- 1Department of Experimental Medicine, Post Graduate School in Hygiene and Preventive Medicine, University of Perugia, Perugia, Italy
- 2Department of Health Sciences (DISSAL), University of Genoa, Genoa, Italy
- 3Digestive Endoscopy Unit, Veneto Institute of Oncology IOV-IRCCS, Padua, Italy
- 4Section of History of Medicine and Ethics, Department of Health Sciences (DISSAL), University of Genoa, Genoa, Italy
- 5Hygiene and Public Health Unit, Local Health Unit 3 of Genoa, Genoa, Italy
- 6Department of Experimental Medicine, University of Perugia, Perugia, Italy
- 7Department of Pharmaceutical Sciences, Unit of Public Health, University of Perugia, Perugia, Italy
Aim: According to the World Health Organization (WHO), communicable tropical and sub-tropical diseases occur solely, or mainly in the tropics, thriving in hot, and humid conditions. Some of these disorders termed as neglected tropical diseases are particularly overlooked. Communicable tropical/sub-tropical diseases represent a diverse group of communicable disorders occurring in 149 countries, favored by tropical and sub-tropical conditions, affecting more than one billion people and imposing a dramatic societal and economic burden.
Methods: A systematic review of the extant scholarly literature was carried out, searching in PubMed/MEDLINE and Scopus. The search string used included proper keywords, like big data, nontraditional data sources, social media, social networks, infodemiology, infoveillance, novel data streams (NDS), digital epidemiology, digital behavior, Google Trends, Twitter, Facebook, YouTube, Instagram, Pinterest, Ebola, Zika, dengue, Chikungunya, Chagas, and the other neglected tropical diseases.
Results: 47 original, observational studies were included in the current systematic review: 1 focused on Chikungunya, 6 on dengue, 19 on Ebola, 2 on Malaria, 1 on Mayaro virus, 2 on West Nile virus, and 16 on Zika. Fifteen were dedicated on developing and validating forecasting techniques for real-time monitoring of neglected tropical diseases, while the remaining studies investigated public reaction to infectious outbreaks. Most studies explored a single nontraditional data source, with Twitter being the most exploited tool (25 studies).
Conclusion: Even though some studies have shown the feasibility of utilizing NDS as an effective tool for predicting epidemic outbreaks and disseminating accurate, high-quality information concerning neglected tropical diseases, some gaps should be properly underlined. Out of the 47 articles included, only 7 were focusing on neglected tropical diseases, while all the other covered communicable tropical/sub-tropical diseases, and the main determinant of this unbalanced coverage seems to be the media impact and resonance. Furthermore, efforts in integrating diverse NDS should be made. As such, taking into account these limitations, further research in the field is needed.
Introduction
According to the World Health Organization (WHO), communicable tropical and sub-tropical diseases “occur solely, or principally, in the tropics” and “thrive in hot, humid conditions.” While some of these disorders, such as malaria, receive adequate treatment and research funding, other infections termed as “neglected tropical diseases” are relatively overlooked (1). Communicable tropical/sub-tropical diseases represent a diverse group of communicable disorders occurring in 149 countries, favored by tropical and sub-tropical conditions, affecting more than one billion people and imposing a dramatic societal and economic burden (2). Moreover, problems related to communicable tropical diseases control are mainly due to (i) tropical climate, that favors the spread of these diseases and (ii) the poverty of the regions affected by these diseases (1). Among the tropical/sub-tropical diseases there are also neglected tropical diseases that include a subset of 17 infectious disorders (caused by viruses, such as dengue, Chikungunya, and rabies, by prokaryotic organisms, such as Buruli ulcer, leprosy, trachoma, treponematoses, or by eukaryotic organisms, like Chagas disease, human African trypanosomiasis, leishmaniases, dracunculiasis, lymphatic filariasis, onchocerciasis, cysticercosis/teniasis, echinococcosis, foodborne trematodiases, and schistosomiasis) (3).
In the contemporary globalized society, the emergence/re-emergence of old and new infectious diseases, due to rapid human development in terms of demographics, populations, and environment, represent a serious public health concern (3). Communicable tropical diseases generate a relevant burden that disproportionately impacts on the world’s poorest, constituting, as such, a major barrier to development efforts in order to alleviate poverty and improve human health status and condition in the developing areas. Malaria and neglected tropical diseases kill more than 800,000 people annually and create long-term disability in millions more (4).
For communicable tropical disorders, the WHO, together with the United Nations Children’s Fund, the United Nations Development Programme, and the World Bank, has launched a “Special Programme for Research and Training in Tropical Diseases” (TDR), which represents a global program of scientific collaborations (1). Furthermore, the WHO has defined a Road Map for controlling and eliminating neglected tropical diseases by 2020 and has suggested some steps, which are fundamental in order to achieve these ambitious goals, including improved diagnostics, treatment strategy, and surveillance systems (5, 6).
Within the era of e-health, characterized by the diffusion of the new information and communication technologies (ICTs), non conventional, or novel data streams (NDS), such as web searches generated data or social media updates, are emerging as a new promising approach in enhancing/complementing traditional surveillance systems (7, 8) and/or supporting public health decision making (9). Actually, Big data, despite their promises and their potential, should not be considered or utilized as a substitute for traditional data sources, but, rather, as a valuable complementary approach. Algorithms and computational techniques they are built and rely on still need to be carefully refined, tuned, and calibrated, in order to avoid the risk of overfitting in Big data inference. For instance, this happened with “Google Flu Trend” (GFT), which failed to provide accurate predictions concerning influenza-like-illness (ILI) cases. GFT predicted, indeed, more than double the proportion of doctor visits for ILI than the centers for disease control and prevention (CDC) (10). Due to these concerns, GFT decided to no longer publish influenza estimates. Similarly, Google Dengue Trends, a web-based tool for predicting dengue cases, is not currently available.
“Infodemiology” (a port-manteau of information and epidemiology) and “infoveillance” (a combination of information and surveillance) have been coined by Gunther Eysenbach to indicate the new emerging “science of distribution and determinants of information in an electronic medium, specifically the Internet, or in a population, with the ultimate aim to inform and improve public health and public policy” (11). Systematically tracking and monitoring, collecting and analyzing health-related demand data generated by NDS could have the potential to predict events relevant for public health purposes, such as epidemic outbreaks, as well as to investigate the effect of media coverage in terms of potential distortions, misinformation and biases—the so-called “epidemics of fear” (12). Details are shown in Figure 1.
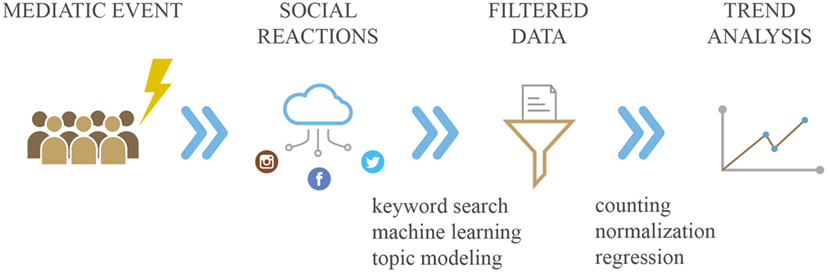
Figure 1. Collection and analysis of health-related demand data generated by novel data streams. Example of media event influencing society and consequent social reactions caught by social networks. Analysis of these data is represented by trend report.
