- 1Department of Biology, University of British Columbia Okanagan, Kelowna, BC, Canada
- 2Agriculture and Agrifood Canada, Lethbridge, AB, Canada
- 3TRUGen Applied Genomics Laboratory, Thompson Rivers University, Kamloops, BC, Canada
- 4Biodiversity Conservation Centre, Kings Park, WA, Australia
- 5School of Agriculture and Environment, UWA Institute of Agriculture, The University of Western Australia, Perth, WA, Australia
- 6ARC Centre for Mine Site Restoration, School of Molecular and Life Sciences, Curtin University, Perth, WA, Australia
The restoration of vegetation post-mining is particularly challenging in extreme conditions such as Mediterranean systems where soil moisture is limiting, soil temperature fluctuates dramatically, and soil carbon is very low. In such systems, soil microbial communities may play an important role in attenuating extreme conditions. Thus, vegetation establishment on such sites may be curtailed by depauperate soil communities. Soil fungal communities, in particular, are essential for nutrient turn over but we know very little about how these communities respond to mining and post-mining restoration. Fungi may be significantly affected by restoration practices. For example, the inclusion of deeper soil profiles (i.e., “overburden”) into restoration events is rare, but may expedite fungal community development. We studied a successional gradient of sand mine restoration in a former Banksia woodland in SW Australia to determine whether soil fungal communities recovered after 13 years. We also asked whether the inclusion of overburden into restoration sites improved soil fungal community development. Overall, fungal communities did not return to a pre-disturbance state by 13 years, nor did the inclusion of overburden affect their trajectory. Longer term studies are need to determine when, if ever, fungal communities are restored, and what effect this has nascent vegetation.
Introduction
Base raw material extraction, including sand and gravel, represents the largest and fastest growing share of the global mining industry (United Nations Environment Program, 2014). Sand extraction, in particular, represents a large ecological threat because sand resources are typically vegetated with unique, stress tolerant flora that have developed adaptations to local environments and may be difficult to restore (De Souza et al., 2013). Such soils are extremely low in organic material, highly leached, and can reach extremely high temperatures (Nussbaumer et al., 2016). Coadaptation between plants and soil microbes to cope with edaphic stresses is an essential part of the stability of all ecosystems, in particular these ecosystems which experience extreme environmental stress. Soil fungi, for example, have been shown to enhance plant drought tolerance (Gehring et al., 2017). Thus, rehabilitation of fungal rhizospheres may be an essential component of vegetation recovery, yet we know little about how to achieve it.
In general, mining of all types significantly reduces soil biota abundance and diversity (Ohsowski et al., 2012). Fungi are key determinants of soil physical, chemical and biological properties as they cycle nutrients, decompose organic matter, and form mycorrhizas with plants supporting their growth (De Boer et al., 2005; Gessner et al., 2010; Dickie et al., 2013). However, fungi are not considered or monitored in ecosystem restoration efforts as consistently as bacterial communities (Jones et al., 2018; Yan et al., 2018), despite the tight link between soil fungi and higher trophic levels (Stevens et al., 2018; Yan et al., 2018). Consideration of soil fungal communities is therefore an important aspect of successful sand mine restoration, but there are few guidelines for their successful restoration.
Soil fungal communities are sensitive to changes in abiotic conditions, therefore substrate conditions in a rehabilitation event may greatly affect the fungal establishment and persistence (Ohsowski et al., 2012). The most common and cost effective approach among mining operations is topsoil transfer from donor to recipient sites that have similar plant communities and ecosystem characteristics (Koch, 2007; Koch and Samsa, 2007). Direct transfer of topsoil (top 10 cm of soil profile) has been practiced for over 40 years, and while it is clearly important, it does not guarantee successful restoration (Koch, 2007). Biota are “inoculated” into the restored site via the topsoil—only if the important components of the soil food web are found within the top 10 cm of the soil profile. However, fungal taxa are well-known to be depth stratified (Rosling et al., 2003; Jumpponen and Jones, 2014), and may occur far below the top 10 cm of soil. This is particularly true in Mediterranean systems due to the great rooting depth of plants (Pickles and Pither, 2014). Unlike surface dwelling soil microbes, these fungal taxa would be less likely to disperse from neighboring, undisturbed sites, and their absence may hinder plant establishment. Including deeper layers of the soil profile in a restoration practice may improve fungal community re-establishment, and could improve restoration outcomes.
Overburden, in the context of sand mining, is the layer of substrate immediately below the top soil (10–100 cm). Overburden is typically not incorporated into restoration practice; however, it may contain a significant proportion of the rhizosphere which represents the most biologically active component in soil ecosystems (Rokich et al., 2001). For seasonally dry soils such as Banksia woodlands, where soil mining is commonly practiced, this layer can be considerably deeper than surface soils (Pickles and Pither, 2014). Failing to include this layer may delay or prevent the restoration of soil microbes, including fungi.
Presently, it is not known if incorporating rhizosphere soils into top soil improves restoration success. We evaluated fungal communities following restoration of a former sand mine location in a trial spanning 15 years, with and without the inclusion of overburden. We hypothesized that all soil fungal communities would become more similar to reference site over time, as fungal propagules from top soil transfer and dispersal from adjacent areas established over time (chronosequence study). Further, we hypothesized that the addition of overburden would hasten the recovery of soil fungal communities to reference conditions compared to plots that did not receive overburden due to inclusion of taxa not present in surface layers (overburden study).
Methods
Restoration Site
The Banksia woodlands are a floristic region in SW Australia with a Mediterranean climate and represents a global biodiversity hotspot of over 80% endemic species (Myers et al., 2000). Less than 35% of historical Banksia woodlands remain, due to large scale land clearing associated with rapid urbanization and to a lesser extent sand mining to support infrastructure development (Rokich, 2016). While much effort has been devoted to post-mine rehabilitation of this threatened ecosystem since the 1990s (Stevens et al., 2016), most restoration efforts achieve an unsatisfactory level of biodiversity return (Mueller et al., 2018).
