- 1Department of Environment and Planning, Centre for Environmental and Marine Studies (CESAM), University of Aveiro, Aveiro, Portugal
- 2Crop and Environment Sciences Department, Harper Adams University, Newport, United Kingdom
- 3Marine, Environment and Technology Centre (MARETEC), LARSyS, Instituto Superior Técnico, University of Lisbon, Lisbon, Portugal
Escherichia coli is a facultative anaerobic bacterium mainly found in the lower intestines of mammals. It is transferred to soils via animal feces and can splash onto plant surfaces during rainfall or be taken up through roots into the plant tissue. Subsequently, it can pose a risk to farm workers handling the soil or the vegetation, or groundwater quality through leaching. Many inoculation studies show that E. coli can survive in soils for weeks to months. The few field studies that have been published suggest E. coli can survive for longer in cold and wet soils. We analyzed 20 topsoil samples from a relatively warm and aerobic Mediterranean pasture soil where grazing, and therefore manure inputs, had been excluded for 24 months, using the shotgun DNA sequencing method. We found that E. coli had not only survived for 2 years, but it had also comparatively thrived, as evidenced by its relative abundance of 17.0% in the control treatment and 20.2% in the biochar-amended treatment (no significant difference). Within-treatment variation was low for the control soil (coefficient of variation of 0.15) and approximately double that for the biochar-amended soil (0.28). There were no correlations between the relative abundance of E. coli and soil organic matter content, available P, soil texture, pH, or other soil nutrients. Our results suggest the existence of a naturalized E. coli soil population, with potentially important implications for farm management, as well as for environmental and public health under the One Health umbrella.
1 Introduction
Escherichia coli is a facultative anaerobic bacterium. It is a mesophilic species with optimum growth from 37°C to 42°C, with growth greatly impeded at lower temperatures and declining rapidly at higher temperatures. E. coli has been intensively studied as it serves as a model organism for understanding fundamental biological processes, including gene expression, replication, and metabolism. It is also widely used in medicine for producing proteins and enzymes, as well as containing several pathogenic strains.
E. coli in soils has mostly been investigated in survival studies under laboratory conditions, with soil samples from the field inoculated with specific strains of the bacterium. In these studies, E. coli abundance, commonly measured in CFUs, tends to decrease rapidly after inoculation before levelling off until the bacterium is found to be below detection limits. Reported survival periods range from weeks to months to 336 days, with longer survival times found for soils that were wetter and for generic rather than pathogenic strains (Murphy et al., 2024).
Traditionally, E. coli is considered an indicator of recent fecal contamination. Survival rates of E. coli in soils have been found to decline with high temperatures and aerobic conditions and low soil clay and dissolved organic carbon (DOC) contents (van Elsas et al., 2011). Microbial community structure and soil management practices were also found to have complex influences on E. coli survival rates. Nonetheless, evidence has suggested that certain strains of E. coli can persist in soil environments, particularly in relatively wet and cold pasture soils of Ireland (Brennan et al., 2010a) and the Alps (Texier et al., 2008). Studies have shown that E. coli can survive for extended periods of time in soils amended with manure, suggesting potential for establishment of naturalized populations. The timeframe of survival is an important consideration, given E. coli’s pathogenic potential, and that restrictions on manure and sewage sludge application to soil are time restricted when growing fresh produce (i.e., soft fruits and vegetables; Department for Environment, Food & Rural Affairs, 2018). Factors such as soil moisture, clay content, vegetation density, and rooting can contribute to enhanced persistence of E. coli in soils. Some hydrochars have been shown to adsorb E. coli in column experiments (Chung et al., 2017). However, the impact of biochar on E. coli survival in soil is variable, ranging from no effect with six different biochar types (Alegbeleye and Sant’Ana, 2023) to having biocidal properties against the pathogenic strain E. coli O157:H7, with the efficacy of the biocidal properties being dependent on biochar production temperature and biochar age (Gurtler et al., 2020). In rainfed pastures, the presence of grazing livestock can contribute further to the introduction and maintenance of E coli populations in soil (Texier et al., 2008; Muirhead, 2023). Naturalization of such communities may affect farm soil productivity, as well as have broader off-farm implications for environmental quality and public health.
