Abstract
Despite increasing appreciation of the speciation continuum, delimiting and describing new species is a major yet necessary challenge of modern phylogeography to help optimize conservation efforts. In amphibians, the lack of phenotypic differences between closely-related taxa, their complex, sometimes unresolved phylogenetic relationships, and their potential to hybridize all act to blur taxonomic boundaries. Here we implement a multi-disciplinary approach to evaluate the nature of two deeply-diverged mitochondrial lineages previously documented in Italian tree frogs (Hyla intermedia s. l.), distributed north and south of the Northern Apennine Mountains. Based on evidence from mitochondrial phylogenetics, nuclear phylogenomics, hybrid zone population genomics, niche modeling analyses, and biometric assessments, we propose that these lineages be considered distinct, cryptic species. Both mitochondrial and nuclear data affirm that they belong to two monophyletic clades of Pliocene divergence (~3.5 My), only admixing over a relatively narrow contact zone restricted to the southeast of the Po Plain (50–100 km). These characteristics are comparable to similarly-studied parapatric amphibians bearing a specific status. Inferred from their current geographic distribution, the two Italian tree frogs feature distinct ecological niches (<15% of niche overlap), raising questions regarding potential adaptive components contributing to their incipient speciation. However, we found no diagnostic morphological and bioacoustic differences between them. This system illustrates the speciation continuum of Western-Palearctic tree frogs and identifies additional cryptic lineages of similar divergence to be treated as separate species (H. cf. meridionalis). We recommend combined approaches using genomic data as applied here for the future taxonomic assessment of cryptic diversity in alloparapatric radiations of terrestrial vertebrates, especially in controversial taxa. Finally, we formally described the northern Italian tree frogs as a new species, Hyla perrini sp. nov.
urn:lsid:zoobank.org:pub:C446334C-3D94-4820-A851-99BD95084F6D
Introduction
Three decades of molecular biogeography have shed light on unsuspected amounts of genetic diversity among terrestrial vertebrate species that were once considered monotypic. In the Western Palearctic, this diversity was shaped by the combined actions of geological and climatic events (e.g., Mountain chain uplifts, the Messinian Salinity crisis, Quaternary glaciations), which have promoted allopatric diversifications between and within Mediterranean regions (e.g., Iberian, Italian, and Balkan Peninsulas, Northwestern Africa, Anatolia), and isolated off-shore islands (e.g., Crete, Cyprus) (Schmitt, 2007; Dufresnes, 2019). Yet, the resulting lineages often remained remarkably cryptic, sharing similar ecologies and featuring no obvious phenotypic differentiation, be it morphology, coloration or behavior, despite millions of years of independent evolution (Bickford et al., 2007, e.g., Wielstra et al., 2013; Diaz-Rodriguez et al., 2017).
What taxonomic status do such cryptic lineages deserve? This question is a topical controversy among zoologists and one of the major challenges in modern phylogeography (Bickford et al., 2007). Mayr's universal “biological species concept,” referring to pre- or post-zygotic barriers to reproductive isolation (Mayr, 1942), does not necessarily involved measurable morphological changes. Fortunately, secondary contact zones between pairs of taxa represent natural laboratories where cryptic species are put to the test (Hewitt, 2011). Pending opportunity for dispersal (Waters et al., 2013), a hybrid zone mediated by selection against hybrids will be narrower than expected under dispersal alone, which can be measured through analyses of genetic introgression. In many instances however, and notably when species have non-overlapping distribution ranges and do not naturally meet, the amount of genetic divergence (e.g., divergence time) is rather used as a proxy for their level of reproductive isolation. However, this indirect approach has several shortcomings. First, nuclear markers may fail to accurately resolve phylogenetic relationships, because of incomplete lineage sorting (ILS) and/or past hybridization events (e.g., during former interglacial expansions), especially for recent divergences (e.g., Pabijan et al., 2012). Second, the time and mode along which genetic divergence accumulates may greatly differ between vertebrate radiations and should therefore be independently assessed for the groups of interest. For instance, parapatric taxa may stop hybridizing after few million years in some systems (e.g., <3 My in Bufotes, Colliard et al., 2010), but long after in others (e.g., >10 Mya in Salamandrina, Hauswaldt et al., 2011; Mattoccia et al., 2011). Third, molecular clocks are extremely sensible to the calibration used in phylogenies, biasing divergence times by millions of years between equally plausible evolutionary scenarios (e.g., Veith et al., 2016). A standardized approach accounting for these limitations is still missing to harmonize species limits in cryptic polytypic radiations.
This issue is particularly relevant for Western Palearctic amphibians (Shaffer et al., 2015), which form so called “species complexes” that diversified throughout the late Miocene (11.0-5.3 Mya), Pliocene (5.3-2.6 Mya) and early Pleistocene (2.6-1.8 Mya), giving birth to evolutionary lineages of different ages, resulting in unresolved taxonomic situations. In recent years, genetic studies have proposed that some of these taxa be considered new, distinct species (e.g., Hyla felixarabica, Gvoždík et al., 2010; Pelodytes hespericus and P. atlanticus, Diaz-Rodriguez et al., 2017; Bufo spinosus, Arntzen et al., 2013; Lissotriton graecus, L. schmidtleri, L. lantzi and L. kosswigi, Pabijan et al., 2017; Triturus karelinii, T. anatolicus and T. ivanbureschi, Wielstra et al., 2013) or, on the contrary, disproved their potential specific status (e.g., Bufo (Bufotes) variabilis, Dufresnes et al., 2018). Many lineages have yet to receive a similar assessment, upon which their taxonomic fate will depend.
With at least eight taxa distributed around the Mediterranean Basin, the Western Palearctic radiation of hylid tree frogs (Hyla arborea species complex) is an ideal system for the study of cryptic speciation (Stöck et al., 2008b, 2012). Most of these lineages are remarkably similar in appearance, mating calls and ecology, despite their Mio-Pliocene origins and signs of reproductive isolation as revealed by narrow transitions at their parapatric margins (see Discussion). Cryptic taxa were further suspected within the north-African populations of H. meridionalis (Recuero et al., 2007; Stöck et al., 2008b, 2012), as well as across the Apennine Peninsula in the Italian endemic H. intermedia (sensu lato, Canestrelli et al., 2007a,b), pending in-depth characterization.
Here we focus on the latter, H. intermedia s. l., which ranges from Sicily to the southern slopes of the Alps in northern Italy and the Swiss canton of Ticino. This taxon features a typical phylogeographic pattern for Italian amphibians, with distinct mitochondrial clades distributed north and south of the Northern Apennine Mountains (clades N and C/S in Canestrelli et al., 2007b), tentatively split as H. intermedia sensu stricto and Hyla “new taxon 2” by Stöck et al. (2008b, 2012). Both mitotypes are found in sympatry along the Apennine foothills from central Italy (Emilia-Romagna), over a transition that appears no wider than tens of kilometers (Canestrelli et al., 2007b). Their nuclear identity, however, has remained cumbersome. Population genetics using allozyme markers accordingly suggested latitudinal population differentiation, yet with massive allele sharing, indicative of ancestral polymorphism or widespread admixture (Canestrelli et al., 2007a). Similarly, subsequent phylogenies based on slowly-evolving nuclear sequences showed little support for monophyletic clades (Stöck et al., 2008b, 2012), supposedly because of past or present gene flow (Gvoždík et al., 2015) or purifying selection on gene markers (e.g., rag-1, Stöck et al., 2008b). Therefore, their evolutionary and taxonomic situation is still to be determined.
In the present study, we provide several lines of evidence that Italian tree frogs belong to two separate species, and we formally describe the northern one as Hyla perrini sp. nov. (see specific section in the Discussion). We combined population genomics and nuclear phylogenomic inferences to show that both species are monophyletic, deeply-diverged, and only admix across a well-defined hybrid zone restricted to their post-glacial contact in Emilia-Romagna, north-central Italy. We further implement analyses of ecological niche modeling, morphometrics, and bioacoustics to explore phenotypic variation between these two cryptic species.
Methods
DNA collection
A total of 171 tissue samples of tree frogs were obtained from non-invasive buccal swabs (adults) or tail tips (tadpoles). Some were already available from our previous genetic studies (Canestrelli et al., 2007a,b; Stöck et al., 2008b, 2011, 2012; Dufresnes et al., 2011, 2013, 2015b,c, 2016b,c) while others were collected during fieldwork campaigns in Northern Italy and Southern Switzerland in the springs of 2013 and 2018. DNA was extracted using the Biosprint robotic workstation (Qiagen) or the DNeasy Blood and Tissue Kit (Qiagen).
RAD-sequencing genomic libraries
Two separate genomic libraries were prepared using the double digest RAD (ddRAD) protocol detailed in Brelsford et al. (2015). Briefly, genomic DNA (5–15 ng/μL, 30–90 ng in total) was digested using Sbf1 and Mse1 restriction enzymes. Adapters containing PCR primer sites and individual barcodes (4–8 nucleotides) were then ligated to the Sbf1 end of the fragment. Barcoded fragments were purified and amplified using PCR with Illumina primers. The amplified libraries were then size selected to enrich for fragments between 400 and 500 bp using gel electrophoresis (2.5% agarose gel run for 2 h 30 at 80 V) and finally, the libraries were purified once more and sequenced (single-end only) on an Illumina Hi-Seq 2500.
Library 1 was designed to reconstruct the nuclear phylogeny of this radiation. It included 51 samples representative of all Western-Palearctic taxa (plus Hyla (Dryophytes) japonica as an outgroup; Table S1B). Particular focus was given to H. perrini sp. nov. (n = 12, including the holotype and two paratypes) and H. intermedia s. s. (n = 7); 3–6 individuals for every other species (Table S1B). Raw data from one sequencing lane was processed as follows. Raw, single end Illumina sequences were quality checked using FastQC v0.10.1 (Andrews, 2010) and the individuals of the library were demultiplexed by individual sample barcodes with the process_radtags.pl module of STACKS v 1.42 (Catchen et al., 2013). The resulting individual fastq files were mapped on the draft genome assembly of H. arborea from Brelsford et al. (2015) with bowtie2 v 2.3.1 (Langmead and Salzberg, 2012; using –local –highly-sensitive specifications. Resulting sam files were subsequently filtered for mapping quality >20 with samtools v 0.1.19 by Li (2011) and the bam files were used as input for ref_map.pl module of STACKS with default parameters. The STACKS catalog obtained consisted of 181,040 loci merged across all individuals (mean of 67,000 loci/individual with average coverage of 31 reads/locus), which was then whitelisted down to 621 loci (biallelic per individual) present across at least 50 individuals. The populations module of STACKS was then run with –phylip_var_all to obtain individual-based alignment (using a population map comprising all 51 individuals as separate “populations”) of 42,834 bp. This alignment was then translated into nexus format with Geneious v8.1.9 (https://www.geneious.com/). Finally, in order to compute genome-wide diversity and divergence between H. perrini sp. nov. and H. intermedia s. s., we outputted a second alignment containing all tags sequenced in both species (2.9 Mb from 29,294 loci).
Library 2 was designed for population genomic analyses of the hybrid zone between the two Italian lineages and included 120 samples from 12 populations sampled across the Apennine Peninsula (Table S1C). Raw, single end illumina sequences from two lanes were again quality checked using FastQC v0.10.1 and demultiplexed using the process_radtags module of STACKS. RAD loci were constructed and SNPs were called using STACKS. We first ran initial tests of parameter values for the core STACKS parameters [Ustacks: -M (2-4), -m (3-6) and cstacks: -n (2-4)] to test for any major effects on the number of loci, average coverage, and number of heterozygous genotype calls (a proxy for genotype call accuracy). The default values (-M 2, -m 3 and -n 2) provided a good balance between data quality and quantity according to the criteria set out in Paris et al. (2017). Modules ustacks, cstacks, sstacks, and populations were then run separately using these parameter values. The STACKS catalog contained 670,396 loci; these were filtered using the populations module and loci were retained if they had a minor allele frequency filter (–min_maf) of >0.05, a maximum observed heterozygosity filter (–max_obs_het) < 0.75 (to accounts for cases of “over merging” of paralogous loci), if they were present in at least 80% of individuals from all 12 populations and had a coverage of at least 6 reads per locus. The resulting dataset contained 2,431 SNPs from 1,639 RAD tags, with an average coverage of 30.45 reads (SD = 11.77).
Mitochondrial data
For phylogenetic purposes, we harvested published sequences for the mitochondrial genes cytochrome b (cyt-b, 840 bp) and cytochrome c oxidase subunit I (COI, 766 bp) in 33 individuals representative of the Western Palearctic diversity of tree frogs (2–6 individuals per taxa), as well as 3 outgroups (H. japonica, H. cinerea, H. squirella). Details on their origin can be found in Table S1. Importantly, we only selected individuals with sequences available for both genes that could thus be concatenated for analysis (total alignment length = 1,606 bp).
Furthermore, distribution data of the two Italian mitochondrial lineages (H. perrini sp. nov. and H. intermedia s. s.) were gathered from previous studies (Canestrelli et al., 2007a,b; Verardi et al., 2009; Stöck et al., 2012; Dufresnes et al., 2015c), totaling 45 populations.
