- Department of Soil Biology, Institute of Soil Science and Land Evaluation, University of Hohenheim, Stuttgart, Germany
Microbial assimilation and stabilization of soil organic carbon (SOC) is an important process in global carbon cycling. For an improved understanding of climate-induced changes in ecosystem C dynamics, it is important to know the group-specific turnover of microbial C. Consequently, we wanted to answer the questions if fungi store newly assimilated C longer than bacteria and if climatic and edaphic properties of different regions affect microbial C assimilation and its subsequent release. This study presents results from a 112-day long field experiment where endogenous microorganisms from two agricultural soils were labeled with 13C labeled glucose to follow the dynamics of newly assimilated C. Both soils were representative for the respective study regions (Kraichgau and Swabian Alb). Whereas, microbial assimilation of newly added C was higher in Kraichgau than in Swabian Alb, the opposite result was obtained for the mean residence time (MRT) of microbial biomass C (76 days in Kraichgau and 93 days in Swabian Alb). The accelerated turnover rates of microbial C in the warmer soil of Kraichgau with lower clay content might be an important mechanism explaining the differences in SOC content between both regions. Gram-positive bacteria assimilated more 13C-glucose into their biomass than fungi and the MRT of C was higher in bacteria as compared to fungi in both regions. Beside these dynamic substrate utilization strategies, we observed possible cross feeding by gram-negative bacteria. Carbon MRT in fungi was region specific and was best represented by a two-pool model; the initial MRT ranged between 5 and 39 days and was, in the end, higher than 4 years. This hints toward a functional separation of the fungal community; fast growing fungi dominant in the early phase and internal redistribution of C in the second phase of decomposition. Our study identified microbial group and region specific MRT of freshly assimilated C as an important parameter, which might help to explain the commonly found variation in soil respiration and SOC stabilization.
Introduction
Microbial assimilation of soil organic carbon and its subsequent turnover are important processes in global carbon cycling (Bardgett et al., 2008; Xu et al., 2014), since they determine the magnitude of microbial biomass in soils and control processes leading to soil carbon stabilization. Mechanisms and controls in microbial carbon assimilation and turnover play also a fundamental role in regulating land-atmosphere interactions (Bardgett et al., 2008; Trivedi et al., 2013). Although microbial-derived C may play an important role in SOC stabilization (Sokol et al., 2019), the factors regulating the release of C from the microbial biomass are still poorly understood (Throckmorton et al., 2012; Wieder et al., 2013).
Microbial biomass turnover represents a significant source of carbon in soils since a major amount of C input into soils is recycled through microbial metabolism with microbial biomass-derived C contributing significantly to formation of recalcitrant soil organic carbon (Kindler et al., 2006, 2009; Miltner et al., 2009). It is widely acknowledged that fungi are more important for SOC stabilization than soil bacteria, which is related to (1) a more efficient C utilization by fungi, and (2) a greater decay-resistance of fungal biomass than bacterial biomass (Jastrow et al., 2007; Throckmorton et al., 2012). Conceptual models that describe soil C turnover dynamics often consider only pool sizes and their average turnover rates (Schweigert et al., 2015); however, turnover times of individual microbial groups representing different pathways of in situ SOC decomposition and stabilization might improve our conceptual understanding of SOC dynamics.
Differential turnover rates of microbial groups in soil systems makes it possible to associate bacteria and fungi to fast and slow energy channels in soil food webs (Moore et al., 2005; Rousk and Bååth, 2007). Bacterial biomass turns over rapidly with turnover times reported between 2.3–33.7 and 2.1–13.1 days as compared to fungi with turnover times between 130 and 150 days (Bååth, 1998; Rousk and Bååth, 2007). Similar differentiation with respect to persistence of bacterial and fungal biomass C has been shown elsewhere; low bacterial biomass C half-lives (6.9 days & 1.3 years) compared to fungi (11.3 days & 3.8 years) (Kindler et al., 2009; Schweigert et al., 2015). However, contradictory group-specific biomass C turnover rates have also been found, which adds uncertainty to the fate of soil C processed by microbial decomposers and their role in C stabilization. For instance, equal or lower mean residence time of fungal biomass C than bacteria have been reported (Balasooriya et al., 2014; Chemidlin Prévost-Bouré et al., 2014; Müller et al., 2017). Nevertheless, studies investigating group-specific microbial turnover rates under in situ conditions are scarce while turnover times of microbial decomposers investigated under lab conditions may vary significantly from in situ conditions since growth rates of microbial decomposers under natural environments could be very slow (Chapman and Gray, 1986).
Turnover times of microbial biomass carbon may also differ considerably between regions and variations in climatic and soil properties including soil temperature, soil water capacity, pH, soil texture, soil carbon content, and microbial community composition are important in this context (Cheng, 2009; Strickland and Rousk, 2010; Blagodatskaya et al., 2011; Rousk and Bengtson, 2014; Wang et al., 2018). Regions with distinct edaphic properties have been shown to differ in turnover rates of soil bacteria and fungi and even microbial necromass has shown different turnover times in regions varying in e.g., soil clay content (Throckmorton et al., 2012; Chemidlin Prévost-Bouré et al., 2014). Clay content controls microbial biomass C turnover with high clay content being associated with slow carbon turnover (Gregorich et al., 1991). In contrast to the regional effects on the turnover of microbial C, land-use did not change incorporation of added substrates into microbial biomass (Dungait et al., 2011), adding uncertainty to the fate of carbon assimilated by soil microorganisms.
The aim of the present study was, therefore, to understand the turnover of newly assimilated microbial C under in situ conditions in agricultural soils of two regions that differed in climatic and edaphic properties. We hypothesized that soil fungi would retain newly assimilated carbon longer than bacteria due to slower turnover rates. We expected also that region-specific soil properties and environmental factors will control mean residence times of fungal and bacterial carbon. We used in situ labeling of bacteria and fungi with 13C labeled glucose to follow the dynamic of newly assimilated C in a 112-day long field experiment. Microbial carbon assimilation and subsequent turnover rates were assessed by measuring 13C in microbial biomass and in bacterial and fungal phospholipid fatty acids.