In March 2016, soils were sampled from Hanson Construction Materials Gaskell Avenue Mine, Lexia, in southwestern Australia (32°14′35.4″S 115°46′38.5″E). The site is located in the Banksia woodlands ecosystem and has a history of 20+ years of rehabilitation. Reference soils samples were taken from sites adjacent to restored sites that were either restored using standard topsoil transfer protocols (Stevens et al., 2016), or amended with overburden. In total, we sampled 5 sites (2003, 2005, 2009, 2011, 2015), two of which had overburden amendments (2011, 2015) plus a reference site. For topsoil only (Stevens et al., 2016), fresh topsoil collected from the donor site was placed directly on top of the underlying substrate on the receptor site. For overburden plus topsoil profiles, soil 10–100 cm was freshly transferred from the intact donor site to the exposed underlying substrate of the mined receptor site, with fresh topsoil (the upper 0–10 cm) subsequently placed on top (Stevens et al., 2016). This profile mimics typical depth stratification of undisturbed reference soil profiles.
After topsoil addition, all restored sites were hand sown with a standard, locally sourced seed mix of species represented already in the seed bank (Mounsey, 2014). These included species routinely used in restoration and native to Banksia woodland such as Allocasuarina fraseriana, Allocasuarina humilis, Banksia attenuata, and Banksia menziesii and the under-story species Acacia pulchella, Beaufortia elegans, Bossiaea eriocarpa, Gompholobium tomentosum, Hovea pun-gens, Hovea trisperma, Jackso-nia densiflora, Nemcia capitata, and Xanthorrhoea preissii (Rokich et al., 2002).
Reference Site Characteristics
The dominant canopy vegetation of Banksia woodlands is comprised of B. attenuata and B. menziesii, with Eucalyptus todtiana and Nuytsia floribunda occurring less frequently (Dodd et al., 1984). The understory is represented by woody species in Myrtaceae, Epacridaceae, Proteaceae, and Papilionaceae, and non-woody species in the Anthericaceae, Stylidiaceae, Haemodoraceae, Cyperaceae, and Dasypogonaceae (Trudgen, 1997). The site is located within the Bassendean dunes, characterized by low-nutrient, leached acidic podzols with high acidity and low water holding capacities (McArthur and Bettenay, 1960; Dodd and Heddle, 1989). In general, reconstructed soils in these sites have lower organic carbon content than native woodland. At our site, reference cites have been shown to have ~0.47% organic carbon (OC), and pH of 6.1, while restored sites have 0.37% OC and pH 7.4 (please refer to Benigno et al., 2013 for more details).
Soil Sampling
Chronosequence
For the chronosequence experiment, soils were sampled from standard topsoil transfer restoration events which occurred in 2003, 2005, 2009, 2011, 2015, as well as a reference site, in a space-for-time substitution design. In all rehabilitated sites, after topsoil addition, sites were hand sown with seeds from 12 common native species and planted with seedlings of three dominant plant species (B. menziesii, B. attenuata and Eucalyptus toditiana) (Dodd et al., 1984) as well as the seed mixture (above).
Influence of Overburden Amendment
Soil samples were collected from two overburden amended sites (2011 and 2015) that also received standard topsoil addition. In addition, we sampled from the adjacent undisturbed site which included reference topsoil (top 10 cm) and reference overburden (10–100 cm). At each site, we randomly sampled five soil cores within a 10 m2 plot. Each soil core was sampled adjacent to an Acacia pulchella shrub to standardize for differences in soil fungal composition. We chose to sample rhizosphere soil of Acacia pulchella because it is native to the area and occurred in all plots. Soils were then stored at 4°C and shipped to UBC Okanagan where DNA was extracted from soils using MoBio Powersoil (MAX, Qiagen Inc.).
Fungal DNA Isolation and Identification
Amplicon Preparation
Fungal ITS amplicons were generated using gITS86F (5′ GTGARTCATCGARTCTTTGAA) and ITS4r (5′ TCCTCCGCTTATTGATATGC) (Vancov and Keen, 2009), and sequencing adaptor barcoded gITS86F and ITS4r primers in the second round. All reactions contained 10 μl of 2X GoTaq Green Master Mix (Promega Corporation, Madison, WI), 2 μl forward primer (1 μM final concentration), 2 μl reverse primer (1 μM final concentration), 1 μl of template DNA, and water for a total volume of 20 μl. The thermocycler program for first round bacterial and fungal amplicon generation consisted of: 95°C for 4 min followed by 25 cycles of 95°C for 30 s, 53.4°C for 45 s, and 72°C for 2 min, with a final extension at 72°C for 5 min. For generating sequencing barcode adapted bacterial and fungal amplicons, the thermocycler program was the same except that 20 cycles were used with an annealing temperature of 65°C. After each round of PCR, amplicons were cleaned using an E.Z.N.A. Cycle Pure Kit (Omega Bio-Tek, Norcross, GA), quantified using a Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific, Mississauga, ON), and visualized on an agarose gel. Template-free and positive controls were included for all reactions.
