Abstract
Stick and leaf insects (Phasmatodea) are large, tropical, predominantly nocturnal herbivores, which exhibit extreme masquerade crypsis, whereby they morphologically and behaviorally resemble twigs, bark, lichen, moss, and leaves. Females employ a wide range of egg-laying techniques, largely corresponding to their ecological niche, including dropping or flicking eggs to the forest floor, gluing eggs to plant substrate, skewering eggs through leaves, ovipositing directly into the soil, or even producing a complex ootheca. Phasmids are the only insects with highly species-specific egg morphology across the entire order, with specific egg forms that correspond to oviposition technique. We investigate the temporal, biogeographic, and phylogenetic pattern of evolution of egg-laying strategies in Phasmatodea. Our results unequivocally demonstrate that the ancestral oviposition strategy for female stick and leaf insects is to remain in the foliage and drop or flick eggs to the ground, a strategy that maintains their masquerade. Other major key innovations in the evolution of Phasmatodea include the (1) hardening of the egg capsule in Euphasmatodea; (2) the repeated evolution of capitulate eggs (which induce ant-mediated dispersal, or myrmecochory); (3) adapting to a ground or bark dwelling microhabitat with a corresponding shift in adult and egg phenotype and egg deposition directly into the soil; and (4) adhesion of eggs in a clade of Necrosciinae that led to subsequent diversification in oviposition modes and egg types. We infer at minimum 16 independent origins of a burying/inserting eggs into soil/crevices oviposition strategy, 7 origins of gluing eggs to substrate, and a single origin each of skewering eggs through leaves and producing an ootheca. We additionally discuss the systematic implications of our phylogenetic results. Aschiphasmatinae is strongly supported as the earliest diverging extant lineage of Euphasmatodea. Phylliinae and Diapheromerinae are both relatively early diverging euphasmatodean taxa. We formally transfer Otocrania from Cladomorphinae to Diapheromerinae and recognize only two tribes within Diapheromerinae: Diapheromerini sensu nov. and Oreophoetini sensu nov. We formally recognize the clade comprising Necrosciinae and Lonchodinae as Lonchodidae stat. rev. sensu nov.
Introduction
Natural History
Stick and leaf insects (Phasmatodea) are among the most charismatic animals on earth. They are large, tropical, predominantly nocturnal herbivores, which exhibit extreme masquerade crypsis, whereby they phenotypically and behaviorally resemble twigs, bark, lichen, moss, and leaves (Bedford, 1978). Examples include the exquisite leaf-mimicking Phyllium, (Figure 1A), the robust, bark-like “tree lobsters” (e.g., Dryococelus australis, The Lord Howe tree lobster—likely the rarest documented insect on the planet, Figure 1C) (Priddel et al., 2003), and a large number of twig, stick, and branch mimics, including the recently described giant, Phobaeticus chani, the world's largest extant insect at just over two feet (56.7 cm) including legs (Hennemann and Conle, 2008). Despite being masters of masquerade among insects, phasmids are often chemically defended through a wide array of noxious chemicals secreted from specialized prothoracic glands when under threat (Eisner et al., 1997), and a few species have developed aposematic coloration (e.g., Achrioptera fallax, Oreophoetes peruana, Orthonecroscia pulcherrima). They are known for their ability to regenerate lost limbs (e.g., resulting from predation or complications during previous molts) during juvenile molts (Brusca et al., 2016). Many phasmids are facultatively parthenogenic, a feature likely tied to the poor dispersal ability of adults, particularly females, which are slow moving, often flightless, and when gravid, even winged species exhibit limited motility (Bradler and Buckley, 2018).
Figure 1
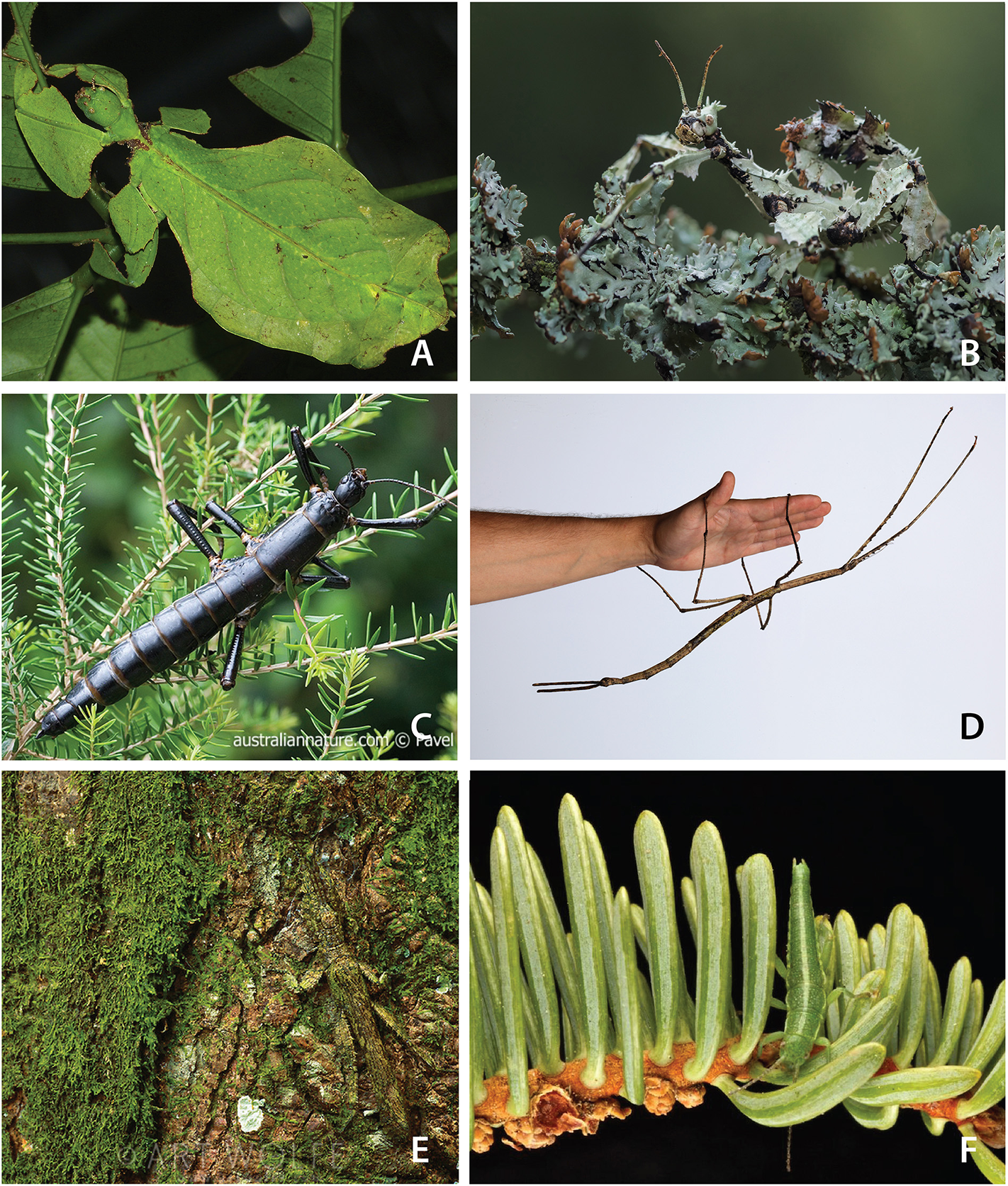

A glimpse of stick and leaf insect diversity and masquerade crypsis. (A) A leaf insect, Phyllium (photograph by Bernard Dupont, used by permission). (B)Extatasoma tiaratum (photograph by Miroslaw Wasinski, used by permission). (C) The Lord Howe stick insect, Dryococelus australis, the rarest known insect on the planet (photograph by Pavel German, used by permission). (D)Ctenomorpha gargantuan, measuring 56.5 cm in length (photograph by the Museum of Victoria, used by permission). (E)Prisopus sp. (photograph by Art Wolfe, used by permission). (F)Timema cristinae (photograph by Moritz Muschick, used by permission).
Female stick and leaf insects employ a wide range of egg-laying techniques, largely corresponding to their ecological niche (Carlberg, 1983; Sellick, 1997a,b) (Figure 2). Many species remain in the foliage during oviposition and passively drop or actively flick single eggs from their ovipositor to the ground. Others carefully fix their eggs to plant substrate with glue (Figure 2D) or even by skewering them through leaves (Figure 2E). Some species, including the robust tree lobsters, carefully place their eggs directly into crevices (e.g., bark) or the soil (Figure 2C). An anomaly for the order, a recently discovered species produces a complex egg case or ootheca that contains numerous eggs in a highly ordered fashion (Figure 2F) (Goldberg et al., 2015).