Materials and Methods
Experimental Sites and Soil Sampling
The present study was conducted in two geographically distinct regions in southwest Germany that also differed in climatic and edaphic conditions. The Swabian Alb is characterized by humid and cold climate, whereas the Kraichgau is characterized by a drier and warmer climate compared to the Swabian Alb. The selected agricultural soils are representative for each region, respectively, based on the soil development expected for the dominant substrates and mean climatic conditions. Characteristic soil physiochemical and biological properties of both soils are given in Table 1. Bulk soil samples (10–15 cm) from an agricultural field in each region were collected, visible roots were removed and soils were sieved to <2 mm mesh size.
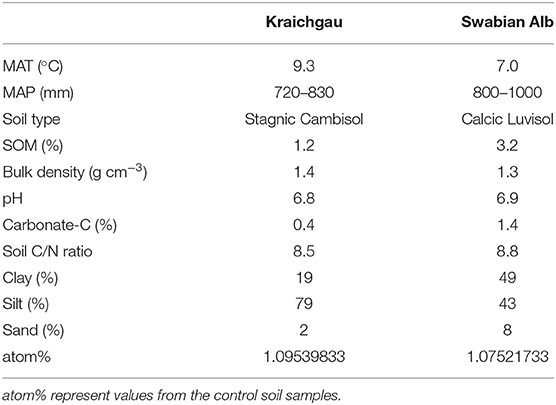
Table 1. Basic soil properties (up to 30 cm soil depth) of Kraichgau and Swabian Alb region. MAT and MAP represent mean annual temperature and precipitation, respectively.
Setup of the Field Experiment
Soils were pre-incubated at 15°C for 1 week for microbial acclimation and to avoid sieving disturbance effects. To label indigenous soil microorganisms, 13C-glucose (99 atom% 13C, Sigma-Aldrich, USA) and nitrogen (ammonium nitrate) were added in solution form corresponding to 10% of initial C and N in microbial biomass. This level of C and N addition was tested in pre-experiments for both soils individually to avoid excessive microbial growth. Later on, thirty PVC cores (Ø 10 cm, H 1 cm) were filled with 80 g (dry weight) soil per core from each region at specific bulk density (see Table 1). Since both soils differed in microbial biomass amount, we added 2.90 mg 13C-glucose per core for Kraichgau and 5.53 mg 13C-glucose for Swabian Alb. Soil-specific solution amounts were added dropwise to the cores using a pipet to bring cores up to 60% water holding capacity. Soil cores were covered from top and bottom with a micromesh PA-material (500 μm mesh size) to avoid macro fauna intrusion but allowing water and microbial exchange. Later on, soils were incubated at room temperature (22°C) for > 26 h. Both the relative amount of glucose addition and time of incubation were tested to avoid excessive microbial growth and to ensure complete microbial uptake of the added glucose, respectively. After initial incubation in the lab, undisturbed soil cores now containing labeled indigenous microorganisms were buried in the top 10–15 cm soil depth in the field. Six cores were sampled from each of the two agricultural fields (one per region) after 1, 7, 27, 55, and 112 days' exposure to in situ conditions, respectively. 1-day sampling corresponds to the soils frozen directly after >26 h initial incubation in the lab (see above). Soil samples from each core were stored at −24°C until further analysis. Soil water content was measured gravimetrically by drying soils at 60°C for 3 days.
Lab Analyses
Microbial Biomass Carbon (13Cmic)
The chloroform-fumigation-extraction method was used to determine microbial biomass carbon with 5 g (fresh weight) soil extracted with 20 ml of 0.025 M K2SO4 solution. See Marhan et al. (2010) for method-specific steps. For isotopic measurement of microbial biomass, the methodology as adapted by Müller et al. (2017) was followed. We used the following equation to calculate 13C in microbial biomass:
where “Cnf” and “Cf” corresponds to the extracted organic C contents of the non-fumigated and the fumigated samples (μg C g−1) and “AT%nf” and “AT%f” are their respective atom% 13C values. Calculation of the substrate-derived C (%) was done using the following equation:
where “AT% Sample” are the respective atom% values of the samples, “AT% glucose” the atom% value of the glucose (99 atom%), and “AT% FS” the atom% value of the SOC. Subsequently, the total amount of substrate-derived carbon in the microbial biomass was calculated (mg 13C g−1 added substrate).
Microbial 13C Phospholipid Fatty Acid (PLFA)
Microbial PLFAs were extracted from 2 × 6 g (fresh weight) soil samples and further transformed into fatty acid methyl esters (FAMEs) by alkaline methanolysis (Frostegård et al., 1991). For methodological details see Kramer et al. (2013). The two lab replicates of each sample were combined into one column before fractionation. For determination of atom% 13C, FAMEs were further fractionated with Ag+-SPE cartridges (6 ml, Supelco, Palo Alto, USA). The first and the fourth fractions, containing saturated and dienoic FAMEs, respectively, were retained in the cartridges and 13C values of the FAMEs were determined as described by Müller et al. (2016). A methyl correction was done for all the FAMEs using a mass balance equation (Denef et al., 2007). PLFAs including a15:0, i15:0, i16:0, and i17:0 represented gram-positive bacteria and cy17:0 and cy19:0 represented gram-negative bacteria. Fungal biomass was represented by PLFA 18:2ω6,9. Substrate-derived C (%) was calculated as described for microbial biomass C (Equation 2). Calculation of mean 13C incorporation into different bacterial groups was done considering the relative proportion of the respective fatty acids to the total amount of group-specific PLFAs as described by Preusser et al. (2017).
Mean Residence Time (MRT) and Statistical Analysis
Effects of region and sampling date (days after exposure in the field) on total microbial biomass and individual microbial groups were analyzed using analysis of variance (ANOVA). The same procedure was applied to the absolute 13C values in individual microbial groups. Homogeneity of variance was tested by Lavene's test and Tukey's HSD test was used for pairwise comparisons. In order to determine how microbial communities differed after substrate addition and over the whole exposure time between Kraichgau and Swabian Alb, principal component analysis was performed using data of substrate-derived 13C into microbial biomass as well as of 13C assimilated into fungal and bacterial PLFAs (R package stats).