Sequencing
Barcoded amplicons were pooled in equimolar amounts and purified following agarose gel electrophoresis using an E.Z.N.A Gel Extraction Kit (Omega Bio-Tek) and, prior to sequencing on an Ion Torrent PGM (Life Technologies Inc., Carlsbad, CA), the library dilution factor was determined using an Ion Library Quantitation Kit. An Ion PGM Template Hi-Q OT2 400 kit on an Ion OneTouch 2 system was used to prepare DNA for sequencing, and enriched Ion Sphere Particles were quantified using an Ion Sphere Quality Control Kit. Sequencing was performed on a 316v2 chip with an Ion PGM Hi-Q View Sequencing Kit. Sequences were deposited in European Nucleotide Archive (accessions ERS3048307-ERS3048349). Demultiplexed FASTQ files were generated using Torrent Suite 5.0.5. Split_libraries_fastq.py in QIIME (v 1.9.1; Caporaso et al., 2010) was used to merge reads into a single FASTA file (phred quality score Q20, 3 low quality bases read truncation, zero Ns allowed, phred offset of 33). Qiime was used for open reference operational taxonomic unit (OTU) picking (pick_open_reference_otus.py) and generating summary plots (summarize_taxa_through_plots.py). Illegal characters were removed from the merged FASTA file before using the PIPITS pipeline (Gweon et al., 2015) for dereplication, removal of unique sequences, OTU picking, chimera removal, and taxonomy assignment with the pipits_funits and pipits_process commands.
Data Analysis
Here we use the term phylotype to describe OTUs for the fungi identified using the above methodology. Differences in fungal phylotypes were assessed using parametric and multi-response permutation procedure (MRPP) using Bray-Curtis similarity matrices. To determine differences in phylotype richness, data were Box Cox transformed (MASS, Venables and Ripley, 2002) prior to ANOVA with site age as a fixed factor. MRPP is a permutation procedure, this process compares the occurrence of phylotypes in each soil group to multiple randomizations of the entire data set, determining whether the pattern observed is likely different to random. An A-value of than 0.3 is considered a significant effect size for ecological data (McCune and Grace, 2002). All values were FDR (false-discover rate) corrected to account for multiple comparisons. Communities were also compared using permANOVA in Primer-E v6 (Anderson, 2001, Clarke and Gorley, 2006) to assess community level differences among years and overburden treatments, and to test for differences in dispersion. PERMADISP values are given for each comparison, indicating, when significant, that differences detected in the PERMANOVA are due to dispersion (heterogeneity) of the data rather than differences in fungal community structure. Indicator species analysis, a permutational procedure which determines which species occur in one soil type to the exclusion of all other treatments was performed in PC-ORD (McCune and Grace, 2002). All analyses were performed in R unless otherwise indicated (R 3.4.2, R Core Team, 2017).
Results
Chronosequence Study
Did soil fungal communities become more similar to reference site over time?
Fungal communities did not become more similar to reference sites over time, but reference soils were distinct from all other treatments (Tables S1). Alpha diversity was highest in the most recently restored sites (2015) (p < 0.05), and lowest in the oldest rehabilitation site (Figure 1). Concurrently, the proportion of phylotypes shared between with reference site also decreased over time (Figure 2). The most recently restored site shared the most (70%), while the oldest site (2003) shared the least (35%).
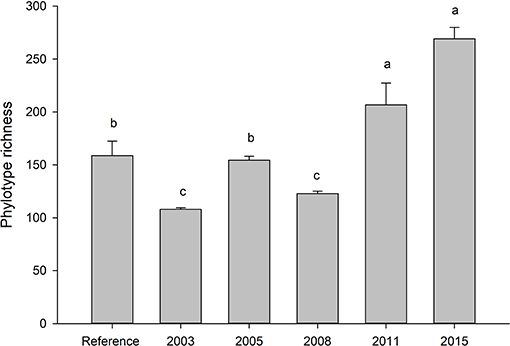
Figure 1. Average (SE) number of fungal phylotypes across the restoration chronosequence. Letters indicate significance at p < 0.05 (Tukey). Values represent a mean of five samples. Ref indicates soils taken from an adjacent, undisturbed “reference” site.
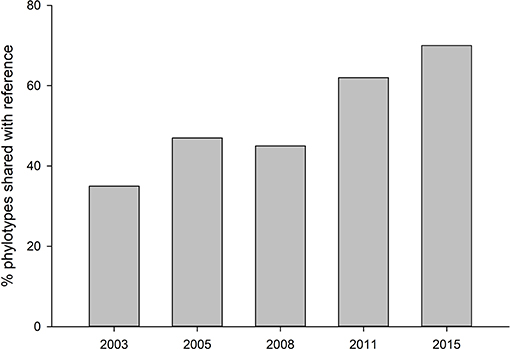
Figure 2. Proportion of fungal phylotypes shared with reference site across the restoration chronosequence.
Across the chronosequence, fungal communities were significantly different (p = 0.001) (Figure 3). The reference site was least similar to 2015 (Bray Curtis similarity 47.2) and most similar to 2003 (44.1), 2005 (44.2), and 2011 (44.3). Thus, there was no directional response for increasing similarity to reference site over time. In fact, variation within sites was often higher than between sites. The reference site had lowest within samples homogeneity (55.8) compared to all restored sites, meaning 2003 and 2008 were significantly less dispersed than the reference site (p = 0.02 and 0.08, respectively; Table S1).
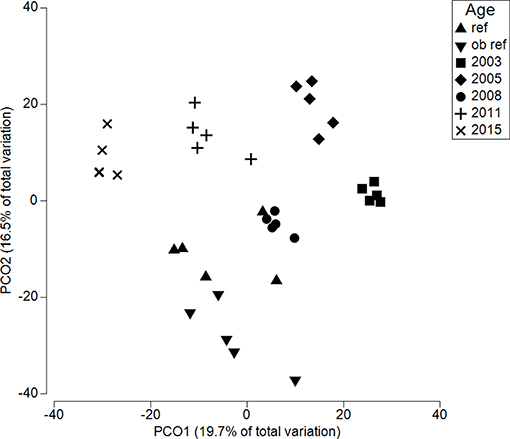
Figure 3. Principle coordinate analysis for soils along a restoration chronosequence. Each symbol represents the fungal community within a single soil core. Ref refers to an adjacent, undisturbed site. All sites are significantly different from each other (p < 0.0001).
