- 1UK Centre for Ecology & Hydrology, Lancaster Environment Centre, Lancaster, United Kingdom
- 2Lancaster Environment Centre, Lancaster University, Lancaster, United Kingdom
- 3Institute of Biological and Environmental Sciences, University of Aberdeen, Aberdeen, United Kingdom
- 4Environmental and Rural Science, University of New England, Armidale, NSW, Australia
- 5Centre for Ecology & Hydrology, Wallingford, United Kingdom
- 6Forest Research Centre, Sandakan, Malaysia
Industrial logging and agricultural expansion are driving rapid transformations of tropical ecosystems, modifying patterns in above-ground plant and below-ground microbial communities. However, the extent to which these changes in biodiversity drive modifications of ecosystem process rates such as leaf litter decomposition is poorly understood. To determine the relative effects of changes to the chemical quality of litter and shifts in microbial decomposers on leaf litter decomposition rates, we performed a controlled, reciprocal transplant, litter decomposition experiment across a tropical land-use disturbance gradient. Litter mixtures and soils were collected from old growth forest, moderately logged forest, heavily logged forest, and oil palm plantation in Sabah, Malaysia, and combined in a fully crossed, factorial microcosm experiment maintained under controlled environmental conditions. We found that whilst litter quality was the most important predictor of litter mass loss, soil origin was also significant, explaining between 5.17 and 15.43% of total variation. Microbial decomposer communities from old growth forest had greater functional breadth relative to those from logged forests and oil palm plantation as all litter types decomposed faster when combined with old growth soil. The most chemically recalcitrant litter (lowest N, highest lignin, lignin:N, and C:N ratio) from moderate logged forest decomposed faster when combined with its “home” soil (Home-Field Advantage) whilst the most labile litter from oil palm decomposed slowest when combined with its “home” soil. This was correlated with lower total soil microbial biomass. Taken together, these findings demonstrate that whilst litter quality regulated rates of litter decomposition across the disturbance gradient, soil microbial decomposer communities were functionally dissimilar between land uses and explained a significant proportion of variation. The impact of disturbance on soil, including microbial community structure, should be considered alongside changes to plant communities when assessing effects on crucial ecosystem processes such as decomposition.
Introduction
The decomposition of non-living organic matter is a fundamental process in the carbon cycle, releasing CO2 whilst recycling nutrients to support ecosystem productivity. The chemical quality of organic matter, abiotic environmental conditions, and microbial decomposer communities all influence decomposition dynamics (Currie et al., 2010; Makkonen et al., 2012; Bradford et al., 2016). Microclimate and litter traits are widely considered as key constraints regulating rates of litter decomposition (Meentemeyer, 1978; Gholz et al., 2000; Cornwell et al., 2008; Wall et al., 2008; Zhang et al., 2008; Makkonen et al., 2012; Bradford et al., 2014; Djukic et al., 2018). However, litter decomposition is ultimately a microbially-mediated process. Therefore, the abundance and structure of soil decomposer communities may also drive local variation in rates of decomposition, but the role of microbial community composition is currently not well-understood (Bradford et al., 2017). Identification of the factors regulating litter decomposition rates is necessary to inform biogeochemical models on how ecosystem organic matter stocks and thus the magnitude of biosphere-atmosphere feedbacks will respond to anthropogenic disturbance and future environmental change (Bonan et al., 2013).
Globally, tropical forests are threatened by industrial logging, fire, and deforestation for agricultural expansion with degraded and secondary forest now comprising over half of all remaining tropical forests (Laurance et al., 2014; Potapov et al., 2017). Southeast Asia is a deforestation hotspot, driven primarily by agricultural expansion of oil palm and commercial timber plantations (Gibbs et al., 2010; Estoque et al., 2019). Remaining lowland tropical forests in the region are heavily degraded, due in part to the high density of commercially valuable, dipterocarp timber species (Achard et al., 2002; Sodhi et al., 2004). Degradation of tropical forests modifies regional above-ground species distributions and community composition across the tropics, driving changes to functional diversity, and leaf litter traits (Bakker et al., 2011; Baraloto et al., 2012; Carreño-Rocabado et al., 2012; Both et al., 2019; Ding et al., 2019). In South East Asian dipterocarp forests, logging disturbance appears to shift tree communities from those with resource conservative traits, prioritizing structural and defense tissues to those with more resource acquisitive traits promoting carbon capture and growth (Both et al., 2019). Given the important role of leaf traits in determining litter quality, this shift may drive changes to rates of litter decomposition within tropical forests (Cornwell et al., 2008). However, evidence from a field experiment showed that subtle differences in microclimate rather than litter quality explained most variation in litter decomposition rates (Both et al., 2017). In contrast to tropical forests where litter inputs are continuous and distributed across the forest floor, oil palm plantation litter is comprised primarily of senesced fronds. These are typically pruned manually twice a year and green fronds may also be pruned during harvesting to access fruit bunches (Corley and Tinker, 2015). These are stacked in piles or rows, therefore significant aboveground C inputs occur only within the vicinity of these frond piles in mature plantations (Ruegg et al., 2019). Leguminous cover crops may also be planted in in young plantations to stabilize soils and improve fertility (Luke et al., 2019).
Tropical land-use change can also drive parallel shifts in soil microbial decomposer communities (Tripathi et al., 2012, 2016; Rodrigues et al., 2013; Kerfahi et al., 2014; McGuire et al., 2015; Mueller et al., 2016; Shi et al., 2019). Theory proposes that the immense diversity of microbial communities should confer a high level of functional redundancy, particularly with regard to bulk processes such as litter decomposition (Nannipieri et al., 2003; Allison and Martiny, 2008). However, experimental work suggests that changes to bacterial and fungal community composition can alter rates of litter decomposition (Strickland et al., 2009b; Allison et al., 2013; Cleveland et al., 2014; Martiny et al., 2017). The specific mechanisms underpinning functional dissimilarity are not well-understood, but two hypotheses have been proposed. The functional breadth hypothesis (FBH) states that decomposer communities from recalcitrant litter environments have wider functional abilities. This confers the capability to degrade a wide range of substrates efficiently, at similar rates with low enzyme activities and microbial biomass per unit of decomposition (Van Der Heijden et al., 2008; Keiser et al., 2011). FBH has been demonstrated with decomposer communities from forest habitats decomposing low quality litter (e.g., low N, high lignin content, lignin:N, and C:N ratios) faster than communities from other ecosystems whilst high quality litter was decomposed equivalently across ecosystems (Strickland et al., 2009a,b; Fanin et al., 2016). Alternatively, the home-field advantage hypothesis (HFA) states that litter decomposes more rapidly in an environment dominated by, or in the vicinity of the species it is derived from (Home) rather than in a foreign environment (Away) due to functional specialization of decomposer communities (Gholz et al., 2000). HFA has been demonstrated in numerous ecosystems but is not universal and appears to be highly context dependent (Ayres et al., 2009b; Wang et al., 2013; Veen et al., 2018; Lin et al., 2019). It has also been shown that the HFA effect becomes more pronounced as home and away litters become more dissimilar (Veen et al., 2015).
In South East Asian dipterocarp forests, most trees, and plants form arbuscular mycorrhizal associations but the canopy is often dominated by a subset of commercially valuable tree species from the Dipterocarpaceae family, all of which form ectomycorrhizal (EcM) associations (Brearley, 2012). Selective removal of these tree species can lead to a reduction in abundance and shifts in community composition of EcM fungi (Kerfahi et al., 2014; McGuire et al., 2015), which may influence decomposition processes through changes to competitive dynamics with saprotrophic fungi (Fernandez and Kennedy, 2016; Corrales et al., 2018). In oil palm plantations, roots are heavily colonized by arbuscular mycorrhizal fungi (AMF) (Phosri et al., 2010), whilst EcM fungi are almost absent (McGuire et al., 2015). Although AMF are not thought to have saprotrophic capabilities, they can enhance rates of litter decomposition (Gui et al., 2017).
To test whether human modification of tropical ecosystems altered the functional capabilities of microbial decomposer communities, and to quantify the relative effects of litter quality and microbial decomposers on litter decomposition rates, we conducted a controlled decomposition experiment with reciprocal litter transplants in microcosms under controlled environmental conditions. Using soil and litter mixtures from a tropical land-use disturbance gradient spanning old growth (OG) forest, moderately logged (ML) forest, heavily logged (HL) forest, and oil palm plantation, we tested three non-exclusive hypotheses that: (1) Litter quality would explain the majority of variation in litter decomposition rates and thus be largely independent of the decomposer community; (2) Soil microbial decomposer communities from forests would have greater functional breadth relative to oil palm plantations (i.e., microbial decomposers from forests would decompose all litters faster than those from oil palm) but be functionally equivalent between OG and logged forests; (3) Higher rates of litter decomposition would be observed where litter and soil shared a common origin (HFA).