Figure 2
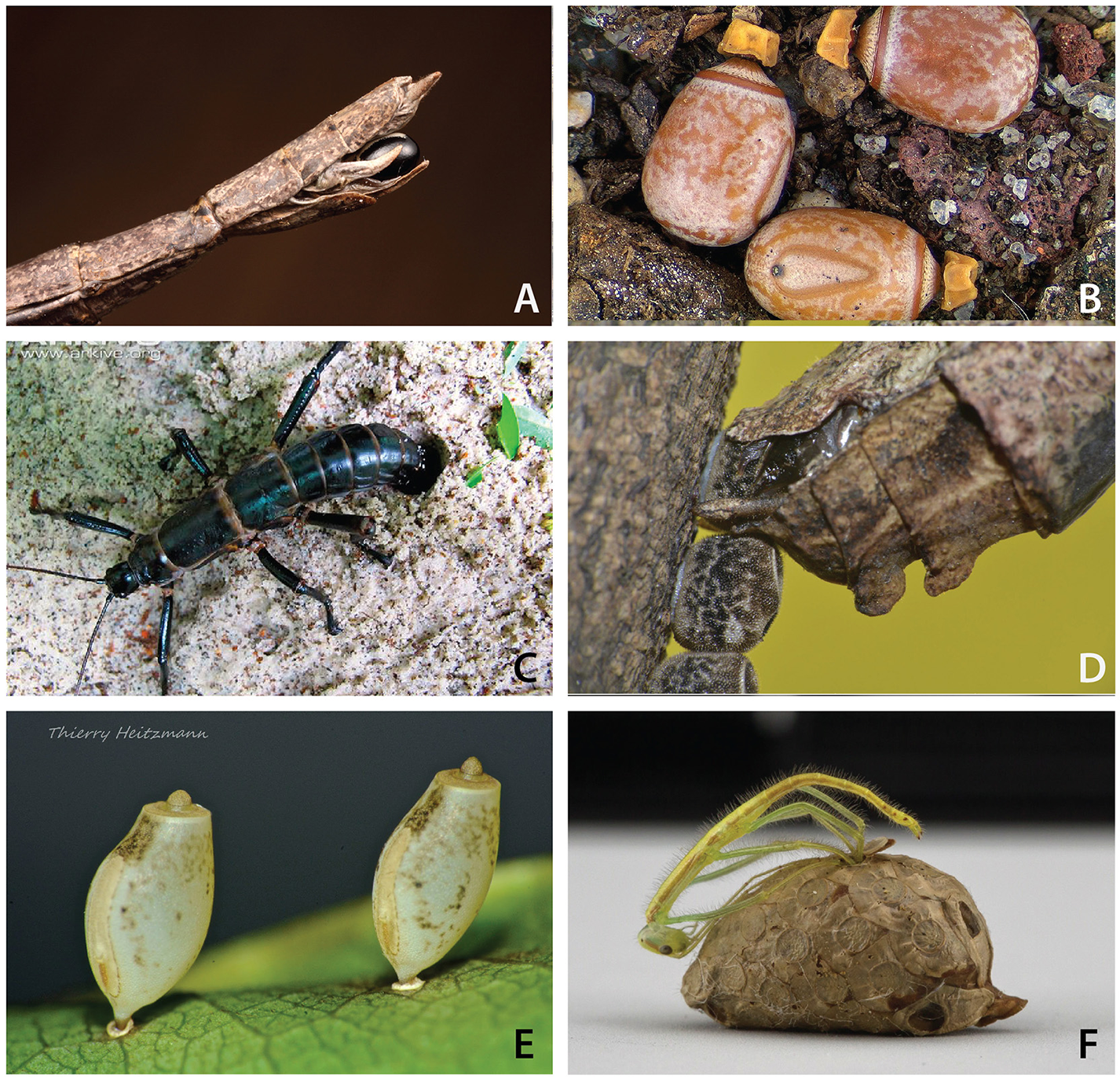

Phasmid oviposition techniques and eggs. (A) Ovipositor and mature egg of Diapheromera femorata. Photo © Alex Wild, used by permission. (B) Capitulate eggs of Eurycnema goliah. Photo by Piotr Naskrecki, used by permission. (C)Dryococelus australis depositing eggs in soil. Photo by Rohan Cleave, Zoos Victoria, Australia, used by permission. (D)Neoclides sp. gluing eggs to branch. Photo by Albert Kang, used by permission. (E)Asceles eggs skewered into leaf. Photo by Thierry Heitzmann, used by permission. (F) Korinninae nymph emerging from ootheca. Photo by Bruno Kneubühler, used by permission.
Whereas, the diversity of oviposition strategies within Phasmatodea is ecologically fascinating and likely evolutionarily significant, the diversity of form exhibited in the eggs themselves is simply extraordinary (Figure 3). Phasmids are the only insects with blatant species-specific egg morphology across the entire order, with specialized egg forms adapted to oviposition technique. For example, eggs that are directly buried into the soil (Figure 3E') are typically tapered at the posterior pole in a bullet-like fashion, modified for ease of burial (e.g., Orxines, Acacus, Centrophasma, and Diesbachia) or are slightly tapered and cylindrical in shape (e.g., Heteropteryx) (Sellick, 1997a). Eggs skewered to leaves are modified with a collard spine-like extension at the posterior pole that is used to pierce the leaf and hold up the egg (Figure 3S). A major guild of eggs exclusive to groups of species that drop or flick their eggs to the ground bear a capitulum—a lipid-rich, detachable, knob-like extension of the operculum (lid of the egg) that serves as the signal adaptation for inducing ant-mediated dispersal (Figures 3A–N) (Moore, 1993; Stanton et al., 2015). Capitulate eggs represent a key innovation for phasmid dispersal and occur in disparate lineages and biogeographic regions worldwide (Compton and Ware, 1991; Hughes and Westoby, 1992; Windsor et al., 1996). Eggs that are attached to plant substrate (usually leaves) with glue are more variable in form. Generally, adhesion is on the ventral surface of the egg with the eggs attached singly (e.g., Sceptrophasma, Sipyloidea). But in a few genera (e.g., Calvisia, Trachythorax) eggs are attached in groups, with eggs additionally glued to the anterior or posterior surface of adjacent eggs (Sellick, 1997a).
Figure 3
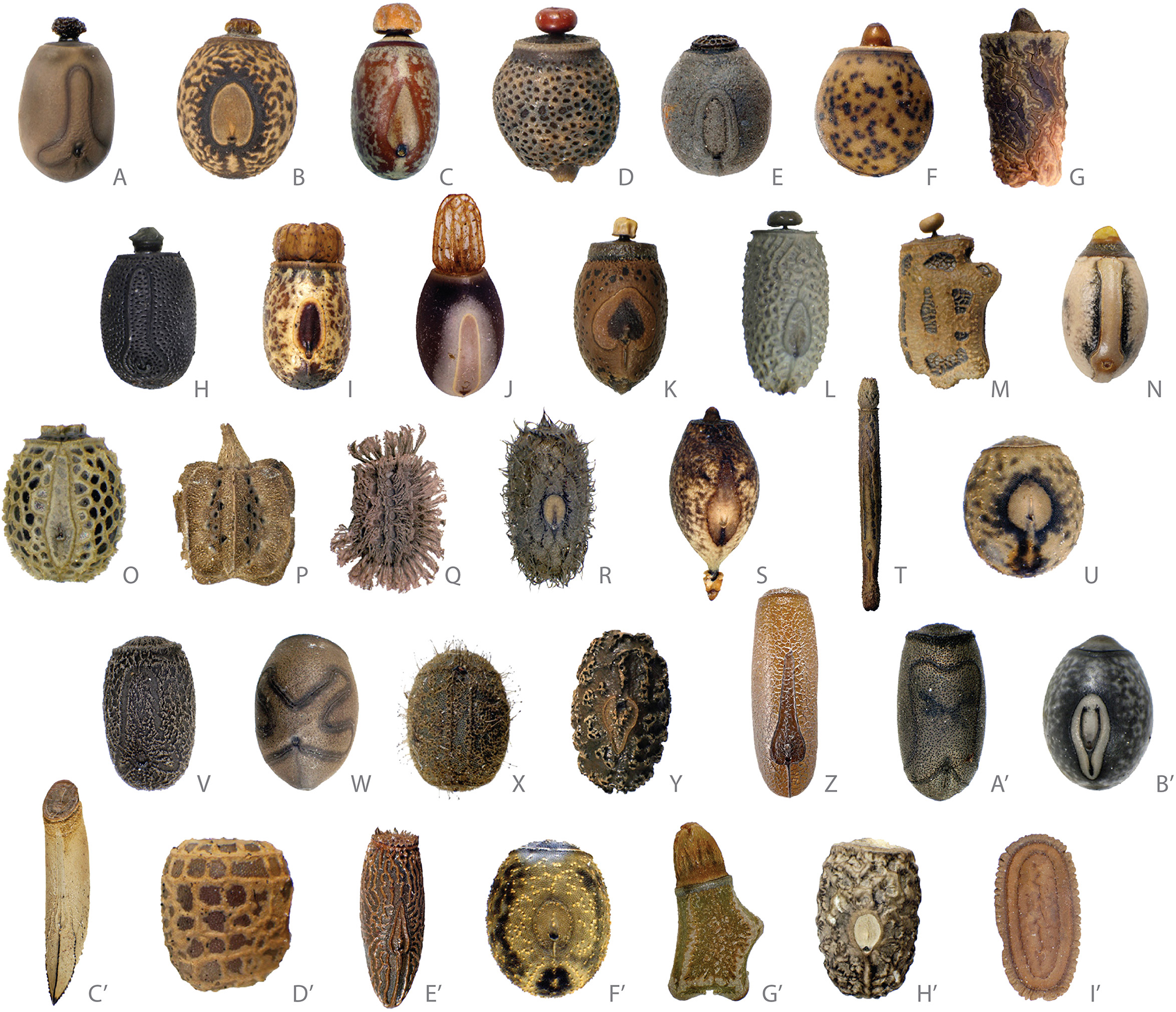

Stick and leaf insect eggs illustrating the breadth of their phenotypic diversity. Eggs depicted in the top two rows have capitula. Corresponding taxonomic names are provided in Supplementary Table S3. Photos by François Tetaert, used by permission.
Given the taxonomic occurrence of phasmid oviposition strategies, multiple origins of various techniques are likely throughout the order. Goldberg et al. (2015) provided an initial investigation of phasmid oviposition modes primarily to place the anomalous ootheca-producing species in an ecological and phylogenetic context. Their analysis strongly supported the recovery of this species among the ecologically- and species-rich Necrosciinae. However, their taxon sampling was not crafted to explore the evolution of egg-laying techniques across phasmid diversity. The evolutionary significance and the temporal, biogeographic, and phylogenetic pattern of evolution of egg-laying strategies in Phasmatodea remain unclear.
Systematics
Phasmatodea stands out as one of the few insect orders to lack a phylogenetically based classification (Bradler and Buckley, 2018). Günther (1953) classified Phasmatodea into a number of subfamilies and tribes, and this classification gained broad acceptance (Bradley and Galil, 1977; Kevan, 1977, 1982). However, subsequent phylogenetic analyses based on morphological (Tilgner, 2002; Bradler, 2009) and molecular data (Whiting et al., 2003; Buckley et al., 2009; Bradler et al., 2014, 2015; Büscher et al., 2018) clearly demonstrate that we lack a meaningful higher-level classification for phasmids. Due to the widespread phenotypic convergence associated with extreme masquerade crypsis and corresponding microhabitat (Buckley et al., 2009; Bradler et al., 2015), it is not surprising that morphology-based classification schemes have failed to reflect the evolutionary history of the group.
Molecular-based phylogenetic studies have largely been limited and focused on biogeographically-restricted taxa such as New Zealand stick insects (e.g., Trewick et al., 2008; Buckley et al., 2010), Mascarene phasmids (Bradler et al., 2015), Mediterranean taxa (e.g., Ghiselli et al., 2007; Scali et al., 2012, 2013), the genus Timema in Southern North America (Law and Crespi, 2002; Schwander et al., 2011), and the primarily Indo-Malayan subfamily Necrosciinae (Bradler et al., 2014; Goldberg et al., 2015). In the present study we use the most broadly sampled molecular data set to date to investigate the temporal, biogeographic, and phylogenetic pattern of evolution of egg-laying strategies in Phasmatodea and infer their evolutionary significance. We further discuss systematic implications and make formal emendations to phasmid classification in light of the current molecular phylogenetic consensus.
