- 1Department of Veterinary Public Health and Pharmacology, Faculty of Veterinary Medicine and Animal Science, University of Peradeniya, Peradeniya, Sri Lanka
- 2Department of Infectious Diseases and Immunology, Utrecht University, Utrecht, Netherlands
- 3Department of Microbiology, Faculty of Medicine, University of Peradeniya, Peradeniya, Sri Lanka
- 4Wageningen Bioveterinary Research, Lelystad, Netherlands
Livestock-associated methicillin-resistant Staphylococcus aureus (LA-MRSA) is widely spread in intensive farming systems and considered an occupational risk for humans. MRSA is a common nosocomial pathogen in Sri Lanka, but information about prevalence of MRSA in pig farming in Sri Lanka is scarce. Farming is largely a small-scale confined system, and antimicrobial use in these systems is poorly regulated with no veterinary oversight for use. This study identified on 100 pig farms a MRSA prevalence of 10%, with MRSA-positive samples in pigs, farm workers, and dust of 1.2% (6/493), 2.2% (5/228), and 0.8% (1/119), respectively. The genotypes of these strains were compared with 22 human MRSA strains from a hospital; identified in pig farms were CC1/ST1/t127, CC5/ST5/t002, CC6/ST6/t304, or t4403, singleton ST3841/t10744, of which CC1/ST1/t127 and CC/ST5/t002 were present both in isolates from pigs and humans, suggesting a human origin. LA-MRSA types associated with intensive farming (ST398, ST9) were not detected. The low MRSA prevalence at farm level (10% vs. up to 70% in intensive farming systems) might be due to the management of these farms—open air and low dust. We conclude that in Sri Lanka the occupational risk for MRSA acquisition of people working with pigs in the described management systems is negligible.
Introduction
Methicillin-resistant Staphylococcus aureus (MRSA) has become a serious concern as it increases morbidity and mortality in affected humans compared to infections with methicillin-susceptible S. aureus. Furthermore, it is a financial burden for health services. Methicillin resistance is caused by the presence of the mecA or mecC gene, which encodes an alternative penicillin-binding protein with low affinity for beta-lactam antimicrobials (Vanderhaeghen et al., 2010). The healthcare associated MRSA clones are commonly referred to as hospital-associated MRSA (HA-MRSA). Other clones of MRSA without a link to healthcare settings are denoted as community-associated MRSA (CA-MRSA), with its first report in 1993 in a remote area of Western Australia (Udo et al., 1993; Chambers, 2001; Stefani et al., 2012; Chatterjee and Otto, 2013).
MRSA in animals gained much attention in the context of public health importance due to the emergence of MRSA clones in food-producing animals, which have an identified zoonotic potential. In 2004, pigs were identified as the reservoir of livestock-associated (LA- MRSA) (Voss et al., 2005) that is now present on many continents (Witte et al., 2007; Khanna et al., 2008; Cui et al., 2009; EFSA, 2009; Guardabassi et al., 2009; Neela et al., 2009; Wagenaar et al., 2009; Battisti et al., 2010; Fluit, 2012; Lim et al., 2012; Sun et al., 2015). Remarkably, the type of LA-MRSA differs with the geographical location, with strains belonging to Clonal Complex (CC)398 in Europe (EFSA, 2009), whilst CC9 is more common in Asian countries (Guardabassi et al., 2009; Neela et al., 2009; Wagenaar et al., 2009; Ye et al., 2016). Multiple MLST types, dominated by ST398, have been reported in the US and Canada (Khanna et al., 2008; Frana et al., 2013; Sun et al., 2015).
Colonization of LA-MRSA in humans has been reported in the Netherlands (Voss et al., 2005) and Canada (Khanna et al., 2008). The LA-MRSA prevalence rate is higher in people working with live animals—up to 50% in pig farmers—compared to the general community in the Netherlands (van Cleef et al., 2010). Although nosocomial spread has been described, ST398 has less capacity to spread in healthcare facilities compared to HA-MRSA (Hetem et al., 2013). Studies have shown that MRSA can be isolated from pork meat (de Boer et al., 2009). For LA-MRSA the most likely route of transmission to meat is during slaughter. The risk for humans in the general population of handling and consumption of LA-MRSA contaminated meat is estimated to be low as even in a highly exposed population like butchers, the human contamination rate is low (de Jonge et al., 2010). Recent reports from Denmark show the presence of LA-MRSA in people with no direct contact with livestock (Liu et al., 2015; Larsen et al., 2016), suggesting the possible adaptation to infect humans and transmission through food has been suggested for these cases.
In Sri Lanka, MRSA was first reported in the Peradeniya Teaching Hospital in 1994 (Thevanesam et al., 1994). There are many reports from Sri Lankan hospitals describing individual clinical cases as well as outbreaks caused by MRSA (Corea et al., 2003; Dayasena et al., 2011; Jayatilleke and Bandara, 2012; Fernando et al., 2015). Globally, Sri Lanka has the highest reported rate (85%) of MRSA as a proportion of hospital-associated S. aureus infections (Stefani et al., 2012). CA-MRSA is endemic in Sri Lanka (Chatterjee and Otto, 2013) and the proportion of MRSA in community-associated S. aureus infections is 38.8% (Song et al., 2011).