Materials and Methods
Study Sites
The study sites were located on orthic Acrisols in the Malaysian state of Sabah on the island of Borneo. The regional climate is moist tropical with annual daily mean temperature of 26.7°C and annual precipitation totals of ~2,600–2,700 mm (Walsh and Newbery, 1999; Kumagai and Porporato, 2012). Four 1 ha research plots comprised of 25 20 × 20 m subplots were selected that form part of the Global Ecosystem Monitoring (GEM) network (http://gem.tropicalforests.ox.ac.uk/). These were chosen to represent a tropical land-use disturbance gradient including OG tropical forest, ML tropical forest, HL tropical forest, and monoculture oil palm plantation. The OG forest plot was located within Danum Valley Conservation Area (DVCA). This forest has experienced little modern disturbance and has been protected since 1976 (Marsh and Greer, 1992). The most common tree genera within the plot were Shorea (Dipterocarpaceae) and Diospyros (Ebenaceae) with a low abundance of pioneer trees (0.1 ± 0.0% of basal area) and high basal area of standing trees (30.6 ± 3.37 m2/ha) (Riutta et al., 2018). Plots representing ML forest, HL forest and oil palm were within research sites located in the Kalabakan Forest Reserve and neighboring Benta Wawasan oil palm estate. These were established as part of the Stability of Altered Forest Ecosystems (SAFE) project and within the Yayasan Sabah Forest Management Area (Ewers et al., 2011) (www.safeproject.net). In the logged forest sites, the most common tree genera were Macaranga (Euphorbiaceae), Shorea (Dipterocarpaceae), and Syzygium (Myrtaceae). The ML forest plot had medium abundance of pioneer trees (6.9 ± 2.2% of basal area) and medium basal area of standing trees (19.3 ± 1.7 m2/ha) whilst the HL forest plot had the highest abundance of pioneer trees (28.1 ± 4.3% of basal area) and lowest standing tree basal area (6.81 ± 1.0 m2/ha) of the 3 forest plots (Riutta et al., 2018). Both logged forest plots (ML, HL) were subjected to one round of logging during the 1970's. The ML forest plot was subsequently logged once whilst the HL forest plot was logged three times between 1990 and 2008. Logging operations targeted primarily hardwoods from the genera Dryobalanops, Dipterocarpus, Shorea, and Parashorea. For more detailed site characteristics and logging history of forest plots refer to Riutta et al. (2018). For further information on tree communities and functional traits, refer to Both et al. (2019) with reference to the following plots: OG Forest = DAN-05, ML Forest = SAF-04, HL Forest = SAF-01. The oil palm plot was on its first rotation with standing palms ~7 years old at the time of sampling. Fertilizer (compound mixture comprised of diammonium phosphate, potassium chloride, ammonium sulfate, magnesium sulfate, and borax) was applied twice a year at a rate of 3–4 kg/palm (personal communication). Senesced fronds were stacked in rows between palms whilst inter-row and palm circle soils were kept weed free. No cover crops were present.
Sample Collection and Preparation
Each 1 ha plot was subdivided into 25 20 × 20 m subplots with five randomly selected for sampling. In forest plots, freshly senesced mixed litter was first collected from a 40 × 40 cm sampling area on the soil surface followed by removal of the organic soil to the horizon with underlying mineral soil. Strongly humified litter was considered as part of the organic soil layer. In the oil palm plantation, surface soils were collected at the edge of the frond stacks. Fresh litter was taken from senesced fronds within the adjacent frond stacks. To account for fronds of different ages we randomly selected 5 fronds for sampling and collected all leaflets from each frond. Leaf litter samples were carefully sorted by hand to remove woody debris, partially humified leaves, and reproductive material. Only intact leaves were retained, which were air-dried until constant weight. Fresh soils were homogenized by passing through a 4 mm sieve to remove stones, coarse woody, and litter debris and a subsample frozen for phospholipid fatty acid (PLFA) extraction and microbial sequencing. Soil and dried litter samples were sealed in plastic bags and transported on ice in a cool box to the UK. Fresh soil was stored at 4°C prior to establishment of the experiment. Soil subsamples for PLFA analysis were freeze-dried, passed through a 2 mm sieve, and ground to a fine powder using a pestle and mortar. Subsamples for total C, N, and inorganic P analysis were dried at 65°C for 48 h and ground as described above. Air-dried litter was further hand sorted to remove any remaining coarse woody debris and partially decomposed leaves and then chopped into small (<5 mm) pieces to create a homogenous litter mixture capturing the high species diversity of these tropical forests. Litter was then sterilized by autoclaving twice at 121°C and then dried at 65°C to constant weight. We acknowledge that autoclaving would likely change the initial chemical composition of litter. To quantify this, a subsample of each initial litter mixture (autoclaved and non-autoclaved) was ground to a fine powder using a pestle and mortar for chemical analysis (Table S1 for non-autoclaved litter chemistry results).
Microcosm Design
Microcosms were constructed in 9 cm diameter plastic petri dishes (Wardle et al., 1998). Fresh soils were adjusted to 70% of maximum water holding capacity and 10 g dry weight equivalent soil was added to each microcosm. A fine mesh (1 mm) disk was laid on top of the soil and 1 g of the autoclaved, sterilized litter mixture was then added. Microcosms were then partially sealed by taping the lid around the edge to minimize moisture loss. A small section was left open to allow for gas exchange. Our experimental design was 4 × 4 fully factorial with all litters being crossed with all soils. This reciprocal transplant design was used in order to separate the relative effects of soil microbial decomposer community and litter quality as well as interactive effects (Reed and Martiny, 2007). Each soil and litter had five true field replicates (n = 80 microcosms). This was replicated three times to allow destructive harvesting at three time intervals to capture progressive stages of litter decomposition. Microcosms were maintained in a controlled temperature incubator at 25°C and weighed weekly to adjust back to 70% WHC by the addition of de-ionized water. Microcosms were destructively harvested after 31, 105, and 398 days. Remaining litter was removed from the mesh disk with tweezers, dried at 65°C for 24 h and weighed. The percentage mass loss was used as a proxy for litter decomposition and calculated as the remaining dry litter weight divided by the initial dry litter weight.
Litter and Soil Analysis
The chemical composition of initial leaf litter (autoclaved and non-autoclaved) was measured as follows: Total C and N were quantified on an elemental analyser (NCS 2500, CE Instruments, UK). Total P was extracted using a sulfuric acid/hydrogen peroxide digest and analyzed on a flow injection auto analyzer (FIAStar 5000, Foss Tecator, Denmark). Ca, K, Mg, and Al were extracted by sulfuric acid digestion and measured using atomic absorption spectroscopy (AAS, Perkin Elmer AAnalyst 100, MA, USA). Fiber analysis (soluble cell content, hemicellulose + proteins, cellulose, lignin + recalcitrants) was performed using automated sequential digestion (ANKOM Technology, Macedon, NY, USA).
Soil pH was measured on fresh subsamples in a 2.5:1 water: soil slurry suspension, allowed to rest for 30 min and measured using a pH meter calibrated between pH 4–7 (pH210 Meter, Hanna Instruments, UK). Total soil C and N was measured using a LECO Truspec Micro elemental analyser (LECO Corporation, USA). Inorganic P was extracted from soils using a Bray No 1 extractant and analyzed using colorimetry on a SEAL AutoAnalyzer 3 (Seal Analytical, UK).
PLFAs were extracted from freeze-dried soils as part of the total lipid extract using a modified Bligh-Dyer extraction (White et al., 1979). Identification of PLFA's was carried out on a GC (Agilent Technologies 6890) fitted with a flame ionization detector (Agilent Technologies 5973). Sample PLFA peaks were identified based on known relative retention times. The terminal and mid-chain branched fatty acids C15:0i, C15:0a, C16:0i C17:0i, and C17:0a were used as indicators of Gram positive bacteria (Whitaker et al., 2014). Cyclopropyl saturated and monounsaturated fatty acids 16:1ω7c, 7,8 cyclic C17:0, C18:1ω7c, and 7,8 cy-C19:0 were used as indicators of Gram negative bacteria (Rinnan and Bååth, 2009). The fatty acids C18:2ω6,9c and C18:1ω9c were taken as indicators of fungi (Willers et al., 2015). Total microbial biomass was calculated as the sum of all identified PLFA's (C13:0, C14:0, C14:1ω5c, C15:0, C15:1ω5c, C16:0, 10Me-C16:0, C16:1ω7t, C16:1ω9c, C16:1ω5c, C17:0, 10Me-C17:0, C18:0i, C17:1ω7c, C18:0a, C18:0, 10Me-C18:0, C18:1ω7t, C18:1ω12c, C18:1ω5c, C18:2ω6t, 9,10-cyC19:0, C19:1ω12c, C20:0, C18:3ω6c, C20:1ω9c, C18:3ω3c, C20:2ω6c, C22:0, C20:3ω6c, C20:4ω6c, C20:5ω3c, C24:0; plus those listed above).
