- Guangzhou Jinan Biomedicine Research and Development Center, College of Life Science and Technology, Jinan University, Guangzhou, China
The outbreaks of COVID-19 due to SARS-CoV-2 has caused serious physical and psychological damage to global human health. COVID-19 spread rapidly around the world in a short time. Confronted with such a highly infectious respiratory disease, the research and development of anti-COVID-19 drugs became an urgent work due to the lack of specific drugs for the treatment of COVID-19. Nevertheless, several existing drugs are available to relieve the clinical symptoms of COVID-19. We reviewed information on selected anti-SARS-CoV-2 candidate therapeutic agents published until June 2, 2020. We also discussed the strategies of the development of anti-COVID-19 drugs in the future. Our review provides a novel insight into the future development of a safer, efficient, and toxic-less anti-COVID-19 drug.
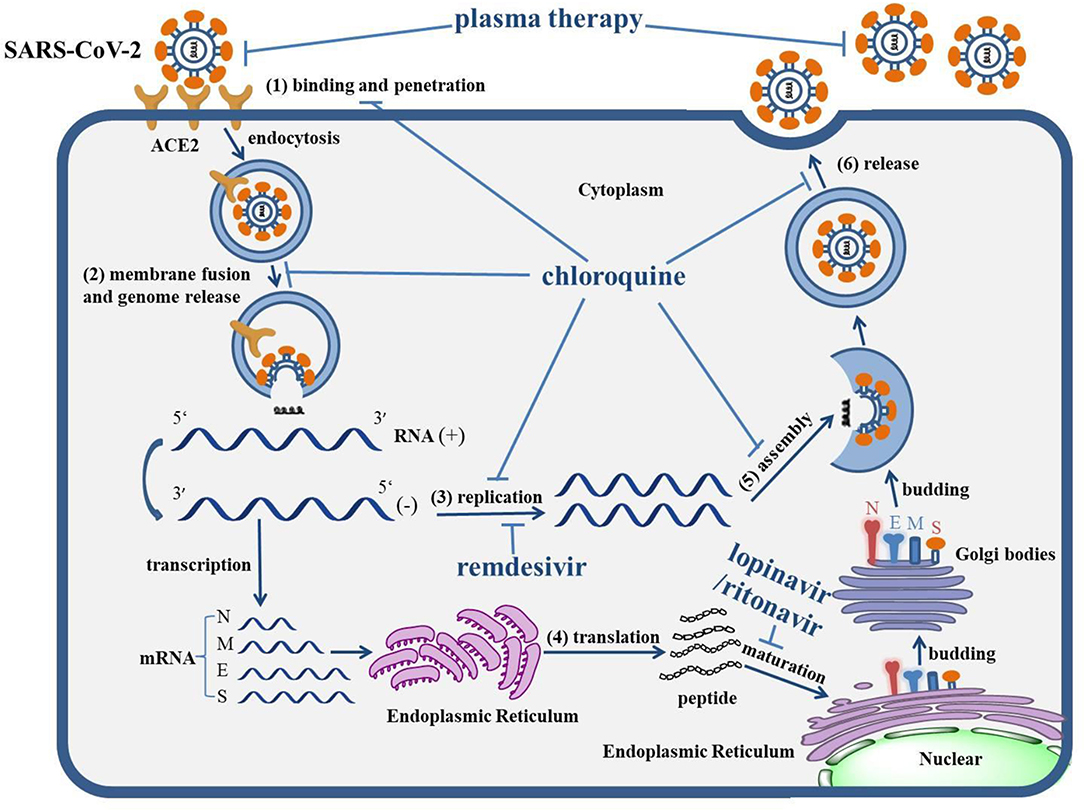
Graphical Abstract. The life cycle of SARS-CoV-2 and the mechanism of actions of anti-COVID-19 drugs. The life cycle of SARS-CoV-2 in host cells includes: (1) Binding and penetration: SARS-CoV-2 binds to the ACE2 receptors on the cell membrane and entry into host cells through endocytosis. (2) Genome release: the genome of SARS-CoV-2 will be released following the process of membrane fusion. (3) Genome replication: the positive (+)-sense genomic RNA directs the synthesis of negative (−)-sense RNA, which can act as the template to synthesize the RNA chain of progeny virus. (4) Protein biosynthesis: negative (−)-sense RNA acts as a template, with mRNAs transcribed to direct the protein biosynthesis of SARS-CoV-2 via the translation process in the cytoplasm. (5) Assembly: the genomic RNA and virion proteins are reassembled to form a mature virion. (6) Release: the progeny viral particles are released through exocytosis. The levels of actions of corresponding drugs were also depicted. Remdesivir can inhibit the replication of the SARS-CoV-2 genome. The combination of lopinavir/ritonavir can block the maturation of protein. Chloroquine virtually interrupts the whole life cycle of SARS-CoV-2. The antibody within plasma can directly neutralize SARS-CoV-2.
Key Points
- The outbreak of COVID-19 caused by SARS-CoV-2 has presented a challenge to global human health. However, there is no specific drug against COVID-19. It is imperative to summarize the mechanism of action and the therapeutic effect of currently used drugs. Moreover, the side effects of existing drugs against COVID-19 need to be recognized.
- Based on the clinical effects and characteristics of existing drugs, the strategies to develop toxic-less, and more effective anti-COVID-19 drugs were also summarized and posed.