Being an agricultural country, questions rise whether livestock farming contributes to MRSA in humans or vice versa. Low biosecurity and the poorly regulated use of antimicrobials in animal production may lead to the selection and spread of MRSA. MRSA has been phenotypically identified in livestock in Sri Lanka, but if these strains belonged to LA-MRSA is unknown (Jayaweera and Kumbukgolla, 2017). With HA-MRSA and CA-MRSA widely spread in the country it is questionable whether LA-MRSA proportionally contributes to the burden of MRSA in humans.
Except for a few farms scattered in different parts of the country, pig farming in Sri Lanka is largely confined to the western coastal part of the country. This area is commonly known as the “pig belt” and includes four districts, namely Colombo, Gampaha, Kaluthara, and Puttlam. About 60% of the pig farms are small-scale (up to 50 animals), 25% are medium-scale (51–100 animals), and the remaining 15% are large-scale (Kothalawala et al., 2007). All pork is produced for the Sri Lankan market and not imported routinely. In general, the biosecurity on these farms is of poor standards (personal observations).
In this context, the aims of the study were (i) to identify the presence of MRSA in pig farming (pigs, farm staff, and adjacent environment/dust), (ii) to compare the genotypes of MRSA in pig farming to the types found in human clinical isolates from a hospital.
Materials and Methods
Selection of Pig Farms
This study was conducted from January-October 2015 on 100 pig farms distributed in the districts of Puttalam (n = 11), Kurunegala (n = 13), Gampaha (n = 31), Colombo (n = 22), and Kalutara (n = 23) of Sri Lanka. On the day of sample collection, any of the pig farms along the route of the veterinarians' mobile clinic for the particular day were asked to participate, thus farmers did not have prior knowledge on sampling. None of the farmers refused our request to participate in the study. The number of farms visited varied from 5 to 10 farms per day.
Sampling of Pig Farms
Each farm was visited by two veterinarians of the research team. Data on farm management practices, including biosecurity measures, were gathered through a structured questionnaire by interviewing the farm owner along with observations made on the farm premises. Samples were collected from pigs, pig pens, the farmer himself, farm employees, and their family members. Informed written consent was obtained from each person. On each farm, whenever possible, nasal swabs from five pigs were collected using one swab rubbing both nostrils. When <5 pigs were present on a farm then all animals were sampled. Transport swabs (CITOswab, CITOTEST Labware Manufacturing Co., Ltd, Jiangsu, China) without transport medium were utilized. Suckling piglets and sick animals on antimicrobial therapy were avoided but other parameters like age, sex, or the purpose of the animals (breeder/fattener) were not considered as criteria for sample collection. The animal handler, usually one of the farm workers, chose the animals for sampling from a pen indicated by researchers. Farm employees and their family members who volunteered to provide samples and who were in good health and did not use antimicrobials at the time of sampling, were included in the study. They were provided with sterile swabs and were instructed on how to collect a nasal swab themselves. Dust on walls and other surfaces (10 × 10 cm) of the pig pens were collected using sterile dry gauze pads. Two dust samples were collected from each farm. Samples were transported to the laboratory in a cool box and processed within 6–12 h. In total 840 samples; 493 samples from pigs, 228 from humans, and 119 dust samples (including pooled samples) were analyzed (Supplementary Table 1).
Human MRSA Strains
From the Peradeniya Teaching Hospital in the Kandy district, human clinical MRSA isolates were obtained from both in- and out-patients, that were collected in the same period (January-October 2015) as sampling the pig farms. In the hospital laboratory various clinical samples, including pus aspirates and pus swabs, are cultured for diagnostic purposes using conventional microbiological techniques. Twenty-two MRSA isolates were included for molecular analysis, as described below. For privacy reasons we did not trace back patient history or any other data related to those isolates.
Isolation and Identification of MRSA in Pig Samples
Nasal swabs and dust samples from the first 19 farms were analyzed individually; for the remaining 81 farms both dust samples were pooled and analyzed. Samples were enriched in 5 mL Mueller Hinton broth (Thermofisher Diagnostics, Nieuwegein, the Netherlands) containing 6.5% NaCl. After overnight incubation cultures were plated on Oxacillin Resistance Screening Agar Base (ORSAB, Thermofisher Diagnostics, Nieuwegein, the Netherlands). After 48 h incubation at 37°C plates were read according to the recommendations of the manufacturer. Suspected colonies were subcultured on blood agar (BA; Thermofisher Diagnostics, Nieuwegein, the Netherlands) and initially identified as S. aureus by colony morphology on BA, Gram staining, by production of free coagulase that converts fibrinogen in plasma to fibrin, and the simultaneous production of deoxyribonuclease (DNase). Identification was confirmed by Matrix-Assisted Laser-Desorption/Ionization—Time of Flight Mass Spectrometry (MALDI-TOF MS; Bruker Daltonik, Leipzig, Germany).
Molecular Analysis
DNA was extracted from the S. aureus isolates. Briefly: a loop full of culture was suspended in 100 μl of TE (10 mM Tris-HCl, 1 mM EDTA, pH 8.0), then incubated at 95°C for 10 min. and cooled on ice. The mecA-gene was detected with a conventional PCR as described previously (Poulsen et al., 2003). MLST and spa-typing were performed as described (Enright et al., 2000; Harmsen et al., 2003). PCR products were sent for sequence analysis (BaseClear, Leiden, the Netherlands). Sequences were assembled using BioNumerics v7.5 (Applied Maths NV, Sint-Martens-Latem, Belgium). Allelic profiles and STs were assigned using the MLST database (http://pubmlst.org/saureus/). Spa-types were assigned using the spa-typing website (http://spaserver.ridom.de/).