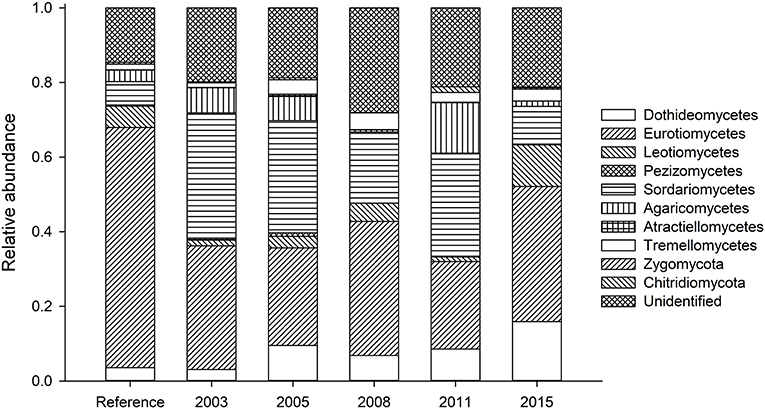
In terms of composition, restored soils had decreased Eurotiomycetes (opportunistic plant pathogens) and increased Sordariomycetes (saprobes and plant pathogens) compared to reference soils (Figure 4, Table S3). Relative abundance of Agaricomycetes (saprobes and mycorrhizal fungi) varied between soils (Figure S1). The 2011 soils show a spike in a phylotype identified only as Agaricaceae (Figure S1a). This phylotype represents 8.3% of all sequences recovered for the 2011 soils. For the Dothidiomycetes (plant endophytes and pathogens; Figure S1b), younger restored soils show more sequences from this Phylum, particularly those identifying as Capnodiales (some plant pathogens), Tubeufiaceae (plant pathogens), and Pleosporales (sapbrobes/endophytes/parasistes). Eurotiomycetes (Figure S1c) represent approximately 1/3 of all sequences recovered. These fungi were dominated largely by Aspergillus (saprobe), Trichocomaceae (saprobes found in extreme conditions), and Chaetothyriales (extremetolerant, saprobe/biotroph).
Many phylotypes detected in the reference soils were much less abundant in the restored soils, Leotiomycete (plant pathogens) phylotype relative abundance appeared to recover by 2008 (Figure S1d), with, and Oidiodendron (saprobe) sp. recovering but Arachnopeziza (saprobe) disappearing by 2015. In the Sordariomycetes (Figure S1e), a phylotype as Sordariales sp. was dominant in all soil, but less abundant in reference soils which was dominated by Coniochaeta (tree pathogens). Tremellomycetes (saprobe) was represented by only three phylotypes, but in different relative abundance to the reference site, by 2003 (Figure S1f).
Overburden Study: Does the Inclusion of Overburden Increase the Similarity of Fungal Communites to Reference Site?
Both years, regardless of overburden amendment differed in terms of phylotype richness (p = 0.04), and sites were significantly different from each other (p = 0.001). The phylotype richness was highest in the most recently restored sites with overburden (271) but overburden amended sites did not differ from topsoil only sites for 2011, 2015, or reference site (p > 0.05) (Figure 5). The proportion of phylotypes shared with reference site was similar for overburden treatments in 2011 (62% overburden vs. 64%) and 2015 (75% overburden vs. 74%) (Figure 6).
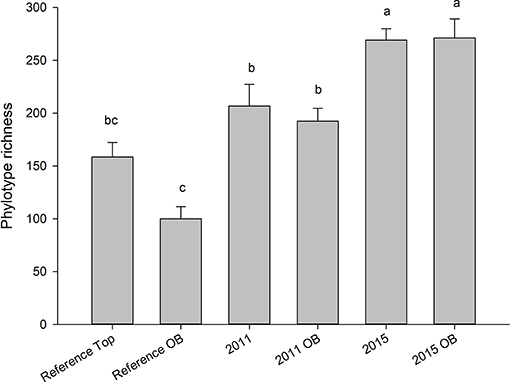
Figure 5. Average (SE) number of fungal phylotypes in sites receiving both standard topsoil transfer and overburden (OB). Letters indicate significance at p < 0.05 (Tukey). Values represent a mean of five samples. Ref indicates soils taken from an adjacent, undisturbed “reference” site.
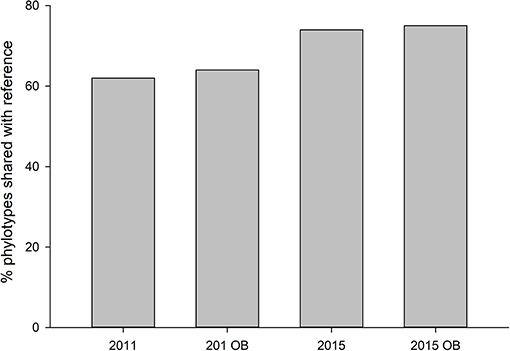
Figure 6. Proportion of fungal phylotypes shared with reference site in sites receiving both standard topsoil transfer and overburden (OB).
Overburden inclusion significantly change fungal community composition (p = 0.001) (Figure 7) but there was also an interaction between overburden and time of restoration (p = 0.01). Overall, top soils were more similar to each other than overburden soil (Bray Curtis similarity 50.8 (top) vs. 46.9 (overburden). This was most pronounced for reference sites (Bray Curtis similarity top 55.8 vs. 45.6 overburden) and less for 2015 sites [BC similarity 57.1 (top) vs. 54.2 (overburden)]. All sites showed significant pairwise differences among overburden and topsoil only amendments (see Table S2). Both reference and 2011 sites had similar levels of phylotype turnover (dispersion) when comparing overburden to topsoil only sites (p = 0.82 reference, p = 0.93 2011), but 2015 overburden sites had more turnover than 2015 topsoil only sites (p = 0.07; Table S4, Figure 7).