Molecular Analyses of Soil Microbial Communities
DNA was extracted from 0.2 g frozen soil using a Powersoil® DNA Isolation Kit according to the manufacturer's instructions. Amplicon libraries were constructed according to a dual indexing strategy with each primer consisting of the appropriate Illumina adapter, 8-nt index sequence, a 10-nt pad sequence, a 2-nt linker and the amplicon specific primer (Kozich et al., 2013). For bacteria, V3-V4 16S rRNA amplicon primers were used (CCTACGGGAGGCAGCAG and GCTATTGGAGCTGGAATTAC) (Kozich et al., 2013). Fungi were targeted by amplifying the ITS2 region using amplicon primers GTGARTCATCGAATCTTTG and TCCTCCGCTTATTGATATGC (Ihrmark et al., 2012). Amplicons were generated using PCR and a high fidelity DNA polymerase (Q5 Taq, New England Biolabs). Amplicon sizes were determined using an Agilent 2200 TapeStation system and libraries normalized using SequalPrep Normalization Plate Kit (Thermo Fisher Scientific) and quantified using a Qubit dsDNA HS kit (Thermo Fisher Scientific). Each amplicon library was sequenced separately on Illumina MiSeq using V3 600 cycle reagents at concentrations of 8 pM with a 5% PhiX Illumina control library.
Sequences were processed in R using DADA2 to quality filter, merge (where appropriate), de-noise, and assign taxonomies (Callahan et al., 2016). Briefly, 16S forward reads were trimmed to 250 bases. ITS amplicons reads were trimmed to 225 and 160 bases, forward, and reverse, respectively. Filtering settings were maximum number of Ns (maxN) = 0, maximum number of expected errors (maxEE) = 1. Sequences were dereplicated and the DADA2 core sequence variant inference algorithm applied. ITS sequences were merged using mergePairs function, whilst forward reads were used for 16S amplicons. Chimeric sequences were removed using removeBimeraDenovo default settings. The actual sequence variants (ASV) were subject to taxonomic assignment using assignTaxonomy at default settings; the training databases used were GreenGenes v13.8 and Unite v7.2 for 16S and ITS, respectively. Prior to analysis, samples were normalized by rarefying to 28,859 reads for 16S and 4,349 reads for ITS. Fungal taxa were classified as saprotrophic using FunGuild (Nguyen et al., 2016). Only those taxa with non-ambiguous classifications and assignments classified as “probable” or “highly probable” were retained.
Data Analysis
All analyses were performed using R statistical software (version 3.5.2) or SAS version 9.4 (SAS Institute, Cary NC). Differences in initial leaf litter chemistry, soil properties, and PLFA concentrations were tested using ANOVA with land use as the independent variable. Tukey's honestly significant difference post-hoc test was used to report pairwise differences between land-uses. Initial PLFA profiles were represented in two dimensions using principal components analysis (PCA). Bacterial, fungal, and saprotrophic fungal community composition was represented in two dimensions by principal co-ordinates analysis (PCoA) ordination of Bray–Curtis dissimilarities. Differences in initial microbial community composition were tested using PERMANOVA. ANOVA was used to test the significance of litter and soil origin as predictors of litter mass loss at each time point. Litter mass loss was the dependent variable whilst litter type and soil origin were included as independent variables with an interaction term. To determine which measured soil covariates were the best predictors of litter mass loss we replaced the factor “soil origin” with measured soil variables (soil chemical properties, PLFA concentrations, bacterial, fungal, and saprotrophic fungal richness) and performed multiple linear regression. Highly correlated variables were removed a priori and by inspecting variance inflation factors. The best model was selected by forward and backward stepwise model selection using AIC as a criterion. To determine whether initial microbial community composition was predictive of litter mass loss at the final time point (398 day) we used the two axis scores from PCoA on bacteria, fungal and saprotrophic fungal community composition alongside litter type as independent variables in multiple linear regression, with litter mass loss as the dependent variable. For all regression models, the effect size (relative importance) of each independent variable was estimated by averaging over orderings of regressors using the relaimpo package and is presented as a percentage of total variance (Groemping, 2006). The Decomposer Ability Regression Test (DART) was used to determine the relative contribution of litter quality, functional breadth of microbial decomposers, and home-field advantage in explaining rates of litter mass loss (Keiser et al., 2014).
Yi is the litter mass loss for observation i, βl is the ability of litter l, γs is the ability of the soil decomposer community s, ηh is the HFA of h where Home = Litterl × Soils when l and s are home field pairings. The intercept α represents the average mass loss for all observations in the dataset after controlling for litter, soil and home-field pairings and ε represents the error term. The parameters to be estimated are βl (Litter Quality), γs (Decomposer Ability) and ηh (Home-Field Advantage).
Results
Litter Quality and Soil Microbial Communities
Multiple litter chemistry parameters were different between land uses (Table 1). Foliar P [F(3,16) = 296.55, p ≤ 0.001], hemicellulose + proteins [F(3,16) = 140.72, p ≤ 0.001], and cellulose [F(3,16) = 30.07, p ≤ 0.001] were higher in oil palm litter relative to all forest sites, whilst soluble cell content [F(3,16) = 21.85, p ≤ 0.001] and lignin + recalcitrant [F(3,16) = 54.09, p ≤ 0.001] were lower. Lignin:N [F(3,16) = 8.51, p ≤ 0.001] and C:N [F(3,16) = 22.41, p ≤ 0.001] ratios were highest in ML forest litter relative to OG forest, HL forest, and oil palm (Table 1). Base cations varied between land uses [K: F(3,16) = 9.48, p ≤ 0.001, Mg: F(3,16) = 32.83, p ≤ 0.001, Ca: F(3,16) = 49.23, p ≤ 0.001]. K and Mg were lowest in ML forest whilst Ca was highest in HL forest (Table 1). Foliar C, N, and Al were not different between land uses (Table 1).
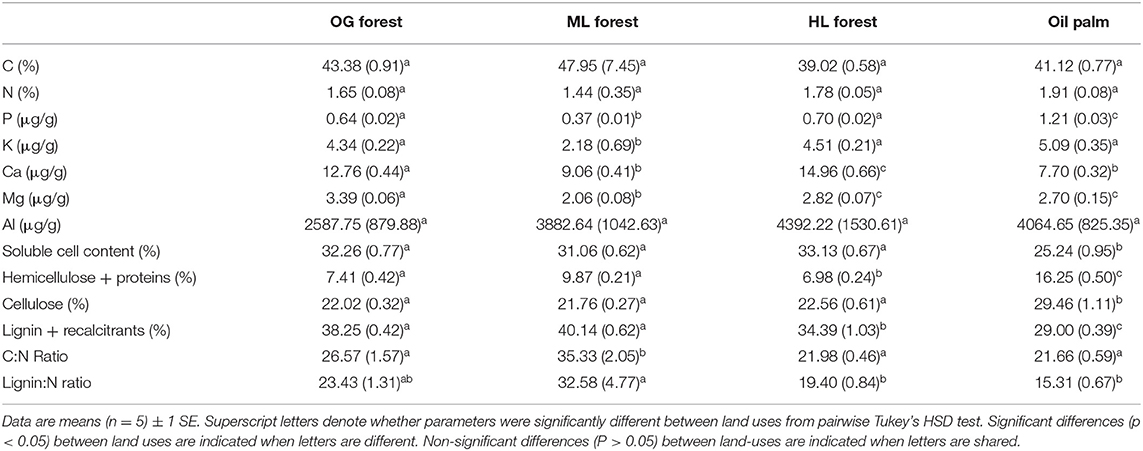
Table 1. Chemical properties of autoclaved litter mixtures taken from old growth (OG) forest, moderate logged (ML) forest, heavily logged (HL) forest, and oil palm plantation.