Antimicrobial Susceptibility Testing
Antimicrobial susceptibility patterns of all MRSA isolates, were identified by determining the minimum inhibitory concentrations (MIC), using a semi-automated broth microdilution method, Micronaut-S (MERLIN Diagnostika GmbH, Berlin, Germany). Each isolate was tested against twelve antimicrobial agents, namely clindamycin, chloramphenicol, enrofloxacin, erythromycin, fusidic acid, gentamicin, kanamycin, neomycin, nitrofurantoin, rifampicin, trimethoprim/sulfamethoxazole, and tetracycline.
Results
Management Practices and the Usage of Antimicrobials
Of the sampled farms, 88 farmers provided information on the number of pigs on their farm. The number of sampled farms categorized as small-scale, medium-scale, and large-scale were 57, 17, and 14, respectively. Among the 14 large-scale farms there was only one farm with more than 500 animals. Some farmers maintained a few animals as breeding stock. They produced piglets for their own farm and sold excess animals. Farmers with only fatteners obtained piglets from state-owned breeder farms or from nearby private breeder farms. All the farms had open houses with half-high walls to separate pens, fresh air circulation, and direct contact with the environment, which allows birds and rodents to enter the pens. Swill feeding was the common practice (88/100) while some farmers did not provide relevant information. Three farmers provided commercial feed particularly for the breeding stocks. Biosecurity levels in the pig farms were poor. For instance, 5% of farms applied access through foot baths, boots were changed on 6% of the farms, and 4% of the farms prevented unauthorized visitors from entering the premises. Free access of wild animals, uncontrolled movement of workers, together with swill feeding hindered biosecurity.
Antimicrobials were present in the storage cupboards of 30 farms and the brand names of the antimicrobials were recorded. Conversations with farm owners revealed that they kept antimicrobials to be used on sick animals, particularly piglets with diarrhea. The antimicrobials were categorized under their generic names and the different antimicrobial groups were identified. Accordingly, penicillins (penicillin G, penicillin + streptomycin, amoxicillin, amoxicillin/clavulanate), were the most common, followed by sulphonamides (trimethoprim/sulfonamide and sulfathiazole), fluoroquinolones (enrofloxacin), tetracyclines (tetracycline and tetracycline + neomycin), and macrolides (tylosin). Except one drug belonging to the macrolides, which was available in both the injectable form and powder form, all other antimicrobials seen on the farms were injectable. The remaining 67 farms did not have any antimicrobials. No data were obtained from three farms.
Isolation and Molecular Analysis of MRSA
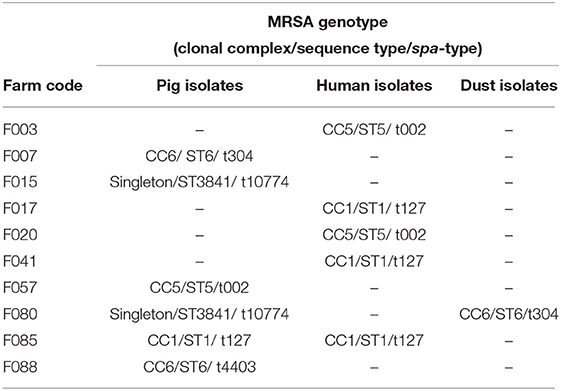
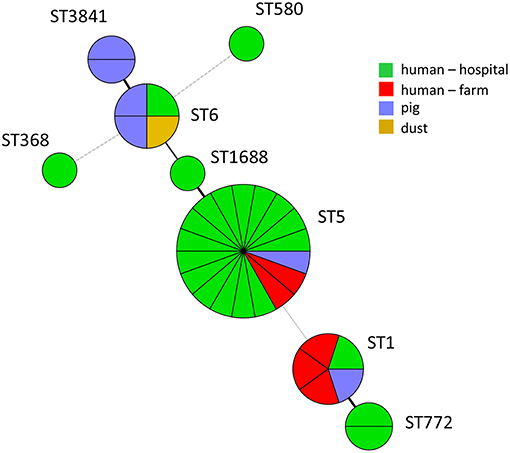
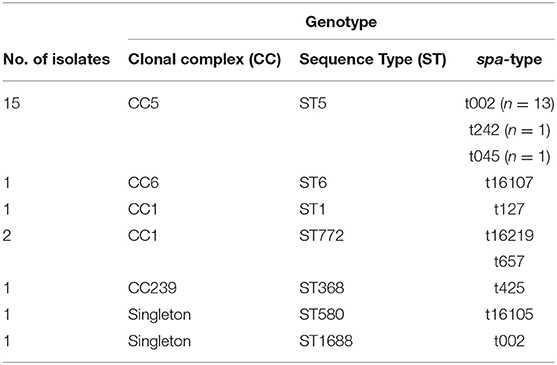
A farm was considered positive for MRSA if any of the three types of samples contained MRSA. Twelve samples out of the 840 (1.4%) were positive for MRSA. MRSA-positive samples in pigs, humans, and dust were 1.2% (6/493), 2.2% (5/228), and 0.8% (1/119), respectively (Supplementary Table 1). Ten farms had one or more MRSA-positive samples (10%). One farm had MRSA in a pig and a human sample, whereas another farm had MRSA in a pig and a dust sample. Out of the human MRSA carriers, one was an employee of a farm having MRSA-positive pigs. The other four humans worked on farms where MRSA was not detected in pigs. The majority of MRSA-positive farms were located in Western province and the highest percentage of positivity was detected in the Kalutara District (21.7%), followed by Colombo (9 %) (Table 1). The MRSA isolates belonged to ST1 (n = 4), ST5 (n = 3), ST6 (n = 3), ST3841—a singleton (n = 2)—and were distributed among different types of samples (Table 2). The isolates belonged to five spa-types (Table 2, Figure 1). On one farm MRSA was isolated from a pig and a farm worker that both belonged to MLST type ST1 and spa-type t127. On another farm MRSA was detected in a pig and dust, but these belonged to different sequence types. Interestingly, ST3841 was isolated from two pigs from two farms situated in different districts, namely Kalutara and Kurunegala. MLST analysis of 22 MRSA isolates from the hospital patients revealed that 15/22 isolates belonged to ST5 (CC5), and the remaining isolates belonged to a variety of STs and spa-types as shown in Table 3.