In their mini-review, van Elsas et al. (2011) questioned the assumption that E. coli shows natural declines in open environments and stated that any environmental reservoir of the bacterium can cause risks to human health: “The factors that determine the rate of survival of particular E. coli strains, and thus the risks (to human health), are not (yet) easily predictable, and our capability to understand the effects of the secondary habitat (especially soil and water resources) on E. coli behaviour will be paramount to our abilities to manage the organism from both environmental and public health perspectives.”
Therefore, to help reduce this knowledge gap, in this study, we aimed to quantify the relative abundance of E. coli in topsoil samples of a rainfed biodiverse pasture, collected 24 months after exclusion of grazing animals, and to investigate correlations between E. coli survival and soil physicochemical variables, including biochar amendment, in a Mediterranean pasture soil.
2 Methods and materials
2.1 Study site and auxiliary data
The study site was located in the rain shadow of the Serra da Estrela mountain range in Portugal, that is, in the foothills directly to the east of Covilhã on a 500 ha livestock production farm called Quinta da França, which is managed both commercially and as a pilot farm by Terraprima (https://www.terraprima.pt/en/). The study site has a hot-summer Mediterranean climate (Csa in the Köppen classification) with wet and mild winters (annual rainfall in the range 600–1,200 mm). Soils are formed from granitic parent material, resulting in relatively deep (70–80 cm) dystric Cambisols (IUSS, 2014) of a sandy loam texture, with a topsoil organic matter content of 2.1%–3.6%, and strongly acidic (pH in 1 M KCl: 3.8–4.3).
The field site (40°16′25.51″N, 7°25′22.32″W) covered an area of 0.25 ha and was located at 440 m.a.s.l. on a 2° slope facing south. It was part of a vineyard until about 1985, when it was converted into a permanent pasture with grazing predominantly by cows. Since 2002, it has been managed as a sown biodiverse pasture a.k.a. SBP (see Teixeira et al., 2015 for a detailed description). Antibiotics were used sparingly in the cattle; that is, on average, an estimated one cow per year (in a herd of ∼100 animals) is treated with antibiotics, prescribed by the veterinarian, which are administered for 3 days while keeping the cow in the stables, including for an extra week after administration.
In April 2022, the field site was completely fenced off, thereby excluding grazing animals. After approximately 6 months, we noticed some rabbit droppings in a few places, so we rabbit-proofed the fence with a finer mesh to a height of 0.4 m. Since then, we have not observed rabbit droppings again, with only mouse droppings found occasionally inside plots (estimated, but not recorded, to be <1% cover). In October 2022, the fenced area was disked to 20 cm depth, and 20 plots of 3 m × 3 m were installed in a randomized block design, 10 control plots and 10 biochar-amended plots where biochar was mixed with topsoil (3% v/v) in a cement mixer to ensure a homogeneous distribution before being placed back into each plot (see Jongen et al., 2025 for a full description of the installation procedure). Blocks were 7 m apart, and plots within blocks were at least 3 m apart. Each plot was instrumented with a microclimate probe (TOMST extreme) in October 2023, which recorded topsoil temperature and volumetric soil moisture content at −6 cm, both logged at 15-min intervals. During the wet part of the year—October to April—the topsoil temperature ranged from 2.5°C to 24.3°C. The median volumetric topsoil moisture content during this period was 37%. We did not record topsoil temperatures during the dry part of the year—May to September—but we did record temperatures in other plots at the same field site. The median topsoil temperature was 24°C, while the maximum was 42°C (the topsoil reached temperatures warmer than 35°C on 43 days). The median volumetric topsoil moisture content during the summer period was 16%.
2.2 Soil sampling
During the approximate peak of biomass development of the SBP, on 24 April 2024, that is, 24 months after grazing exclusion, we took six topsoil samples (0–20 cm) from each of the plots using an auger, taking two samples under each plant functional group; that is, grasses (patches with Lolium multiflorum, L. perenne, Vulpia bromoides, Agrostis sp.), legumes (patches with Trifolium subterraneum), and forbs (patches with Chamaemelum sp., Leontodon taraxacoides, Coleostephus myconis, and Bellardia trixago), and mixed them into one composite sample per plot (20 samples in total). In the biochar-amended plots, this corresponded to the biochar incorporation depth. Each composite sample was passed through a 2-mm sieve, homogenized again with a sterile spoon, and ∼50 g of the <2 mm subsample was transferred to a plastic zip lock bag, closed, and frozen at −25°C in a mobile field freezer. After each plot was sampled, the auger and sieve were cleaned with 70% ethanol before continuing to the next plot. We used a portable electricity-powered freezer to keep the samples at −25°C until arrival at the laboratory. Upon arrival, samples were frozen at −75°C. After 30 days, samples were sent for DNA extraction and shotgun metagenomic analysis to Macrogen Inc., South Korea, in a box with four icepacks.