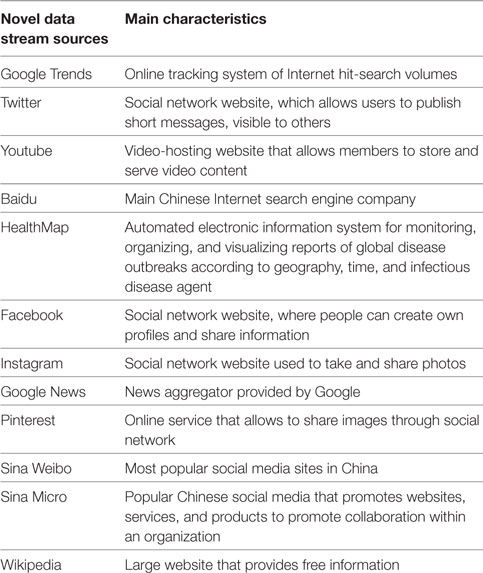
The aim of the current investigation was to systematically assess the feasibility of exploiting NDS for surveillance purposes and/or their potential for capturing public reaction to epidemic outbreaks. The main characteristics of NDS analyzed in this paper are briefly overviewed in Box 1.
Box 1. Main characteristics of novel data streams (NDS) analyzed.
Materials and Methods
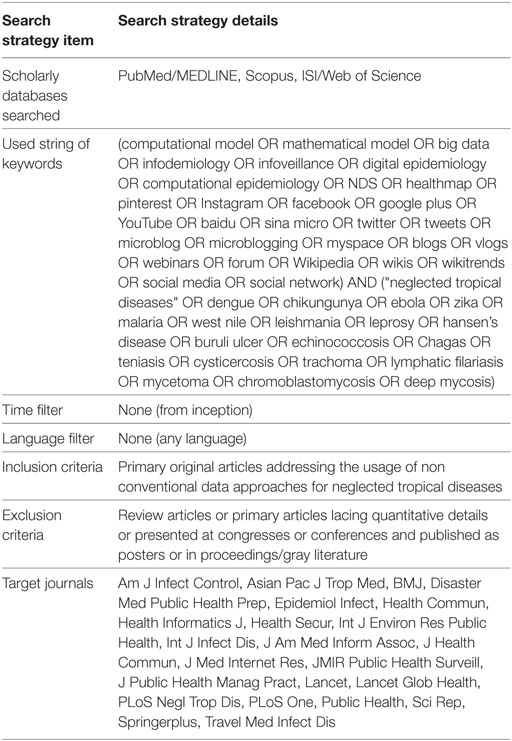
The following systematic review was conducted according to the “Preferred Reporting Items for Systematic Reviews and Meta-Analyses” (PRISMA) guidelines (13). The literature search was performed in July–August 2017 using pre-established ad hoc key words and updated in September–October 2017. The search strategy is detailed in Table 1.
Inclusion and Exclusion Criteria
Articles were included in the present systematic review whether they met the following inclusion criteria: (i) full text available; (ii) original articles; (iii) focused on communicable tropical and sub-tropical disorders, including neglected tropical diseases; and (iv) assessing novel sources of data, such as Twitter, web searches monitoring tools like Google Trends, Facebook, Google Plus, Wikipedia access logs, and traffic tracking tools, such as WikiTrends, and so on.
Exclusion criteria were: (i) studies without original data (abstract, letters to editor, editorials, comments, commentaries, expert opinions, reviews) and (ii) studies published in congress proceedings and gray literature.
No time and language filter was applied. Two researchers (NLB and VG), independently, screened title and abstract in order to verify the articles relevance. Possible disagreements were resolved through discussion or third reviewer consultation. The full text was downloaded only for the selected titles, and reference lists of included studies were also checked in order to identify any other potential relevant paper.
Data Extraction
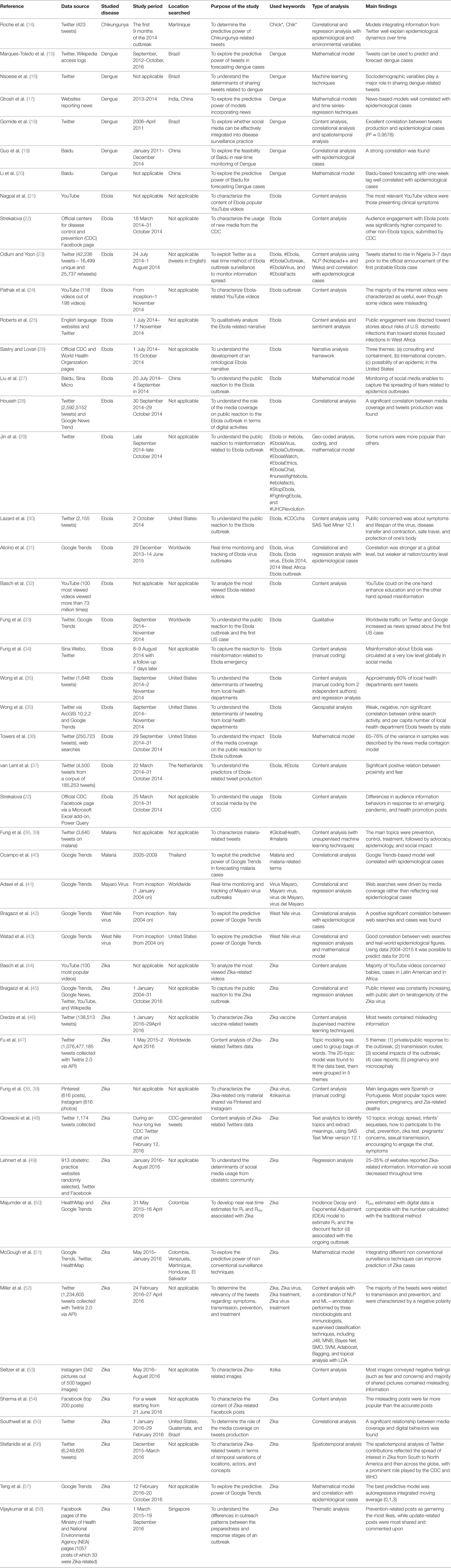
Main information, from the included studies, were extracted independently from two authors (VG and NLB) and collected in a pre-defined ad hoc spreadsheet. The collected data included: (i) surname of the first author, (ii) year of publication, (iii) data source, (iv) studied disease, (v) study period, (vi) location searched, (vii) used keywords, (viii) aim of the study, and (ix) main findings.