Methods
Taxon and Gene Sampling
A global sampling comprising 284 phasmid exemplars representing all major lineages and ca. 32% (150 of 473) of the generic diversity and a single representative of Embioptera (Metoligotoma) as an outgroup (see Misof et al., 2014) was used in the present study (Supplementary Table S1). We sampled regions of 7 genes to infer the evolutionary history of Phasmatodea: nuclear 18S rRNA (18S), 28S rRNA (28S) and histone subunit 3 (H3), and mitochondrial 12S rRNA (12S), 16S rRNA (16S), cytochrome-c oxidase subunit I (COI) and cytochrome-c oxidase subunit II (COII). Molecular data were acquired via traditional PCR and Sanger sequencing following methods outlined previously (Bradler et al., 2014; Robertson et al., 2015), or from previously published data available on GenBank (Buckley et al., 2009, 2010; Bradler et al., 2014, 2015; Goldberg et al., 2015). Sequences new to this study are deposited in GenBank under accession numbers MK291527-MK291924, MK296739-MK296749, and MK297240-MK297296.
Alignment and Data Partitions
The protein encoding genes H3 and COI were length invariant and thus alignment of these genes was trivial, based on conservation of amino acid (AA) reading frame. COII however contained a length variable region in the coding sequence. Using Mesquite 3.31 (Maddison and Maddison, 2017), COII was translated into AA sequence and aligned using MUSCLE (Edgar, 2004) as implemented in Mesquite. The COII nucleotide sequences were then aligned via Mesquite to match the aligned AA sequences. Ribosomal genes were aligned using an online implementation of MAFFT 7 (Katoh and Toh, 2008) http://mafft.cbrc.jp/alignment/server/ applying the G-INS-I strategy. Resulting alignments were visually inspected to check for ambiguously aligned regions and alignment artifacts. Multiple ambiguously aligned expansion regions were removed from the 28S alignment using GBlocks (Talavera and Castresana, 2007) with options for a “less stringent selection.”
We used PartitionFinder 2 (Lanfear et al., 2016) to simultaneously select the best-fit partitioning scheme and corresponding nucleotide substitution models for our data. The data were initially partitioned with ribosomal markers partitioned by gene and protein encoding genes partitioned by codon position. The analysis was run using a greedy search scheme (search = greedy), with all models considered (models = all). The corrected Akaike information criteria (AICc) was used to evaluate the fit of competing models. Alignments of the individual markers (12S: 463 bp; 16S: 600 bp; COI: 762 bp; COII: 621 bp; 18S: 1871 bp; 28S: 1608 bp; H3: 327 bp) were concatenated using Sequence Matrix 1.7.8 (Vaidya et al., 2011). PartitionFinder separated the data into 5 subsets as follows: 12S+16S (HKY+I+G+X), 18S+28S (GTR+I+G+X), codon positions 1+2 (GTR+I+G+X), COX position 3 (GTR+I+G), and H3 position 3 (GTR+I+G+X). Subsequent analyses were performed using this partitioned combined data set comprising 6,252 bp.
Phylogeny and Divergence Time Estimation
Phylogenetic analyses were performed on the Cipres Science Gateway (Miller et al., 2010) (www.phylo.org/). Exploratory RAxML analyses (Stamatakis et al., 2005) were executed on initial data sets including individual gene alignments to monitor potential contamination and assess gene performance, and on the combined data to assess terminal inclusion for constrained nodes in the divergence time analyses (see below). We performed RAxML rapid bootstrapping with a subsequent ML search (Stamatakis et al., 2005) executing 500–1000 bootstrap inferences using a GTR+G model [as recommended in Stamatakis et al. (2008; the RAxML 7.0.3 manual)]. FigTree 1.4 (Rambaut, 2012) was used to visualize the resulting trees.
Phylogeny and divergence time estimation was performed simultaneously in BEAST 2 (v2.4.5) (Bouckaert et al., 2014) using 5 unambiguous crown-group phasmid fossils as minimum calibration points that we were able to assign to specific nodes on the tree using synapomorphy-based anatomical evidence (Bradler et al., 2015) (Supplementary Table S2). The analysis was run using a lognormal distributed relaxed clock model with a Yule speciation prior. Clock and tree parameters were linked across all data partitions. Fossil calibrations were implemented using a lognormal prior distribution with a log-mean of 1.0 and log-SD of 1.0. Five separate analyses comprising 80–100 million generations each, with parameters sampled every 4,000 generations, were run to ensure convergence. Convergence and sufficiently high effective sample sizes (ESSs) of parameter estimates were verified in Tracer v1.6 (Rambaut et al., 2014). The initial 5,000 trees sampled were discarded as burnin and TreeAnnotator v1.8.4 (Rambaut and Drummond, 2016) was used to calculate a maximum credibility tree from the posterior 15001 trees. FigTree 1.4 (Rambaut, 2012) was used to visualize the resulting tree.
Oviposition Strategies
We scored oviposition strategies for our taxonomic sampling using the following 5 (unordered) states (Supplementary Table S1):
0: Drop or flick eggs
1: Bury/insert eggs into soil or crevices
2: Glue eggs to substrate
3: Pierce eggs into leaves
4: Produce ootheca
Twenty-two of the 285 (ca. 7%) taxa investigated were scored as missing data (?) for oviposition technique due to unknown natural history information for these species. Even so, our data provide more than sufficient taxonomic and geographic coverage to infer general patterns regarding the timing and geographic pattern of evolution of egg laying techniques in stick and leaf insects. Ancestral state reconstruction of egg-laying strategies was performed in Mesquite 3.31 (Maddison and Maddison, 2017) using parsimony unambiguous optimization.
Results and Discussion
The Evolution of Oviposition Strategies
The diversity exhibited in phasmid ova and oviposition strategies is striking. Our results unequivocally demonstrate that the ancestral oviposition strategy for stick and leaf insects is to remain in the foliage and drop or flick eggs to the ground (Figures 4–6) as suggested previously (Bradler, 2009; Goldberg et al., 2015), contradicting earlier assumptions that favored a ground-dwelling ecomorph to represent the ancestral character state (Carlberg, 1987). The phasmid egg capsule is a defining phenotypic character for Phasmatodea. It includes the presence of both a detachable lid-like operculum at its anterior pole through which the first instar nymph emerges, and an interior, air-containing plate that is demarcated externally by the micropylar plate (Sellick, 1997b). In Euphasmatodea (all phasmids excluding Timema) the egg capsule is greatly hardened. The eggs of Timema are soft but the females coat them with pulverized soil or sand that hardens to provide modest outer protection. The greatly hardened egg capsule of Euphasmatodea represents a key innovation in phasmid evolution, equipping the eggs to withstand an otherwise potentially perilous fall from the canopy, float extended periods of time (e.g., >1 year) on sea water (Kobayashi et al., 2014) and pass through the intestine of birds (Suetsugu et al., 2018) without compromising viability. The Western Nearctic genus Timema includes 21 species and Euphasmatodea just over 3,000 species (Bradler and Buckley, 2018). The striking disparity in species richness between these sister clades strongly suggests significant ecological opportunity [e.g., acquisition of key innovations, colonization of new habitat, extinction of antagonists (Yoder et al., 2010)] obtained in Euphasmatodea, and whereas there are certainly many selective factors at play (e.g., horizontal gene transfer of pectinases Shelomi et al., 2016), the evolution of a hardened egg capsule in Euphasmatodea is foremost.
Figure 4
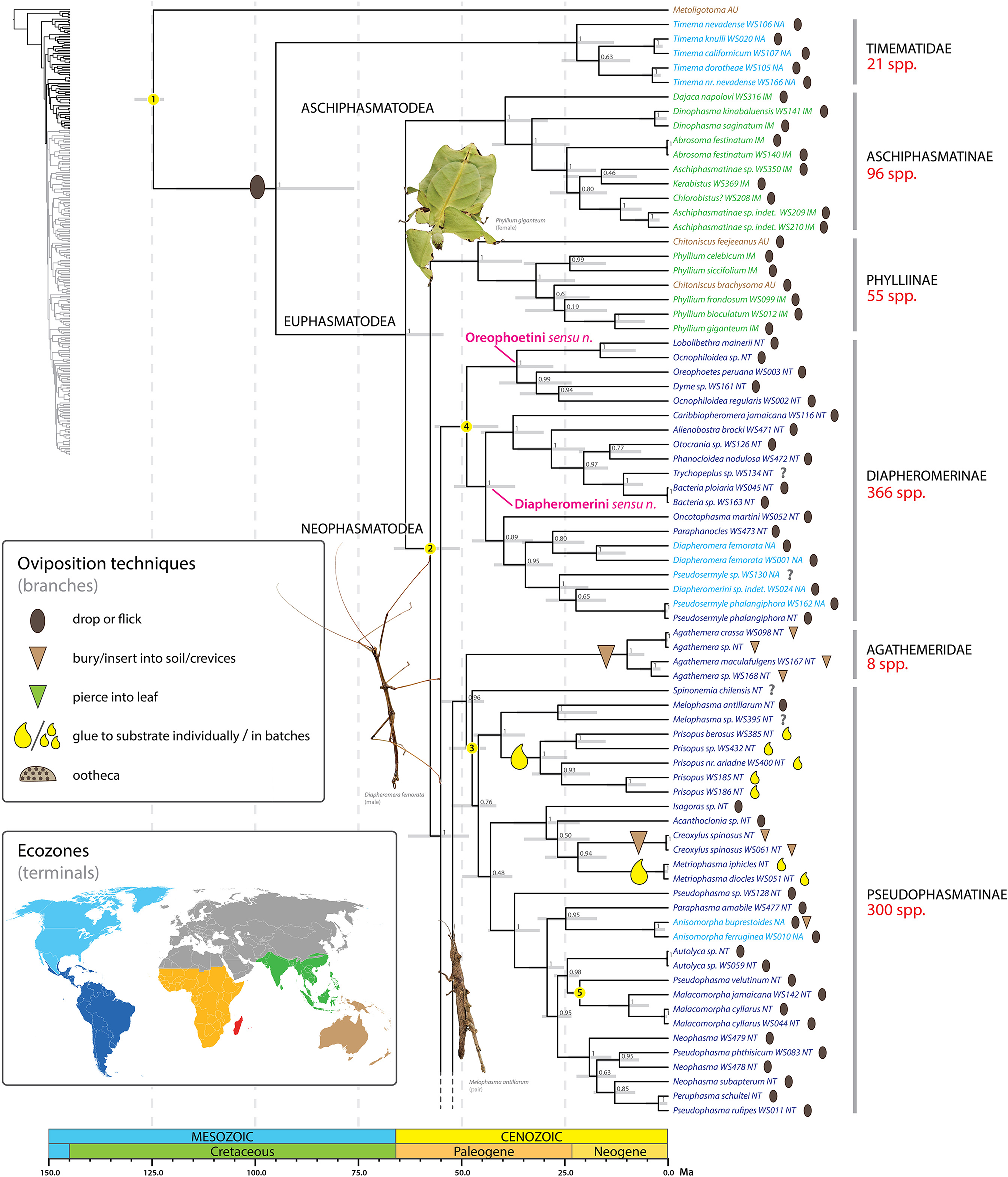
Chronogram of Phasmatodea (part 1 of 3). The full time tree is shown to the left with the emboldened region enlarged and colored for discussion. Fossil calibration points are denoted with numbered yellow circles (see Supplementary Table S2). Ninety-five percentage confidence intervals around node ages are indicated by gray bars. Bayesian posterior probability confidence intervals are listed for each node. Terminals are colored by ecozone following the inset. Oviposition technique for each sampled species is shown to the right of the terminal according to the inset, with inferred transitions optimized and marked on the branches. Photos of stick and leaf insects by Melinda Fawver (Diapheromera femorata), Isselee (Phyllium giganteum), Bruno Kneubühler (Melophasma antillarum) modified for use, used by permission.