2.3 Shotgun metagenomic sequencing
The E. coli relative abundance (RA) data came as part of a metagenomic shotgun sequencing. The sequencing aimed at quantifying the relative abundances of bacteria and Archaea present in the topsoil during the peak biomass development of SBP, and therefore, it does not inform on their potential pathogenicity or virulence. The sequencing library was prepared by DNA fragmentation and ligation using ligate adapters. Adapter-ligated fragments were amplified by PCR. Clusters were generated using bridge amplification, and sequencing was performed with Illumina synthesis (SBS) technology. After sequencing was completed, Illumina HiSeq/NovaSeq raw data were demultiplexed by index sequences, and paired-end FASTQ files were generated for each sample. Adapter sequences and data with an average phred quality score less than 20 were removed using Trimmomatic (v0.39) (Bolger et al., 2014) of the Kneaddata (v0.10.0) (Beghini et al., 2021) pipeline (option: SLIDINGWINDOW:4:20). The pre-processed data were analyzed using MetaPhlAn4 (v4.0.0) (Blanco-Miguez et al., 2023) for ∼1 million microorganisms composed of NCBI reference genomes and species-level genome bins (SGBs). The reads were mapped to the specific marker genes of the microbial species using Bowtie2 (Langmead and Salzberg, 2012), and the species abundances were calculated based on the average number of reads mapped to the marker genes. At this time, marker genes that mapped to less than 33% were removed as the species were considered absent.
2.4 Statistical analysis
A linear mixed-effects model (LME) with restricted maximum likelihood was used to analyze the response of E. coli relative abundance to treatments with blocks as a random factor, using the R packages lme4 and lmerTest. Estimated marginal means, along with their confidence intervals, were extracted from the LME model using the “modelbased” package and visualized as a violin plot using the “ggplot2” and “see” packages. Correlation graphs between E. coli RA, soil moisture, and temperature parameters were made using the “ggpubr” package.
3 Results
E. coli was present in composite topsoil samples from all 20 plots (Supplementary Figure S1). The estimated marginal means were 17.0% and 20.2% for control and biochar-amended treatments, respectively, without any significant difference between treatments (Figure 1a). No significant correlations were found between E. coli relative abundance (RA) and soil pH or soil organic matter contents before or after biochar application (Figures 1b–d) or metrics of soil temperature during the growing season (Supplementary Figure S1). For biochar-amended plots, E. coli RA significantly increased with increasing soil moisture content, whereas there was no significant trend for the control plots. Potential indicators of E. coli inputs from animal feces did not show any significant correlations with E. coli RA (Figures 1e,f).
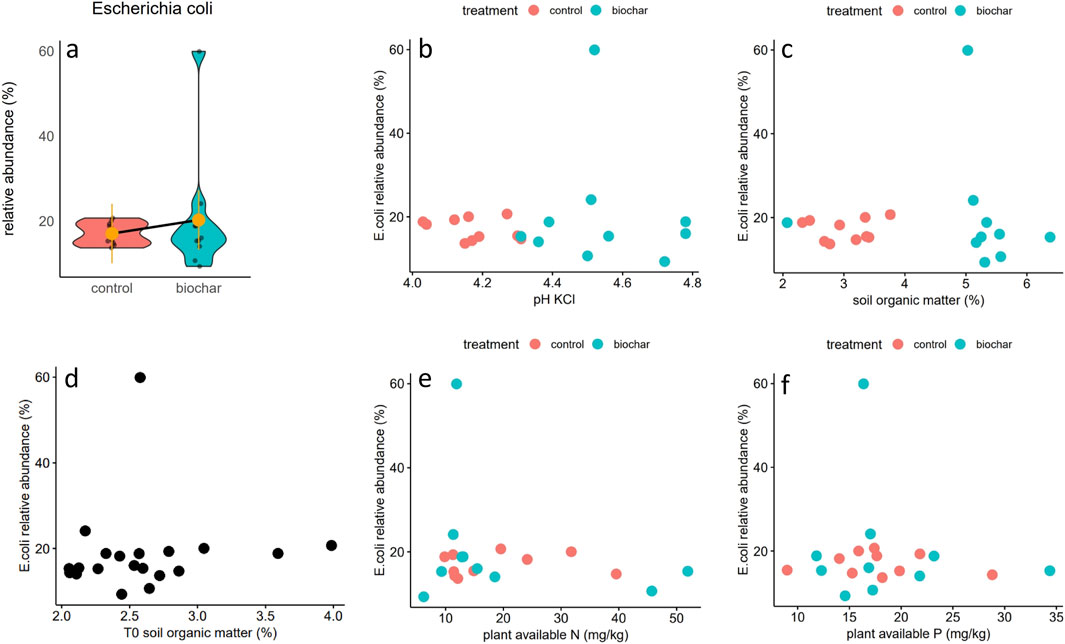
Figure 1. (a) Violin plot of E. coli RA in topsoil samples. Yellow dots represent the estimated marginal means of its relative abundance in each treatment, along with confidence intervals, extracted from the linear mixed effects model; (b) scatterplot between soil pH measured in KCl and E. coli RA; (c) scatterplot between soil organic matter content and E. coli RA; (d) scatterplot between soil organic matter content in the plots before biochar application and E. coli RA; (e) scatterplot between total available N in the soil and E. coli RA; (f) scatterplot between plant available P (represented as P2 from P2O5) and E. coli RA.
