- 1G. Eliava Institute of Bacteriophages, Microbiology, and Virology, Tbilisi, Georgia
- 2United States Army Medical Research Institute of Infectious Diseases (USAMRIID), Fort Detrick, MD, USA
- 3Institute for Advanced Computer Studies, University of Maryland College Park, College Park, MD, USA
- 4Department of Cell Biology and Molecular Genetics, Maryland Pathogen Research Institute, University of Maryland College Park, College Park, MD, USA
Among the more than 70 different Vibrio species inhabiting marine, estuarine, and freshwater ecosystems, 12 are recognized as human pathogens. The warm subtropical climate of the Black Sea coastal area and inland regions of Georgia likely provides a favorable environment for various Vibrio species. From 2006 to 2009, the abundance, ecology, and diversity of clinically important Vibrio species were studied in different locations in Georgia and across seasons. Over a 33-month period, 1,595 presumptive Vibrio isolates were collected from the Black Sea (n = 657) and freshwater lakes around Tbilisi (n = 938). Screening of a subset of 440 concentrated and enriched water samples by PCR-electrospray ionization/mass spectrometry (PCR-ESI/MS) detected the presence of DNA from eight clinically important Vibrio species: V. cholerae, V. parahaemolyticus, V. vulnificus, V. mimicus, V. alginolyticus, V. harveyi, V. metschnikovii, and V. cincinnatiensis. Almost 90% of PCR/ESI-MS samples positive for Vibrio species were collected from June through November. Three important human-pathogenic Vibrio species (V. cholerae, V. parahaemolyticus, and V. vulnificus) were detected in 62.8, 37.8, and 21.4% of samples testing positive for Vibrios, respectively. The results of these activities suggest that natural reservoirs for human-pathogenic Vibrios exist in Georgian aquatic environments. Water temperature at all sampling sites was positively correlated with the abundance of clinically important Vibrio spp. (except V. metschnikovii), and salinity was correlated with species composition at particular Black Sea sites as well as inland reservoirs.
Introduction
Vibrio species are ubiquitous and abundant in aquatic environments. They exist suspended in the water column, attached to plankton, and in the tissues or organs of various marine animals (1). The genus Vibrio includes more than 70 species, which are characterized as halophilic or non-halophilic according to their need for sodium chloride for growth (2, 3). At least 12 Vibrio species, including V. cholerae, V. parahaemolyticus, and V. vulnificus, are pathogenic for humans and important to public health (4–7).
Among more than 200 known V. cholerae serotypes, serotypes O1 and O139 have epidemic and clinical importance. These serotypes are the causative agents of cholera, a severe diarrheal disease with 50–60% mortality in untreated cases. Cholera is contracted through consumption of contaminated food and water. Non-O1 and non-O139 Vibrio serotypes (also called non-agglutinable or NAG, serotypes) have also been implicated as etiologic agents of moderate to severe gastroenteritis in humans (2, 8, 9). V. parahaemolyticus, a halophilic Vibrio species, mainly causes gastroenteritis associated with consumption of contaminated seafood. In the last decade, epidemics of gastroenteritis in Southeast Asia, Japan, and North America have been linked to V. parahaemolyticus pandemic serotype O3:K6 (10, 11). Wound infections and septicemia caused by V. vulnificus carry a case fatality rate (CFR) as high as 50% in healthy patients; the CFR of V. vulnificus infections is even higher among immuno-compromised patients or those with liver disease (6). V. metschnikovii can cause gastroenteritis and wound infections leading to septicemia, while V. fluvialis, Grimontia hollisae (formerly V. hollisae), and V. furnissii typically cause gastroenteritis (12, 13). V. alginolyticus is commonly associated with ear infections, but can also cause respiratory infections and bacteremia (14, 15). Photobacterium damselae subsp. damselae (formerly V. damsela) causes opportunistic wound infections that can rapidly progress to a necrotizing condition, and ultimately to sepsis, which is fatal in 30–50% of human cases (16). A few cases of human illness (e.g., septicemia, meningitis, and diarrhea) have been associated with V. cincinnatiensis (17). V. alginolyticus, P. damselae, and V. harveyi are also opportunistic pathogens of economic significance in aquaculture, responsible for high mortality in cultured fish and shellfish, sometimes destroying an entire aquaculture operation (18).
Prevention and control of infections caused by Vibrio species pathogenic for humans depend on understanding their ecology, pathogenicity, and modes of transmission. A warm, subtropical climate – such as in the Black Sea coastal region of the South Caucasus – may support growth and multiplication of most Vibrio species (2, 19–21). A limited number of short reports are available on the abundance of NAG Vibrio species (more specifically, V. parahaemolyticus and V. alginolyticus) in some coastal areas of the Black Sea (9, 22–24). In addition, environmental surveillance studies have been performed to fill gaps in the understanding of the ecology of V. cholerae and other Vibrio species of public health importance in Georgia and Azerbaijan (25–29). However, the full range of disease-causing Vibrio species in aquatic environments in this region has never been comprehensively examined.
The Black Sea coastal zones of Georgia and water reservoirs around Tbilisi have traditionally been popular recreational zones; the number of international visitors to the country has drastically increased in the last decade. The combination of climate change (in particular, elevated air and surface water temperatures) and the increasing anthropogenic effects of tourism may increase the risk of emergence and spread of water-borne and food-borne infections (30). An increase in the frequency of enteric diseases with diarrhea has been registered in Georgia, especially in the Ajara region, which is near the Black Sea sampling sites used in this study (31); a significant portion of these infections with idiopathic etiology may have been caused by pathogenic Vibrio species.
In light of these factors, the aim of this study was to conduct environmental surveillance to assess the abundance and diversity of clinically important Vibrio species along the Georgian coastal zone of the Black Sea and in freshwater reservoirs near Tbilisi. The study also aimed to evaluate the efficacy of conventional methods used by Georgian public health laboratories to detect and identify pathogenic Vibrios in the aquatic environment. To our knowledge, this report presents the first detailed description of environmental parameters in relation to the prevalence and community structure of pathogenic Vibrio species in recreational waters in Georgia.
Materials and Methods
Sample Sites and Collection
From June 2006 through October 2008, surface water samples were collected bi-weekly from July through September and monthly the rest of the year. Water samples were collected along the Georgian Black Sea coastal zone at four permanent stations located 50–100 m from the shore. Sample sites included two estuaries: Supsa (N 42°00.008′ E 41° 41.01′) and Chorokhi (N 41°36.116′ E 41° 34.021′); and two popular recreational/tourist attraction areas: Green Cape (N 41°41.91′ E 41° 42.01′) and Batumi Boulevard (N 41°39.570′ E 41°38.006′). Water samples were also collected 2–5 m from the shore of three inland reservoirs around Tbilisi: Kumisi Lake (Site 1: N 41°35.153′ E 044°51.591′; Site 2: N 41°34.839′ E 044°51.304′); Lisi Lake (Site 1: N 41°44.440′ E 044°44.261′; Site 2: N 41°44.483′ E 044°44.326′); and Tbilisi Sea (Site 1: N 41°46.150′ E 044°48.904′; Site 2: N 41°45.765′ E 044°50.308′). Sample locations are displayed in Figure S1 in Supplementary Material with coordinates listed in Table S1 in Supplementary Material.
Temperature, salinity, pH, conductivity, dissolved oxygen, and total dissolved solids were measured using a portable multi-log environmental meter (YSI Model 556 MPS, Yellow Springs, OH, USA), according to manufacturer’s instructions. The physicochemical parameters of each sample were measured in triplicate, and the mean was recorded for laboratory analysis; seasonal temperature and salinity measurements for the sampling sites are given in Table S2 in Supplementary Material. At the time of sample collection at each site, 100 L of water was filtered through plankton nets (200, 64, and 10-μm mesh size) to collect concentrated plankton samples and plankton-free water (32, 33).
Bacterial Strains
Five Vibrio strains were obtained from the collection of the Institute Pasteur (CIP): V. cholerae CIP 62.13; V. cholerae CIP 55.91 (O1 classical); V. cholerae CIP 106855 (O1 El Tor); V. cholerae CIP 104151 (O139); and V. natriegens CIP 103193.
The National Center for Disease Control (NCDC) in Tbilisi, Georgia kindly provided two strains of V. cholerae: V. cholerae N145/P-1 (O1 classical) and V. cholerae N890/M-878 (O1 El Tor).
