- 1Department of Environmental Sciences, University of California, Riverside, Riverside, CA, United States
- 2IISD Experimental Lakes Area (IISD-ELA), Winnipeg, MB, Canada
- 3Department of Chemistry, University of Manitoba, Winnipeg, MB, Canada
- 4Institute of Environmental Health, College of Environmental and Resource Sciences, Zhejiang University, Hangzhou, China
Transport of diluted bitumen (dilbit) from Canada’s oil sands region poses risk for leaks and spills of petroleum-derived contaminants into the environment. Exposure of fish to dilbit is known to cause cardiotoxicity, developmental deformities, and impairment in swim performance. However, previous studies have examined the toxicity of dilbit in laboratory settings which does not account for environmental and biological food-web variables that may alter exposure and/or toxicity of dilbit. Moreover, most methods of assessing organism health following oil exposure require lethal sampling. This work is a part of a larger set of experiments where dilbit spills were simulated within enclosures on a lake; the present study assesses the impacts of residual levels of dilbit that may have entered the surrounding lake environment from the enclosures following model spill cleanup. In order to understand the impacts of residual dilbit in an ecosystem setting without use of lethal sampling, epidermal mucus was collected and sequenced from lake charr (Salvelinus namaycush) exposed to residual dilbit in a boreal lake. While concentrations reached a maximum of 2.29 μg/L total polycyclic aromatic compounds (ΣPAC) within surface waters, surface water ΣPAC concentrations generally remained below 1 μg/L. Results of RNA sequencing were compared to sequencing data from mucus collected prior to dilbit additions. Differential gene expression and pathway analyses indicated dysregulation of genes associated with intermediary and energy metabolism as well as a trend in upregulation of cyp1a3 in epidermal mucus following dilbit exposure. Thus, results of the present study suggest that lake charr undergo consistent biological responses after exposure to residual levels of dilbit following a model spill, and that mRNA-based analysis of mucus may be a viable method for non-lethal oil exposure assessment. Overall, the results provide insight on the response of wild fish to very dilute dilbit exposures after a model spill cleanup.
Introduction
The oil sands region of Alberta, Canada has one of the largest known reserves of bitumen, containing an estimated 50 billion cubic meters of the heavy type crude oil (Canada National Energy Board, 2006). Due to its viscous nature, bitumen is mixed with varying amounts of light oil diluent (e.g., naptha-based condensate) to decrease overall viscosity for more efficient transport, thus forming diluted bitumen (dilbit). Dilbit is primarily exported from the oil sands region via extensive transcontinental pipeline networks, raising concern for potential leaks and spills into lakes and rivers located along transportation routes (Dupuis and Ucan-Marin, 2015; Canada Energy Regulator, 2018). Although export by rail has increased in recent years, volumes remain small compared to pipelines but present an additional potential spill vector (Canada Energy Regulator, 2018). Multiple significant dilbit spills have occurred in North America, with the largest being the 2010 Enbridge Line 6B pipeline rupture which released 1.15 million gallons of dilbit into the Kalamazoo River in Michigan (Crosby et al., 2013; National Academies of Sciences Medicine and Engineering, 2016). The spill impacted 1,560 acres of in-stream habitats as well as floodplain and upland areas and resulted in ecological injury to fish, benthic invertebrates, birds, mammals, and other wildlife (USFWS, 2015). Although major dilbit spills have occurred in the past and future spills are inevitable, there remain significant research gaps on the behavior of dilbit in the environment and its toxicity to aquatic organisms. Furthermore, oil sands dilbit production and export is projected to continue to expand due to significant economic incentives (Crosby et al., 2013; Boufadel et al., 2015). Continual increases in transport of dilbit and proposed expansions of existing pipeline networks make it critical to fully characterize the effects of dilbit on aquatic ecosystems for adequate risk assessment of future spills and proper guidance in spill response decision making.
As bitumen is heavily biodegraded over geological time, dilbit contains a high proportion of heavy molecular weight compounds such as resins, asphaltenes, and high molecular weight polycyclic aromatic compounds (PACs) and alkylated PACs (Yang et al., 2011; National Academies of Sciences Medicine and Engineering, 2016). Moreover, the chemical fingerprint of dilbit (as quantified by n-alkanes, PACs and their alkylated homologues, terpanes, steranes, bicyclic sesquiterpanes, etc.) is highly variable and dependent on extraction and refining processes, geographic source, and the composition and volume of diluent added by the producer (Adams et al., 2013; Lee et al., 2016). Following a spill, the chemical properties of dilbit are further modified by complex weathering processes such as photo-oxidation, biodegradation, evaporation, emulsification, dispersion, and sedimentation (Dew et al., 2015; National Academies of Sciences Medicine and Engineering, 2016). Due to its complex chemical composition and multiple factors affecting its chemical properties, it may be difficult to understand the fate and behavior of spilled dilbit in freshwater environments. In order to better understand dilbit’s behavior in low-energy lake systems, Stoyanovich et al., 2019 conducted experimental releases of dilbit in outdoor microcosms containing natural lake water and sediments (Stoyanovich et al., 2019). As volatile hydrocarbons evaporated, dilbit increased in density and viscosity, resulting in its submergence after 8 days. Furthermore, there was a rapid accumulation of PACs in the water column, which tend to be drivers of oil toxicity in aquatic organisms.
The toxicity of dilbit in freshwater environments has been minimally studied and has only recently begun to receive more extensive research (Dupuis and Ucan-Marin, 2015; National Academies of Sciences Medicine and Engineering, 2016). Similar to toxic endpoints associated with conventional crude oil, dilbit exposure in fish has shown to induce mortality, decreased utilization of energy stores, metabolic alterations, reduced growth, oxidative stress, behavioral changes, developmental malformations, delayed hatching, changes in heart rate, abnormal inflation or complete un-inflation of the swim bladder, among other adverse effects (Madison et al., 2015, 2017; Philibert et al., 2016; Alsaadi F. et al., 2018, Alsaadi et al., 2018 F. M.; Alderman et al., 2018; Avey et al., 2020). Impairments induced by dilbit exposure may have impacts on organismal fitness and, thus, population health. For example, exposure of sockeye salmon parr (Oncorhynchus nerka) to water soluble fractions of dilbit (3.5–66.7 μg/L ΣPAC) resulted in concentration-dependent cardiac remodeling that correlated to reductions in swimming performance (Alderman et al., 2017b). Dilbit may also impede exercise recovery, as dilbit-exposed sockeye salmon experienced increased cellular damage following exercise as suggested by an increase in tissue leakage proteins, such as creatine kinase, in serum compared to controls (Alderman et al., 2017a). As reduced swim performance may impair migratory success, dilbit spills may pose risk to sensitive salmonid populations in freshwater ecosystems.
Following a freshwater oil spill, there is a need for rapid exposure assessment of biota in the impacted area. Currently, many methods used to assess ecological damage to organisms, such as those used following the Kalamazoo River spill, require lethal sampling which may result in further impairment (USFWS, 2015). Use of exposure assessment tools which are nonlethal and minimally invasive is important to reduce further perturbations on animal populations. In fish, molecular analysis of epidermal mucus may be a viable tool for monitoring effects of oil exposure as it is nonlethal, rapid, inexpensive, and sample procurement requires little training. Following exposure to Deepwater Horizon slick oil, RNA sequencing of mucus from juvenile mahi-mahi (Coryphaena hippurus) identified molecular changes in mucus associated with oil toxicity, such as upregulation of cytochrome P450 1A (CYP1A) and dysregulation of transcripts associated with cardiotoxicity and immunosuppression (Greer et al., 2019). Moreover, exposure of dusky splitfin (Goodea gracilis) to crude oil resulted in increased antioxidant defense response in the epidermal mucus (Dzul-Caamal et al., 2016). Results from these previous studies suggest that epidermal mucus may be a useful target for environmental monitoring following oil spills. However, previous studies were laboratory-based and may not reflect changes in chemical properties of oil brought about by environmental conditions such as a evaporation, photo-oxidation, emulsification, dispersion, biodegradation, and sedimentation (National Academies of Sciences Medicine and Engineering, 2016).