One of the samples was a statistical outlier (2.7 standard deviations removed from the mean in the biochar-amended treatment). Within-treatment variation was low for the control soil (coefficient of variation (CV) of 0.15) and approximately double that for the biochar-amended soil (0.28), after omission of the outlier.
4 Discussion and recommendations
The prevalence of E. coli (100%) in our pasture topsoil samples is higher than in previous work, which did not use a “without grazing” exclusion, that found forest and pasture land use to be the most prevalent for the presence of E. coli with 73% prevalence, compared to 10% prevalence in cropland and 15% in shrubland (Dusek et al., 2018). The analysis used in this study can only report on relative abundance rather than population growth per se. Nevertheless, the results demonstrate that the E. coli population was still maintained at significant levels, as a proportion of the total microbial community investigated in this study over timeframes previously thought to be untenable without further inoculation from inputs such as manures or sewage sludges, especially in such a warm environment (Tran et al., 2020). The higher prevalence in our study may be because the previous study focused on the landscape scale, whereas our study was in a relatively homogeneous, small area, that is, 0.25 ha. In addition, direct observations of runoff during an intense rainfall event, as well as observed soil surface evidence of erosion/deposition at the <1 m2 scale throughout the field site following rainfall, suggest that bacteria may have been distributed from cow pats to other areas of the field site before grazing exclusion, thereby contributing to the observed E. coli prevalence. This is also supported by the lack of observed significant correlations between indicators of animal feces inputs (available N and P) and E. coli RA. Texier et al. (2008) found >97% E. coli prevalence in Alpine grassland soils without grazing exclusion using the 16S rRNA method. Another grazing-exclusion field study, in Ireland, using lysimeters, found the prevalence of E. coli in leachate samples to vary from 3% to 48%, with the higher value being for a poor-draining Stagnic Luvisol (Brennan et al., 2010b). One possible reason may be that their use of the CFU method underestimated E. coli presence when it was in a viable but non-culturable (VBNC) state, to which our DNA sequencing method is not sensitive.
One of the biochar-amended treatment plots was a clear statistical extreme outlier for which we could not find any explanation in our auxiliary data. It is also unlikely to be caused by a point source, for example, wild animal excrement, because of the sampling design of six subsamples across each 9-m2 plot. Potentially, it was caused by an auxiliary variable that we did not measure, or by a DNA analysis error. Apart from the extreme outlier, the remaining 19 plots showed relatively consistent E. coli RA, that is, 9.3%–24.1%, with no significant difference between the control and biochar-amended treatments. Other biochar amendment studies have found increases, decreases, and no effect in E. coli survival depending on combinations of biochar and soil properties (Ye et al., 2018; Ahmadimoghadam et al., 2023; Alegbeleye and Sant’Ana, 2023). Although the biochar treatment in our study had similar E. coli RA after 24 months of grazing/manure exclusion as the control treatment, it did have double the range and almost double the within-treatment variation (after omitting the outlier). However, increases in range and within-treatment variation are not uncommon for studies where the treatment consists of incorporating a foreign material into the soil (Olmo et al., 2016), and a CV of 0.28 for a field study is still quite low.