The following reference strains were obtained from the American Type Culture Collection (ATCC): V. vulnificus ATCC 27562; V. parahaemolyticus ATCC 17802; V. metschnikovii ATCC 700040; V. fluvialis ATCC 33809; V. furnissii ATCC 35016; V. alginolyticus ATCC 17749 serotype XII; Photobacterium damselae subsp. damselae ATCC 33539; Grimontia (Vibrio) hollisae ATCC 33564; V. cincinnatiensis ATCC 35912; and V. harveyi ATCC 14126.
Isolation of Vibrio Species
Different volumes of plankton-free water (10–100 mL) were passed through 0.45-μm membrane filters to concentrate bacteria. The filters were placed onto thiosulfate-citrate-bile salts-sucrose (TCBS) agar, and incubated at 35°C for 24 h. After incubation, presumptive Vibrio colonies were counted. Filters were also incubated in 1% alkaline peptone water (APW), pH 8.6, for 18 h at 35°C to enhance growth of Vibrios. After incubation, a sterile inoculating loop was used to streak samples onto TCBS agar plates. Similarly, concentrated plankton samples were spread onto TCBS agar plates and/or inoculated in APW concentrate Vibrios. Plates were then incubated at 35–37°C for 24 h. Presumptive Vibrio colonies (yellow, green, and olive-green) that grew on TCBS plates were counted and sub-cultured onto T1N1 (1% trypticase, 1% sodium chloride) agar plates.
Biochemical Identification of Vibrio Species Isolates
Presumptive Vibrio isolates were sub-cultured onto Luria-Bertani agar and screened for the following biochemical properties: gelatinase production; oxidase activity; salt requirement/tolerance (growth in T1N1–T1N8 solutions); glucose oxidation/fermentation; arginine dehydrolase utilization; lysine decarboxylase utilization; and utilization of sucrose, arabinose, lactose, and mannose (4, 32).
Biochemical identification parameters were analyzed using a software algorithm designed for this study. The algorithm was used to compare biochemical tests’ results for presumptive Vibrio isolates with those of 12 clinically important, well-characterized, standard strains of Vibrio species. The algorithm also included existing data on the physiological and biochemical properties of various Vibrio species (4, 33, 34). If the properties of a Vibrio isolate were similar to those of a reference strain, a percentage of affinity was calculated using different weighted factors for the particular biochemical parameter(s) (P, [P]).
Identification of Vibrio Species Isolated by PCR
Bacterial DNA was extracted using an AquaPure genomic DNA isolation kit (Bio-Rad Laboratories, Hercules, CA, USA) according to manufacturer’s instructions. Species-specific PCR (internal transcribed spacer PCR) and collagenase-targeted PCR were used to confirm the identification of presumptive V. cholerae, V. mimicus, V. parahaemolyticus, V. alginolyticus, and V. vulnificus isolates (32, 33).
To determine the serogroup (O1 or O139) of V. cholerae isolates, a multiplex PCR was performed as described by Huq et al. (32, 33). The assay was also used to detect virulence factor genes (e.g., ctxA) in confirmed V. cholerae isolates. PCR primers and target genes used to detect clinically important Vibrio species are listed in Table S3 in Supplementary Material.
For identification of all other presumptive Vibrio species not identified by species-specific PCR, 16S rRNA was amplified and sequenced, and the sequences were compared to corresponding data in NCBI and RDPII databases (33).
Direct Detection of Vibrio Species Isolates by PCR/ESI-MS
Vibrio species were directly detected in water samples with a Vibrio-specific PCR-electrospray ionization (ESI)/mass spectrometry (MS) assay (26). Briefly, total community DNA was amplified from water samples using an eight-reaction broad-range PCR assay targeted to members of the Vibrio genus and Vibrionaceae family. After PCR, a purified aliquot from each reaction was sprayed into a Bruker Daltonics microTOF (Billerica, MA, USA) mass spectrometer. Because of the high mass accuracy (mass measurement error <1 ppm) of the spectrometer, the mass of each PCR amplicon could be accurately determined and a base composition could be assigned with confidence (e.g., xA, xT, xC, and xG) (35). Because the assay included eight primers, multiple base counts were assigned to each sample from various parts of the genome. For confirmatory identification, base compositions of the samples were added to the Ibis database (National Center for Biotechnology Information, Bethesda, MD, USA) for comparison with base compositions of Vibrio reference strains and related bacteria.
Direct Fluorescent Antibody Test
A direct fluorescent antibody (DFA) test was used to detect V. cholerae O1 and O139 in enriched and concentrated water and plankton samples (32, 33). Preparation of specimens was performed using a DFA test kit (New Horizon Diagnostics, Columbia, MD, USA) according to the manufacturer’s instructions. Specimens were examined with an epi-fluorescent microscope (Axioskop 40, Opton Zeiss, Germany) at 100× magnification.
Data Analysis
Each water sample was tested in triplicate for all biochemical tests outlined above, and the mean values and standard errors were calculated for each variable on a given sampling date. For the purpose of analysis, seasons were defined as follows:
• Winter: December through February;
• Spring: March through May;
• Summer: June through August;
• Fall: September through November.
Statistical analysis was carried out using the Statistical Toolpak for Microsoft Excel 2010. Correlations (Pearson’s r) between factors described herein were significant at the 0.05 level.
Results and Discussion
Isolation and Phenotypic Characterization of Clinically Important Vibrio Species
During the study period, 1,440 water samples were collected and analyzed; from these, 1,595 presumptive Vibrio isolates were collected. Among these samples, 657 presumptive Vibrio isolates were collected from four sites on the Black Sea coast and 938 from three inland water reservoirs. Over 70% of the isolates were assigned to 10 clinically important Vibrio species based on biochemical characteristics. These isolates were recovered from concentrated and enriched water samples, as well as from plankton. Of the 1,595 isolates, 856 were identified as presumptive non-halophilic Vibrios (specifically, V. cholerae and V. mimicus) and 739 were identified as halophilic Vibrio species. As expected, the Black Sea Vibrio population was most the diverse: nine species of clinically important Vibrios were recovered (V. parahaemolyticus, V. vulnificus, V. cholerae, V. metschnikovii, V. alginolyticus, V. harveyi, V. furnissii, V. fluvialis, and V. cincinnatiensis). Only seven Vibrio species (V. cholerae, V. vulnificus, V. mimicus, V. metschnikovii, V. alginolyticus, V. harveyi, and V. fluvialis) were collected from the inland water reservoirs. In general, halophilic Vibrio species were more commonly found in the Black Sea coastal zones (47.8%), than in freshwater lakes (6.2%). Among the halophilic Vibrio species, V. parahaemolyticus was the most prevalent. V. cholerae comprised more than 65% of isolates from inland reservoirs, and 23% of those from Black Sea sites. It should be noted that up to 28% of isolates were identified only to genus level. In a separate study by Mitaishvili et al., some of these isolates were identified as non-human-pathogenic Vibrio species, including V. natriegens, V. splendidus, and V. estuarinus (36).
PCR Identification of Vibrio Species
To establish the diversity of the Vibrio species more accurately and also for comparison purposes, a subset of 274 halophilic Vibrio isolates (attributed to eight Vibrio species, mostly V. parahaemolyticus) and 520 non-halophilic Vibrio isolates (512 V. cholerae and 8 V. mimicus) were subjected to either PCR with species-specific primers; or amplification, sequencing, and comparison of 16S rRNA signatures, employing data available in public databases. A high level of agreement (98.5%, or 503 isolates out of 512) was observed between standard biochemical and PCR identifications for V. cholerae. Similar results were observed among 144 isolates of V. parahaemolyticus (95.8%).
The agreement between the two methods was relatively low (38.5–46.5%) for other Vibrio species. For example, some of the V. vulnificus isolates presumptively identified based on biochemical analysis were found to be V. cholerae by PCR. The majority of presumptive V. alginolyticus and V. cincinnatiensis isolates were identified as V. parahaemolyticus by PCR. Additionally, several isolates of V. fluvialis identified by biochemical tests were determined to be V. furnissii by genetic analysis. These results most likely are due to the significant genetic variability of biochemical features (e.g., salt tolerance, sucrose, lactose, and arabinose utilization) and especially of the halophilic Vibrio species.
Among 794 presumptive Vibrio isolates, 10 of the clinically important Vibrio species were genetically identified. The distribution of species identified by biochemical analysis was different than that indicated by genetic testing. Among PCR-confirmed Vibrio isolates, V. cholerae was most prevalent (64.6%) among tested isolates, followed by V. parahaemolyticus (21.4%), V. vulnificus (4.0%), and V. alginolyticus (2.6%). Combined, the remaining six species (V. metschnikovii, V. harveyi, V. furnissii, V. fluvialis, V. cincinnatiensis, and V. mimicus) accounted for 4.4% of samples. Genetic identification was not definitive for 3% of the Vibrio isolates. The distributions of the 10 Vibrio species confirmed by PCR are presented in Figure 1.