Soil properties also differed between land uses (Table 2). Soil moisture [F(3,16) = 39.32, p ≤ 0.001], total C [F(3,16) = 3.64, p = 0.04], and C:N ratio [F(3,16) = 27.76, p ≤ 0.001] were lowest in oil palm and highest in ML forest whilst soil pH [F(3,16) = 20.73, p ≤ 0.001] was lowest in ML forest and highest in OG forest (Table 2). Total PLFA [F(3,16) = 4.38, p = 0.02], bacterial PLFA [F(3,16) = 4.25, p = 0.02], and G- PLFA [F(3,16) = 7.66, p = 0.002] concentrations were all highest in ML forest and lowest in oil palm (Table 2). Bacterial, fungal and saprotrophic fungal richness varied according to land use [Bacteria: F(3,16) = 7.48, p = 0.002, Fungi: F(3,16) = 3.59, p = 0.04, Saprotrophic Fungi: F(3,16) = 5.21, p = 0.01] (Table 2) as did initial bacterial (16S) and fungal (ITS) community composition (Figures 1A–C) [PERMANOVA: Bacteria: F(3,19) = 1.87, p ≤ 0.001, R2 = 0.26, Fungi: F(3,19) = 2.47, p ≤ 0.001, R2 = 0.32, Saprotrophic Fungi: F(3,19) = 1.58, p ≤ 0.001, R2 = 0.23]. PLFA profiles were also significantly different between land uses (Figure 1D) [PERMANOVA: PLFA: F(3,19) = 5.16, p ≤ 0.001, R2 = 0.49]. Initial bacterial communities across all land uses were comprised primarily of Acidobacteria, Actinobacteria, and Proteobacteria whilst fungal communities were dominated by taxa from the phyla Ascomycota, Basidiomycota, and Mortierellomycota (Figures 2A,B).
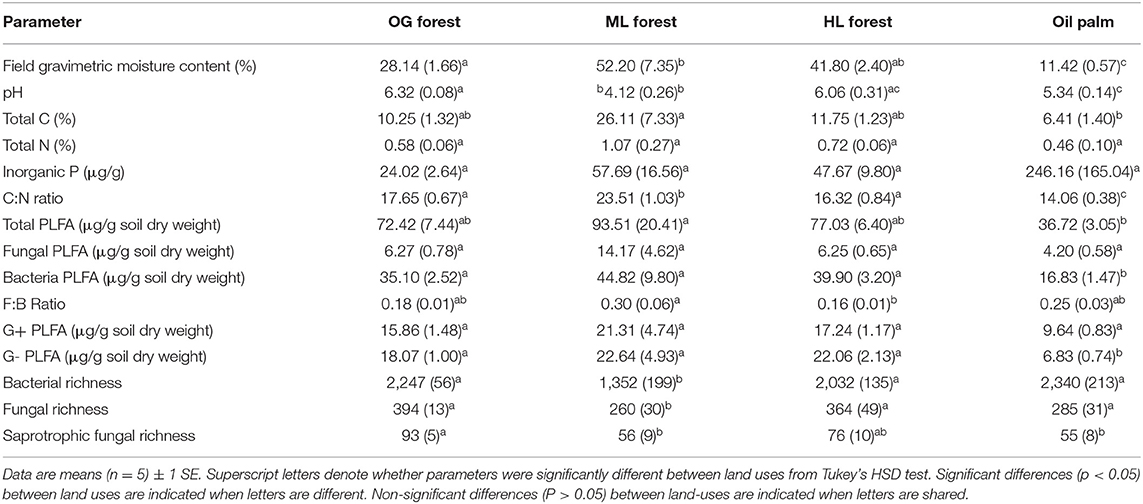
Table 2. Soil properties, PLFA concentrations, and microbial richness (number of observed ASV's) measured across old growth (OG) forest, moderate logged (ML) forest, heavily logged (HL) forest, and oil palm plantation.
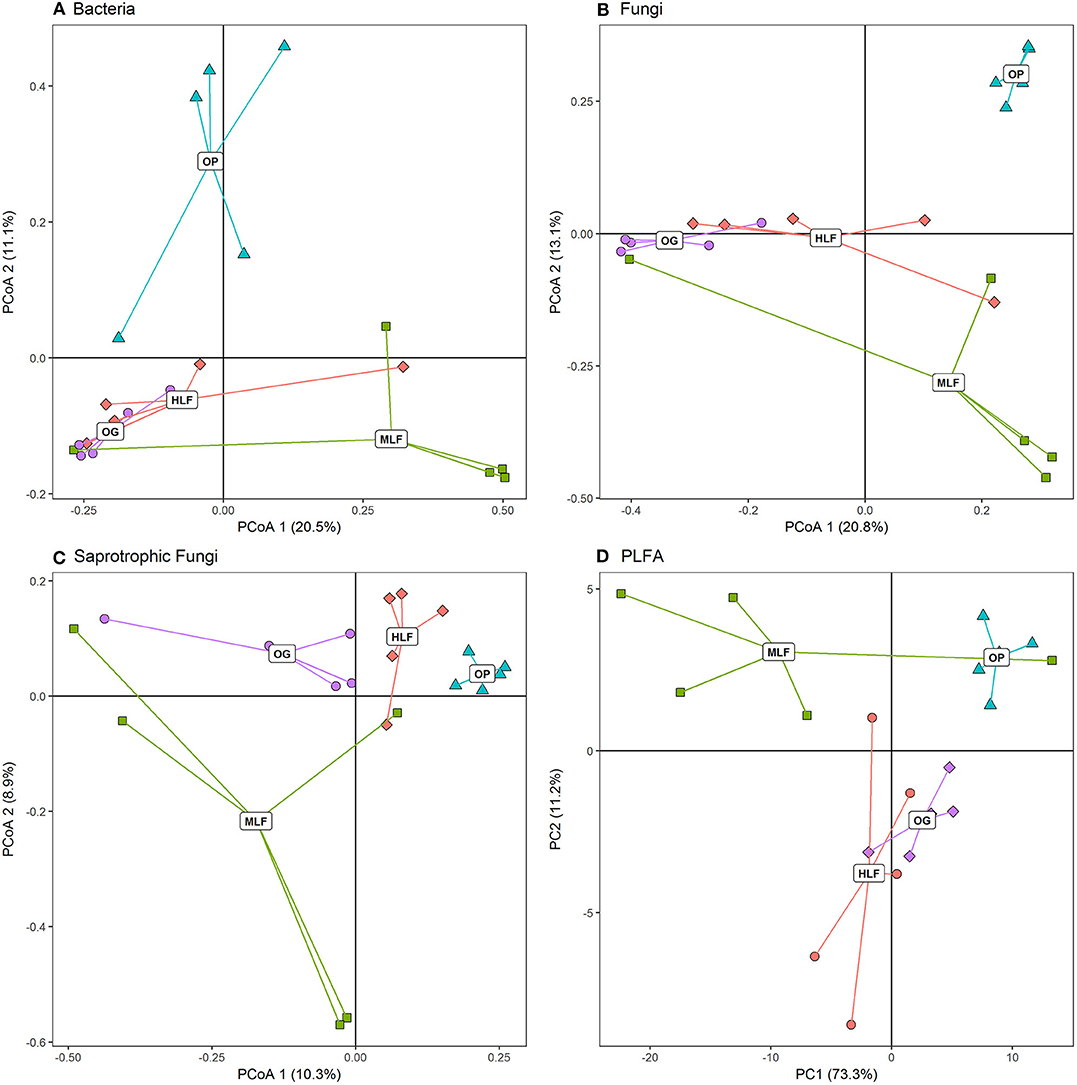
Figure 1. (A–C) Principal co-ordinates analysis (PCoA) ordinations of Bray–Curtis dissimilarities of bacteria, fungal, and saprotrophic fungal microbial community composition. (D) Principal components analysis of PLFA concentrations across old–growth forest (OG), moderate logged forest (MLF), heavily logged forest (HLF), and oil palm plantation (OP).