Our findings do not conform with the reported environmental causes linked to E. coli decline, that is, high temperatures, aerobic conditions, low soil clay contents, and low DOC contents (van Elsas et al., 2011). If anything, two factors indicate reduced survivability. Specifically, clay content, which was 18% in the soil studied here, can be considered relatively low in this context, because soils with <18% clay are classified as “coarse” (Feeney et al., 2024). Aerobic conditions can be considered relatively high because the soil is not compacted (exclusion of grazing animals), and the good drainage conditions cause a quick decrease in SMC during dry periods. Nevertheless, we still found evidence indicating that in this relatively aerobic soil, there was a trend for E. coli to increase with increasing soil moisture content during the growing season. We only found this trend for biochar-amended plots, which may be in part because of the twice-as-large range in E. coli RA compared to the control treatment. More field data, covering a wider range of soil moisture contents, are required to explore whether this trend is specific to biochar-amended soil or also occurs in control soil. Although we do not have data on DOC, the relatively low SOM content (before biochar amendment) would likely mean that DOC was also likely low, with this factor also likely to decrease E. coli abundance in the soil over time.
Due to the restricted inputs through exclusion of grazing by livestock and rabbits, the observed 17%–20% relative abundance of E. coli after 24 months is suggestive of not only survival, but naturalization of E. coli. The field site has been under the same management since its conversion from a vineyard around 40 years before this study. Antibiotic use in the grazing animals is so sparse that the potential of an antibiotic-resistant strain of E. coli entering the soil that way is extremely unlikely. Various bacterial adaptation mechanisms have been suggested recently to potentially favor E. coli, for example, stress resistance in nutrient-poor environments (Somorin et al., 2016), such as the soil environment included in this study, and other microbiological DNA or transcriptional mechanisms (Sazykin et al., 2024; Nakamoto et al., 2025), which are beyond the scope of this brief research report.
While our results demonstrate that E. coli can persist and even thrive relative to other microorganisms in rainfed pasture soils long after the removal of grazing animals, we acknowledge that our study did not include strain-level identification or functional gene profiling. As such, we cannot determine whether the E. coli populations observed harbor virulence factors or antibiotic resistance genes, which are critical for assessing their potential risks to public and environmental health. As such, any discussion of environmental and public health implications is, therefore, speculative at this point. Nevertheless, the long-term persistence of E. coli in soils raises important questions about its ecological adaptation and the possibility of these populations becoming naturalized components of the soil microbiome.
While the specific impacts are not fully understood, E coli naturalization in these pastoral ecosystems can have both on-farm and off-farm implications (Jones, 1999; Muirhead, 2023), which should be taken into consideration under a One Health approach. At the farm level, this could reflect in altered microbial and nutrient dynamics, thus impacting soil productivity and other soil-based ecosystem services (Cheong et al., 2024). Environmental health implications include potential soil/water contamination (Jones, 1999), splashing during rainfall events, as well as surface runoff into downstream freshwater systems and leaching into groundwater reservoirs (Muirhead, 2023). In addition to directly affecting human communities that rely on these resources, other direct human exposure routes are linked to occupational health and safety for individuals working in or near contaminated soil or water via ingestion, inhalation, or dermal contact, thus potentially leading to gastrointestinal illness if pathogenic strains are present. Concerns in relation to food safety may also arise upon E. coli entering the food chain, particularly in integrated crop-livestock systems (Jones, 1999; Muirhead, 2023; Cheong et al., 2024).
Specific recommendations for research and management involve filling current knowledge gaps in elucidating the potential long-term consequences to soil ecosystems, livestock, and public health, as well as the underlying mechanisms for E. coli survival and human/animal exposure. It is also necessary to further understand the ecological role of naturalized E. coli populations in soil, their interactions with native microbes, and the factors influencing their persistence and pathogenicity. Specifically, we recommend that future studies incorporate metagenomic, whole-genome sequencing, or functional gene assays to explore the phylogenetic diversity, pathogenic potential, and resistance profiles of soil-resident E. coli strains. Such approaches would significantly strengthen risk assessments and guide appropriate monitoring and management strategies.
