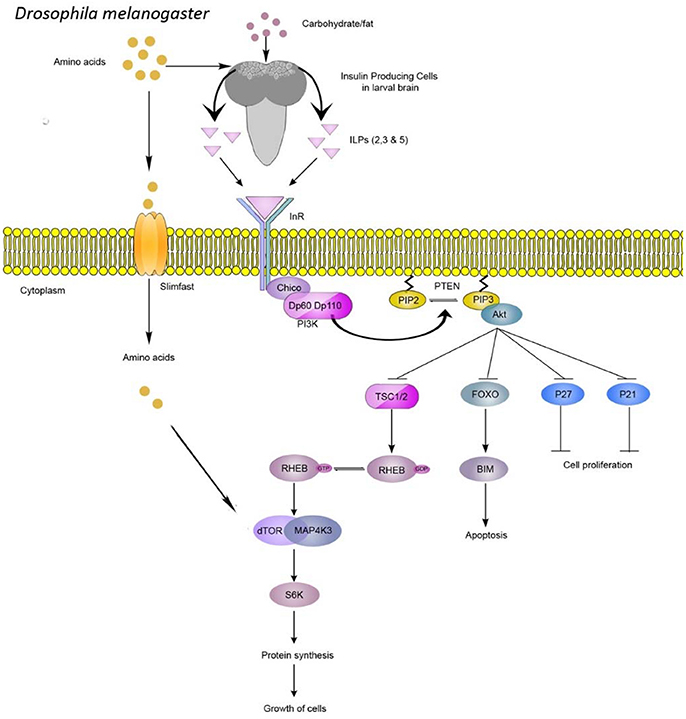
Figure 1. PI3K/Akt pathway plays a central role in nutrient-mediated cell proliferation and growth. In Drosophila, amino acids, sugars, and fats control the release of insulin-like–peptides (ILPs), which bind the insulin receptor homolog (InR). Downstream signaling is mediated by insulin receptor substrate (Chico) binding to InR, followed by recruitment of phosphoinositide-3-kinase (PI3K), which leads to enzymatic conversion of phosphatidylinositol (3,4)-bisphosphate (PIP2) to PIP3. Akt kinase recognizes PIP3 to control cell proliferation and growth. TOR kinase can also be regulated by the availability of extracellular amino acids which enter the cell via Slimfast. TOR kinase activation leads to phosphorylation of effectors such as S6 kinase, which stimulates protein translation and cellular growth.
Genome-wide expression profiles revealed overlapping changes in gene transcript levels between TOR kinase null mutants and starved control animals (Li et al., 2010). Hence, the effects of short-term nutritional stress might be partially mediated by reduced TOR signaling. Despite these studies on acute starvation, it remains unknown whether PI3K/Akt signaling is also altered under chronic nutritional stress. Future studies using targeted genomic approaches might reveal the selection of this pathway and unravel malnutrition-mediated evolutionary responses that underlie tolerance to nutritional stress. Here, we review the studies that have evolved phenotypic responses under chronic malnutrition and highlight the possibility that the PI3K/Akt pathway is under selection during prolonged nutritional adversity in flies. These perspectives inform our understanding of the molecular basis of the physiological and behavioral changes associated with adaptation to malnutrition.
Body Size and Weight
Evolved body size is associated with available nutrition. Flies selected for 29 generations on low concentration sugar and protein medium exhibit a number of adaptive consequences, including smaller body size and reduced weight (Kolss et al., 2009). Conversely, a quasi-natural selection experiment using a protein-rich diet favored large flies with greater dry mass and reduced lipid content compared to controls (Kristensen et al., 2011). Interestingly, mutations in components of the PI3K/Akt pathway impede larval growth and also reduce body size. Genetic manipulations of ILPs, InR, Chico, PI3K, Akt, TOR, S6K, FOXO, and Slimfast (Figure 1) have all been found to reduce body weight or size in larvae or adults (Böhni et al., 1999; Montagne et al., 1999; Oldham et al., 2000; Zhang et al., 2000, 2009; Britton et al., 2002; Colombani et al., 2003; Hietakangas and Cohen, 2007; Mattila et al., 2009), mimicking development under poor nutritional conditions (Kolss et al., 2009; Vijendravarma et al., 2011, 2012a). Such parallels in the literature bolster our hypothesis that the nutrition-sensing PI3K/Akt pathway might be under selective pressure during prolonged malnutrition. Changes in body size or weight might be associated with multiple factors, including the critical size required for initiation of metamorphosis (Tennessen and Thummel, 2011). Fly growth during development is characterized by two distinct phases: (1) a pre-critical-weight phase that can vary in time depending on the nutritional conditions and (2) a post-critical-weight phase where nutritional stress can no longer delay pupariation. During the pre-critical-weight phase, insulin signaling affects development time but not body size whereas, in the post-critical-weight period, it affects body size and not development time (Mirth et al., 2005; Shingleton et al., 2005). Hence, any changes in critical weight threshold can have a direct impact on both development time and body size. Interestingly, flies selected under chronic malnutrition develop more quickly (Kolss et al., 2009), possibly due to reduced critical size threshold (Vijendravarma et al., 2012a). Although mutations in PI3K/Akt pathway components result in a developmental delay (Zhang et al., 2000; Rulifson et al., 2002; Layalle et al., 2008), shortened development time in malnutrition-selected populations might be caused by altered prothoracicotropic hormone signaling, which has been proposed to regulate critical size upstream of TOR function (Layalle et al., 2008).