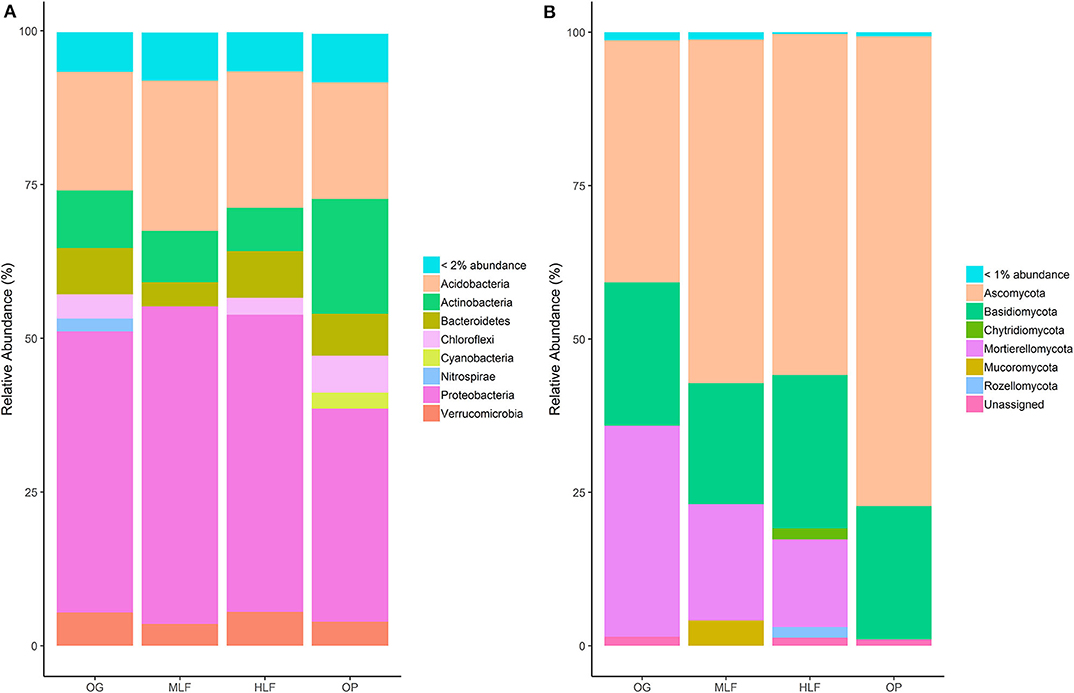
Figure 2. Relative abundance of phyla belonging to kingdoms Bacteria (A) and Fungi (B) across old-growth (OG), moderately logged forest (MLF), heavily logged forest (HLF), and oil palm (OP). Bacterial phyla representing <2% abundance were grouped into one category (<2% Abundance) whilst fungal phyla representing <1% abundance were also grouped into one category (<1% Abundance).
Controls on Rates of Litter Decomposition
Over the course of the 398-day experiment, significant differences in the rates of mass loss were observed between litter mixtures. Overall, oil palm litter decomposed faster than all forest litters losing 12.48 ± 0.59% 1SE after 31 days, 34.97 ± 1.49% 1SE after 105 days, and 42.51 ± 1.48% 1SE after 398 days (Figure 3) in line with its higher chemical quality (low lignin: N ratio). ML forest litter decomposed slowest losing 8.31 ± 0.54% 1SE after 31 days, 19.05 ± 0.54% 1SE after 105 days, and 23.85 ± 1.82% 1SE after 398 days. Both litter and soil origin were significant predictors of litter mass loss across all three time points (Table 3). Litter type accounted for between 34.51 and 66.28% of total variance whilst soil origin accounted for between 5.17 and 15.43% (Table 3). To explore whether soil properties, microbial abundance or richness drove the effect of soil origin, we “decomposed” the factor soil origin into measured soil properties (soil chemical properties, PLFA concentrations, bacterial, fungal, and saprotrophic fungal richness) using multiple regression. This showed that at the 398 day time point, soil inorganic P, total PLFA concentrations, and bacterial richness best explained variation in litter mass loss rates (Table 4). Initial bacterial, fungal, and saprotrophic fungal community composition was also predictive of litter mass loss at 398 day. However, these models explained less variation than the previous model (Table 5).
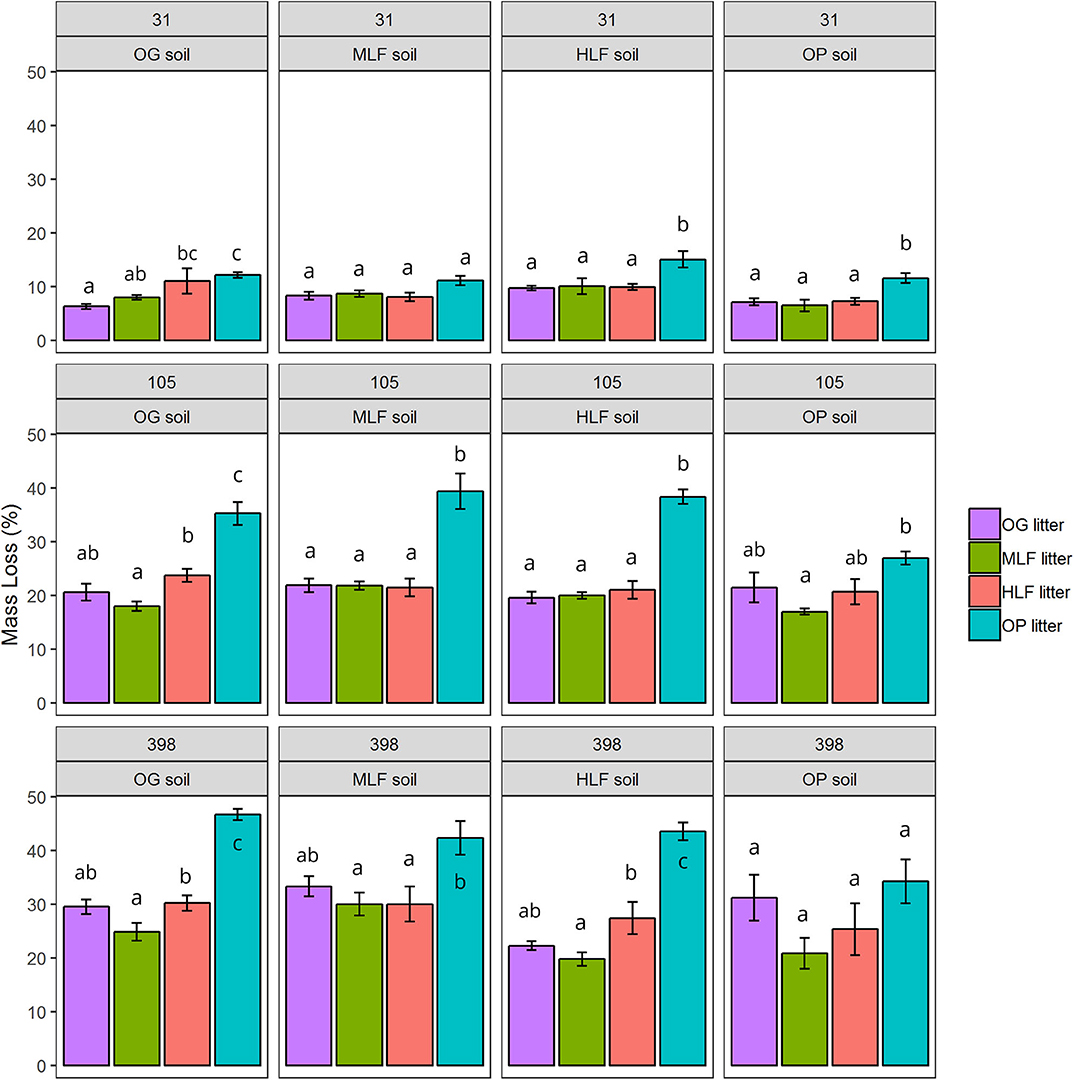
Figure 3. Mean litter mass loss from microcosms consisting of soils and mixed litter collected from old growth (OG) forest, moderate logged forest (MLF), heavily logged forest (HLF), and oil pam (OP). The four litter mixtures and soils were combined in a fully—factorial design (16 unique combinations) and destructively harvested after 31, 105, and 398 days. Values are mean percentage mass loss ± 1 SE of OG, MLF, HLF, and OP litter after 31, 105, and 398 days of decomposition. Superscript letters denote pairwise significant differences at P < 0.05 within each panel from ANOVA and Tukey's honestly significant difference post-hoc tests.
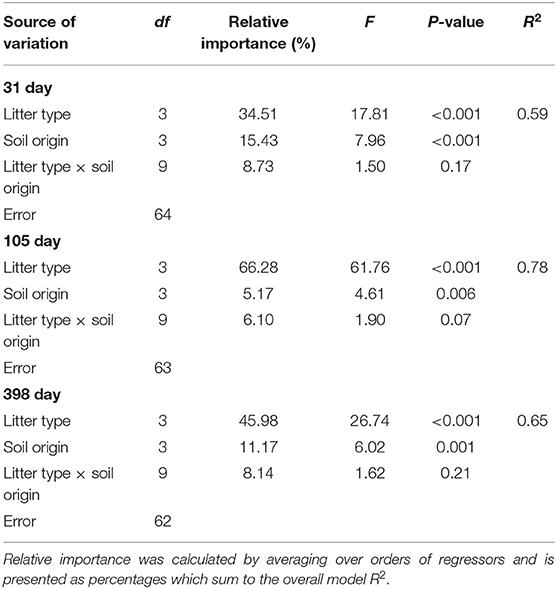
Table 3. Two way ANOVA results for the effects of soil origin, litter type, and an interaction term on litter mass loss (log-transformed) after 31, 105, and 398 days.