Phylogenetic analyses
For each mitochondrial (1,606 bp from two partitioned genes) and genomic (42,834 bp from 621 RAD tags concatenated in a supermatrix) alignments separately, we independently performed time-calibrated Bayesian phylogenetic reconstructions in BEAST 2.4.8 (module StarBEAST; Bouckaert et al., 2014), with a relaxed lognormal molecular clock and a birth-death tree prior. Site substitution models were GTR+G (nuclear), TN93+G+I (COI) and HKY+G (cyt-b), selected with bModelTest (Bouckaert and Drummond, 2017). We used three calibration points with normally distributed priors. The first two were inferred from multiple fossil calibrations, independently applied by Smith et al. (2005) and Li et al. (2015): the split between the H. japonica group and Western Palearctic Hyla (36 Mya ± 6 My), and the time since the most recent common ancestor of Western Palearctic tree frogs (20 Mya ± 3 My). In order to stabilize the clock, we used a third, younger calibration: the Messinian divergence of the Tyrrhenian endemic H. sarda (set to 5.33 Mya ± 2 My), which is the most plausible origin for this insular species, based on phylogeographic (Stöck et al., 2008b, 2012; Bisconti et al., 2011) and fossil evidence (Kotsakis, 1980). The chains were run for 50 million (nuclear) and 250 million iterations (mtDNA), sampling every 5,000 (nuclear) and 25,000 iterations (mtDNA), and checked for stationarity and effective sample sizes (ESS) of parameters >200 with Tracer 1.5. The first 10% of trees were discarded as burnin, and the remaining 9,000 were analyzed by TreeAnnotator 2.4.8, and visualized by DensiTree 2.2.6 (Bouckaert and Heled, 2014).
Furthermore, we explored the relationships between Western-Palearctic hylids with a Principal Component Analysis (PCA) on the nuclear genomic dataset (haplotype VCF file analyzed in adegenet R package), discarding the early-diverged H. japonica and H. meridionalis. Finally, we estimated nucleotide diversity (Π) for Hyla perrini sp. nov. and H. intermedia, as well as the percentage of fixed differences between them.
Population structure analyses
In order to characterize the nuclear population structure between Hyla perrini sp. nov. and H. intermedia s. s. and within the contact zone, we performed STRUCTURE (Pritchard et al., 2000) analyses using the SNP dataset from library 2. STRUCTURE was run from a k of 1–12, with an MCMC chain length of 10,000,000 (burn-in 1,000,000) and three iterations were performed for each value of k. STRUCTURE outputs were fed to STRUCTURE HARVESTER (Earl and vonHoldt, 2012) to identify the most appropriate value of k using the Evanno method (Evanno et al., 2005). We further computed pairwise genetic distance (Fst) between pure populations (loc. 1 and 2 for H. perrini sp. nov.; loc. 12 for H. intermedia s. s.), as well as their respective diversity (observed and expected heterozygosity Ho and He) with VCFtools (http://vcftools.sourceforge.net/).
Additionally, we analyzed the modality of the Hyla perrini sp. nov./H. intermedia s. s. transition trough cline analyses, which are powerful tools to understand the dynamic of hybrid zones in the geographic space, through the estimation of cline width (w) and center (c). To this purpose, we fitted sigmoid clines to mitochondrial frequency data (n = 10 populations, totaling 70 individuals) and nuclear hybrid index (STRUCTURE admixture coefficient; n = 12 populations, totaling 120 individuals) along a transect running from Southern Switzerland (pure H. perrini sp. nov.) to Abruzzo, Italy (pure H. intermedia s. s.), using the R package hzar (Derryberry et al., 2014). We performed model selection between two-parameter clines (w and c) and models with exponential tails (up to eight parameters) and kept the model with the best AIC score.
Niche modeling analyses
To examine potential ecological differences between both Hyla species, as well as to obtain an approximate age for the contact zone, we modeled their present and past (Last Glacial Maximum) distributions with MaxEnt (v. 3.3.3k; Phillips et al., 2006). We combined 84 localities of H. intermedia and 154 localities of Hyla perrini sp. nov. (Figure S1), comprising our own and previously published records, as well as localities from the Global Biodiversity Information Facility database information (http://www.gbif.org/). Duplicated localities were removed by ENMTools v. 1.3 (Warren et al., 2010).
Nineteen bioclimatic layers were extracted from the WorldClim 1.4 database (http://www.worldclim.org). An additional nine uncorrelated variables were included: Altitude (WorldClim 1.4), Global Aridity and Potential Evapo-Transpiration (aridity index; http://www.cgiar-csi.org/data/global-aridity-and-pet-database), GlobCover 2009 (Global Land Cover Map; due.esrin.esa.int/globcover/), spatial heterogeneity of global habitat (EarthEnv, http://www.earthenv.org/texture.html), percent tree cover (Github, https://github.com/globalmaps/gm_ve_v1), as well as four topographic layers (aspect, exposition, slope, and terrain roughness index) calculated in QGIS v. 2.18.1 (http://www.qgis.org/).
To eliminate predictor collinearity prior to generating the models, we calculated Pearsons's correlation coefficients for all pairs of bioclimatic variables using the ENMTools. We excluded the variable of a correlated pair with |r| > 0.8 that we considered to be the less biologically important of the pair, based on known preferences of Hyla species. The resulting dataset hence contained 10 uncorrelated bioclimatic variables, plus the nine additional variables mentionned above. We used a jackknife analysis for estimating their relative contributions to the MaxEnt model (Table S2) and applied a mask that extends from 35° to 48° N and 5° to 20° E.
These 19 variables were implemented in the final models for both species (30 arc-seconds resolution). We used 70% of the occurrence localities as training data, and the remaining 30% were kept for testing the resulting models. Model performance was measured using the Area Under the Curve (AUC) derived from the Receiver Operating Characteristic (ROC) plots. The plots represent the model's ability to discriminate species locations from pseudo-absences by plotting sensitivity against 1 - specificity. AUC values range from 0.5 to 1.0, with 0.5 indicating no greater fit than expected by chance and 1.0 indicating a perfect model fit. Models with test AUC values above 0.75 are considered useful and above 0.90 very good (Swets, 1988; Elith, 2000). To properly parameterize the model, we evaluated the performance of various combinations of ten regularization multipliers (from 0.5 to 5, in increments of 0.5). The best-fit models were parameterized with a regularization multiplier of 0.5 for H. intermedia and 1.5 for H. perrini sp. nov. We used the default auto feature setting within the MaxEnt software to generate the model. The niche overlap between species was then estimated using Schoener's D distance in ENMTools (Schoener, 1968), which is the best suited to compute niche overlaps from potential distributions derived from species distribution models (Rödder and Engler, 2011).
The ten selected uncorrelated bioclimatic layers (2.5 min resolution, extracted from WorldClim 1.4) were further used to predict the Last Glacial Maximum (LGM) distributions. Two general atmospheric circulation models were used to generate LGM climate scenarios: the Community Climate System Model (CCSM; http://www.cesm.ucar.edu/models/ccsm4.0/ccsm/) and the Model for Interdisciplinary Research on Climate (MIROC; Watanabe et al., 2011). Models were generated by MaxEnt using the above described settings.
Biometric analyses
Morphometrics
We tested whether the newly described taxon H. perrini sp. nov. exhibits significant differences in morphology and breeding calls compared to its sister species H. intermedia s. s. and its other parapatric relative H. arborea. In spring 2018, male tree frogs from two Hyla perrini sp. nov. (n = 24, including the type locality), four H. intermedia s. s. (n = 15) and two H. arborea populations (n = 22), were captured, photographed and measured for nine variables commonly used in amphibian taxonomy: snout-vent length (SVL), hind leg length (HL), tarsus + tibia length (TTL), head width (HW), eye diameter (ED), tympanum diameter (TD), inter-eye distance (EE), inter-nostril distance (NN) and eye-nostril distance (EN). We used a 1 mm accurate ruler with a blocking end for the first three variables, and a digital caliper (0.1 mm accuracy) for the remaining six variables. For the sake of consistency, all individuals were measured by the same person (CD), except for six of the H. intermedia s. s. individuals (DC). Measures were corrected by the relative size of individuals (i.e., divided by SVL) and compared between the three species by means of Principal Component Analyses (PCA) and Multivariate Analysis of Variance (MANOVA), which allowed for the inclusion of between-population intraspecific variation as a cofactor. Variables showing significant difference between species were further compared by non-parametric Tukey tests.
Bioacoustics
Mating calls were recorded directly at the breeding sites mentionned above using a portable digital recorder (EVISTR L57, setup to 48,000 Hz). A total of 11 different males of Hyla perrini sp. nov., 10 of H. intermedia and 10 of H. arborea could be recorded. We pre-edited the calling sequences of each individual with the software Audacity (http://audacity.fr/), and used the R package Seewave (Sueur et al., 2008) for downstream processing and data analyses.
The calls of most European hylids, including Italian taxa, consist of series of short monotonous notes, each made of partially-fused pulses (Rosso et al., 2006). For each male, we visualized and analyzed a set of ten consecutive notes in the middle of a call series, using the built-in functions of Seewave (spectro, oscillo, a.timer, and fpeaks). We averaged note duration (ND), duration of pause between two consecutive notes (PD), pulse rate (PUL) and measured the frequency peaks of the two main energy bursts (fundamental frequency Ff at ~1 kHz and dominant frequency Fd, at ~2 kHz). The actual band-width of the call is wider but cannot be reliably analyzed because of background noise and/or because the signal was too weak in some recordings. Temperature, known to affect call properties (Köhler et al., 2017), was recorded. Other important abiotic variables such as time of the day (early evening: 21−23 h), moment in the breeding season (main peak of activity: mid-April 2018 and early May 2018 for south-alpine and north-alpine populations, respectively), and motivation context (chorus of >15 males) were all similar and should have limited effect on our comparisons. In the same ways as for morphometric analyses, we explored and tested species differences with PCA and MANOVA, considering populations and temperature as co-variables for the latter.
Collection of type specimens
Two males and two females were collected in April and May 2018 from the type locality of H. perrini sp. nov, in the Swiss canton of Ticino (see details in Discussion). A third male, found dead but in a good state of preservation, was also included in the type series. All five were measured and DNA was collected with buccal swabs; the holotype and two paratypes were included in the phylogenomic dataset (Table S1B). Specimens were subsequently anesthetized and euthanized by overdose of MS-222 0.015% (Sandoz) buffered in 0.03% sodium bicarbonate, and inventoried in the herpetological collection of the Cantonal Zoological Museum of Lausanne (MZL42340-42344). The type series was later examined post-mortem, including photography and 14 standard morphometric measurements with a digital caliper (0.1 mm accuracy), made by a single one of us (GM).
Data accessibility
The raw sequence reads from both RAD libraries are available on the NCBI SRA archive under BioProject PRJNA485248. Phylogenomic, population genomic, ecological niche modeling and biometric data files used in the analyses are provided in Data Sheet S1.
Results
Phylogeny of western-palearctic tree frogs
Time-calibrated phylogenies of mitochondrial (1,606 bp from two genes) and nuclear sequences (42,834 bp from 621 RAD tags) congruently recovered all known Hyla taxa from the Western Palearctic (Figures 1, 2). The nuclear tree is fully resolved (Figure 2), suggesting no signs of past admixture or incomplete lineage sorting, while the mtDNA tree differs slightly at a few unresolved nodes (notably H. sarda) and larger uncertainty in the molecular dating (Figure 1). Both trees fully support the split of H. intermedia s. s. and Hyla perrini sp. nov. as two monophyletic clades, with divergence pointing to the upper Pliocene (nuDNA: 3.5 Mya, 95% HPD: 5.0-2.2; mtDNA: 2.5, 95% HPD: 4.2-1.0). The first two axes of the PCA (based on individual genotypes) recovered most known species, including H. intermedia and H. perrini sp. nov. which form adjacent yet distinct clusters (Figure S2). These two Italian taxa featured 9.8% of fixed difference at COI and 8.7% at cyt-b (9.2% for the concatenated alignment). At the nuclear level, they differ by 0.3%, based on 2.9 Mb. Nucleotide diversity (Π) was estimated at 0.0061 for H. intermedia s. s. and 0.0069 for H. perrini sp. nov.
Figure 1
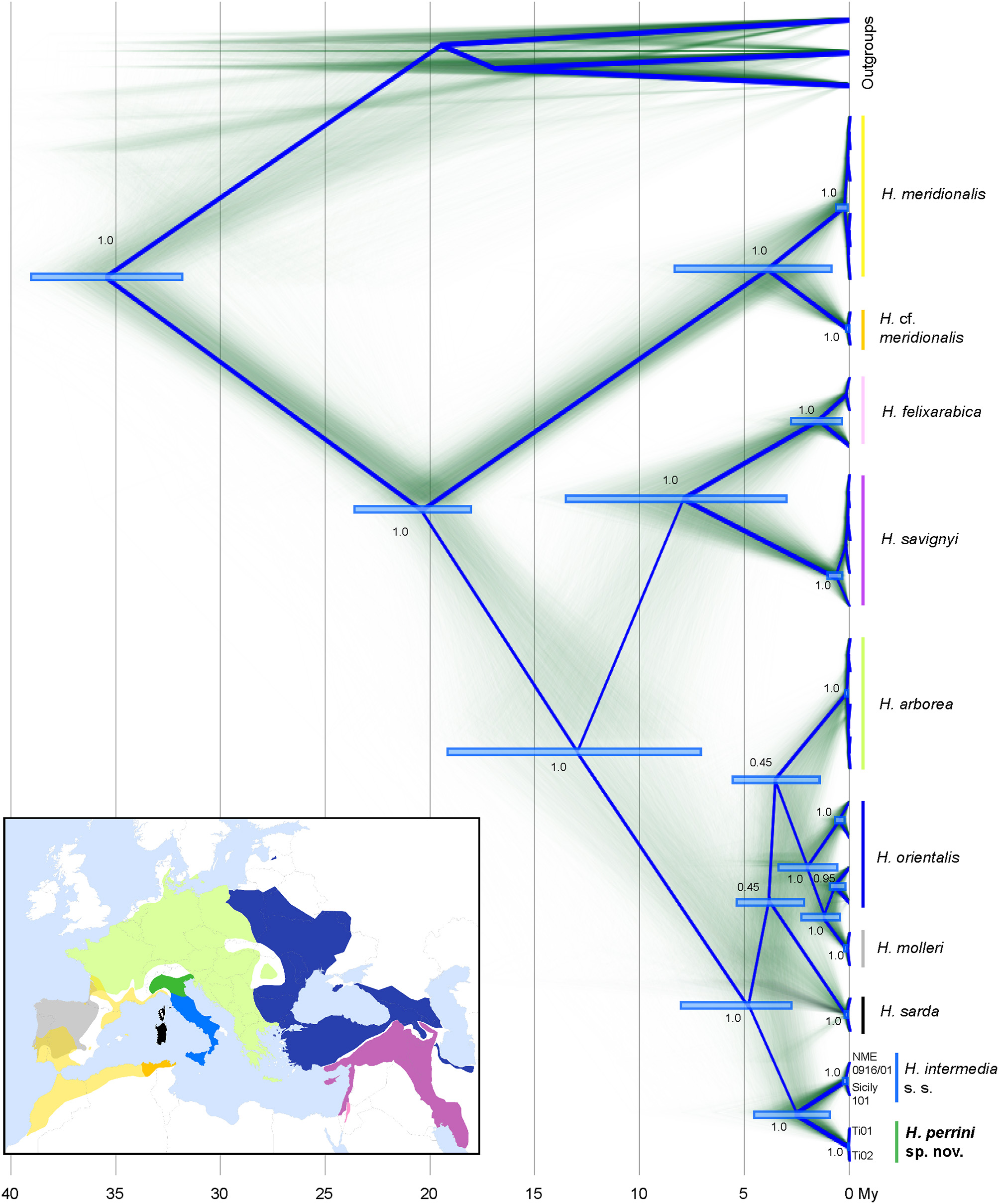
Time-calibrated phylogeny of mitochondrial sequences from cyt-b (840 bp) and COI (766 bp). Clock rates were nearly identical between markers (2.5e−2 substitutions/site/My). Diffuse lines show branches of each of the sampled trees, summarized by the root canal (blue framework). For major nodes, the Bayesian posterior probabilities and confidence intervals of divergence times are indicated. Species distributions use the color codes shown on the tree. The map was built in QGIS 2.18 using map layers from ESRI and the IUCN red list (adapted by Dufresnes, 2019). Hyla japonica, H. cinerea and H. squirella were used as outgroups (Table S1A).