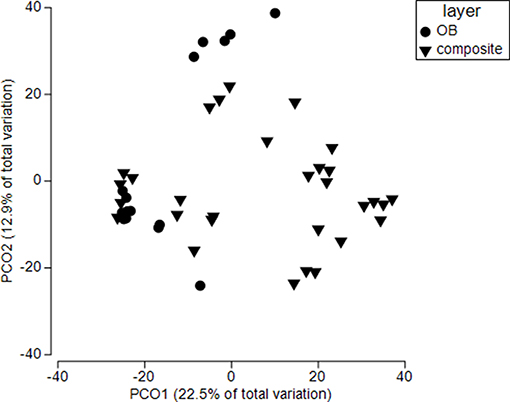
Figure 7. Principle coordinate analysis for soils in sites receiving both standard topsoil transfer and overburden (OB). Each symbol represents the fungal community within a single soil core. Ref refers to an adjacent, undisturbed site. All sites are significantly different from each other (p < 0.0001), but sites responded differently to overburden treatment (site X overburden, p < 0.05). See Table S2 for more details.
In terms of composition, there were few differences among overburden amended soils and those with topsoil only (Figure 8). The mycorrhizal fungus Pisolithus was more abundant in overburden reference soils. Also, Gymnopilus allantopus (saprobe) was more abundant in overburden soils (2011) (Figure S2a). Ceraporia, a crust fungus, was extremely common in 2011 (topsoil only). There were fewer Dothidiomycetes in the reference overburden soil than the reference topsoil, and its abundance increased in restored soils (Figure S2b). Eurotiomycetes, on the other hand, were found more often in the reference soils relative to the restored soils (Figure S2c). Particularly Chaetothriales (non-lichen epiphytes), which comprised 30% of reference overburden samples. Leohumicola, a heat tolerant crust fungus, was an important component of the Leotiomycetes for reference overburden, but not top soil (Figure S2d). Reference overburden had among the lowest abundance of Sodiaromycetes (Figure S2e). Tremellomyctes were almost completely absent from reference overburden, but common in other sites particularly 2015 (Figure S2f).
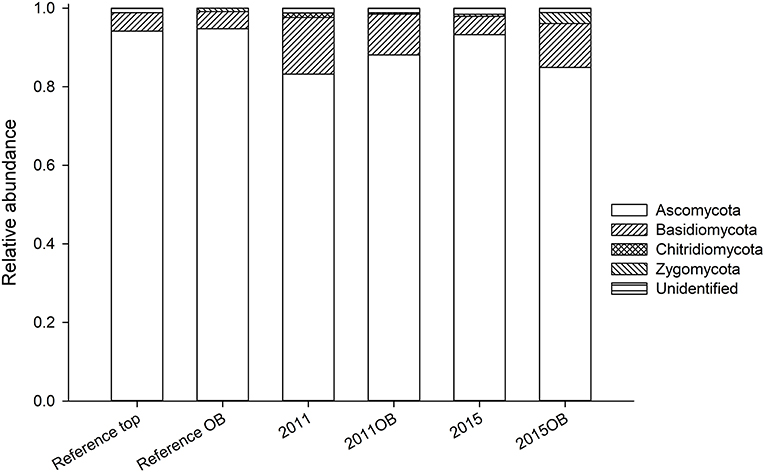
Figure 8. Relative abundance of fungal phyla in samples receiving topsoil transfer compared to those receiving overburden.
Discussion
We investigated the recovery of soil fungal communities post-sand mining over time in former Banksia woodlands in SW Australia. After 13 years, restored soil fungal communities were significantly different from the reference site, indicating a lack of recovery. As we hypothesized, the addition of overburden significantly changed fungal communities, but, contrary to our hypothesis, did not result in a faster recovery back to the reference state.
Were Older Sites Most Similar to Reference Sites?
We found little evidence that rehabilitated soils were approaching a reference state even 13 years post-rehabilitation. One study in a nearby system showed that the soil fungal community did not return to pre-bauxite mining conditions after 18 years (Banning et al., 2011), suggesting that our results are consistent for that area. In our study, recently transferred soils harbored significantly more phylotypes than reference soils, whereas the oldest sites had the fewest. The higher number of phylotypes early in succession could be due to greater fluctuations in the types of food available for the fungi (Frankland, 1998). Or it may reflect the overall species pool on which environmental filtering has yet to select suitable taxa. If species richness is due to a large species pool, our data support heavy environmental filtering which resulted in the lowest phylotype richness overall by 2003. Studies have shown decreased soil biodiversity and functioning post-mining (Gil-Sotres et al., 1992), while others show no change in richness (Yan et al., 2018). In this study it is not clear why numbers dropped below reference levels but indicates that either the community is still in flux or that reduced substrate complexity cannot support the same level of fungal diversity. Given that beta-diversity was also lowest in the oldest site (2013) and highest in the reference site, indicates a lack of complexity in restored sites may take a very long time to reach reference levels, if ever.
However, ecosystems rarely recover to reference conditions. In a meta-analysis of 400 studies, Jones et al. (2018) report a rate of recovery of 1.1–10% per year (with a median of 2.9%), with forest vegetation recovering most slowly than other ecosystems, and mining restoration recovery falling near the median of disturbance types considered. The relationship between soil community richness and ecosystem functions is well-demonstrated (i.e., van der Heijden et al., 2008), but it is not yet clear if soil “recovery” can be evaluated through measures of soil microbial community metrics such as alpha-diversity or composition. Whether such soils achieve complete recovery may not be necessary for vegetation success, but this remains to be shown over long-term experiments which are lacking. A more appropriate recovery time-frame might be decades, rather than years.
Did Overburden Change Recovery Trajectory?