Once critical weight is attained, the prothoracic gland (PG) induces pupariation through release of ecdysone (Mirth et al., 2005). Hence, a lowered critical weight threshold results in earlier release of ecdysone. PG cells grow in size due to an organ-specific augmentation in PI3K/Akt signaling that results in strikingly reduced critical weight culminating in an increase in the basal level of ecdysteroid biosynthesis (Colombani et al., 2005; Mirth et al., 2005). This consequently results in accelerated development and reduced body size due to a decrease in cell number (Colombani et al., 2005; Mirth et al., 2005). Furthermore, it was shown that the growth rate of flies with an enlarged PG was dependent on nutritional conditions (Mirth et al., 2005). Together, these results indicate that PI3K/Akt-mediated growth of the PG negatively regulates the developmental time and body size of the fly. These observations might suggest that increased PG-mediated ecdysone signaling underlies the physiological phenotypes observed in fly populations under chronic malnutrition (Kolss et al., 2009; Vijendravarma et al., 2011, 2012a; Figure 2).
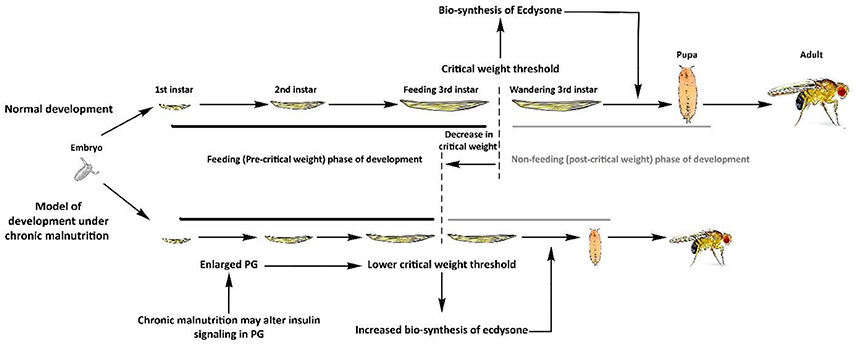
Figure 2. Proposed model for the development of fly populations selected under chronic malnutrition. Under normal rearing conditions, larvae feed during the pre-critical phase to achieve critical weight, which results in release of an ecdysone pulse. This hormonal shift ceases larval feeding and drives the larval-to-pupal transition. Under chronic malnutrition, the prothoracic gland (PG) might become nutritionally desensitized through elevated PI3K/Akt signaling, which consequently enlarges its size. This lowers the critical weight threshold and results in increased synthesis of ecdysone, driving an early larval-to pupal transition. This shift in the critical weight threshold results in accelerated development time and lower body size and weight.
How might PG-mediated ecdysone signaling be enhanced under malnourished conditions, when PI3K/Akt signaling is typically reduced? Specific tissues differ in their sensitivity to nutrients and PI3K/Akt signaling (Shingleton et al., 2005) to result in organ-specific growth—a phenomenon known as allometry. Some organs, such as the male genitalia of the fly, do not reduce in size in response to nutritional stress; they are insensitive to the nutritional environment (Shingleton et al., 2005; Shingleton, 2010; Tang et al., 2011). This reduced sensitivity to nutrition is attained by maintaining high TOR signaling, or through tissue-specific reduction in the FOXO transcription factor under low nutritional conditions (Tang et al., 2011; Koyama et al., 2013). This mechanism would allow organs to become less sensitive to nutrition in order to mollify the effects of poor nutrition. Hence, it is possible that, under chronic nutritional stress, the PG might be less nutritionally sensitive and maintain or increase PI3K/Akt signaling, resulting in a relatively enlarged PG. It will be of interest to test if the expression of PI3k/Akt signaling components is indeed upregulated under chronic nutritional stress.
Broad regulatory functions of Akt also include cell growth, control of apoptosis, and cell proliferation (Manning and Cantley, 2007). Hence, the PI3K/Akt pathway can directly impact cell size and number to potentially regulate body size (Goberdhan et al., 1999; Verdu et al., 1999; Zhang et al., 2000; Miron et al., 2001). Indeed, mutants of chico show reduced body size due to a reduction in both cell number and size (Böhni et al., 1999). In contrast, evolutionary changes in wing size under chronic malnutrition were mediated entirely by reduced cell number (Vijendravarma et al., 2011). These studies might imply that pathway components upstream or at the level of Akt might be the targets of selection under nutritional stress.