Figure 2
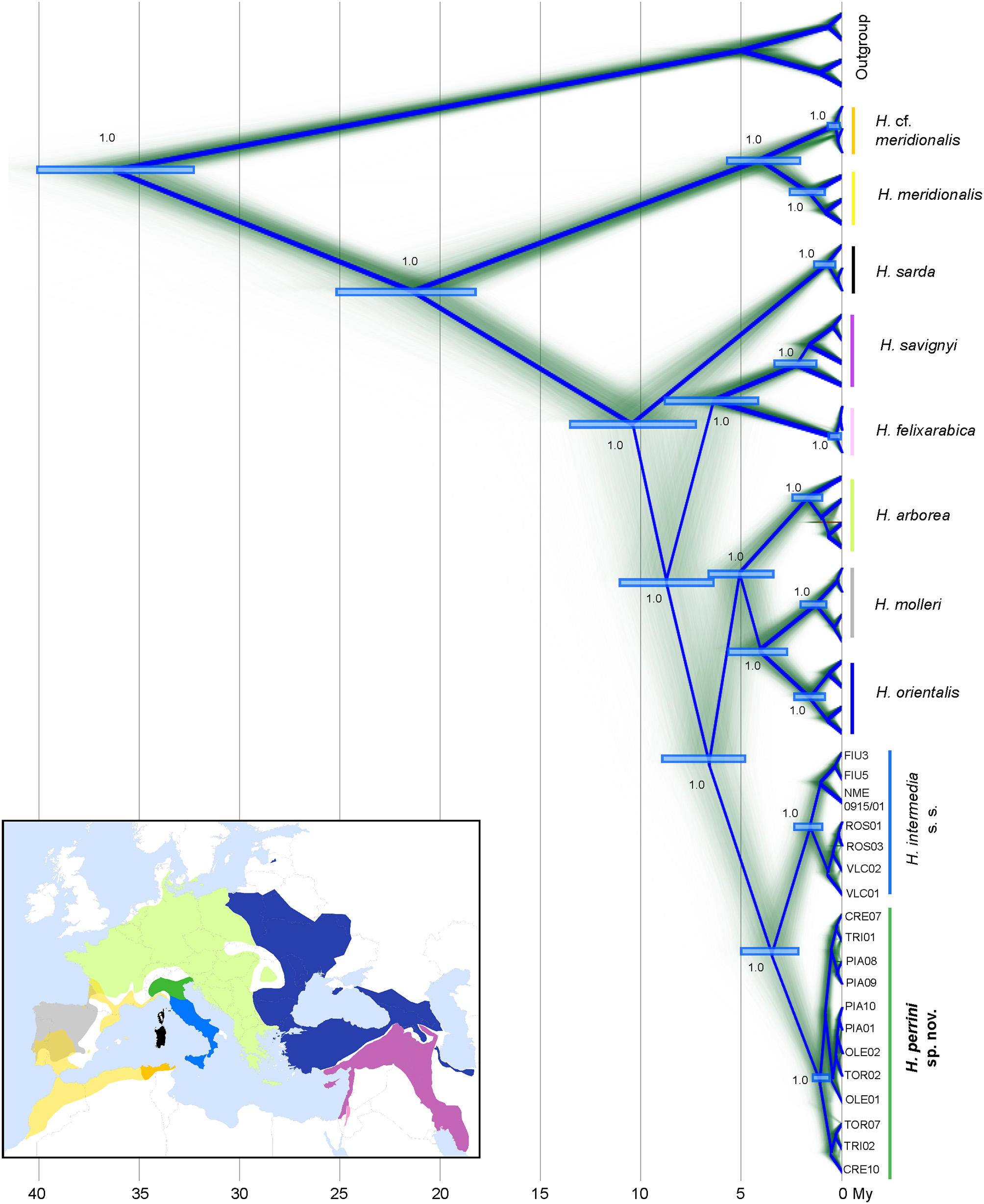
Time-calibrated phylogeny of nuclear loci (42,834 bp from 621 RAD tag sequences). Clock rate averaged 3.8e−4 substitutions/site/My. Diffuse lines show branches of the sampled trees, summarized by the root canal (blue framework). For major nodes, the Bayesian posterior probabilities and confidence intervals of divergence times are indicated. Species distributions use the color codes shown on the tree. Species distributions use the color codes shown on the tree. The map was built in QGIS 2.18 using map layers from ESRI and the IUCN red list (adapted by Dufresnes, 2019). The polytypic H. japonica was used as outgroup (Table S1B).
Hybrid zone between apennine tree frogs
MtDNA data in across the distribution clearly delineates the ranges of Hyla perrini sp. nov. (Northern Italy and Southern Switzerland) and H. intermedia s. s. (Central-Southern Italy and Sicily) (Figure 3), which are mostly kept apart by the Apennines, but locally meet in southeastern Emilia-Romagna. Population genomics (RAD data) yielded a similar nuclear signal (based on 2,431 SNPs from 1,639 RAD tags, Figure 4): STRUCTURE analyses with k = 2 (the most likely number of k) perfectly recover the two gene pools, with a spectrum of intermediate coefficients for frogs sampled at the transition, indicative of genetic admixture (loc. 4–11 in Figure 4).
Figure 3
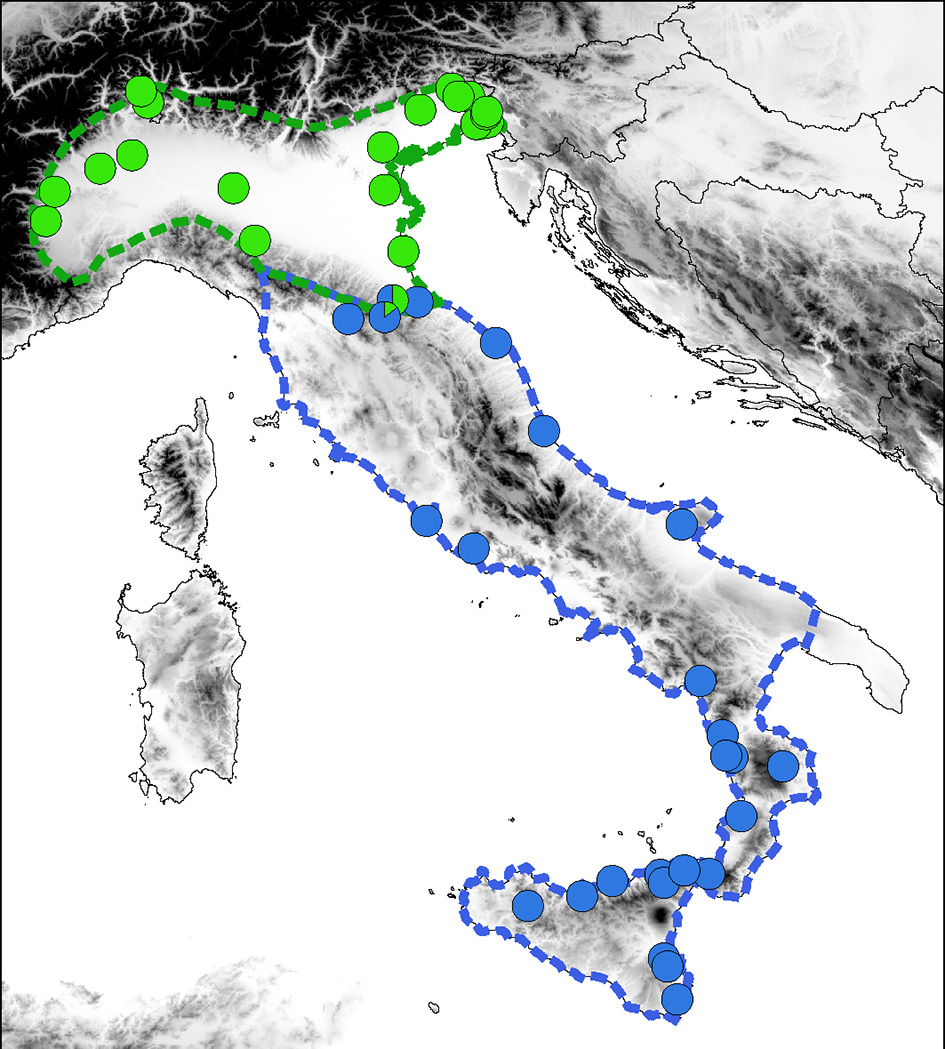
Distribution of the mitochondrial lineages of H. perrini sp. nov. (green) and H. intermedia s. s. (blue) across their natural ranges, based on records from previous studies. Dashed lines delineate range margins. The map was built in QGIS 2.18 using map layers from ESRI.
Figure 4
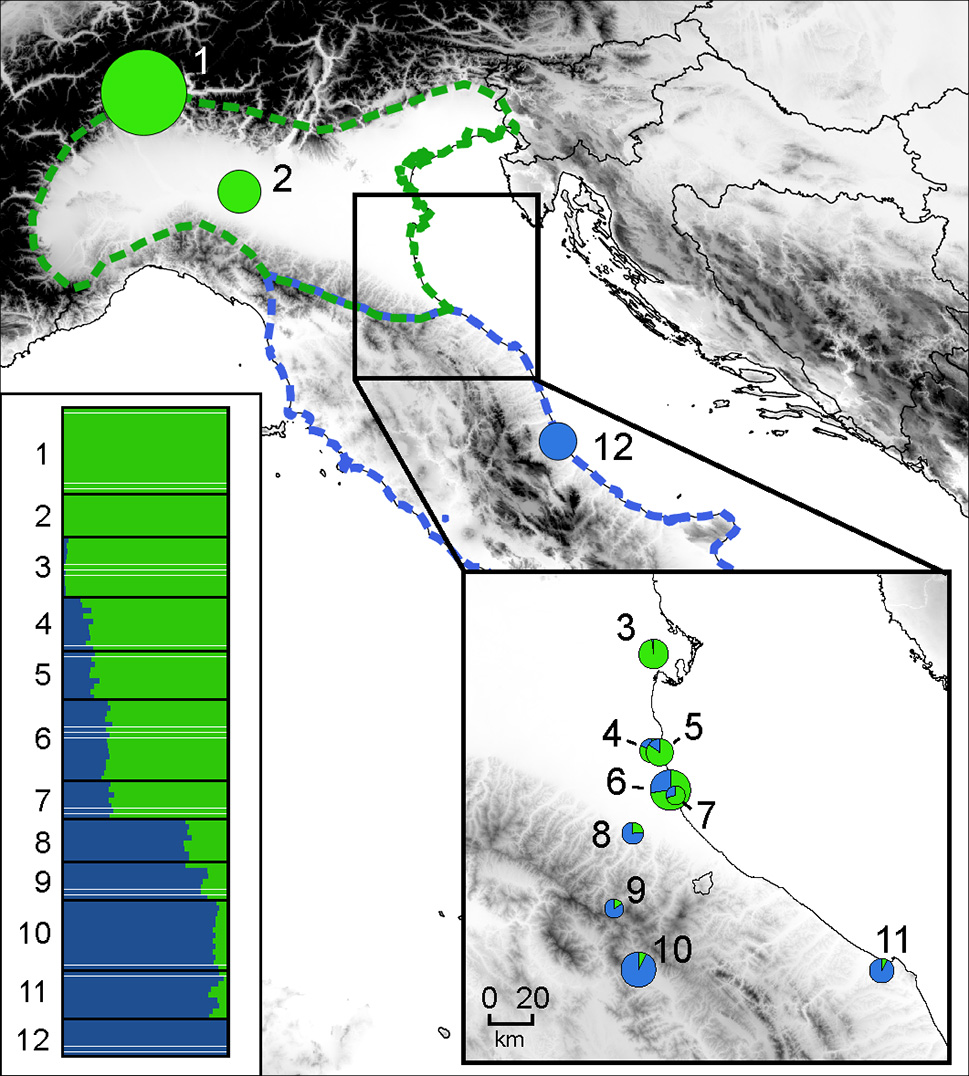
Genetic structure across the H. perrini sp. nov. / H. intermedia s. s. transect, inferred from Bayesian clustering of individuals (n = 120 from 12 populations) in k = 2 groups (STRUCTURE), based on 2,431 loci. Horizontal barplots show individuals assignments to the H. perrini sp. nov. (green) or H. intermedia s. s. (blue) gene pools; pie charts feature the population averages, with size proportional to sample size. The map was built in QGIS 2.18 using map layers from ESRI.
