- 1Institute of Ecology and Evolution, University of Oregon, Eugene, OR, United States
- 2Department of Integrative Biology, University of California, Berkeley, Berkeley, CA, United States
Bacterial communities are a major component of global diversity and are intimately involved in most terrestrial biogeochemical processes. Despite their importance, we know far less about the response of bacteria to human-induced environmental change than we do about other organisms. Understanding the response of organisms to land use change is especially pressing for tropical rainforests, which are being altered at a higher rate than any other ecosystem. Here, we conduct a meta-analysis of studies performed in each of the major tropical rainforest regions to ask whether there are consistent responses of belowground bacterial communities to the conversion of tropical rainforest to agriculture. Remarkably, we find common responses despite wide variation across studies in the types of agriculture practiced and the research methodology used to study land use change. These responses include changes in the relative abundance of phyla, most notably decreases in Acidobacteria [−1.94 ± 1.1 fold (average ± 95% CI)] and Proteobacteria (−1.38 ± 1.0 fold) and increases in Actinobacteria (1.55 ± 1.1 fold), Chloroflexi (3.47 ± 1.2 fold), and Firmicutes (6.6 ± 1.3 fold). We also find that alpha diversity (at the scale of single soil cores) consistently increases (1.17 ± 1.0 fold) with ecosystem conversion. These consistent responses suggest that, while there is great diversity in agricultural practices across the tropics, common features such as the use of slash-and-burn tactics have the potential to alter bacterial community composition and diversity belowground.
Introduction
Deforestation of the world's remaining tropical rainforests is occurring at a rapid rate and it is important to understand how the loss of tropical forest will change belowground bacterial communities. Soil bacteria mediate most biogeochemical cycles (Madsen, 2011) and comprise the majority of global biodiversity (Hug et al., 2016). When ecosystems undergo conversion, such as from forest to agriculture, biogeochemical cycles are often profoundly altered (Neill et al., 1997, 2005; DeFries et al., 1999; Verchot et al., 2000) and the diversity of plant and animal communities can decrease, along with changes in species composition (Perfecto and Snelling, 1995; Banks et al., 2009; Bernard et al., 2009; Gardner et al., 2009; Gibson et al., 2011; Newbold et al., 2015). No ecosystem type is experiencing conversion at a faster rate than tropical rainforests (Dirzo and Raven, 2003), which are declining primarily due to agricultural expansion (De Moraes et al., 1996). From 1980 to 2012, approximately 1.3 million km2 of tropical rainforest were cleared for agriculture, an area the size of France, Spain and the UK combined (Gibbs et al., 2010; Hansen et al., 2013).
Across the tropics, forest-to-agriculture conversion occurs through a similar mechanism, slash-and-burn, but differs in eventual agricultural characteristics. In slash-and-burn, valuable trees are removed for lumber and the remaining vegetation is burned to clear the land for agricultural use. The predominant agricultural use varies by region; oil palm plantations in southeast Asia (Koh and Wilcove, 2008), beef cattle pastures or soybean plantation in the Amazon Basin (Morton et al., 2006) and small scale subsistence farming of plantain or manioc in the Congo Basin (Sunderlin et al., 2000; Zhang et al., 2006). Duration of agricultural use also varies by region, from one-three years for plantations in the Congo (Zhang et al., 2002) to decades of continual use in Southeast Asia and the Amazon (Rodrigues et al., 2013; Tripathi et al., 2016). While there is great diversity in agricultural characteristics across the tropics, consistent patterns in bacterial response should nonetheless emerge if community change is driven by common aspects of ecosystem conversion, such as slash-and-burn tactics.
Earlier research (prior to the advent of high-throughput molecular approaches) on the effect of land use change in the tropics on belowground microbial communities observed that land use change led to changes in microbial community composition and often increases in diversity (e.g., Borneman and Triplett, 1997; Da Silveira et al., 2006; Pereira et al., 2006; Jesus et al., 2009). These earlier studies inspired a growing number of recent studies utilizing next-generation sequencing to document the effects of land use change on belowground microbial communities. We took advantage of this growing body of literature to ask whether there are consistent responses of belowground microbial communities to ecosystem conversion. Specifically, we conducted a meta-analysis to ask whether there were consistent effects of forest to agriculture conversion on the diversity and composition of belowground bacterial communities and soil chemical properties across the world's three largest tropical rainforests (the Amazon Basin, the Congo Basin, and Southeast Asia). Despite regional differences in agricultural practices, and a diversity of research methodologies across studies, we observed consistent changes in diversity and phylum level composition.