Adult female stick insects lay eggs over a period of several months at a rate of one to several per day (Goldberg et al., 2015). During periods of active oviposition the sound of phasmid eggs dropping to the forest floor is reminiscent of the onset of rain (Beier, 1968). The ground plan oviposition technique of dropping/flicking eggs represents a boon for maintaining their masquerade as females are not required to move in search of specific oviposition sites, but can remain put, invisible to threat. Dropping/flicking eggs is not only employed by the earliest diverging (extant) lineages (e.g., Aschiphasmatinae, Phylliinae, Diapheromerinae) but occurs in the majority of phasmid species worldwide. In fact, all inferred transitions away from this ancestral oviposition strategy to a new technique occur in taxa well nested within major phasmid lineages (see below). Droppers and flickers occur in all biogeographic regions where phasmids occur.
Phasmids have limited dispersal ability (Compton and Ware, 1991) and are generally considered non-volant (Bradler et al., 2015). Roughly 60% of phasmid species exhibit greatly reduced wings or lack wings altogether in the adult form (Whiting et al., 2003) and even winged forms may not exhibit sustained flight (Maginnis, 2006). Various stick and leaf insects have evolved three key innovations to compensate for the loss of motility associated with the phenotypic and behavioral consequences of extreme masquerade: polyphagy, parthenogenesis, and myrmecochory. In general, flightless species are polyphagous (linked to their limited motility) whereas volant species (e.g., Necrosciinae) exhibit a more restricted diet and are regarded as host plant specialists (Blüthgen et al., 2006). Roughly 10% of phasmids exhibit varying forms of facultative parthenogenesis (e.g., Parabacillus, Bacillus, Clonopsis) including species hybrid parthenogens (Note that hybridogenetic females can escape hybridity back to Mendelian reproduction in just one generation Scali, 2009).
Another major key innovation in phasmid diversification appears to be the evolution of capitulate eggs, which facilitate ant-mediated dispersal, or myrmecochory (Compton and Ware, 1991; Hughes and Westoby, 1992; Windsor et al., 1996; Stanton et al., 2015). Capitula are phenotypically and functionally analogous to elaiosomes of angiosperm seeds. In fact, capitula and elaiosomes have a similar chemical profile, thus stick insects, and flowering plants utilize the same chemical signaling pathway to exploit ant behavior (Stanton et al., 2015). Selective benefits for myrmecochorous stick insects overlap with those for angiosperms including distance dispersal as a way of escaping clumping of offspring and rapid removal as a protection against predation and fire, but also parasitism by wingless parasitoid wasps that specialize on phasmid eggs (Compton and Ware, 1991). Like elaiosomes (Lengyel et al., 2009, 2010), capitula vary tremendously in form and given their taxonomic occurrence have likely evolved independently in geographically disparate lineages (Sellick, 1997a,b).
We infer at minimum 16 independent shifts to a burying/inserting eggs into soil/crevices oviposition strategy (Figures 4–6). All but one of these transitions represents a shift from the ancestral condition of dropping or flicking eggs (the one exception is Diesbachia and allies, see below). Depositing eggs into the soil or crevices of bark represents a major departure from simply dropping eggs from the canopy. Such major shifts in oviposition mode typically correspond to a major shift in microhabitat, egg phenotype, and often, adult phenotype. For example, most of the shifts to deposition in soil/crevices correspond to adapting to a ground or bark dwelling microhabitat, with a corresponding robust adult phenotype and egg deposition directly into the soil. These eggs are typically cylindrical, bullet-shaped, or otherwise modified for ease of burial (e.g., Figures 3Z,A',E'). Extreme examples include two independent shifts, one in Dryococelus australis and another within a New Zealand clade including Trapezaspis, Microcanachus, and Canachus (Figure 6). These are clear instances of adapting to a ground dwelling microhabitat, with a corresponding robust tree lobster phenotype (Buckley et al., 2009) and egg deposition directly into the soil. This trend is also evident in several other taxa including the Eurycanthomorpha (Bradler, 2009) of Lonchodinae (Figure 5) and, to a lesser degree, the Heteropteryginae, and the South American genus Agathemera (Figure 4).
Figure 5
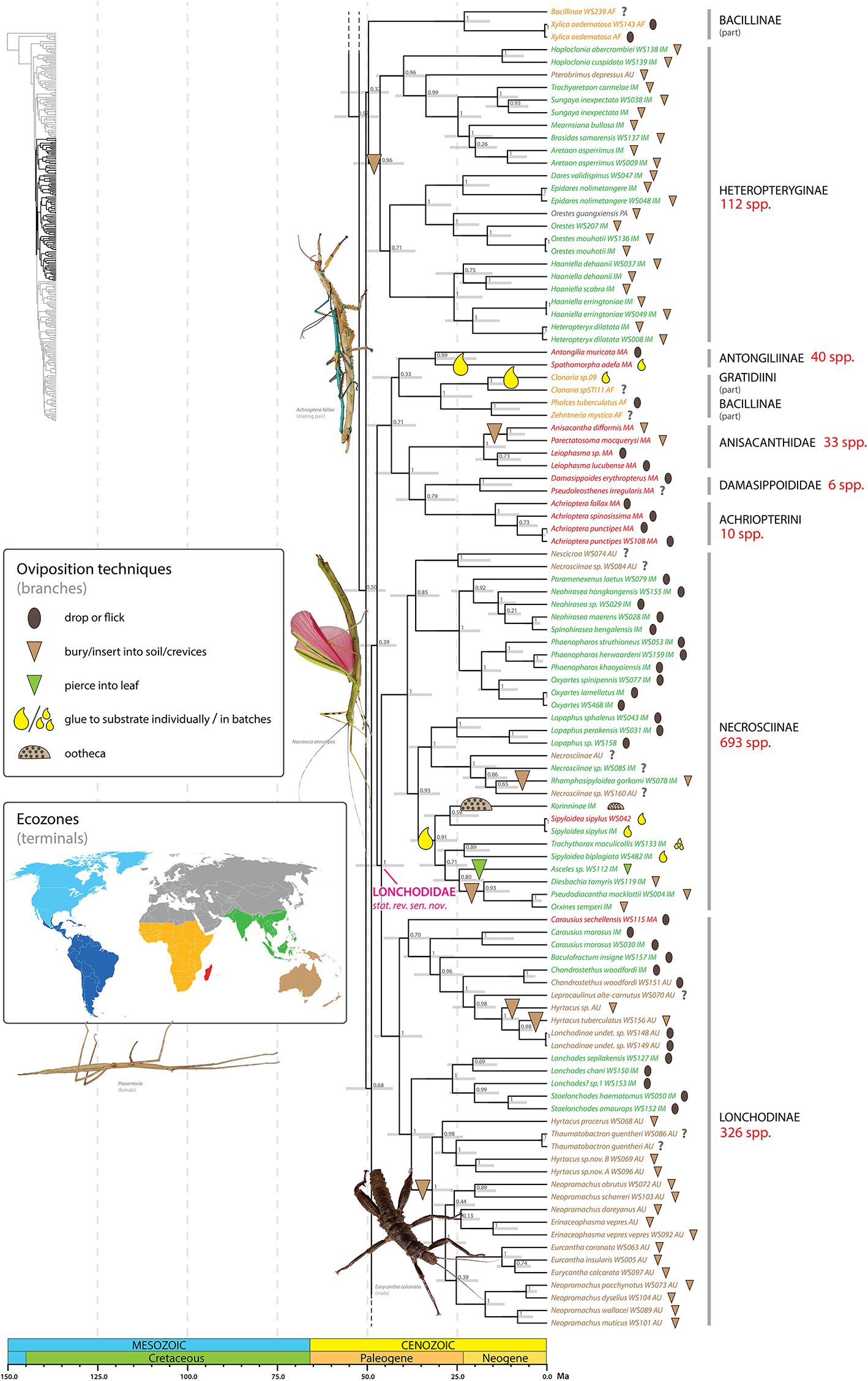
Chronogram of Phasmatodea (part 2 of 3). The full time tree is shown to the left with the emboldened region enlarged and colored for discussion. See Figure 1 caption for details. Photos of stick and leaf insects by Igor Siwanowicz (Achrioptera fallax), Isselee (Necroscia annulipes), Bruno Kneubühler (Prosentoria) modified for use, Valentino2 (Eurycantha calcarata), used by permission.
At least one shift to egg deposition into soil/crevices, inferred in the most recent common ancestor of the Heteropteryginae, resulted in a radiation of over 100 species (Figure 5). Heteropteryginae are distributed primarily in Indo-Malaysia, with only a few Australasian or Palearctic species. We estimate Heteropteryginae diverged from its sister taxon some time between 56 and 44 Ma (ca. 50 Ma) and began to radiate between 53 and 40 Ma (ca. 46.4 Ma). This lineage comprises species generally exhibiting a robust phenotype compared to the quintessential stick-like phasmids and includes the massive Jungle Nymph, Heteropteryx dilatata.
Adhering eggs to substrate evolved at least seven times throughout phasmid diversification (Figures 4–6). We infer two independent shifts to gluing eggs among the Neotropical Pseudophasmatinae in the taxa Prisopus and Metriophasma (Figure 4). Other shifts occur in Antongiliinae, two disparate clades of Gratidiini (Pachymorphinae), Necrosciinae, and Mascarene Lanceocercata. Most of the inferred shifts to a gluing oviposition strategy apparently did not result in marked subsequent diversification. However, one exception may be the Afrotropical and Indomalaysian clade comprising species of Clonaria, and Sceptrophasma (Gratidiini) (Figure 6). It is difficult to estimate the species-richness of this clade due to the fact that Clonaria (or Gratidiini) is recovered as polyphyletic, with some Afrotropical species distantly removed, recovered sister to Phalces and Zehntneria. In addition to the problematic generic boundaries, the species diversity of Clonaria is not well-accounted for.