Foraging Behavior
In addition to physiological responses, foraging behavior is also influenced in Drosophila exposed to poor nutritional conditions (Sokolowski et al., 1997; Vijendravarma et al., 2012b). Drosophila larvae spend most of their time feeding and naturally exhibit a foraging polymorphism driven by the foraging (for) gene (Osborne et al., 1997). for encodes a cGMP-dependent protein kinase (PKG) (Osborne et al., 1997) and has been shown to alter larval nutrient acquisition in response to food deprivation (Kaun et al., 2007, 2008). In the presence of food, rovers (forR) have a longer foraging path and a higher propensity to leave a food patch as compared to sitters (forS). However, in the absence of food, these variants do not exhibit any difference in activity (Sokolowski et al., 1984; Kaun et al., 2007).
The evolution of Drosophila foraging behavior has been reported under density-dependent selection whereby high larval density reduces food access and induces nutritional stress on unsuccessful competitors (Ruiz-Dubreuil et al., 1996; Sokolowski et al., 1997). Interestingly, the rover phenotype was selected under high-density rearing (Sokolowski et al., 1997). In contrast, populations evolved under chronic larval malnutrition without crowding showed a shorter foraging path length compared to unselected controls (Vijendravarma et al., 2012b). Thus, the context of how malnutrition is applied plays a role in the evolution of behavior. In a competitive environment, greater mobility increases the likelihood of exploiting heterogeneous food or finding a better nutritional niche. However, the energetic cost of locomotion in Drosophila larvae (Berrigan and Pepin, 1995) might not be favorable in a non-competitive context.
Similar to larvae, adult rovers and sitters show variation in their feeding behaviors. Rover adults exhibit a greater proboscis extension response to sucrose than sitters under food deprivation (Scheiner et al., 2004; Hughson et al., 2017). After sucrose feeding, rover and sitter adults exhibit different walking patterns—rovers travel further compared to sitters (Pereira and Sokolowski, 1993). In addition, rovers are more sensitive to starvation and show extended food search patterns following food deprivation (Hughson et al., 2017). Genomic and metabolomic analyses of rover and sitter adults demonstrated that the insulin-mediated PI3K/Akt pathway might be responsible for the rover- and sitter-dependent feeding behaviors under food deprivation (Kent et al., 2009). Compared to sitters, rovers show higher PI3K/Akt signaling (Kent et al., 2009). Combining for alleles with PI3K/Akt signaling mutants changed rover-like behavior into sitter-like behavior in transheterozygote animals, further suggesting an epistatic link between the PI3K/Akt and for pathways (Kent et al., 2009). This also indicates that PI3K/Akt signaling attenuation might lead to sitter-like behavior under nutrient deprivation, further supporting our speculation that PI3K/Akt pathway components might experience selective pressures under chronic malnutrition. Future work could help assess potential molecular interactions between the PKG and PI3K/Akt pathways.
Learning and Memory
Selection under long-term malnutrition was also reported to affect the learning ability of Drosophila (Kolss and Kawecki, 2008). Flies reared on a high carbohydrate-low protein diet exhibit impaired visual learning and memory formation (Xia et al., 1997). Conversely, fly populations selected for 51 generations on conditions that improve learning and memory (Mery and Kawecki, 2002) exhibited reduced larval competitive ability for a limited food source (Mery and Kawecki, 2003). This suggests a direct relationship between the availability of nutrients and learning and memory. However, the underlying mechanism of the effect of nutritional stress on memory formation is still unclear.
PI3K/Akt signaling has also been implicated in long-term memory formation (Chambers et al., 2015). Genetic impairment of this signaling cascade in Drosophila larval mushroom bodies, a pair of structures responsible for memory formation, hampers feeding behavior by causing a reduction in food intake (Zhao and Campos, 2012). Interestingly, the for rover/sitter polymorphism has also been shown to mediate associative learning and memory in Drosophila (Mery et al., 2007). forR flies tend to have weaker long-term memory compared to that of forS (Mery et al., 2007). Taken together, these studies further support the idea that there is a molecular link between the PI3K/Akt and PKG pathways (Kent et al., 2009) and that PI3K/Akt might be a possible selection target under chronic malnutrition.
Cannibalism
Drosophila melanogaster is an herbivorous and detritivorous species which primarily feeds on a rotten vegetative matter and associated microbes (Carson, 1971; Yamada et al., 2015). However, under nutritional stress, larvae can cannibalize and feed on conspecifics (Vijendravarma et al., 2013; Ahmad et al., 2015). Larvae selected on chronic malnutrition locate and prey upon conspecific victims more efficiently than non-selected controls, suggesting that cannibalism is an adaptive trait (Vijendravarma et al., 2013). Typically, Drosophila is morphologically ill-equipped to scavenge on a carnivorous diet. However, feeding exclusively on cannibalistic diet induces larval mouth hook plasticity by increasing the number of teeth (Vijendravarma et al., 2013). This allows larvae to puncture a victim's cuticle during cannibalism. The adaptive significance of this plastic phenotype, as well as its molecular basis, are still not known. Since cannibalistic behavior is reported to evolve under chronic malnutrition, it will be interesting to probe if there exists a link between the PI3K/Akt pathway and cannibalism.