Results
A total of 17,945 articles were retrieved: two articles were found by means of extensive manual hand-searching and cross-referencing. After a preliminary screening, a total of 14,996 articles were excluded because they did not meet the inclusion criteria. Two more articles were retrieved from additional sources, finally 57 remaining articles were analyzed in full. 10 of them were excluded with reasons and last 47 articles were included in the present systematic review. Results are syntetized in (Table 2). The screening process is shown in Figure 2.
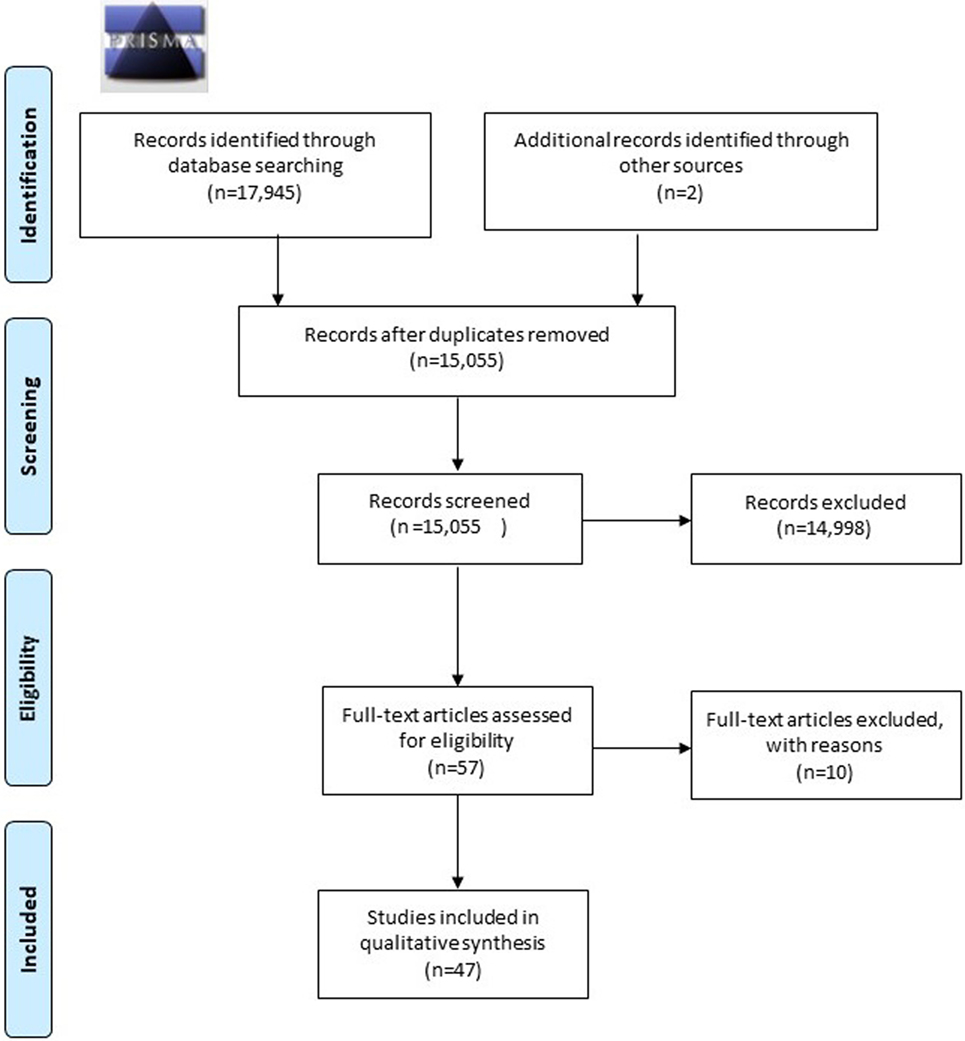
Figure 2. Flow diagram with screening process, according to the preferred reporting items for systematic reviews and meta-analyses guidelines.
Out of 47 articles included in the review, 1 on Chikungunya, 6 on dengue, 19 were focused on Ebola, 2 on malaria, 1 on Mayaro virus, 2 on West Nile virus, and 16 on Zika. Looking at the non conventional data approaches used, 11 studies were searching on Google Trends, 5 on You tube, 2 on Wikipedia, 1 on Google News, 2 on HealthMap, 1 on Sina Weibo, and 1 on Sina Micro, 1 on Pinterest, 2 on Instagram, 3 on Web sites, 3 on Baidu, and 5 on Facebook. The most used data source was Twitter with 25 studies. However, some studies analyzed data on several sources, such as Fung and colleagues who search information about Zika virus on Pinterest and Instagram (39), or Househ et al. who searched on Twitter and Google Trends (28). Fifteen were dedicated on developing and validating forecasting techniques for real-time monitoring of neglected tropical diseases, while the remaining studies investigated public reaction to infectious outbreaks (Figure 3), in terms, for example, of sentiment analysis and spreading of fake news related to tropical disorder outbreaks.
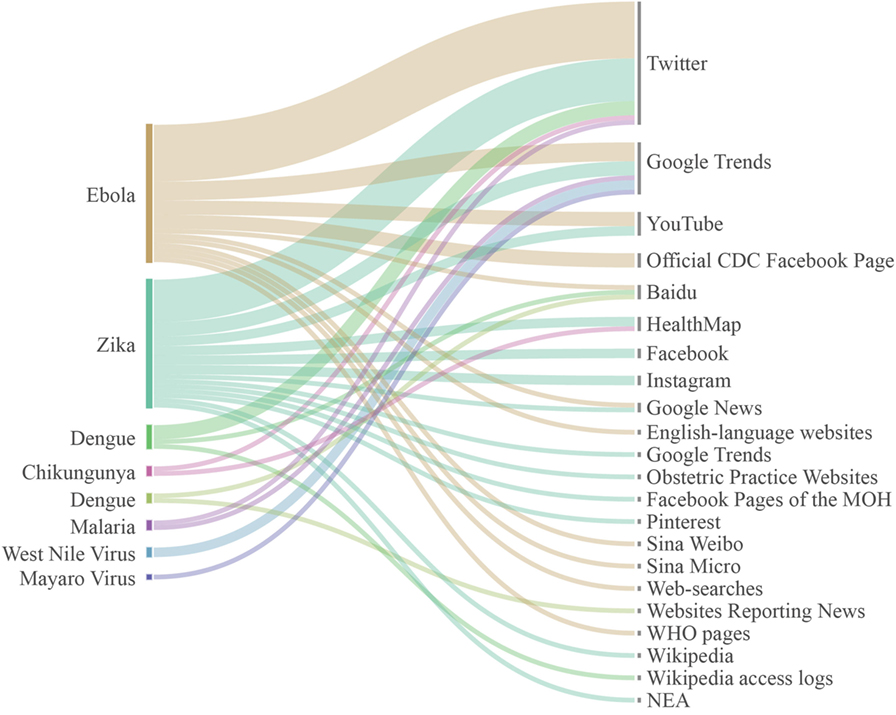
Figure 3. Communicable tropical/sub-tropical disorders and neglected tropical diseases investigated and non conventional sources analyzed, according to our results. Line thicknesses represent the volume of studies retrieved.