Although research suggests that mucus may be a promising method for nonlethal sampling, the efficacy of mucus as an oil exposure assessment tool has not yet been assessed on wild populations in the field. In the present study, epidermal mucus was collected from lake charr (Salvelinus namaycush) residing in a pristine Canadian boreal shield lake at the International Institute for Sustainable Development Experimental Lakes Area (IISD-ELA) where experimental dilbit spills and cleanups were conducted within enclosures on the lake (detailed in Rodriguez-Gil et al., 2021). While experimental spills were conducted within enclosures, minute levels of dilbit entered the surrounding lake environment leading to potential exposure of resident lake charr. To minimize impacts on the small and vulnerable lake charr population, lethal or invasive sampling of tissue or blood was not used and, instead, epidermal mucus was collected. RNA sequencing was conducted on lake charr mucus collected prior to and after dilbit additions to identify altered molecular pathways associated with dilbit exposure. Lake charr may serve as a model species for the study as many dilbit pipelines are concentrated in areas of Canada where they intersect freshwater ecosystems containing sensitive salmonid species. Results of the present study will identify molecular pathways altered by dilbit in a whole-lake setting and further contribute to the development of mucus as a nonlethal tool for oil exposure assessment. Overall, the present study provides insight on the response of wild fish to very dilute µg/L dilbit exposure after a model spill cleanup.
Methods
Study Area and Diluted Bitumen Additions
The study was conducted at IISD-ELA, a freshwater research center located near Kenora, Ontario, Canada (49°40′N, 93°44′W) (https://www.iisd.org/ela/). Being located in a sparsely populated region, the lakes at IISD-ELA are mostly unaffected by anthropogenic influences. IISD-ELA’s Lake 260 was selected for the present study (49°42′N, 93°46′W) which has an area of 331,328 m2, a volume of 1,975,971 m3, an average depth of 5.9 m, and a maximum depth of 15.7 m. Lake 260 is a boreal lake and serves as a model for freshwater water bodies in Canada’s Boreal Shield ecozone.
The present study was a part of a larger series of dilbit experiments at IISD-ELA. In order to examine the fate, behavior and potential impacts of spilled dilbit in freshwater lakes, dilbit spills were simulated within 10 m diameter, ∼100 m3 volume limnocorrals on Lake 260 beginning in 2018 (Orihel, 2019). The experimental design of the simulated dilbit spills are described in a previous publication (Rodriguez-Gil et al., 2021). Briefly, limnocorrals consist of a floatation collar that supports an impermeable curtain that extends to the bottom of the lake where it is sealed to the sediments using sandbags, providing an enclosed ecosystem to model whole lake processes. A series of 9 limnocorrals were constructed and deployed in water depths of ∼1.5 m. After installation and pre-spill monitoring, Cold Lake winter Blend dilbit was added to seven limnocorrals using a regression with added volumes ranging from 1:100,000–1:100 dilbit to water (v:v). The total volume of dilbit added to the enclosures was ∼382 L. Chemical and physical weathering, as well as potential toxicity, of the dilbit were studied for 70 days, after which residual surface dilbit was removed from the limnocorrals. The limnocorral curtains were then cut at mid water column depth prior to winter freeze up to allow continued study of the dilbit that remained on the lake bottom. As a result, there was potential for dilbit to enter the lake water column throughout the duration of the winter months and prior to final sediment cleaning, which took place in August 2019. Additionally, in June 2019 fourteen shoreline enclosures (5 × 15 m and enclosing volumes of 25,000–30,000 L of lake water) were treated with 1.5 L of weathered dilbit (Palace, 2021). While the environments were enclosed, tracer studies indicated that some leakage to the outside lake environment occurred, potentially contributing to enriched PAC concentrations in the lake. The present study thus examined the potential effects that residual dilbit entering the surrounding lake environment following cleanup had on resident fish.
Water Extractions and Analytical Chemistry
Surface water samples (1 L) were collected from the lake at 10 time points during the study: 3 days before oil addition and on days 1,2,3,5,9,13, 18, 53, and 80 after application. Samples were filtered first using Whatman GF/C filters and then transferred to 2 L separatory funnels. Each funnel was spiked with 20 µl of a 5 ppm recovery internal standard which consisted of d8-naphthalene, d8-acenaphthylene, d10-acenaphthene, d10-fluorene, d10-phenanthrene, d10-pyrene, d12-Benz(a)anthracene, d12-chrysene, d12-benzo (b)fluoranthene, d12-benzo (k)fluoranthene, d12-benzo (a)pyrene, d12-indeno (1,2,3-c,d)pyrene, d14-dibenzo (a,h)anthracene, and d14-benzo (g,h,i)perylene. Ten grams of NaCl was dissolved into each sample prior to the liquid:liquid extraction. The samples were extracted by adding 50 ml of dichloromethane (DCM) to each funnel and gently swirling for 1 min. The DCM was drained into a round bottom flask and the water extraction was repeated. The combined 100 ml DCM extract was then evaporated to a final volume of 1 ml and spike with 20 µl of 5 ng/μl d10-anthracene as an instrument performance internal standard. Samples were vortexed and transferred to 2 ml amber glass autosampler vials for storage at 4°C until analysis by GC-MS/MS.
An Agilent 7,890 GC coupled to a 7,000°C triple quadrupole mass spectrophotometer fitted with an electron ionization (EI) source used for the native and d-labelled PAC and alkyl PAC quantification. Helium was used as the carrier gas at a flow rate of 1.2 ml/min with an Agilent J&W HP-5ms ultra inert column (30 m × 0.25 mm, 0.25 μm film thickness). Sample volumes of 1 μl were injected by a PAL RSI 85 auto sampler at a temperature of 60°C. GC conditions and details of the multiple reaction monitoring precursor and product ion transitions for the 16 priority PACs and alkylated PACs that were analyzed in the samples are included in Idowu et al., 2018.
Samples Collection
The comparisons of interest in the study were 1) 2017 and 2018 to determine yearly gene expression changes in baseline samples and 2) 2018 and dilbit-exposed samples (2019) to determine gene expression changes following dilbit exposure. Thus, prior to dilbit additions into enclosures in September 2017, seven lake charr were collected from the surrounding lake environment and sampled for epidermal mucus. In September 2018, seven additional mucus samples were collected prior to cutting of limnocorral curtains and exposure of the lake water column to sunken dilbit. Finally, mucus was collected in September 2019 following exposure of dilbit to the lake water column. Thus, the experimental design is a temporal comparison pre- and post-exposure. Fish were captured from the lake from a boat using hook and line and were placed in separate bins of fresh lake water until processing (held <5 min). Retrieval and post-capture air exposure were minimized to reduce physiological stress. Water was renewed for each fish to prevent cross-contamination. Lake charr were placed in a solution of 0.1 g/L pH buffered (7.0) tricaine methanesulfonate (MS-222) mixed in lake water until fin movement ceased and fish were unresponsive to light pressure on the caudal fin. Total length, fork length, and weight of the individuals were recorded. Sex was determined through external morphological characteristics if possible. Mucus was collected by gently scraping the skin with a stainless-steel spatula, avoiding displacement of scales, to aggregate the mucus along the body axis in the anterior to posterior direction. Mucus was aspirated using a syringe, dispensed into a microcentrifuge tube, placed on dry ice, and stored at −80°C until analyzed. Lake charr were then released back into the lake. As the lake charr population is small and, thus, vulnerable at IISD-ELA, additional tissue types or blood via lethal/invasive sampling were not collected. Tools and work area were cleansed with ethanol and then lake water between fish to prevent cross-contamination. Condition factor (K) was calculated for each individual using formula K = weight/length3 × 100. Statistical differences in average weight, total length, and condition factor among 2017, 2018, and 2019 lake charr were tested using a one-way ANOVA and Tukey multiple comparisons test at a significance threshold of p ≤ 0.05. All capture and handling procedures were conducted in accordance with the Canadian Council on Animal Care approved protocols.