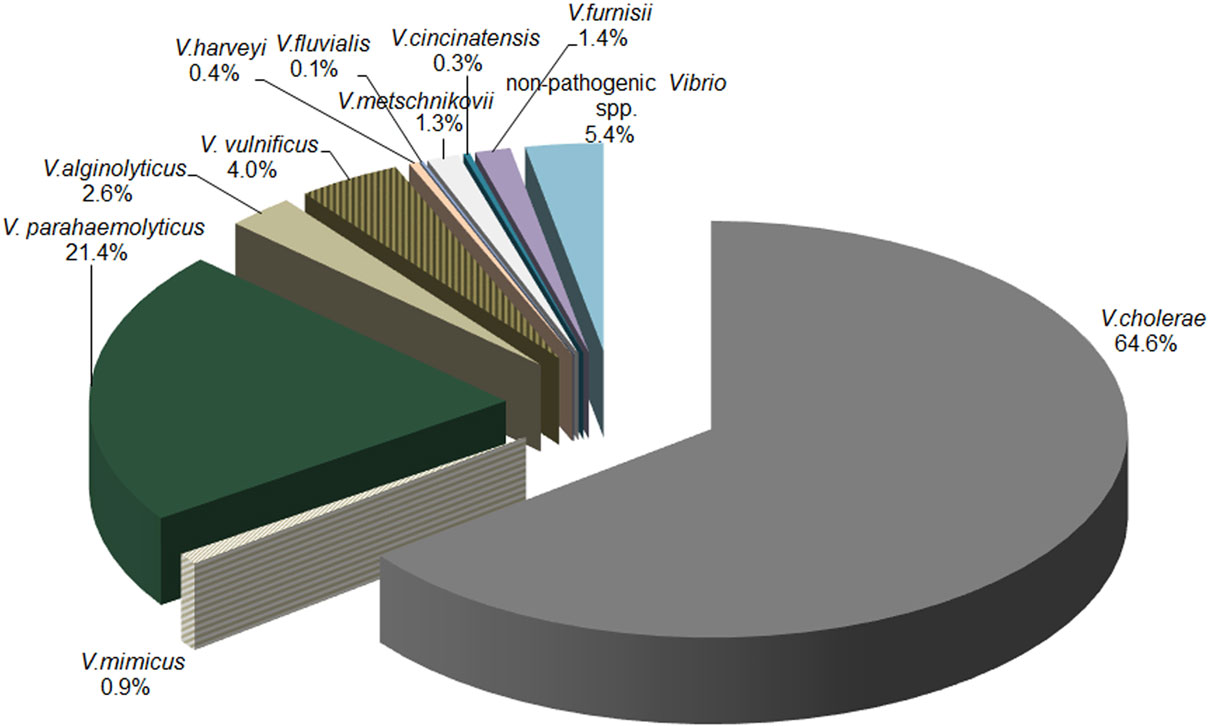
Figure 1. Distribution of Vibrio species via PCR. The distribution of Vibrio species detected by PCR in water samples collected across Georgia.
Direct Detection of Clinically Important Vibrio Species in Water Samples
In parallel with culture isolation, Vibrio species were directly detected in water samples collected during 2006–2008 from marine and freshwater sites (n = 440 samples; 220 from the Black Sea and 220 from lakes) using a PCR/ESI-MS method verified by Whitehouse et al. in a previous study employing standard and environmental Vibrio strains (26). Eight clinically important Vibrio species were detected by this method. Six of the eight Vibrio species were halophilic (V. parahaemolyticus, V. vulnificus, V. alginolyticus, V. harveyi, V. metschnikovii, and V. cincinnatiensis); the remaining two Vibrio species were non-halophilic (V. cholerae and V. mimicus), that is, not requiring additional salt for growth (all Vibrios have an absolute requirement for NaCl for growth). Two non-pathogenic Vibrio species [V. natriegens and V. (Comamonas) neocistes] were also detected.
The distribution of samples positive for specific Vibrio species by PCR/ESI-MS is presented in Figure 2. In alignment with culture results, V. cholerae was the most prevalent pathogenic Vibrio species. In water samples containing the DNA of at least one Vibrio species, 62.8% contained V. cholerae (n = 304).
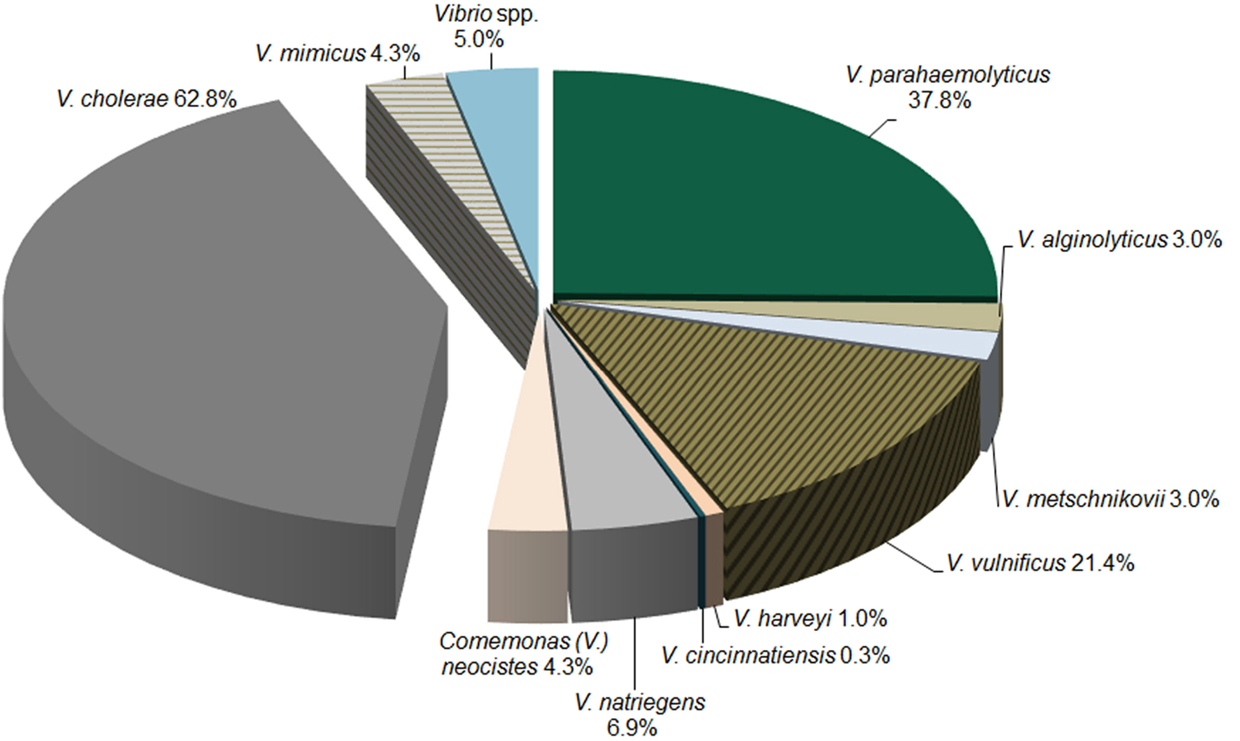
Figure 2. Distribution of Vibrio species via PCR/ESI-MS. The distribution of Vibrio species detected by PCR/ESI-MS in water samples collected across Georgia.
Population diversity of Black Sea Vibrio species as determined by PCR/ESI-MS was similar to that observed among biochemically identified species (Figure 3A). Halophilic Vibrio species were more abundant in samples collected at Black Sea sites (72.6% of all Vibrio-positive samples) than those from freshwater lakes (29.1%); this is in agreement with culture results. Among Vibrio-positive Black Sea water samples (n = 153), V. parahaemolyticus was most frequently detected (53.6%), followed by V. cholerae and V. vulnificus (48.4 and 36.6%, respectively). Other Vibrio species (V. alginolyticus, V. metschnikovii, V. fluvialis, and V. harveyi) comprised between 0.7 and 3.9% of those identified in Black Sea samples.