We found no evidence for our hypothesis that overburden amendments would improve fungal community recovery. The reasons for this are not clear, and there has been no other investigations of the effect of overburden on soil fungal communities. In our study, soil collected from the “overburden” layer in reference sites differed from reference top soil, both in terms of phylotype richness and community composition. Contrary to our hypothesis, however, overburden reference soil was less diverse than reference top soil. This was surprising, given the lack of organic material and extreme temperatures of the top soil.
Whether our results reflect a general trend among dune soil systems is not clear. Little is known about fungal diversity in extreme soils (such as those found in Banksia woodlands), but fungal communities in other systems have been shown to be stratified based on soil depth (Jumpponen et al., 2010). Plantss in arid ecosystems tend to root deeper, on average, than in temperate or tropical climates (Canadell et al., 1996; Pickles and Pither, 2014), thus we would expect increased fungal activity in deeper, rhizosphere zones. We found some support for this with Pisolithus, a mycorrhizal fungus, found predominately in the reference overburden, and almost completely absent in the topsoil. Given that ectomycorrhizal plants show strong host preference (Dickie, 2007), lack of suitable host may prevent some plants from establishing. In our study, we identified indicator fungi unique to reference topsoil, including oily yeasts, non-lichenised rock inhabiting fungi, and fungi previously isolated or identified from challenging environments such as deserts or industrial tailings (Table 1). Indicator fungi unique to overburden soils were also mostly saprobes, yeast-like, and extremophilic, but contained two putative mycorrhizal fungal species. Restoration practices which increase the biodiversity of symbiotic fungi may hasten revegetation (Harris, 2009). To fully understand the importance of fungal taxa in restored soils, future studies must evaluate the relationship between fungal composition and plant establishment over long time scales.
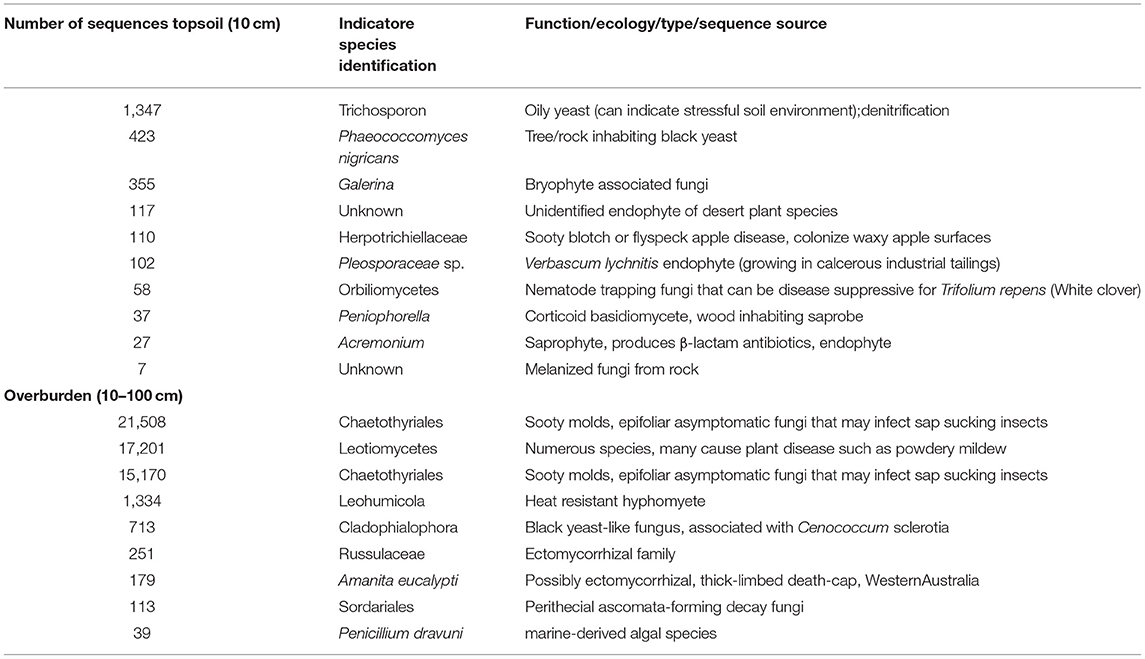
Table 1. Inferred ecological function based on indicator species identity (species found in these samples to the exclusion of others) found in undisturbed reference overburden and topsoil.
Conclusion
Restoration of soil fungal communities in a former sand mine did not recover to reference conditions over a 13-year time period. The inclusion of overburden in the restoration profile resulted in changes to the fungal community which may, over time, lead to different trajectories. However, given the paucity of data concerning the restoration of soil microbial communities in ecosystem restoration, these changes may not affect vegetation restoration. This study highlights the need for exemplar restoration projects that include appropriate balanced design and multiple suitable reference sites. Such sites will enable rigorous tests of key ecological management questions and are critical in ecological restoration research, where imperfect study systems are often the location of research simply because they are all that exist (Prober et al., 2019). Good design of monitoring programs will also be important for stakeholders (e.g., mining companies) that need to demonstrate restoration success to regulators.
Ultimately, evaluating fine-scale changes to soil microbial communities may not be the most effective predictor for restoration success. It may be that soil microbial communities are functional even if they do not return to a “reference state.” Thus, studies should also consider functional responses, such as microbial biomass, substrate utilization and nutrient cycling. Longer experiments are needed, particularly in stressful conditions such as the Banksia woodland, where recovery may take longer than in mesic systems. Successful restoration efforts depend on additional research, particularly for understudied vegetation types and climates.
Data Availability Statement
The sequence data used to support the findings of this study have been deposited in the European Nucleotide Archive (https://www.ebi.ac.uk/ena).
Author Contributions
MH, PN, and JS conceived of study. MH and PN collected field samples. BM and JV analyzed data. MG, LA, AW, MH, PN, JS, and JV wrote manuscript.