Neglected Tropical Diseases
Chikungunya Virus
Only one study was related to Chikungunya. Roche et al. (14) harnessed tweets related to Chikungunya posted during the outbreak in Martinique (14) and, performing a regression analysis with epidemiological and environmental variables, found that the integration of model and tweets contents well explained epidemiological dynamics over time.
Dengue
Five studies were related to dengue. Four of them relied on predictive models to predict dengue outbreaks. In Brazil, Gomide et al. (18) exploited Twitter and performed extensive content, correlation, and spatiotemporal analyses (18). Authors were able to find an excellent association between tweets production and epidemiological cases (R2 = 0.9578). Always in Brazil, Marques-Toledo et al. (15) utilized both Twitter and Wikipedia access logs in building predictive mathematical models for forecast dengue cases. In China, Guo et al. (19) leveraged Baidu for real-time monitoring and tracking of dengue cases (19). A strong correlation with epidemiological cases was found. In India and in China, Ghosh et al. (17) explored the predictive power of models incorporating websites reporting news related to dengue, carrying out mathematical models, and time series-regression techniques (17). News-based models were found to well correlate with epidemiological cases.
One article harnessed big data to explore the determinants of sharing tweets related to dengue. In particular, Nsoesie et al. (16), using machine learning techniques, found that sociodemographic variables played a major role in producing and sharing dengue-related tweets (16).
Communicable Tropical/Subtropical Diseases
Ebola
Twenty articles were related to Ebola. All of them exploited big data sources to capture public reaction to Ebola outbreaks, both in terms of sentiments, fears, and concerns and of knowledge, beliefs, and attitudes. More in detail, four studies exploited Twitter. van Lent et al. (37) investigated the predictors of Ebola-related tweet production and found a significant positive relation between proximity and fear for Ebola virus (37). Jin et al. (29) harnessed Twitter to understand the public reaction to misinformation related to Ebola outbreak, performing an extensive geo-coded analysis, coding, and mathematical modeling (29). Authors found that some Ebola-related rumors were more popular than others Lazard et al. (30) found that the public was mainly concerned with symptoms and lifespan of the virus, disease transfer and contraction, safe travel, and protection of one’s body (30). Interestingly, Wong et al. (35) aimed at understanding the determinants of tweeting from local health departments. Approximately 60% of local health departments sent tweets (35).
Three studies utilized YouTube. Nagpal et al. (21) analyzed the most popular Ebola-related videos and found that the most relevant ones were those presenting clinical symptoms (21). Pathak et al. (24) found that the majority of the internet videos about Ebola were useful, even though some videos were misleading (24). Basch et al. (32) analyzed the 100 most viewed videos on YouTube with more than 73 million of visualizations and concluded that YouTube has a Yin–Yang nature, in that it could, on the one hand, enhance education and, on the other hand, spread misinformation (32).
Three studies utilized Facebook. Sastry and Lovari (26) analyzed the material posted on the official CDC and WHO pages (26). The following major themes were identified: (a) consulting and containment, (b) international concern, and (c) the possibility of an epidemic in the United States. Strekalova (22), reviewing the official CDC page, found that the CDC submitted fewer posts about Ebola than about non-Ebola topics, even though audience engagement was significantly higher (22). Furthermore, men were more interested in Ebola posts and submitted more comments per user. Moreover, Strekalova (22) found that there were differences in audience information behaviors in response to the emerging Ebola pandemic and health promotion posts (22).
Seven studies utilized more than one big data source. Fung et al. (33) used both Twitter and GT to understand the public reaction to the Ebola outbreak and the first US case (33). Fung et al. (34) combined Sina Weibo and Twitter to capture the reaction to misinformation related to Ebola emergency (34). Liu et al. (27) harnessed Baidu and Sina Micro to investigate the public reaction to the Ebola outbreak in China, performing a mathematical model (27). Roberts et al. (25) mined both English language websites and Twitter to qualitatively analyze the Ebola-related narrative, carrying out content and sentiment analysis (25). Househ (28), using Twitter and Google News Trend, found a significant correlation between media coverage and tweets production (28). Towers et al. (36) integrated Twitter and web searches to understand the impact of the media coverage on the public reaction to Ebola outbreak in the United States in terms of digital activities, performing a mathematical model (36). Wong et al. (35) exploited both Twitter and GT to understand the determinants of tweeting from local health departments Ebola, by means of a geospatial analysis (35). Authors found a weak, negative, non-significant correlation between online search activity and per capita number of local health department Ebola tweets by state.
Besides capturing public reaction to Ebola epidemic, three studies attempted to perform also predictive models and analyses. Alicino et al. (31) explored the feasibility of exploiting GT for a real-time monitoring and tracking of Ebola virus outbreaks, carrying out correlation and regression analysis with epidemiological cases (31). Authors found that correlation was stronger at a global level, but weaker at nation/country level, probably due to unbalanced, biased media coverage, and to digital divide. Odlum and Yoon (23) utilized Ebola-related tweets as a real-time method of Ebola outbreak surveillance to monitor information spread, capture early epidemic detection, as well as to examine content of public knowledge and attitudes (23). Authors found that tweets began to start to rise in Nigeria 3–7 days prior to the official announcement of the first probable Ebola case. Topics discussed included risk factors, prevention education, disease trends, and human compassion.
Malaria
GT was used for forecasting malaria cases by Ocampo et al. in 2013 (40). This study was performed using data related to Thailand in the period 2005–2009. Authors developed four Google search query-based models: namely, the so-called “microscopy model” (which uses terms associated with official data), the “automatic model” (based on automated selection algorithm), the “physician model” (generated from terms selected by surveyed Thai physicians), and the “stepwise model.” GT-based models well correlated with epidemiological cases.
Fung et al. (38, 39) used Twitter and performed a content analysis of the Malaria-related tweets (38). The main topics were: prevention, control, and treatment, followed by advocacy, epidemiological information, and societal impact.
Mayaro Virus
Only one study was related to Mayaro virus and exploited GT. Adawi et al. (41) explored the feasibility of utilizing GT for a real-time monitoring and tracking of Mayaro virus outbreaks (41). Correlational and regression analysis were performed with epidemiological cases and with other NDS, including Google News, PubMed/MEDLINE. Authors found that web searches were driven by media coverage rather than reflecting real epidemiological cases.