The findings of this study support the tentative conclusions from two previous field studies that E. coli may become naturalized in the soil (Brennan et al. , 2010a; Texier et al., 2008). This is contrary to a wealth of laboratory and greenhouse studies that report weeks to months of E. coli survival in soil (van Elsas et al., 2011; Murphy et al., 2024). Considering the implications for OneHealth, it would seem prudent to further investigate E. coli presence and survival in field soils. Specifically, the hypothesis that E. coli survives for longer, or has an increased chance of establishing itself in the soil microbial community, in soils that are year-round predominantly wet, is questioned by the results from this study, in which the soils are dry for extended periods of time. Therefore, we recommend that E. coli, including generic and pathogenic strains, be monitored in soils of other Mediterranean field sites, under any land use, that receive organic fertilizers where E. coli may be present (manures, slurries, and low-grade treated sewage), and pastures with and without the presence of grazers up to 2 years after the final amendment with those fertilizers. In addition, monitoring E. coli in the same samples with multiple measurement techniques, for example, culturing and various types of differential abundance methods (Nearing et al., 2022), may provide further insights into how these quantifications relate to each other.
5 Conclusion
We conclude that E. coli not only survived, but apparently thrived, either growing or maintaining its prevalence relative to the abundance of other microorganisms characterized in this study, in both the control and biochar-amended pasture soil, 24 months after grazing/manure input exclusion. This has implications for monitoring and management requirements as well as policy implications, whereby currently the exclusion period for growing fresh produce globally varies between 90 days and 365 days following manure application. Future research incorporating genomic or functional profiling is key for determining whether soil-resident E. coli represent a benign, naturalized community or a latent risk to human and animal health.
Data availability statement
The authors acknowledge that the data presented in this study must be deposited and made publicly available in an acceptable repository, prior to publication. Frontiers cannot accept a manuscript that does not adhere to our open data policies.
Author contributions
FV: Formal Analysis, Writing – original draft, Methodology, Writing – review and editing, Conceptualization, Validation. AJ: Writing – original draft, Methodology, Data curation, Conceptualization, Investigation, Visualization, Validation, Writing – review and editing, Formal Analysis. SJ: Writing – review and editing, Writing – original draft, Supervision. TD: Writing – review and editing, Writing – original draft. ZK: Data curation, Writing – original draft, Writing – review and editing. AB: Resources, Writing – original draft, Funding acquisition, Conceptualization, Methodology, Writing – review and editing, Supervision.
Funding
The author(s) declare that financial support was received for the research and/or publication of this article. This work is funded by national funds through FCT–Fundação para a Ciência e a Tecnologia I.P., under the project/grant UID/50006 + LA/P/0094/2020 (doi.org/10.54499/LA/P/0094/2020). We acknowledge the support of projects POLLINATE (https://doi.org/10.54499/PTDC/EAM-AMB/1509/2021) and HABITAT project 15341 (COMPETE2030- FEDER-00901600) through COMPETE 2030, FEDER, and FCT. We also acknowledge funding of authors FV (https://doi.org/10.54499/2023.06689.CEECIND/CP2840/CT0011) and A.C. Bastos through OE (https://doi.org/10.54499/DL57/2016/CP1482/CT0006). We also wish to thank the European Commission Joint Program SOIL for the funding of the project TRUESOIL (https://doi.org/10.54499/EJPSoils/0001/2021). La Caixa Foundation 100010434 is acknowledged for funding A. Jelinčić through the PhD fellowship Inphinit Incoming 2022 (PASTURE+, LCF/BQ/DI22/11940011).
Acknowledgments
We wish to acknowledge Oscar Gonzalez-Pelayo and Victoria Arcenegui-Baldo for assistance with sampling.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Correction note
A correction has been made to this article. Details can be found at: 10.3389/fenvs.2025.1708183.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fenvs.2025.1638383/full#supplementary-material
References
Ahmadimoghadam, Z., Tabatabaei, S. H., and Nourmahnad, N. (2023). Hydrophobic biochar enhances Escherichia coli retention in a sandy loam soil influenced by irrigation management. Int. J. Environ. Sci. Technol. 20 (8), 8399–8410. doi:10.1007/s13762-023-05002-8
Alegbeleye, O., and Sant’Ana, A. S. (2023). Survival behavior of six enterotoxigenic Escherichia coli strains in soil and biochar-amended soils. Environ. Res. 223, 115443. doi:10.1016/j.envres.2023.115443
Beghini, F., McIver, L. J., Blanco-Míguez, A., Dubois, L., Asnicar, F., Maharjan, S., et al. (2021). Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. Elife 10, e65088. doi:10.7554/elife.65088
Blanco-Miguez, A., Beghini, F., Cumbo, F., McIver, L. J., Thompson, K. N., Zolfo, M., et al. (2023). Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. 41, 1633–1644. doi:10.1038/s41587-023-01688-w
Bolger, A. M., Marc, L., and Bjoern, U. (2014). Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120. doi:10.1093/bioinformatics/btu170
Brennan, F. P., Abram, F., Chinalia, F. A., Richards, K. G., and O’Flaherty, V. (2010a). Characterization of environmentally persistent Escherichia coli isolates leached from an Irish soil. Applied and environmental microbiology, 76(7), 2175–2180.