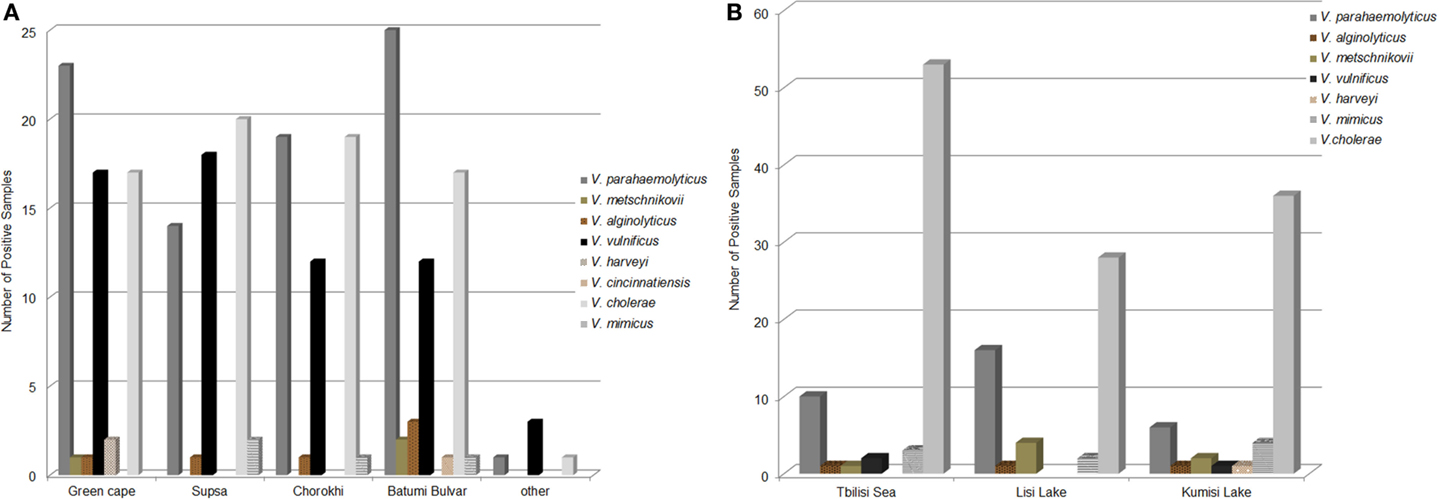
Figure 3. Abundance of Vibrio species in Black Sea and lake water samples. The abundance of Vibrio species in water samples as determined by PCR/ESI-MS from (A) The Black Sea and (B) Freshwater and brackish lakes.
Of the seven Vibrio species identified by conventional culture, six were detected in fresh and brackish lake samples, using PCR/ESI-MS (Figure 3B). V. cholerae was most frequently isolated from water samples collected at all three inland reservoirs (77.5% of all positive samples). DNA from other Vibrio spp. was detected less frequently, including: V. parahaemolyticus (21.2%), V. mimicus (6.0%), V. metschnikovii (4.6%), and V. vulnificus (2.0%). Among the lake samples, abundance and diversity of Vibrio species were highest in Kumisi Lake, with six species identified by phenotypic characterization and five by PCR. This diversity is most likely related to the relatively high salinity of the lake.
Also, in agreement with results of our previously published study, 20.7% of Black Sea and lake samples contained V. natriegens and/or V. neocistes. Also as observed in our previous report, V. natriegens was cultured from all four Black Sea sites along with other Vibrio species that are not pathogenic for humans (36).
Seasonality in Abundance of Clinically Important Vibrio Species
Seasonal patterns in abundance of clinically important Vibrio species were reflected in the isolation frequency and results of PCR/ESI-MS. Ninety-five percent of Vibrio spp. were isolated from April through November, with the highest detection rates in July, August, and September (Figures 4A,B); water temperatures for these time periods (per location) are reported in Table S2 in Supplementary Material. In our earlier reports, we described a positive correlation between total Vibrio counts and water temperature at Black Sea and inland lake sites (37, 38). In addition, in the current study, we observed a correlation between isolation frequency of distinct groups of Vibrio species and water temperature at marine and freshwater sites. Interestingly, in this study, the correlation coefficient was higher for halophilic Vibrio species (r = 0.85 for Black Sea sites and r = 0.74 for lakes) than for V. cholerae (r = 0.45 for Black Sea sites and r = 0.64 for lakes). A different pattern of seasonal abundance was observed for V. metschnikovii isolates, which were collected during the cold season and were detected by PCR/ESI-MS from October through April (Figures 4A,B). This is indicative of cold tolerance of V. metschnikovii in comparison with the other pathogenic Vibrio species (39, 40).
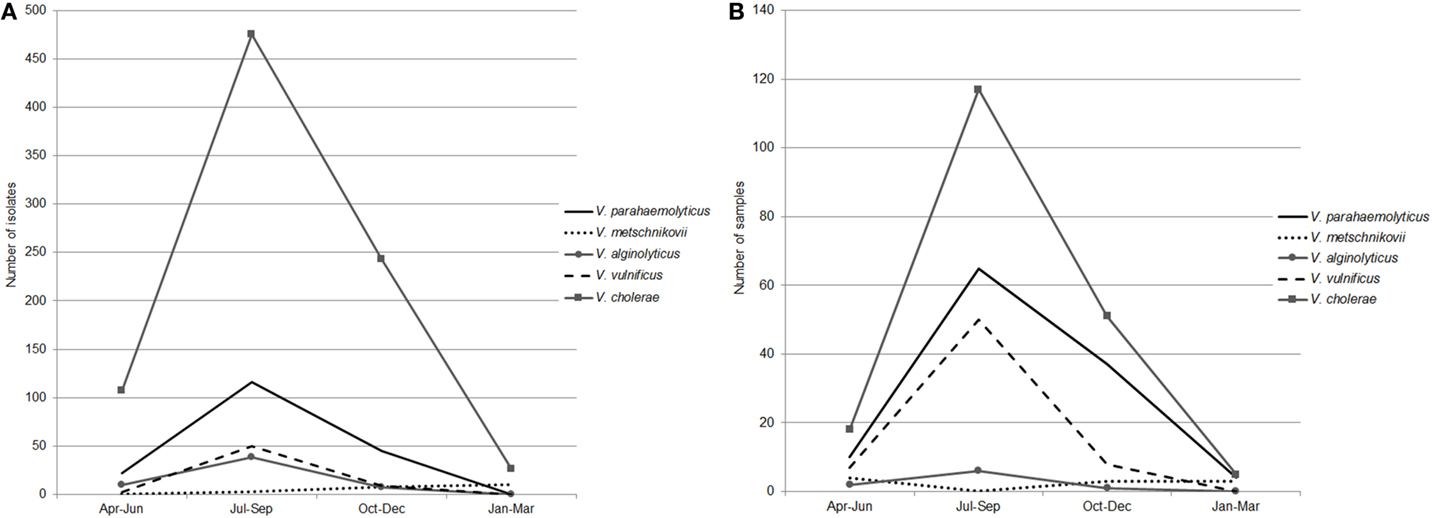
Figure 4. Seasonal abundance of five Vibrio species. The seasonal abundance of five Vibrio species (V. parahaemolyticus, V. metschnikovii, V. alginolyticus, V. vulnificus, and V. cholerae) as determined by (A) Biochemical identification and, (B) PCR/ESI-MS. Seasonal water temperature is reported in Table S2 in Supplementary Material.
Salinity as an Environmental Factor Influencing Vibrio Community Composition
Salinity, an important environmental factor, can influence microbial community composition and abundance in aquatic environments (1, 2, 7). In an earlier published study, we reported no significant correlation between total Vibrio counts and salinity for Black Sea samples; instead, species diversity (by culturing and direct detection techniques) appears to be linked to salinity (37). For example, PCR/ESI-MS direct detection (Figure 3A) showed less diversity among Vibrio species in the Supsa and Chorokhi estuaries (five species at each site) than at the Green Cape and Batumi Boulevard sites (six and seven species, respectively). In the samples collected from Batumi Boulevard sites where the salinity is higher (16–20%), V. parahaemolyticus DNA was detected more frequently by PCR/ESI-MS than in samples from the Supsa estuary (salinity 3–10.6%), although a similar pattern of seasonal temperature dependence was observed at both Black Sea sampling stations (Figures 5A,B).