Funding
Research was funded by the Gledden Foundation and the Institute for Advanced Studies University of Western Australia.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
MH was funded by the Robert and Maude Gledden Visiting Fellowship. PN was funded by the Australian Government through the Australian Research Council Industrial Transformation Training Centre for Mine Site Restoration (project number ICI150100041).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/ffgc.2019.00078/full#supplementary-material
References
Anderson, M. J. (2001). A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46. doi: 10.1111/j.1442-9993.2001.01070.pp.x
Banning, N. C., Gleeson, D. B., Grigg, A. H., Grant, C. D., Andersen, G. L., Brodie, E. L., et al. (2011). Soil microbial community successional patterns during forest ecosystem restoration. Appl. Environ. Microbiol. 77, 6158–6164. doi: 10.1128/AEM.00764-11
Benigno, S. M., Dixon, K. W., and Stevens, J. C. (2013). Increasing soil water retention with native-sourced mulch improves seedling establishment in postmine Mediterranean sandy soils. Restor. Ecol. 2, 617–626. doi: 10.1111/j.1526-100X.2012.00926.x
Canadell, J., Jackson, R. B., Ehleringer, J. B., Mooney, H. A., Sala, O. E., and Schulze, E. D. (1996). Maximum rooting depth of vegetation types at the global scale. Oecologia 108, 583–595. doi: 10.1007/BF00329030
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. doi: 10.1038/nmeth.f.303
De Boer, W., Folman, L. B., Summerbell, R. C., and Boddy, L. (2005). Living in a fungal world: Impact of fungi on soil bacterial niche development. FEMS Microbiol. Rev. 29, 795–811. doi: 10.1016/j.femsre.2004.11.005
De Souza, R. F., Da Silva, K. A., De Mello, C. M. A., Goto, B. T., Da Silva, F. S. B., Sampaio, E. V. S. B., et al. (2013). Arbuscular mycorrhizal fungi in revegetated mine dunes. Land Degred. Dev. 24, 147–155. doi: 10.1002/ldr.1113
Dickie, I. A. (2007). Host preference, niches and fungal diversity. N. Phytol. 174, 230–233. doi: 10.1111/j.1469-8137.2007.02055.x
Dickie, I. A., Martínez-García, L. B., Koele, N., Grelet, G.-A., Tylianakis, J. M., Peltzer, D. A., et al. (2013). Mycorrhizas and mycorrhizal fungal communities throughout ecosystem development. Plant Soil 367, 11–39. doi: 10.1007/s11104-013-1609-0
Dodd, J., and Heddle, E. (1989). Water relations of Banksia woodlands. J. R. Soc. Western Austral. 71, 91–92.
Dodd, J., Heddle, E. M., Pate, J. S., and Dixon, K. W. (1984). Rooting patterns of sandplain plants and their functional significance, in Kwongan, Plant Life of the Sandplain: Biology of a South-West Australian Shrubland Ecosystem. eds J. S. Pate and J. S. Bear (Nedlands, WA: University of Western Australia Press).
Frankland, J. (1998). Fungal succession—unravelling the unpredictable. Mycol. Res. 102, 1–15. doi: 10.1017/S0953756297005364
Gehring, C. A., Sthultz, C. M., Flores Renteria, L., Whippple, A. V., and Whitham, T. G. (2017). Tree genetics defines fungal partner communitis that may confer drought tolerance. Proc. Natl. Acad. Sci. U.S.A. 114, 11169–11174. doi: 10.1073/pnas.1704022114
Gessner, M. O., Swan, C. M., Dang, C. K., McKie, B. G., Bardgett, R. D., Wall, D. H., et al. (2010). Diversity meets decomposition. Trends Ecol. Evol. 25, 372–380. doi: 10.1016/j.tree.2010.01.010
Gil-Sotres, F., Trasar-Cepeda, C. C., Ceccanti, B., and Leiros, M. C. (1992). Biochermical characterization of biological activity in very young mine soils. Biol. Fert. Soils 12, 25–30. doi: 10.1007/BF00337233
Gweon, H. S., Oliver, A., Taylor, J., Booth, T., Gibbs, M., Read, D. S., et al. (2015). PIPITS : an automated pipeline for analyses of fungal internal transcribed spacer sequences from the Illumina sequencing platform. Methods Ecol. Evol. 6, 973–980. doi: 10.1111/2041-210X.12399
Harris, J. (2009). Soil microbial communities and restoration ecology: facilitators or followers? Science 325, 573–574. doi: 10.1126/science.1172975
Jones, H. P., Jones, P. C., Barbier, E. B., Blackburn, R. C., Benayas, J. M. R., Holl, K. D., et al. (2018). Restoration and repair of Earth's damaged ecosystems. Proc. Biol. Sci. 285:20172577. doi: 10.1098/rspb.2017.2577.
Jumpponen, A., Jones, K., and Blair, J. (2010). Vertical distribution of fungal communities in tallgrass prairie soil. Mycologia 102, 1027–1041. doi: 10.3852/09-316
Jumpponen, A., and Jones, K. L. (2014). Tallgrass prairie soil fungal communities are resilient to climate change. Fungal Ecol. 10, 44–57.
Koch, J. M. (2007). Alcoa's mining and restoration process in South Western Australia. Restor. Ecol. 15, 11–16. doi: 10.1111/j.1526-100X.2007.00288.x
Koch, J. M., and Samsa, G. P. (2007). Restoring jarrah forest in southwestern Australia after bauxite mining. Restor. Ecol. 15, S17–S25. doi: 10.1111/j.1526-100X.2007.00289.x
McArthur, W. M., and Bettenay, E. (1960). The Deverlopment and Distribution of the Soils of the Swan Cosyal Plain, Western Australia. CSIRO.
McCune, B., and Grace, J. (2002). Analysis of Ecological Communities. Gleneden Beach, OR: MjM Software Design.