Antimicrobial Resistance Profiles of MRSA Isolated From Pig Farms and Hospital Patients
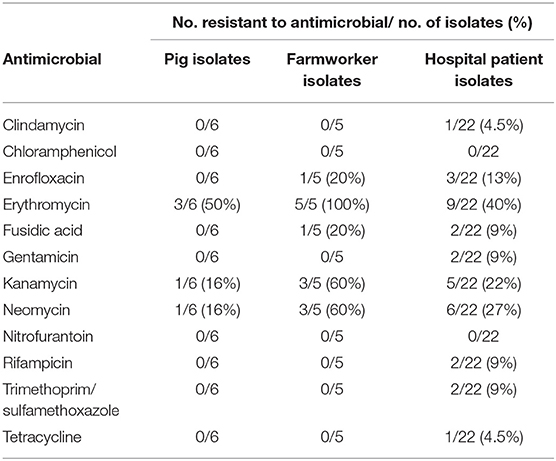
All MRSA isolates were susceptible to chloramphenicol and nitrofurantoin (Table 4). MRSA isolates from pigs and farm workers were also susceptible to an additional five of the tested antimicrobials (n = 12). Conversely, resistance against 10/12 antimicrobials was observed among MRSA isolates from hospital patients.
Discussion
This study provides the first information on molecular characterization of MRSA in association with livestock in Sri Lanka. During the study period of 10 months, 100 farms from five districts spanning the western coast of the country, with the highest proportion of pig farms of the country, were sampled. The total pig population in Sri Lanka in 2014 was 71,838 (Department of animal production, 2014), of which 88% (63,388/ 71,838) of pigs were reared in the studied districts. MRSA was identified in only six pigs, five farm workers, and a dust sample. Accordingly, the farm prevalence was 10%, which included four farms each having a colonized worker in the absence of positive pigs. Since these workers are restricted to work on one farm it is not possible that they obtained MRSA from another pig farm. In an earlier study from Sri Lanka considerable higher positivity rates were reported (Jayaweera and Kumbukgolla, 2017). The identification and characterization methods were different, and therefore comparison is difficult, in particular regarding the MRSA types that were present. The identified farm prevalence is considerably lower compared to prevalence of 70% in the Netherlands (Broens et al., 2011), 38% in Italy (Battisti et al., 2010), and 45% in Canada (Khanna et al., 2008). The MRSA colonization rates of farm workers in our study was 2%, which is much lower than those found in conventional farming in other countries, which reported prevalence in pig farm employees of 37, 20, 7.4, and 5.5% in the Netherlands (van Cleef et al., 2016), Canada (Khanna et al., 2008), China (Ye et al., 2016), and Malaysia (Neela et al., 2009), respectively. The fact that at the six farms with MRSA-positive pigs only one farm worker was positive is clearly different from strong correlations between positive animals and farm workers observed with intensive pig farming. In four farms, only farm workers were MRSA positive, although presence in animals cannot be excluded as only 5 animals were sampled. This shows that the presence at a farm, either in humans or animals, does not lead to widespread colonization in pigs. We hypothesize that the lower farm prevalence and the lower within-farm prevalence, with limited transmission between pigs and humans, may be due to various reasons. One important factor is persistence of MRSA in the housing environment and air of pig barns (Friese et al., 2012). Transmission through air is an important route of exposure for MRSA (Bos et al., 2016) and high concentrations of staphylococci and endotoxins in pig farm air have been identified as occupational risks (Masclaux et al., 2013). In Sri Lanka, most of the farms are small to medium-scale and pigs are reared in open houses with half-high walls, enabling air circulation and some sunlight. Farmers routinely washed the pens at least twice a day, with ample water to clean the floors after swill feeding and to cool the animals.
A majority of the published studies reporting MRSA, particularly LA-MRSA (ST398 and ST9), was from animals reared in intensive farming systems and studies in non-intensive systems are scarce. A study from Germany reported the absence of LA-MRSA (Cuny et al., 2012) on a pig farm with an alternative rearing system, and a study from Peru showed that MRSA colonization in backyard scavenging pigs is lower compared to intensively farmed animals (Arriola et al., 2011). Similarly, prevalence of MRSA in Dutch organic pig farms is 21% in comparison to 70% in conventional farms (van de Vijver et al., 2014). In line with those findings, in the USA, people working with backyard pigs did not have an increased risk for being MRSA positive compared to the general population (Gordoncillo et al., 2012). The aforementioned studies support the notion that intensification might be a risk factor for MRSA colonization in pig farms and occupational transmission to humans.