A non-linear model using generalized least square (gnls function of the nlme package) was applied to the substrate-derived 13C values in microbial biomass and in individual microbial groups to derive mean residence times of microbial biomass C. A single exponential decay model was fitted to calculate the C turnover in microbial biomass C and in gram-positive bacteria in both Kraichgau and Swabian Alb. For improving model fit, samples after 7 days of incubation were removed from the model fit to the microbial biomass C data because soil processing during preparation of the field incubation most probably induced an increase in microbial biomass and a concomitant decrease in the 13C label. A double exponential decay model was superior to a single exponential decay model for fitting fungal PLFA data in Kraichgau and Swabian Alb (model superiority was tested with model AIC value). MRT of 13C in different microbial pools (in days) were estimated from the inverse of the decay constants whose significance was tested with ANOVA. All statistical analyses were performed in R program for statistical computing - version 3.5.1 (R Core Team, 2018).
Results
Microbial Biomass and Community Composition
Microbial biomass C generally increased during exposure of soil cores in the field [F(1, 56) = 49.98, P < 0.001] and the two soils differed from each other with the soil from Swabian Alb having a significantly higher microbial biomass than the one from Kraichgau [F(1, 56) = 136.91, P < 0.001, Figure 1]. Like microbial biomass C, absolute amounts of fungal and bacterial (gram-positive and gram-negative bacteria) PLFAs were higher in Swabian Alb than Kraichgau over the whole exposure time (Table S1). The two soils differed from each other along both PC1 [F(1, 56) = 38.75, P < 0.001] and PC2 [F(1, 56) = 67.53, P < 0.001] and also differentiated from each other in substrate utilization (Figure 2). The first two principal components (PCs) together explained 80% of the observed variance in microbial community composition.
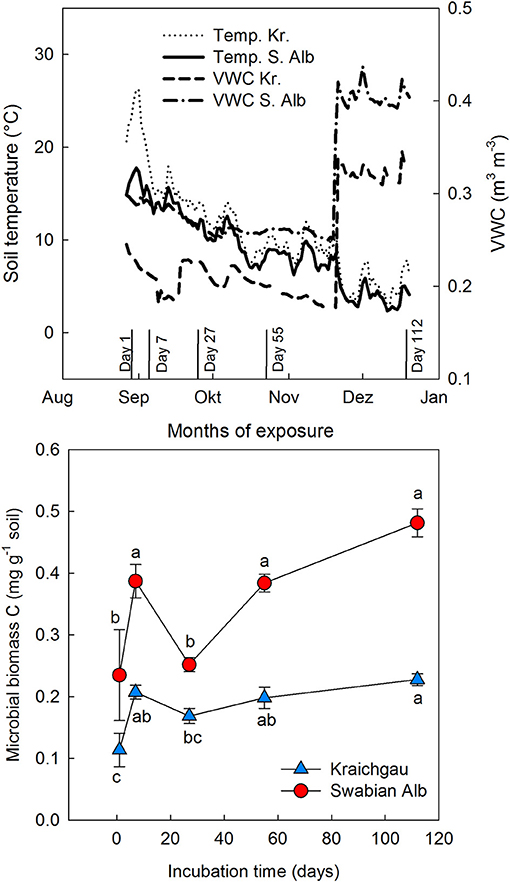
Figure 1. Dynamics of soil temperature, soil moisture (VWC), and microbial biomass C during exposure of soil cores in the field for Kraichgau (Kr.) and Swabian Alb (S. Alb). For the lower panel, bars represent standard deviation (n = 6) with different letter showing significant differences between sampling dates within each region (P < 0.05).
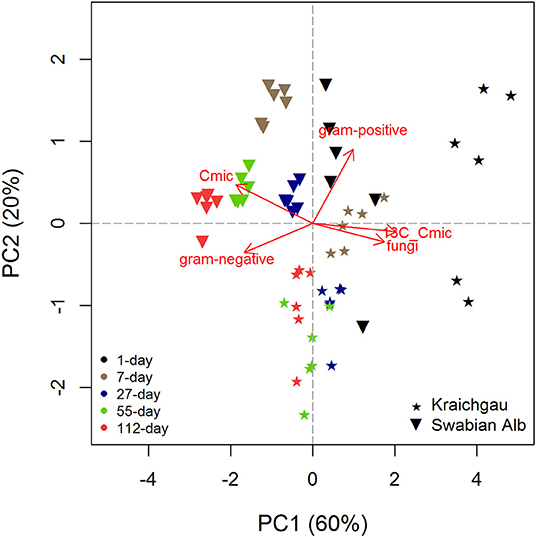
Figure 2. PCA analysis for showing shifts in microbial community composition from Kraichgau and Swabian Alb over the exposure time of 112 days. Gram-positive and gram-negative represent the two bacterial groups, Cmic is the absolute amount of microbial biomass C, and 13C-Cmic is the substrate-derived 13C in microbial biomass.
Incorporation of 13C Into Microbial Biomass
After initial incubation in the lab (26 h after substrate addition), microbial biomass in the soil of Kraichgau assimilated 86% more of the added 13C-glucose than in the soil of Swabian Alb (Figure 3A). A sharp decrease in 13C was observed in both regions after 7 days, which continued at a slower rate until the end of the experiment. The total decrease in assimilated 13C was similar for both soils with 65 and 66% decrease in Kraichgau and Swabian Alb, respectively (Figure 3A). The single exponential decay model resulted in 76 days MRT for 13C in microbial biomass in Kraichgau, which was lower than the MRT of 93 days in Swabian Alb (Table 2).