cDNA Library Generation and Illumina Sequencing
Total RNA was extracted from bulk mucus samples (n = 4 per year) using the RNeasy Lipid Tissue Mini Kit (Qiagen, Qiagen, Germantown, MD). Total RNA concentrations were quantified using an Invitrogen™ Qubit™ 4 Fluorometer (Invitrogen, Carlsbad, CA) and a NanoDrop™ ND-1000 Spectrophotometer (Thermo Fisher Scientific, Waltham, MA). RNA integrity was assessed on an Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA). Sequencing cDNA libraries were prepared using the NEBNext® Ultra™ II Directional RNA Library Prep Kit for Illumina® (New England Biolabs, Ipswich, MA) with the NEBNext® rRNA Depletion Kit using 500 ng of total RNA as input. NEBNext® Multiplex Oligos for Illumina® Index Primer Sets were used to multiplex cDNA libraries. Final cDNA libraries were assessed on an Agilent 2100 Bioanalyzer to verify proper fragment sizes and to quantify final library concentrations. The 2018 and dilbit-exposed samples (2019) cDNA libraries (n = 8) were pooled according to treatment group and the resulting samples were assessed on an Agilent 2100 Bioanalyzer. Pooled, multiplexed libraries underwent 1 × 75 bp sequencing on an Illumina NextSeq™ 500 at the Institute for Integrative Genome Biology at the University of California, Riverside. The 2017 cDNA libraries (n = 4) were processed using the same methods but were analyzed on a separate run prior to running the 2018 and 2019 libraries.
Bioinformatics Analysis
Raw sequencing reads were submitted to the NCBI Sequence Read Archive under the BioProject ID PRJNA729809. Quality of raw read data was assessed with FastQC v.0.11.7 (Andrews, 2010) prior to downstream analysis. Removal of adapter sequences and poor quality reads was performed using Trimmomatic v.0.36 (Bolger et al., 2014) with the following parameters: ILLUMINACLIP: TruSeq3-SE.fa:2:30:10 LEADING: 3 TRAILING: 3 SLIDINGWINDOW: 4:15 MINLEN: 36. Read quality was again assessed with FastQC following trimming. Using Bowtie2 (Langmead and Salzberg, 2012), read data for each sample was aligned to a de novo brook trout (Salvelinus fontinalis) transcriptome available for download at the PhyloFish database (http://phylofish.sigenae.org/index.html) (Pasquier et al., 2016). The transcriptome was annotated against the UniProtKB/Swiss-Prot database (http://www.ebi.ac.uk/uniprot/) using NCBI-BLASTx v.2.8.1 + reporting only the top hit (-max_target_seqs 1).
The comparisons of interest in the study were 1) 2017 and 2018 to determine yearly gene expression changes in baseline samples and 2) 2018 and dilbit-exposed samples (2019) to determine gene expression changes following potential dilbit exposure. RSEM (Li and Dewey, 2011) was used to quantify abundance of transcripts in each sample. To visualize transcriptome-wide patterns among 2018 baseline and 2019 dilbit-exposed samples, unsupervised hierarchical clustering was used to generate a sample-to-sample Euclidian distance heat map using the RSEM count matrix that underwent a regularized-logarithm transformation. Principal component analysis (PCA) was also performed on the transformed RSEM count matrix for baseline and dilbit-treated samples. Unsupervised hierarchical clustering was used to produce a heat map of the top 500 most-variable transcripts among 2018 and dilbit-treated samples. The top 500 most-variable transcripts were extracted from the transformed count matrix using the genefilter R package (Gentleman et al., 2019). The non-transformed transcript count matrix was input into DESeq2 v.1.23.6 (Love et al., 2014) to determine differentially expressed transcripts among 2018 and dilbit-treated samples at p ≤ 0.05 after Benjamini–Hochberg false discovery rate (FDR) correction. Differential expression analysis was also conducted among 2017 and 2018 baseline data to examine yearly changes in gene expression. Differentially-expressed transcripts were input into DAVID for gene ontology (GO) and KEGG pathway analysis as well as into Ingenuity Pathway Analysis (IPA).
qPCR Validation of RNA Sequencing Results
CDNA was created for 2018 baseline samples and dilbit-exposed samples using the Promega Reverse Transcription System (Promega, Madison, WI) with 500 ng of total RNA as input. Melt curve analysis and agarose gel electrophoresis were performed prior to qPCR to confirm primer specificity. The reactions for qPCR consisted of 10 μL SsoAdvanced universal SYBR Green Supermix (Bio-Rad, Hercules, CA), 0.2 μM forward and reverse primers, and 2 μL of cDNA in a final volume of 20 μl. Samples were run in triplicate for each gene on a CF× Connect Real-Time PCR Detection System (Bio-Rad, Hercules, CA) with the following thermal cycler conditions: 95°C for 10 min, 40 cycles of 95°C for 15 s, 58°C for 30 s, and 72°C for 30 s. Relative expression was determined by the 2−ΔΔCt method (Livak and Schmittgen, 2001) with β-actin as the normalizing gene. Primer sequences were designed using IDT PrimerQuest Tool® with transcript sequences from the brook trout assembly. Primer information is listed in Supplementary Table S1.
Results and Discussion
Soluble Dilbit Constituents in Lake 260
The present study was a part of a larger series of experiments at IISD-ELA where dilbit spills and cleanups were carried out within limnocorrals and shoreline enclosures in a boreal lake (Rodriguez-Gil et al., 2021). There were two potential vectors for dilbit constituents entering the surround lake environment. First, following cutting of limnocorral curtains on 27 September 2018, sunken dilbit remained on the sediment and open to the lake environment for 323 days until cleanup on 16 August 2019. Second, following the completion of dilbit additions to shoreline enclosures on 22 June 2019, tracer studies indicated that some leakage from the enclosures to the outside lake environment occurred which may have contributed water soluble dilbit compounds into the lake. Thus, the present study assessed the potential impacts of residual dilbit following a spill cleanup on wild fish residing in the lake outside of the enclosures.
ΣPAC concentrations of 0.058 μg/L were detected in lake surface waters prior to dilbit additions into shoreline enclosures in June 2019 (Figure 1); these levels could be linked to residual dilbit in sediments following previous 2018 dilbit experiments, through atmospheric sources, or from background PACs from organic sediment inputs. Following the additions of dilbit into shoreline enclosures on June 22 (Day 0), there was an increase in PACs within surface waters, reaching a maximum concentration of 2.29 μg/L ΣPAC on Day 5 (Figure 1). ΣPAC concentrations in surface waters declined to 1.05 μg/L on Day 6 and remained under 1 μg/L for the remainder of the exposure period (Figure 1). By Day 66, ΣPAC concentrations in surface waters declined to 0.082 μg/L ΣPAC concentrations at the time of sampling mucus from dilbit-exposed lake charr was approximately 0.067 μg/L (interpolation from Day 66 and Day 81 data points) (Figure 1). Three- and 4-ring compounds were the most dominant of the total PACs, with phenanthrenes, dibenzothiophenes, pyrenes, retene, and napthalenes being the most abundant compound groups detected in lake water outside of the enclosures (Supplementary Material; additional manuscript forthcoming further detailing these chemistry data). As dilbit was exposed to weathering in the field, it is likely that lighter molecular weight compounds volatilized, leaving an increased proportion of higher molecular weight 3- and 4-ring PACs. Chemistry results are supported by previous studies simulating dilbit spills in outdoor tanks containing lake water and sediments where rapid desorption of PACs into the water column was observed, most notably phenanthrenes, dibenzothiophenes, and fluorenes (Stoyanovich et al., 2019).