Out of the 100 farms, 30 had antimicrobials on the farm premises at the time of sampling and some farmers stated using these drugs for diseased animals without consulting a veterinarian. However, group medication of animals, which has been identified as a risk factor for LA-MRSA in veal calf farming (Graveland et al., 2010), was not a common practice. This is also clear from the fact that almost only injectable drugs were present. The farmers were unaware of antimicrobial stewardship and should be educated to prevent unnecessary use. The most common feeding practice in the studied farms was swill feeding (88/100). This type of feed does not contain antimicrobials in contrast to several types of commercial feed and it is not a risk for selection of resistant bacteria. Furthermore, the levels of resistance seen in farm MRSA isolates toward tested antimicrobials were low, and remarkably, isolates from farm workers also showed low levels of resistance compared to those of human patient isolates (Jayatilleke and Bandara, 2012). The dogma is that there is a direct association of antimicrobial usage with MRSA prevalence (Tacconelli et al., 2008; Graveland et al., 2010), as has been shown in association with pig farming in the Netherlands (Dorado-García et al., 2015). A limitation of this study could be the omission of sampling humans or animals on antimicrobial therapy, because there is clear association between MRSA positivity and antimicrobial usage in the same herd (Tacconelli et al., 2008). In the present study, no risk factors could be defined, as histories of antimicrobial use were not available due to poor record keeping, and the low MRSA prevalence.
Molecular characterization showed four different sequence types and five spa-types in farm isolates. An important finding was the absence of predominant LA-MRSA types ST398 and ST9. Since its first isolation from Asia in 2009 on Chinese farms, LA-MRSA belonging to ST9 was reported from farms in Malaysia and Hong Kong (Guardabassi et al., 2009; Neela et al., 2009). The infrequent import of animals from positive countries might explain the absence of CC9 and CC398 in Sri Lanka. Nevertheless, there were six pigs colonized with MRSA of various sequence types. Among those, there is a new sequence type (ST3841) of spa-type t10774. Two pigs carried this new type and were reared on two different farms, at two distant districts—Kaluthara and Kurunegala- so that a common origin could not be explained. The other STs that were identified were ST5 and ST1. One pig was colonized with ST5 and spa-type t002, which belongs to the most common CA-MRSA type described in humans (Song et al., 2011; Williamson et al., 2013), and which has also been identified in pigs, horses, dogs, and cats in Canada (Khanna et al., 2008; Frana et al., 2013). It has been suggested that this type originates from colonized humans. This was also described in a Korean study where on 66 pig farms, besides CC398, occasionally the HA-MRSA ST72 type was identified (Lim et al., 2012). What is unclear in these studies, is if this represents contamination of nasal passages or that these MRSA types can establish colonization in pigs. Colonization of farm workers with ST1 of spa-type t127 was the most common (3/5 MRSA isolates) followed by ST5 of spa-type t002 (2/5 MRSA isolates). Both spa-types have been reported among human HA- and CA-MRSA isolates in Sri Lanka (Song et al., 2011). In a recent study from India, ST5 was isolated from pork, confirming the presence of this type in the food chain (Zehra et al., 2018). In the present study, 15/22 MRSA isolates from human patients belonged to ST5, of which 13 belonged to spa-type t002. As this ST belongs to predominant CA-MRSA types it cannot be excluded that patients acquired this strain outside the hospital. Interestingly, on one farm both a farm worker and pig carried a MRSA ST1, spa-type t127 isolate with the same resistance profile that suggested transmission. Given the low prevalence of MRSA on pig farms, it is highly unlikely that the workers obtained MRSA from the pigs, and rather suggests a possible transmission from humans to pigs. This study was performed in the live production of the pork production chain. No carcass or meat samples were analyzed for the presence of MRSA and therefore the role of the food chain in exposure of consumers with MRSA cannot be assessed. Given the high prevalence in humans and the findings in this study with CA-MRSA and HA-MRSA present in pigs, it will be impossible to quantify the role of pork in the transmission of MRSA to humans in Sri Lanka.
In conclusion, MRSA has a remarkably low prevalence in the pig farming community in Sri Lanka. Predominant LA-MRSA types are absent on pig farms, but pigs and farm workers share sequence types that are associated with CA-MRSA and HA-MRSA strains.
Ethics Statement
The study was approved by the Ethical Review Committee of the Faculty of Medicine and the Ethics Committee of the Faculty of Veterinary Medicine and Animal Science, University of Peradeniya, 20400 Peradeniya, Sri Lanka in 2014. All study participants signed an informed consent, and any personal information obtained during the study is kept confidential.
Author Contributions
RK, CG, ND, and LR conceived the study and collected the samples. KV, LR, and HG performed the laboratory experiments. RK, BD, KV, and JW analyzed the data. KV, RK, BD, and JW wrote the paper. All authors revised and approved the final version of the manuscript.