Accordingly, steep clines of less than 100 km (Figure 5) were inferred for the nuclear hybrid index (w = 96 km; 95% CI: 64–153) and the frequencies of mitotypes (w = 41 km; 95% CI 20–92). Both clines remarkably coincided, with centers at the northeastern foothills of the Apennines (between population 7 and 8, Figure 5), shifted by only 2 km (for nuclear: c = 378 km along north-south transect; 95% CI: 365–394; for mtDNA: c = 380 km: 95% CI: 365–392). No trace of genetic introgression was found apart from the contact zone.
Figure 5
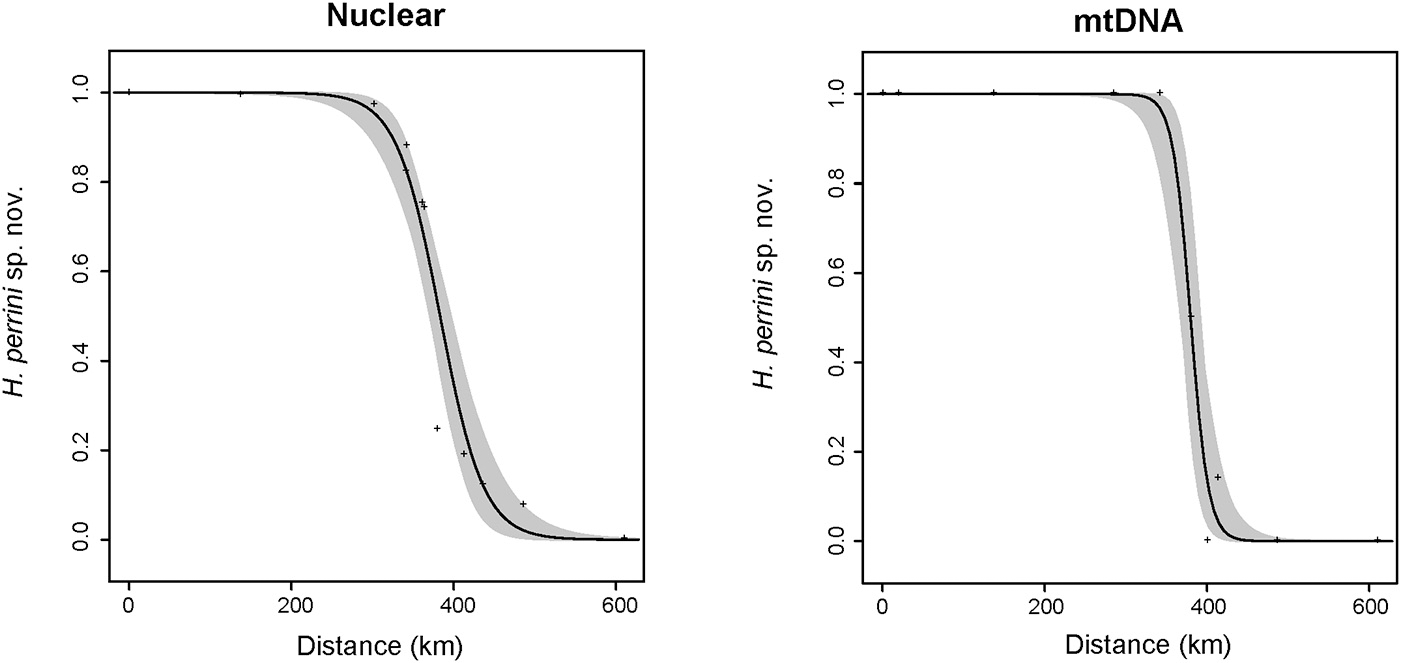
Geographic clines fitted to nuclear hybrid index data (STRUCTURE admixture coefficient, left panel) and mitochondrial frequency data (right panel) along the H. perrini sp. nov./H. intermedia s. s. transect. The best selected models featured only two parameters (width and center), while parental frequencies could be set to 0 and 1. Nuclear cline: width w = 96 km and center c = 378 km; mitochondrial cline: w = 40 km and c = 380 km.
Pure populations of H. perrini sp. nov. featured nearly identical observed (Ho) and expected (He) heterozygosity (loc. 1, the type locality: Ho and He = 0.18; loc. 2: Ho and He = 0.17). Similar values were obtained for H. intermedia s. s. (Ho = 0.16, He = 0.15). Pairwise genetic distance between the two species was high (loc. 1–12, Fst = 0.49; loc. 2–12, Fst = 0.46), compared to between the two H. perrini sp. nov. populations (loc. 1-2, Fst = 0.09).
Ecological niche modeling
The present time MaxEnt models for H. intermedia and Hyla perrini sp. nov. received robust evaluation metrics (AUCtest = 0.899 ± 0.023 and 0.939 ± 0.017 respectively) and showed significance for the binomial omission test, indicating a good performance of the model. The predicted niches under the current climate conditions are shown in Figure 6, and the contribution of variables shown in Table S2. These remarkably differ between the two species (D = 0.092), and only overlap on the northern slopes of the Apennine Mountains.
Figure 6
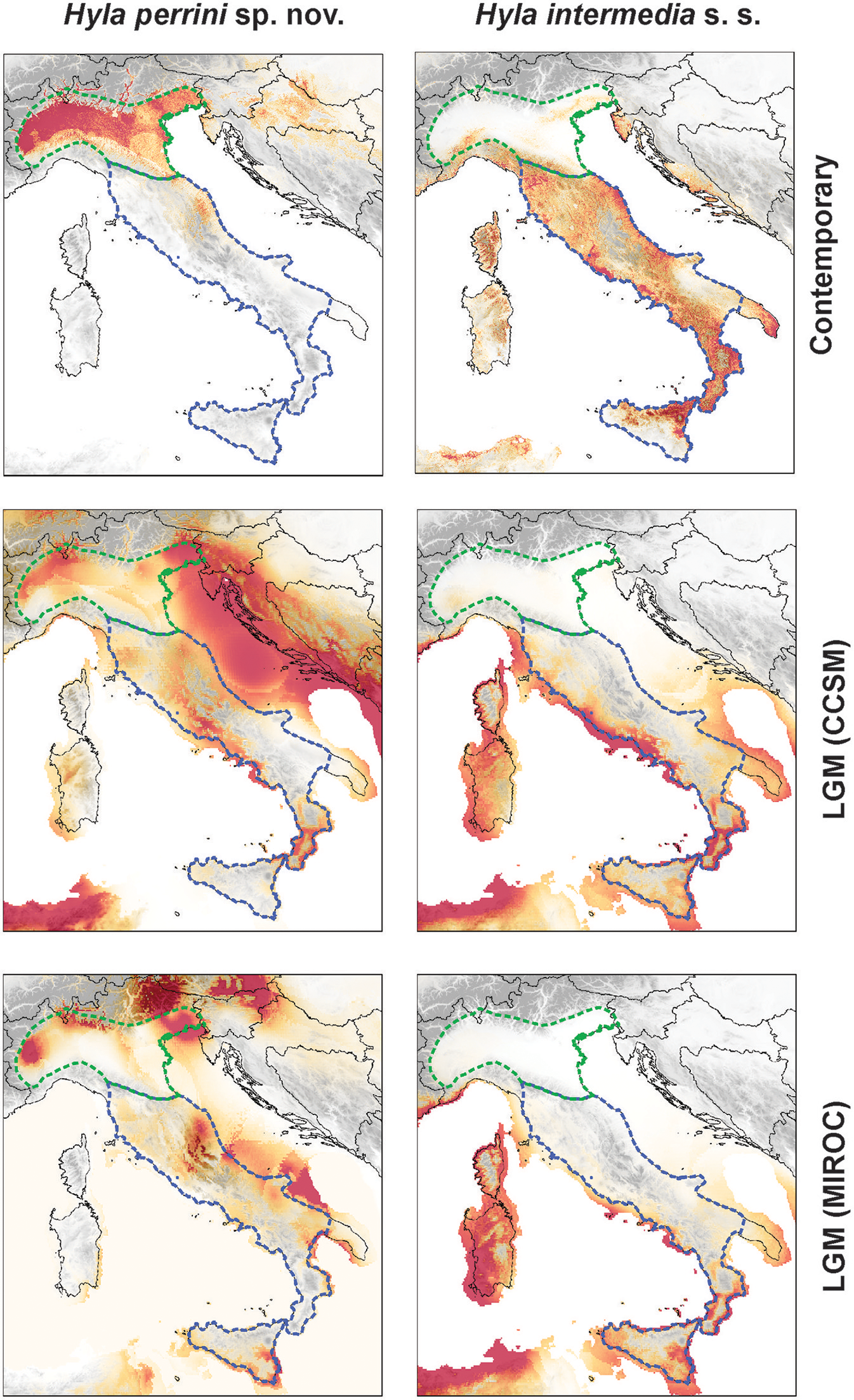
Projection of the ecological niches for H. perrini sp. nov. (Left) and H. intermedia s. s. (Right) under current (top) and Last Glacial Maximum (LGM; 21'000 years ago) environmental conditions from the Community Climate System Model (CCSM, middle) and the Model for Interdisciplinary Research on Climate (MIROC, bottom). Parameter contributions are available in Table S2. The maps were built in QGIS 2.18 using map layers from ESRI.
The CCSM and MIROC models both suggested that the LGM environmental conditions were most suitable along the Tyrrhenian and southern Adriatic coasts for H. intermedia s. s., while the inlands seem unfit (Figure 6). In contrast, within current range limits, the predicted distribution of glacial Hyla perrini sp. nov. populations is essentially restricted along the Alpine edge of Po Plain (Figure 6). Additional potential areas highlighted by the models (especially the CCSM), like the Tyrrhenian islands and the Balkan coast, are irrelevant given that these are inhabited by other Hyla species (H. sarda and H. arborea). The current area of contact between the two species does not seem to have hosted suitable LGM conditions.
Biometrics
Morphometric variation
Analyses of nine variables (corrected by SVL) showed little morphological differentiation between Hyla perrini sp. nov., H. intermedia s. s., and H. arborea (Figure 7A, Figure S3). Multivariate analyses of variance (MANOVA), accounting for intra-population variation, nevertheless highlighted a significant species effect for five variables after Bonferroni correction: HW, TD, EN, NN, and HL (Figure S3). Among them, Hyla perrini sp. nov. differs from H. intermedia s. s. by a smaller head (HW) and shorter hind legs (HL) (Tukey tests). Both Italian taxa differ from H. arborea by a smaller tympanum (TD) and a shorter eye-nostril distance (EN) (Figure S3) (Tukey tests).
Figure 7
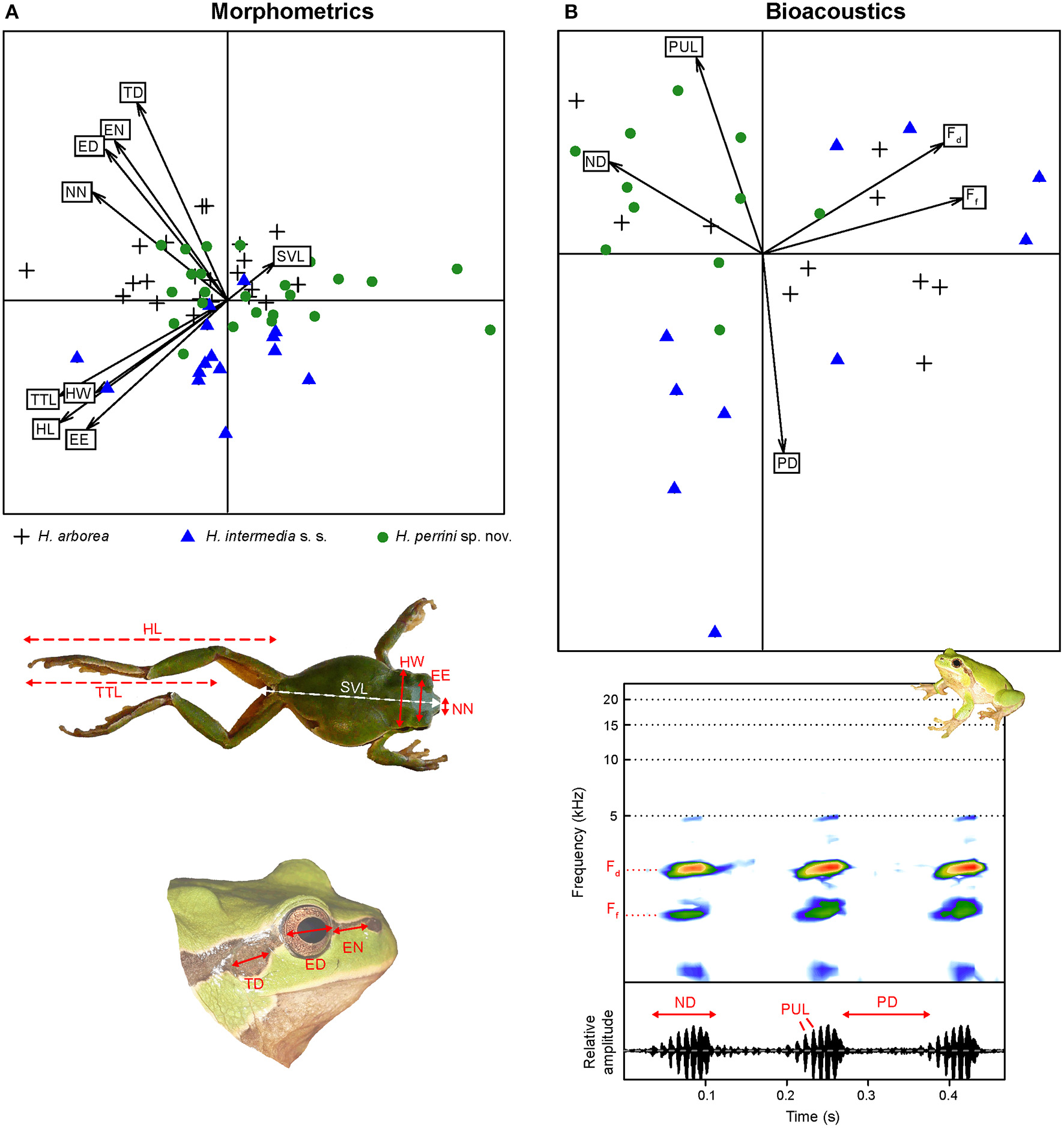
Principal Component Analyses (PCA) on biometric variables of H. perrini sp. nov. (green circles), H. intermedia s. s. (blue triangles) and their close relative H. arborea (black crosses). (A) PCA on morphometric data. The photos illustrate the nine measurements included (see section Methods). (B) PCA on bioacoustic data. The lower graphs illustrate three notes of a call from H. perrini sp. nov., showing the spectral of temporal features included. Note that the temperature was lower (12°C) for the six H. intermedia s. s. with a longer pause duration and lower pulse rate (negative values on axis 2), while all other individuals were recorded at >15°C.