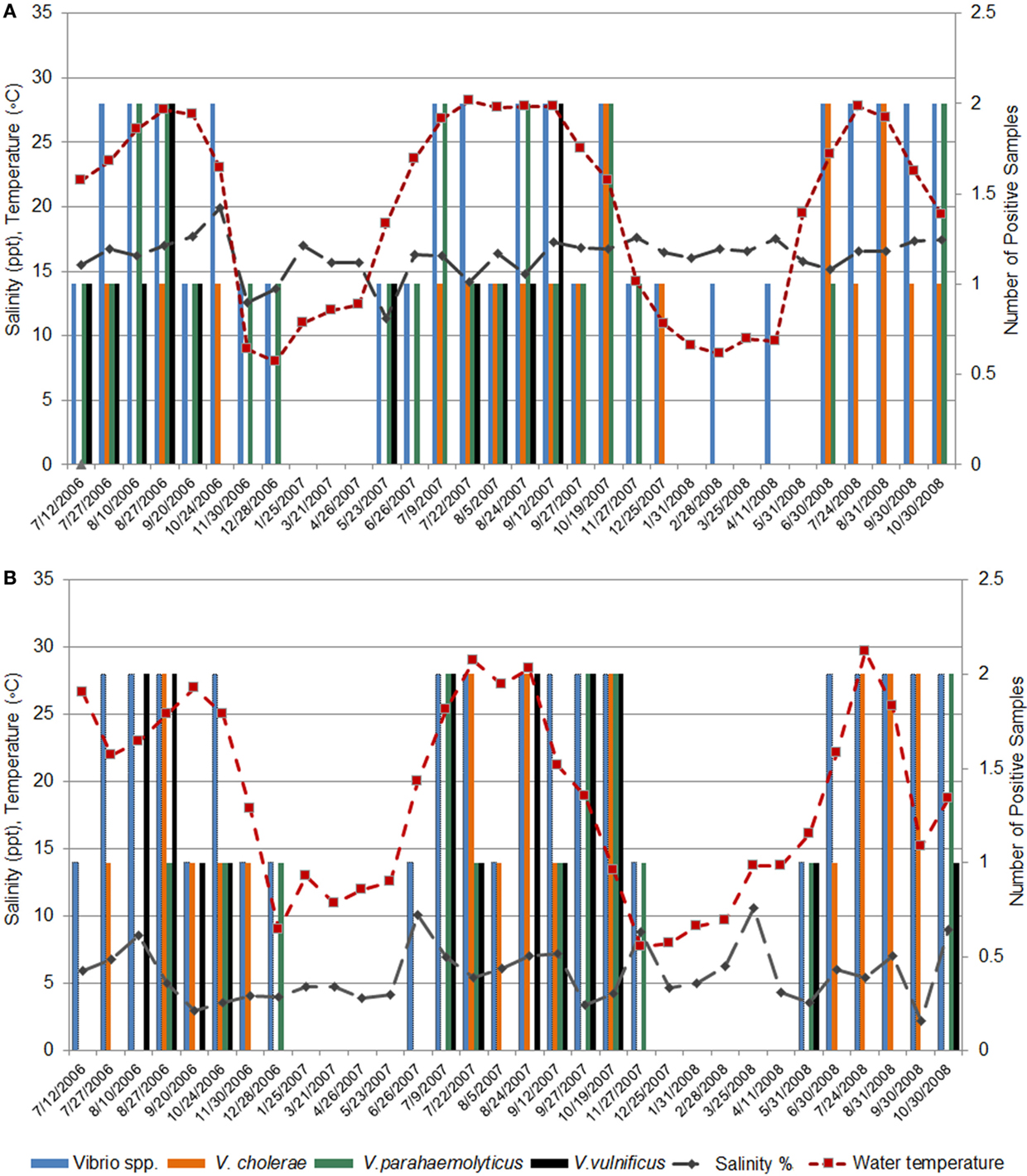
Figure 5. Influence of water temperature on the detection of Vibrio species. The influence of water temperature on the overall detection rates of Vibrio species by PCR/ESI-MS for (A) Batumi Boulevard, (B) Supsa estuary.
Among inland reservoirs, a more diverse Vibrio population was detected by both culture and PCR/ESI-MS in Kumisi Lake (five clinically important Vibrio species in total) than the other two lakes (Figure 3B). It was surprising to detect a number of pathogenic halophilic Vibrio species in inland lakes, where non-halophilic Vibrios were predicted (and ultimately isolated) to be more prevalent; for example, V. alginolyticus was detected in these reservoirs, and was most abundant in the Tbilisi Sea (average salinity of 0.1%). This may be explained by the proximity of this artificially created reservoir (fed by the Iori River) to three small saltwater lakes that existed in the area before the creation of the Tbilisi Sea in the 1950s (41).
V. cholerae: A Prevalent Human-Pathogenic Vibrio in Georgian Aquatic Environments
The most important human-pathogenic Vibrio species, V. cholerae, was demonstrated by culture and direct detection to be prevalent in Georgian aquatic environments. According to PCR/ESI-MS, V. cholerae was present in the majority of lake samples (77.5% of positive samples). V. cholerae was detected less frequently in Black Sea samples (48.4%). This aligned with culture results, among which V. cholerae comprised 23 and 75.2% of Vibrio species isolates from the Black Sea and lake samples, respectively. According to our earlier report, the vast majority (94.6%) of Georgian isolates of V. cholerae were attributed to the non-O1/non-O139 group (28). These groups are generally non-pathogenic to humans, although some serotypes can cause mild to severe gastroenteritis (8, 9).
Epidemic V. cholerae serotype O1 was directly detected by DFA in the concentrated and enriched water and plankton samples collected at all four sites in the coastal zone of the Black Sea during the study period (Figure 6). Toxigenic V. cholerae O1 was also detected in lake samples (specifically from Kumisi Lake and the Tbilisi Sea) collected in 2008 and 2009 (Figure 6). In addition, signals indicative of V. cholerae O139 were detected in eight marine samples collected in the summer of 2008. These data, in combination with results of our earlier DFA investigations on lake samples collected in 2006 and 2007, indicate that toxigenic V. cholerae is common in all of the target water bodies in this study (25). These serotypes were detected most frequently in Black Sea samples and often associated with plankton fractions (Figure 7). This is in agreement with previously published serological data and multiplex PCR data from other investigators studying V. cholerae (28). Forty-six isolates (most of which were collected from enriched marine samples) were confirmed as V. cholerae serotype O1 and six isolates were presumptively identified as V. cholerae serotype O139. Some of the V. cholerae O1 strains belonged to the El Tor biotype, while others revealed characteristics of hybrid variants. Two O1 and six non-O1 strains carried ctx genes.
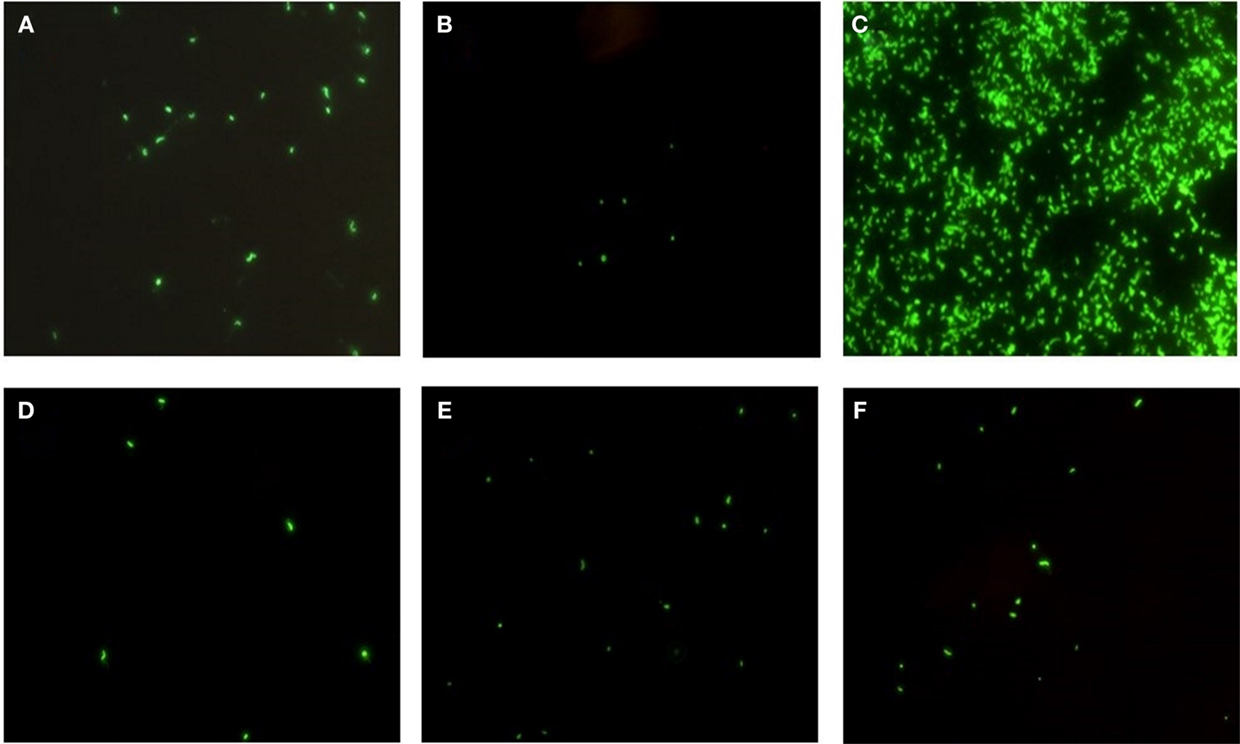
Figure 6. Detection of V. cholerae O1 by DFA in Black Sea and lake samples. Detection of V. cholerae O1 in water samples from (A–C) Green Cape sample sites on the Black Sea, and (D–F) Kumisi Lake. Concentrated water samples are shown in (A,D), plankton samples in (B,E) and enriched water samples in (C,F). Organisms were detected and visualized using a direct fluorescent-monoclonal antibody (DFA) kit.