Introduction
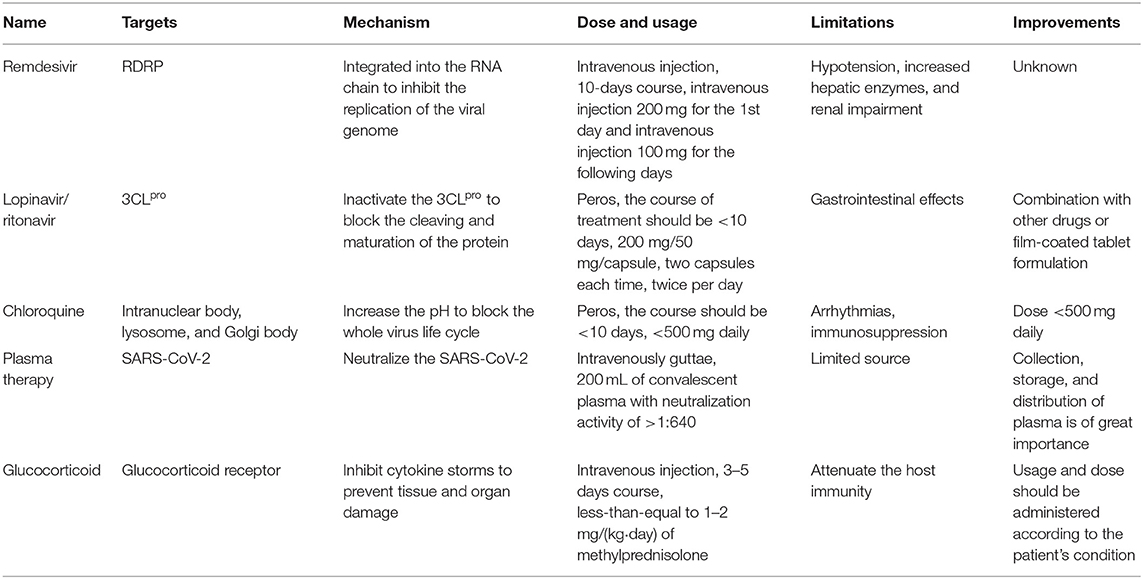
It was reported that a group of patients with pneumonia from an unknown cause were hospitalized at the end of 2019 (1). Most of them had respiratory symptoms, such as fever, cough, muscle soreness, headache, sore throat, chest pain, diarrhea, nausea, and vomiting, and among whom some even developed the complication of acute respiratory distress syndrome (ARDS) (2). This pneumonia was caused by a novel coronavirus termed severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), a sister of SARS-CoV, as revealed by the subsequent result of gene sequencing (3). The corresponding disease caused by SARS-CoV-2 was named 2019 coronavirus disease (COVID-19) (4). Of note, SARS-CoV-2 exhibited a high level of person-to-person transmission (5), which may be due to the strong affinity with its receptor Angiotensin-Converting Enzyme 2 (ACE2) (6). The ongoing outbreak of COVID-19 had been announced as a global pandemic by the WHO on 11 March, 2020. According to the report from the Center for Systems Science and Engineering at Johns Hopkins University (last updated on 4/26/2020), the global cumulative number of confirmed cases of COVID-19 has reached 2,856,771, with 202,473 deaths (7). Confronted with such a situation, the development of effective drugs for the treatment of COVID-19 has become a crucial and urgent work (8). However, there is currently no specific drugs against SARS-CoV-2, despite some treatments that had been used in the clinical treatment of COVID-19 (9). The mechanisms of actions of these drugs need to be elucidated and discussed. In this review, we summarized and discussed the currently available clinical treatment measures according to their mechanism of action and therapeutic effect (Table 1). The studies presented in this review were obtained from Google Scholar search engines and the PubMed database from searches up to June 2, 2020. Search terms include “COVID-19,” “SARS-CoV-2,” “anti-COVID-19 drugs,” “COVID-19 clinical trials,” and “development strategies of anti-COVID-19 drugs” in abstract, title, and keywords. The strategies of the future development of anti-COVID-19 drugs were also discussed. Our review would be beneficial for the development of more effective and toxic-less anti-COVID-19 drugs.
Remdesivir
The nucleoside analogs are important reagents for combating virus infection (10). As one of the well-characterized adenosine analogs, remdesivir can restrain the proliferation of SARS-CoV, MERS-CoV, and Ebola virus in vitro (11). Remdesivir can be integrated into the RNA chain of progeny virus as the substrate of the RNA-dependent RNA polymerase (RdRp), which inhibits the replication of viral genomes and thereby causes the mature termination of the virus (12). It has also been verified that remdesivir can strongly interfere with the accomplishment of the SARS-CoV-2 life cycle in host cells (13). The latest report indicated a clinical improvement of severe COVID-19 patients from multiple countries in 36 of 53 patients (68%) after treatment with remdesivir (14). Due to such an excellent efficacy, remdesivir has entered into multiple clinical trials (15). The results of a randomized, double-blind, placebo-controlled, multicentre trial suggested that whether intravenous remdesivir could decrease the time to clinical improvement in those treated earlier needs to be confirmed by further clinical studies. However, no statistically significant clinical benefits were observed in the remdesivir group compared with the placebo group in this clinical trial (16). Indeed, a patient with COVID-19 successfully recovered after receiving remdesivir intravenously in the United States (17), which further indicates that remdesivir would be rapidly applied as a clinical treatment for COVID-19 in the future. However, remdesivir has been found to cause side effects in the clinic, such as hypotension, increased hepatic enzymes, and renal impairment (14). The mechanism responsible for the side effects of remdesivir is not clear. Further study is needed to address the mechanism of the side effects caused by remdesivir. Collectively, remdesivir is a relatively promising anti-SARS-COV-2 candidate therapeutic agent (18).