West Nile Virus
Two studies focused on the West Nile virus (42), and both of them used GT. Bragazzi et al. (42) aimed at exploiting the predictive power of GT (42) in Italy, performing a correlation analysis with epidemiological cases. Authors found a positive significant correlation between web searches and cases. Watad et al. (43) explored the predictive power of GT in the United States, carrying out correlation and regression analyses as well as mathematical modeling (43). Results showed a good correlation between web searches and real-world epidemiological figures. The best seasonal autoregressive integrated moving average model with explicative variable (SARIMAX) computed was (0,1,1)X(0,1,1)4, that is to say a “seasonal exponential smoothing” model. Moreover, using data from 2004 to 2015 it was possible to predict data for 2016.
Zika Virus
Sixteen studies focused on Zika and nine of them used Twitter as non conventional data source. In the majority of the cases (4 papers), the type of performed analysis was content analysis (46–48, 52), even though carried out with various research purposes. More in detail, Miller et al. (52) conducted a tweets analysis during the period of the hosting of the Olympics games and captured public reaction in terms of sentiments and concerns related to the potential association between Zika infection, microcephaly, and Guillain–Barrè syndrome, an association probable, but not yet confirmed at that time. Although the total polarity was negative, the percentage of positive tweets was higher than expected. An imbalance in the volume of tweets focusing on treatment was found. Similarly, a study by Fu et al. (47) lead to the emergence of five major themes: (1) government, private, and public sector, and general public response to the outbreak; (2) transmission routes; (3) societal impacts of the outbreak; (4) case reports; and (5) pregnancy and microcephaly. Glowacki et al. (48) investigated the use of new ICTs by healthcare authorities and organisms and, for the purpose, collected tweets during an hour-long live CDC Twitter chat, identifying 10 major topics. Some of them were related to the virology of Zika, spread, infants’, and pregnants’ sequelae, sexual transmission, and symptomatology. Dredze et al. (46) focused on the spreading of conspiracy theories and pseudo-scientific claims and found that tweets disseminating misleading information were concentrated almost all during the first week of pandemic (46).
Three studies used quantitative approaches, namely correlation and regression analysis (45, 49, 55), mathematical modeling (51), and spatiotemporal analysis (56). Southwell et al. (55) found strong positive correlations between news coverage, social media mentions, and online search behavior (55). Bragazzi et al. (45) found a constantly increasing public interest toward Zika, with the public opinion being particularly worried by the alert of teratogenicity of the Zika virus (45). In particular, the most frequent queries were about symptoms, transmission, and possible sequelae, such as microcephaly. Lehnert et al. (49) performed a regression analysis in order to understand the determinants of social media usage from obstetric community (49). The percentage of obstetric practice websites increased the number of information posted about Zika virus throughout the time, however, the proportion of practice sites posting Zika virus content on Facebook and Twitter declined. Practice websites related to university hospitals were more likely to post information on Zika virus compared to independent practice sites. McGough et al. (51) through a mathematical model, integrated different non conventional surveillance data (51), such as Google searches, Twitter microblogs, and the HealthMap digital surveillance system, and found that models relying on Google and Twitter showed the best 2- and 3-week ahead predictions. Last, Stefanidis et al. (56) performed a spatiotemporal analysis in order to characterize Zika-related tweets in terms of temporal variations of locations, actors, and concepts (56). The spatiotemporal analysis of the different Twitter contributions reflected the spread of interest in Zika from South America to North America and, then, across the globe. Healthcare institutional bodies, such as the CDC and the WHO, played a major role in tweet production.
Other type of big data sources explored in Zika studies were Facebook (54, 58), Google trends (50, 57), YouTube (44), Pinterest, and Instagram (39, 53). Vijaykumar et al. (58) analyzed the Facebook material posted on the public page of Ministry of Health (MOH) of Singapore and the Facebook page of National Environmental Agency (NEA), in order to evaluate the outreach and the engagement during the Zika pandemic. Generally speaking the MOH’s posts were more shared and received much more like compared to NEA’s post, however, the NEA’s posts were much more commented. Looking at the content, the NEA’s posts were more focused on prevention and intervention compared to the MOH’s posts with, in their turns, were more related to updates and investigations. Sharma et al. (54) analyzed the top 200 Facebook posts collected for 1 week starting from 21 June 2016 (54). The misleading posts were far more popular than the posts dispersing accurate, relevant public health information about the disease. Actually, the most popular relevant posts were published by the WHO, and obtained 43,000 views with 964 shares. The most popular misleading posts obtained, instead, more than 530,000 views, around 20,000 combined shares, and hundreds of comments.
Another big data source was GT. Actually, two studies examined GT-generated volume data in order to build predictive models. Teng et al. (57) aimed at predicting the number of infection cases (57). Authors constructed an autoregressive integrated moving average model (0, 1, 3) for the dynamic estimation of ZIKV outbreaks. Majumder et al. (50), using nontraditional digital data, such as HealthMap and Google Trends, tried to estimate the R0 and Robs parameters of Zika virus spreading in Colombia. Authors observed an initially low, but increasing awareness and interest toward Zika. Google search was used in order to distribute more realistical over time, cumulative reported case counts. The ranges for Robs estimated using digital data were well comparable with the figures calculated with the traditional method, even though a little lower. Transmission parameters can be estimated in real time using digital surveillance data, especially when traditional methods are not available.
Only one study assessed the content of YouTube videos on Zika (44). Basch et al. (44) analyzed the 100 most viewed English ZIKV-related videos. Among them, the majority were consumer-generated and Internet-based news videos. According to the contents, the majority of the videos concerned babies, cases in Latin American and in Africa.
Also Pinterest and Instagram were exploited, however, only two studies were conducted and both of them performed a content analysis (39, 53). Fung et al. (38, 39) analyzed more than 600 posts and photos on Facebook and Pinterest, respectively (39). The most popular topics were: prevention, pregnancy, and Zika-related deaths. Seltzer et al. (53) analyzed images posted on Instagram (53) and found that, even though the majority of posts focused on transmission and prevention, most of them conveyed negative feelings (such as fear and concerns) and contained misleading information.
Discussion
In the past years, there has been a growing interest from the scholarly community in big data sources and their impact on public health. This was parallel to the interest toward neglected and communicable tropical diseases. Currently, communicable tropical diseases—including also the subset of neglected ones—represent re-emerging infections. However, re-emergence is not a completely new phenomenon occurring only in the past decades, actually it is happening since centuries. On the other hand, today re-emergence and dispersion of infectious agents are more rapid and geographically extensive, mainly due to globalization, and to arthropods or other vectors adaptation to its effects (59).
Novel data streams appear to be promising tools for predicting the spread of infectious agents, and, as such, can potentially aid and inform early decision support for when and how to employ public health interventions within a certain community. Emergency situations, being urgent scenarios, need accurate, reliable, and fast predictive models (60). Traditional surveillance systems are often plagued by a number of shortcomings and drawbacks, such as a significant delay in releasing official government-reported case counts (51). NDS seem to offer a real-time way to track and monitor outbreak dynamics, as well as to capture relevant information and parameters related to infection rates when these details are scarcely known or not available.