Figure 6
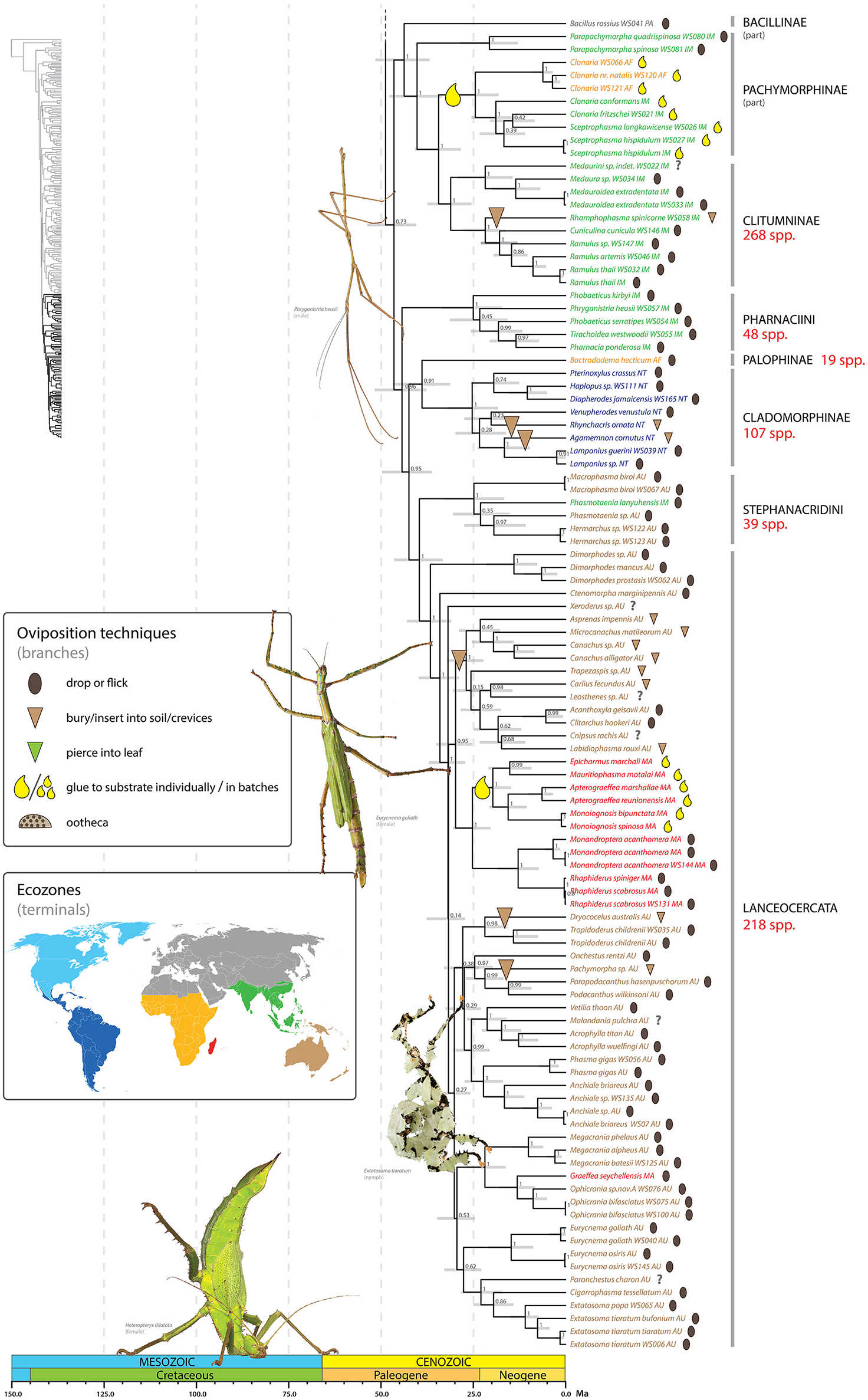
Chronogram of Phasmatodea (part 3 of 3). The full time tree is shown to the left with the emboldened region enlarged and colored for discussion. See Figure 1 caption for details. Photos of stick and leaf insects by Finalion (Eurycnema goliath), Laetitia Bourgois (Extatosoma tiaratum), Magnus Forsberg (Heteropteryx dilatata), Bruno Kneubühler (Phryganistria heusii) modified for use, used by permission.
Necrosciinae is the most species-rich and ecologically diverse lineage in the order treated at the subfamily rank; the ca. 700 species collectively employ all modes of phasmid oviposition (Figure 5). Rhamphosipyloidea gorkomi buries its eggs, but the oviposition mode of closely related taxa is unknown and thus so are the evolutionary transitions leading to this shift to depositing eggs in soil. One lineage comprising Korinninae sp., Trachythorax, Diesbachia and allies, Asceles, and several species of Sipyloidea exhibits a diverse range of oviposition strategies. It is unclear what selective pressures may have promoted this diversification, but we infer an initial shift from dropping/flicking to adhesion at the base of this lineage. Following this shift to gluing eggs singly, multiple instances of evolutionary novelty are evident. The general reproduction strategy of Phasmatodea regardless of oviposition mode is to deposit eggs singly thereby avoiding aggregating their offspring. In some taxa, including Trachythorax, Neoclides (not sampled in present study), and others, the females glue their eggs in loose single layer clusters (Goldberg et al., 2015). We recovered the ootheca producing Korinninae sister to Sipyloidea spp. that glue their eggs singly. Because the production of ootheca is a remarkable departure from the remaining phasmid oviposition strategies, reconstructing the ancestral states and evolutionary transitions leading to this shift is of great interest. The analyses of Goldberg et al. (2015) support Asceles, which skewer their eggs into leaves, as the sister to Korinninae. It is unclear under this sister group hypothesis what the ancestral oviposition condition might be. The evolution of an ootheca oviposition strategy from an ancestral mode of adhesion as supported by our results is an intuitive pathway since the sticky glandular secretion would already be in place (see Goldberg et al., 2015). We recovered a clade with moderate support (PP = 80) formed by sister taxa each exhibiting a novel mode of oviposition from the ancestral adhesion strategy. One of these lineages includes taxa that insert their eggs into soil or crevices of bark (e.g., Diesbachia, Pseudodiacantha, Orxines). Interestingly, the other lineage is Asceles, which pierces their eggs into leaves (Figure 2E).
Phasmatodea Systematics
Our molecular phylogeny represents the most extensively sampled estimate of the phasmid tree of life to date, in terms of the unique lineages, number of terminals, and number of loci. Our results present many significant relationships, some proposed previously and some novel. Corroborating earlier studies (Buckley et al., 2009; Bradler et al., 2014, 2015; Goldberg et al., 2015; Büscher et al., 2018), our results indicate a general pattern of strong branch support for the major lineages and toward the tips of the tree but only moderate support at some of the deeper relationships (Figures 4–6). Even so, we are encouraged that compared to previous molecular analyses (e.g., Buckley et al., 2009; Bradler et al., 2014, 2015; Goldberg et al., 2015; Büscher et al., 2018) the backbone relationships are in general recovered with higher support, an observation likely tied to increased taxon and gene sampling. The observed pattern of major diversification of lineages over a relatively short period of time may be the signature of an ancient rapid radiation of major phasmatodean lineages (see also Bradler, 2015; Bradler and Buckley, 2018).
We recovered an early subdivision of the Euphasmatodea into Aschiphasmatinae and all remaining stick and leaf insects with maximum support (PP = 1) (Figure 4). Aschiphasmatinae are enigmatic stick insects. They have been recovered as the earliest or one of the earliest diverging extant lineages of Euphasmatodea based on morphological (Tilgner, 2002) and molecular phylogenetic analyses (Buckley et al., 2009; Bradler et al., 2015). Engel et al. (2016) regarded this lineage as Aschiphasmatodea, with the remainder of the Euphasmatodea forming the Neophasmatodea. Within Aschiphasmatinae, Dajaca is the sister group to all remaining Aschiphasmatinae, thus reflecting its subdivision into Dajacini and Aschiphasmatini as suggested by Bragg (2001). However, Aschiphasmatidae sensu Bragg (2001) (= Aschiphasmatinae + Korinninae) is not supported since the species-poor Korinninae is recovered as a subordinate lineage of Necrosciinae as shown previously (Goldberg et al., 2015). Anatomical features supporting Neophasmatodea include galealobulus (maxillary lobe at base of galea) present (absent in Timema and Aschiphasmatinae), pro-spina absent (present in Timema and Aschiphasmatinae) (Tilgner, 2002).
The Phylliinae, or leaf insects, are strongly supported as the earliest diverging lineage of Neophasmatodea (Figure 4). This placement is consistent with the recent molecular study of Bradler et al. (2015), and additional studies employing a narrower taxon sampling regime also recover Phylliinae as a relatively early diverging lineage within Euphasmatodea (Buckley et al., 2009; Kômoto et al., 2011). This Old World lineage includes the 47 million year old fossil Eophyllium messelense (Wedmann et al., 2007), one of the oldest known extinct euphasmatodeans. Members of Phylliinae possess a unique egg morphology, making them distinct from most other phasmids. The edge of the micropylar plate is broken up into strips creating a fringe around its border (Sellick, 1997b). Fringed plates also occur in the Datamini (Heteropteryginae).
In contrast to some earlier studies that recovered paraphyletic Diapheromerinae at the euphasmatodean base (Whiting et al., 2003; Bradler et al., 2014 in part), we observe monophyletic New World Diapheromerinae (with the inclusion of Otocrania, see below) (Figure 4). However, our topology does not reflect the current classification (Phasmida Species File, Brock et al., 2017) of Diapheromerinae into the three subgroups Oreophoetini (Oreophoetes), Ocnophilini (Ocnophiloidea), and Diapheromerini (the remaining Diapheromerinae taxa in our tree). Instead, we recover with maximum support (PP = 1) a clade comprising Lobolibethra, Dyme, Oreophoetes, and polyphyletic Ocnophiloidea and this clade forms the sister group to the remaining Diapheromerinae. Perhaps a more significant result is the placement of Otocrania (formerly incorrectly assigned to Cladomorphinae) among the Diapheromerinae with strong support indicating its true affinity within Diapheromerinae. Further work is needed to delimit meaningful groups within Diapheromerinae reflective of their evolutionary history, but for now we formally transfer Otocrania to Diapheromerinae and recognize only two tribes within the subfamily, Diapheromerini sensu nov. and Oreophoetini sensu nov.
Diapheromerini, Kirby 1904 sensu nov.
Type genus. Diapheromera Gray, 1835: [18]
Diagnosis. The tribe Diapheromerini sensu nov. is well-supported by molecular data (see above), but to date very few anatomical apomorphies have been identified for the group. Most Diapheromerini sensu nov. produce eggs with a matrix (non-stalked) capitulum.
Included taxa. Diapheromerini sensu nov. comprises the majority of the species diversity within the subfamily, including most taxa previously classified as Diapheromerini (except Dyme and Lobolibethra; see below) and Otocrania, formerly regarded as Cladomorphinae.
Oreophoetini, Zompro 2001 sensu nov.
Type genus. Oreophoetes Rehn, 1904: [56]
Diagnosis. The tribe Oreophoetini sensu nov. is well-supported by molecular data (see above), but to date no morphological characters have been identified that unite the group. Oreophaetini sensu nov. produce eggs lacking a capitulum and are restricted to the Neotropics.