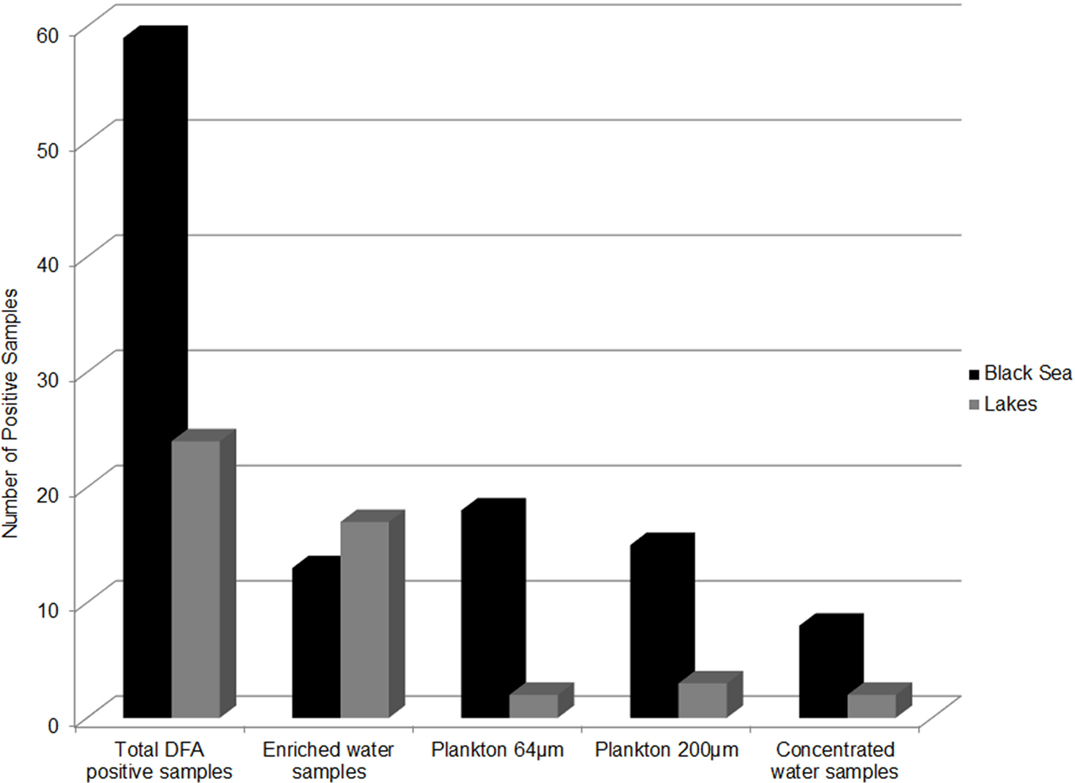
Figure 7. Distribution of V. cholerae O1 DFA positive samples. The distribution by sample type and sampling site of V. cholerae O1 positive samples as detected by DFA.
Interestingly, V. cholerae was observed along with two other important Vibrio species in the same ecological niche: V. parahaemolyticus (33.1% of PCR-ESI/MS samples) and V. vulnificus (11.6%). The increased probability of co-mingling between V. cholerae and these important Vibrio species could facilitate transfer of genetic material (e.g., virulence factors) between the species.
Comparative Ecology of V. parahaemolyticus in Marine and Lake Reservoirs
The second most important human-pathogenic Vibrio species, V. parahaemolyticus, was detected by PCR/ESI-MS in samples from the Black Sea. Most surprisingly, V. parahaemolyticus was detected in all sampled lakes, but no confirmed V. parahaemolyticus isolates were recovered from these sites. Interestingly, V. parahaemolyticus was more frequently detected by PCR/ESI-MS in concentrated lake samples than enriched lake samples; the reverse of this was observed for Black Sea samples (Figure 8). For comparison, as expected, DNA from V. cholerae was found most frequently in enriched lake samples, and in equal quantities in enriched and concentrated marine samples. The abundance of V. parahaemolyticus DNA in fresh and brackish water sites is likely explained by the lower salinity of lake water, which is a stressor for V. parahaemolyticus, a moderately halophilic bacterium. The low frequency of isolation of V. parahaemolyticus from lake water, coupled with the relatively high frequency of direct molecular detection, suggests that the species may be present in a viable but non-culturable state. Isolation of V. parahaemolyticus-specific bacteriophages from Lake Kumisi and Lake Lisi samples also supports the conclusion that this Vibrio species is present (42). The possibility of V. parahaemolyticus existing in a transitional state in freshwater reservoirs, in association with freshwater fish, has been proposed by Sarkar et al. and other researchers and is important, as this may serve as a potential exposure route to the bacterium (43, 44).
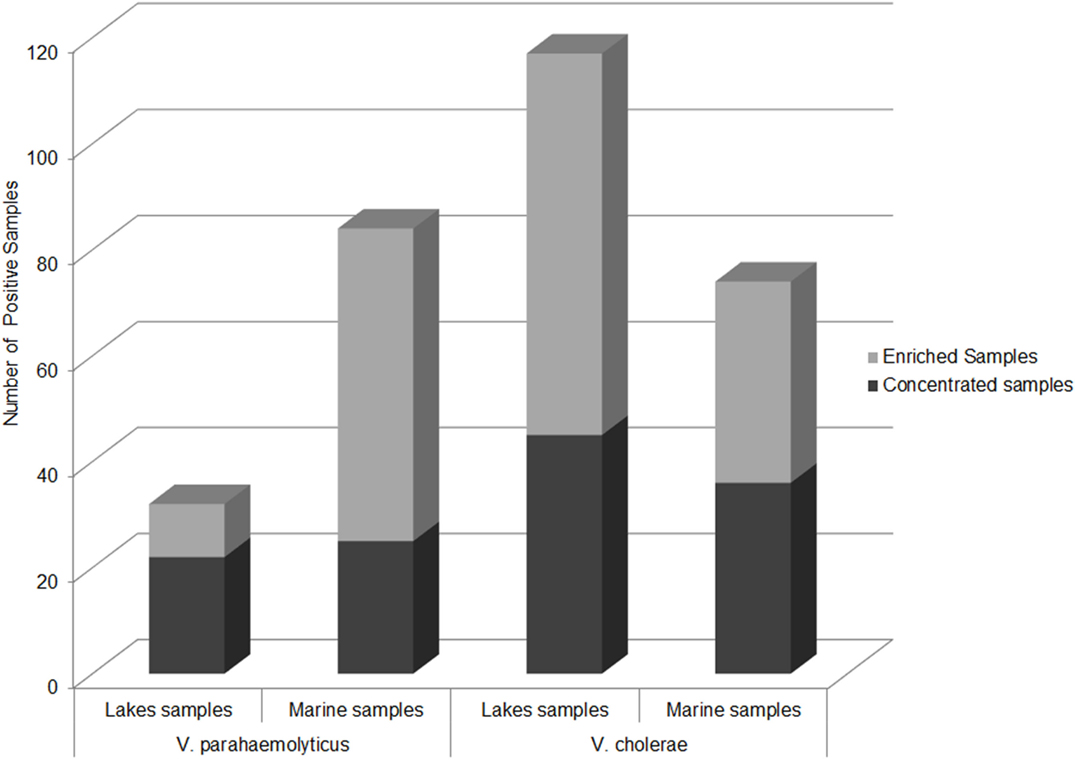
Figure 8. Abundance of PCR/ESI-MS positive samples of V. parahaemolyticus and V. cholerae. Number of samples by sample type and sample site that yielded detectable levels of V. parahaemolyticus and V. cholerae via PCR/ESI-MS.