Novel data streams are also versatile tools in that they can be exploited to capture public reactions to epidemic outbreaks, in terms of emotion and fears, and of knowledge, attitudes, and practices. Some studies have harnessed big data sources to understand the spread of misinformation. Years of researches in the field of health communication and psychology have shown that opinion change represents a much more challenging issue than opinion formation, since, once people believe something wrong or misleading, it is difficult to dissuade them from such rooted beliefs (46). With respect to this topic, some studies have shown that NDS have a Yin–Yang nature, being, on the one hand, useful resources for promoting health education and being, on the other hand, vehicles of potentially dangerous information and content. In the era of the “post-truth,” the dissemination of fake news, alleged claims, and not evidence-based rumors could have serious implications in terms of public health. Techniques of social bookmarking and the direct involvement of healthcare workers and practitioners (in producing health-related websites, posting and sharing online material, tweeting, chatting, and so on) could be useful strategies (61).
Stakeholders and health authorities should be aware of the new ICTs, in that they could usefully exploit Internet-based tools for collecting the concerns of public opinion and replying to them, re-ensuring, and disseminating accurate, high-quality information (45). However, some studies included in the current systematic reviews have stressed gaps in usage of NDS by official healthcare organisms and bodies. Efforts should be made to convey a proper and effective health communication, utilizing ICTs and borrowing approaches from social marketing, making their posted material and delivered information more appealing, in terms of public outreach and engagement.
Another important point that should be stressed is that the value of each paper included in the current systematic review does not appear equal with respect to the field of public health. For example, the studies by Gomide and co-workers, McGough and collaborators, Odlum and Yoon, Roche and co-workers, and Teng and coauthors are highly relevant to public health outcomes (14, 18, 23, 43, 51, 57), while the others relate primarily to social networks. As such, only few papers with respect to the overall number of articles included in the present systematic review are directly relevant for public health outcomes. This definitely deserves further investigation and research in the field.
Our systematic review has some major strength, including the breadth of the search performed. However, even though efforts have been made in order to ensure completeness of the findings, alternate spellings/misspellings of keywords could have affected the results [for example, there are nine articles returned for “chikugunya” (an incorrect spelling of the disease chikungunya) returned recently on PubMed/MEDLINE]. On the other hand, reference lists of included articles have been extensively hand-searched, to increase the chance of getting all potentially relevant studies Relatedly, a variety of computational, “big data”-related terms (such as machine learning, collective intelligence or deep learning) were not included. Ad hoc search strings are, of course, finite in length, however, we expect to have included all relevant investigations meeting with inclusion/exclusion criteria on the basis that we have carried out extensive cross-referencing and additional hand-searching.
Conclusion
Even though some studies have shown the feasibility of utilizing NDS as an effective tool for predicting epidemic outbreaks and disseminating accurate, high-quality information concerning communicable tropical diseases, some gaps should be properly underlined. Actually, among 47 studies included in our systematic review, only 7 studies focused on neglected tropical diseases (Chikungunya and dengue), while all the others were focusing on communicable tropical diseases (19 on Ebola, 2 on Malaria, 1 on Mayaro virus, 2 on West Nile virus, and 16 on Zika). In particular, out of the 17 groups of neglected tropical diseases individuated by the WHO, only two types of infectious diseases (namely, dengue and Chikungunya) were covered, and the main determinant of this unbalanced coverage seems to be the media impact and resonance, as well as the fear of the spreading of epidemic agents to Western countries. Furthermore, efforts in integrating diverse NDS should be made. As such, taking into account these limitations, further research in the field is needed.
Ethical Statement
No ethical approval is required.
Author Contributions
VG and NB contributed in conception and design of the study, data extraction and analysis. DN contributed in assembly and data interpretation. VG, NB, and DN drafted the manuscript. MM, RR, LM, and MM contributed in manuscript revision. All the authors approved the final version of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank Ms. Valeria Parisi, University of Milan, for assistance with figures.
Funding
This research received no grant from any funding agency, commercial, or not-for-profit sectors.
References
1. World Health Organization. (2018). Available from: http://www.who.int/tdr/en/ (accessed February 4, 2018).
2. World Health Organization. Neglected Tropical Diseases (2017). Available from: http://www.who.int/neglected_diseases/en/ (accessed February 4, 2018).
3. Mackey TK, Liang BA, Cuomo R, Hafen R, Brouwer KC, Lee DE. Emerging and reemerging neglected tropical diseases: a review of key characteristics, risk factors, and the policy and innovation environment. Clin Microbiol Rev (2014) 27(4):949–79. doi:10.1128/CMR.00045-14
4. Hotez PJ. The poverty-related neglected diseases: why basic research matters. PLoS Biol (2017) 15(11):e2004186. doi:10.1371/journal.pbio.2004186
5. Nii-Trebi NI. Emerging and neglected infectious diseases: insights, advances, and challenges. Biomed Res Int (2017) 2017:5245021. doi:10.1155/2017/5245021
6. Peeling RW, Boeras DI, Nkengasong J. Re-imagining the future of diagnosis of neglected tropical diseases. Comput Struct Biotechnol J (2017) 15:271–4. doi:10.1016/j.csbj.2017.02.003
7. Brownstein JS, Freifeld CC, Madoff LC. Digital disease detection – harnessing the Web for public health surveillance. N Engl J Med (2009) 360(21):2153–5. doi:10.1056/NEJMp0900702
8. Wilson K, Brownstein JS. Early detection of disease outbreaks using the Internet. CMAJ (2009) 180(8):829–31. doi:10.1503/cmaj.090215
9. Althouse BM, Scarpino SV, Meyers LA, Ayers JW, Bargsten M, Baumbach J, et al. Enhancing disease surveillance with novel data streams: challenges and opportunities. EPJ Data Sci (2015) 4:17. doi:10.1140/epjds/s13688-015-0054-0
10. Lazer D, Kennedy R, King G, Vespignani A. Big data. The parable of Google Flu: traps in big data analysis. Science (2014) 343(6176):1203–5. doi:10.1126/science.1248506