Lopinavir/Ritonavir
Lopinavir is an inhibitor of Human Immunodeficiency Virus 1 (HIV-1) protease (19). The metabolism of lopinavir can be delayed by ritonavir to enhance the anti-HIV-1 effect of lopinavir; therefore, these two drugs are often used in combination (20). The brand name of such a combined drug is Kaletra (21), which displays a broad-spectrum antiviral activity, including on SARS-CoV-2 (22). Mechanism studies suggested that the lopinavir/ritonavir combination may inactivate the 3-chymotrypsin-like cysteine protease (3CLpro) that cleaves protein precursors into a variety of active proteins required for the life cycle of SARS-CoV-2 (23). A non-comparative case series of 10 patients suggested that lopinavir may ameliorate the symptoms of COVID-19 (24). After receiving lopinavir/ritonavir with arbidol combination therapy, the negative conversion rate of COVID-19 on the 7 and 14th days was significantly increased (25). Indeed, the viral load of a COVID-19 patient who received lopinavir/ritonavir combination therapy was gradually decreased and even completely cleared within the next few days in Korea (26). A retrospective analysis further supported that lopinavir is an effective drug for the treatment of COVID-19 (27). However, no benefit was observed in COVID-19 patients who were receiving lopinavir/ritonavir combination therapy as revealed by a randomized, controlled, open-label trial (28). Importantly, lopinavir/ritonavir combination (200 mg/50 mg/capsule, two capsules each time, twice per day for adults, the course of treatment should be <10 days) was recommended for the treatment of COVID-19 by the National Health Commission of China. However, the lopinavir/ritonavir combination can induce severe gastrointestinal effects for the treatment of COVID-19, the cause of which remains unknown (28). Of note, the lopinavir/ritonavir combination can be used in combination with other drugs to alleviate adverse reactions, such as probiotics, soluble fiber, and L-Glutamine (GLN) (29). Besides, the film-coated tablet formulation of lopinavir/ritonavir induces fewer gastrointestinal side effects than when used in tablet formulation (30).
Chloroquine
Chloroquine is a cheap and safe drug that has been used in the clinic for more than 70 years (31). Chloroquine is a first-line drug for the treatment of Plasmodium falciparum infection (32). Importantly, chloroquine also exerts strong antiviral effects (33). Mechanically, chloroquine can increase the pH of the intranuclear body, lysosome, and Golgi body, which jointly prevents virus penetration, genome replication, and assembly of mature viral particles (34). Of note, it was confirmed that chloroquine can suppress the replication of SARS-CoV-2 in vitro with an EC50 of 1.13 μM (13). Interim analysis of preliminary data from 23 ongoing clinical trials reported in a letter suggested that chloroquine phosphate is superior to the control treatment in inhibiting the exacerbation of COVID-19 pneumonia (35). Hydroxychloroquine, a derivative of chloroquine, also can significantly inhibit the infection of SARS-CoV-2 on VeroE6 cells with weak toxicity (36). An uncontrolled, non-comparative, observational study in a cohort of 80 inpatients reported clinical improvements and rapid fall of viral load after receiving hydroxychloroquine and azithromycin combination therapy (37). However, the administration of hydroxychloroquine alone did not significantly increase the negative conversion rate in COVID-19 patients (38). Indeed, chloroquine phosphate is recommended as an effective treatment by the National Health Commission of the People's Republic of China. There are more than 16 clinical trials aimed at determining the effectiveness of chloroquine in the treatment of COVID-19 (39). If the result of the clinical trial supports the efficacy and safety of chloroquine against COVID-19, chloroquine will become one of the most available drugs for the treatment of COVID-19 (40). Specifically, chloroquine and hydroxyquinoline could impair host immunity by inhibiting toll-like receptor 7 (TLR7) and toll-like receptor 9 (TLR9) signaling due to the increased pH (41). In particular, a high dose of chloroquine and hydroxyquinoline can cause arrhythmias and even death by interfering with the polarization and depolarization of the heart (42). Indeed, it was recommended that <500 mg of chloroquine and hydroxyquinoline is used as a daily dose for adults and it is not advocated for long-term use; if long-term use is needed, the toxic-less hydroxychloroquine should be given priority (43).
Plasma Therapy
The antibodies against SARS-CoV-2 produced by plasma cells can neutralize the virus to reduce its pathogenicity (44). Scientists have been devoted to the development of antibodies against SARS-CoV-2 since the outbreak of COVID-19 (45). SARS-CoV-2 contains four conserved structural proteins—the spike (S) protein, the membrane (M) protein, the nucleocapsid (N) protein, and the small envelope (E) protein (46)—in which the S protein shows excellent antigenicity (47). Of note, the SARS-specific human monoclonal antibody CR3022 can bind to the S protein of SARS-CoV-2 as determined by enzyme-linked immunosorbent assay (ELISA) and biolayer interferometry binding (BLI) assay, whereas the clinical efficacy of CR3022 needs to be further verified (48). Additionally, it has been reported that the antibodies within convalescent plasma can neutralize SARS-CoV-2 efficiently and rapidly (49). As revealed by a clinical trial with a small sample size, the convalescent plasma may be a potential treatment for COVID-19 patients who were admitted to the intensive care unit (ICU) (50). Similarly, another clinical trial also revealed the remarkable efficacy and feasibility of plasma therapy for the treatment of COVID-19 (51). However, plasma therapy is limited by the shortage of its sources because an ideal therapeutic plasma should be compatible with the recipients (52). Therefore, the collection, storage, and distribution of plasma would be crucial work for the development of plasma therapy (52). Collectively, plasma therapy for the treatment of COVID-19 patients with systemic, severe, and critical conditions requires confirmation in larger studies.