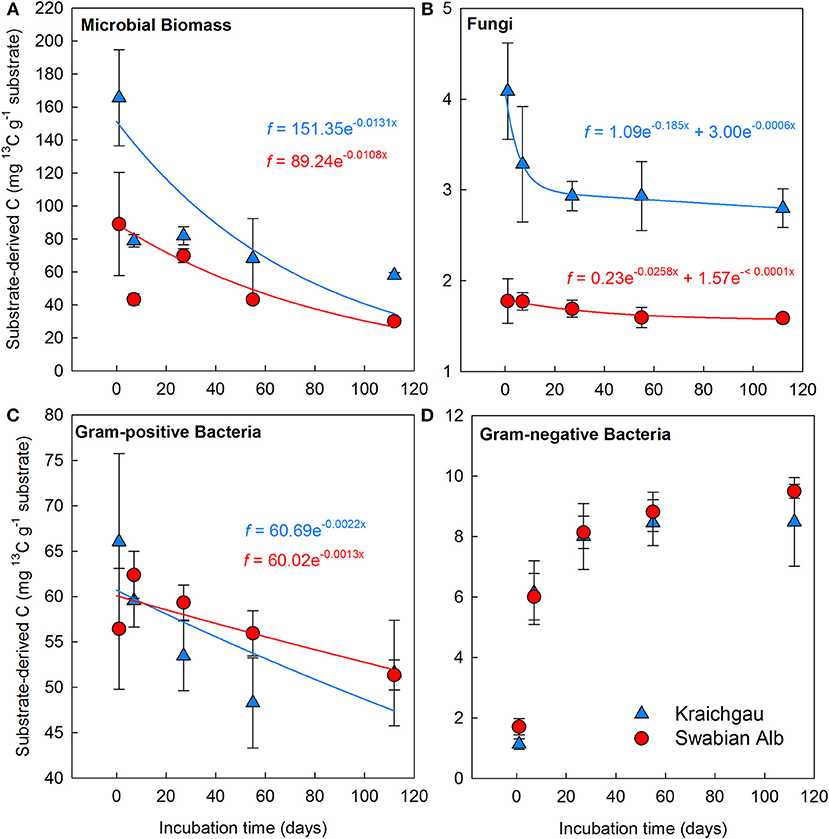
Figure 3. Substrate-derived 13C assimilated into microbial biomass (A), and into the PLFAs of soil fungi (B), gram-positive bacteria (C), and gram-negative bacteria (D) from Kraichgau and Swabian Alb. Lines represent fitted exponential decay functions. Bars represent standard deviation (n = 6).
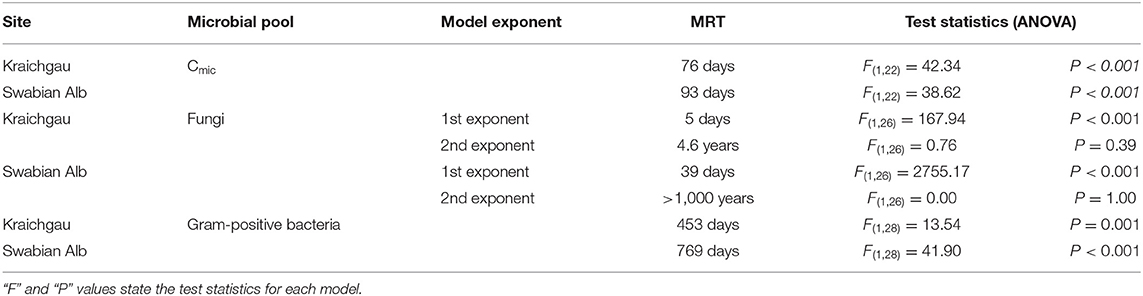
Table 2. Mean residence time (MRT) of microbially assimilated 13C, calculated by fitting single or double exponential decay functions to microbial biomass and PLFA data in Kraichgau and Swabian Alb.
Incorporation of 13C in Microbial PLFAs
Even though the amount of fungal PLFA was lower in Kraichgau (Table S1), the proportion of assimilated 13C was larger than in Swabian Alb; substrate-derived 13C in fungal PLFAs in the soil of Kraichgau was initially ~130% higher than in Swabian Alb [F(1, 56) = 235.13, P < 0.001, Figure 3B]. In Kraichgau, a sharp decrease in assimilated 13C was observed after 7 days of exposure whereas only a minor loss was observed between days 27 and 112. In contrast, no such sudden changes in assimilated 13C in fungal PLFAs were observed in Swabian Alb (Figure 3B). Over the period of 112 days, a decrease of 32% was observed in assimilated 13C in fungal PLFA in Kraichgau as compared to a decrease of only 11% in Swabian Alb. These differences in the temporal patterns of the assimilated 13C resulted in much faster turnover times of the fungal biomass C in Kraichgau than in Swabian Alb (Table 2). The double exponential decay model provided MRT for the fast pool of 5 days for Kraichgau and of 39 days for Swabian Alb. For the slow pool MRT of 4.6 years for Kraichgau and of > 1,000 years for Swabian Alb were fitted (Table 2).
As for fungal PLFAs, 13C assimilation by gram-positive bacteria differed between both soils (Figure 3C). At day 1, 13C incorporation into gram-positive bacterial PLFAs was 1.2 times higher in Kraichgau than in Swabian Alb [F(1, 56) = 24.79, P < 0.001]. With increasing exposure time, assimilated 13C generally decreased in gram-positive bacterial PLFAs in Kraichgau (Figure 3C). On the contrary, in Swabian Alb, amounts of assimilated 13C in gram-positive bacterial PLFAs increased after 7-days exposure and subsequently decreased constantly toward the end of field exposure. Over the whole exposure time, amounts of assimilated 13C decreased up to 22% in Kraichgau and up to 9% in Swabian Alb (Figure 3C). Regional trends in turnover times of gram-positive bacterial biomass C were similar to what was observed for fungal biomass C; MRT calculated from the single exponential decay model was lower in Kraichgau (453 days) than in Swabian Alb (769 days; Table 2).
In contrast to gram-positive bacteria and fungi, initial enrichment of 13C was lower in gram-negative bacterial PLFAs (Figure 3D and Figure S2) and 13C assimilation in gram-negative bacterial PLFAs increased continuously [F(1, 56) = 51.83, P < 0.001] with similar trends between Kraichgau and Swabian Alb (Figure 3D and Figure S2). Trends in loss / gain of assimilated 13C in individual bacterial PLFAs and their turnover times are shown in Figures S1, S2 and Table S2, respectively.
Discussion
The role of microbial decomposers in SOC decomposition, especially bacteria and fungi, has received considerable attention in carbon sequestration as a large proportion of SOC is of microbial origin (Banerjee et al., 2016). We studied the C turnover within soil microorganisms, specifically key players of SOC decomposition i.e., bacteria and fungi, in order to better understand soil C dynamics under in situ conditions. Performing field incubations in two regions differing in climatic and edaphic conditions allowed investigating the role of soil biotic and abiotic properties in controlling the fate of carbon in microbial biomass.