11. Eysenbach G. Infodemiology and infoveillance: framework for an emerging set of public health informatics methods to analyze search, communication and publication behavior on the Internet. J Med Internet Res (2009) 11(1):e11. doi:10.2196/jmir.1157
12. Eysenbach G. Infodemiology: the epidemiology of (mis)information. Am J Med (2002) 113(9):763–5. doi:10.1016/S0002-9343(02)01473-0
13. Moher D, Liberati A, Tetzlaff J, Altman DG, Group P. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS Med (2009) 6(7):e1000097. doi:10.1371/journal.pmed.1000097
14. Roche B, Gaillard B, Leger L, Pelagie-Moutenda R, Sochacki T, Cazelles B, et al. An ecological and digital epidemiology analysis on the role of human behavior on the 2014 Chikungunya outbreak in Martinique. Sci Rep (2017) 7(1):5967. doi:10.1038/s41598-017-05957-y
15. Marques-Toledo CA, Degener CM, Vinhal L, Coelho G, Meira W, Codeco CT, et al. Dengue prediction by the web: tweets are a useful tool for estimating and forecasting Dengue at country and city level. PLoS Negl Trop Dis (2017) 11(7):e0005729. doi:10.1371/journal.pntd.0005729
16. Nsoesie EO, Flor L, Hawkins J, Maharana A, Skotnes T, Marinho F, et al. Social media as a sentinel for disease surveillance: what does sociodemographic status have to do with it? PLoS Curr (2016) 8:1–17. doi:10.1371/currents.outbreaks.cc09a42586e16dc7dd62813b7ee5d6b6
17. Ghosh S, Chakraborty P, Nsoesie EO, Cohn E, Mekaru SR, Brownstein JS, et al. Temporal topic modeling to assess associations between news trends and infectious disease outbreaks. Sci Rep (2017) 7:40841. doi:10.1038/srep40841
18. Gomide J, Veloso A, Meira W, Almeida V, Benevenuto F, Ferraz F, et al. Dengue surveillance based on a computational model of spatio-temporal locality of Twitter. ACM Web Sci Conf (2011) 14(17):1–8. doi:10.1145/2527031.2527049
19. Guo P, Liu T, Zhang Q, Wang L, Xiao J, Zhang Q, et al. Developing a dengue forecast model using machine learning: a case study in China. PLoS Negl Trop Dis (2017) 11(10):e0005973. doi:10.1371/journal.pntd.0005973
20. Li Z, Liu T, Zhu G, Lin H, Zhang Y, He J, et al. Dengue Baidu Search Index data can improve the prediction of local dengue epidemic: a case study in Guangzhou, China. PLoS Negl Trop Dis (2017) 11(3):1–13. doi:10.1371/journal.pntd.0005354
21. Nagpal SJ, Karimianpour A, Mukhija D, Mohan D, Brateanu A. YouTube videos as a source of medical information during the Ebola hemorrhagic fever epidemic. Springerplus (2015) 4:457. doi:10.1186/s40064-015-1251-9
22. Strekalova YA. Health risk information engagement and amplification on social media. Health Educ Behav (2017) 44(2):332–9. doi:10.1177/1090198116660310
23. Odlum M, Yoon S. What can we learn about the Ebola outbreak from tweets? Am J Infect Control (2015) 43(6):563–71. doi:10.1016/j.ajic.2015.02.023
24. Pathak R, Poudel DR, Karmacharya P, Pathak A, Aryal MR, Mahmood M, et al. YouTube as a source of information on Ebola virus disease. N Am J Med Sci (2015) 7(7):306–9. doi:10.4103/1947-2714.161244
25. Roberts H, Seymour B, Fish SA II, Robinson E, Zuckerman E. Digital health communication and global public influence: a study of the Ebola epidemic. J Health Commun (2017) 22(Sup 1):51–8. doi:10.1080/10810730.2016.1209598
26. Sastry S, Lovari A. Communicating the ontological narrative of Ebola: an emerging disease in the time of “epidemic 2.0”. Health Commun (2017) 32(3):329–38. doi:10.1080/10410236.2016.1138380
27. Liu K, Li L, Jiang T, Chen B, Jiang Z, Wang Z, et al. Chinese public attention to the outbreak of Ebola in West Africa: evidence from the online big data platform. Int J Environ Res Public Health (2016) 13(8):E780. doi:10.3390/ijerph13080780
28. Househ M. Communicating Ebola through social media and electronic news media outlets: a cross-sectional study. Health Informatics J (2016) 22(3):470–8. doi:10.1177/1460458214568037
29. Jin F, Wng W, Zhao L, Dougherty E, Cao Y, Lu CT, et al. Misinformation propagation in the age of Twitter. Browse J Mag (2014) 47(12):90–4. doi:10.1109/MC.2014.361
30. Lazard AJ, Scheinfeld E, Bernhardt JM, Wilcox GB, Suran M. Detecting themes of public concern: a text mining analysis of the Centers for Disease Control and Prevention’s Ebola live Twitter chat. Am J Infect Control (2015) 43(10):1109–11. doi:10.1016/j.ajic.2015.05.025
31. Alicino C, Bragazzi NL, Faccio V, Amicizia D, Panatto D, Gasparini R, et al. Assessing Ebola-related web search behaviour: insights and implications from an analytical study of Google Trends-based query volumes. Infect Dis Poverty (2015) 4:54. doi:10.1186/s40249-015-0090-9
32. Basch CH, Basch CE, Ruggles KV, Hammond R. Coverage of the Ebola virus disease epidemic on YouTube. Disaster Med Public Health Prep (2015) 9(5):531–5. doi:10.1017/dmp.2015.77
33. Fung IC, Tse ZT, Cheung CN, Miu AS, Fu KW. Ebola and the social media. Lancet (2014) 384(9961):2207. doi:10.1016/S0140-6736(14)62418-1
34. Fung IC, Fu KW, Chan CH, Chan BS, Cheung CN, Abraham T, et al. Social media’s initial reaction to information and misinformation on Ebola, August 2014: facts and rumors. Public Health Rep (2016) 131(3):461–73. doi:10.1177/003335491613100312
35. Wong R, Harris JK, Staub M, Bernhardt JM. Local health departments tweeting about Ebola: characteristics and messaging. J Public Health Manag Pract (2017) 23(2):e16–24. doi:10.1097/PHH.0000000000000342
36. Towers S, Afzal S, Bernal G, Bliss N, Brown S, Espinoza B, et al. Mass media and the contagion of fear: the case of Ebola in America. PLoS One (2015) 10(6):e0129179. doi:10.1371/journal.pone.0129179
37. van Lent LG, Sungur H, Kunneman FA, van de Velde B, Das E. Too far to care? Measuring public attention and fear for Ebola using Twitter. J Med Internet Res (2017) 19(6):e193. doi:10.2196/jmir.7219
38. Fung IC-H, Jackson AM, Ahweyevu JO, Grizzle JH, Yin J, Tse ZTH, et al. #Globalhealth Twitter conversations on #Malaria, #HIV, #TB, #NCDS, and #NTDS: a cross-sectional analysis. Ann Global Health (2017) 83(3–4):682–90. doi:10.1016/j.aogh.2017.09.006
39. Fung IC, Blankenship EB, Goff ME, Mullican LA, Chan KC, Saroha N, et al. Zika-virus-related photo sharing on Pinterest and Instagram. Disaster Med Public Health Prep (2017) 11(6):656–9. doi:10.1017/dmp.2017.23
40. Ocampo AJ, Chunara R, Brownstein JS. Using search queries for malaria surveillance, Thailand. Malar J (2013) 12:390. doi:10.1186/1475-2875-12-390
41. Adawi M, Bragazzi NL, Watad A, Sharif K, Amital H, Mahroum N. Discrepancies between classic and digital epidemiology in searching for the Mayaro virus: preliminary qualitative and quantitative analysis of Google trends. JMIR Public Health Surveill (2017) 3(4):e93. doi:10.2196/publichealth.9136
42. Bragazzi NL, Bacigaluppi S, Robba C, Siri A, Canepa G, Brigo F. Infodemiological data of West-Nile virus disease in Italy in the study period 2004-2015. Data Brief (2016) 9:839–45. doi:10.1016/j.dib.2016.10.022
43. Watad A, Watad S, Mahroum N, Higazi T, Brigo F, Igwe S, et al. Now-Casting/Forecasting the West-Nile Virus in the USA: An Extensive Novel Data Streams-Based Time Series Analysis and Structural Equation Modeling of Related Digital Searching Behavior. EPJ Data Science (2018) (in press).