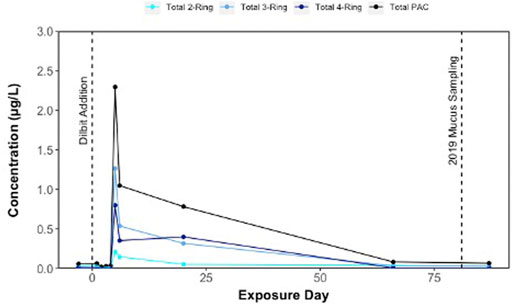
FIGURE 1. Concentrations (µg/L) of polycyclic aromatic compounds (PACs) in surface waters of Lake 260 following dilbit addition into fourteen shoreline enclosures on the lake on Day 0 (22 June 2019). Lake charr epidermal mucus was sampled on Day 81 (10 September 2019).
Routes of exposure to oil constituents in fish include ventilation through gills, dermal uptake, and ingestion (Nichols et al., 1996; Almeda et al., 2013). As dilbit constituents were detected within the water column, absorption of waterborne compounds through skin and gills was one probable route of exposure. Moreover, lake charr may also have ingested oil-contaminated organisms, such as zooplankton and forage fish which were more likely to have contact with sunken dilbit (Almeda et al., 2013; National Academies of Sciences Medicine and Engineering, 2016). Through consumption of oil-contaminated material, dilbit constituents such as PACs may be transferred to apex predatory species through trophic transfer within the food web. Thus, exposure to PACs may continue even after most constituents have volatilized from the water column. Finally, there was also potential for exposure via direct contact with sunken dilbit that remained on the sediments. Additional studies on the trophic movement of dilbit constituents in lakes are needed to better understand bioaccumulation processes in boreal lake systems.
Bioinformatics Output and Transcriptomic Responses
Lake charr were collected by hook and line angling from Lake 260 in September of 2017 and 2018 (prior to dilbit exposure) and September of 2019 after dilbit exposure (Supplementary Table S2). Average weights for 2017, 2018, and 2019 lake charr were 644.3 ± 87.9 g, 814.3 ± 92.8 g, and 961.9 ± 71.5 g, respectively (Figure S1). Average total lengths were 42.59 ± 1.9 cm in 2017, 46.8 ± 1.7 cm in 2018, and 48.74 ± 1.3 cm in 2019 (Supplementary Figure S1). Moreover, average condition factors were 0.80 ± 0.02 in 2017, 0.78 ± 0.02 in 2018, and 0.82 ± 0.02 in 2019 (Supplementary Figure S1). Average weight and total lengths were significantly lower in 2017 compared to 2019 (p ≤ 0.04 for both comparisons), but there were no significant differences in weight and length among 2018 baseline and 2019 dilbit-treated individuals. There were no significant differences in condition factor among all years. Mucus samples from four individuals from each year were sequenced. The 12 FASTQ files generated from Illumina sequencing encompassed a total of 616 M single-end raw reads, with an average of 51 M reads per sample (Supplementary Table S3).
To determine year-year differences in background transcriptional signatures, comparisons were made in fish sampled prior to dilbit exposure in 2017 and 2018. A total of 1,576 transcripts were differentially expressed among 2017 and 2018 baseline years at a FDR adjusted p-value ≤ 0.05 (Supplementary Material). The top GO biological processes were “translation”, “ribosomal small unit assembly”, and “in utero embryonic development”, whereas the top GO molecular functions were “structural constituent of ribosome”, “poly(A) RNA binding”, and “mRNA binding” (Supplementary Material).
Comparing animals pre- and post-dilbit exposure, the sample-to-sample Euclidian distance heat map (Supplementary Figure S2) and PCA of transcriptome-wide count data (Figure 2A) indicated clustering of individuals based on treatment. Unsupervised hierarchical clustering of the top 500 most-variable transcripts also clustered individuals by treatment (Figure 2B). Thus, results of clustering analyses indicate that there were clear differences in the mucus transcriptomic profiles of lake charr following additions of dilbit. PCA of transcriptome-wide count data did not show distinct clustering patterns based on sex, suggesting that transcriptomic changes among years were driven by treatment (Supplementary Figure S3).
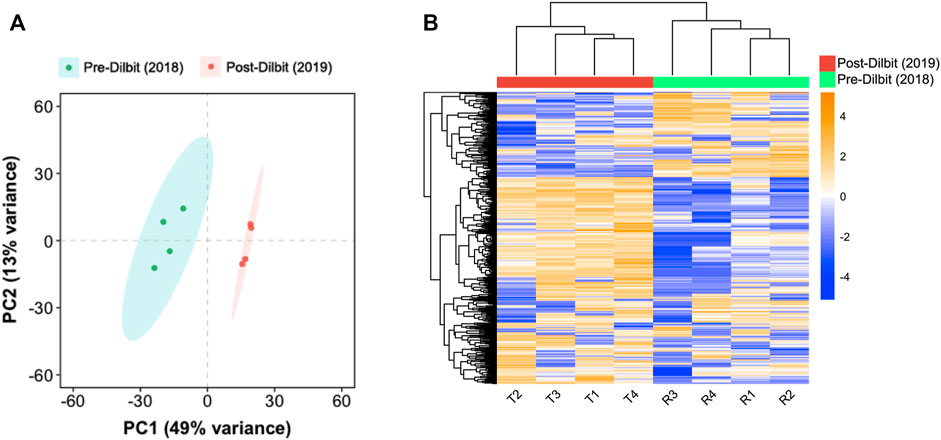
FIGURE 2. Comparison of pre- and post-dilbit exposed lake charr using (A) principal component analysis of transcriptome-wide count data and (B) unsupervised hierarchical clustering of the top 500 most variable transcripts.
A total of 714 transcripts were differentially expressed at a FDR adjusted p-value ≤ 0.05 among individuals sampled in pre-and post-dilbit (Supplementary Table S4). The RNA sequencing results were confirmed through qPCR (Supplementary Figure S4). Among all pathway analyses, there was prevalence of biological pathways related to intermediary metabolism, more specifically relating to carbohydrates and amino acids. For example, of the 12 KEGG pathways identified, three were related to intermediary metabolism including ‘arginine biosynthesis’ (rank 3), ‘alanine, aspartate, and glutamate metabolism’ (rank 5), and ‘carbon metabolism’ (rank 11) (Figure 3; Supplementary Table S5). Intermediary metabolism pathways were also identified in the GO analysis, including the biological process ‘urea cycle’ (rank 11) as well as the molecular functions ‘glucose binding’ (rank 9) and ‘carbamoyl-phosphate synthase activity’ (rank 18) (Figure 3; Supplementary Table S6; Supplementary Table S7). This trend was also seen in the IPA results, where the top canonical pathways (ranked by p value) included ‘superpathway of citrulline metabolism’ (rank 1), ‘arginine degradation 1’ (rank 10), ‘urea cycle’ (rank 18), and ‘arginine degradation VI (arginase 2 pathway)’ (rank 19) (Figure 4). Taken together, results of the pathway analyses suggest molecular changes in metabolic and energetic processes in lake charr following dilbit exposure. It should also be pointed out that the number of differentially expressed genes was greater among baseline year comparisons, suggesting that differences in environmental variables among years contribute a significant amount of variation. While field-based toxicological studies may be beneficial as they capture effects from multiple stressors, additional laboratory studies may aid in discerning effects from potential confounding environmental variables (discussed further in text).
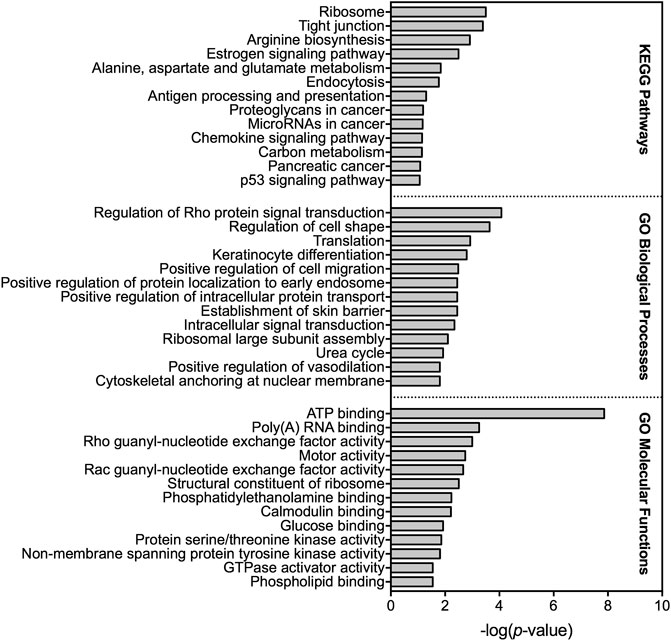
FIGURE 3. Top KEGG pathways as well as gene ontology (GO) biological processes and molecular functions altered based on differential gene expression analysis among 2018 pre-dilbit lake charr and 2019 post-dilbit lake charr.