13C Incorporation Into Key Players of Carbon Decomposition and Its Turnover
The first process defining the role of microbial groups for C sequestration is the uptake and assimilation of C from the soil. In our study, bacteria as compared to fungi incorporated higher amounts of added 13C-glucose into their biomass, observed as higher substrate-derived C after initial incubation, and this trend remained visible over the 112-day's exposure time (Figures 3B,C and Figure S2). Dungait et al. (2011) reported similar observation over highest 13C-glucose incorporation into gram-positive bacteria (30%) and lowest incorporation into fungi (<1%), even at a much shorter time-scale (10 days). Similarly, Brant et al. (2006) also reported higher glucose incorporation by bacteria (gram-positive and gram-negative) than fungi. However, care must be taken in not over interpreting these results as we investigated only one fungal PLFA in comparison to four PLFAs for gram-positive bacteria. Furthermore, even though incorporation of 13C-glucose into bacterial and microeukaryote populations has been shown earlier through rRNA stable isotope probing (Kramer et al., 2016), a non-uniform labeling of soil microbial biomass through the glucose must be kept in mind while interpreting our results (Papp et al., 2020). Nevertheless, our results support the established idea of bacteria as dominant consumers of highly labile C (Castro et al., 2010; Vries and de Caruso, 2016), although it was recently shown that specific fungi quickly utilize labile rhizodeposits (Hünninghaus et al., 2019). Moreover, in the context of bacterial association with fast C cycling pathways as compared to fungal association with slow C cycling pathways (Moore et al., 2005), such higher bacterial incorporation of available labile C may have important implications for soil C dynamics, for instance, their contribution to non-living soil organic matter (Kindler et al., 2006). Within bacterial community, gram-positive bacteria initially incorporated and retained higher substrate-derived 13C into their PLFAs than gram-negative bacteria (Figures 3C,D and Figure S2). Müller et al. (2017) found opposite results, i.e., higher 13C-substrate incorporation in gram-negative bacterial PLFAs in the early stages of litter decomposition. This discrepancy in observed response of the bacterial community in our study might just be due to higher abundance (biomass) of gram-positive bacteria in both regions leading to higher added-substrate uptake and incorporation, and absolute abundances of bacterial PFLAs supported this argument (Table S1).
After soil microorganisms have assimilated C, its turnover within the soil microorganisms further controls the fate of the C as being stabilized within the microbial C pool or being released either as CO2 or stabilized SOC. Turnover of microbial biomass C assessed by loss in assimilated 13C over time was 76 and 93 days in Kraichgau and Swabian Alb, respectively (Table 2). These turnover times agree to the reported MRT of microbial biomass C in previous studies, for instance, 110 day's microbial biomass C turnover time reported by Cheng (2009) and 3.3 to 125 day's MRT in early stages of litter decomposition reported by Müller et al. (2017). The loss of assimilated 13C in microbial biomass was much higher than carbon loss in individual bacterial and fungal PLFAs (Figures 3A–C), which was also reflected in a reduced MRT (Table 2). This observation was also previously found in the literature (Malik et al., 2015; Müller et al., 2017) and might be explained by the fact that PLFAs represent only a specific proportion of the microbial biomass. Even though microbial fatty acids are degenerated very fast after cell death (Willers et al., 2015), intensive recycling of fast turning over components, such as PLFAs, may lead to their longer turnover times (Gunina et al., 2017). Furthermore, chemical or physical protection / stabilization of fatty acids or their residues might be one mechanism explaining these results (Lützow et al., 2006; Miltner et al., 2009).
However, although PLFAs represent only a specific proportion of the microbial biomass, group-specific turnover of PLFAs can be used to clarify the role of microbial groups for SOC stabilization. We observed a rather continuous decrease in assimilated 13C in gram-positive PLFAs (Figure 3C). Contrary to this observation, the 13C label increased continuously in gram-negative bacterial PLFAs in both soils (Figure 3D). These results may be explained by the cross-feeding behavior of soil bacteria and may hint toward possible translocation of carbon between different microbial groups (Balasooriya et al., 2014). However, such cross-feeding may be uni- or bi-directional (D'Souza et al., 2018), as evident from a small increase in gram-positive bacterial 13C incorporation in Swabian Alb after 7 days and in Kraichgau after 112 days (Figure 3C). Müller et al. (2017) suggested that the cross-feeding aspect must be taken into account while interpreting C turnover times.
Two aspects mainly affect carbon storage in soil bacterial and fungal biomass. First aspect is the difference in their physiology or cell resistance to decomposition; fungal cell walls are more resistant to decomposition than bacterial cell walls, therefore, fungal contribution to C sequestration and stabilization itself is important (Jastrow et al., 2007). The second aspect deals with the difference in fungal and bacterial life spans with fungi turning over more slowly than bacteria (Moore et al., 2005). We fitted a double exponential decay model (decision based on model AIC value) to fungal PLFA representing turnover of rapid and slow carbon pools. The MRT of carbon in fungal PLFA representing the rapid pool (first exponent) was much lower than MRT of carbon in gram-positive bacterial PFLAs. In contrast, the MRT of the more stable C pool (second exponent) in fungal PLFA was larger than bacterial carbon MRT (Table 2). This only partly confirms our hypothesis that carbon stored in fungi has longer residence time than bacteria. Our results are in accordance to other studies performed at microbial hotspots in either agricultural or grassland soils (Balasooriya et al., 2014; Müller et al., 2017), reporting shorter PLFA 13C residence time in fungi than bacteria. It is important to mention here that we calculated residence time of carbon assimilated in gram-positive bacterial PLFAs from a single exponential decay model, which may not be suitable since gram-positive bacterial PLFAs may also consist of slow and fast carbon pools. However, assimilated 13C increased in most individual gram-positive PLFAs after 7 days in Swabian Alb and after 112 days in Kraichgau (Figure 3C and Figure S1). Intensive recycling (see also above) of assimilated 13C within gram-positive bacteria may have led, therefore, to a more even carbon distribution in PLFAs over time (Rinnan and Bååth, 2009), making a single decay model more suitable. The short MRT of the fast fungal C pool may also be explained by differences in the fungal community; for instance, some fast-growing fungi like yeasts may have exploited the added substrate and released the assimilated C as soon as the substrate pool was depleted (Müller et al., 2016). Such an effect of fungal community composition may explain the observed sharp decrease in fungal assimilated 13C within 20 days of exposure in Kraichgau (Figure 3B, see discussion below).