Methods
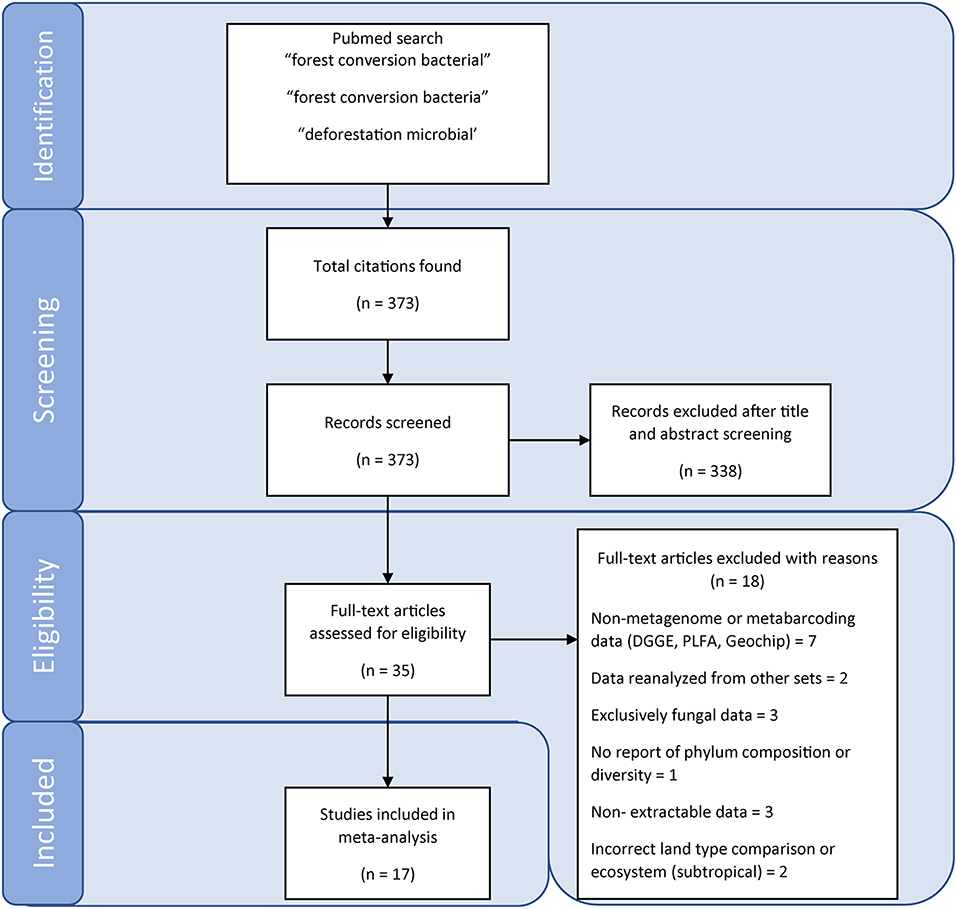
We conducted a literature search on Web of Science and Google Scholar on the 23 January 2019. Search terms included: forest conversion bacteria, forest conversion bacterial, and deforestation microbial. Titles and abstracts of articles matching our search terms were then screened for studies that directly compare soil samples from primary tropical rainforest with proximal agricultural sites, analyze metabarcode or metagenome sequences and report at least one diversity metric, composition at the phylum level and soil chemical characteristics. While studies using pre-next generation sequencing approaches [such as denaturing gradient gel electrophoresis (DGGE), terminal restriction length polymorphism (tRFLP), phospholipid fatty acids (PLFA), or Sanger sequencing of amplicon clone libraries] provide valuable insights, they were not included in our meta-analysis because of the incommensurability and/or uncertainty of taxonomic affiliation (e.g., DGGE, tRFLP, PLFA) or low sampling depth (e.g., clone libraries), which limits their utility in the quantitative meta-analyses used here. The literature search workflow is presented in Figure 1.
Following identification, means (X), standard deviations (SDs), and sample sizes (n) were extracted from the published studies. If not reported, these statistics were derived from other metrics; standard error, 95% confidence interval, or diversity metrics and composition were derived and summarized from OTU tables using the vegan package in R 3.4.4. The meta-analysis was then conducted following Borenstein et al. (2009). Briefly, for each study a response ratio was determined for diversity, phylum-level relative abundance and soil chemical factors using the following equation
Where for study i, the means of forest and agricultural samples are represented, respectively, by xfi and xai. To assess responses across all studies a weighting value (w) was calculated.
The summary response ratio (rrw) was then calculated by averaging individual response ratios and weighting values.
To determine if the summary response ratio was significantly different from zero, we then conducted two tailed t-tests (α = 0.05). For phyla that we found to have a significant response, we then examined the consistency of this response at finer taxonomic levels (order, class) using findings reported in the published papers or by deriving information from the underlying OTU tables.
Recent studies (Lajeunesse, 2015) have stressed the importance of verifying that logRR is a valid metric for ecological meta-analysis when studies have (1) small sample sizes, (2) large differences in means between treatment and control groups, and (3) means near zero. To determine whether effect sizes were valid and accurate approximations we used a diagnostic proposed by (Hedges et al., 1999):
and a modified diagnostic by Lajeunesse (2015),
which improves the confirming accuracy of effect sizes for small sample sizes that also lie outside the problematic ranges near zero. Samples failing the Geary test (<3 in either equation) were identified as more likely to be inaccurate. To account for the influence of these samples, response ratios were recalculated excluding samples that failed the Geary test and compared to results including all samples.
Results
Our literature search identified 17 studies (Supplementary Table 1), with representation from all three major rainforests (Figure 2). A diverse set of agriculture types were represented, including cattle pasture (predominantly in the Amazon), oil palm plantation (exclusively in Southeast Asia), and banana/manioc plantation in the Congo basin (Supplementary Table 1). A variety of sampling techniques were used; with soil cores collected along transects, quadrats or nested sampling schemes, and sequencing of samples done individually or as pooled samples.
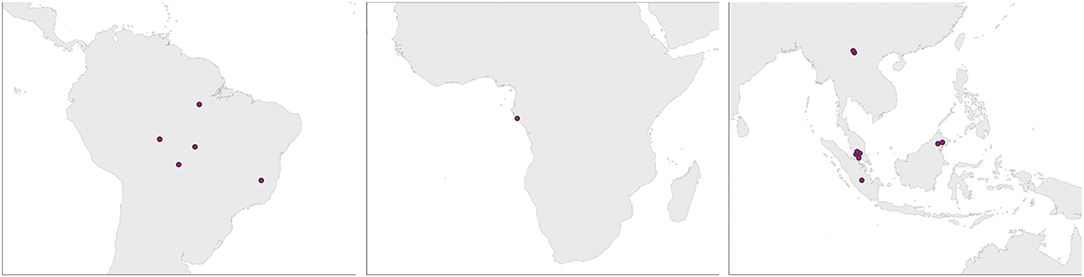
Figure 2. Study locations from the Amazon, Congo, and SE Asia. Studies from the same site are represented by a single dot.
The V3 and/or the V4 region of 16S rRNA gene was sequenced in all studies using 454 pyrosequencing or the Illumina Hiseq or Miseq platforms. Three studies sequenced metagenomes, with one study solely analyzing metagenomic data and the other two pairing metagenomes with 16S metabarcoding (Supplementary Table 1). A wide variety of diversity metrics were used to quantify alpha diversity across studies. The Shannon index was the most commonly reported, followed by richness, with other metrics (Simpson, Faiths PD, Hills effective species number) included in only a few studies. There was not a single diversity metric reported in every study, but since response ratio is a unitless metric, differing metrics were considered together as alpha diversity.