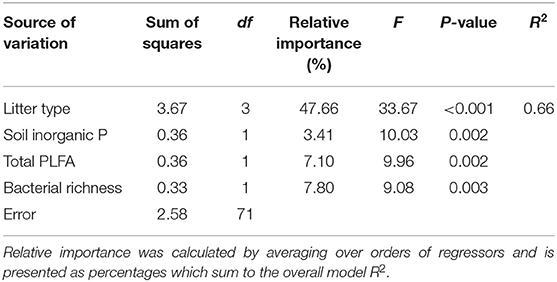
Table 4. Multiple linear regression results from a model with litter mass loss (log-transformed) at 398 day as the dependent variable and litter type, soil inorganic P, total PLFA concentrations, and bacterial richness as independent variables.
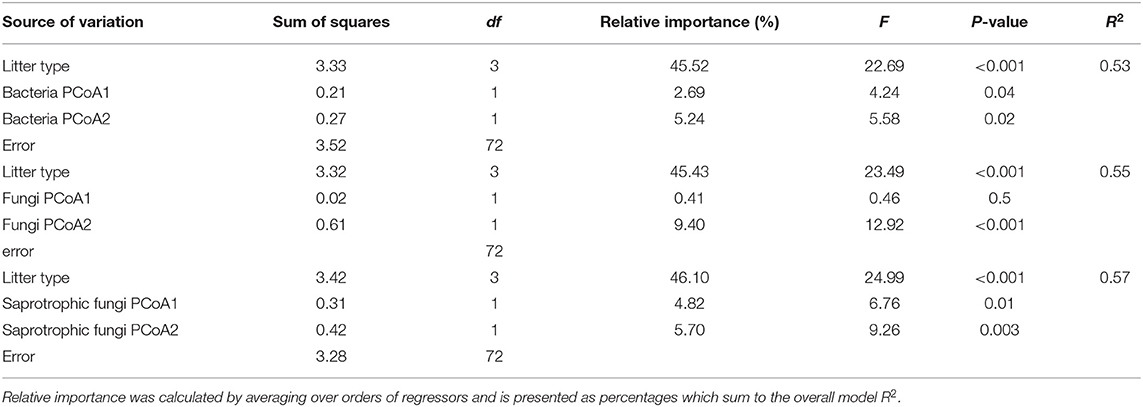
Table 5. Multiple linear regression results from a model with litter mass loss (log-transformed) at 398 day as the dependent variable, litter type, and axis scores 1 and 2 from principal co-ordinates analysis (PCoA) of initial bacteria, fungal, and saprotrophic fungal community composition as independent variables.
Final litter mass loss (after 398 day) of the lowest and highest chemical quality litters from ML forest and oil palm, respectively, was significantly correlated with initial bacterial and fungal community composition whilst PLFA profiles were significantly correlated with mass loss of all litters (Table S2). To assign a quality index to each litter and determine whether microbial decomposer communities were different in terms of their functional breadth and whether they perceived their “home” litter differently (HFA), we applied the DART regression framework. This showed that oil palm litter was of the highest quality whilst ML forest litter was the lowest (Table 6, Figure 4). OG and HL forest litter mixtures had intermediate qualities. After 398 day, the functional breadth of decomposer communities was significantly higher in OG forest and lower in HL forest relative to the average across all sites (Table 6, Figure 4). A positive HFA was detected for ML forest litter as it decomposed significantly faster when combined with its “home” soil whilst a negative HFA was observed for oil palm litter which decomposed slower in combination with its “home” soil (Table 6, Figure 4).
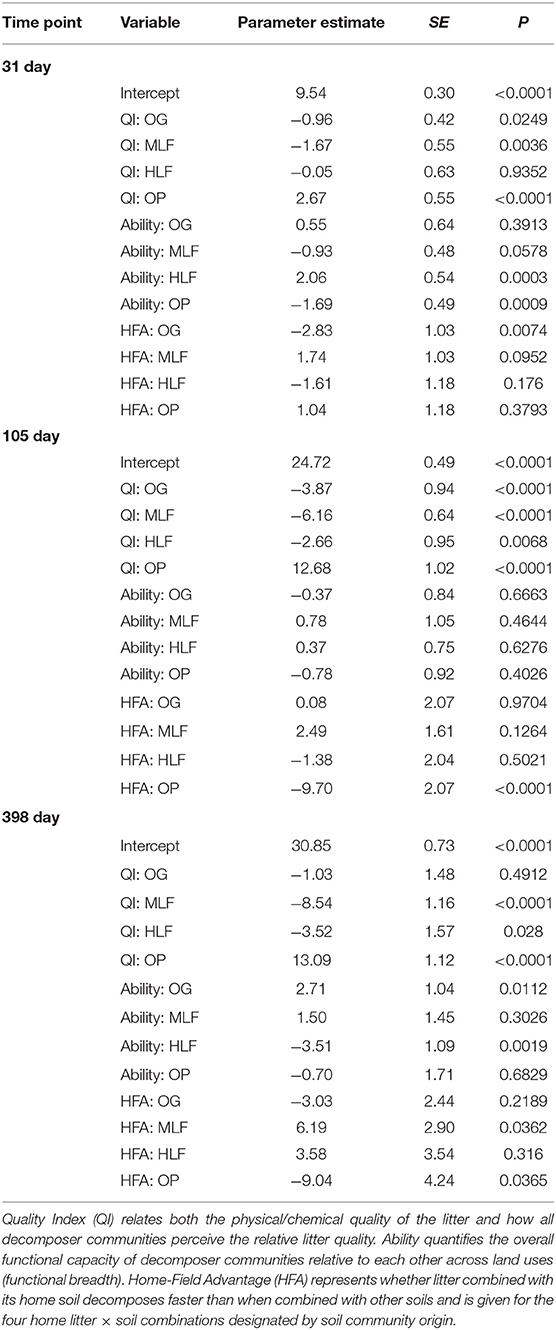
Table 6. Output from the decomposer ability regression test (DART) using mass loss of litter mixtures from old growth (OG) forest, moderate logged (ML) forest, heavily logged (HL) forest, and oil palm after 31, 105, and 398 days as a proxy for decomposition.
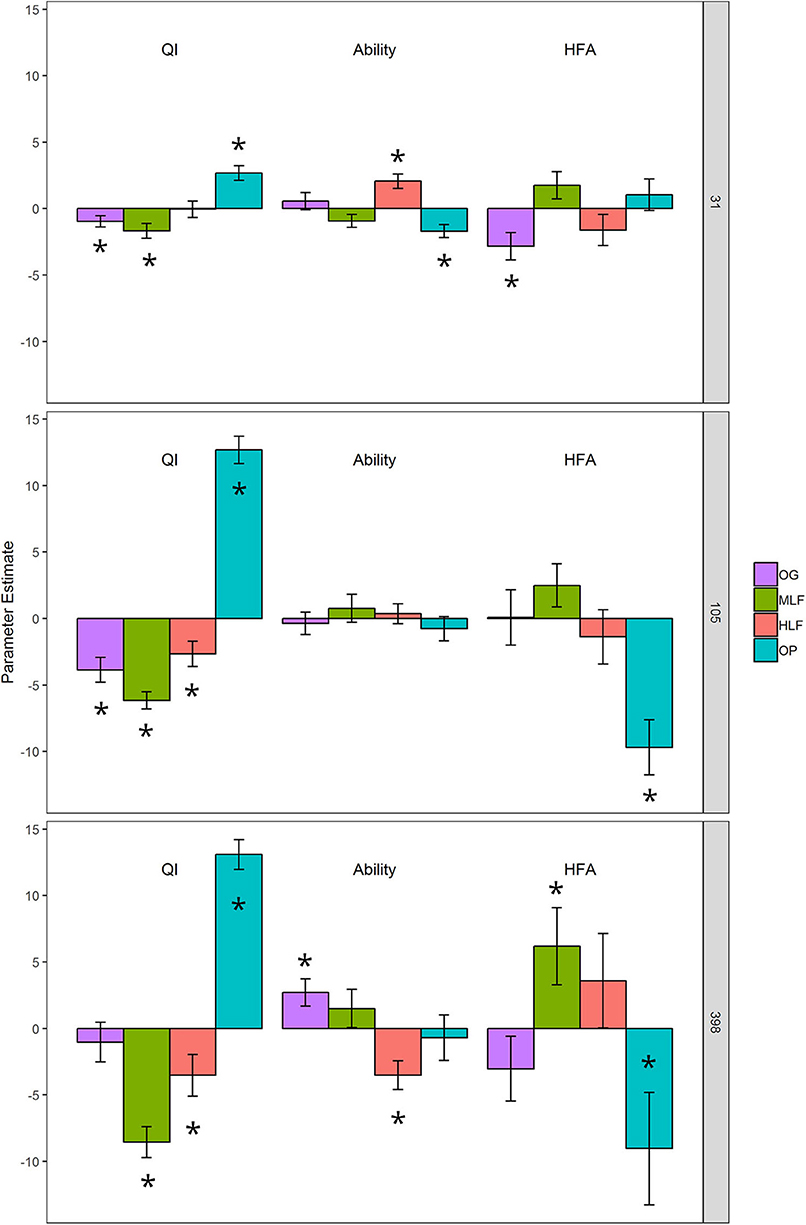
Figure 4. DART output calculated from % litter mass loss from litter decomposition microcosms after 31, 105, and 398 days: QI (litter quality index ranking the chemical quality of litters within the study), Ability (the functional capabilities of the microbial decomposer community to decompose all litters), and HFA (home-field advantage or the relative advantage of a decomposer community decomposing its “home” litter type). The bars are parameter estimate means ± 1 SE. “*” above or within bars represent a statistically significant difference from the intercept (y = 0) at p < 0.05. The intercept represents the average litter mass loss for all observations in the dataset after controlling for litter, soil and home-field pairings.