Regional Differences in Soil Properties Control Turnover Times of Microbial Communities
Both soils in our study differed significantly in 13C-glucose incorporation into microbial biomass; incorporation in Kraichgau was twice as high as in Swabian Alb despite of higher microbial biomass in Swabian Alb (Figures 1, 3A), which was also reflected in substrate-derived carbon in bacterial and fungal PLFAs (Figures 3B,C). Low amounts of labile SOC in Kraichgau under equilibrium conditions (Ali et al., 2018) may explain the observed trends. We argue that microbial biomass in Kraichgau reacted fast upon labile substrate availability, possibly creating microbial hotspots (Kuzyakov and Blagodatskaya, 2015), and assimilated more substrate-derived carbon into their biomass. However, in Swabian Alb where SOC is higher than in Kraichgau, native labile carbon may have also been utilized for growth leading to lower observed 13C incorporation. Although soil types and soil properties were not replicated in our experimental design, both study sites were representative for the respective region. Our results, therefore, provided insights into mechanisms explaining regional differences in turnover of microbial biomass C.
The enhanced C incorporation into microbial biomass in the Kraichgau was also reflected in lower MRT of C in microbial biomass as well as in fungal and bacterial PLFAs in Kraichgau compared to Swabian Alb (Table 2). In addition to the above-mentioned differences in SOC contents and its implications for substrate preferences and internal C turnover, these differences in 13C MRT may also be explained by climatic differences between both regions. During exposure in the field, soil temperature in Kraichgau was on average 2°C higher than in Swabian Alb and the differences in temperature were highest, up to 7°C, during the first week of field exposure (Figure 1), leading to a faster decline in incorporated 13C due to temperature-sensitive microbial cell production and death (Castro et al., 2010; Hagerty et al., 2014). Furthermore, warmer soils of Kraichgau might have supported increased abundance of microbial predators and an accelerated shift in microbial community composition (Hagerty et al., 2014) than for the microbial community in the Swabian Alb (Figure 2). Long MRT of microbial biomass C as well as of the C incorporated into specific bacterial and fungal PLFAs in Swabian Alb might also be controlled by soil texture (Gregorich et al., 1991); soil clay content was about 2.5 times higher in Swabian Alb than in Kraichgau (see Table 1). Association of microbial biomass and microbial compounds (e.g., peptides and proteins) with negatively charged sites in soils, for instance clay (Six et al., 2006; Miltner et al., 2009), might, therefore, explain the relative stability of assimilated 13C in the Swabian Alb.
Conclusions
Our study provides important information on the turnover times of individual microbial groups representing different pathways of in situ SOC decomposition and stabilization, which might improve our conceptual understanding of SOC dynamics. We focused on answering the questions if fungi store newly assimilated C longer than bacteria and if regional differences in climatic and edaphic properties affect microbial C assimilation and its subsequent release. We observed dynamic substrate utilization strategies/strong cross-feeding behavior adapted by microbial groups reflected through depletion of assimilated 13C in gram-positive bacteria and fungi but its further use by gram-negative bacteria. Longer turnover times of fungal assimilated 13C for the second phase of C turnover emphasizes greater fungal role in carbon stabilization. The accelerated turnover rates of microbial C in the warmer soil of Kraichgau with lower clay content might be an important mechanism explaining the differences in SOC content between both regions. Our results also imply that regions with short MRT of freshly assimilated C in microbial biomass might produce high respiration from relatively higher SOC decomposition resulting in less SOC stabilization. However, if the observed differences in MRT of microbial C are related to differences in SOC, stabilization depends also on the fate of microbial C after its release from the microbial biomass (i.e., after becoming necromass).
Data Availability Statement
All datasets generated for this study are included in the article/Supplementary Material.
Author Contributions
CP and EK perceived the general concept of the study. RA conducted field experiment, analyzed soil samples, performed data analysis, and wrote the manuscript. All authors listed have carefully reviewed the manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Sabine Rudolf for supporting PLFAs analyses, Wolfgang Armbruster for GC-IRMS 13C analysis and Joachim Ingwersen for providing basic data on soil properties. We are thankful to the reviewers for their constructive comments, which helped to improve the quality of this manuscript. This study was funded by DFG (Deutsche Forschungsgemeinschaft) under the framework of Research Unit FOR1695 Agricultural landscapes und global climate change – Processes and feedbacks on a regional scale within subproject P9 microbial regulation of soil organic matter decomposition at the regional scale (KA 1590/10-1, KA 1590/ 10-2, and PO 617 1578/3-2).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fenvs.2020.00033/full#supplementary-material
References
Ali, R. S., Kandeler, E., Marhan, S., Demyan, M. S., Ingwersen, J., Mirzaeitalarposhti, R., et al. (2018). Controls on microbially regulated soil organic carbon decomposition at the regional scale. Soil Biol. Biochem. 118, 59–68. doi: 10.1016/j.soilbio.2017.12.007