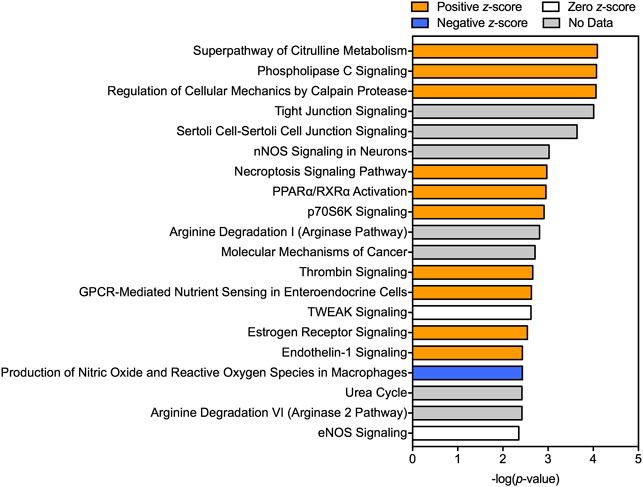
FIGURE 4. Top canonical pathways predicted by Ingenuity Pathway Analysis (IPA) to be affected following dilbit exposure. A positive z-score indicates predicted activation whereas a negative z-score indicates a predicted inhibition.
Several studies in fish have shown that exposure to PACs and oil, can alter energy metabolism mechanics and induce energy mobilization causing increased energy demands due to stress (Wendelaar Bonga, 1977; Frasco and Guilhermino, 2002; Tintos et al., 2008; Vijayan et al., 2010; Ings et al., 2012; Hook et al., 2018). Stress-induced changes in metabolism are associated with a surge of cortisol, which results in activation of gluconeogenesis and, consequently, increased plasma glucose concentrations. Moreover, gluconeogenesis relies on non-carbohydrate sources such as lactate, glycerol, and amino acids for the formation of glucose. Due to the multiple pathways associated with amino acid metabolism, it is suggested that lake charr catabolized and degraded protein for energy production (Figure 3; Figure 4). Metabolism of amino acids for energy is further supported by the upregulation of alanine aminotransferase 2 (gpt2) which converts alanine into pyruvate for glucose production and has been shown to be enhanced in response to stress for gluconeogenesis (Figure 5) (Mommsen et al., 1999; Kumar et al., 2010; Qian et al., 2015). Other gluconeogenic and amino acid catabolism transcripts were also upregulated, including glutamate dehydrogenase (glud1), glutamine-fructose-6-phosphate aminotransferase (gfpt1), and ornithine aminotransferase (oat) (Figure 5). Furthermore, several of these transcripts are also involved in the urea cycle, likely enhancing the urea cycle to convert excess ammonia following breakdown of amino acids (Figure 3; Figure 4; Figure 5). Cortisol also elicits decreased potential for glycolysis in fish through decreased activities of pyruvate kinase, hexokinase, fructose 1,6-bisphosphate, and glucose-6-phosphate 1-dehydrogenase (Laiz-Carrión et al., 2003). In the present study, glycolytic transcripts such hexokinase-1 (hk-1) and glucose-6-phosphate 1-dehydrogenase (g6pd) were downregulated in lake charr mucus following dilbit exposure (Figure 5). As hexokinase is a rate-limiting enzyme of glycolysis, decreased transcription of hk-1 and other glycolytic transcripts supports the utilization of gluconeogenesis during stress-induced energy production. Although molecular changes suggested a shift in energetics, there were no differences in condition factor pre- and post-dilbit exposure, suggesting that the disruption was not enough to elicit changes in body mass.
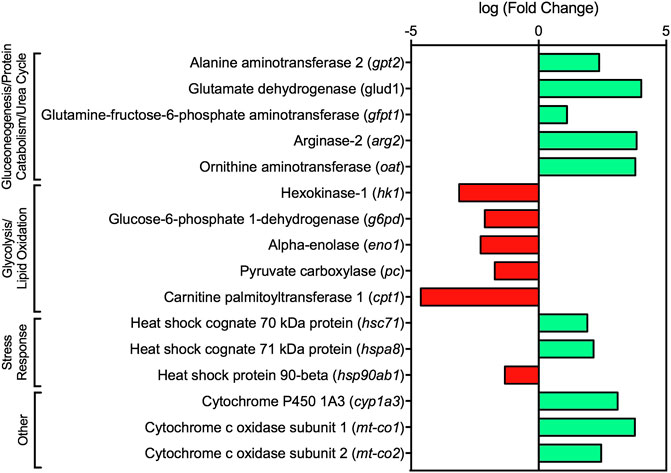
FIGURE 5. Fold change of select transcripts dysregulated in dilbit-exposed lake charr relating to energy metabolism and stress response from RNA sequencing results. All transcripts were significant at FDR adjusted p-value ≤ 0.05, except cyp1a3 which was significant at a FDR adjusted p-value ≤ 0.1.
Shifts in energy metabolism have been observed in fish following dilbit exposure. Previous studies exposing Atlantic salmon (Salmo salar) smolts and sockeye salmon embryos to dilbit resulted in elevated cardiac and total body triglyceride levels, suggesting decreased lipid utilization for energy compared to controls (Alderman et al., 2018; Avey et al., 2020). In the present study, decreased lipid utilization was suggested by downregulation of the transcript encoding for carnitine palmitoyltransferase I (cpt1a) which is the rate limiting step in long-chain fatty acid oxidation (Figure 5) (Frøyland et al., 1998; Qu et al., 2016). Furthermore, expression of genes involved in protein degradation was increased in muscle of dilbit-exposed Atlantic salmon smolts, namely ubiquitin which labels protein for proteasomal degradation (Avey et al., 2020). Although ubiquitin was not upregulated in lake charr mucus, several transcripts encoding for enzymes involved in ubiquitination of proteins were upregulated, including polyubiquitin, ubiquitin conjugating enzyme E2 D2, and others (Supplementary Table S4). Thus, results from the present and previous studies together suggest that dilbit may induce shifts in energy metabolism, such as increased protein degradation and potentially greater utilization of amino acids for energy. Although dilbit exposure has been shown to increase muscle damage in sockeye salmon (Alderman et al., 2017b), it is not known whether muscle damage is correlated to transcriptional changes involved in muscle protein degradation and catabolism. This hypothesis could be further investigated through analysis of energy stores within muscle in subsequent sampling events via non-lethal biopsy collection.