Glucocorticoid
Glucocorticoid, also known as an adrenocortical hormone, is a steroid hormone secreted by the human adrenal gland (53). As one of the most important physiological hormones, glucocorticoids can regulate the biosynthesis and metabolism of the host (54). Significantly, glucocorticoid also shows a strong activity of anti-inflammation (55). However, the long-term use of glucocorticoids also induces severe side effects, such as the increased risk of osteonecrosis, endocrine disorders, and heart failure (56). During the outbreak of SARS in 2003, the clinical application of glucocorticoids was an inevitable choice for critically ill patients in China (57). Glucocorticoids can inhibit cytokine storms and chemokines caused by SARS-CoV-2 to prevent acute lung injury and acute respiratory distress syndrome (58). Although the clinical evidence does not support glucocorticoid treatment for SARS-CoV-2 infection (59), glucocorticoid can serve as adjuvant therapy for critical patients with COVID-19 (60). The 6th edition of the Diagnosis and treatment plan of Corona Virus Disease 2019 recommended glucocorticoid [≤1–2 mg/(kg·day) of methylprednisolone] as an alternative therapy. Of note, glucocorticoid can attenuate the host immunity by inhibiting toll-like receptor 4 (TLR4) signaling and T cell activation, which may cause the secondary infection of other pathogens (61). However, such side effects can be partly restored by the combination of thalidomide and glucocorticoid with a reduced dose of glucocorticoids (62). Therefore, the usage and dose should be administered moderately according to the patient's condition when glucocorticoid is used to relieve inflammation of COVID-19 patients. Collectively, further randomized controlled trials are needed to determine the safety and feasibility of glucocorticoids in relieving inflammatory symptoms of COVID-19 patients (63).
Other Treatments
In addition to the drugs mentioned above, some drugs with fewer reports showed the activity of anti-SARS-CoV-2. Specifically, cinanserin, another HIV-1 protease inhibitor, may be a potential drug against COVID-19 as indicated by molecular docking and antiviral activity assay (64). It has been recently reported that baricitinib is also a potential option for COVID-19 patients via blocking the ACE2 receptor-mediated endocytosis, although its efficacy remains to be clinically tested (65). Oseltamivir has been widely used for COVID-19 patients although the therapeutic effect on COVID-19 remains to be further explored (66). Some nucleoside analogs, including favipiravir, penciclovir, and ribavirin, can significantly inhibit the proliferation of SARS-CoV-2 in vitro (13). Among these, ribavirin can inhibit the replication of both DNA and RNA viruses (67). The combination of ribavirin and lopinavir/ritonavir or interferon can be used to treat COVID-19 as recommended by the 6th edition of Diagnosis and treatment plan of Corona Virus Disease 2019. Arbidol can prevent viral replication by interrupting the virus life cycle and enhancing the immune response (68). The clinical trial of arbidol in the treatment of COVID-19 (NCT04246242) has been registered. Besides, interferon is a broad-spectrum antiviral factor secreted by the host upon the invasion of pathogens. SARS-CoV-2 was more sensitive to interferon than SARS-CoV in vitro (69). IFN-α spray is also recommended for the treatment of COVID-19 according to 6th edition of the Diagnosis and treatment plan of Corona Virus Disease 2019. Viral genome editing is also an emerging therapeutic strategy for combating SARS-CoV-2. For example, the CRISPR/Cas13d system delivered by an adeno-associated virus (AAV) serotypes harboring a high affinity with the lungs can accurately excise the genome of SARS-CoV-2 (70). Traditional Chinese medicine is also a tremendous source for anti-COVID-19 drugs (71). For instance, lianhuaqingwen can inhibit the proliferation of SARS-CoV-2 and reduce the induction of inflammatory factors by SARS-CoV-2 (72).
Conclusions and Future Perspective
The outbreak of COVID-19 raises a serious challenge to the global economy and human health; therefore, the development of an effective treatment for COVID-19 has become an urgent work (73). Indeed, there has been no specific drug against coronavirus since the outbreak of SARS in 2003. The drugs currently used for the treatment of COVID-19 partly refer to those for the treatment of SARS and MERS (74), which are still assessed in clinical trials. We should pay attention to the following aspects in the future development of anti-COVID-19 drugs.
For one thing, the drugs capable of blocking any step of the virus life cycle can be designed as antiviral drugs. For example, camostat mesylate can significantly block the penetration of SARS-CoV-2 by inhibiting the activity of the serine protease TMPRSS2, which is a factor mediating the penetration of SARS-CoV-2 (75). EK1, a pan-coronavirus fusion inhibitor, can interfere with the membrane fusion of SARS-CoV-2 with the host cell by targeting the S protein (76). Similar to remdesivir, sofosbuvir, galidesivir, and tenofovir may also act as the substrate of RdRp to inhibit the SARS-CoV-2 genome replication as revealed by the molecular docking results (77). The 3CLpro enzyme is responsible for cleaving polymeric protein precursors to produce many non-structural proteins that are required for the replication of SARS-CoV-2, which indicates that 3CLpro inhibitors, including celecoxib and alprazolam, can be used to combat COVID-19 (78). Of note, based on targeting the ACE2 receptor, a previous study had discovered several potential anti-COVID-19 drugs by using computational methods, such as xanthones and hesperidin (79). Indeed, the lack of specific drugs for combating SARS-CoV-2 was largely due to the incomprehensive recognition of the mechanism of SARS-CoV-2 infection in host cells (80). Therefore, future work should focus on exploring the life cycle of SARS-CoV-2 in human cells and the detailed mechanism of the pathogenesis of SARS-CoV-2.
For another, the alleviation of host inflammation is an essential and urgent work for COVID-19 patients with cytokine storm (81). Of note, in addition to glucocorticoids mentioned above, there are other agents with an anti-inflammatory effect, such as tocilizumab and jakotinib, an interleukin-6-receptor antagonist and Janus kinase (JAK) inhibitor, respectively (82). Further, screening from FDA-approved drugs based on computational methods would be an ideal strategy to ensure the efficiency of anti-COVID-19 drugs development (83). Indeed, the vaccine is crucial for the prevention and control of COVID-19. Scientists need to have a better understanding of the interaction between SARS-CoV-2 and the immune system. The S protein is an ideal antigen for the development of vaccines due to its high affinity with the ACE2 receptor (84). In particular, the receptor-binding domain (RBD) element of the S protein may be applied to vaccine development (85).