Conclusion
In this study, we describe the occurrence, diversity, and seasonal distribution of clinically important Vibrio species in recreational waters in the South Caucasus. In total, 10 clinically important Vibrio species (of 12 known species) were detected among these sites; nine of which were detected in Black Sea coastal waters and six in inland sites. The Black Sea Vibrio populations were dominated by V. parahaemolyticus, followed by V. cholerae, V. vulnificus, and V. alginolyticus. V. cholerae was readily detected in fresh and brackish water samples collected at sites near Tbilisi.
V. cholerae was observed in the same ecological niche as two other important Vibrio species: V. vulnificus and V. alginolyticus. This finding is significant for several reasons. First, the possibility of acquiring infections caused by these pathogens will likely increase as the concentration/diversity of pathogenic Vibrio species located in a single niche increases. Second, such cohabitation increases the opportunity for interspecies interactions and potential exchanges of genetic material such as virulence factor genes.
As observed in previous studies, isolation of target Vibrio species and their direct detection using DNA-based methods for detection in marine and lake samples revealed seasonal patterns related to temperature that suggested that temperature affects Vibrio abundance, while salinity affects Vibrio species composition (37, 38).
Data obtained by conventional bacteriology, DFA, and PCR/ESI-MS analysis indicate that environmental reservoirs for toxigenic V. cholerae O1 (possibly O139 at some marine sites) are present in the recreational waters of Georgia, including freshwater lakes near Tbilisi, and the Georgian coastal zone of the Black Sea. Of particular note is the detection of V. cholerae O1 in the Tbilisi Sea, a freshwater reservoir that serves as a source of drinking water for several districts of Tbilisi. Previous detection of pandemic variants of V. parahaemolyticus (serotypes O3:K6) suggested increased risk for infection with the organism in areas where it is detected but not known to have (or that are not monitored for) outbreaks of seafood-related gastroenteritis or other water-related illnesses (28). This is even more apparent when the high frequency of seasonally reported enteric diseases with idiopathic etiologies in Georgia (especially in the Ajara region) is taken into account (30, 31).
Perpetually changing environmental conditions (e.g., increasing surface water temperatures) can significantly elevate the risk of infections related to potentially human-pathogenic Vibrio species. Such changes may also affect temperate regions with mild subtropical climates such as the South Caucasus. In the South Caucasus, climate change may lead to accelerated ecological problems and raise additional public health issues. In the coming decade, increases in air temperature (with associated long-lasting hot summers) and frequent heavy storms, combined with elevated surface water temperatures (up to ≥30°C) predicted for the Black Sea coastal areas, could trigger massive mortalities of aquatic organisms and possibly a significant rise in the incidence of diarrheal disease (30). Similarly, we can expect to observe such effects of climate change (with corresponding consequences for public health) associated with the recreational water bodies around Tbilisi.
In summary, although the occurrence of potentially pathogenic autochthonous water bacteria such as Vibrios cannot be controlled in natural ecosystems, the likelihood of the incidence of human illness can be reduced. This can be accomplished by limiting exposure to recreational water bodies with suspected elevated numbers of pathogenic Vibrio species. Furthermore, exposure can also be limited in areas where changes in ecological parameters have the potential to trigger proliferation of clinically important Vibrio species. Therefore, regular monitoring of water reservoirs for possible microbial pathogens is recommended to allow for early response by public health authorities (e.g., prevention and treatment measures to combat relevant diseases). The results presented in this report, as well as those in our previous publications (17, 27, 30) provide a foundation for effective monitoring of target Vibrio species, as well as for creation of predictive models based on environmental indicators.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This research was made possible by financial support provided by the U.S. Department of Defense (DoD), Defense Threat Reduction Agency (DTRA), through the Cooperative Biological Engagement Program’s (CBEP) Cooperative Biological Research Project (CBR) GG-13, in Georgia. The research was also partially supported by the National Science Foundation grant (No. 0813066) and the National Institutes of Health grant (No. 2RO1A1039129-11A2) awarded to Drs. Anwar Huq and Rita Colwell of the University of Maryland. The findings, opinions, and views expressed herein belong to the authors and do not reflect an official position of any organization listed. The authors are grateful to George Mikuchadze for his assistance in algorithm development for biochemical identification of Vibrio species. The authors also thank Attimo Research and Development, and Sonya Narodny for editorial assistance.
Supplementary Material
The Supplementary Material for this article can be found online at http://journal.frontiersin.org/article/10.3389/fpubh.2015.00232
References
1. Belkin S, Colwell RR. Oceans and Health: Pathogens in the Marine Environment. New York, NY: Springer (2005). 474 p.
2. Thompson FL, Iida T, Swings J. Biodiversity of Vibrios. Microbiol Mol Biol Rev (2004) 68(3):403–31. doi: 10.1128/MMBR.68.3.403-431.2004
3. Barbieri E, Falzano L, Fiorentini C, Pianetti A, Baffone W, Fabbri A, et al. Occurrence, diversity, and pathogenicity of halophilic Vibrio spp. and non-O1 Vibrio cholerae from estuarine waters along the Italian Adriatic coast. Appl Environ Microbiol (1999) 65(6):2748–53.
4. Farmer JJ, Janda J, Birkhead K. Vibrio: Manual of Clinical Microbiology. 8 ed. Washington, DC: ASM Press (2003). p. 706–18.
5. Johnson CN, Bowers JC, Griffitt KJ, Molina V, Clostio RW, Pei S, et al. Ecology of Vibrio parahaemolyticus and Vibrio vulnificus in the coastal and estuarine waters of Louisiana, Maryland, Mississippi, and Washington (United States). Appl Environ Microbiol (2012) 78(20):7249–57. doi:10.1128/AEM.01296-12
6. Hlady WG, Klontz KC. The epidemiology of Vibrio infections in Florida, 1981-1993. J Infect Dis (1996) 173(5):1176–83. doi:10.1093/infdis/173.5.1176
7. Pfeffer CS, Hite MF, Oliver JD. Ecology of Vibrio vulnificus in estuarine waters of eastern North Carolina. Appl Environ Microbiol (2003) 69(6):3526–31. doi:10.1128/AEM.69.6.3526-3531.2003
8. Morris JG Jr. Cholera – modern pandemic disease of ancient lineage. Emerg Infect Dis (2011) 17(11):2099–104. doi:10.3201/eid1711.111109
9. Onishenko G, Ganin V, Golubinsky E. Vibrios of Non-O1 Serological Group and Their Importance in Human Pathology. Moscow: Ministry of Health of the Russian Federation (2001).
10. Okuda J, Ishibashi M, Hayakawa E, Nishino T, Takeda Y, Mukhopadhyay AK, et al. Emergence of a unique O3:K6 clone of Vibrio parahaemolyticus in Calcutta, India, and isolation of strains from the same clonal group from Southeast Asian travelers arriving in Japan. J Clin Microbiol (1997) 35(12):3150–5.
11. DePaola A, Ulaszek J, Kaysner CA, Tenge BJ, Nordstrom JL, Wells J, et al. Molecular, serological, and virulence characteristics of Vibrio parahaemolyticus isolated from environmental, food, and clinical sources in North America and Asia. Appl Environ Microbiol (2003) 69(7):3999–4005. doi:10.1128/AEM.69.7.3999-4005.2003
12. Varghese MR, Farr RW, Wax MK, Chafin BJ, Owens RM. Vibrio fluvialis wound infection associated with medicinal leech therapy. Clin Infect Dis (1996) 22(4):709–10. doi:10.1093/clinids/22.4.709a
13. Hickman FW, Farmer JJ, Hollis DG, Fanning GR, Steigerwalt AG, Weaver RE, et al. Identification of Vibrio hollisae sp. nov. from patients with diarrhea. J Clin Microbiol (1982) 15(3):395–401.
14. Uh Y, Park JS, Hwang GY, Jang IH, Yoon KJ, Park HC, et al. Vibrio alginolyticus acute gastroenteritis: report of two cases. Clin Microbiol Infect (2001) 7(2):104–6. doi:10.1046/j.1469-0691.2001.00207.x
15. Hornstrup MK, Gahrn-Hansen B. Extraintestinal infections caused by Vibrio parahaemolyticus and Vibrio alginolyticus in a Danish county, 1987-1992. Scand J Infect Dis (1993) 25(6):735–40. doi:10.3109/00365549309008571
16. Shin JH, Shin MG, Suh SP, Ryang DW, Rew JS, Nolte FS. Primary Vibrio damsela septicemia. Clin Infect Dis (1996) 22(5):856–7. doi:10.1093/clinids/22.5.856
17. Nishibushi M. Miscellaneous human pathogens. In: Thompson F, Austin B, Swings J, editors. The Biology of Vibrios. ASM Press (2006). 423 p.