Diversity
Alpha diversity results were reported in all but one study. In these studies, the majority (9), reported increases in alpha diversity with conversion (p > 0.001), six reported no change, and one study reported a decrease (Figure 3). In contrast, beta diversity was not reported in a majority of studies. Among the eight studies that reported beta diversity, a wide variety of sampling and statistical approaches were used, and the results were highly variable. This precluded a quantitative meta-analysis of beta diversity shifts. Among studies that did analyze beta diversity, authors reported significant decreases in beta diversity following conversion in three studies (Rodrigues et al., 2013; Mendes et al., 2015; Meyer et al., 2019), while two studies (de Carvalho et al., 2016; Cuer et al., 2018) reported significant increases, and the remainder reported no response.
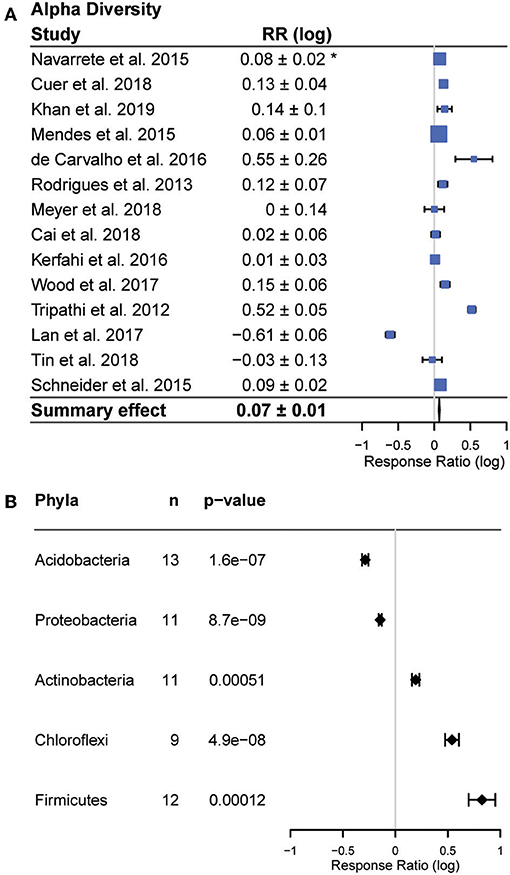
Figure 3. Effects of forest to agriculture conversion on bacterial diversity and composition. (A) Changes in alpha diversity measured as the response ratio (RR). Positive values indicate higher diversity in agricultural areas and negative values indicate higher diversity in forest areas. The bars represent the 95% confidence intervals (CIs) and box size corresponds to study weight in the summary effect size. Summary response ratio was significantly different from zero (two tailed t-tests, p < 0.001). (B) Changes in abundance of phyla reported in a majority of studies, including the number of studies reporting each phylum out of 16 studies. *Confidence interval (95%) of the response ratio.
Composition
Bacterial community composition varied across studies, but members of five phyla (Actinobacteria, Firmicutes, Chloroflexi, Acidobacteria, Proteobacteria) were reported in a majority of studies (Figure 3). We observed consistent responses to land use change in each of these five phyla (all p < 0.01). The relative abundances of Actinobacteria (fold change (average ± 95% CI) 1.55 ± 1.1 Chloroflexi (average fold change 3.47 ± 1.2) and Firmicutes (average fold change 6.6 ± 1.3) increased with conversion, while the relative abundance of Acidobacteria (average fold change −1.94 ± 1.1) and Proteobacteria (average fold change −1.38 ± 1.0) decreased (Figure 3). In addition to having overall significant effects, these responses were also remarkably universal across studies (Figure 3). Only two studies reported any changes in abundance that were opposite to the summary effect; Firmicutes were reported to decrease in Cuer et al. (2018), and Schneider et al. (2015) reported a decrease in Actinobacteria and increase in Acidobacteria with conversion (Figure 4).
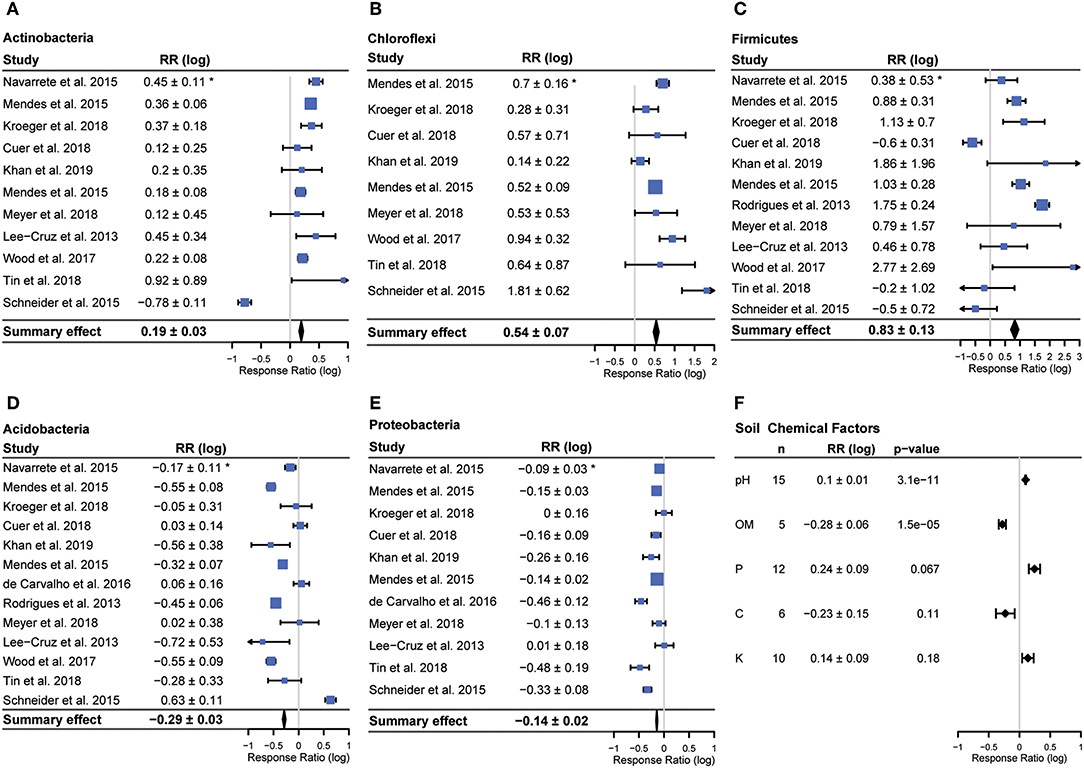
Figure 4. Forest plot for each phylum with a significant summary effect (all p < 0.001) (A–E) and summary effects for soil chemical data (F). Bars denote 95% confidence intervals, boxes are centered on means and box size corresponds to study weight for calculating summary effect. *Confidence interval (95%) of the response ratio.