Conclusion
Recent studies have exploited the advantages of using Drosophila to demonstrate evolutionary responses to chronic nutritional stress (Kolss and Kawecki, 2008; Kolss et al., 2009; Vijendravarma et al., 2011, 2012a,b, 2013, 2015; Zajitschek and Zajitschek, 2016). However, the genetic and molecular mechanisms underlying changes in physiology and behavior are still mostly unknown. By reviewing phenotypic studies of fly populations under chronic malnutrition, we speculate that components of the nutrient-sensing PI3K/Akt pathway might be among the selection targets under nutritional stress; a number of physiological and behavioral traits that emerge under nutritional stress parallel those of PI3K/Akt pathway mutants. If the PI3K/Akt pathway is a target of selective pressure under chronic malnutrition, genetic signatures of selection and altered expression of its components could be revealed by genomic sequencing and expression profiling. Additionally, proteomic profiles of animals selected under chronic nutritional stress could be used to identify post-translational modifications that contribute to alterations of the PI3K/Akt pathway.
Next-generation sequencing technologies have greatly facilitated evolve-and-resequence studies, which attempt to identify genetic variants causative of evolved traits. Recently, resequenced D. melanogaster strains evolved under low oxygen levels showed signatures of selection acting on Notch pathway signaling; follow-up analyses on genetic mutants supported the idea that the Notch pathway is involved in hypoxic responses (Zhou et al., 2011). Furthermore, D. melanogaster strains sequenced after multi-generational exposure to the natural fly virus, Drosophila C virus (DCV) identified candidate DCV resistance genes that were subsequently verified in mutational analyses (Martins et al., 2014). While genome-wide approaches might uncover pathways that are under selective pressure, including or independent of PI3K/Akt signaling, there will still be challenges in identifying candidate genes involved in polygenic traits. In fact, many evolve-and-resequence studies using D. melanogaster have highlighted the difficulties in characterizing experimental evolutionary changes given the complexities revealed in associating phenotypic output with specific alterations within the genome-wide databases (Turner et al., 2011; Turner and Miller, 2012; Reed et al., 2014; Griffin et al., 2016). Thus, although genomic sequencing could facilitate future studies of the molecular basis of adaptation to nutritional stress, a more targeted approach informed by a comprehensive examination of relevant studies might be more fruitful.
A major technical issue to overcome is the lack of a defined standard medium, resulting in the use of different fly food recipes across labs (Marx, 2015). Varying different components of fly food can alter the carbohydrate-protein ratio (Bruce et al., 2013; Tatar et al., 2014; Lee, 2015). These changes might have large impacts on studies dealing with selection over multiple generations. Furthermore, altering the concentration of different nutrients can result in a range of responses in flies (Lee et al., 2008; Andersen et al., 2010; Sisodia et al., 2015) and preclude the identification of specific nutritional changes associated with adaptive responses. Future studies using chemically defined diets will improve the comparison of empirical findings across labs (Piper et al., 2014). A nutritional geometry approach (Simpson and Raubenheimer, 1999, 2012) might be needed to comprehensively quantify the specific contributions of macro- or micro-nutrients. Fortunately, the large number of diets needed for these types of studies are feasible using Drosophila. The classical components of the PI3K/Akt pathway are evolutionarily and functionally conserved in mammals (Oldham, 2011). Therefore, studies using the fly model may help us better understand the molecular basis of development and evolution across species.
Author Contributions
MA drafted the paper. All authors contributed equally to the conception and revision of the paper.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by grant R01AG045036 from the National Institutes of Health (WJ) and the Higher Education Commission of Pakistan (HEC/2228 and HEC/5909, MT). EK is an O'Keeffe Neuroscience Scholar of the Esther B. O'Keeffe Charitable Foundation.
References
Ahmad, M., Chaudhary, S. U., Afzal, A. J., and Tariq, M. (2015). Starvation-induced dietary behaviour in Drosophila melanogaster larvae and adults. Sci. Rep. 5:14285. doi: 10.1038/srep14285
Ahuja, A., Tanwar, R. S., Tyagi, P. K., Sarsar, V., Paudwal, M., and Kumar, P. (2012). Local adaption in life history traits of Drosophila melanogaster in extreme conditions of humidity. Int. J. Anim. Biotechnol. 2, 1–5.
Andersen, L. H., Kristensen, T. N., Loeschcke, V., Toft, S., and Mayntz, D. (2010). Protein and carbohydrate composition of larval food affects tolerance to thermal stress and desiccation in adult Drosophila melanogaster. J. Insect Physiol. 56, 336–340. doi: 10.1016/j.jinsphys.2009.11.006
Badyaev, A. V. (2005). “Role of stress in evolution: from individual adaptability to evolutionary adaptation,” in Variation: A Central Concept in Biology, eds B. Hallgrímsson and B. K. Hall (Burlington, MA; San Diego, CA; London, UK: Elsevier Academic Press), 277–302.