Bioacoustic analyses
Call structure and parameters were globally similar between species (Figure 7B). Among the five bioacoustic variables measured, two showed a significant species effect (Ff and PD), while accounting for populations and temperature as co-variables (which were both significant). Species-by-species comparisons failed to find a significant pair for Ff; pause duration (PD) was longer in H. intermedia s. s. than Hyla perrini sp. nov. (Figure S4). This is illustrated by the second axis of the PCA (Figure 7B), although the six males driving this axis were all recorded on a colder night (11°C) compared to the rest of the samples (>15°C).
Discussion
Cryptic speciation in apennine tree frogs
Two lines of evidence indicate that the Italian tree frog Hyla intermedia s. l. consists of distinct evolutionary entities, distributed across the southern and the northern parts of the Northern Apennine Peninsula, hereby named Hyla intermedia s. s. and Hyla perrini sp. nov., respectively (see its description below). First, the mitochondrial clades previously identified (Canestrelli et al., 2007a,b; Stöck et al., 2008b, 2012) are fully supported by our nuclear loci (Figure 2). Both lineages are monophyletic and diverged at Pliocene times (Figures 1, 2). Second, population genomic analyses unambiguously distinguished the two gene pools across the Italian Peninsula, and only detected admixture over less than 100 km along a well-defined corridor between the northeastern Apennines and the Adriatic coast (Figures 3–5). These results imply that the weak nuclear identity reported by previous studies (Canestrelli et al., 2007b; Stöck et al., 2008b, 2012; Gvoždík et al., 2015) stemmed from low marker resolution and persistent ancestral polymorphism, rather than frequent gene flow and widespread introgression.
Italian Hyla most likely differentiated due to the continued uplift of the Apennines since the lower Miocene, some 20 Mya (Picotti and Pazzaglia, 2008). Moreover, what is now the Po Plain was flooded as part of the Apennine foredeep after the Messinian (Pellen et al., 2017), which may have contributed to the initial stages of divergence. Thermophilic species like tree frogs survived the ensuing Pleistocene glaciations in isolated refugia. Phylogeographic analyses identified a single one for Hyla perrini sp. nov. (northern Italy) but at least two for H. intermedia s. s. (Sicily/Calabria and Central Italy; Canestrelli et al., 2007b). According to our ecological niche modeling, these were probably nested in subalpine valleys (Hyla perrini sp. nov.) and along the paleo-coasts (H. intermedia s. s.; Figure 5). Their secondary contact in north-central Italy is thus mostly likely post-glacial, following demographic expansions after the LGM (Canestrelli et al., 2007b). Tree frogs are a text-book example of the phylogeography of peninsular Italy, where the Apennines triggered various north/south divergences in many widespread amphibians, eventually resulting in different species/subspecies (e.g., Pelophylax lessonae / bergeri, Canestrelli and Nascetti, 2008; Bombina variegata / pachypus; Canestrelli et al., 2006b; Salamandrina perspicillata/terdigitata, Canestrelli et al., 2006a; Triturus carnifex, Canestrelli D Salvi et al., 2012; Lissotriton vulgaris meridionalis, Maura et al., 2014; Rana dalmatina, Canestrelli et al., 2013).
Our complementary genetic approaches suggest that these two taxa are worthy of a specific status, in light of the current taxonomic divisions of other European amphibians. First, their deep, congruent mtDNA and nuclear divergence (averages: 2.5 and 3.5 Mya) is comparable to or older than that of several already established species, such as the Iberian and Oriental tree frogs, H. molleri and H. orientalis (~3 Mya; Stöck et al., 2012, this study); the Parsley frogs Pelodytes hespericus, P. atlanticus and P. punctatus (2–3 Mya; Diaz-Rodriguez et al., 2015, 2017); the green toads Bufo (Bufotes) viridis, B. siculus and B. balearicus (2–3 Mya; Stöck et al., 2008a; Dufresnes et al., 2014); the pool frogs Pelophylax lessonae and P. bergeri (1–2 Mya, Canestrelli and Nascetti, 2008), the Cyprus water frog P. cypriensis (2–3 My, Lymberakis et al., 2007, but 5.3 Mya in Plötner et al., 2012), the Caucasian toad Bufo verrucossisimus (1–2 Mya; Recuero et al., 2012); it is barely younger than many Messinian taxa, e.g., H. arborea (5–6 Mya, Stöck et al., 2012, this study), Middle Eastern Triturus newts (Wielstra and Arntzen, 2011). Note that molecular dating is inherently sensitive to the marker(s) and calibration(s) used (e.g., Veith et al., 2016) and should obviously be interpreted cautiously. Here the mitochondrial phylogeny is not fully resolved, particularly for H. sarda (one of our calibration points); hence the nuclear estimate is probably more trustworthy than the mitochondrial one, which also stands out from their respective confidence intervals.
Second, the restricted introgression along a well-defined contact zone and clear-cut clines argue for limited gene flow between the two gene pools. While the Apennine Mountains kept the two lineages apart over most of the range, the area of contact features no obvious barriers to dispersal. Under a model of neutral diffusion (Barton and Gale, 1993; see Dufresnes et al., 2015a), and assuming a dispersal distance σ of 1.5 km per year (Vos et al., 2000), the cline width w measured from our nuclear (w = 96 km) and mitochondrial data (w = 41 km) would be exceeded in about 650 and 120 generations since first contact, respectively. Accounting for a generation time of 1–2 years, these timing estimates are shorter than the presumed post-glacial contact, suggesting partial reproductive barriers, and that the hybrid zone has reached equilibrium. Selection coefficients s for an averaged nuclear gene, estimated from w and σ (w ∞ σ/√s; Barton and Gale, 1993) were relatively weak, i.e., s = 0.002 under intrinsic selection (heterozygote disadvantage) and s = 0.001 under extrinsic selection (ecological disadvantage). These modest estimates reflect the fact that most genes probably face little or no selection against hybrids, and so that reproductive isolation may involve only a few genes (that have lost linkage with neutral ones) since the post-glacial contact. Male-biased dispersal, expected in polygynous, lek-breeding animals such as tree frogs (Lampert et al., 2003; and documented in European Hyla, Vos et al., 2000), could account for the wider nuclear transition compared to the mitochondrial one. Because both clines coincide, asymmetric fitness between crosses appears unlikely. Analyses of sex-linked vs. autosomal markers may shed some light on the nature of potential hybrid incompatibilities with respect to sex-biased dispersal (Sciuchetti et al., 2018). In parallel, estimation of selection at the locus level should inform on the genomic architecture of speciation and identify the genes/genomic regions involved.
This neat transition falls within the range of those reported between other interspecific lineages that still hybridize at their parapatric margins, i.e., Bufo (Bufotes) viridis and B. balearicus (50–100 km, Dufresnes et al., 2014), Bufo bufo and B. spinosus (50–100 km, Arntzen et al., 2018), Triturus ivanbureschi and T. anatolicus (<50 km, but with hundreds of kilometers of introgression due to hybrid zone movement, Wielstra et al., 2017). In European hylids, it is similar to that of the Messinian diverged taxa, H. arborea and H. orientalis (40 km in the Balkans but >100 km in Poland, Dufresnes et al., 2015a, 2016a), H. arborea and H. molleri (>100 km, Stöck et al., 2012, CD and MS unpublished data), suggesting that the speciation continuum in tree frogs spans from the Pliocene to the upper Miocene (Dufresnes et al., 2015a). Given such a timeframe, the Algerian/Tunisian lineage H. cf. meridionalis (“new taxon 1” in Stöck et al., 2008b, 2012) also deserves taxonomic attention, according to our phylogenomic analysis (Figure 2).
The clear-cut Hyla perrini sp. nov./intermedia s. s. transition might be bounded by differential ecological adaptation. The ecological niches of the two taxa are highly disruptive, with projected distributions overlapping only at range margins (Figure 6). A similar pattern was evidenced for other parapatric amphibians meeting in Central Italy (e.g., Lissotriton vulgaris and L. italicus, Iannella et al., 2017). Yet, these differences might also stem from their narrow realized niches, bounded to different geography, i.e., H. intermedia s. s. inhabiting the southern, dry Mediterranean Italy while Hyla perrini sp. nov. is restricted to the more temperate Po Plain. European Hyla species can be quite generalist, e.g., the genetically homogeneous Hyla arborea ranges from southern Greece to Northern Germany (Stöck et al., 2012; Dufresnes et al., 2013). Populations from Hyla perrini sp. nov. also persisted for decades north of the Alps following illegal introductions (Dufresnes et al., 2015c, as H. intermedia).
Are these two species really cryptic? Unsurprisingly, we quantified little morphometric and bioacoustic differences (Figure 7). Amphibians are known for remarkable resemblances between sibling taxa, hence the need for molecular tools to identify them (Dubey et al., 2018). Morphological variation is rather associated to ecological conditions such as climate and geography (as in H. intermedia s. l., Rosso et al., 2004, and other Hyla, Gvoždík et al., 2008), stemming from plasticity or convergent adaptation (e.g., the “spinosus” morphotype found in southeastern European Bufo bufo, Garcia-Porta et al., 2012; Recuero et al., 2012), and therefore not reflecting genetic divergence and taxonomic levels. Surprisingly however, we found the tympanum of Italian taxa to be smaller than that of H. arborea, while the inverse was initially reported (Nascetti et al., 1995; based on Balkan populations of H. arborea). Given the overlapping morphological variation between these species, we discourage the use of any external criteria for identification purposes. Further investigations should be conducted at the contact zone, where character displacement, especially in breeding calls, could have evolved following reinforcement. More generally, our study should motivate future assessment of the phenotypic variation (notably bioacoustics) between Hyla perrini sp. nov. and H. intermedia s. s. species across their respective ranges, to test whether their genetic differentiation is also adaptive.
We thereby conclude that Italian tree frogs consist of two cryptic species, and describe the northern one, Hyla perrini sp. nov., in the next section. We recommend such multidisciplinary approaches for future taxonomic evaluation of cryptic lineages, as also conducted elsewhere (e.g., Diaz-Rodriguez et al., 2017). Particularly, we stress the need for genomic resources to resolve nuclear divergence and assess patterns of introgression, especially in young sibling species where traditional genetic markers (e.g., allozyme, microsatellite, or few nuclear sequences) might lead to erroneous conclusions.
Description of Hyla perrini sp. nov.
Identity and diagnosis
Populations of this species have previously been considered as H. intermedia, due to phenotypic similarity and phylogenetic relationships. Based on mtDNA variation, H. perrini sp. nov. was previously called “clade N” by Canestrelli et al. (2007a,b) and “new taxon 2” by Stöck et al. (2008b, 2012). Hyla perrini is a medium-sized tree frog, morphologically similar to the parapatric H. intermedia s. s. (its sister species) and H. arborea. According to the populations analyzed here, it subtly differs from H. intermedia s. s. by a narrower head and shorter hind legs, yet without diagnostic differences. Like most European hylids, H. perrini displays a green or brownish back coloration and an immaculate belly, separated by a lower black and upper white lateral stripe ending near the groin in an inguinal loop of varying shapes. Other typical characteristics include a horizontal pupil, adhesive disks on fingers and toes, and a yellowish vocal sac on the male throat. The advertisement calls have a similar pattern and structure as H. intermedia s. s. and H. arborea: singular notes made of 7–9, partially fused pulses, totaling ~60–80 ms, interrupted by pauses of 80–120 ms, emitted over the 0.5–5 kHz range, with main bursts of energy at 0.95–1.05 kHz (fundamental frequency) and 2.0–2.4 kHz (dominant frequency); a sonogram and a spectrogram of H. perrini from the type locality is provided in Figure 7B, as well as an audio file (Audio S1). To our knowledge, the tadpoles have not been systematically compared but are supposedly similar between these species (Nöllert and Nöllert, 2003): spiracle sinistral; cloaca dextral; pointy tail tip and high upper fin reaching the eyes, with a single line of dark patches on the tail muscle; Labial Tooth Row Formula (LTRF) 2/3. Hyla perrini has evolved since the upper Pliocene, 2.5–3.5 Mya, and can be distinguished from its relatives, notably H. intermedia s. s., by concordant differences at genetic markers, i.e., ~9% of mitochondrial divergence (COI, cyt-b), and ~0.3% of nuclear divergence (RAD tags).