Author Contributions
LH participated in data collection, data analysis, drafting, and editing of the manuscript. YilW contributed to the critical review and revision of the manuscript. All authors reviewed, supported the final manuscript, and agreed to this publication.
Funding
This review was supported by the grant of Research and Development and Industrialization of Artemisia Argyi-derived disinfectant for the prevention and control of COVID-19 of universities in Guangdong provinces (No. 2020KZDZX1025).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
ARDS, acute respiratory distress syndrome; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; COVID-19, 2019 coronavirus disease; ACE2, Angiotensin-Converting Enzyme 2; RdRp, RNA-dependent RNA polymerase; HIV-1, Human Immunodeficiency Virus 1; 3CLpro, 3-chymotrypsin-like cysteine protease; GLN, L-Glutamine; TLR7, toll-like receptor 7; TLR9, toll-like receptor 9; S, spike; M, membrane; N, nucleocapsid; E, envelope; ELISA, enzyme-linked immunosorbent assay; BLI, biolayer interferometry binding; ICU, intensive care unit; TLR4, toll-like receptor 4; AAV, adeno-associated virus; JAK, Janus kinase; RBD, receptor-binding domain.
References
1. Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. (2020) 382:727–33. doi: 10.1056/NEJMoa2001017
2. Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. (2020) 395:507–13. doi: 10.1016/S0140-6736(20)30211-7
3. Jiang S, Shi Z, Shu Y, Song J, Gao GF, Tan W, et al. A distinct name is needed for the new coronavirus. Lancet. (2020) 395:949. doi: 10.1016/S0140-6736(20)30419-0
4. Chang L, Yan Y, Wang L. Coronavirus disease 2019: coronaviruses and blood safety. Transfusion Med Rev. (2020) 34:75–80. doi: 10.1016/j.tmrv.2020.02.003
5. Xu X, Chen P, Wang J, Feng J, Zhou H, Li X, et al. Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci China Life Sci. (2020) 63:457–60. doi: 10.1007/s11427-020-1637-5
6. Luan J, Lu Y, Jin X, Zhang L. Spike protein recognition of mammalian ACE2 predicts the host range and an optimized ACE2 for SARS-CoV-2 infection. Biochem Biophys Res Commun. (2020) 526:165–9. doi: 10.1016/j.bbrc.2020.03.047
7. Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infectious Dis. (2020) 20:533–4. doi: 10.1016/S1473-3099(20)30120-1
8. Sohrabi C, Alsafi Z, O'Neill N, Khan M, Kerwan A, Al-Jabir A, et al. World Health Organization declares global emergency: a review of the 2019 novel coronavirus (COVID-19). Int J Surgery. (2020) 76:71–6. doi: 10.1016/j.ijsu.2020.02.034
9. Han Q, Lin Q, Jin S, You L. Recent insights into 2019-nCoV: a brief but comprehensive review. J Infection. (2020) 80:373–7. doi: 10.1016/j.jinf.2020.02.010
10. Pruijssers AJ, Denison MR. Nucleoside analogues for the treatment of coronavirus infections. Curr Opin Virol. (2019) 35:57–62. doi: 10.1016/j.coviro.2019.04.002
11. Agostini ML, Andres EL, Sims AC, Graham RL, Sheahan TP, Lu X, et al. Coronavirus susceptibility to the antiviral remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. mBio. (2018) 9:e00221–18. doi: 10.1128/mBio.00221-18
12. Warren TK, Jordan R, Lo MK, Ray AS, Mackman RL, Soloveva V, et al. Therapeutic efficacy of the small molecule GS-5734 against Ebola virus in rhesus monkeys. Nature. (2016) 531:381–5. doi: 10.1038/nature17180
13. Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. (2020) 30:269–71. doi: 10.1038/s41422-020-0282-0
14. Grein J, Ohmagari N, Shin D, Diaz G, Asperges E, Castagna A, et al. Compassionate use of remdesivir for patients with severe covid-19. N Engl J Med. (2020) 382:2327–36. doi: 10.1056/NEJMc2015312
15. Singh AK, Singh A, Singh R, Misra A. Remdesivir in COVID-19: a critical review of pharmacology, pre-clinical and clinical studies. Diabetes Metabolic Syndrome Clin Res Rev. (2020) 14:641–8. doi: 10.1016/j.dsx.2020.05.018
16. Wang Y, Zhang D, Du G, Du R, Zhao J, Jin Y, et al. Remdesivir in adults with severe COVID-19: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet. (2020) 395:1569–78. doi: 10.1016/S0140-6736(20)31022-9
17. Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, et al. First case of 2019 novel coronavirus in the United States. N Engl J Med. (2020) 382:929–36. doi: 10.1056/NEJMoa2001191
18. Jean S-S, Lee P-I, Hsueh P-R. Treatment options for COVID-19: the reality and challenges. J Microbiol Immunol Infection. (2020) 53:436–43. doi: 10.1016/j.jmii.2020.03.034
19. Badaya A, Sasidhar YU. Inhibition of the activity of HIV-1 protease through antibody binding and mutations probed by molecular dynamics simulations. Sci Rep. (2020) 10:5501. doi: 10.1038/s41598-020-62423-y