Berrigan, D., and Pepin, D. J. (1995). How maggots move - allometry and kinematics of crawling in larval diptera. J. Insect Physiol. 41, 329–337. doi: 10.1016/0022-1910(94)00113-U
Bochdanovits, Z., and de Jong, G. (2003). Experimental evolution in Drosophila melanogaster: interaction of temperature and food quality selection regimes. Evolution 57, 1829–1836. doi: 10.1111/j.0014-3820.2003.tb00590.x
Böhni, R., Riesgo-Escovar, J., Oldham, S., Brogiolo, W., Stocker, H., Andruss, B. F., et al. (1999). Autonomous control of cell and organ size by CHICO, a Drosophila homolog of vertebrate IRS1-4. Cell 97, 865–875. doi: 10.1016/S0092-8674(00)80799-0
Britton, J. S., Lockwood, W. K., Li, L., Cohen, S. M., and Edgar, B. A. (2002). Drosophila's insulin/PI3-kinase pathway coordinates cellular metabolism with nutritional conditions. Dev. Cell 2, 239–249. doi: 10.1016/S1534-5807(02)00117-X
Bruce, K. D., Hoxha, S., Carvalho, G. B., Yamada, R., Wang, H. D., Karayan, P., et al. (2013). High carbohydrate-low protein consumption maximizes Drosophila lifespan. Exp. Gerontol. 48, 1129–1135. doi: 10.1016/j.exger.2013.02.003
Carson, H. L. (1971). The Ecology of Drosophila Breeding Sites. Honolulu, HI: University of Hawaii Press.
Chambers, D. B., Androschuk, A., Rosenfelt, C., Langer, S., Harding, M., and Bolduc, F. V. (2015). Insulin signaling is acutely required for long-term memory in Drosophila. Front. Neural Circ. 9:8. doi: 10.3389/fncir.2015.00008
Chen, C., Jack, J., and Garofalo, R. S. (1996). The Drosophila insulin receptor is required for normal growth. Endocrinology 137, 846–856. doi: 10.1210/endo.137.3.8603594
Colombani, J., Bianchini, L., Layalle, S., Pondeville, E., Dauphin-Villemant, C., Antoniewski, C., et al. (2005). Antagonistic actions of ecdysone and insulins determine final size in Drosophila. Science 310, 667–670. doi: 10.1126/science.1119432
Colombani, J., Raisin, S., Pantalacci, S., Radimerski, T., Montagne, J., and Léopold, P. (2003). A nutrient sensor mechanism controls Drosophila growth. Cell 114, 739–749. doi: 10.1016/S0092-8674(03)00713-X
Goberdhan, D. C., Paricio, N., Goodman, E. C., Mlodzik, M., and Wilson, C. (1999). Drosophila tumor suppressor PTEN controls cell size and number by antagonizing the Chico/PI3-kinase signaling pathway. Genes Dev. 13, 3244–3258. doi: 10.1101/gad.13.24.3244
Griffin, P. C., Hangartner, S. B., Fournier-Level, A., and Hoffmann, A. A. (2016). Genomic trajectories to desiccation resistance: convergence and divergence among replicate selected Drosophila lines. Genetics 205, 871–890. doi: 10.1534/genetics.116.187104
Hietakangas, V., and Cohen, S. M. (2007). Re-evaluating AKT regulation: role of TOR complex 2 in tissue growth. Genes Dev. 21, 632–637. doi: 10.1101/gad.416307
Hughson, B. N., Anreiter, I., Jackson Chornenki, N. L., Murphy, K. R., Ja, W. W., Huber, R., et al. (2017). The adult foraging assay (AFA) detects strain and food-deprivation effects in feeding-related traits of Drosophila melanogaster. J. Insect Physiol. doi: 10.1016/j.jinsphys.2017.08.011. [Epub ahead of print].
Kaun, K. R., Chakaborty-Chatterjee, M., and Sokolowski, M. B. (2008). Natural variation in plasticity of glucose homeostasis and food intake. J. Exp. Biol. 211, 3160–3166. doi: 10.1242/jeb.010124
Kaun, K. R., Riedl, C. A., Chakaborty-Chatterjee, M., Belay, A. T., Douglas, S. J., Gibbs, A. G., et al. (2007). Natural variation in food acquisition mediated via a Drosophila cGMP-dependent protein kinase. J. Exp. Biol. 210, 3547–3558. doi: 10.1242/jeb.006924
Keller, A. (2007). Drosophila melanogaster's history as a human commensal. Curr. Biol. 17, R77– R81. doi: 10.1016/j.cub.2006.12.031
Kent, C. F., Daskalchuk, T., Cook, L., Sokolowski, M. B., and Greenspan, R. J. (2009). The Drosophila foraging gene mediates adult plasticity and gene-environment interactions in behaviour, metabolites, and gene expression in response to food deprivation. PLoS Genet. 5:e1000609. doi: 10.1371/journal.pgen.1000609
Kolss, M., and Kawecki, T. J. (2008). Reduced learning ability as a consequence of evolutionary adaptation to nutritional stress in Drosophila melanogaster. Ecol. Entomol. 33, 583–588. doi: 10.1111/j.1365-2311.2008.01007.x