Discussion
We measured how litter quality and microbial decomposer communities varied across a tropical forest land-use disturbance gradient and their relative importance in terms of litter decomposition rates under controlled microclimatic conditions (Hypothesis 1). We also tested whether microbial decomposer communities from forests were functionally broader than those from oil palm plantations regarding the decomposition of litter mixtures (Hypothesis 2) and whether decomposer communities had a functional specialization to decompose litter from their native environments (Home-Field Advantage) (Hypothesis 3).
Alongside climate, litter quality has been shown to be a dominant control on rates of litter decomposition (Cornwell et al., 2008; Makkonen et al., 2012; Cleveland et al., 2014; Fanin et al., 2016). Our results broadly agree with these studies and lend support to Hypothesis 1 as we found that litter type explained the most variation in litter mass loss at all stages of decomposition across OG, ML, HL forest, and oil palm plantation (34.51–66.28%) (Table 3). Variation in forest chemical litter quality was not aligned with logging history as ML forest litter had lower quality litter than HL and OG forest, which had similar litter qualities (Table 1, Table S1). This probably reflects the higher heterogeneity in functional diversity of logged tropical forests as it does not agree with a recent study of tree community-weighted mean traits across this gradient (Both et al., 2019). This showed that whilst leaf tissue nutrient concentrations were primarily controlled by soil nutrient availability, logging did drive shifts from resource conservative to acquisitive traits (Both et al., 2019). Oil palm litter had the highest foliar P concentrations and lowest lignin: N ratios (Table 2) and decomposed ~40% faster than all forest litters (Figure 2).
Understanding whether microbial decomposer communities of distinct compositions are functionally equivalent or dissimilar concerning their ability to perform ecosystem processes such as litter decomposition is critical for predicting ecosystem responses to future environmental change (Allison and Martiny, 2008; Allison et al., 2013). We found that, in addition to litter quality, soil origin was also a significant predictor of litter mass loss at all time points and across all land-uses explaining 5.17–15.43% of total variation (Table 3). This finding is in agreement with similar studies in temperate ecosystems and indicates that litter decomposition rates, whilst largely regulated by litter quality, are not decoupled from microbial decomposer community composition under controlled, microclimatic conditions (Ayres et al., 2009b; Strickland et al., 2009b; Cleveland et al., 2014; Fanin et al., 2016).
Hypothesis 2 was not supported as microbial decomposer communities from OG forest had significantly higher functional breadth (i.e., they decomposed all 4 litter types faster than the average of decomposer communities from other land-uses) whilst decomposer communities from HL forest had the lowest functional breadth (i.e., they decomposed all 4 litter types slower than the average of decomposer communities from other land-uses) (Figure 4). OG and HL forest were the most similar in terms of both microbial decomposer community composition (Figure 1), microbial biomass and soil abiotic properties (Table 1) suggesting that even subtle shifts to the compositional make-up of microbial communities can lead to significant functional dissimilarity. Microbial decomposer communities from the oil palm plantation had a functional ability that was not significantly different from the average across all plots (Figure 4). This was surprising as total microbial biomass (Total PLFA), which has been positively associated with rates of litter decomposition and is represented in most decomposition models (McGuire and Treseder, 2010; Bradford et al., 2017), was much lower in oil palm (Table 1) relative to all forest plots. However, this result may have been confounded by past fertilization with inorganic nutrients as fertilization with N, P, K, and micronutrients has been shown to increase both cellulose and leaf litter decomposition rates in tropical forest (Hobbie and Vitousek, 2000; Kaspari et al., 2008). Soil inorganic P concentrations were higher in oil palm than forests (Table 2). When soil origin was replaced with soil properties, inorganic P concentrations as well as total PLFA and bacterial richness were positively associated with litter mass loss after 398 day. As P is thought to be a key limiting nutrient in lowland tropical forest ecosystems, alleviation of this limitation by fertilization may have increased rates of forest litter decomposition in oil palm, despite a lower abundance of microbial decomposers (Camenzind et al., 2018).
Numerous previous studies have shown that microbial communities decompose litter from their “home” site more quickly than exogenous litter (the Home-Field Advantage Hypothesis) (Gholz et al., 2000; Ayres et al., 2009b; Austin et al., 2014; Veen et al., 2015; Lin et al., 2019). However, evidence for this appears variable and context dependent with other studies showing no or even negative HFA (McGuire et al., 2010; Gießelmann et al., 2011; St. John et al., 2011). In partial support of Hypothesis 3, we found that after 398 day, the lowest quality litter mixture from ML forest decomposed significantly faster than all other litters when combined with its home soil (Figure 4). This is consistent with findings suggesting that HFA effects are stronger for recalcitrant litter and become more pronounced as litter types become more dissimilar (Veen et al., 2015). This may be explained by the higher concentrations of complex compounds in low quality litter, which require specialist decomposers (i.e., lignocellulolytic fungi) that can secrete specific enzymes to break them down (Ayres et al., 2009b; Milcu and Manning, 2011; Veen et al., 2015; Lin et al., 2019). We found evidence for this specialization as initial soil bacterial, fungal, saprotrophic fungal community composition, and PLFA profiles were correlated to rates of ML forest litter mass loss after 398 day (Table S2). A HFA effect can also be present for labile litter decomposition, explained by stimulation of competition for resources amongst copiotrophic microbial decomposers (Wang et al., 2013; Lin et al., 2019). Between 31–105 day and 105–398 day, labile oil palm litter decomposed slower in its “home” soil environment relative to all forest soils indicating a negative HFA effect at this stage of decomposition (Figure 4). The decay of labile compounds is thought to be primarily mediated by generalist microbial decomposers, which may be abundant in soil microbial biomass (Ayres et al., 2009a; Milcu and Manning, 2011; López-Mondéjar et al., 2018; Lin et al., 2019). Therefore, this negative HFA may be explained by lower microbial biomass (total PLFA) and thus abundance of generalist decomposers in oil palm relative to all forest sites (Table 1). In support of this, initial total PLFA and PLFA profiles were correlated with final mass loss of all litter mixtures (Table 4, Table S2). This suggests that generalist species found across land uses, rather than specialist decomposers regulated the decomposition of oil palm litter.
Conclusions
These data provide evidence that under controlled climatic conditions, litter quality regulates rates of litter decomposition across tropical OG forest, logged forests, and oil palm plantations. However, differences in soil microbial decomposer community explained a smaller but significant portion of variation in litter mass loss with OG forest decomposer communities retaining greater functional breadth, decomposing all litter mixtures faster relative to communities under logged forest and oil palm. We found evidence for a HFA in the most recalcitrant litter mixture from ML forest and significant correlations between initial decomposer community composition, microbial abundance, richness, and litter mass loss after 398 day. These findings suggest that shifts in soil microbial decomposer community structure should be considered when assessing the impacts of disturbance on crucial process rates, such as decomposition, in tropical forest ecosystems.