Mounsey, C. M. (2014). A compositional, functional, and structural assessment of a 20 year post-mine restoration chronosequence. (PhD dissertation), University of Western Australia, Perth, WA, Australia.
Mueller, A. L., Canham, C. A., van Etten, E. J. B., Stock, W. D., and Forend, R. H. (2018). Using a functional ecology approach to assist plant selection for retoration of Mediterranean woodlands. For. Ecol. Manag. 424, 1–10. doi: 10.1016/j.foreco.2018.04.032
Myers, N., Mittermeiter RA Mittermeier, C. G., da Fonseca, G. A. B., and Kent, J. (2000). Biodiversity hotspots for conservation priorities. Nature 403, 853–858. doi: 10.1038/35002501
Nussbaumer, Y., Cole, M. A., Offler, C. E., and Patrick, J. W. (2016). Identifying and ameliorating nutrient limiatiaons to reconstructin a forest ecosystem on mined land. Restor. Ecol. 24, 202–211. doi: 10.1111/rec.12294
Ohsowski, B. M., Klironomons, J. N., Dunfield, K. E., and Hart, M. M. (2012). The potential of soil amendments for restoring severely disturbed grasslands. Appl. Soil Ecol. 60, 7–83. doi: 10.1016/j.apsoil.2012.02.006
Pickles, B. J., and Pither, J. (2014). Still scratching the surface: how much of the ‘black box’of soil ectomycorrhizal communities remains in the dark? N. Phytol. 201, 1101–1105. doi: 10.1111/nph.12616
Prober, S. M., Doerr, V. A. J., Broadhurst, L. M., Williams, K. J., and Dickson, F. (2019). Shifting the conservation paradigm: a synthesis of options for renovating nature under climate change. Ecol. Monogr. 89:e01333. doi: 10.1002/ecm.1333
R Core Team (2017). R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing. Available online at: http://www.R-project.org/ (accessed November 27, 2017).
Rokich, D. P. (2016). Melding of research and practise to improve restoration of Banskia woodlands after sand extraction Perth WA. Ecol. Manag. Restor. 17, 112–122. doi: 10.1111/emr.12214
Rokich, D. P., Dixon, K. W., Sivasithamparam, K., and Meney, K. A. (2002). Smoke, mulch, and seed broadcasting effects on woodland restoration in Western Australia. Restor. Ecol. 10, 185–194. doi: 10.1046/j.1526-100X.2002.02040.x
Rokich, D. P., Meney, M. A., Dixon, K. W., and Sivasithampram, K. (2001). The impact of soil disturbance on root development in woodland communities of Western Australa. Austr. J. Botany 49, 169–183. doi: 10.1071/BT00015
Rosling, A., Landeweert, R., Lindahl, B. D., Larsson, K.-H., Kuyper, T. W., Taylor, A. F. S., et al. (2003). Vertical distribution of ectomycorrhizal fungal taxa in a podzol soil profile. N. Phytol. 159, 775–783. doi: 10.1046/j.1469-8137.2003.00829.x
Stevens, B. M., Propster, J., Wilson, G. W. T., Abraham, A., Ridenour, C., Doughty, C., et al. (2018). Mycorrhizal symbioses influence the trophic structure of the Serengeti. J. Ecol. 106, 536–546. doi: 10.1111/1365-2745.12916
Stevens, J. C., Rokich, P. D., Newton, V. J., Barrett, R. L., and Dixon, K. W. (2016). Banksia Woodlands: A Restoration Guide for the Swan Coastal Plain (UWA Publishing), 353.
Trudgen, M. (1997). A Report on the Flora and Floristic Community Types Found on Parts of Sand Mining Leases Held by Rocla Inthe Gnangara Area. Report prepared for Rocla Quarry Products, Perth, WA.
United Nations Environment Program (2014). Sand, Rarer Than One Thinks. Global Environmental Alert Service. Available online at: http://hdl.handle.net/20.500.11822/8665 (accessed February 24, 2019).
van der Heijden, M. G., Bardgett, R. D., and van Straalen, N. M. (2008). The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol. Lett. 11, 296–310. doi: 10.1111/j.1461-0248.2007.01139.x
Vancov, T., and Keen, B. (2009). Amplification of soil fungal community DNA using the ITS86F and ITS4 primers. FEMS Microbiol. Lett. 296, 91–96. doi: 10.1111/j.1574-6968.2009.01621.x
Venables, W. N., and Ripley, B. D. (2002). Modern Applied Statistics with S, 4th Edn. New York, NY: Springer.
Keywords: fungi, sand mine, rehabilitation, mycorrhizas, plant pathogens, extreme conditions
Citation: Hart M, Gorzelak MA, McAmmond BM, Van Hamme JD, Stevens J, Abbott L, Whiteley AS and Nevill P (2019) Fungal Communities Resist Recovery in Sand Mine Restoration. Front. For. Glob. Change 2:78. doi: 10.3389/ffgc.2019.00078
Received: 02 September 2019; Accepted: 04 November 2019;
Published: 26 November 2019.
Edited by:
Stephanie N. Kivlin, The University of Tennessee, Knoxville, United StatesReviewed by:
Michala Phillips, United States Geological Survey (USGS), United StatesKendall Beals, The University of Tennessee, Knoxville, United States
Catherine Zabinski, Montana State University, United States
Copyright © 2019 Hart, Gorzelak, McAmmond, Van Hamme, Stevens, Abbott, Whiteley and Nevill. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Miranda Hart, bWlyYW5kYS5oYXJ0QHViYy5jYQ==
 Miranda Hart
Miranda Hart Monika A. Gorzelak
Monika A. Gorzelak Breanne M. McAmmond3
Breanne M. McAmmond3 Lynette Abbott
Lynette Abbott Andrew S. Whiteley
Andrew S. Whiteley Paul Nevill
Paul Nevill