Funding
This study was funded by a grant (No: 14-105) from the National Research Council of Sri Lanka. We would like to thank the veterinarians, field officers and the farmers for the support extended during the study.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fsufs.2019.00025/full#supplementary-material
References
Arriola, C. S., Güere, M. E., Larsen, J., Skov, R. L., Gilman, R. H., Gonzalez, A. E., et al. (2011). Presence of methicillin-resistant Staphylococcus aureus in pigs in Peru. PLoS ONE 6:e28529. doi: 10.1371/journal.pone.0028529
Battisti, A., Franco, A., Merialdi, G., Hasman, H., Iurescia, M., Lorenzetti, R., et al. (2010). Heterogeneity among methicillin-resistant Staphylococcus aureus from italian pig finishing holdings. Vet. Microbiol. 142, 361–366. doi: 10.1016/j.vetmic.2009.10.008
Bos, M. E. H., Verstappen, K. M., van Cleef, B. A. G. L., Dohmen, W., Dorado-Garcia, A., Graveland, H., et al. (2016). Transmission through air as a possible route of exposure for MRSA. J. Expos. Sci. Environ. Epidemiol. 26, 263–269. doi: 10.1038/jes.2014.85
Broens, E. M., Graat, E. A. M., van der Wolf, P. J., van de Giessen, A. W., and de Jong, M. C. M. (2011). Prevalence and risk factor analysis of livestock associated MRSA-positive pig herds in the Netherlands. Prev. Vet. Med. 102, 41–49. doi: 10.1016/j.prevetmed.2011.06.005
Chambers, H. F. (2001). “The Changing Epidemiology of Staphylococcus aureus” Emerging Infect. Dis. 7, 178–182. doi: 10.3201/eid0702.700178
Chatterjee, S. S., and Otto, M. (2013). Improved understanding of factors driving methicillin-resistant Staphylococcus aureus epidemic waves. Clin. Epidemiol. 5, 205–217. doi: 10.2147/CLEP.S37071
Corea, E., de Silva, T., and Perera, J. (2003). Methicillin-resistant Staphylococcus aureus: prevalence, incidence and risk factors associated with colonization in Sri Lanka. J. Hosp. Infect. 55, 145–148. doi: 10.1016/S0195-6701(03)00256-1
Cui, S., Li, J., Hu, C., Jin, S., Li, F., Guo, Y., et al. (2009). Isolation and characterization of methicillin-resistant Staphylococcus aureus from swine and workers in China. J. Antimicrob. Chemother. 64, 680–683. doi: 10.1093/jac/dkp275
Cuny, C., Friedrich, A. W., and Witte, W. (2012). Absence of livestock-associated methicillin-resistant Staphylococcus aureus clonal complex CC398 as a nasal colonizer of pigs raised in an alternative system. Appl. Environ. Microbiol. 78, 1296–1297. doi: 10.1128/AEM.07260-11
Dayasena, R. P., Dayasiri, M. B. K. C., Jayasuriya, C., and Perera, D. S. C. (2011). Aetiological agents in chronic suppurative otitis media in Sri Lanka. Austr. Med. J. 4, 101–104. doi: 10.4066/AMJ.2011.549
de Boer, E., Zwartkruis-Nahuis, J. T. M., Wit, B., Huijsdens, X. W., de Neeling, A. J., Bosch, T., et al. (2009). Prevalence of methicillin-resistant Staphylococcus aureus in meat. Int. J. Food Microbiol. 134, 52–56. doi: 10.1016/j.ijfoodmicro.2008.12.007
de Jonge, R., Verdier, J. E., and Havelaar, A. H. (2010). Prevalence of meticillin-resistant Staphylococcus aureus amongst professional meat handlers in the Netherlands, March–July 2008. Eurosurveillance 15:19712. doi: 10.2807/ese.15.46.19712-en
Dorado-García, A., Dohmen, W., Bos, M. E. H., Verstappen, K. M., Houben, M., Wagenaar, J. A., et al. (2015). Dose-response relationship between antimicrobial drugs and livestock-associated MRSA in pig farming. Emerg. Infect. Dis. 21, 950–959. doi: 10.3201/eid2106.140706
EFSA. (2009). Analysis of the Baseline Survey on the Prevalence of Methicillin-Resistant Staphylococcus aureus (MRSA) in Holdings with Breeding Pigs, in the EU, 2008 - Part A: MRSA Prevalence Estimates. EFSA J. 7:1376. doi: 10.2903/j.efsa.2009.1376
Enright, M. C., Day, N. P., Davies, C. E., Peacock, S. J., and Spratt, B. G. (2000). Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 38, 1008–1015.