Kolss, M., Vijendravarma, R. K., Schwaller, G., and Kawecki, T. J. (2009). Life-history consequences of adaptation to larval nutritional stress in Drosophila. Evolution 63, 2389–2401. doi: 10.1111/j.1558-5646.2009.00718.x
Koyama, T., Mendes, C. C., and Mirth, C. K. (2013). Mechanisms regulating nutrition-dependent developmental plasticity through organ-specific effects in insects. Front. Physiol. 4:263. doi: 10.3389/fphys.2013.00263
Kristensen, T. N., Overgaard, J., Loeschcke, V., and Mayntz, D. (2011). Dietary protein content affects evolution for body size, body fat and viability in Drosophila melanogaster. Biol. Lett. 7, 269–272. doi: 10.1098/rsbl.2010.0872
Layalle, S., Arquier, N., and Léopold, P. (2008). The TOR pathway couples nutrition and developmental timing in Drosophila. Dev. Cell 15, 568–577. doi: 10.1016/j.devcel.2008.08.003
Lee, K. P. (2015). Dietary protein: carbohydrate balance is a critical modulator of lifespan and reproduction in Drosophila melanogaster: a test using a chemically defined diet. J. Insect Physiol. 75, 12–19. doi: 10.1016/j.jinsphys.2015.02.007
Lee, K. P., Simpson, S. J., Clissold, F. J., Brooks, R., Ballard, J. W., Taylor, P. W., et al. (2008). Lifespan and reproduction in Drosophila: new insights from nutritional geometry. Proc. Natl. Acad. Sci. U.S.A. 105, 2498–2503. doi: 10.1073/pnas.0710787105
Li, L., Edgar, B. A., and Grewal, S. S. (2010). Nutritional control of gene expression in Drosophila larvae via TOR, Myc and a novel cis-regulatory element. BMC Cell Biol. 11:7. doi: 10.1186/1471-2121-11-7
Manning, B. D., and Cantley, L. C. (2007). AKT/PKB signaling: navigating downstream. Cell 129, 1261–1274. doi: 10.1016/j.cell.2007.06.009
Martins, N. E., Faria, V. G., Nolte, V., Schlötterer, C., Teixeira, L., Sucena É, E., et al. (2014). Host adaptation to viruses relies on few genes with different cross-resistance properties. Proc. Natl. Acad. Sci. U.S.A. 111, 5938–5943. doi: 10.1073/pnas.1400378111
Mattila, J., Bremer, A., Ahonen, L., Kostiainen, R., and Puig, O. (2009). Drosophila FoxO regulates organism size and stress resistance through an adenylate cyclase. Mol. Cell. Biol. 29, 5357–5365. doi: 10.1128/MCB.00302-09
Matzkin, L. M., Watts, T. D., and Markow, T. A. (2007). Desiccation resistance in four Drosophila species: sex and population effects. Fly 1, 268–273. doi: 10.4161/fly.5293
Mery, F., and Kawecki, T. J. (2002). Experimental evolution of learning ability in fruit flies. Proc. Natl. Acad. Sci. U.S.A. 99, 14274–14279. doi: 10.1073/pnas.222371199
Mery, F., and Kawecki, T. J. (2003). A fitness cost of learning ability in Drosophila melanogaster. Proc. R. Soc. B 270, 2465–2469. doi: 10.1098/rspb.2003.2548
Mery, F., Belay, A. T., So, A. K., Sokolowski, M. B., and Kawecki, T. J. (2007). Natural polymorphism affecting learning and memory in Drosophila. Proc. Natl. Acad. Sci. U.S.A. 104, 13051–13055. doi: 10.1073/pnas.0702923104
Miron, M., Verd,ú, J., Lachance, P. E., Birnbaum, M. J., Lasko, P. F., and Sonenberg, N. (2001). The translational inhibitor 4E-BP is an effector of PI (3) K/Akt signalling and cell growth in Drosophila. Nat. Cell Biol. 3, 596–601. doi: 10.1038/35078571
Mirth, C., Truman, J. W., and Riddiford, L. M. (2005). The role of the prothoracic gland in determining critical weight for metamorphosis in Drosophila melanogaster. Curr. Biol. 15, 1796–1807. doi: 10.1016/j.cub.2005.09.017
Montagne, J., Stewart, M. J., Stocker, H., Hafen, E., Kozma, S. C., and Thomas, G. (1999). Drosophila S6 kinase: a regulator of cell size. Science 285, 2126–2129. doi: 10.1126/science.285.5436.2126
Oldham, S. (2011). Obesity and nutrient sensing TOR pathway in flies and vertebrates: functional conservation of genetic mechanisms. Trends Endocrinol. Metab. 22, 45–52. doi: 10.1016/j.tem.2010.11.002
Oldham, S., Montagne, J., Radimerski, T., Thomas, G., and Hafen, E. (2000). Genetic and biochemical characterization of dTOR, the Drosophila homolog of the target of rapamycin. Genes Dev. 14, 2689–2694. doi: 10.1101/gad.845700
Osborne, K. A., Robichon, A., Burgess, E., Butland, S., Shaw, R. A., Coulthard, A., et al. (1997). Natural behavior polymorphism due to a cGMP-dependent protein kinase of Drosophila. Science 277, 834–836. doi: 10.1126/science.277.5327.834