Brennan, F. P., O’Flaherty, V., Kramers, G., Grant, J., and Richards, K. G. (2010b). Long-term persistence and leaching of Escherichia coli in temperate maritime soils. Applied and environmental microbiology, 76(5), 1449–1455.
Cheong, S., Jay-Russell, M. T., Chandler-Khayd, C., Di, F. J., Haghani, V., Aminanadi, P., et al. (2024). Presence of foodborne pathogens and survival of generic Escherichia coli in an organic integrated crop-livestock system. Front. Sustain. Food Syst. 8. doi:10.3389/fsufs.2024.1343101
Chung, J. W., Edewi, O. C., Foppen, J. W., Gerner, G., Krebs, R., and Lens, P. N. L. (2017). Removal of Escherichia coli by intermittent operation of saturated sand columns supplemented with hydrochar derived from sewage sludge. Appl. Sci. 7 (8), 839. doi:10.3390/app7080839
Department for Environment, Food & Rural Affairs. (2018). Annual report and accounts 2017–2018 (HC 1263). UK Government. Available online at: https://www.gov.uk/government/publications/defras-annual-report-and-accounts-2017-to-2018.
Dusek, N., Hewitt, A. J., Schmidt, K. N., and Bergholz, P. W. (2018). Landscape-scale factors affecting the prevalence of Escherichia coli in surface soil include land cover type, edge interactions, and soil pH. Applied and environmental microbiology, 84 (10), e02714–17.
Feeney, C. J., Bentley, L., De Rosa, D., Panagos, P., Emmett, B. A., Thomas, A., et al. (2024). Benchmarking soil organic carbon (SOC) concentration provides more robust soil health assessment than the SOC/clay ratio at European scale. Sci. Total Environ. 951, 175642. doi:10.1016/j.scitotenv.2024.175642
Gurtler, J. B., Mullen, C. A., Boateng, A. A., Mašek, O., and Camp, M. J. (2020). Biocidal activity of fast pyrolysis biochar against Escherichia coli O157: H7 in soil varies based on production temperature or age of biochar. J. Food Prot. 83 (6), 1020–1029. doi:10.4315/0362-028x.jfp-19-331
International Union of Soil Sciences (IUSS) (2014). World reference base for soil resources 2014: international soil classification system for naming soils and creating legends for soil maps. world soil resources reports no. 106. Rome: FAO.
Jones, D. L. (1999). Potential health risks associated with the persistence of Escherichia coli O157 in agricultural environments. Soil Use Manag. 15 (2), 76–83. doi:10.1111/j.1475-2743.1999.tb00069.x
Jongen, M., Bastos, A. C., Jelinčić, A., Simões, L., Gonzalez Pelayo, O., Niemeyer, J., et al. (2025). Biochar, plant productivity and community structure: a two-year study in a newly sown legume-rich pasture in Portugal under ambient and reduced precipitation. Available online at: https://ssrn.com/abstract=5215014. (Accessed August, 2025).