Fernando, D., Ratnatunga, C., and Athapaththu, R. (2015). Necrotizing soft tissue infection caused by community acquired methicillin-resistant Staphylococcus aureus: an emerging deadly entity. Sri Lanka J. Forensic Med. Sci. Law 5:2. doi: 10.4038/sljfmsl.v5i2.7754
Fluit, A. C. (2012). Livestock-associated Staphylococcus aureus. Clin. Microbiol. Infect. 18, 735–744. doi: 10.1111/j.1469-0691.2012.03846.x
Frana, T. S., Beahm, A. R., Hanson, B. M., Kinyon, J. M., Layman, L. L., Karriker, L. A., et al. (2013). Isolation and characterization of methicillin-resistant Staphylococcus aureus from pork farms and visiting veterinary students. PLoS ONE 8:e53738. doi: 10.1371/journal.pone.0053738
Friese, A., Schulz, J., Hoehle, L., Fetsch, A., Tenhagen, B.-A., Hartung, J., et al. (2012). Occurrence of MRSA in air and housing environment of pig barns. Vet. Microbiol. 158, 129–135. doi: 10.1016/j.vetmic.2012.01.019
Gordoncillo, M. J., Abdujamilova, N., Perri, M., Donabedian, S., Zervos, M., and Bartlett, P. (2012). Detection of methicillin-resistant Staphylococcus aureus (MRSA) in backyard pigs and their owners, Michigan USA. Zoonoses Public Health 59, 212–216. doi: 10.1111/j.1863-2378.2011.01437.x
Graveland, H., Wagenaar, J. A., Heesterbeek, H., Mevius, D., van Duijkeren, E., and Heederik, D. (2010). Methicillin resistant Staphylococcus aureus ST398 in veal calf farming: human MRSA carriage related with animal antimicrobial usage and farm hygiene. PLoS ONE 5:e10990. doi: 10.1371/journal.pone.0010990
Guardabassi, L., O'Donoghue, M., Moodley, A., Ho, J., and Boost, M. (2009). Novel lineage of methicillin-resistant Staphylococcus aureus, Hong Kong. Emerg. Infect. Dis. 15, 1998–2000. doi: 10.3201/eid1512.090378
Harmsen, D., Claus, H., Witte, W., Rothgänger, J., Claus, H., Turnwald, D., et al. (2003). Typing of methicillin-resistant Staphylococcus aureus in a University hospital setting by using novel software for spa repeat determination and database management. J. Clin. Microbiol. 41, 5442–5448. doi: 10.1128/JCM.41.12.5442-5448.2003
Hetem, D. J., Bootsma, M. C. J., Troelstra, A., and Bonten, M. J. M. (2013). Transmissibility of livestock-associated methicillin-resistant Staphylococcus aureus. Emerg. Infect. Dis. 19, 1797–1802. doi: 10.3201/eid1911.121085
Jayatilleke, K., and Bandara, P. (2012). Antibiotic sensitivity pattern of Staphylococcus aureus in a tertiary care hospital of Sri Lanka. Sri Lanka J. Infect. Dis. 2:2. doi: 10.4038/sljid.v2i2.4162
Jayaweera, J. A. A. S., and Kumbukgolla, W. W. (2017). Antibiotic resistance patterns of methicillin-resistant Staphylococcus aureus (MRSA) isolated from livestock and associated farmers in Anuradhapura, Sri Lanka. GERMS 7, 132–139. doi: 10.18683/germs.2017.1118
Khanna, T., Friendship, R., Dewey, C., and Weese, J. S. (2008). Methicillin resistant Staphylococcus aureus colonization in pigs and pig farmers. Vet. Microbiol. 128, 298–303. doi: 10.1016/j.vetmic.2007.10.006
Kothalawala, K. A. C. H. A., Perera, E. R. K., Piyadasa, H. W., and Udugama, H. A. W. M. R. U. W. K. (2007). Effect of different feeding systames on the economics of swine farming in Sri Lanka. Sri Lanka Vet. J. 54–55, 11–14. Available online at: https://journal.slva.org/content/journal/id/5
Larsen, J., Stegger, M., Andersen, P. S., Petersen, A., Larsen, A. R., Westh, H., et al. (2016). Evidence for human adaptation and foodborne transmission of livestock-associated methicillin-resistant Staphylococcus aureus. Clin. Infect. Dis. 63, 1349–1352. doi: 10.1093/cid/ciw532
Lim, S.-K., Nam, H.-M., Jang, G.-C., Lee, H.-S., Jung, S.-C., and Kwak, H.-S. (2012). The first detection of methicillin-resistant Staphylococcus aureus ST398 in pigs in Korea. Vet. Microbiol. 155, 88–92. doi: 10.1016/j.vetmic.2011.08.011
Liu, C. M., Price, L. B., Hungate, B. A., Abraham, A. G., Larsen, L. A., Christensen, K., et al. (2015). Staphylococcus aureus and the ecology of the nasal microbiome. Sci. Adv. 1:e1400216. doi: 10.1126/sciadv.1400216
Masclaux, F. G., Sakwinska, O., Charrière, N., Semaani, E., and Oppliger, A. (2013). Concentration of airborne Staphylococcus aureus (MRSA and MSSA), total bacteria, and endotoxins in pig farms. Ann. Occup. Hyg. 57, 550–557. doi: 10.1093/annhyg/mes098
Neela, V., Zafrul, A. M., Mariana, N. S., van Belkum, A., Liew, Y. K., and Ghaznavi Rad, E. (2009). Prevalence of ST9 methicillin-resistant Staphylococcus aureus among pigs and pig handlers in Malaysia. J. Clin. Microbiol. 47, 4138–4140. doi: 10.1128/JCM.01363-09
Poulsen, A. B., Skov, R., and Pallesen, L. (2003). Detection of low-level methicillin-resistant Staphylococcus aureus with commercially available tests. J. Clin. Microbiol. 41:3458. doi: 10.1128/JCM.41.7.3458.2003
Song, J.-H., Hsueh, P.-R., Chung, D. R., Ko, K. S., Kang, C.-I., Peck, K. R., et al. (2011). Spread of methicillin-resistant Staphylococcus aureus between the community and the hospitals in Asian countries: an {ANSORP} study. J. Antimicrob. Chemother. 66, 1061–1069. doi: 10.1093/jac/dkr024