The Geary test excluded 5 samples, four Firmicutes and one Chloroflexi. Summary effects were very similar for Firmicutes (summary effect 0.83 ± 0.13 vs. 0.9 ± 0.13 excluding Geary < 3 samples) and the summary effect was identical for Chloroflexi (summary effect 0.54 ± 0.07 for both) (Supplementary Figure 1). Given the small number of total studies, the similarity of summary effects when excluding potentially problematic studies and the fact that the goal of this study was ask whether responses were consistent rather than to determine the exact degree of response, the findings reported here include all studies, including those scoring < 3 on the Geary test.
Within the five phyla exhibiting consistent responses, we looked for consistencies at finer taxonomic levels, either by analyzing available OTU tables or by compiling findings reported in the papers. Such data were rare among the studies; only three reported responses of at least some classes or orders, and four provided OTU tables. Within these seven studies, we found no consistent responses at the class or order level within the phyla that exhibited consistent responses. Thus, while we have no evidence for common responses at finer taxonomic levels, it should be noted that this hypothesis was tested using far fewer studies than used for testing phylum level hypotheses.
Soil Chemical Factors
The only soil chemical factor that was both reported in a majority of studies and varied significantly with conversion was pH (p < 0.001), which increased following conversion (Figure 4). The other two soil chemical factors reported in a majority of studies, potassium (K) and phosphorus (P), did not vary significantly with conversion. Total carbon was reported in 6 studies and organic matter was reported in 5 studies. Organic matter decreased significantly with conversion (p < 0.01) in those studies that reported it. While we could have estimated organic matter from total carbon using the van Bemmelen factor (for studies that reported total carbon but not organic matter), we chose not to do this since this factor has been shown to be inconsistent across studies that estimate both total carbon and organic matter (Pribyl, 2010).
Discussion
Tropical rainforests harbor greater biodiversity than any other terrestrial ecosystem and are important contributors to global biogeochemical processes (Gardner et al., 2009). These ecosystems are being lost rapidly, primarily from land use conversion for agriculture (Gibbs et al., 2010; Hansen et al., 2013). The effects of land use change on the biodiversity of plants and animals is well-documented, but effects of land use change on belowground microbial communities, which comprise a majority of global biodiversity and which mediate most biogeochemical cycles (Madsen, 2011; Hug et al., 2016), is not as well-understood. Using a meta-analysis of recent studies spanning the world's tropical zone, we detected consistent changes to the composition and diversity of soil bacterial communities to land use change. While effect sizes were sometimes small, the fact that we detect consistent bacterial responses is remarkable considering the wide geographic range, the immense bacterial diversity of the study sites, and considerable methodological variation.
Diversity
The increase in bacterial alpha diversity following conversion to agriculture is consistent with earlier work using pre-next generation sequencing approaches (da C Jesus et al., 2009; Jesus et al., 2009), and is in contrast to commonly reported decreases in the diversity of plants (Ibrahim et al., 2003), animals (Brook et al., 2003; Gardner et al., 2009; Lim, 2017), and fungi (Mueller et al., 2014; McGuire et al., 2015; Kerfahi et al., 2016; Lan et al., 2017; Cai et al., 2018), following agricultural conversion. This may result from ecological differences between bacteria and eukaryotes; for example, bacterial diversity may respond to changes in soil chemical properties, rather than being directly affected by land use change. We found that pH consistently increased with land use change across the studies we surveyed, and others have observed that pH can be correlated with bacterial richness (Fierer and Jackson, 2006; Lauber et al., 2009). Aboveground productivity could also drive this trend. Agricultural management often promotes high productivity, which could increase soil labile carbon, driving increased bacterial diversity (Dias-Filho et al., 2001). While we found that organic matter decreased with conversion, labile carbon is a subset of organic matter so it is possible for organic matter to decrease while labile carbon increases (Sahoo et al., 2019). Alternatively, the apparent increase in bacterial diversity could be an artifact of sampling. Increased alpha diversity at the level of a soil core may not represent landscape-level responses (i.e., gamma diversity). Bacteria may actually experience a landscape-level loss of diversity, similar to macro-organisms, if decreases in beta diversity offset or overwhelm increases in alpha diversity (as in Rodrigues et al., 2013). In this instance, the apparent differences may be more methodological than ecological. This, however, is not always the case. For instance, de Carvalho et al. (2016) found higher alpha, beta, and gamma diversity in converted sites. These contrasting results thus highlight the importance of increased efforts to characterize the different dimensions of diversity in the context of land use change. Unfortunately, we cannot address this hypothesis because beta diversity was not consistently reported in the studies we surveyed and when measured was assessed with a wide variety of approaches. Consequently, we found that the results of beta diversity assessments were mixed. The difference in responses of eukaryotic and bacterial diversity to land use change should be investigated further to determine if this difference is ecological or an artifact of experimental design.
Composition
The commonality of community composition responses at the phylum level is striking and could suggest ecological trait conservation at the phylum level. Land use change is commonly associated with numerous changes to environmental factors (e.g., pH, P, K, etc.) and biotic factors (mixing of species, introduction of new species, altered dispersal barriers, etc.). The response of an organism to these changes could depend on traits such as pH tolerance, life history strategies, stress tolerance, or dispersal ability. While the response to land use change is not a direct measurement of a particular trait, it could be thought of as the encapsulation of many individual traits, similar to a polygenetic phenotype. Because traits, especially complex traits, tend to be conserved phylogenetically (Martiny et al., 2012, 2015), responses that are driven by complex combinations of traits should be conserved. A recent meta-analysis found Acidobacteria and Verrucomicrobia, for example, were consistently more competitive under oligotrophic conditions (Ho et al., 2017). We detected a consistent decrease in Acidobacteria relative abundance following conversion, a trend which may be related to soil C inputs or changes to the soil environment (Navarrete et al., 2015). We did not identify a consistent response in the Verrucomicrobia, but there was a non-significant decreasing trend (summary effect −0.15 ± 0.06, p = 0.058, n = 9). Verrucomicrobia also were not as commonly reported as other phyla. If land use conversion results in a more copiotrophic environment, it could explain the decreases in the abundances of these phyla as well as the increase in local diversity. Acidobacteria have often been correlated with soil pH, although subgroups respond differently, or even inconsistently, and it remains unclear if pH has a direct and causal relationship or covaries with other environmental factors (Kielak et al., 2016). Regardless of the postulated explanations, the conservation of responses we have found here, along with consistent declines in Acidobacteria and Proteobacteria, and increases in Actinobacteria, Firmicutes, and Chloroflexi, support the notion of phylum-level conservation of traits that mediate land use change responses.