Pereira, H. S., and Sokolowski, M. B. (1993). Mutations in the larval foraging gene affect adult locomotory behavior after feeding in Drosophila melanogaster. Proc. Natl. Acad. Sci. U.S.A. 90, 5044–5046. doi: 10.1073/pnas.90.11.5044
Piper, M. D., Blanc, E., Leitão-Gonçalves, R., Yang, M., He, X., Linford, N. J., et al. (2014). A holidic medium for Drosophila melanogaster. Nat. Methods 11, 100–105. doi: 10.1038/nmeth.2731
Reed, L. K., Lee, K., Zhang, Z., Rashid, L., Poe, A., Hsieh, B., et al. (2014). Systems genomics of metabolic phenotypes in wild-type Drosophila melanogaster. Genetics 197, 781–793. doi: 10.1534/genetics.114.163857
Rion, S., and Kawecki, T. J. (2007). Evolutionary biology of starvation resistance: what we have learned from Drosophila. J. Evol. Biol. 20, 1655–1664. doi: 10.1111/j.1420-9101.2007.01405.x
Romero, N. M., Dekanty, A., and Wappner, P. (2007). Cellular and developmental adaptations to hypoxia: a Drosophila perspective. Meth. Enzymol. 435, 123–144. doi: 10.1016/S0076-6879(07)35007-6
Ruiz-Dubreuil, G., Burnet, B., Connolly, K., and Furness, P. (1996). Larval foraging behaviour and competition in Drosophila melanogaster. Heredity 76 (Pt 1), 55–64. doi: 10.1038/hdy.1996.7
Rulifson, E. J., Kim, S. K., and Nusse, R. (2002). Ablation of insulin-producing neurons in flies: growth and diabetic phenotypes. Science 296, 1118–1120. doi: 10.1126/science.1070058
Scheiner, R., Sokolowski, M. B., and Erber, J. (2004). Activity of cGMP-dependent protein kinase (PKG) affects sucrose responsiveness and habituation in Drosophila melanogaster. Learn. Mem. 11, 303–311. doi: 10.1101/lm.71604
Shingleton, A. W. (2010). The regulation of organ size in Drosophila: Physiology, plasticity, patterning and physical force. Organogenesis 6, 76–87. doi: 10.4161/org.6.2.10375
Shingleton, A. W., Das, J., Vinicius, L., and Stern, D. L. (2005). The temporal requirements for insulin signaling during development in Drosophila. PLoS Biol. 3:e289. doi: 10.1371/journal.pbio.0030289
Simpson, S. J., and Raubenheimer, D. (1999). Assuaging nutritional complexity: a geometrical approach. Proc. Nutr. Soc. 58, 779–789. doi: 10.1017/S0029665199001068
Simpson, S. J., and Raubenheimer, D. (2012). The Nature of Nutrition: A Unifying Framework From Animal Adaptation to Human Obesity. Princeton, NJ: Princeton University Press.
Sisodia, S., Verma, P., and Singh, B. N. (2015). Effect of diet quality and associated metabolic changes in adult stress response and life-history traits in Drosophila ananassae. Curr. Sci. 109, 1687. doi: 10.18520/v109/i9/1687-1696
Sokolowski, M. B., Kent, C., and Wong, J. (1984). Drosophila larval foraging behaviour: developmental stages. Anim. Behav. 32, 645–651. doi: 10.1016/S0003-3472(84)80139-6
Sokolowski, M. B., Pereira, H. S., and Hughes, K. (1997). Evolution of foraging behavior in Drosophila by density-dependent selection. Proc. Natl. Acad. Sci. U.S.A. 94, 7373–7377. doi: 10.1073/pnas.94.14.7373
Stephan, W., and Li, H. (2007). The recent demographic and adaptive history of Drosophila melanogaster. Heredity 98, 65–68. doi: 10.1038/sj.hdy.6800901
Tang, H. Y., Smith-Caldas, M. S., Driscoll, M. V., Salhadar, S., and Shingleton, A. W. (2011). FOXO regulates organ-specific phenotypic plasticity in Drosophila. PLoS Genet. 7:e1002373. doi: 10.1371/journal.pgen.1002373
Tatar, M., Post, S., and Yu, K. (2014). Nutrient control of Drosophila longevity. Trends Endocrinol. Metab. 25, 509–517. doi: 10.1016/j.tem.2014.02.006
Tennessen, J. M., and Thummel, C. S. (2011). Coordinating growth and review maturation — Insights from Drosophila. Curr. Biol. 21, R750– R757. doi: 10.1016/j.cub.2011.06.033
Trotta, V., Calboli, F. C., Ziosi, M., Guerra, D., Pezzoli, M. C., David, J. R., et al. (2006). Thermal plasticity in Drosophila melanogaster: a comparison of geographic populations. BMC Evol. Biol. 6:67. doi: 10.1186/1471-2148-6-67
Turner, T. L., and Miller, P. M. (2012). Investigating natural variation in Drosophila courtship song by the evolve and resequence approach. Genetics 191, 633–642. doi: 10.1534/genetics.112.139337