Stefani, S., Chung, D. R., Lindsay, J. A., Friedrich, A. W., Kearns, A. M., Westh, H., et al. (2012). Meticillin-resistant Staphylococcus aureus (MRSA): global epidemiology and harmonisation of typing methods. Int. J. Antimicrob. Agents 39, 273–282. doi: 10.1016/j.ijantimicag.2011.09.030
Sun, J., Yang, M., Sreevatsan, S., and Davies, P. R. (2015). Prevalence and characterization of Staphylococcus aureus in growing pigs in the USA. PLoS ONE 10:e0143670. doi: 10.1371/journal.pone.0143670
Tacconelli, E., De Angelis, G., Cataldo, M. A., Pozzi, E., and Cauda, R. (2008). Does antibiotic exposure increase the risk of methicillin-resistant Staphylococcus aureus (MRSA) isolation? A systematic review and meta-analysis. J. Antimicrob. Chemother. 61, 26–38. doi: 10.1093/jac/dkm416
Thevanesam, V., Wijeyawardana, W. L., and Ekanayake, E. W. (1994). Methicillin resistant Staphylococcus aureus: the scale of the problem in a Sri Lankan hospital. J. Hosp. Infect. 26, 123–127. doi: 10.1016/0195-6701(94)90054-X
Udo, E. E., Pearman, J. W., and Grubb, W. B. (1993). Genetic analysis of community isolates of methicillin-resistant Staphylococcus aureus in Western Australia. J. Hosp. Infect. 25, 97–108. doi: 10.1016/0195-6701(93)90100-E
van Cleef, B., Van Benthem, B., Verkade, E. J. M., Van Rijen, M. M. L., Kluytmans-Van Den Bergh, M. F. Q., Graveland, H., et al. (2016). Health and health-related quality of life in pig farmers carrying livestock-associated methicillin-resistant Staphylococcus aureus. Epidemiol. Infect. 8, 1774–1783. doi: 10.1017/S0950268815003192
van Cleef, B. A., Verkade, E. J. M., Wulf, M. W., Buiting, A. G., Voss, A., Huijsdens, X. W., et al. (2010). Prevalence of livestock-associated MRSA in communities with high pig-densities in the Netherlands. PLoS ONE 5:e9385. doi: 10.1371/journal.pone.0009385
van de Vijver, L. P. L., Tulinski, P., Bondt, N., Mevius, D., and Verwer, C. (2014). Prevalence and molecular characteristics of methicillin-resistant Staphylococcus aureus (MRSA) in organic pig herds in the Netherlands. Zoonoses Public Health 61, 338–345. doi: 10.1111/zph.12076
Vanderhaeghen, W., Hermans, K., Haesebrouck, F., and Butaye, P. (2010). Methicillin-resistant Staphylococcus aureus (MRSA) in food production animals. Epidemiol. Infect. 138, 606–625. doi: 10.1017/S0950268809991567
Voss, A., Loeffen, F., Bakker, J., Klaassen, C., and Wulf, M. (2005). Methicillin-resistant Staphylococcus aureus in pig farming. Emerg. Infect. Dis. 11, 1965–1966. doi: 10.3201/eid1112.050428
Wagenaar, J. A., Yue, H., Pritchard, J., Broekhuizen-Stins, M., Huijsdens, X., Mevius, D. J., et al. (2009). Unexpected sequence types in livestock associated methicillin-resistant Staphylococcus aureus (MRSA): MRSA ST9 and a single locus variant of ST9 in pig farming in China. Vet. Microbiol. 139, 405–409. doi: 10.1016/j.vetmic.2009.06.014
Williamson, D. A., Roberts, S. A., Ritchie, S. R., Coombs, G. W., Fraser, J. D., and Heffernan, H. (2013). Clinical and molecular epidemiology of methicillin-resistant Staphylococcus aureus in New Zealand: rapid emergence of sequence type 5 (ST5)-SCCmec-IV as the dominant community-associated MRSA clone. PLoS ONE 8:e62020. doi: 10.1371/journal.pone.0062020
Witte, W., Strommenger, B., Cuny, C., Heuck, D., and Nuebel, U. (2007). Methicillin-resistant Staphylococcus aureus containing the panton-valentine leucocidin gene in Germany in 2005 and 2006. J. Antimicrob. Chemother. 60, 1258–1263. doi: 10.1093/jac/dkm384
Ye, X., Fan, Y., Wang, X., Liu, W., Yu, H., Zhou, J., et al. (2016). Livestock-associated methicillin and multidrug resistant, S. aureus in humans is associated with occupational pig contact, not pet contact. Sci. Rep. 6:19184. doi: 10.1038/srep19184
Keywords: MRSA, pigs, Sri Lanka, livestock, small-scale farming, pork, food chain
Citation: Kalupahana RS, Duim B, Verstappen KM, Gamage CD, Dissanayake N, Ranatunga L, Graveland H and Wagenaar JA (2019) MRSA in Pigs and the Environment as a Risk for Employees in Pig-Dense Areas of Sri Lanka. Front. Sustain. Food Syst. 3:25. doi: 10.3389/fsufs.2019.00025
Received: 22 January 2019; Accepted: 04 April 2019;
Published: 08 May 2019.
Edited by:
Ömer Akineden, University of Giessen, GermanyReviewed by:
Abdulwahed Ahmed Hassan, University of Giessen, GermanyAmir Abdulmawjood, University of Veterinary Medicine Hannover, Germany
Copyright © 2019 Kalupahana, Duim, Verstappen, Gamage, Dissanayake, Ranatunga, Graveland and Wagenaar. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Birgitta Duim, Qi5kdWltQHV1Lm5s
 Ruwani S. Kalupahana
Ruwani S. Kalupahana Birgitta Duim
Birgitta Duim Koen M. Verstappen
Koen M. Verstappen Chandika D. Gamage3
Chandika D. Gamage3 Nilanthi Dissanayake
Nilanthi Dissanayake Haitske Graveland
Haitske Graveland Jaap A. Wagenaar
Jaap A. Wagenaar