Functional Consequences
Shifts in the diversity and composition of soil bacterial communities may have important implications for ecosystem function. Higher diversity levels have been associated with increased resistance, resilience, and/or stability of ecosystem function for a variety of aboveground ecosystems (Flynn et al., 2011; Hooper et al., 2012; Tilman et al., 2012). The generalizability of this trend, however, is an ongoing debate (Srivastava and Vellend, 2005; Cadotte et al., 2011). Community composition has also been reported to affect ecosystem function (Swenson et al., 2000a,b; Panke-Buisse et al., 2014). A recent meta-study found that incorporating microbial composition data with environmental data improves predictions of ecosystem process rates in 53% of cases (Graham et al., 2016), suggesting that widespread changes to bacterial diversity and composition could impact the functioning of ecosystems. However, we did not explicitly test this hypothesis in our study.
Recommendations for Future Research
The apparent commonality of bacterial responses warrants a more detailed examination, and we suggest several paths forward. There is a need to increase the geographic representation of key areas, especially the Congo River Basin. Our comparison is limited by the number of studies fitting our criteria. Tropical ecosystems in general have received considerably less attention from microbial ecologists than temperate areas (Pajares et al., 2016). Encouragingly, the application of next generation sequencing to the study of bacterial responses to land use change is an emerging area of research, with every study we compared published since 2012. Increasing the number of studies, particularly in the Congo Basin, where the first metabarcoding study in that region was recently conducted (Meyer et al., 2019), will allow more rigorous testing of the patterns we report, and would provide sufficient power to assess the importance of other characteristics, such as agricultural form and duration of agricultural use, on bacterial responses.
While studies such as the ones in this meta-analysis tend to be rare in higher latitude forests perhaps because many of these forests were converted by the mid-twentieth century (Houghton, 1994), a cross-biome comparison of forested and agricultural areas showed decreases in Acidobacteria and Proteobacteria and increases in Actinobacteria, Chloroflexi, and a number of other phyla (Crowther et al., 2014). Future work could ask whether conversion to agriculture elicits different bacterial responses across latitudes. This could help extend the generality of these responses beyond the tropics.
Inclusion of other soil organisms such as fungi, nematodes and microarthropods will allow for a more complete understanding of how soil ecosystems are affected by land use change (as in Kerfahi et al., 2016). Furthermore, simultaneous sampling of soil biota would allow the direct comparison of responses across large taxonomic groups; this could reveal fundamentally different responses to land use change across these groups. For example, while only five studies of tropical land use change to date have focused on fungi, four reported decreases in alpha diversity with conversion, suggesting that fungi may respond differently to ecosystem conversion than bacteria.
Finally, as more studies are conducted, increased standardization should be adopted for sampling protocols and reporting of findings. The studies assessed here span a breadth of protocols for sampling and a variety of metrics to assess diversity and composition. The generality of findings across this breadth points to the strength of the trends we observed. However, we were hampered in our ability to assess consistent changes in class level abundances and beta diversity due to low levels of reporting and by limited overlap in sampling protocols and analyses. Specifically, we suggest reporting GPS coordinates for every sample, reporting common diversity metrics (e.g., Shannon diversity or taxonomic richness), making OTU tables and raw sequences publicly available, and we recommend not pooling soil samples (so that spatially explicit analyses are possible). A more standardized set of collection and assessment methods will allow for a more conclusive and thorough analysis of the relationship between land use change and community attributes. If the above steps are taken, we believe it will lead to a much richer understanding of how land use change affects tropical soil communities.
Conclusions
We compared studies spanning a global longitudinal range, with representation from the three largest rainforests, each of which harbors vast biodiversity and high levels of endemism. The studies we compared varied in the primary form of agriculture, duration of agricultural use, and research methodology. Despite this variation, we identified consistent shifts in the diversity and composition of bacterial communities in response to the conversion of tropical rainforests to agriculture. Our findings suggest that land use conversion can have predictable effects at a pan-tropical level. With tropical rainforests undergoing dramatic and rapid reductions, it is important to consider the ramifications such changes will have on belowground soil communities and the ecosystem processes they mediate.
Data Availability Statement
The datasets for this manuscript are not publicly available because Data were drawn from multiple published studies, a full list including doi links is available with Supplementary Table 1. Requests to access the datasets should be directed to aWFuLmFiLnBldGVAZ21haWwuY29t.
Author Contributions
IP conducted the literature search, meta-analysis, and wrote the manuscript. IP, KM, and BB designed the study, contributed to manuscript revisions, and read and approved the final version.
Funding
This work was supported by the National Science Foundation—Dimensions of Biodiversity (DEB 14422214).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank the authors of the studies we analyzed, without which this meta-analysis would not be possible and thank the local and native peoples who helped in the completion of these studies and whose land was used for the purpose of these studies. This manuscript has been released as a Pre-Print at doi: 10.1101/600924 (Petersen et al., 2019).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fevo.2019.00391/full#supplementary-material
References
Banks, J. E., Sandvik, P., and Keesecker, L. (2009). Beetle (Coleoptera) and spider (Araneae) diversity in a mosaic of farmland, edge, and tropical forest habitats in western Costa Rica. Pan Pac. Entomol. 83, 152–160. doi: 10.3956/0031-0603-83.2.152
Bernard, H., Fjelds, Å. J, and Mohamed, M. (2009). A case study on the effects of disturbance and conversion of tropical lowland rain forest on the non-volant small mammals in north Borneo: management implications. Mamm. Study 34, 85–96. doi: 10.3106/041.034.0204
Borenstein, M., Hedges, L. V., Higgins, J. P. T., and Rothstein, H. R. (2009). Introduction to Meta-Analysis. Handbook of Set Theoretic Topology. doi: 10.1002/9780470743386
Borneman, J., and Triplett, E. W. (1997). Molecular microbial diversity in soils from eastern Amazonia: evidence for unusual microorganisms and microbial population shifts associated with deforestation. Appl. Environ. Microbiol. 63, 2647–2653.