Distribution
Hyla perrini is endemic to the Po Plain and adjacent valleys in Northern Italy and the Swiss canton of Ticino (Figure 3). Its range borders H. arborea in northeastern Italy/Slovenia, where the two species meet in a narrow contact zone along the Isonzo River (Verardi et al., 2009); it borders and hybridizes with H. intermedia s. s. on the northeastern edge of the Apennines in Emilia Romagna. A now-extinct population was introduced in the 1950s from Ticino to the eastern shores of Lake Geneva (Dufresnes et al., 2015c; Dubey et al., 2018). The species is found from sea level up to about 1,500 m in Italy and 1,100 m in Switzerland.
Holotype
MZL42340, adult male collected by Christophe Dufresnes on April 7th 2018 at Piazzogna, canton of Ticino, Switzerland (46.1361°N, 8.8206°E), subsequently deposited at the Cantonal Museum of Zoology of Lausanne, Switzerland (Figure 8). Full measurements are available in Table S3. SVL of 43.3 mm, head narrower than body, with a rounded snout. Tympanum conspicuous, circular, with diameter 55% of the eye diameter. Nostrils slightly protuberant, located at the same distance to the snout than to the eye. Vomerine teeth present, at the same level than choanae. Long legs, 1.4 × the size of the body. Five long, half-webbed, toes, bearing oval discs; relative lengths from inner to outer toes: 4 > 3 > 5 > 2 > 1; protruding metatarsal tubercle, rounded; subarticular tubercles distinct. Arms slender, with four unwebbed fingers bearing oval discs; subarticular tubercles distinct; thenar tubercle present, elongated. Ventral skin granular; vocal sac folds on the throat; dorsal skin smooth. Coloration in life: dorsum olivaceous green, ventrum white, gular folds of the vocal sac yellowish; prominent black and white lateral stripe running from the nostril to the groin, forming a well-marked inguinal loop. Changes of coloration in alcohol: dorsum bluish gray, vocal sac whitish, black lateral stripe diffused.
Figure 8
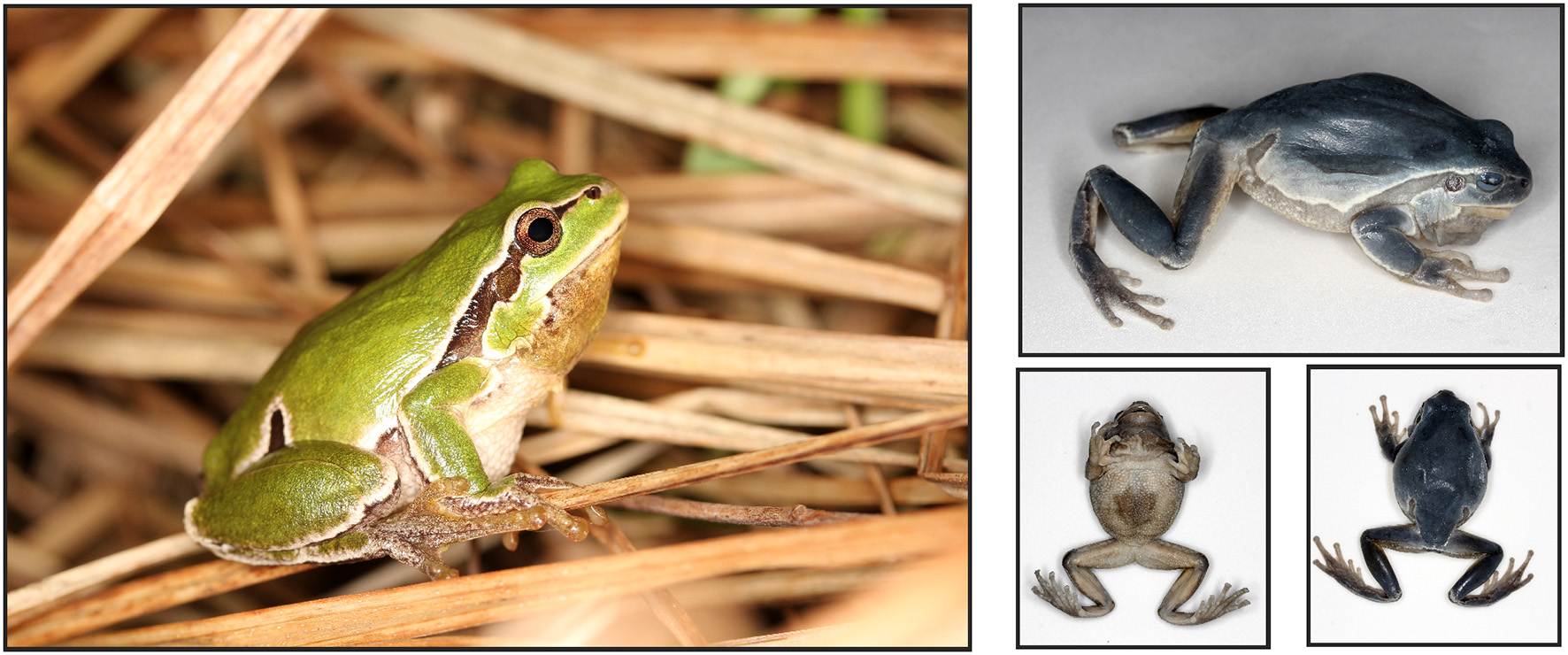

The holotype of Hyla perrini sp. nov. in life and deposited at the Cantonal Museum of Zoology of Lausanne, under voucher MZL42340.
Paratypes
The type series includes four paratypes, two males (MZL42341 and MZL42342) and two females (MZL42343 and MZL42344), collected on April 7th 2018 and May 8th 2018, respectively, by the same collector at the same locality. Their measurements and photos can be found in Table S3 and Figure S5.
Variation
Males body size (SVL) averaged 40.7 mm (35–47 mm) from the two populations of H. perrini surveyed here (Swiss Canton of Ticino and Lombardy, Italy, Table S1D). Rosso et al. (2004) reported mean SVL of 30.7–38.4 mm across five populations of this species. Females are generally larger in tree frogs; few females measured during our 2018 fieldwork were as large as 55 mm SVL. As in related species, back coloration was found to vary from light green to brownish gray, with the tint physiologically changing between environments (CD pers. obs.). The lateral stripe greatly differs between individuals and could be used for individual recognition in mark-capture-recapture studies (e.g., Pellet et al., 2007, on H. arborea). The inguinal loop, especially, can take various shapes, from weak and interrupted dots to heavily marked and elongated commas.
Karyotype
Like other European hylids, Hyla perrini has two sets of 12 chromosomes (2n = 24). A karyotype and idiogram, most likely from this species (specimen K51), is shown by Anderson (1986; p. 61, as “H. arborea”; cf. Anderson, 1991), who analyzed eight metaphases of voucher HGD74640 (Florida State Museum), listed as “? Po River” (p. 347) yet elsewhere detailed “unknown locality in Italy”. Importantly, Anderson (1986) found a secondary constriction in the long arm of chromosome 10 (Anderson, 1986: Figure 2; see Anderson, 1991), interpreted as the position of the nucleolous organizing region (NOR), which is similar to that in the closely related H. arborea (Schmid, 1978). In H. perrini, sex determination is under genetic control by chromosome 1 in an XY manner (LG1, Stöck et al., 2011; Dufresnes et al., 2015b; as H. intermedia), most likely at the dmrt1 gene (Brelsford et al., 2016).
Nomenclatural history
The nomenclatural history of the new species is tightly intertwined with that of H. intermedia Boulenger (1882, p. 381) first described Hyla arborea var. intermedia based on two syntypes: BMNH 52.12.11.55 from Bononia (i.e., Bologna, at the south-eastern edge of the range of the new species), and BMNH 82.7.13.6 from Palermo, Sicily. However, by lectotype designation of BMNH 82.7.13.6, Dubois (1995) made this latter specimen the only name bearer of the epitheton intermedia, and also restricted its type locality to “Palermo,” making intermedia inapplicable to North-Italian tree frogs. In the same work, Dubois (1995) synonymized H. intermedia (i.e., the South-Italian taxon) with H. arborea (Linnaeus, 1758). Using allozymes, Nascetti et al. (1995) found tree frogs from the entire Italian Peninsula to be distinct from H. arborea, and described them as Hyla italica. However, they conducted two additional nomenclatural acts. First, they stated for another old name coined for Italian tree frogs, Hyla variegata (Rafinesque, 1814; type specimens not designated or known to exist; cf. Frost, 2018), an homonym of Rana variegata (Bonnaterre, 1789), hence making it unavailable. Second, they designated a neotype (MZUF 20365) with its type locality near Palermo (Sicily; ca. 37° 00′N, 14° 20′E) and called it H. italica as a replacement name (nomen novum) for H. variegata. By this action and type locality, H. italica became a junior subjective synonym of H. intermedia (as stated by Dubois, 1995), and thus inapplicable to our new species.
Therefore, to the best of our knowledge, there has been no available scientific name for North Italian tree frogs (see also Frost, 2018). In this region, several other authors (cf. Bonato et al., 2007) have previously used either H. arborea or, like De Betta (1857, p. 265), Hyla viridis (= Rana arborea Linnaeus, 1758; cf. Stejneger, 1907), for the herpetofauna of Veneto and Tirol Provinces. Phylogenetic studies did not apply any name other than H. intermedia (Canestrelli et al., 2007a,b). Stöck et al. (2008b, 2012) considered the northern taxon (preliminary called “Hyla new taxon 2”; “n. t. 2”), as a subspecies of H. intermedia (Stöck et al., 2008b, 2012).
To conclude, the name Hyla intermedia applies for the southern lineage (H. intermedia s. s. in this paper), and we have hereby proposed a new name for the tree frogs from Northern Italy.
Etymology
We name this new species Hyla perrini after Nicolas Perrin, professor of ecology, who recently retired from the University of Lausanne (Switzerland). He made countless contributions to the field of evolutionary biology, especially on Palearctic tree frogs, and led numerous studies on phylogeography, population genetics and sex chromosome evolution in this species group, several of them involving this taxon (as H. intermedia or H. n. t. 2). Beyond his scientific excellence, we acknowledge his exceptional mentoring abilities and dedicated support throughout many years of great collaborations.
As a vernacular name in english, we propose H. perrini to be called the “Po's tree frog,” which reflects its geographic distribution and translates as “Raganella padana” (Italian), “Rainette du Po” (French), “Po-Laubfrosch” (German), “Рахкавка з По/Rakhkavka z Po” (Ukrainian), “Квакшаиз По/Kvaksha iz Po” (Russian), “Rela do Pó” (Portuguese), “Trädgroda från Po” (Swedish), “Rainette du Pôô” (Vaudois), and “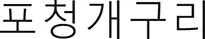 /Po cheong-keguri” (Korean).
/Po cheong-keguri” (Korean).
Natural history and threats
Hyla perrini is a thermophilic amphibian breeding in April-May in shallow standing waters such as small temporary ponds or flooded meadows. Its lifestyle is globally similar to related European Hyla for which detailed information are available in naturalist publications (e.g., Nöllert and Nöllert, 2003; Dufresnes, 2019, where it is treated). In the latest assessments, Italian tree frogs (H. intermedia s. l.) are currently listed as “Least Concern” across the whole range (Andreone et al., 2009) as well as across Italy (Rondinini et al., 2013). Separate re-evaluations for each taxon will be needed, especially for H. perrini which might suffer from agriculture pollution and habitat loss across the heavily-impacted Po Plain, its main range. This new species appears on the Swiss red list of amphibians as “Endangered” (Schmidt and Zumbach, 2005), with relatively stable populations but a restricted distribution in the country (TM pers. comm.). For the time being, we recommend H. perrini to receive the IUCN red list status “Data Deficient”, pending further specific assessment.
Statements
Ethics statement
This study was carried our in accordance of the recommendations of the Ufficio della natura e del paesaggio (Ticino, Switzerland) and the Direzione generale per la protezione della natura e del mare (Italy). The protocol was approved by these committees.
Author contributions
CD, NR, and DJ designed the study. CD, GM, NR, AlB, DC, and DJ conducted fieldwork. RSe conducted labwork. CD, GM, AlB, SL, SD, and DJ analyzed the data. CB-C, OB, AmB, EC, GF, KG, CG, AH, GL, JL, TM, BP, PS, RSa, and MS contributed samples and ideas. CD drafted the main part of the manuscript, supplemented by SL, GM, MS, and DJ, and subsequently improved by all co-authors.
Acknowledgments
The first and main person to thank is Nicolas Perrin, for his scientific support and positive influence, which guided our research for years, on Hyla and other systems. We wish him successful fieldtrips to observe the new species. We are also grateful to E. Geffen, R. Rouag, and N. Bruyndonckx for unpublished samples included in the phylogeny; M. Poujai for supplying bioacoustic equipment; Z. Daeppen for help on the field. Olivier Glaizot and André Keiser for their help at the Cantonal Museum of Zoology of Lausanne. Frogs were captured under collecting permits from the Ufficio della natura e del paesaggio to CD (Ticino, Switzerland) and the Direzione generale per la protezione della natura e del mare to DC (Italy). CD was funded by a fellowship from the Swiss National Science Foundation (fellowship P2LAP3_171818).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fevo.2018.00144/full#supplementary-material
Figure S1Localities used in the niche modeling analyses.
Figure S2PCA on the phylogenomic dataset.
Figure S3Distribution of morphometric data for H. perrini, H. intermedia s. s. and H. arborea.
Figure S4Distribution of bioacoustic variables for H. perrini, H. intermedia s. s. and H. arborea.
Figure S5Ventral and dorsal view of the paratypes of H. perrini.
Table S1Samples included in this study.
Table S1ASequences included in the mitochondrial phylogeny.
Table S1BSamples included in the nuclear phylogeny.