Included taxa. Oreophoetini sensu nov. includes Oreophoetes, Ocnophiloidea Zompro, Lobolibethra, and Dyme. Other taxa formerly included in Oreophoetini (e.g., Oreophoetophasma Zompro) Ocnophiloidea (Dubiophasma Zompro, Exocnophila Zompro, Ocnophila Brunner von Wattenwyl, Parocnophila Zompro) but not sampled in this or previous molecular phylogenetic studies are considered incertae sedis.
Another strongly supported Neotropical clade (PP = 0.96) comprises the species poor South American Agathemera and Heteronemiini + the species rich Pseudophasmatidae (Figure 4), often referred to as Pseudophasmatinae (Bradler et al., 2014, 2015; Goldberg et al., 2015). These taxa were considered as Pseudophasmatinae by Günther (1953), but its subgroups Agathemera, Heteronemiini (= Bacunculini sensu Günther, 1953) and Prisopodini were subsequently removed from Pseudophasmatinae (or Pseudophasmatidae, Zompro, 2004). Based on morphological evidence, Agathemera was repeatedly placed as sister group to all remaining Euphasmatodea (Klug and Bradler, 2006; Bradler, 2009; Friedemann et al., 2012), but this hypothesis is in sharp contrast to all molecular studies which support Agathemera as a subordinate euphamatodean taxon (Whiting et al., 2003; Buckley et al., 2009; Bradler et al., 2014, 2015; Goldberg et al., 2015; Büscher et al., 2018), albeit with no consensus regarding its placement among the remaining stick insects. Zompro (2004) erected the subfamily Prisopodinae for Prisopus and related species (e.g., Melophasma) and the family Prisopodidae for Prisopodinae + Korinninae. Prisopus and allies are enigmatic taxa. We recovered Prisopodini nested within Pseudophasmatinae (see also Goldberg et al., 2015), but in some analyses it was supported as an early divergent neophasmatodean lineage far removed from Pseudophasmatinae. The internal phylogeny of Pseudophasmatidae does not reflect the current classification following the Phasmida Species File (Brock et al., 2017, mainly based on Zompro, 2004). The subgroup Xerosomatinae is supported (PP = 1) but not its subdivision into tribes, e.g., Xerosomatini (represented here by Acanthoclonia and Creoxylus) and Prexaspini (represented by Isagoras and Metriophasma) (see also Bradler et al., 2015). It is also evident from our results that the eponymous and species-rich genus Pseudophasma is polyphyletic.
Represented in the present study by 24 exemplars, the Heteropteryginae (or Heteropterygidae, Zompro, 2004) are supported as monophyletic (Figure 5). Multiple heteropterygine taxa (e.g., Datamini) exhibit relatively high rates of sequence divergence across the sampled genes as reflected in their long branches and unstable position in Euphasmatodea when the taxon sampling is limited. With increased taxon sampling of Heteropteryginae we observe increased support for its monophyly. For example, early molecular analyses that included three to four exemplars (Whiting et al., 2003; Buckley et al., 2009) did not recover a monophyletic Heteropteryginae, whereas subsequent studies with increased sampling have (Goldberg et al., 2015; Büscher et al., 2018). The heteropterygine tribes Datamini, Heteropterygini, and Obrimini are each recovered with high support (PP = 1, 1, 0.96 respectively), but the relationships between these three tribes are not well-supported, with Datamini forming the sister group to Heteropterygini (PP = 0.71). The other two studies that included exemplars of all three tribes each recovered a unique scenario of tribal relationships (Bradler et al., 2015; Goldberg et al., 2015; Büscher et al., 2018). Our results further illustrate several taxonomic shortcomings within Heteropteryginae on a lower scale. For example, the Tisamenini as recently erected by Hennemann et al. (2016), represented here by Hoploclonia and Pterobrimus, are paraphyletic. What's more, Hoploclonia abercrombiei was recently synonimyzed with H. cuspidata (Seow-Choen, 2016), but the genetic distance spanning these two taxa (reflected by relative branch lengths) clearly indicates they are distinct species (Figure 5); H. abercrombiei should be brought out of synonymy accordingly. Similarly, the genus Haaniella is paraphyletic with respect to Heteropteryx dilatata (see also Goldberg et al., 2015), the later strongly supported as the sister taxon to H. erringtoniae. If these relationships are confirmed in subsequent analyses Haaniella or parts of it will need to be synonymized with Heteropteryx. Only a few superficial characters have been used to separate the monotypic Heteropteryx from Haaniella including Heteropteryx having a strongly conical and elevated head, distinct spines on the abdominal terga of females, a shortened mesothorax in the male form, and specific coloring in both sexes (Hennemann et al., 2016). The remarkable similarity in form of the eggs of some Haaniella species and Heteropteryx strongly indicates their close affinity.
We recovered an African-Malagasy clade with low to moderate support (PP = 0.71) (Figure 5). The clade comprises two Malagasy lineages recovered with high support: Antongiliinae (Antongilia + Spathomorpha) (PP = 0.99) and Anisacanthidae + Damasippoidini + Achriopterini (PP = 1). The one subordinate African lineage (PP = 1) consists of the Gratidiini (Pachymorphinae) taxa Clonaria (part) and Zehntneria and the Bacillinae taxon Phalces. Bradler et al. (2015) also recovered this African clade with maximum support. Neither Bacillinae nor Pachymorphinae are recovered as monophyletic. The polyphyly of these two subfamilies has been demonstrated in previous studies as well (Buckley et al., 2009). Some of the sampled Bacillinae species (or Bacillidae species sensu Zompro, 2004) (e.g., Xylica oedematosa, Bacillus rossius) exhibit long branches and their respective phylogenetic positions tend to be unstable across analyses. For example, one African Antongiliinae/Bacillinae clade containing Xylica and allies (PP = 1) was recovered with negligible support (PP = 0.32) as the sister group to Heteropteryginae. In contrast, previous molecular studies have recovered Xylica oedematosa as the earliest divergent lineage of Euphasmatodea (Bradler et al., 2015), or part of that lineage (Buckley et al., 2009). Note that the clade comprising Xylica oedematosa and allies was recovered in preliminary analyses of these data with Parapachymorpha, Gratidiini (in part), and Clitumninae (excluding Pharnaciini); this instability is reflected by the poor branch support along the base of the tree spanning these alternative placements. Antongiliinae as erected by Zompro (2004) and currently recognized in the Phasmida Species File (Brock et al., 2017), represented here by Antongilia and Xylica, is not supported. In contrast to Bradler et al. (2015), we recovered Antongilia muricata and Spathomorpha adefa as sister taxa (PP = 0.99) lending support to a single colonization hypothesis rather than independent colonization of Madagascar by these taxa.
A sister group relationship between Necrosciinae and Lonchodinae has been repeatedly shown (Bradler et al., 2014; Goldberg et al., 2015) and is corroborated here based on a significantly enlarged taxon sampling with maximum support (PP = 1) (Figure 5). Necrosciinae and Lonchodinae are both characterized by the presence of long antennae and the absence of the area apicalis on the tibiae (Günther, 1953; Bradley and Galil, 1977). Both subfamilies are currently treated as members of different families (Necrosciinae as part of Diapheromeridae and Lonchodidae as part of Phasmatidae) in the Phasmida Species File (Brock et al., 2017), highlighting the inadequacy of this database to represent the higher-level phylogenetic relationships among stick and leaf insects despite its utility at the species level. We formally recognize the clade comprising Necrosciinae and Lonchodinae at the family level as Lonchodidae stat. rev. sensu nov.
Lonchodidae Brunner von Wattenwyl, 1893 stat. rev. sensu nov.
Type genus. Lonchodes Gray, 1835: [19]
Diagnosis. Lonchodidae stat. rev. sensu nov. are strongly supported by molecular data (see above) but only vaguely characterized by morphology including the following combination of features: Adults with antennae long, and area apicalis on tibiae absent. Lonchodidae stat. rev. sensu nov. are distributed in Indomalaysia, Australasia, Madagascar, and related islands, and with very few species occurring in the Palaearctic.
Included taxa. Lonchodidae stat. rev. sensu nov. comprises all taxa included in the current concept of the constituent subfamilies Lonchodinae and Necrosciinae as set forth in Bradler et al. (2014).
The species-rich genus Sipyloidea is clearly polyphyletic with species recovered in multiple clades within Necrosciinae (Figure 5). Most higher phasmid taxa exhibit high levels of endemism. In sharp contrast, the genus Sipyloidea as presently constituted is widely distributed, with species occurring in Madagascar, Africa, Indo-Malaysia, and Australasia. Our results clearly reject the hypothesis that Sipyloidea is a species-rich genus with a broad geographic distribution, but rather represents several unique lineages with restricted geographic distributions.
Within Lonchodinae, the Eurycanthomorpha comprising the New Guinean tree lobsters (Eurycantha spp.) and allies (Buckley et al., 2009; Bradler et al., 2015; Büscher et al., 2018) are recovered with maximum support (PP = 1). However, our results indicate that several lonchodine genera are polyphyletic, including Carausius, Hyrtacus, and Neopromachus.
We recovered with maximum support a wingless lineage comprising Bacillus rossius, Parapachymorpha, Gratidiini (in part: Clonaria spp., Sceptrophasma), Clitumnini and Medaurini (Figure 6). Whereas, this clade represents a grouping that has not been formally recognized, variations of it have been suggested repeatedly (Hennemann and Conle, 2008; Bradler, 2009; Buckley et al., 2009; Bradler et al., 2014, 2015). Based on phylogenetic inference of morphological data, Bradler (2009) recovered two disparate clades, one comprising Bacillus rossius + Gratidiini (Clonaria, Sceptrophasma) and the other formed by Clitumnini, Medaurini, and Parapachymorpha. Molecular phylogenetic studies strongly support the grouping of Clitumnini (e.g., Ramulus), Medaurini (without Parapachymorpha), and African and Southeast Asian Gratidiini (Bradler, 2009; Buckley et al., 2009; Bradler et al., 2014, 2015; Büscher et al., 2018). Traditionally, Gratidiini was classified as Pachymorphinae based on the presence of short antennae, whereas Clitumnini and Medaurini were placed in Phasmatinae. Both subfamilies are clearly polyphyletic (Bradler, 2009; Buckley et al., 2009; Bradler et al., 2014). Hennemann and Conle (2008) established the subfamily Clitumninae, comprising Clitumnini + Medaurini (including Parapachymorpha) + Pharnaciini. However, Clitumninae are not recovered as monophyletic in the present analysis. Instead Clitumnini and Medaurini (without Parapachymorpha) are sister taxa (PP = 1) with the African and Southeast Asian Gratidiini subtending them (PP = 1). Parapachymorpha forms the sister taxon (PP = 1) of this undisputed clade and Bacillus rossius is recovered as the sister to the whole group (PP = 1). Pharnaciini is strongly supported as more closely related to Lanceocercata and allies than to Medaurini and Clitumnini (see below).