Perturbations in metabolism following dilbit exposure was also suggested by the upregulation of cytochrome c oxidase subunit 1 (mt-co1) and cytochrome c oxidase subunit 2 (mt-co2) which encode for catalytic subunits of cytochrome C oxidase (COX) (Figure 5) (Capaldi, 1990). COX functions as an aerobic respiratory enzyme and catalyzes the final step of the mitochondrial electron transport chain (Capaldi, 1990; Barrientos et al., 2002). In fish, the enzymatic activity of COX is an indicator of metabolic capacity and a biomarker of metabolic perturbation following exposure to petroleum hydrocarbons (Gagnon, 1999; Cohen et al., 2005). Following waterborne exposures to crude oil water-accommodated fractions, COX activity in the gills and liver of Australian bass (Macquaria novemaculeata) either increased or decreased compared to controls depending on concentration (14–134 μg/L Σpetroleum hydrocarbons) (Cohen et al., 2005). Similarly, COX activity was altered in the heart and red muscle of Atlantic salmon smolts following subchronic exposure to dilbit, being either increased or decreased relative to controls depending on tissue type and concentration (9.65 and 67.9 μg/L ΣPAC) (Avey et al., 2020). Thus, dysregulation of mt-co1 and mt-co2 in dilbit-exposed lake charr corroborates previous studies suggesting that crude oil induces alterations in COX and, thus, changes in energy metabolism. Although there are many abiotic and biotic factors that may contribute to stress response in the environment, the lack of molecular evidence for energetic changes among baseline years 2017–2018 provides a degree of support that the suggested energy metabolism changes in 2019 were a result of dilbit additions. For example, there were no GO pathways related to intermediary metabolism or energetics among 2017 and 2018, suggesting no molecular changes in energy metabolism year-to-year at baseline conditions. Furthermore, only two genes (g6pd and gpt2) relating to energy metabolism were dysregulated among baseline conditions of the genes identified to be altered following dilbit additions (Figure 5).
Identification of Biomarkers
The expression patterns of certain genes may be used as biomarkers of exposure and can help researchers quickly assess whether an individual has been exposed to a contaminant of interest. The induction of CYP1A has commonly been used as biomarker of exposure to petroleum hydrocarbons and other organic pollutants (Goksøyr, 1995; Woodin et al., 1997; Whyte et al., 2000) and exposure to dilbit has also been shown to induce CYP1A activity in several fish species (Madison et al., 2015, 2017, Alderman et al., 2017a, 2017b, 2018; Alsaadi F. M. et al., 2018; McDonnell et al., 2019). In the present study, there was a trend of cyp1a3 upregulation in lake charr following dilbit additions (FDR-adj p = 0.08) (Figure 5; Supplementary Table S4). Although not statistically significant at a p = 0.05, a trend in upregulation of cyp1a3 may have biological significance and may be a viable biomarker in mucus, especially at higher PAC concentrations seen following an oil spill. A study exposing Japanese medaka embryos to water accommodated fractions (WAF) of Cold Lake dilbit indicated that cyp1a mRNA synthesis increased monotonically with concentration in hatched fish (Madison et al., 2017). In the present study, ΣPAC levels within surface waters were relatively low at the period of sampling (approximately 0.067 μg/L) and a greater cyp1a response would likely be expected at higher concentrations observed during a spill. Our results are consistent with findings of a previous study in which cyp1a1 and cyp1b1 were the top upregulated transcript in mucus of mahi-mahi following exposure to Deepwater Horizon slick oil (16.55 μg/L ΣPAC) (Greer et al., 2019). In the present study, there was a clear trend of cyp1a3 upregulation which was confirmed by qPCR at relatively low environmental concentrations and confirms that this may be a sensitive biomarker even within mucus of animals.
Applications and Implications
Overall, results of RNA sequencing suggest stress following dilbit exposure, as indicated by upregulation of transcripts involved in cortisol-induced energy metabolism. More specifically, evidence suggests that there was a potential increased reliance on amino acid metabolism and decreased reliance on fatty acid oxidation for energy production in dilbit-exposed lake charr compared to controls. The changes in energy metabolism and utilization support previous studies which have also shown that crude oil and dilbit specifically may induce changes in energy metabolism (Alderman et al., 2018; Avey et al., 2020). Moreover, cyp1a3 was upregulated in lake charr mucus, suggesting a molecular response to dilbit. Further laboratory studies should be conducted to confirm these molecular changes however, coupled with assessment of phenotypic endpoints. Laboratory studies may also be beneficial as potential confounding variables in the field can be eliminated, such as varying environmental variables among sampling years. Along with this, future studies should utilize greater replication for a more definitive results, as this preliminary study utilized 4 individuals per treatment group. As RNA sequencing identified molecular changes related to energy metabolism and PAC exposure in lake charr mucus, results of the present study indicate that transcriptomic analysis of epidermal mucus is a promising vector for development of nonlethal oil exposure assessment methods.
There are some realities that should be considered when using molecular analysis of mucus for exposure assessment following an oil spill. Although RNA sequencing is a holistic approach to capturing molecular changes in an organism, use of targeted qPCR for genes such as cyp1a and ahr may be a more rapid and inexpensive method for monitoring exposure following an oil spill. Along with this, mucus from reference individuals will be needed for either RNA sequencing or qPCR analysis. Although use of baseline mucus samples from the population of interest would be ideal to minimize confounding variables, it may not be feasible. Instead, mucus from individuals from a well-characterized reference location may be used. Use of other nonlethal endpoints in conjunction with analysis of mucus for target genes of interest may provide a more complete insight on organism health. Examples include analysis of blood for enzymatic activity or use of muscle plugs for contaminant loading assessment. There are also logistical issues that require further research before the method may be applied for spill monitoring to determine magnitude and geographic extent of effects. For example, further research is needed to identify how rapidly the mucus transcriptome responds to external conditions as well as to determine the turnover rate of the mucus transcriptome. Furthermore, the relationship among gene expression in actively metabolizing tissues, such as liver, and the mucus should be identified in order to better understand the physiological significance of differential expression in the mucus. While additional research is needed, use of epidermal mucus is a promising method for oil exposure assessment without the use of lethal sampling.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA729809.
Ethics Statement
The animal study was reviewed and approved by the Canadian Council on Animal Care.
Author Contributions
NA contributed to study design, collected samples, conducted bioinformatics and statistical analyses, and wrote the manuscript. VP contributed to study design, collected samples, provided field support, and edited the manuscript. LH collected samples, provided field support, and edited the manuscript. LP conducted chemistry analysis. DS contributed to study design, provided field support, and edited the manuscript. All authors contributed final refinements to the manuscript.
Funding
This project received funding and research support as a part of a large multidisciplinary program that includes participation from the Canadian Association of Petroleum Producers (CAPP), the Canadian Energy Pipelines Association (CEPA), Environment and Climate Change Canada, Fisheries and Oceans Canada, Natural Resources Canada, Ontario Municipal Employees Coordinating Committee, Ontario Ministry of Natural Resources and Forestry, National Energy Board, Institut national de la recherche scientifique, University of Manitoba, University of Ottawa, Queen’s University, University of Calgary, University of Saskatchewan, and McGill University.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Acknowledgments
Additionally, we would like to thank Gregg Tomy and the Centre for Oil and Gas Research and Development (COGRAD).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fenvs.2022.836640/full#supplementary-material
References
Adams, J., Larter, S., Bennett, B., Huang, H., Westrich, J., Kruisdijk, C. v., et al. (2013). The Dynamic Interplay of Oil Mixing, Charge Timing, and Biodegradation in Forming the Alberta Oil SandsInsights from Geologic Modeling and Biogeochemistry. Heavy-oil Oil-sand Pet. Syst. Alberta Beyond. 64, 23–102. doi:10.1306/13371578St643552
Alderman, S. L., Dindia, L. A., Kennedy, C. J., Farrell, A. P., and Gillis, T. E. (2017a). Proteomic Analysis of Sockeye salmon Serum as a Tool for Biomarker Discovery and New Insight into the Sublethal Toxicity of Diluted Bitumen. Comp. Biochem. Physiol. D: Genomics Proteomics 22, 157–166. doi:10.1016/j.cbd.2017.04.003
Alderman, S. L., Lin, F., Farrell, A. P., Kennedy, C. J., and Gillis, T. E. (2017b). Effects of Diluted Bitumen Exposure on Juvenile Sockeye salmon: From Cells to Performance. Environ. Toxicol. Chem. 36, 354–360. doi:10.1002/etc.3533
Alderman, S. L., Lin, F., Gillis, T. E., Farrell, A. P., and Kennedy, C. J. (2018). Developmental and Latent Effects of Diluted Bitumen Exposure on Early Life Stages of Sockeye salmon (Oncorhynchus nerka). Aquat. Toxicol. 202, 6–15. doi:10.1016/j.aquatox.2018.06.014
Almeda, R., Wambaugh, Z., Chai, C., Wang, Z., Liu, Z., and Buskey, E. J. (2013). Effects of Crude Oil Exposure on Bioaccumulation of Polycyclic Aromatic Hydrocarbons and Survival of Adult and Larval Stages of Gelatinous Zooplankton. PLoS One 8, e74476–21. doi:10.1371/journal.pone.0074476
Alsaadi, F., Hodson, P. V., and Langlois, V. S. (2018a). An Embryonic Field of Study: The Aquatic Fate and Toxicity of Diluted Bitumen. Bull. Environ. Contam. Toxicol. 100, 8–13. doi:10.1007/s00128-017-2239-7
Alsaadi, F. M., Madison, B. N., Brown, R. S., Hodson, P. V., and Langlois, V. S. (2018b). Morphological and Molecular Effects of Two Diluted Bitumens on Developing Fathead Minnow (Pimephales promelas). Aquat. Toxicol. 204, 107–116. doi:10.1016/j.aquatox.2018.09.003
Avey, S. R., Kennedy, C. J., Farrell, A. P., Gillis, T. E., and Alderman, S. L. (2020). Effects of Diluted Bitumen Exposure on Atlantic salmon Smolts: Molecular and Metabolic Responses in Relation to Swimming Performance. Aquat. Toxicol. 221, 105423. doi:10.1016/j.aquatox.2020.105423
Barrientos, A., Barros, M. H., Valnot, I., Rötig, A., Rustin, P., and Tzagoloff, A. (2002). Cytochrome Oxidase in Health and Disease. Gene 286, 53–63. doi:10.1016/s0378-1119(01)00803-4
Bolger, A. M., Lohse, M., and Usadel, B. (2014). Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 30, 2114–2120. doi:10.1093/bioinformatics/btu170
Boufadel, M., Chen, B., Foght, J., Hodson, P., Lee, K., Swanson, S., et al. (2015). The Behaviour and Environmental Impacts of Crude Oil Released into Aqueous Environments Fall.