Table S1CLocalities included in the hybrid zone analyses.
Table S1DPopulations sampled for the biometric analyses.
Table S2Variables used in the ecological niche modeling and their contributions.
Table S3Measurements of the type series.
Data Sheet S1Data files analyzed in this study.
Audio S1Chorus of H. perrini from the type locality.
References
1
Anderson K. (1986). A Cytotaxonomic Analysis of the Holarctic Tree Frogs of the Genus Hyla. Ph.D. dissertation, New York University, New York, NY.
2
Anderson K. (1991). Chromosome evolution in Holarctic Hyla tree frogs, in Amphibian Cytogenetics and Evolution, eds D. M.GreenS. K.Sessions (New York, NY: Academic Press). 299–322. 10.1016/B978-0-12-297880-7.50017-2
3
Andreone F. Schmidt B. Vogrin M. Corti C. Sindaco R. Romano A. (2009). Hyla Intermedia. The IUCN Red List of Threatened Species 2009:e.T55517A11323168.
4
Andrews S. (2010). FastQC: a Quality Control Tool for High Throughput Sequence Data. Babraham Bioinformatics. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc
5
Arntzen J. W. McAtear J. Butôt R. Martinez-Solano I. (2018). A common toad hybrid zone that runs from the Atlantic to the Mediterranean. Amphib. Reptil. 39, 41–50. 10.1163/15685381-00003145
6
Arntzen J. W. Recuero E. Canestrelli D. Martinez-Solano I. (2013). How complex is the Bufo bufo species group?Mol. Phylogenet. Evol. 69, 1203–1208. 10.1016/j.ympev.2013.07.012
7
Barton N. Gale K. S. (1993). Genetic analysis of hybrid zones, in Hybrid Zones and the Evolutionary Process, ed HarrisonR. (New York, NY: Oxford University Press), 13–45.
8
Bickford D. Lohman D. J. Sodhi N. S. Ng P. K. L. Meier R. Winker K. et al . (2007). Cryptic species as a window on diversity and conservation. Trends Ecol. Evol.22, 148–155. 10.1016/j.tree.2006.11.004
9
Bisconti R. Canestrelli D. Colangelo P. Nascetti G. (2011). Multiple lines of evidence for demographic and range expansion of a temperature species (Hyla sarda) during the last glaciation. Mol. Ecol. 20, 5313–5327. 10.1111/j.1365-294X.2011.05363.x
10
Bonato L. Fracasso G. Pollo R. Richard J. Semenzato M. (2007). Atlante degli Anfibi e dei Rettili del Veneto, ed Nuovadimensione, Venice: Associazione Faunisti Veneti.
11
Bouckaert R. Heled K. Khünert D. Vaughan T. Wu C. H. Xie D. et al . (2014). BEAST 2: a software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 10:e1003537. 10.1371/journal.pcbi.1003537
12
Bouckaert R. R. Drummond A. J. (2017). bModelTest: bayesian phylogenetic site model averaging and model comparison. BMC. Evol. Biol. 17:42. 10.1186/s12862-017-0890-6
13
Bouckaert R. R. Heled J. (2014). DensiTree 2: seeing trees through the forest. 10.1101/012401
14
Boulenger G. A. (1882). Catalogue of the Batrachia Salientia s. Ecaudata in the Collection of the British Museum, 2nd Edn. London: Taylor and Francis.
15
Brelsford A. Dufresnes C. Perrin N. (2015). High-density sex-specific linkage maps of a European tree frog (Hyla arborea) identify the sex chromosome without information on offspring sex. Heredity116, 177–181. 10.1038/hdy.2015.83
16
Brelsford A. Dufresnes C. Perrin N. (2016). Trans-species variation in Dmrt1 is associated with sex determination in four European tree-frog species. Evolution70, 840–847. 10.1111/evo.12891
17
Canestrelli D Salvi D. Maura M. Bologna M. A. Nascetti G. (2012). One species three Pleistocene evolutionary histories: phylogeography of the Italian crested newt, Triturus carnifex. PLoS ONE7:e41754. 10.1371/journal.pone.0041754
18
Canestrelli D. Bisconti R. Sacco F. Nascetti G. (2013). What triggers the rising of an intraspecific biodiversity hotsport? hints from the agile frog. Sci. Rep.4:5042. 10.1038/srep05042
19
Canestrelli D. Cimmaruta R. Costantini V. Nascetti G. (2006b). Genetic diversity and phylogeography of the Apennine yellow-bellied toad Bombina pachypus, with implications for conservation. Mol. Ecol. 15, 3741–3754. 10.1111/j.1365-294X.2006.03055.x
20
Canestrelli D. Cimmaruta R. Nascetti G. (2007b). Phylogeography and historical demography of the Italian treefrog, Hyla intermedia, reveals multiple refugia, population expansions and secondary contacts within peninsular Italy. Mol. Ecol. 16, 4808–4821. 10.1111/j.1365-294X.2007.03534.x
21
Canestrelli D. Nascetti G. (2008). Phylogeography of the pool frog Rana (Pelophylax) lessonae in the Italian peninsula and Sicily: multiple refugia, glacial expansions and nuclear mitochondrial discordance. J. Biogeogr. 35, 1923–1936. 10.1111/j.1365-2699.2008.01946.x
22
Canestrelli D. Verardi A. Nascetti G. (2007a). Genetic differentiation and history of populations of the Italian treefrog Hyla intermedia: lack of concordance between mitochondrial and nuclear markers. Genetica130, 241–255. 10.1007/s10709-006-9102-9
23
Canestrelli D. Zangari F. Nascetti G. (2006a). Genetic evidence for two distinct species within the Italian endemic Salamandrina terdigitata (Bonnaterre, 1789) (Amphibia: Urodela: Salamandridae). Herpetol. J.16, 221–227.
24
Catchen J. Hohenlohe P. Bassham S. Amores A. Cresko W. (2013). Stacks: an analysis tool set for population genomics. Mol. Ecol. 22, 3124–3140. 10.1111/mec.12354
25
Colliard C. Sicilia A. Turrisi G. F. Arculeo M. Perrin N. Stöck M. (2010). Strong reproductive barriers in a narrow hybrid zone of West-Mediterranean green toads (Bufo viridis subgroup) with Plio-Pleistocene divergence. BMC Evol. Biol. 10:232. 10.1186/1471-2148-10-232
26
De Betta E. (1857). Herpetologia delle provincie venete e del tirolo meridonale. Atti Accad. Agric. Arti Comm.35, 1–365. (Vincenti i Franchini, Verona).
27
Derryberry E. P. Derryberry G. E. Maley J. M. Brumfield R. T. (2014). HZAR: hybrid zone analysis using an R software package. Mol. Ecol. Resour. 14, 652–663. 10.1111/1755-0998.12209
28
Diaz-Rodriguez J. Gehara M. Marquez R. Vences M. Goncalves H. Sequeira F. et al . (2017). Integration of molecular, bioacoustical and morphological data reveals two new cryptic species of Pelodytes (Anura, Pelodytidae) from the Iberian Peninsula. Zootaxa4243, 1–41. 10.11646/zootaxa.4243.1.1
29
Diaz-Rodriguez J. Goncalves H. Sequeira F. Sous-neves T. Tejedo M. Ferrand N. et al . (2015). Molecular evidence for cryptic candidate species in Iberian Pelodytes (Anura, Pelodytidae). Mol. Phylogenet. Evol.83, 224–241. 10.1016/j.ympev.2014.12.007
30
Dubey S. Lavanchy G. Thiébaud J. Dufresnes C. (2018). Herps without border: a new newt case and a review of transalpine alien introductions in Western Europe. Amphib. Reptil. 10.1163/15685381-20181028
31
Dubois A. (1995). The valid scientific name of the Italian treefrog, with comments on the status of some early scientific names of Amphibia Anura, and on some articles of the code concerning secondary homonyms. Dumerilia2, 55–71.
32
Dufresnes C. (2019). Amphibians of Europe, North Africa and the Middle East.London: Bloomsbury, 224.
33
Dufresnes C. Bonato L. Novarini N. Betto-Colliard C. Perrin N. Stöck M. (2014). Inferring the degree of incipient speciation in secondary contact zones of closely related lineages of Palearctic green toads (Bufo viridis subgroup). Heredity1, 9–20. 10.1038/hdy.2014.26
34
Dufresnes C. Borzée A. Horn A. Stöck M. Ostini M. Sermier R. et al . (2015b). Sex-chromosome homomorphy in Palearctic tree frogs results from both turnovers and X-Y recombination. Mol. Biol. Evol. 32, 2328–2337. 10.1093/molbev/msv113
35
Dufresnes C. Brelsford A. Crnobrnja-Isailovic J. Tzankov N. Lymberakis P. Perrin N. (2015a). Timeframe of speciation inferred from secondary contact zones in the European tree frog radiation (Hyla arborea group). BMC Evol. Biol.15:155. 10.1186/s12862-015-0385-2
36
Dufresnes C. Dubey S. Ghali K. Canestrelli D. Perrin N. (2015c). Introgressive hybridization of threatened European tree frogs (Hyla arborea) by introduced H. Intermedia in Western Switzerland. Conserv. Genet.16, 1507–1513. 10.1007/s10592-015-0745-x
37
Dufresnes C. Gangoso L. Perrin N. Stöck M. (2011). Stripeless tree frogs (Hyla meridionalis) with stripes on the Canary Islands. Salamandra47, 232–236.
38
Dufresnes C. Litvinchuk S. N. Borzée A. Jang Y. Li J. T. Miura I. et al . (2016c). Phylogeography reveals an ancient cryptic radiation in East-Asian tree frogs (Hyla japonica group) and complex relationships between continental and island lineages. BMC Evol. Biol. 16:253. 10.1186/s12862-016-0814-x
39
Dufresnes C. Litvinchuk S. N. Leuenberger J. Ghali K. Zinenko O. Stöck M. et al . (2016b). Evolutionary melting pots: a biodiversity hotspot shaped by ring diversifications around the Black Sea in the Eastern tree frog (Hyla orientalis). Mol. Ecol.25, 4285–4300. 10.1111/mec.13706
40
Dufresnes C. Lymberakis P. Kornilios P. Savary R. Perrin N. Stöck M. (2018). Phylogeography of Aegean green toads (Bufo viridis subgroup): continental hybrid swarm vs. insular diversification with discovery of a new island endemic. BMC Evol. Biol. 18:67. 10.1186/s12862-018-1179-0
41
Dufresnes C. Majtyka T. Baird S. J. E. Gerchen J. F. Borzée A. Savary R. et al . (2016a). Empirical evidence for large X-effects in animals with undifferentiated sex chromosomes. Sci. Rep. 6:21029. 10.1038/srep21029
42
Dufresnes C. Wassef J. Ghali K. Brelsford A. Stöck M. Lymberakis P. et al . (2013). Conservation phylogeography: does historical diversity contribute to regional vulnerability in European tree frogs (Hyla arborea)?Mol. Ecol.22, 5669–5684. 10.1111/mec.12513
43
Earl D. A. vonHoldt B. M. (2012). STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour.4, 359–361. 10.1007/s12686-011-9548-7
44
Elith J. (2000). Quantitative methods for modeling species habitat: comparative performance and an application to Australian plants, in Quantitative Methods for Modeling Species Habitat, eds FersonS.BurgmanM. (New York, NY: Springer), 39–58. 10.1007/0-387-22648-6_4
45
Evanno G. Regnaut S. Goudet J. (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620. 10.1111/j.1365-294X.2005.02553.x
46
Frost D. R. (2018). Amphibian Species of the World: an Online Reference. Version 6.0. New York NY: American Museum of Natural History.
47
Garcia-Porta J. Litvinchuk S. N. Crochet P. A. Romano A. Geniez P. H. Lo-Valvo M. et al . (2012). Molecular phylogenetics and historical biogeography of the west-palearctic common toads (Bufo bufo species complex). Mol. Phylogenet. Evol.63, 113–130. 10.1016/j.ympev.2011.12.019
48
Gvoždík V. Canestrelli D. Garcia-Paris M. Moravec J. Nascetti G. Recuero E. et al . (2015). Speciation history and widespread introgression in the European short-call tree frogs (Hyla arborea sensu lato, H. intermedia and H. sarda).Mol. Phylogenet. Evol.83:143155. 10.1016/j.ympev.2014.11.012
49
Gvoždík V. Moravec J. Klütsch C. Kotlik P. (2010). Phylogeography of the Middle Eastern tree frogs (Hyla, Hylidae, Amphibia) as inferred from nuclear and mitochondrial DNA variation, with a description of a new species. Mol. Phylogenet. Evol. 55, 1146–1166. 10.1016/j.ympev.2010.03.015
50
Gvoždík V. Moravec J. Kratochvil L. (2008). Geographic morphological variation in parapatric Western Palearctic tree frogs, Hyla arborea and Hyla savignyi: are related species similarly affected by climatic conditions?Biol. J. Linn. Soc. 95, 539–556. 10.1111/j.1095-8312.2008.01056.x
51
Hauswaldt J. S. Angelini C. Pollok A. Steinfartz S. (2011). Hybridization between two ancient salamander lineages: molecular evidence for endemic spectacled salamanders on the Apennine peninsula. J. Zool. 284, 248–256. 10.1111/j.1469-7998.2011.00807.x
52
Hewitt G. M. (2011). Quaternary phylogeography: the roots of hybrid zones. Genetica139, 617–638. 10.1007/s10709-011-9547-3
53
Iannella M. Cerasoli F. Biondi M. (2017). Unravelling climate influences on the distribution of the parapatric newts Lissotriton vulgaris meridionalis and L. italicus.Front. Zool.14:55. 10.1186/s12983-017-0239-4
54
Köhler J. Jansen M. Rodriguez A. Kok P. J. R. Toledo L. F. Emmrich M. et al . (2017). The use of bioacoustics in anuran taxonomy: theory, terminology, methods and recommendations for best practice. Zootaxa4251, 1–124. 10.11646/zootaxa.4251.1.1
55
Kotsakis T. (1980). Osservazioni sui vertebrati quaternari della sardegna. B. Soc. Geol. Ital. 99, 151–165.