Turner, T. L., Stewart, A. D., Fields, A. T., Rice, W. R., and Tarone, A. M. (2011). Population-based resequencing of experimentally evolved populations reveals the genetic basis of body size variation in Drosophila melanogaster. PLoS Genet. 7:e1001336. doi: 10.1371/journal.pgen.1001336
Verdu, J., Buratovich, M. A., Wilder, E. L., and Birnbaum, M. J. (1999). Cell-autonomous regulation of cell and organ growth in Drosophila by Akt/PKB. Nat. Cell Biol. 1, 500–506. doi: 10.1038/70293
Vijendravarma, R. K., and Kawecki, T. J. (2013). Epistasis and maternal effects in experimental adaptation to chronic nutritional stress in Drosophila. J. Evol. Biol. 26, 2566–2580. doi: 10.1111/jeb.12248
Vijendravarma, R. K., Narasimha, S., and Kawecki, T. J. (2011). Plastic and evolutionary responses of cell size and number to larval malnutrition in Drosophila melanogaster. J. Evol. Biol. 24, 897–903. doi: 10.1111/j.1420-9101.2010.02225.x
Vijendravarma, R. K., Narasimha, S., and Kawecki, T. J. (2012a). Chronic malnutrition favours smaller critical size for metamorphosis initiation in Drosophila melanogaster. J. Evol. Biol. 25, 288–292. doi: 10.1111/j.1420-9101.2011.02419.x
Vijendravarma, R. K., Narasimha, S., and Kawecki, T. J. (2012b). Evolution of foraging behaviour in response to chronic malnutrition in Drosophila melanogaster. Proc. R. Soc. B 279, 3540–3546. doi: 10.1098/rspb.2012.0966
Vijendravarma, R. K., Narasimha, S., and Kawecki, T. J. (2013). Predatory cannibalism in Drosophila melanogaster larvae. Nat. Commun. 4:1789. doi: 10.1038/ncomms2744
Vijendravarma, R. K., Narasimha, S., Chakrabarti, S., Babin, A., Kolly, S., Lemaitre, B., et al. (2015). Gut physiology mediates a trade-off between adaptation to malnutrition and susceptibility to food-borne pathogens. Ecol. Lett. 18, 1078–1086. doi: 10.1111/ele.12490
Xia, S. Z., Liu, L., Feng, C. H., and Guo, A. K. (1997). Nutritional effects on operant visual learning in Drosophila melanogaster. Physiol. Behav. 62, 263–271. doi: 10.1016/S0031-9384(97)00113-3
Yamada, R., Deshpande, S. A., Bruce, K. D., Mak, E. M., and Ja, W. W. (2015). Microbes promote amino acid harvest to rescue undernutrition in Drosophila. Cell Rep. 10, 865–872. doi: 10.1016/j.celrep.2015.01.018
Zajitschek, F., and Zajitschek, S. R. (2016). Evolution under dietary restriction increases male reproductive performance without survival cost. Proc. R. Soc. B Biol. Sci. 283:20152726. doi: 10.1098/rspb.2015.2726
Zhang, H., Liu, J., Li, C. R., Momen, B., Kohanski, R. A., and Pick, L. (2009). Deletion of Drosophila insulin-like peptides causes growth defects and metabolic abnormalities. Proc. Natl. Acad. Sci. U.S.A. 106, 19617–19622. doi: 10.1073/pnas.0905083106
Zhang, H., Stallock, J. P., Ng, J. C., Reinhard, C., and Neufeld, T. P. (2000). Regulation of cellular growth by the Drosophila target of rapamycin dTOR. Genes Dev. 14, 2712–2724. doi: 10.1101/gad.835000
Zhao, X. L., and Campos, A. R. (2012). Insulin signalling in mushroom body neurons regulates feeding behaviour in Drosophila larvae. J. Exp. Biol. 215, 2696–2702. doi: 10.1242/jeb.066969
Zhou, D., Udpa, N., Gersten, M., Visk, D. W., Bashir, A., Xue, J., et al. (2011). Experimental selection of hypoxia-tolerant Drosophila melanogaster. Proc. Natl. Acad. Sci. U.S.A. 108, 2349–2354. doi: 10.1073/pnas.1010643108
Keywords: adaptation, artificial selection, evolved traits, experimental evolution, malnutrition, nutritional deprivation, PI3K/Akt, starvation
Citation: Ahmad M, Keebaugh ES, Tariq M and Ja WW (2018) Evolutionary Responses of Drosophila melanogaster Under Chronic Malnutrition. Front. Ecol. Evol. 6:47. doi: 10.3389/fevo.2018.00047
Received: 29 October 2017; Accepted: 04 April 2018;
Published: 19 April 2018.
Edited by:
Adalbert Balog, Sapientia Hungarian University of Transylvania, RomaniaReviewed by:
Wendy Hood, Auburn University, United StatesZsolt Czekes, Babeş-Bolyai University, Romania
Copyright © 2018 Ahmad, Keebaugh, Tariq and Ja. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Muhammad Tariq, bS50YXJpcUBsdW1zLmVkdS5waw==
William W. Ja, d2phQHNjcmlwcHMuZWR1
†Shared co-first authorship.
 Muhammad Ahmad
Muhammad Ahmad Erin S. Keebaugh
Erin S. Keebaugh Muhammad Tariq
Muhammad Tariq William W. Ja
William W. Ja