20. Cvetkovic RS, Goa KL. Lopinavir/ritonavir: a review of its use in the management of HIV infection. Drugs. (2003) 63:769–802. doi: 10.2165/00003495-200363080-00004
21. Tobaiqy M, Qashqary M, Al-Dahery S, Mujallad A, Hershan AA, Kamal MA, et al. Therapeutic management of patients with COVID-19: a systematic review. Infection Prev Practice. (2020) 2:100061. doi: 10.1016/j.infpip.2020.100061
22. McKee DL, Sternberg A, Stange U, Laufer S, Naujokat C. Candidate drugs against SARS-CoV-2 and COVID-19. Pharmacol Res. (2020) 157:104859. doi: 10.1016/j.phrs.2020.104859
23. Nukoolkarn V, Lee VS, Malaisree M, Aruksakulwong O, Hannongbua S. Molecular dynamic simulations analysis of ritronavir and lopinavir as SARS-CoV 3CLpro inhibitors. J Theoretical Biol. (2008) 254:861–7. doi: 10.1016/j.jtbi.2008.07.030
24. Liu F, Xu A, Zhang Y, Xuan W, Yan T, Pan K, et al. Patients of COVID-19 may benefit from sustained lopinavir-combined regimen and the increase of eosinophil may predict the outcome of COVID-19 progression. Int J Infectious Dis. (2020) 95:183–91. doi: 10.1016/j.ijid.2020.03.013
25. Deng L, Li C, Zeng Q, Liu X, Li X, Zhang H, et al. Arbidol combined with LPV/r versus LPV/r alone against Corona Virus Disease 2019: a retrospective cohort study. J Infection. (2020) 81:e1–5. doi: 10.1016/j.jinf.2020.03.002
26. Lim J, Jeon S, Shin H-Y, Kim MJ, Seong YM, Lee WJ, et al. Case of the index patient who caused tertiary transmission of Coronavirus disease 2019 in Korea: the application of lopinavir/ritonavir for the treatment of COVID-19 pneumonia monitored by quantitative RT-PCR. J Korean Med Sci. (2020) 35:e88. doi: 10.3346/jkms.2020.35.e89
27. Yao TT, Qian JD, Zhu WY, Wang Y, Wang GQ. A systematic review of lopinavir therapy for SARS coronavirus and MERS coronavirus–a possible reference for coronavirus disease-19 treatment option. J Med Virol. (2020) 92:556–63. doi: 10.1002/jmv.25729
28. Cao B, Wang Y, Wen D, Liu W, Wang J, Fan G, et al. A trial of lopinavir-ritonavir in adults hospitalized with severe covid-19. N Engl J Med. (2020) 382:1787–99. doi: 10.1056/NEJMc2008043
29. Heiser CR, Ernst JA, Barrett JT, French N, Schutz M, Dube MP. Probiotics, soluble fiber, and L-Glutamine (GLN) reduce nelfinavir (NFV)- or lopinavir/ritonavir (LPV/r)-related diarrhea. J Int Association Physicians AIDS Care. (2004) 3:121–9. doi: 10.1177/154510970400300403
30. Sáez de la Fuente J, Granja V, Escobar I, Collada de la Fuente E, Moreno V, Rubio R. Study of the gastrointestinal tolerance of a new tablet formulation of the lopinavir/ritonavir antiretroviral in HIV-infected patients. J Acquired Immune Deficiency Syndromes. (2009) 50:294–8. doi: 10.1097/QAI.0b013e3181989a74
31. Trape J-F. The public health impact of chloroquine resistance in Africa. Am J Tropical Med Hygiene. (2001) 64:12–7. doi: 10.4269/ajtmh.2001.64.12
32. Bonilla-Ramirez L, Rios A, Quiliano M, Ramirez-Calderon G, Beltrán-Hortelano I, Franetich JF, et al. Novel antimalarial chloroquine-and primaquine-quinoxaline 1, 4-di-N-oxide hybrids: design, synthesis, Plasmodium life cycle stage profile, and preliminary toxicity studies. Eur J Med Chem. (2018) 158:68–81. doi: 10.1016/j.ejmech.2018.08.063
33. Savarino A, Boelaert JR, Cassone A, Majori G, Cauda R. Effects of chloroquine on viral infections: an old drug against today's diseases. Lancet Infectious Dis. (2003) 3:722–7. doi: 10.1016/S1473-3099(03)00806-5
34. Rolain J-M, Colson P, Raoult D. Recycling of chloroquine and its hydroxyl analogue to face bacterial, fungal and viral infections in the 21st century. Int J Antimicrobial Agents. (2007) 30:297–308. doi: 10.1016/j.ijantimicag.2007.05.015
35. Gao J, Tian Z, Yang X. Breakthrough: chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci Trends. (2020) 14:72–3. doi: 10.5582/bst.2020.01047
36. Liu J, Cao R, Xu M, Wang X, Zhang H, Hu H, et al. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov. (2020) 6:16. doi: 10.1038/s41421-020-0156-0
37. Gautret P, Lagier JC, Parola P, Hoang VT, Meddeb L, Sevestre J, et al. Clinical and microbiological effect of a combination of hydroxychloroquine and azithromycin in 80 COVID-19 patients with at least a 6-days follow up: a pilot observational study. Travel Med Infect Dis. (2020) 34:101663. doi: 10.1016/j.tmaid.2020.101663
38. Tang W, Cao Z, Han M, Wang Z, Chen J, Sun W, et al. Hydroxychloroquine in patients with mainly mild to moderate coronavirus disease 2019: open label, randomised controlled trial. BMJ. (2020) 369:m1849. doi: 10.1136/bmj.m1849
39. Touret F, de Lamballerie X. Of chloroquine and COVID-19. Antiviral Res. (2020) 177:104762. doi: 10.1016/j.antiviral.2020.104762