Langmead, B., and Salzberg, S. L. (2012). Fast gapped-read alignment with bowtie 2. Nat. Methods 9 (4), 357–359. doi:10.1038/nmeth.1923
Muirhead, R. W. (2023). Escherichia coli runoff from sheep and dairy cow grazed pasture: a plot scale simulation. J. Environ. Qual. 52 (2), 287–295. doi:10.1002/jeq2.20448
Murphy, C. M., Weller, D. L., Bardsley, C. A., Ingram, D. T., Chen, Y., Oryang, D., et al. (2024). Survival of twelve pathogenic and generic Escherichia coli strains in agricultural soils as influenced by strain, soil type, irrigation regimen, and soil amendment. J. food Prot. 87 (10), 100343. doi:10.1016/j.jfp.2024.100343
Nakamoto, S., Kobayashi, I., Watanabe, K., Kikuta, T., Imamura, S., and Shimada, T. (2025). Identification of a comprehensive set of transcriptional regulators involved in the long-term survivability of Escherichia coli in soil. Sci. Rep. 15 (1), 4279. doi:10.1038/s41598-025-85609-8
Nearing, J. T., Douglas, G. M., Hayes, M. G., MacDonald, J., Desai, D. K., Allward, N., et al. (2022). Microbiome differential abundance methods produce different results across 38 datasets. Nat. Commun. 13 (1), 342. doi:10.1038/s41467-022-28034-z
Olmo, M., Lozano, A. M., Barrón, V., and Villar, R. (2016). Spatial heterogeneity of soil biochar content affects soil quality and wheat growth and yield. Sci. Total Environ. 562, 690–700. doi:10.1016/j.scitotenv.2016.04.089
Sazykin, I., Naumova, E., Azhogina, T., Klimova, M., Karchava, S., Khmelevtsova, L., et al. (2024). Glyphosate effect on biofilms formation, mutagenesis and stress response of E. сoli. J. Hazard. Mater. 461, 132574. doi:10.1016/j.jhazmat.2023.132574
Somorin, Y., Abram, F., Brennan, F., and O’Byrne, C. (2016). The general stress response is conserved in long-term soil-persistent strains of Escherichia coli. Applied and Environmental Microbiology, 82 (15), 4628–4640.
Texier, S., Prigent-Combaret, C., Gourdon, M. H., Poirier, M. A., Faivre, P., Dorioz, J. M., et al. (2008). Persistence of culturable Escherichia coli fecal contaminants in dairy alpine grassland soils. Journal of Environmental Quality, 37(6), 2299–2310.
Teixeira, R. F., Proenca, V., Crespo, D., Valada, T., and Domingos, T. (2015). A conceptual framework for the analysis of engineered biodiverse pastures. Ecol. Eng. 77, 85–97. doi:10.1016/j.ecoleng.2015.01.002
Tran, D. T., Bradbury, M. I., Van Ogtrop, F. F., Bozkurt, H., Jones, B. J., and McConchie, R. (2020). Environmental drivers for persistence of Escherichia coli and salmonella in manure-amended soils: a meta-analysis. J. food Prot. 83 (7), 1268–1277. doi:10.4315/0362-028x.jfp-19-460
Van Elsas, J. D., Semenov, A. V., Costa, R., and Trevors, J. T. (2011). Survival of Escherichia coli in the environment: fundamental and public health aspects. ISME J. 5 (2), 173–183. doi:10.1038/ismej.2010.80
Ye, M., Sun, M., Zhao, Y., Jiao, W., Xia, B., Liu, M., et al. (2018). Targeted inactivation of antibiotic-resistant Escherichia coli and Pseudomonas aeruginosa in a soil-lettuce system by combined polyvalent bacteriophage and biochar treatment. Environ. Pollut. 241, 978–987. doi:10.1016/j.envpol.2018.04.070
Keywords: soil, biochar, E. coli, grassland, shotgun DNA
Citation: Verheijen F, Jelinčić A, Jeffery S, Domingos T, Khodaparast Z and Bastos AC (2025) Escherichia coli thrives in soil 24 months after grazing exclusion in a rainfed Mediterranean biodiverse pasture. Front. Environ. Sci. 13:1638383. doi: 10.3389/fenvs.2025.1638383
Received: 30 May 2025; Accepted: 24 July 2025;
Published: 29 August 2025; Corrected: 26 September 2025.
Edited by:
Lyudmyla Symochko, University of Coimbra, PortugalReviewed by:
Xiaomeng Chen, Northeast Agricultural University, ChinaMitiku Mihiret Seyoum, University of Arkansas, United States
Copyright © 2025 Verheijen, Jelinčić, Jeffery, Domingos, Khodaparast and Bastos. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Antun Jelinčić, YW50dW5qZWxpbmNpY0BnbWFpbC5jb20=
 Frank Verheijen
Frank Verheijen Antun Jelinčić
Antun Jelinčić Simon Jeffery2
Simon Jeffery2 Zahra Khodaparast
Zahra Khodaparast