Bååth, E. (1998). Growth rates of bacterial communities in soils at varying pH: a comparison of the thymidine and leucine incorporation techniques. Microbial Ecol. 36, 316–327. doi: 10.1007/s002489900118
Balasooriya, W. K., Denef, K., Huygens, D., and Boeckx, P. (2014). Translocation and turnover of rhizodeposit carbon within soil microbial communities of an extensive grassland ecosystem. Plant Soil 376, 61–73. doi: 10.1007/s11104-012-1343-z
Banerjee, S., Kirkby, C. A., Schmutter, D., Bissett, A., Kirkegaard, J. A., and Richardson, A. E. (2016). Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol. Biochem. 97, 188–198. doi: 10.1016/j.soilbio.2016.03.017
Bardgett, R. D., Freeman, C., and Ostle, N. J. (2008). Microbial contributions to climate change through carbon cycle feedbacks. ISME J. 2, 805–814. doi: 10.1038/ismej.2008.58
Blagodatskaya, E., Yuyukina, T., Blagodatsky, S., and Kuzyakov, Y. (2011). Turnover of soil organic matter and of microbial biomass under C3–C4 vegetation change. Consideration of 13C fractionation and preferential substrate utilization. Soil Biol. Biochem. 43, 159–166. doi: 10.1016/j.soilbio.2010.09.028
Brant, J. B., Sulzman, E. W., and Myrold, D. D. (2006). Microbial community utilization of added carbon substrates in response to long-term carbon input manipulation. Soil Biol. Biochem. 38, 2219–2232. doi: 10.1016/j.soilbio.2006.01.022
Castro, H. F., Classen, A. T., Austin, E. E., Norby, R. J., and Schadt, C. W. (2010). Soil microbial community responses to multiple experimental climate change drivers. Appl. Environ. Microbiol. 76, 999–1007. doi: 10.1128/AEM.02874-09
Chapman, S. J., and Gray, T. R. G. (1986). Importance of cryptic growth, yield factors and maintenance energy in models of microbial growth in soil. Soil Biol. Biochem. 18, 1–4. doi: 10.1016/0038-0717(86)90095-7
Chemidlin Prévost-Bouré, N., Dequiedt, S., Thioulouse, J., Lelièvre, M., Saby, N. P. A., Jolivet, C., et al. (2014). Similar processes but different environmental filters for soil bacterial and fungal community composition turnover on a broad spatial scale. PLoS ONE 9:e111667. doi: 10.1371/journal.pone.0111667
Cheng, W. (2009). Rhizosphere priming effect. Its functional relationships with microbial turnover, evapotranspiration, and C–N budgets. Soil Biol. Biochem. 41, 1795–1801. doi: 10.1016/j.soilbio.2008.04.018
Denef, K., Bubenheim, H., Lenhart, K., Vermeulen, J., van Cleemput, O., Boeckx, P., et al. (2007). Community shifts and carbon translocation within metabolically-active rhizosphere microorganisms in grasslands under elevated CO2. Biogeosciences 4, 769–779. doi: 10.5194/bg-4-769-2007
D'Souza, G., Shitut, S., Preussger, D., Yousif, G., Waschina, S., and Kost, C. (2018). Ecology and evolution of metabolic cross-feeding interactions in bacteria. Nat. Product Rep. 35, 455–488. doi: 10.1039/C8NP00009C
Dungait, J. A. J., Kemmitt, S. J., Michallon, L., Guo, S., Wen, Q., Brookes, P. C., et al. (2011). Variable responses of the soil microbial biomass to trace concentrations of 13C-labelled glucose, using 13C-PLFA analysis. Eur. J. Soil Sci. 62, 117–126. doi: 10.1111/j.1365-2389.2010.01321.x
Frostegård, Å., Tunlid, A., and Bååth, E. (1991). Microbial biomass measured as total lipid phosphate in soils of different organic content. J. Microbiol. Methods 14, 151–163. doi: 10.1016/0167-7012(91)90018-L
Gregorich, E. G., Voroney, R. P., and Kachanoski, R. G. (1991). Turnover of carbon through the microbial biomass in soils with different texture. Soil Biol. Biochem. 23, 799–805. doi: 10.1016/0038-0717(91)90152-A
Gunina, A., Dippold, M., Glaser, B., and Kuzyakov, Y. (2017). Turnover of microbial groups and cell components in soil: 13C analysis of cellular biomarkers. Biogeosciences 14, 271–283. doi: 10.5194/bg-14-271-2017
Hagerty, S. B., van Groenigen, K. J., Allison, S. D., Hungate, B. A., Schwartz, E., Koch, G. W., et al. (2014). Accelerated microbial turnover but constant growth efficiency with warming in soil. Nat. Clim. Change 4, 903–906. doi: 10.1038/nclimate2361
Hünninghaus, M., Dibbern, D., Kramer, S., Koller, R., Pausch, J., Schloter-Hai, B., et al. (2019). Disentangling carbon flow across microbial kingdoms in the rhizosphere of maize. Soil Biol. Biochem. 134, 122–130. doi: 10.1016/j.soilbio.2019.03.007
Jastrow, J. D., Amonette, J. E., and Bailey, V. L. (2007). Mechanisms controlling soil carbon turnover and their potential application for enhancing carbon sequestration. Clim. Change 80, 5–23. doi: 10.1007/s10584-006-9178-3
Kindler, R., Miltner, A., Richnow, H.-H., and Kästner, M. (2006). Fate of gram-negative bacterial biomass in soil–mineralization and contribution to SOM. Soil Biol. Biochem. 38, 2860–2870. doi: 10.1016/j.soilbio.2006.04.047
Kindler, R., Miltner, A., Thullner, M., Richnow, H.-H., and Kästner, M. (2009). Fate of bacterial biomass derived fatty acids in soil and their contribution to soil organic matter. Organic Geochem. 40, 29–37. doi: 10.1016/j.orggeochem.2008.09.005
Kramer, S., Dibbern, D., Moll, J., Huenninghaus, M., Koller, R., Krueger, D., et al. (2016). Resource partitioning between bacteria, fungi, and protists in the detritusphere of an agricultural soil. Front. Microbiol. 7:1524. doi: 10.3389/fmicb.2016.01524
Kramer, S., Marhan, S., Haslwimmer, H., Ruess, L., and Kandeler, E. (2013). Temporal variation in surface and subsoil abundance and function of the soil microbial community in an arable soil. Soil Biol. Biochem. 61, 76–85. doi: 10.1016/j.soilbio.2013.02.006
Kuzyakov, Y., and Blagodatskaya, E. (2015). Microbial hotspots and hot moments in soil. Concept & review. Soil Biol. Biochem. 83, 184–199. doi: 10.1016/j.soilbio.2015.01.025
Lützow, M., v., Kögel-Knabner, I., Ekschmitt, K., Matzner, E., Guggenberger, G., et al. (2006). Stabilization of organic matter in temperate soils. Mechanisms and their relevance under different soil conditions? A review. Eur. J. Soil Sci. 57, 426–445. doi: 10.1111/j.1365-2389.2006.00809.x