Canada Energy Regulator (2018). Crude Oil Annual Export Summary – 2018. Available at: https://www.cer-rec.gc.ca/nrg/sttstc/crdlndptrlmprdct/stt/crdlsmmr/crdlsmmr-eng.html.
Canada National Energy Board (2006). Canada’s Oil Sands Opportunities and Challenges to 2015: An Update.
Capaldi, R. A. (1990). Structure and Function of Cytochrome C Oxidase. Annu. Rev. Biochem. 59, 569–596. doi:10.1146/annurev.bi.59.070190.003033
Cohen, A., Gagnon, M. M., and Nugegoda, D. (2005). Alterations of Metabolic Enzymes in Australian Bass, Macquaria Novemaculeata, after Exposure to Petroleum Hydrocarbons. Arch. Environ. Contam. Toxicol. 49, 200–205. doi:10.1007/s00244-004-0174-1
Dew, W. A., Hontela, A., Rood, S. B., and Pyle, G. G. (2015). Biological Effects and Toxicity of Diluted Bitumen and its Constituents in Freshwater Systems. J. Appl. Toxicol. 35, 1219–1227. doi:10.1002/jat.3196
Dupuis, A., and Ucan-Marin, F. (2015). A Literature Review on the Aquatic Toxicology of Petroleum Oil: an Overview of Oil Properties and Effects to Aquatic Biota. DFO Can. Sci. Advis. Secr. Res. Doc. 2015/007, 51.
Dzul-Caamal, R., Salazar-Coria, L., Olivares-Rubio, H. F., Rocha-Gómez, M. A., Girón-Pérez, M. I., and Vega-López, A. (2016). Oxidative Stress Response in the Skin Mucus Layer of Goodea Gracilis (Hubbs and Turner, 1939) Exposed to Crude Oil: A Non-invasive Approach. Comp. Biochem. Physiol. A: Mol. Integr. Physiol. 200, 9–20. doi:10.1016/j.cbpa.2016.05.008
Frasco, M. F., and Guilhermino, L. (2002). Effects of Dimethoate and Beta-Naphthoflavone on Selected Biomarkers of Poecilia reticulata. Fish. Physiol. Biochem. 26, 149–156. doi:10.1023/A:1025457831923
Frøyland, L., Madsen, L., Eckhoff, K. M., Lie, Ø., and Berge, R. K. (1998). Carnitine Palmitoyltransferase I, Carnitine Palmitoyltransferase II, and Acyl-CoA Oxidase Activities in Atlantic salmon (Salmo salar). Lipids 33, 923–930. doi:10.1007/s11745-998-0289-4
Gagnon, M. M., and Holdway, D. A. (1999). Metabolic Enzyme Activities in Fish Gills as Biomarkers of Exposure to Petroleum Hydrocarbons. Ecotoxicology Environ. Saf. 44, 92–99. doi:10.1006/eesa.1999.1804
Gentleman, R., Carey, V., Huber, W., and Hahne, F. (2019). Genefilter: Methods for Filtering Genes from High-Throughput Experiments. version 1.68.0.R. Packag
Goksøyr, A. (1995). Use of Cytochrome P450 1A (CYP1A) in Fish as a Biomarker of Aquatic Pollution BT - Toxicology in Transition. in, eds. G. H. Degen, J. P. Seiler, and P. Bentley (Berlin, Heidelberg: Springer Berlin Heidelberg), 80–95.
Greer, J. B., Andrzejczyk, N. E., Mager, E. M., Stieglitz, J. D., Benetti, D., Grosell, M., et al. (2019). Whole-Transcriptome Sequencing of Epidermal Mucus as a Novel Method for Oil Exposure Assessment in Juvenile Mahi-Mahi (Coryphaena hippurus). Environ. Sci. Technol. Lett. 6, 538–544. doi:10.1021/acs.estlett.9b00479
Hook, S. E., Mondon, J., Revill, A. T., Greenfield, P. A., Stephenson, S. A., Strzelecki, J., et al. (2018). Monitoring Sublethal Changes in Fish Physiology Following Exposure to a Light, Unweathered Crude Oil. Aquat. Toxicol. 204, 27–45. doi:10.1016/j.aquatox.2018.08.013
Idowu, I., Francisco, O., Thomas, P. J., Johnson, W., Marvin, C., Stetefeld, J., et al. (2018). Validation of a Simultaneous Method for Determining Polycyclic Aromatic Compounds and Alkylated Isomers in Biota. Rapid Commun. Mass. Spectrom. 32, 277–287. doi:10.1002/rcm.8035
Ings, J. S., Vijayan, M. M., and Servos, M. R. (2012). Tissue-specific Metabolic Changes in Response to an Acute Handling Disturbance in Juvenile Rainbow trout Exposed to Municipal Wastewater Effluent. Aquat. Toxicol. 108, 53–59. doi:10.1016/j.aquatox.2011.09.009
Kumar, V., Sahu, N. P., Pal, A. K., Kumar, S., Sinha, A. K., Ranjan, J., et al. (2010). Modulation of Key Enzymes of Glycolysis, Gluconeogenesis, Amino Acid Catabolism, and TCA Cycle of the Tropical Freshwater Fish Labeo Rohita Fed Gelatinized and Non-gelatinized Starch Diet. Fish. Physiol. Biochem. 36, 491–499. doi:10.1007/s10695-009-9319-5
Laiz-carrión, R., Martín Del Río, M. P., Miguez, J. M., Mancera, J. M., and Soengas, J. L. (2003). Influence of Cortisol on Osmoregulation and Energy Metabolism in Gilthead seabreamSparus Aurata. J. Exp. Zool. 298A, 105–118. doi:10.1002/jez.a.10256
Langmead, B., and Salzberg, S. L. (2012). Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 9, 357–359. doi:10.1038/nmeth.1923
Lee, K., Boufadel, M., Chen, B., Foght, J., Hodson, P., Swanson, S., et al. (2016). High-priority Research Needs for Oil Spills in Canada: Summary of a Royal Society Expert Panel Report on the Behaviour and Environmental Impacts of Crude Oil Released into Aqueous Environments. 39th AMOP Tech. Semin. Environ. Contam. Response.