Brook, B. W., Sodhl, N. S., and Ng, P. K. L. (2003). Catastrophic extinctions follow deforestation in Singapore. Nature 424, 420–423. doi: 10.1038/nature01795
Cadotte, M. W., Carscadden, K., and Mirotchnick, N. (2011). Beyond species: functional diversity and the maintenance of ecological processes and services. J. Appl. Ecol. 48, 1079–1087. doi: 10.1111/j.1365-2664.2011.02048.x
Cai, Z., Zhang, Y., Yang, C., and Wang, S. (2018). Land-use type strongly shapes community composition, but not always diversity of soil microbes in tropical China. Catena 165, 369–380. doi: 10.1016/j.catena.2018.02.018
Crowther, T. W., Maynard, D. S., Leff, J. W., Oldfield, E. E., Mcculley, R. L., Fierer, N., et al. (2014). Predicting the responsiveness of soil biodiversity to deforestation: a cross-biome study. Global Change Biol. 20, 2983–2994. doi: 10.1111/gcb.12565
Cuer, C. A., de Rodrigues, R. A. R., Balieiro, F. C., Jesus, J., Silva, E. P., Alves, B. J. R., et al. (2018). Short-term effect of Eucalyptus plantations on soil microbial communities and soil-atmosphere methane and nitrous oxide exchange. Sci. Rep. 8:15133. doi: 10.1038/s41598-018-33594-6
da C Jesus, E., Marsh, T. L., Tiedje, J. M., and de S Moreira, F. M. (2009). Changes in land use alter the structure of bacterial communities in Western Amazon soils. ISME J. 3, 1004–1011. doi: 10.1038/ismej.2009.98
Da Silveira, É. L., Pereira, R. M., Scaquitto, D. C., Pedrinho, E. A. N., Val-Moraes, S. P., Wickert, E., et al. (2006). Bacterial diversity of soil under eucalyptus assessed by 16S rDNA sequencing analysis. Pesqui. Agropecu. Bras. 41, 1507–1516. doi: 10.1590/S0100-204X2006001000008
de Carvalho, T. S., da C Jesus, E., Barlow, J., Gardner, T. A., Soares, I. C., Tiedje, J. M., et al. (2016). Land use intensification in the humid tropics increased both alpha and beta diversity of soil bacteria. Ecology 97, 2760–2771. doi: 10.1002/ecy.1513
De Moraes, J. F. L., Volkoff, B., Cerri, C. C., and Bernoux, M. (1996). Soil properties under Amazon forest and changes due to pasture installation in Rondônia, Brazil. Geoderma 70, 63–81. doi: 10.1016/0016-7061(95)00072-0
DeFries, R. S., Field, C. R., Fung, I., Collatz, G. J., and Bounoua, L. (1999). Combining satellite data and bioegeochemical models to estimate global effects of human-induced land cover change on carbon emissions and primary productivity. Global Biogeochem. Cycles 13, 803–815. doi: 10.1029/1999GB900037
Dias-Filho, M. B., Davidson, E. A., and de Cláudio, C. J. R. (2001). "Linking biogeochemical cycles to cattle pasture management and sustainability in the Amazon basin,” in The Biogeochemistry of the Amazon Basin, eds M. E. McClain, R. L. Victoria, and J. E. Richey (Oxford University Press), 84–105.
Dirzo, R., and Raven, P. H. (2003). Global state of biodiversity and loss. Annu. Rev. Environ. Res. 28, 137–167. doi: 10.1146/annurev.energy.28.050302.105532
Fierer, N., and Jackson, R. (2006). The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. U.S.A. 103, 626–631. doi: 10.1073/pnas.0507535103
Flynn, D. F. B., Mirotchnick, N., Jain, M., Palmer, M. I., and Naeem, S. (2011). Functional and phylogenetic diversity as predictors of biodiversity–ecosystem-function relationships. Ecology 92, 1573–1581. doi: 10.1890/10-1245.1
Gardner, T. A., Barlow, J., Chazdon, R., Ewers, R. M., Harvey, C. A., Peres, C. A., et al. (2009). Prospects for tropical forest biodiversity in a human-modified world. Ecol. Lett. 12, 561–582. doi: 10.1111/j.1461-0248.2009.01294.x
Gibbs, H. K., Ruesch, A. S., Achard, F., Clayton, M. K., Holmgren, P., Ramankutty, N., et al. (2010). Tropical forests were the primary sources of new agricultural land in the 1980s and 1990s. Proc. Natl. Acad. Sci. U.S.A. 107, 16732–16737. doi: 10.1073/pnas.0910275107
Gibson, L., Lee, T. M., Koh, L. P., Brook, B. W., Gardner, T. A., Barlow, J., et al. (2011). Primary forests are irreplaceable for sustaining tropical biodiversity. Nature 478, 378–381. doi: 10.1038/nature10425
Graham, E. B., Knelman, J. E., Schindlbacher, A., Siciliano, S., Breulmann, M., Yannarell, A., et al. (2016). Microbes as engines of ecosystem function: when does community structure enhance predictions of ecosystem processes? Front. Microbiol. 7:214. doi: 10.3389/fmicb.2016.00214
Hansen, M. C. C., Potapov, P. V., Moore, R., Hancher, M., Turubanova, S. A. A., Tyukavina, A., et al. (2013). High-resolution global maps of 21st-century forest cover change. Science 342, 850–854. doi: 10.1126/science.1244693
Hedges, L. V., Gurevitch, J., and Curtis, P. S. (1999). The meta-analysis of response ratios in experimental ecology. Ecology 80, 1150–1156. doi: 10.1890/0012-9658(1999)080[1150:TMAORR]2.0.CO;2
Ho, A., Lonardo, D. P., and Di Bodelier, P. L. E. (2017). Revisiting life strategy concepts in environmental microbial ecology. FEMS Microbiol. Ecol. 93:fix006. doi: 10.1093/femsec/fix006