40. Colson P, Rolain JM, Raoult D. Chloroquine for the 2019 novel coronavirus SARS-CoV-2. Int J Antimicrobial Agents. (2020) 55:105923. doi: 10.1016/j.ijantimicag.2020.105923
41. Ponticelli C, Moroni G. Hydroxychloroquine in systemic lupus erythematosus (SLE). Expert Opin Drug Saf. (2017) 16:411–9. doi: 10.1080/14740338.2017.1269168
42. White NJ. Cardiotoxicity of antimalarial drugs. Lancet Infectious Dis. (2007) 7:549–58. doi: 10.1016/S1473-3099(07)70187-1
43. Wong YK, Yang J, He Y. Caution and clarity required in the use of chloroquine for COVID-19. Lancet Rheumatol. (2020) 2:e255. doi: 10.1016/S2665-9913(20)30093-X
44. Crowe JE. Jr Principles of broad and potent antiviral human antibodies: insights for vaccine design. Cell Host Microbe. (2017) 22:193–206. doi: 10.1016/j.chom.2017.07.013
45. Zhou G, Zhao Q. Perspectives on therapeutic neutralizing antibodies against the Novel Coronavirus SARS-CoV-2. Int J Biol Sci. (2020) 16:1718–23. doi: 10.7150/ijbs.45123
46. Chan JF, Kok KH, Zhu Z, Chu H, To KK, Yuan S, et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerging Microbes Infections. (2020) 9:221–36. doi: 10.1080/22221751.2020.1719902
47. Zheng M, Song L. Novel antibody epitopes dominate the antigenicity of spike glycoprotein in SARS-CoV-2 compared to SARS-CoV. Cell Mol Immunol. (2020) 17:536–8. doi: 10.1038/s41423-020-0385-z
48. Tian X, Li C, Huang A, Xia S, Lu S, Shi Z, et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerging Microbes Infections. (2020) 9:382–5. doi: 10.1080/22221751.2020.1729069
49. Chen L, Xiong J, Bao L, Shi Y. Convalescent plasma as a potential therapy for COVID-19. Lancet Infectious Dis. (2020) 20:398–400. doi: 10.1016/S1473-3099(20)30141-9
50. Shen C, Wang Z, Zhao F, Yang Y, Li J, Yuan J, et al. Treatment of 5 critically Ill patients with COVID-19 with convalescent plasma. JAMA. (2020) 323:1582–9. doi: 10.1001/jama.2020.4783
51. Duan K, Liu B, Li C, Zhang H, Yu T, Qu J, et al. Effectiveness of convalescent plasma therapy in severe COVID-19 patients. Proc Natl Acad Sci USA. (2020) 117:9490–6. doi: 10.1073/pnas.2007408117
52. Roback JD, Guarner J. Convalescent plasma to treat COVID-19: possibilities and challenges. JAMA. (2020) 323:1561–2. doi: 10.1001/jama.2020.4940
53. Kroon J, Pereira AM, Meijer OC. Glucocorticoid sexual dimorphism in metabolism: dissecting the role of sex hormones. Trends Endocrinol Metab. (2020) 31:357–67. doi: 10.1016/j.tem.2020.01.010
54. Scherholz ML, Schlesinger N, Androulakis IP. Chronopharmacology of glucocorticoids. Adv Drug Delivery Rev. (2019) 151–152:245–61. doi: 10.1016/j.addr.2019.02.004
55. Coutinho AE, Chapman KE. The anti-inflammatory and immunosuppressive effects of glucocorticoids, recent developments and mechanistic insights. Mol Cell Endocrinol. (2011) 335:2–13. doi: 10.1016/j.mce.2010.04.005
56. Oray M, Abu Samra K, Ebrahimiadib N, Meese H, Foster CS. Long-term side effects of glucocorticoids. Expert Opin Drug Safety. (2016) 15:457–65. doi: 10.1517/14740338.2016.1140743
57. Zhao R, Wang H, Wang X, Feng F. Steroid therapy and the risk of osteonecrosis in SARS patients: a dose-response meta-analysis. Osteoporosis Int. (2017) 28:1027–34. doi: 10.1007/s00198-016-3824-z
58. Sanders JM, Monogue ML, Jodlowski TZ, Cutrell JB. Pharmacologic treatments for coronavirus disease 2019 (COVID-19): a review. JAMA. (2020) 323:1824–36. doi: 10.1001/jama.2020.6019
59. Russell CD, Millar JE, Baillie JK. Clinical evidence does not support corticosteroid treatment for 2019-nCoV lung injury. Lancet. (2020) 395:473–5. doi: 10.1016/S0140-6736(20)30317-2
60. Zhou W, Liu Y, Tian D, Wang C, Wang S, Cheng J, et al. Potential benefits of precise corticosteroids therapy for severe 2019-nCoV pneumonia. Signal Transduction Targeted Therapy. (2020) 5:18. doi: 10.1038/s41392-020-0127-9
61. Wang H, Ding Y, Li X, Yang L, Zhang W, Kang W. Fatal aspergillosis in a patient with SARS who was treated with corticosteroids. N Engl J Med. (2003) 349:507–8. doi: 10.1056/NEJM200307313490519
62. Kumar S, Zhi K, Mukherji A, Gerth K. Repurposing antiviral protease inhibitors using extracellular vesicles for potential therapy of COVID-19. Viruses. (2020) 12:E486. doi: 10.3390/v12050486
63. Shang L, Zhao J, Hu Y, Du R, Cao B. On the use of corticosteroids for 2019-nCoV pneumonia. Lancet. (2020) 395:683–4. doi: 10.1016/S0140-6736(20)30361-5
64. Jin Z, Du X, Xu Y. Structure of M(pro) from SARS-CoV-2 and discovery of its inhibitors. Nature. (2020) 582:289–93. doi: 10.1038/s41586-020-2223-y
65. Richardson P, Griffin I, Tucker C, Smith D, Oechsle O, Phelan A, et al. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet. (2020) 395:e30–1. doi: 10.1016/S0140-6736(20)30304-4