Li, B., and Dewey, C. N. (2011). RSEM: Accurate Transcript Quantification from RNA-Seq Data with or without a Reference Genome. BMC Bioinformatics 12. doi:10.1186/1471-2105-12-323
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 25, 402–408. doi:10.1006/meth.2001.1262
Love, M. I., Huber, W., and Anders, S. (2014). Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 15. doi:10.1186/s13059-014-0550-8
Madison, B. N., Hodson, P. V., and Langlois, V. S. (2017). Cold Lake Blend Diluted Bitumen Toxicity to the Early Development of Japanese Medaka. Environ. Pollut. 225, 579–586. doi:10.1016/j.envpol.2017.03.025
Madison, B. N., Hodson, P. V., and Langlois, V. S. (2015). Diluted Bitumen Causes Deformities and Molecular Responses Indicative of Oxidative Stress in Japanese Medaka Embryos. Aquat. Toxicol. 165, 222–230. doi:10.1016/j.aquatox.2015.06.006
Marett-Crosby, M., Fay, R., Groark, C., Kani, A., Smith, J., and Sullivan, T. (2013). Transporting Alberta’s Oil Sands Products: Defining the Issues and Assessing the Risks. NOAA Tech. Memo. NOS OR&R 44. Wonderful Demons. Available at: http://scholar.google.com/scholar?hl=en&btnG=Search&q=intitle:Transporting+Alberta+Oil+Sands+Products+:+Defining+the+Issues+and+Assessing+the+Risks#0.153166doi:10.1007/978-1-4614-6800-4_13
McDonnell, D., Madison, B. N., Baillon, L., Wallace, S. J., Brown, S. R., Hodson, P. V., et al. (2019). Comparative Toxicity of Two Diluted Bitumens to Developing Yellow Perch (Perca flavescens). Sci. Total Environ. 655, 977–985. doi:10.1016/j.scitotenv.2018.11.199
Mommsen, T. P., Vijayan, M. M., and Moon, T. W. (1999). Cortisol in Teleosts: Dynamics, Mechanisms of Action, and Metabolic Regulation. Rev. Fish. Biol. Fish. 9, 211–268. doi:10.1023/A:1008924418720
National Academies of Sciences Medicine and Engineering (2016). Spills of Diluted Bitumen from Pipelines: A Comparative Study of Environmental Fate, Effects, and Response. Washington, DC: The National Academies Press. doi:10.17226/21834
Nichols, J. W., McKim, J. M., Lien, G. J., Hoffman, A. D., Bertelsen, S. L., and Elonen, C. M. (1996). A Physiologically Based Toxicokinetic Model for Dermal Absorption of Organic Chemicals by Fish. Toxicol. Sci. 31, 229–242. doi:10.1006/faat.1996.009510.1093/toxsci/31.2.229
Orihel, D. (2019). Why We Spilled Oil for Science. Can. Geogr. Available at: https://www.canadiangeographic.ca/article/why-we-spilled-oil-science.
Palace, V. (2021). “Session PS8-05 Unconvetional Products: Title: The Freshwater Oil Spill Remediation Study (FOReSt): 2018 Pilot Study at the Experimental Lakes Area, Canada,” in International Oil Spill Conference.
Pasquier, J., Cabau, C., Nguyen, T., Jouanno, E., Severac, D., Braasch, I., et al. (2016). Gene Evolution and Gene Expression after Whole Genome Duplication in Fish: The PhyloFish Database. BMC Genomics 17, 1–10. doi:10.1186/s12864-016-2709-z
Philibert, D. A., Philibert, C. P., Lewis, C., and Tierney, K. B. (2016). Comparison of Diluted Bitumen (Dilbit) and Conventional Crude Oil Toxicity to Developing Zebrafish. Environ. Sci. Technol. 50, 6091–6098. doi:10.1021/acs.est.6b00949
Qian, K., Zhong, S., Xie, K., Yu, D., Yang, R., and Gong, D.-W. (2015). Hepatic ALT Isoenzymes Are Elevated in Gluconeogenic Conditions Including Diabetes and Suppressed by Insulin at the Protein Level. Diabetes Metab. Res. Rev. 31, 562–571. doi:10.1002/dmrr.2655
Qu, Q., Zeng, F., Liu, X., Wang, Q. J., and Deng, F. (2016). Fatty Acid Oxidation and Carnitine Palmitoyltransferase I: Emerging Therapeutic Targets in Cancer. Cell Death Dis 7, e2226. doi:10.1038/cddis.2016.132
Rodriguez-Gil, J. L., Stoyanovich, S., Hanson, M. L., Hollebone, B., Orihel, D. M., Palace, V., et al. (2021). Simulating Diluted Bitumen Spills in Boreal lake Limnocorrals - Part 1: Experimental Design and Responses of Hydrocarbons, Metals, and Water Quality Parameters. Sci. Total Environ. 790, 148537. doi:10.1016/J.SCITOTENV.2021.148537
Stoyanovich, S. S., Yang, Z., Hanson, M., Hollebone, B. P., Orihel, D. M., Palace, V., et al. (2019). Simulating a Spill of Diluted Bitumen: Environmental Weathering and Submergence in a Model Freshwater System. Environ. Toxicol. Chem. 38, 2621–2628. doi:10.1002/etc.4600
Tintos, A., Gesto, M., Míguez, J. M., and Soengas, J. L. (2008). β-Naphthoflavone and Benzo(a)pyrene Treatment Affect Liver Intermediary Metabolism and Plasma Cortisol Levels in Rainbow trout Oncorhynchus mykiss. Ecotoxicology Environ. Saf. 69, 180–186. doi:10.1016/j.ecoenv.2007.03.009
U.S. Fish and Wildlife Service (USFWS) (2015). Final Damage Assessment and Restoration Plan/Environmental Assessment for the July 25-26, 2010 Enbridge Line 6B Oil Discharges Near Marshall. Washington DC: MI, 104.
Vijayan, M. M., Aluru, N., and Leatherland, J. F. (2010). “Stress Response and the Role of Cortisol,” in Fish Diseases and Disorders. Editors J. F. Leatherland, and P. T. K. Woo (Wallingford: CAB International), Vol. 2.
Whyte, J. J., Jung, R. E., Schmitt, C. J., and Tillitt, D. E. (2000). Ethoxyresorufin-O-deethylase (EROD) Activity in Fish as a Biomarker of Chemical Exposure. Crit. Rev. Toxicol. 30, 347–570. doi:10.1080/10408440091159239
Woodin, B. R., Smolowitz, R. M., and Stegeman, J. J. (1997). Induction of Cytochrome P4501A in the Intertidal Fish Anoplarchus Purpurescens by Prudhoe Bay Crude Oil and Environmental Induction in Fish from Prince William Sound. Environ. Sci. Technol. 31, 1198–1205. doi:10.1021/es9607190
Keywords: nonlethal approach, transcriptomics, ecotoxicology, diluted bitumen, aquatic toxicology
Citation: Andrzejczyk NE, Palace VP, Hrenchuk L, Peters L and Schlenk D (2022) RNA Sequencing of Lake Charr Epidermal Mucus to Assess Molecular Effects of Diluted Bitumen Exposure in a Boreal Lake. Front. Environ. Sci. 10:836640. doi: 10.3389/fenvs.2022.836640
Received: 15 December 2021; Accepted: 17 January 2022;
Published: 04 February 2022.
Edited by:
Nsikak U. Benson, Covenant University, NigeriaCopyright © 2022 Andrzejczyk, Palace, Hrenchuk, Peters and Schlenk. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Nicolette E. Andrzejczyk, bmFuZHIwMDlAdWNyLmVkdQ==, bmlra2kuYW5kcnplamN6eWtAbmF2eS5taWw=
 Nicolette E. Andrzejczyk
Nicolette E. Andrzejczyk Vince P. Palace
Vince P. Palace Lee Hrenchuk2
Lee Hrenchuk2