44. Basch CH, Fung IC, Hammond RN, Blankenship EB, Tse ZT, Fu KW, et al. Zika virus on YouTube: an analysis of English-language video content by source. J Prev Med Public Health (2017) 50(2):133–40. doi:10.3961/jpmph.16.107
45. Bragazzi NL, Alicino C, Trucchi C, Paganino C, Barberis I, Martini M, et al. Global reaction to the recent outbreaks of Zika virus: insights from a Big Data analysis. PLoS One (2017) 12(9):e0185263. doi:10.1371/journal.pone.0185263
46. Dredze M, Broniatowski DA, Hilyard KM. Zika vaccine misconceptions: a social media analysis. Vaccine (2016) 34(30):3441–2. doi:10.1016/j.vaccine.2016.05.008
47. Fu KW, Liang H, Saroha N, Tse ZT, Ip P, Fung IC. How people react to Zika virus outbreaks on Twitter? A computational content analysis. Am J Infect Control (2016) 44(12):1700–2. doi:10.1016/j.ajic.2016.04.253
48. Glowacki EM, Lazard AJ, Wilcox GB, Mackert M, Bernhardt JM. Identifying the public’s concerns and the Centers for Disease Control and Prevention’s reactions during a health crisis: an analysis of a Zika live Twitter chat. Am J Infect Control (2016) 44(12):1709–11. doi:10.1016/j.ajic.2016.05.025
49. Lehnert JD, Ellingson MK, Goryoka GW, Kasturi R, Maier E, Chamberlain AT. Use of obstetric practice web sites to distribute Zika virus information to pregnant women during a Zika virus outbreak. J Public Health Manag Pract (2017) 23(6):608–13. doi:10.1097/PHH.0000000000000537
50. Majumder MS, Santillana M, Mekaru SR, McGinnis DP, Khan K, Brownstein JS. Utilizing nontraditional data sources for near real-time estimation of transmission dynamics during the 2015-2016 Colombian Zika Virus disease outbreak. JMIR Public Health Surveill (2016) 2(1):e30. doi:10.2196/publichealth.5814
51. McGough SF, Brownstein JS, Hawkins JB, Santillana M. Forecasting Zika incidence in the 2016 Latin America outbreak combining traditional disease surveillance with search, social media, and news report data. PLoS Negl Trop Dis (2017) 11(1):e0005295. doi:10.1371/journal.pntd.0005295
52. Miller M, Banerjee T, Muppalla R, Romine W, Sheth A. What are people tweeting about Zika? An exploratory study concerning its symptoms, treatment, transmission, and prevention. JMIR Public Health Surveill (2017) 3(2):e38. doi:10.2196/publichealth.7157
53. Seltzer EK, Horst-Martz E, Lu M, Merchant RM. Public sentiment and discourse about Zika virus on Instagram. Public Health (2017) 150:170–5. doi:10.1016/j.puhe.2017.07.015
54. Sharma M, Yadav K, Yadav N, Ferdinand KC. Zika virus pandemic-analysis of Facebook as a social media health information platform. Am J Infect Control (2017) 45(3):301–2. doi:10.1016/j.ajic.2016.08.022
55. Southwell BG, Dolina S, Jimenez-Magdaleno K, Squiers LB, Kelly BJ. Zika virus-related news coverage and online behavior, United States, Guatemala, and Brazil. Emerg Infect Dis (2016) 22(7):1320–1. doi:10.3201/eid2207.160415
56. Stefanidis A, Vraga E, Lamprianidis G, Radzikowski J, Delamater PL, Jacobsen KH, et al. Zika in Twitter: temporal variations of locations, actors, and concepts. JMIR Public Health Surveill (2017) 3(2):e22. doi:10.2196/publichealth.6925
57. Teng Y, Bi D, Xie G, Jin Y, Huang Y, Lin B, et al. Dynamic forecasting of Zika epidemics using Google trends. PLoS One (2017) 12(1):e0165085. doi:10.1371/journal.pone.0165085
58. Vijaykumar S, Meurzec RW, Jayasundar K, Pagliari C, Fernandopulle Y. What’s buzzing on your feed? Health authorities’ use of Facebook to combat Zika in Singapore. J Am Med Inform Assoc (2017) 24(6):1155–9. doi:10.1093/jamia/ocx028
59. Gould E, Pettersson J, Higgs S, Charrel R, de Lamballerie X. Emerging arboviruses: why today? One Health (2017) 4:1–13. doi:10.1016/j.onehlt.2017.06.001
60. Cooper KM, Bastola DR, Gandhi R, Ghersi D, Hinrichs S, Morien M, et al. Forecasting the spread of mosquito-borne disease using publicly accessible data: a case study in Chikungunya. AMIA Annu Symp Proc (2016) 2016:431–40.
Keywords: big data, Zika, Ebola, Chikungunya, West Nile virus, dengue, Mayaro virus, communicable tropical diseases
Citation: Gianfredi V, Bragazzi NL, Nucci D, Martini M, Rosselli R, Minelli L and Moretti M (2018) Harnessing Big Data for Communicable Tropical and Sub-Tropical Disorders: Implications From a Systematic Review of the Literature. Front. Public Health 6:90. doi: 10.3389/fpubh.2018.00090
Received: 20 December 2017; Accepted: 07 March 2018;
Published: 21 March 2018
Edited by:
Pierpaolo Cavallo, Università degli Studi di Salerno, ItalyReviewed by:
Greg Morrison, University of Houston, United StatesMonica Catarina Botelho, Instituto Nacional de Saúde Doutor Ricardo Jorge (INSA), Portugal
Copyright: © 2018 Gianfredi, Bragazzi, Nucci, Martini, Rosselli, Minelli and Moretti. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Vincenza Gianfredi, dmluY2VuemEuZ2lhbmZyZWRpQHN0dWRlbnRpLnVuaXBnLml0
†These authors have contributed equally to this work.
 Vincenza Gianfredi
Vincenza Gianfredi Nicola Luigi Bragazzi
Nicola Luigi Bragazzi Daniele Nucci
Daniele Nucci Mariano Martini
Mariano Martini Roberto Rosselli
Roberto Rosselli Liliana Minelli
Liliana Minelli Massimo Moretti
Massimo Moretti