66. Lu H. Drug treatment options for the 2019-new coronavirus (2019-nCoV). Biosci Trends. (2020) 14:69–71. doi: 10.5582/bst.2020.01020
67. Sidwell RW, Huffman JH, Khare LGP, Allen B, Witkowski RJT, Robins K. Broad-spectrum antiviral activity of virazole: 1-f8- D-Ribofuranosyl- 1,2,4-triazole- 3-carboxamide. Science. (1972) 177:705–6. doi: 10.1126/science.177.4050.705
68. Blaising J, Polyak SJ, Pécheur EI. Arbidol as a broad-spectrum antiviral: an update. Antiviral Res. (2014) 107:84–94. doi: 10.1016/j.antiviral.2014.04.006
69. Mantlo E, Bukreyeva N, Maruyama J, Paessler S, Huang C. Antiviral activities of type I interferons to SARS-CoV-2 infection. Antiviral Res. (2020) 179:104811. doi: 10.1016/j.antiviral.2020.104811
70. Nguyen TM, Zhang Y, Pandolfi PP. Virus against virus: a potential treatment for 2019-nCov (SARS-CoV-2) and other RNA viruses. Cell Res. (2020) 30:189–90. doi: 10.1038/s41422-020-0290-0
71. Ling CQ. Traditional Chinese medicine is a resource for drug discovery against 2019 novel coronavirus (SARS-CoV-2). J Integrative Med. (2020) 18:87–8. doi: 10.1016/j.joim.2020.02.004
72. Runfeng L, Yunlong H, Jicheng H, Weiqi P, Qinhai M, Yongxia S, et al. Lianhuaqingwen exerts anti-viral and anti-inflammatory activity against novel coronavirus (SARS-CoV-2). Pharmacol Res. (2020) 156:104761. doi: 10.1016/j.phrs.2020.104761
73. Khan Z, Karataş Y, Rahman H. Anti COVID-19 drugs: need for more clinical evidence and global action. Adv Therapy. (2020) 37:2575–9. doi: 10.1007/s12325-020-01351-9
74. Yang Y, Peng F, Wang R, Guan K, Jiang T, Xu G, et al. The deadly coronaviruses: the 2003 SARS pandemic and the 2020 novel coronavirus epidemic in China. J Autoimmunity. (2020) 109:102434. doi: 10.1016/j.jaut.2020.102434
75. Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. (2020) 181:271–80. doi: 10.1016/j.cell.2020.02.052
76. Xia S, Liu M, Wang C, Xu W, Lan Q, Feng S, et al. Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. (2020) 30:343–55. doi: 10.1038/s41422-020-0305-x
77. Elfiky AA. Ribavirin, Remdesivir, Sofosbuvir, Galidesivir, and Tenofovir against SARS-CoV-2 RNA dependent RNA polymerase (RdRp): a molecular docking study. Life Sci. (2020) 253:117592. doi: 10.1016/j.lfs.2020.117592
78. Gimeno A, Mestres-Truyol J, Ojeda-Montes MJ, Macip G, Saldivar-Espinoza B, Cereto-Massagué A, et al. Prediction of novel inhibitors of the main protease (M-pro) of SARS-CoV-2 through consensus docking and drug reposition. Int J Mol Sci. (2020) 21:E3793. doi: 10.3390/ijms21113793
79. Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y, et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharmaceutica Sinica B. (2020) 10:766–88. doi: 10.1016/j.apsb.2020.02.008
80. Rismanbaf A. Potential treatments for COVID-19; a narrative literature review. Archiv Academic Emergency Med. (2020) 8:e29. doi: 10.22037/aaem.v8i1.596
81. Stebbing J, Phelan A, Griffin I, Tucker C, Oechsle O, Smith D, et al. COVID-19: combining antiviral and anti-inflammatory treatments. Lancet Infectious Dis. (2020) 20:400–2. doi: 10.1016/S1473-3099(20)30132-8
82. Zhang W, Zhao Y, Zhang F, Wang Q, Li T, Liu Z, et al. The use of anti-inflammatory drugs in the treatment of people with severe coronavirus disease 2019 (COVID-19): the experience of clinical immunologists from China. Clin Immunol. (2020) 214:108393. doi: 10.1016/j.clim.2020.108393
83. Sukumar N, Das S. Current trends in virtual high throughput screening using ligand-based and structure-based methods. Combinatorial Chem High Throughput Screening. (2011) 14:872–88. doi: 10.2174/138620711797537120
84. Walls AC, Park Y-J, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. (2020) 181:281–92. doi: 10.1016/j.cell.2020.02.058
Keywords: COVID-19, SARS-CoV-2, anti-COVID-19 drugs, remdesivir, lopinavir/ritonavir, chloroquine, plasma therapy, glucocorticoid
Citation: Huang L, Chen Y, Xiao J, Luo W, Li F, Wang Y, Wang Y and Wang Y (2020) Progress in the Research and Development of Anti-COVID-19 Drugs. Front. Public Health 8:365. doi: 10.3389/fpubh.2020.00365
Received: 26 April 2020; Accepted: 25 June 2020;
Published: 07 July 2020.
Edited by:
Herbert Leonel de Matos Guedes, Federal University of Rio de Janeiro, BrazilReviewed by:
Zuliang Jie, University of Texas MD Anderson Cancer Center, United StatesZakir Khan, Çukurova University, Turkey
Copyright © 2020 Huang, Chen, Xiao, Luo, Li, Wang, Wang and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yiliang Wang, d2FuZ195aWxpYW5nQHNpbmEuY24=; Yifei Wang, dHdhbmcteWZAMTYzLmNvbQ==
†These authors have contributed equally to this work
 Lianzhou Huang
Lianzhou Huang Yuanqiu Chen†
Yuanqiu Chen† Yifei Wang
Yifei Wang