18. Chatterjee SH, Haldar S. Vibrio related diseases in aquaculture and development of rapid and accurate identification methods. J Marine Sci Res Dev (2012) S1:002. doi:10.4172/2155-9910.S1-002
19. Cavallo RA, Stabili L. Culturable Vibrios biodiversity in the Northern Ionian Sea (Italian coasts). Sci Mar (2004) 68(S1):23–9. doi:10.3989/scimar.2004.68s123
20. Danovaro R, Fonda Umani S, Pusceddu A. Climate change and the potential spreading of marine mucilage and microbial pathogens in the Mediterranean Sea. PLoS One (2009) 4(9):e7006. doi:10.1371/journal.pone.0007006
21. Cantet F, Hervio-Heath D, Caro A, Le Mennec C, Monteil C, Quemere C, et al. Quantification of Vibrio parahaemolyticus, Vibrio vulnificus and Vibrio cholerae in French Mediterranean coastal lagoons. Res Microbiol (2013) 164(8):867–74. doi:10.1016/j.resmic.2013.06.005
22. Aldova E, Zakhariev ZA, Dinev TS, Zlatanov ZT. [Vibrio parahaemolyticus in the Black Sea]. Zentralbl Bakteriol Orig (1971) 218(2):176–88.
23. Libinzon AE, Domaradskii IV, Us ZI, Demina AI, Nagornaia AF. [Parahemolytic Vibrios and related halophilic microorganisms of the Black Sea]. Zh Mikrobiol Epidemiol Immunobiol (1974) 00(5):80–4.
24. Libinzon AE, Gal’tseva GV. [Taxonomic position of representatives of the species Vibrio cholerae]. Zh Mikrobiol Epidemiol Immunobiol (1981) 11:41–4.
25. Grim CJ, Jaiani E, Whitehouse CA, Janelidze N, Kokashvili T, Tediashvili M, et al. Detection of toxigenic Vibrio cholerae O1 in freshwater lakes of the former Soviet Republic of Georgia. Environ Microbiol Rep (2010) 2(1):2–6. doi:10.1111/j.1758-2229.2009.00073.x
26. Whitehouse CA, Baldwin C, Sampath R, Blyn LB, Melton R, Li F, et al. Identification of pathogenic Vibrio species by multilocus PCR-electrospray ionization mass spectrometry and its application to aquatic environments of the former Soviet Republic of Georgia. Appl Environ Microbiol (2010) 76(6):1996–2001. doi:10.1128/aem.01919-09
27. Akhmedov R, Haley BJ, Rajabov M, Ahmadova S, Gurbanov S, Colwell RR, et al. Detection of Vibrio cholerae in environmental waters including drinking water reservoirs of Azerbaijan. Environ Microbiol Rep (2013) 5(1):30–8. doi:10.1111/j.1758-2229.2012.00369.x
28. Kokashvili T, Elbakidze T, Jaiani E, Janelidze N, Kamkamidze G, Whitehouse C, et al. Comparative phenotypic characterization of Vibrio cholerae isolates collected from aquatic environments of Georgia. Georgian Med News (2013) 224:55–62.
29. Haley BJ, Kokashvili T, Tskshvediani A, Janelidze N, Mitaishvili N, Grim CJ, et al. Molecular diversity and predictability of Vibrio parahaemolyticus along the Georgian coastal zone of the Black Sea. Front Microbiol (2014) 5:45. doi:10.3389/fmicb.2014.00045
30. UND Programme. Climate Change Strategy of Ajara. Tbilisi: The United Nations Development Programme (2013).
31. Ministry of Labor H, Social Affairs of Georgia. Health care: statistical yearbook. Georgia 2010. In: NCfDCaP, editor. Health. Atlanta, GA: Department of Medical Statistics of National Centre for Disease Control and Public Health (2011) 75–87.
32. Huq A, Grim C, Colwell RR, Nair GB. Detection, isolation, and identification of Vibrio cholerae from the environment. Curr Protoc Microbiol (2006) Chapter 6:6A.5. doi:10.1002/9780471729259.mc06a05s02
33. Huq A, Haley BJ, Taviani E, Chen A, Hasan NA, Colwell RR. Detection, isolation, and identification of Vibrio cholerae from the environment. Curr Protoc Microbiol (2012) Chapter 6:6A.5. doi:10.1002/9780471729259.mc06a05s26
34. Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST. Bergey’s Manual of Determinative Bacteriology. Baltimore, MD: Williams & Wilkins (1994).
35. Muddiman DC, Anderson GA, Hofstadler SA, Smith RD. Length and base composition of PCR-amplified nucleic acids using mass measurements from electrospray ionization mass spectrometry. Anal Chem (1997) 69(8):1543–9. doi:10.1021/ac961134r
36. Mitaishvili N, Tskhvediani A, Elbakidze T, Kokashvili T, Natroshvili G, Kajaia G, et al. The abundance and diversity of Vibrio species not pathogenic to humans Georgian aquatic environment. Proc Georgian Acad Sci (2013) 9:43–9.
37. Janelidze N, Jaiani E, Lashkhi N, Tskhvediani A, Kokashvili T, Gvarishvili T, et al. Microbial water quality of the Georgian coastal zone of the Black Sea. Mar Pollut Bull (2011) 62(3):573–80. doi:10.1016/j.marpolbul.2010.11.027
38. Jaiani E, Kokashvili T, Mitaishvili N, Elbakidze T, Janelidze N, Lashkhi N, et al. Microbial water quality of recreational lakes near Tbilisi, Georgia. J Water Health (2013) 11(2):333–45. doi:10.2166/wh.2013.057
39. Lee J, Donovan T, Furniss AL. Characterization, taxonomy, and emended description of Vibrio metschnikovii. Int J Syst Bacteriol (1978) 28(1):99–111. doi:10.1099/00207713-28-1-99
40. Arunagiri K, Jayashree K, Sivakumar T. Isolation and identification of Vibrios from marine food resources. Int J Curr Microbiol App Sci (2013) 2(7):217–32.
42. Tskhvediani A, Khukhunashvili T, Porchkhidze K, Tsertsvadze G, Tediashvili M. The possible use of V. parahaemolyticus – specific bacteriophages for prevention and therapy of infections caused by V. parahaemolyticus. Georgian Med News (2014) 6(231):82–8.
43. Sarkar BL, Nair GB, Banerjee AK, Pal SC. Seasonal distribution of Vibrio parahaemolyticus in freshwater environs and in association with freshwater fishes in Calcutta. Appl Environ Microbiol (1985) 49(1):132–6.
Keywords: aquatic environment, Black Sea, lakes, infection, Vibrios, conventional culture, direct detection, diversity
Citation: Kokashvili T, Whitehouse CA, Tskhvediani A, Grim CJ, Elbakidze T, Mitaishvili N, Janelidze N, Jaiani E, Haley BJ, Lashkhi N, Huq A, Colwell RR and Tediashvili M (2015) Occurrence and diversity of clinically important Vibrio species in the aquatic environment of Georgia. Front. Public Health 3:232. doi: 10.3389/fpubh.2015.00232
Received: 16 June 2015; Accepted: 28 September 2015;
Published: 13 October 2015
Edited by:
Nathan Wolfe, Metabiota, USACopyright: © 2015 Kokashvili, Whitehouse, Tskhvediani, Grim, Elbakidze, Mitaishvili, Janelidze, Jaiani, Haley, Lashkhi, Huq, Colwell and Tediashvili. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Marina Tediashvili, bV90ZWRpYXNoLmlibXZAY2F1Y2FzdXMubmV0
†Opinions, interpretations, conclusions, and recommendations are those of the author and are not necessarily endorsed by the US Army.
‡Present address: Christopher J. Grim, U.S. Food and Drug Administration, Laurel, MD, USA;
Nino Mitaishvili, Branch of Battelle Memorial Institute in Georgia, Tbilisi, Georgia
 Tamar Kokashvili
Tamar Kokashvili Chris A. Whitehouse
Chris A. Whitehouse Ana Tskhvediani
Ana Tskhvediani Christopher J. Grim
Christopher J. Grim Tinatin Elbakidze
Tinatin Elbakidze Nino Mitaishvili
Nino Mitaishvili Nino Janelidze
Nino Janelidze Ekaterine Jaiani
Ekaterine Jaiani Bradd J. Haley4
Bradd J. Haley4 Anwar Huq
Anwar Huq Marina Tediashvili
Marina Tediashvili