56
Lampert K. P. Rand A. S. Mueller U. G. Ryan M. J. (2003). Fine-scale genetic pattern and evidence for sex-biased dispersal in the túngara frog, Physalaemus pustulosus. Mol. Ecol.12, 3325–3334. 10.1046/j.1365-294X.2003.02016.x
57
Langmead B. Salzberg S. L. (2012). Fast gapped-read alignment with Bowtie 2. Nat. Methods9, 357–359. 10.1038/nmeth.1923
58
Li H. (2011). A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics27, 2987–2993. 10.1093/bioinformatics/btr509
59
Li J. T. Wang J. S. Nian H. H. Litvinchuk S. N. Wang J. Li Y. et al . (2015). Amphibians crossing the Bering land bridge: evidence from holarctic treefrogs (Hyla, Hylidae, Anura). Mol. Phylogenet. Evol. 87, 80–90. 10.1016/j.ympev.2015.02.018
60
Lymberakis P. Poulakakis N. Manthalou G. Tsigenopoulos C. S. Magoulas A. Mylonas M. (2007). Mitochondrial phylogeography of Rana (Pelophylax) populations in the Eastern Mediterranean region. Mol. Phylogenet. Evol. 44, 115–125. 10.1016/j.ympev.2007.03.009
61
Mattoccia M. Marta S. Romano A. Sbordoni V. (2011). Phylogeography of an Italian endemic salamander (genus Salamandrina): glacial refugia, postglacial expansions, and secondary contact. Biol. J. Linn. Soc. 104, 903–922. 10.1111/j.1095-8312.2011.01747.x
62
Maura M. Salvi D. Bologna M. A. Nascetti G. Canestrelli D. (2014). Northern richness and cryptic refugia: phylogeography of the Italian smooth newt Lissotriton vulgaris meridionalis. Biol. J. Linn. Soc. 113, 590–603. 10.1111/bij.12360
63
Mayr E. (1942). Systematics and the Origin of Species, from the Viewpoint of a Zoologist. Cambridge: Harvard University Press.
64
Nascetti G. Lanza B. Bullini L. (1995). Genetic data for the specific status of the Italian treefrog (Amphibia: Anura: Hylidae). Amphib. Reptil.16, 215–227. 10.1163/156853895X00019
65
Nöllert A. Nöllert C. (2003). Guide des Amphibiens d'Europe. Paris: Delachaux et Niestlé.
66
Pabijan M. Crottini A. Reckwell D. Irisarri I. Hauswaldt J. S. Vences M. (2012). A multigene species tree for Western Mediterranean painted frogs (Discoglossus). Mol. Phylogenet. Evol.64, 690–696. 10.1016/j.ympev.2012.05.017
67
Pabijan M. Zielinski P. Dudek K. Stuglik M. Babik W. (2017). Isolation and gene flow in a speciation continuum in newts. Mol. Phylogenet. Evol. 116, 1–12. 10.1016/j.ympev.2017.08.003
68
Paris J. R. Stevens J. R. Catchen J. M. (2017). Lost in parameter space: a road map for stacks. Methods Ecol. Evol. 8, 1360–1373. 10.1111/2041-210X.12775
69
Pellen R. Popescu S. M. Suc J. P. Melinte-Dobrinescu M. C. Rubino J. L. Rabineau M. et al . (2017). The apennine foredeep (Italy) during the latest messinian: lago mare reflects competing brackish and marine conditions based on calcareous nannofossils and dinoflagellate cysts. Geobios50, 237–257. 10.1016/j.geobios.2017.04.004
70
Pellet J. Helfer V. Yannic G. (2007). Estimating population size in the European tree frog (Hyla arborea) using individual recognition and chorus counts. Amphib. Reptil. 28, 287–294. 10.1163/156853807780202530
71
Phillips S. Anderson R. Schapire R. (2006). Maximum entropy modeling of species geographic distributions. Ecol. Modell. 190, 231–259. 10.1016/j.ecolmodel.2005.03.026
72
Picotti V. Pazzaglia F. J. (2008). A new active tectonic model for the construction of the Northern Apennines mountain front near Bologna (Italy). J. Geophys. Res. B Solid Earth 113:B08412. 10.1029/2007JB005307
73
Plötner J. Baier F. Akin C. Mazepa G. Schreiber R. Beerli P. et al . (2012). Genetic data reveal that waterfrogs of Cyprus (genus Pelophylax) are an endemic species of Messinian origin. Zoosyst. Evol. 88, 261–283. 10.1002/zoos.201200021
74
Pritchard J. K. Stephens M. Donnelly P. (2000). Inference of population structure using multilocus genotype data. Genetics155, 945–959.
75
Recuero E. Canestrelli D. Vörös J. Szabo K. Poyarkov N. A. Arntzen J. W. et al . (2012). Multilocus species tree analyses resolve the radiation of the widespread Bufo bufo species group. Mol. Phylogent. Evol. 62, 71–86. 10.1016/j.ympev.2011.09.008
76
Recuero E. Iraola A. Rubio X. Machordom A. Garcia-Paris M. (2007). Mitochondrial differentiation and biogeography of Hyla meridionalis (Anura: Hylidae): an unusual phylogeographical pattern. J. Biogeogr. 34, 1207–1219. 10.1111/j.1365-2699.2007.01688.x
77
Rödder D. Engler J. O. (2011). Quantitative metrics of overlaps in Grinnellian niches: advances and possible drawbacks. Global Ecol. Biogeogr. 20, 915–927. 10.1111/j.1466-8238.2011.00659.x
78
Rondinini C. Battistoni A. Peronace V. Teofili C. (2013). Lista Rossa IUCN dei Vertebrati Italiani. Roma: Comitato Italiano IUCN e Ministero dell'Ambiente e della Tutela del Territorio e del Mare.
79
Rosso A. Castellano S. Giacoma C. (2004). Ecogeographic analysis of morphological and life-history variation in the Italian treefrog. Evol. Ecol.18, 303–321. 10.1007/s10682-004-0925-5
80
Rosso A. Castellano S. Glacoma C. (2006). Preferences for call spectral properties in Hyla intermedia. Ethology112, 599–607. 10.1111/j.1439-0310.2005.01186.x
81
Schmid M. (1978). Chromosome banding in Amphibia. I. Constitutive heterochromatin and nucleolus organizer regions in Bufo and Hyla. Chromosoma66, 361–388. 10.1007/BF00328536
82
Schmidt B. R. Zumbach S. (2005). Liste Rouge des Amphibiens Menaces en Suisse. Edit. Office federal de l'environnement, des forêts et du Paysage (OFEFP), Berne, et Centre de Coordination Pour la Protection des amphibiens et des Reptiles de Suisse (KARCH), Berne: Série OFEFP: L'environnement pratique.
83
Schmitt T. (2007). Molecular biogeography of Europe: pleistocene cycles and postglacial trends. Front. Zool. 4:11. 10.1186/1742-9994-4-11
84
Schoener T. W. (1968). Anolis lizards of Bimini: resource partitioning in a complex faunaEcology49, 704–726.
85
Sciuchetti L. Dufresnes C. Cavoto E. Brelsford A. Perrin N. (2018). Dobzhansky-Muller incompatibilities, dominance drive, and sex-chromosome introgression at secondary contact zones: a simulation study. Evolution72, 1351–1361. 10.1111/evo.13510
86
Shaffer H. B. Gidis M. McCartney-Melstad E. Neal K. M. Oyamaguchi H. M. Tellez M. et al . (2015). Conservation genetics and genomics of amphibians and reptiles. Annu. Rev. Anim. Biosci. 3, 113–138. 10.1146/annurev-animal-022114-110920
87
Smith S. A. Stephens P. R. Wiens J. J. (2005). Replicate patterns of species richness, historical biogeography and phylogeny in Holarctic treefrogs. Evolution59, 2433–2450. 10.1111/j.0014-3820.2005.tb00953.x
88
Stejneger L. (1907). Herpetology of Japan and adjacent territory. Bull. U.S. Natl. Mus. 58, 1–577.
89
Stöck M. Dubey S. Klütsch C. Litvinchuk S. N. Scheidt U. Perrin N. (2008b). Mitochondrial and nuclear phylogeny of circum-Mediterranean tree frogs from the Hyla arborea group. Mol. Phylogenet. Evol. 49, 1019–1024. 10.1016/j.ympev.2008.08.029
90
Stöck M. Dufresnes C. Litvinchuk S. N. Lymberakis P. Biollay S. Berroneau M. et al . (2012). Cryptic diversity among Western Palearctic tree frogs: postglacial range expansions, range limits, and secondary contacts of three European tree frog lineages (Hyla arborea group). Mol. Phylogenet. Evol.65, 1–9. 10.1016/j.ympev.2012.05.014
91
Stöck M. Horn A. Grossen C. Lindtke D. Sermier R. Betto-Colliard C. et al . (2011). Ever-young sex chromosomes in European tree frogs. PLoS Biol. 9:e1001062. 10.1371/journal.pbio.1001062
92
Stöck M. Sicilia A. Belfiore N. M. Buckley D. Lo Brutto S. Lo Valvo M. et al . (2008a). Post-Messinian evolutionary relationships across the sicilian channel: mitochondrial and nuclear markers link a new green toad from sicily to african relatives. BMC Evol. Biol. 8:56. 10.1186/1471-2148-8-56
93
Sueur J. Aubin T. Simonis C. (2008). Equipment review: Seewave, a free modular tool for sound analysis and synthesis. Bioacoustics18, 218–226. 10.1080/09524622.2008.9753600
94
Swets J. (1988). Measuring the accuracy of diagnostic systems. Science240, 1285–1293. 10.1126/science.3287615
95
Veith M. Göçmen B. Sotiropoulos K. Kieren S. Godmann O. Steinfarz S. (2016). Seven at one blow: the origin of major lineages of the viviparous Lycian salamanders (Lyciasalamandra, Veith and Steinfarz, 2004) was triggered by a single paleo-historic event. Amphib. Reptil. 37, 373–387. 10.1163/15685381-00003067
96
Verardi A. Canestrelli D. Nascetti G. (2009). Nuclear and mitochondrial patterns of introgression between the parapatric European tree frogs Hyla arborea and H. intermedia.Ann. Zool. Fennici46, 247–258. 10.5735/086.046.0402
97
Vos C. C. Ter Braak C. J. F. Nieuwenhuizen W. (2000). Incidence function modelling and conservation of the tree frog Hyla arborea in the Netherlands. Ecol. Bull. 48, 165–180. 10.2307/20113255
98
Warren D. L. Glor R. E. Turelli M. (2010). ENMTOOLS: a toolbox for comparative studies of environmental niche models. Ecography. 33, 607–611. 10.1111/j.1600-0587.2009.06142.x
99
Watanabe S. Hajima T. Sudo K. Nagashima T. Takemura H. Okajima H. et al . (2011). MIROC-ESM 2010: model description and basic results of CMIP5-20c3 m experiments. Geosci. Model. Dev. 4, 845–872. 10.5194/gmd-4-845-2011
100
Waters J. M. Fraser C. I. Hewitt G. M. (2013). Fonder takes all: density-dependent processes structure biodiversity. Trends Ecol. Evol. 28, 78–85. 10.1016/j.tree.2012.08.024
101
Wielstra B. Arntzen J. W. (2011). Unravelling the rapid radiation of crested newts (Triturus cristatus superspecies) using complete mitogenomic sequences. BMC. Evol. Biol. 11:162. 10.1186/1471-2148-11-162
102
Wielstra B. Baird A. B. Arntzen J. W. (2013). A multimarker phylogeography of crested newts (Triturus cristatus superspecies) reveals cryptic species. Mol. Phylogenet. Evol. 67, 161–175. 10.1016/j.ympev.2013.01.009
103
Wielstra B. Burke T. Butlin R. K. Avci A. Uzum N. Bozkurt E. et al . (2017). A genomic footprint of hybrid zone movement in crested newts. Evol. Lett. 12, 93–101. 10.1002/evl3.9
Summary
Keywords
amphibians, bioacoustics, hybrid zone, phylogenomics, RAD-seq, taxonomy
Citation
Dufresnes C, Mazepa G, Rodrigues N, Brelsford A, Litvinchuk SN, Sermier R, Lavanchy G, Betto-Colliard C, Blaser O, Borzée A, Cavoto E, Fabre G, Ghali K, Grossen C, Horn A, Leuenberger J, Phillips BC, Saunders PA, Savary R, Maddalena T, Stöck M, Dubey S, Canestrelli D and Jeffries DL (2018) Genomic Evidence for Cryptic Speciation in Tree Frogs From the Apennine Peninsula, With Description of Hyla perrini sp. nov. Front. Ecol. Evol. 6:144. doi: 10.3389/fevo.2018.00144
Received
12 June 2018
Accepted
03 September 2018
Published
02 October 2018
Volume
6 - 2018
Edited by
Mariana Mateos, Texas A&M University, United States
Reviewed by
Wieslaw Babik, Jagiellonian University, Poland; Alberto G. Sáez, Universität Zürich, Switzerland
Updates
Copyright
© 2018 Dufresnes, Mazepa, Rodrigues, Brelsford, Litvinchuk, Sermier, Lavanchy, Betto-Colliard, Blaser, Borzée, Cavoto, Fabre, Ghali, Grossen, Horn, Leuenberger, Phillips, Saunders, Savary, Maddalena, Stöck, Dubey, Canestrelli and Jeffries.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Christophe Dufresnes christophe.dufresnes@unil.ch
This article was submitted to Phylogenetics, Phylogenomics, and Systematics, a section of the journal Frontiers in Ecology and Evolution
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.