Hooper, D. U., Adair, E. C., Cardinale, B. J., Byrnes, J. E. K., Hungate, B. A., Matulich, K. L., et al. (2012). A global synthesis reveals biodiversity loss as a major driver of ecosystem change. Nature 486, 105–108. doi: 10.1038/nature11118
Houghton, R. A. (1994). The worldwide extent of land-use change. BioScience 44, 305–313. doi: 10.2307/1312380
Hug, L. A., Baker, B. J., Anantharaman, K., Brown, C. T., Probst, A. J., Castelle, C. J., et al. (2016). A new view of the tree of life. Nat. Microbiol. 1:16048. doi: 10.1038/nmicrobiol.2016.48
Ibrahim, A. B., Wee, Y. C., Turner, I. M., Corlett, R. T., Chew, P. T., and Tan, H. T. W. (2003). A study of plant species extinction in Singapore: lessons for the conservation of tropical biodiversity. Conserv. Biol. 8, 705–712. doi: 10.1046/j.1523-1739.1994.08030705.x
Jesus, E. C., Marsh, T. L., Tiedje, J. M., and Moreira, F. M. (2009). Changes in land use alter the structure of bacterial communities in Western Amazon soils. ISME J. 3, 1004–1011. doi: 10.1038/ismej.2009.47
Kerfahi, D., Tripathi, B. M., Dong, K., Go, R., and Adams, J. M. (2016). Rainforest conversion to rubber plantation may not result in lower soil diversity of bacteria, fungi, and nematodes. Microb. Ecol. 72, 359–371. doi: 10.1007/s00248-016-0790-0
Kielak, A. M., Barreto, C. C., Kowalchuk, G. A., van Veen, J. A., and Kuramae, E. E. (2016). The ecology of Acidobacteria: moving beyond genes and genomes. Front. Microbiol. 7:744. doi: 10.3389/fmicb.2016.00744
Koh, L. P., and Wilcove, D. S. (2008). Is oil palm agriculture really destroying tropical biodiversity? Conserv. Lett. 1, 60–64. doi: 10.1111/j.1755-263X.2008.00011.x
Lajeunesse, M. J. (2015). Bias and correction for the log response ratio in ecological meta-analysis. Ecology 96, 2056–2063. doi: 10.1890/14-2402.1
Lan, G., Li, Y., Jatoi, M. T., Tan, Z., Wu, Z., and Xie, G. (2017). Change in soil microbial community compositions and diversity following the conversion of tropical forest to rubber plantations in Xishuangbanan, Southwest China. Trop. Conserv. Sci. 10, 1–14. doi: 10.1177/1940082917733230
Lauber, C. L., Hamady, M., Knight, R., and Fierer, N. (2009). Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 75, 5111–5120. doi: 10.1128/AEM.00335-09
Lim, B. K. (2017). “Lessons from Amazonia: The Ecology and Conservation of a Fragmented Forest” edited by Richard O. Bierregaard, Jr. et al. 2001. [book review]. Can. Field Nat. 117:672. doi: 10.22621/cfn.v117i4.794
Madsen, E. L. (2011). Microorganisms and their roles in fundamental biogeochemical cycles. Curr. Opin. Biotechnol. 22, 456–464. doi: 10.1016/j.copbio.2011.01.008
Martiny, A. C., Treseder, K., and Pusch, G. (2012). Phylogenetic conservatism of functional traits in microorganisms. ISME J. 7, 830–838. doi: 10.1038/ismej.2012.160
Martiny, J. B. H., Jones, S. E., Lennon, J. T., and Martiny, A. C. (2015). Microbiomes in light of traits: a phylogenetic perspective. Science 350:aac9323. doi: 10.1126/science.aac9323
McGuire, K. L., D'Angelo, H., Brearley, F. Q., Gedallovich, S. M., Babar, N., Yang, N., et al. (2015). Responses of soil fungi to logging and oil palm agriculture in Southeast Asian tropical forests. Microb. Ecol. 69, 733–747. doi: 10.1007/s00248-014-0468-4
Mendes, L. W., Tsai, S. M., Navarrete, A. A., de Hollander, M., van Veen, J. A., and Kuramae, E. E. (2015). Soil-borne microbiome: linking diversity to function. Microb. Ecol. 70, 255–265. doi: 10.1007/s00248-014-0559-2
Meyer, K. M., Petersen, I. A. B., Tobi, E., Korte, L., and Bohannan, B. J. M. (2019). Use of RNA and DNA to identify mechanisms of bacterial community homogenization. Front. Microbiol. 10:2066. doi: 10.3389/fmicb.2019.02066
Morton, D. C., DeFries, R. S., Shimabukuro, Y. E., Anderson, L. O., Arai, E., del Bon Espirito-Santo, F., et al. (2006). Cropland expansion changes deforestation dynamics in the southern Brazilian Amazon. Proc. Natl. Acad.Sci. U.S.A. 103, 14637–14641. doi: 10.1073/pnas.0606377103
Mueller, R. C., Paula, F. S., Mirza, B. S., Rodrigues, J. L. M., Nüsslein, K., and Bohannan, B. J. M. (2014). Links between plant and fungal communities across a deforestation chronosequence in the Amazon rainforest. ISME J. 8, 1548–1550. doi: 10.1038/ismej.2013.253
Navarrete, A. A., Venturini, A. M., Meyer, K. M., Klein, A. M., Tiedje, J. M., Brendan, B. J., et al. (2015). Differential response of Acidobacteria subgroups to forest-to-pasture conversion and their biogeographic patterns in the western Brazilian Amazon. Front. Microbiol. 6:1443. doi: 10.3389/fmicb.2015.01443
Neill, C., Piccolo, M. C., Cerri, C. C., Steudler, P. A., Melillo, J. M., and Brito, M. (1997). Net nitrogen mineralization and net nitrification rates in soils following deforestation for pasture across the southwestern Brazilian Amazon Basin landscape. Oecologia 110, 243–252. doi: 10.1007/s004420050157
Neill, C., Steudler, P. A., Garcia-Montiel, D. C., Melillo, J. M., Feigl, B. J., Piccolo, M. C., et al. (2005). Rates and controls of nitrous oxide and nitric oxide emissions following conversion of forest to pasture in Rondônia. Nutr. Cycl. Agroecosyst. 71, 1–15. doi: 10.1007/s10705-004-0378-9