Pharnaciini contains a number of giant-sized stick insects including the recently described Phobaeticus chani. Our sampling includes two species of Phobaeticus, P. kirbyi, and P. serratipes, but our results suggest the genus as presently constituted is not monophyletic, with P. serratipes strongly supported (PP = 0.99) as the sister group to Tirachoidea + Pharnacia. In our analysis (see Figure 6) the tribe Pharnaciini forms the sister group (PP = 0.96) to the clade [(Cladomorphinae + Bactrododema) + (Stephanacridini + Lanceocercata)]. This diverse grouping comprises a lineage that has been repeatedly shown in the past, albeit with differing placements of Bactrododema (Buckley et al., 2009, 2010; Bradler et al., 2014, 2015; Goldberg et al., 2015; Büscher et al., 2018).
The internal relationships of Lanceocercata (Figure 6) are recovered with varying support and generally corroborate those recovered in previous studies (Buckley et al., 2009, 2010; Bradler et al., 2014, 2015; Goldberg et al., 2015; Büscher et al., 2018). The genus Dimorphodes forms the sister group (PP = 1) to all remaining Lanceocercata (PP = 1). Ctenomorpha marginipennis and Xeroderus represent additional early divergent taxa. Within Lanceocercata a New Caledonia/New Zealand clade is recovered (PP = 1) (Buckley et al., 2010) as well as a Mascarene lineage (PP = 1). We further find support for monophyletic Platycraninae (PP = 1), the coconut stick insects. The sister taxon to Dryococelus australis, The Lord Howe stick insect, remains unclear with no consensus across analyses (Buckley et al., 2009, 2010; Bradler et al., 2015; Goldberg et al., 2015); candidates include Eurycnema (Buckley et al., 2009), Ctenomorpha + Eurycnema (Buckley et al., 2009) and Platycraninae (Bradler et al., 2015). We recovered a strongly supported (PP = 0.98) sister grouping of Dryococelus australis and the fully winged Tropidoderus childrenii from the East Coast of Australia.
Clearly much work is needed to delimit meaningful groups within Phasmatodea reflective of their evolutionary history. Corroborating previous phylogenetic studies (see above), our results highlight the limitations of the current classification scheme for Phasmatodea and the need for future phylogenetic revision and reassessment of concepts and boundaries of higher taxa. The relationships among several major phasmid lineages remain unsettled. Even so, we are encouraged with our results of having recovered major phasmatodean divergences with significant support [e.g., Aschiphasmatinae and Phylliinae as early divergent taxa of Euphasmatodea; Diapheromerinae sensu nov.; Pseudophasmatinae s.l.; Necrosciinae + Lonchodinae; Clitumnini, Medaurini, Gratidiini and allies; ((Cladomorphinae + Bactrododema) + (Stephanacridini + Lanceocercata))] and look forward to continued progress by the phasmid community.
Statements
Data availability statement
All datasets analyzed for this study are included in the manuscript and the Supplementary Material. Oviposition mode data is provided in Supplementary Table S1. The DNA sequences used for phylogenetic inference can be found at GenBank [http://www.ncbi.nlm.nih.gov/genbank] under accession numbers MK291527-MK291924, MK296739-MK296749, and MK297240-MK297296. The alignment used for the phylogenetic reconstruction is provided as a Nexus input file as Supplementary Data Sheet 1.
Author contributions
JR, SB, and MW designed the research. JR generated and analyzed molecular data. JR, SB, and MW wrote the manuscript. JR generated the artwork. All authors have approved the final version of the manuscript.
Funding
This research was funded in part by the National Science Foundation (DEB- 1557114 to MW, JR, and SB; DEB-1256976 to Wendy Moore and JR) and by the German Science Foundation (BR 2930/5-1 to SB).
Acknowledgments
We recognize and are grateful for the support of the following people and agencies that have enhanced the value of this work. Christoph Seiler (Altlussheim, Germany), Seth Bybee (BYU), Joe McHugh (UGA), Gavin Svenson (CMNH), provided valuable specimens for this research. Yelena Pacheco helped generate sequence data for several critical taxa. Rohan Cleave, Melinda Fawver, Pavel German, Thierry Heitzmann, Isselee, Albert Kang, Bruno Kneubühler, Moritz Muschick, Piotr Naskrecki, Igor Siwanowicz, François Tetaert, Miroslaw Wasinski, Alex Wild, Art Wolfe, Valentino2, and the Museum of Victoria provided use of their stunning photos of stick and leaf insects. Special thanks go to Mark Miller and the CIPRES Science Gateway.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fevo.2018.00216/full#supplementary-material
Supplementary Table S1Taxon and data sampling for the present study. Identification of voucher specimens new to this study was performed or confirmed by SB.
Supplementary Table S2Fossil Calibrations used in the divergence time estimate in BEAST2.
Supplementary Table S3Corresponding taxonomic names for the stick and leaf insect eggs pictured in Figure 3.
Supplementary Data Sheet 1Nexus input file.
References
1
BedfordG. O. (1978). Biology and ecology of the Phasmatodea. Ann. Rev. Entomol.23, 125–149.
2
BeierM. (1968). Phasmida (Stab- oder Gespenstheuschrecken), in Handbuch der Zoologie, IV, 2, Vol. 10, eds HelmckeJ.-G.StarckD.WermuthH. (Berlin: Walter de Gruyter and Company), 1–56.
3
BlüthgenN.MetznerA.RufD. (2006). Food plant selection by stick insects (Phasmida) in a Bornean rain forest. J. Trop. Ecol. 22, 35–40. 10.1017/S0266467405002749
4
BouckaertR.HeledJ.KühnertD.VaughanT.WuC.-H.XieD.et al. (2014). BEAST 2: a software platform for bayesian evolutionary analysis. PLoS Comput. Biol.10:e1003537. 10.1371/journal.pcbi.1003537
5
BradlerS. (2009). Phylogenie der Stab- und Gespenstschrecken (Phasmatodea). Spec. Phyl. Evol.2, 3–139. 10.17875/gup2009-710
6
BradlerS. (2015). Der phasmatodea tree of life: Überraschendes und Ungeklärtes in der Stabschrecken-Evolution. Entomologie heute27, 1–23.
7
BradlerS.BuckleyT. R. (2018). Biodiversity of Phasmatodea in Insect Biodiversity: Science and Society, Volume II, eds FoottitR. G.AdlerP. H. (Hoboken, NJ: Wiley-Blackwell), 281–313.
8
BradlerS.CliquennoisN.BuckleyT. R. (2015). Single origin of Mascarene stick insects:ancient radiation on sunken islands?BMC Evol. Biol. 15:196. 10.1186/s12862-015-0478-y
9
BradlerS.RobertsonJ. A.WhitingM. F. (2014). A molecular phylogeny of Phasmatodea with emphasis on Necrosciinae, the most species-rich subfamily of stick insects. Syst. Entomol.39, 205–222. 10.1111/syen.12055
10
BradleyJ. C.GalilB. S. (1977). The taxonomic arrangement of the Phasmatodea with keys to the subfamilies and tribes. Proc. Entomol. Soc. Washington79, 176–208.
11
BraggP. E. (2001). Phasmids of Borneo.Borneo; Kota: Natural History Publications Kinabalu, 772.
12
BrockP. D.BüscherT. H.BakerE. (2017). Phasmida species file online: phasmida species file version 5.0/5.0, in Species 2000 and ITIS Catalogue of Life, eds RoskovY.OwerG.OrrellT.NicolsonD.BaillyN.KirkP. M.BourgoinT.DeWaltR. E.van NieukerkenW.DecockE.ZarucchiJ.PenevL. (Leiden: Species 2000; Naturalis). Available online at: www.catalogueoflife.org/col
13
BruscaR. C.MooreW.ShusterS. M. (2016). Invertebrates, 3rd Edn. Sunderland, MA: Sinauer Associates, 500.
14
BuckleyT. R.AttanayakeD.BradlerS. (2009). Extreme convergence in stick insect evolution: phylogenetic placement of the Lord Howe Island tree lobster. Proc. R. Soc. Lond. B276, 1055–1062. 10.1098/rspb.2008.1552
15
BuckleyT. R.AttanayakeD.NylanderJ. A. A.BradlerS. (2010). The phylogenetic placement and biogeographical origins of the New Zealand stick insects (Phasmatodea). Syst. Entomol.35, 207–225. 10.1111/j.1365-3113.2009.00505.x
16
BüscherT. H.BuckleyT. R.GrohmannC.GorbS. N.BradlerS. (2018). The evolution of tarsal adhesive microstructures in stick and leaf insects (Phasmatodea). Front. Ecol. Evol.6:69. 10.3389/fevo.2018.00069
17
CarlbergU. (1983). A review of different types of egglaying in the Phasmida in relation to the shape of the eggs and with a discussion on their taxonomic importance (Insecta). Biol. Zentralblatt102, 587–602.
18
CarlbergU. (1987). Evolutionary and ecological aspects on ovarian diversity in Phasmida, in Evolutionary Biology of Orthopteroid Insects, ed BaccettiB. (Chichester: Ellis Horwood Ltd.), 174–176.
19
ComptonS. G.WareA. B. (1991). Ants disperse the elaiosome-bearing Eggs of an African stick insect. Psyche98, 207–213.
20
EdgarR. C. (2004). MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acid Res.32, 1792–1797. 10.1093/nar/gkh340
21
EisnerT.MorganR. C.AttygalleA. B.SmedleyS. R.HerathK. B.MeinwaldJ. (1997). Defensive production of quinoline by a phasmid insect (Oreophoetes peruana). J. Exp. Biol.200, 2493–2500.
22
EngelM. S.WangB.AlqarniA. S. (2016). A thorny, ‘anareolate’ stick-insect (Phasmatidae s.l.) in Upper Cretaceous amber from Myanmar, with remarks on diversification times among Phasmatodea. Cret. Res.63, 45–53. 10.1016/j.cretres.2016.02.015
23
FriedemannK.WipflerB.BradlerS.BeutelR. G. (2012). On the head morphology of Phyllium and the phylogenetic relationships of Phasmatodea (Insecta). Act. Zool.93, 184–199. 10.1111/j.1463-6395.2010.00497.x
24
GhiselliF.MilaniL.ScaliV.PassamontiM. (2007). The Leptynia hispanica species complex (Insecta Phasmida): polyploidy, parthenogenesis, hybridization and more. Mol. Ecol.16, 4256–4268. 10.1111/j.1365-294X.2007.03471.x
25
GoldbergJ.BresseelJ.ConstantJ.KneubühlerB.LeubnerF.MichalikP.et al. (2015). Extreme convergence in egg-laying strategy across insect orders. Sci. Rep.5:7825. 10.1038/srep07825
26
GüntherK. (1953). Über die taxonomische Gliederung und geographische Verbreitung der Insektenordnung der Phasmatodea. Beitr. Entomol.3, 541–563.