Malik, A. A., Dannert, H., Griffiths, R. I., Thomson, B. C., and Gleixner, G. (2015). Rhizosphere bacterial carbon turnover is higher in nucleic acids than membrane lipids. Implications for understanding soil carbon cycling. Front. Microbiol. 6:268. doi: 10.3389/fmicb.2015.00268
Marhan, S., Kandeler, E., Rein, S., Fangmeier, A., and Niklaus, P. A. (2010). Indirect effects of soil moisture reverse soil C sequestration responses of a spring wheat agroecosystem to elevated CO2. Glob. Change Biol. 16, 469–483. doi: 10.1111/j.1365-2486.2009.01949.x
Miltner, A., Kindler, R., Knicker, H., Richnow, H.-H., and Kästner, M. (2009). Fate of microbial biomass-derived amino acids in soil and their contribution to soil organic matter. Organic Geochem. 40, 978–985. doi: 10.1016/j.orggeochem.2009.06.008
Moore, J. C., McCann, K., and de Ruiter, P. C. (2005). Modeling trophic pathways, nutrient cycling, and dynamic stability in soils. Pedobiologia 49, 499–510. doi: 10.1016/j.pedobi.2005.05.008
Müller, K., Kramer, S., Haslwimmer, H., Marhan, S., Scheunemann, N., Butenschön, O., et al. (2016). Carbon transfer from maize roots and litter into bacteria and fungi depends on soil depth and time. Soil Biol. Biochem. 93, 79–89. doi: 10.1016/j.soilbio.2015.10.015
Müller, K., Marhan, S., Kandeler, E., and Poll, C. (2017). Carbon flow from litter through soil microorganisms. From incorporation rates to mean residence times in bacteria and fungi. Soil Biol. Biochem. 115, 187–196. doi: 10.1016/j.soilbio.2017.08.017
Papp, K., Hungate, B. A., and Schwartz, E. (2020). Glucose triggers strong taxon-specific responses in microbial growth and activity: insights from DNA and RNA qSIP. Ecology 101:e02887. doi: 10.1002/ecy.2887
Preusser, S., Marhan, S., Poll, C., and Kandeler, E. (2017). Microbial community response to changes in substrate availability and habitat conditions in a reciprocal subsoil transfer experiment. Soil Biol. Biochem. 105, 138–152. doi: 10.1016/j.soilbio.2016.11.021
R Core Team (2018). R A language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing.
Rinnan, R., and Bååth, E. (2009). Differential utilization of carbon substrates by bacteria and fungi in tundra soil. Appl. Environ. Microbiol. 75, 3611–3620. doi: 10.1128/AEM.02865-08
Rousk, J., and Bååth, E. (2007). Fungal and bacterial growth in soil with plant materials of different C/N ratios. FEMS Microbiol. Ecol. 62, 258–267. doi: 10.1111/j.1574-6941.2007.00398.x
Rousk, J., and Bengtson, P. (2014). Microbial regulation of global biogeochemical cycles. Front. Microbiol. 5:103. doi: 10.3389/fmicb.2014.00103
Schweigert, M., Herrmann, S., Miltner, A., Fester, T., and Kästner, M. (2015). Fate of ectomycorrhizal fungal biomass in a soil bioreactor system and its contribution to soil organic matter formation. Soil Biol. Biochem. 88, 120–127. doi: 10.1016/j.soilbio.2015.05.012
Six, J., Frey, S. D., Thiet, R. K., and Batten, K. M. (2006). Bacterial and fungal contributions to carbon sequestration in agroecosystems. Soil Sci. Soc. Am. 70, 555–569. doi: 10.2136/sssaj2004.0347
Sokol, N. W., Kuebbing, S. E., Karlsen-Ayala, E., and Bradford, M. A. (2019). Evidence for the primacy of living root inputs, not root or shoot litter, in forming soil organic carbon. N. Phytol. 221, 233–246. doi: 10.1111/nph.15361
Strickland, M. S., and Rousk, J. (2010). Considering fungal:bacterial dominance in soils – Methods, controls, and ecosystem implications. Soil Biol. Biochem. 42, 1385–1395. doi: 10.1016/j.soilbio.2010.05.007
Throckmorton, H. M., Bird, J. A., Dane, L., Firestone, M. K., and Horwath, W. R. (2012). The source of microbial C has little impact on soil organic matter stabilisation in forest ecosystems. Ecol. Lett. 15, 1257–1265. doi: 10.1111/j.1461-0248.2012.01848.x
Trivedi, P., Anderson, I. C., and Singh, B. K. (2013). Microbial modulators of soil carbon storage. Integrating genomic and metabolic knowledge for global prediction. Trends Microbiol. 21, 641–651. doi: 10.1016/j.tim.2013.09.005
Vries, F. T., and de Caruso, T. (2016). Eating from the same plate? Revisiting the role of labile carbon inputs in the soil food web. Soil Biol. Biochem. 102, 4–9. doi: 10.1016/j.soilbio.2016.06.023
Wang, J., Sun, J., Xia, J., He, N., Li, M., Niu, S., et al. (2018). Soil and vegetation carbon turnover times from tropical to boreal forests. Funct. Ecol. 32, 71–82. doi: 10.1111/1365-2435.12914
Wieder, W. R., Bonan, G. B., and Allison, S. D. (2013). Global soil carbon projections are improved by modelling microbial processes. Nat. Clim. Change 3, 909–912. doi: 10.1038/nclimate1951
Willers, C., van Jansen Rensburg, P. J., and Claassens, S. (2015). Phospholipid fatty acid profiling of microbial communities-a review of interpretations and recent applications. J. Appl. Microbiol. 119, 1207–1218. doi: 10.1111/jam.12902
Keywords: fungi, bacteria, 13C glucose, stable isotope-probing, turnover rate
Citation: Ali RS, Poll C and Kandeler E (2020) Soil Properties Control Microbial Carbon Assimilation and Its Mean Residence Time. Front. Environ. Sci. 8:33. doi: 10.3389/fenvs.2020.00033
Received: 05 November 2019; Accepted: 05 March 2020;
Published: 24 March 2020.
Edited by:
Maria Luz Cayuela, Spanish National Research Council, SpainReviewed by:
Egbert Schwartz, Northern Arizona University, United StatesRuben Lopez-Mondejar, Academy of Sciences of the Czech Republic, Czechia
Copyright © 2020 Ali, Poll and Kandeler. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Rana Shahbaz Ali, cmFuYUB1bmktaG9oZW5oZWltLmRl; c2hhaGJhenJhbmEucmFuYUBnbWFpbC5jb20=
 Rana Shahbaz Ali
Rana Shahbaz Ali Christian Poll
Christian Poll Ellen Kandeler
Ellen Kandeler