Newbold, T., Hudson, L. N., Hill, S. L. L., Contu, S., Lysenko, I., Senior, R. A., et al. (2015). Global effects of land use on local terrestrial biodiversity. Nature 520, 45–50. doi: 10.1038/nature14324
Pajares, S., Bohannan, B. J. M., and Souza, V. (2016). Editorial: the role of microbial communities in tropical ecosystems. Front. Microbiol. 7:1805. doi: 10.3389/fmicb.2016.01805
Panke-Buisse, K., Poole, A. C., Goodrich, J. K., Ley, R. E., and Kao-Kniffin, J. (2014). Selection on soil microbiomes reveals reproducible impacts on plant function. ISME J. 9, 980–989. doi: 10.1038/ismej.2014.196
Pereira, R. M., da Silveira, É. L., Scaquitto, D. C., Pedrinho, E. A. N., Val-Moraes, S. P., Wickert, E., et al. (2006). Molecular characterization of bacterial populations of different soils. Braz. J. Microbiol. 37, 439–447. doi: 10.1590/S1517-83822006000400007
Perfecto, I., and Snelling, R. (1995). Biodiversity and the transformation of a tropical agroecosystem: ants in coffee plantations. Ecol. Appl. 5, 1084–1097. doi: 10.2307/2269356
Petersen, I. A., Meyer, K. M., and Bohannan, B. J. (2019). Consistent bacterial responses to land use change across the tropics. BioRxiv 600924. doi: 10.1101/600924
Pribyl, D. W. (2010). A critical review of the conventional SOC to SOM conversion factor. Geoderma 156, 75–83. doi: 10.1016/j.geoderma.2010.02.003
Rodrigues, J. L. M., Pellizari, V. H., Mueller, R., Baek, K., Jesus, E. D. C., Paula, F. S., et al. (2013). Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc. Natl. Acad. Sci. U.S.A. 110, 988–993. doi: 10.1073/pnas.1220608110
Sahoo, U. K., Singh, S. L., Gogoi, A., Kenye, A., and Sahoo, S. S. (2019). Active and passive soil organic carbon pools as affected by different land use types in Mizoram, Northeast India. PLoS ONE 14:e0219969. doi: 10.1371/journal.pone.0219969
Schneider, D., Engelhaupt, M., Allen, K., Kurniawan, S., Krashevska, V., Heinemann, M., et al. (2015). Impact of lowland rainforest transformation on diversity and composition of soil prokaryotic communities in sumatra (Indonesia). Front. Microbiol. 6:1339. doi: 10.3389/fmicb.2015.01339
Srivastava, D. S., and Vellend, M. (2005). Biodiversity-ecosystem function research: is it relevant to conservation? Annu. Rev. Ecol. Evol. Syst. 36, 267–294. doi: 10.1146/annurev.ecolsys.36.102003.152636
Sunderlin, W. D., Ndoye, O., Bikie, H., Laporte, N., Mertens, B., and Pokam, J. (2000). Economic crisis, small-scale agriculture, and forest cover change in southern Cameroon. Environ. Conserv. 27, 284–290. doi: 10.1017/S0376892900000321
Swenson, W., Arendt, J., and Wilson, D. S. (2000a). Artificial selection of microbial ecosystems for 3-chloroaniline biodegradation. Environ. Microbiol. 2, 564–571. doi: 10.1046/j.1462-2920.2000.00140.x
Swenson, W., Wilson, D. S., and Elias, R. (2000b). Artificial ecosystem selection. Proc. Natl. Acad. Sci. U.S.A. 97, 9110–9114. doi: 10.1073/pnas.150237597
Tilman, D., Reich, P. B., and Isbell, F. (2012). Biodiversity impacts ecosystem productivity as much as resources, disturbance, or herbivory. Proc. Natl. Acad. Sci. U.S.A. 109, 10394–10397. doi: 10.1073/pnas.1208240109
Tripathi, B. M., Edwards, D. P., Mendes, L. W., Kim, M., Dong, K., Kim, H., et al. (2016). The Impact of tropical forest logging and oil palm agriculture on the soil microbiome. Mol. Ecol. 25, 2244–2257. doi: 10.1111/mec.13620
Tripathi, B. M., Kim, M., Singh, D., Lee-Cruz, L., Lai-Hoe, A., Ainuddin, A. N., et al. (2012). Tropical soil bacterial communities in Malaysia: PH dominates in the Equatorial tropics too. Microb. Ecol. 64, 474–484. doi: 10.1007/s00248-012-0028-8
Verchot, L. V., Davidson, E. A., Cattânio, J. H., and Ackerman, I. L. (2000). Land-use change and biogeochemical controls of methane fluxes in soils of eastern Amazonia. Ecosystems 3, 41–56. doi: 10.1007/s100210000009
Zhang, Q., Justice, C. O., and Desanker, P. V. (2002). Impacts of simulated shifting cultivation on deforestation and the carbon stocks of the forests of central Africa. Agric. Ecosyst. Environ. 90, 203–209. doi: 10.1016/S0167-8809(01)00332-2
Keywords: land use change, bacteria, pan-tropical, meta-analysis, global, soil microbial ecology, rainforest
Citation: Petersen IAB, Meyer KM and Bohannan BJM (2019) Meta-Analysis Reveals Consistent Bacterial Responses to Land Use Change Across the Tropics. Front. Ecol. Evol. 7:391. doi: 10.3389/fevo.2019.00391
Received: 29 April 2019; Accepted: 01 October 2019;
Published: 16 October 2019.
Edited by:
Charles K. Lee, University of Waikato, New ZealandReviewed by:
Ederson Da Conceicao Jesus, Brazilian Agricultural Research Corporation (EMBRAPA), BrazilDavid James Van Horn, University of New Mexico, United States
Copyright © 2019 Petersen, Meyer and Bohannan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ian A. B. Petersen, aWFuLmFiLnBldGVAZ21haWwuY29t
 Ian A. B. Petersen
Ian A. B. Petersen Kyle M. Meyer
Kyle M. Meyer Brendan J. M. Bohannan
Brendan J. M. Bohannan