27
HennemannF. H.ConleO. V. (2008). Revision of Oriental Phasmatodea: the tribe Pharnaciini, Günther, 1953, including the description of the world's longest insect, and a survey of the family Phasmatidae Gray, 1835 with keys to the subfamilies and tribes (Phasmatodea: “Anareolatae”: Phasmatidae). Zootaxa 1906, 1–316. Available online at: https://www.mapress.com/zootaxa/2008/f/z01906p316f.pdf
28
HennemannF. H.ConleO. V.BrockP. D.Seow-ChoenF. (2016). Revision of the Oriental subfamily Heteropteryginae Kirby, 1896, with a re-arrangement of the family Heteropterygidae and the descriptions of five new species of Haaniella Kirby, 1904. (Phasmatodea: Areolatae: Heteropterygidae). Zootaxa4159, 1–219. 10.11646/zootaxa.4159.1.1
29
HughesL.WestobyM. (1992). Capitula on stick insects and elaiosomes on seeds: convergent adaptations for burial by ants. Funct. Ecol.6, 642–648.
30
KômotoN.YukuhiroK.UedaK.TomitaS. (2011). Exploring the molecular phylogeny of phasmids with whole mitochondrial genome sequences. Mol. Phylogenet. Evol.58, 43–52. 10.1016/j.ympev.2010.10.013
31
KatohK.TohH. (2008). Recent developments in the MAFFT multiple sequence alignment program. Brief. Bioinformatics9, 276–285. 10.1093/bib/bbn013
32
KevanD. K. McE. (1977). The higher classification of orthopteroid insects: a general view. Lyman Entomol. Mus. Res. Lab. Memoirs4, 1–31.
33
KevanD. K. McE. (1982). Phasmatoptera, in Synopsis and Classification of Living Organisms, Vol. 2. ed ParkerS. F. (New York, NY: McGraw-Hill), 379–383.
34
KlugR.BradlerS. (2006). The pregenital abdominal musculature in phasmids and its implications for the basal phylogeny of Phasmatodea (Insecta: Polyneoptera). Organ. Divers. Evol.6, 171–184. 10.1016/j.ode.2005.08.004
35
KobayashiS.UsuiR.NomotoK.UshirokitaM.DendaT.IzawaM. (2014). Does egg dispersal occur via the ocean in the stick insect genus Megacrania (Phasmida: Phasmatidae)?Ecol. Res.29, 1025–1032. 10.1007/s11284-014-1188-4
36
LanfearR.FrandsenP. B.WrightA. M.SenfeldT.CalcottB. (2016). PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34, 772–773. 10.1093/molbev/msw260
37
LawJ. H.CrespiB. J. (2002). The evolution of geographic parthenogenesis in Timema walking-sticks. Mol. Ecol.11, 1471–1489. 10.1046/j.1365-294X.2002.01547.x
38
LengyelS.GoveA. D.LatimerA. M.MajerJ. D.DunnR. R. (2009). Ants sow the seeds of global diversification in flowering plants. PLOS ONE4:e5480. 10.1371/journal.pone.0005480
39
LengyelS.GoveA. D.LatimerA. M.MajerJ. D.DunnR. R. (2010). Convergent evolution of seed dispersal by ants, and phylogeny and biogeography in flowering plants: a global survey. Perspect. Plant Ecol. Evol. Syst.12, 43–55. 10.1016/j.ppees.2009.08.001
40
MaddisonW. P.MaddisonD. R. (2017). Mesquite: A Modular System for Evolutionary Analysis. Version 3.31 Available online at: http://mesquiteproject.org
41
MaginnisT. L. (2006). Leg regeneration stunts wing growth and hinders flight performance in a stick insect (Sipyloidea sipylus). Proc. R. Soc. Lond. B273, 1811–1814. 10.1098/rspb.2006.3508
42
MillerM. A.PfeifferW.SchwartzT. (2010). Creating the CIPRES Science Gateway for inference of large phylogenetic trees, in Proceedings of the Gateway Computing Environments Workshop (GCE) (New Orleans, LA), 1–8.
43
MisofB.LiuS.MeusemannK.PetersR. S.DonathA.MayerC.et al. (2014). Phylogenomics resolves the timing and pattern of insect evolution. Science346, 763–767. 10.1126/science.1257570
44
MooreP. D. (1993). How to get carried away. Nature361, 304–305.
45
PriddelD.CarlileN.HumphreyM.FellenbergS.HiscoxD. (2003). Rediscovery of the ‘extinct’ Lord Howe Island stick-insect (Dryococelus australis (Montrouzier)) (Phasmatodea) and recommendations for its conservation. Biodivers. Conserv.12, 1391–1403. 10.1023/A:1023625710011
46
RambautA. (2012). FigTree v1.4. Available online at: http://tree.bio.ed.ac.uk/software/figtree/
47
RambautA.DrummondA. J. (2016). TreeAnnotator v1.8.4. Available online at: http://www.beast2.org/
48
RambautA.SuchardM. A.XieD.DrummondA. J. (2014). Tracer v1.6. Available online at: http://beast.bio.ed.ac.uk/Tracer
49
RobertsonJ. A.SlipinskiA.MoultonM.ShockleyF.GiorgiA.LordN. P.et al. (2015). Phylogeny and classification of Cucujoidea and the recognition of a new superfamily Coccinelloidea (Coleoptera: Cucujiformia). Syst. Entomol.40, 745–778. 10.1111/syen.12138
50
ScaliV. (2009). Stick insects: parthenogenesis, polyploidy and beyond, in Life and Time: The Evolution of Life and its History, eds CasellatoS.BurighelP.MinelliA. (Padova: Cleup), 171–192.
51
ScaliV.MilaniL.PassamontiM. (2012). Revision of the stick insect genus Leptynia: description of new taxa, speciation mechanism and phylogeography. Contrib. Zool.81, 25–42. Available online at:http://www.ctoz.nl/vol81/nr01/a02
52
ScaliV.MilaniL.PassamontiM. (2013). Description and ecology of new Pijnackeria stick insects: four bisexual species and a triploid parthenogen with their phyletic relationships. J. Zool. Syst. Evol. Res.51, 213–226. 10.1111/jzs.12018
53
SchwanderT.HenryL.CrespiB. J. (2011). Molecular evidence for ancient asexuality in Timema stick insects. Curr. Biol.21, 1129–1134. 10.1016/j.cub.2011.05.026
54
SellickJ. T. (1997a). The range of egg capsule morphology within the Phasmatodea and its relevance to the taxonomy of the order. Ital. J. Zool.64, 97–104.
55
SellickJ. T. (1997b). Descriptive terminology of the phasmid egg capsule, with an extended key to the phasmid genera based on egg structure. Syst. Entomol.22, 97–122.
56
Seow-ChoenF. (2016). A Taxonomic Guide to the Stick Insects of Borne.Borneo; Kota Kinabalu: Natural History Publications, 454.
57
ShelomiM.DanchinE. G. J.HeckelD.WipflerB.BradlerS.ZhouX.et al. (2016). Horizontal gene transfer of pectinases from bacteria preceded the diversification of stick and leaf insects. Sci. Rep.6:26388. 10.1038/srep26388
58
StamatakisA.HooverP.RougemontJ. (2008). A rapid bootstrap algorithm for the RAxML Web-Servers. Syst. Biol.57, 758–771. 10.1080/10635150802429642
59
StamatakisA.LudwigT.MeierH. (2005). RAxML-III: a fast program for maximum likelihood-based inference of large phylogenetic trees. Bioinformatics21, 456–463. 10.1093/bioinformatics/bti191
60
StantonA. O.DiasD. A.O'HanlonJ. C. (2015). Egg dispersal in the Phasmatodea: convergence in chemical signalling strategies between plants and animals?J. Chem. Ecol. 41, 689–695. 10.1007/s10886-015-0604-8
61
SuetsuguK.FunakiS.TakahashiA.ItoK.YokoyamaT. (2018). Potential role of bird predation in the dispersal of otherwise flightless stick insects. Ecology99, 1504–1506. 10.1002/ecy.2230
62
TalaveraG.CastresanaJ. (2007). Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol.56, 564–577. 10.1080/10635150701472164
63
TilgnerE. H. (2002). Systematics of Phasmida. Dissertation, University of Georgia, Athens, GA.
64
TrewickS. A.Morgan-RichardsM.CollinsL. J. (2008). Are you my mother? Phylogenetic analysis reveals orphan hybrid stick insect genus is part of a monophyletic New Zealand clade. Mol. Phylogenet. Evol.48, 799–808. 10.1016/j.ympev.2008.05.025
65
VaidyaG.LohmanD.MeierR. (2011). SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics27, 171–180. 10.1111/j.1096-0031.2010.00329.x
66
WedmannS.BradlerS.RustJ. (2007). The first fossil leaf insect: 47 million years of specialized cryptic morphology and behavior. Proc. Natl. Acad. Sci. U.S.A.104, 565–569. 10.1073/pnas.0606937104
67
WhitingM. F.BradlerS.MaxwellT. (2003). Loss and recovery of wings in stick insects. Nature421, 264–267. 10.1038/nature01313
68
WindsorD. M.TrapnellD. W.AmatG. (1996). The egg capitulum of a Neotropical walkingstick, Calynda biscuspis, induces aboveground egg dispersal by the ponerine ant, Ectatomma ruidum. J. Inst. Behav.9, 353–367.
69
YoderJ. B.ClanceyE.Des RochesS.EastmanJ. M.GentryL.GodsoeW.et al. (2010). Ecological opportunity and the origin of adaptive radiations. J. Evol. Biol.23, 1581–1596. 10.1111/j.1420-9101.2010.02029.x
70
ZomproO. (2004). Revision of the genera of the Areolatae, including the status of Timema and Agathemera (Insecta, Phasmatodea). Verhandl. Naturw. Ver. Hamburg37, 1–327.
Summary
Keywords
phylogeny, adaptive radiation, key innovation, Phasmida, classification, systematics, taxonomy
Citation
Robertson JA, Bradler S and Whiting MF (2018) Evolution of Oviposition Techniques in Stick and Leaf Insects (Phasmatodea). Front. Ecol. Evol. 6:216. doi: 10.3389/fevo.2018.00216
Received
06 September 2018
Accepted
29 November 2018
Published
19 December 2018
Volume
6 - 2018
Edited by
Anthony I. Cognato, Michigan State University, United States
Reviewed by
Aaron D. Smith, Arizona State University, United States; Brian I. Crother, Southeastern Louisiana University, United States
Updates
Copyright
© 2018 Robertson, Bradler and Whiting.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: James A. Robertson erotylid@gmail.com
This article was submitted to Phylogenetics, Phylogenomics, and Systematics, a section of the journal Frontiers in Ecology and Evolution
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.