Data Availability Statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation, to any qualified researcher. The molecular data for this study have been deposited in the European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB36064.
Author Contributions
DE, NO, and NM designed the study. DE and SR collected field data with assistance from NM-L in accessing field sites. DE, SB, and TG conducted laboratory analyses. DE analyzed the data and wrote the manuscript with contributions from all co-authors. NO and NM secured the funding with support from NM-L. All authors contributed to the article and approved the submitted version.
Funding
This publication was a contribution from the UK NERC—funded Biodiversity and Land-use Impacts on Tropical Ecosystem Function (BALI) consortium and was funded through NERC's Human Modified Tropical Forests research programme (Grant No. NE/K016377/1 awarded to the BALI consortium) and supported by funding from the Sime Darby Foundation to the SAFE Project.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Glen Reynolds and the South East Asia Rainforest Research Partnership (SEARRP), Sabah Foundation, Benta Wawasan, the State Secretary, Sabah Chief Minister's Department, the Maliau Basin and Danum Valley Management Committees, Sabah Forestry Department, Sabah Forest Research Centre at Sepilok, Sabah Biodiversity Centre (JKM/MBS.1000-2/2 JLD.5 (20)), and the Economic Planning Unit for their support, access to the field sites and for permission to carry out fieldwork in Sabah. The support from Laura Kruitbos, Unding Jami, Terhi Riutta, Rob Ewers and SAFE research assistants is gratefully acknowledged.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/ffgc.2020.00081/full#supplementary-material
References
Achard, F., Eva, H. D., Stibig, H.-J., Mayaux, P., Gallego, J., Richards, T., et al. (2002). Determination of deforestation rates of the world's humid tropical forests. Science 297, 999–1002. doi: 10.1126/science.1070656
Allison, S. D., Lu, Y., Weihe, C., Goulden, M. L., Martiny, A. C., Treseder, K. K., et al. (2013). Microbial abundance and composition influence litter decomposition response to environmental change. Ecology 94, 714–725. doi: 10.1890/12-1243.1
Allison, S. D., and Martiny, J. B. H. (2008). Resistance, resilience, and redundancy in microbial communities. Proc. Natl. Acad. Sci. U.S.A. 105, 11512–11519. doi: 10.1073/pnas.0801925105
Austin, A. T., Vivanco, L., Gonzalez-Arzac, A., and Perez, L. I. (2014). There's no place like home? An exploration of the mechanisms behind plant litter- decomposer affinity in terrestrial ecosystems. New Phytol. 204, 307–314. doi: 10.1111/nph.12959
Ayres, E., Steltzer, H., Berg, S., and Wall, D. H. (2009a). Soil biota accelerate decomposition in high-elevation forests by specializing in the breakdown of litter produced by the plant species above them. J. Ecol. 97, 901–912. doi: 10.1111/j.1365-2745.2009.01539.x
Ayres, E., Steltzer, H., Simmons, B. L., Simpson, R. T., Steinweg, J. M., Wallenstein, M. D., et al. (2009b). Home-field advantage accelerates leaf litter decomposition in forests. Soil Biol. Biochem. 41, 606–610. doi: 10.1016/j.soilbio.2008.12.022
Bakker, M. A., Carreño-Rocabado, G., and Poorter, L. (2011). Leaf economics traits predict litter decomposition of tropical plants and differ among land use types. Funct. Ecol. 25, 473–483. doi: 10.1111/j.1365-2435.2010.01802.x
Baraloto, C., Hérault, B., Paine, C. E. T., Massot, H., Blanc, L., Bonal, D., et al. (2012). Contrasting taxonomic and functional responses of a tropical tree community to selective logging. J. Appl. Ecol. 49, 861–870. doi: 10.1111/j.1365-2664.2012.02164.x
Bonan, G. B., Hartman, M. D., Parton, W. J., and Wieder, W. R. (2013). Evaluating litter decomposition in earth system models with long-term litterbag experiments: an example using the Community Land Model version 4 (CLM4). Glob. Chang. Biol. 19, 957–974. doi: 10.1111/gcb.12031
Both, S., Elias, D. M. O., Kritzler, U. H., Ostle, N. J., and Johnson, D. (2017). Land use not litter quality is a stronger driver of decomposition in hyperdiverse tropical forest. Ecol. Evol. 7, 9307–9318. doi: 10.1002/ece3.3460
Both, S., Riutta, T., Paine, C. E. T., Elias, D. M. O., Cruz, R. S., Jain, A., et al. (2019). Logging and soil nutrients independently explain plant trait expression in tropical forests. New Phytol. 221, 1853–1865. doi: 10.1111/nph.15444
Bradford, M. A., Berg, B., Maynard, D. S., Wieder, W. R., and Wood, S. A. (2016). Understanding the dominant controls on litter decomposition. J. Ecol. 104, 229–238. doi: 10.1111/1365-2745.12507
Bradford, M. A., Veen, G. F., Bonis, A., Bradford, E. M., Classen, A. T., Cornelissen, J. H. C., et al. (2017). A test of the hierarchical model of litter decomposition. Nat. Ecol. Evol. 1, 1836–1845. doi: 10.1038/s41559-017-0367-4
Bradford, M. A., Warren, R. J., Baldrian, P., Crowther, T. W., Maynard, D. S., Oldfield, E. E., et al. (2014). Climate fails to predict wood decomposition at regional scales. Nat. Clim. Chang. 4, 625–630. doi: 10.1038/nclimate2251
Brearley, F. Q. (2012). Ectomycorrhizal associations of the dipterocarpaceae. Biotropica 44, 637–648. doi: 10.1111/j.1744-7429.2012.00862.x
Callahan, B. J., Mcmurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P. (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13:581. doi: 10.1038/nmeth.3869
Camenzind, T., Hattenschwiler, S., Treseder, K. K., Lehmann, A., and Rillig, M. C. (2018). Nutrient limitation of soil microbial processes in tropical forests. Ecol. Monogr. 88, 4–21. doi: 10.1002/ecm.1279
Carreño-Rocabado, G., Peña-Claros, M., Bongers, F., Alarcón, A., Licona, J.-C., and Poorter, L. (2012). Effects of disturbance intensity on species and functional diversity in a tropical forest. J. Ecol.100, 1453–1463. doi: 10.1111/j.1365-2745.2012.02015.x
Cleveland, C. C., Reed, S. C., Keller, A. B., Nemergut, D. R., O'neill, S. P., Ostertag, R., et al. (2014). Litter quality versus soil microbial community controls over decomposition: a quantitative analysis. Oecologia 174, 283–294. doi: 10.1007/s00442-013-2758-9
Corley, R. H. V., and Tinker, P. B. (2015). ‘Growth, flowering and yield,” in The Oil Palm (Chichester: Wiley Blackwell), 89–137. doi: 10.1002/9781118953297.ch5
Cornwell, W. K., Cornelissen, J. H. C., Amatangelo, K., Dorrepaal, E., Eviner, V. T., Godoy, O., et al. (2008). Plant species traits are the predominant control on litter decomposition rates within biomes worldwide. Ecol. Lett. 11, 1065–1071. doi: 10.1111/j.1461-0248.2008.01219.x
Corrales, A., Henkel, T. W., and Smith, M. E. (2018). Ectomycorrhizal associations in the tropics – biogeography, diversity patterns and ecosystem roles. New Phytol. 220, 1076–1091. doi: 10.1111/nph.15151
Currie, W. S., Harmon, M. E., Burke, I. C., Hart, S. C., Parton, W. J., and Silver, W. (2010). Cross-biome transplants of plant litter show decomposition models extend to a broader climatic range but lose predictability at the decadal time scale. Glob. Chang. Biol. 16, 1744–1761. doi: 10.1111/j.1365-2486.2009.02086.x
Ding, Y., Zang, R. G., Lu, X. H., and Huang, J. H. (2019). Functional features of tropical montane rain forests along a logging intensity gradient. Ecol. Indic. 97, 311–318. doi: 10.1016/j.ecolind.2018.10.030
Djukic, I., Kepfer-Rojas, S., Schmidt, I. K., Larsen, K. S., Beier, C., Berg, B., et al. (2018). Early stage litter decomposition across biomes. Sci. Total Environ. 628–629, 1369–1394. doi: 10.1016/j.scitotenv.2018.01.012
Estoque, R. C., Ooba, M., Avitabile, V., Hijioka, Y., Dasgupta, R., Togawa, T., et al. (2019). The future of Southeast Asia's forests. Nat. Commun. 10:1829. doi: 10.1038/s41467-019-09646-4
Ewers, R. M., Didham, R. K., Fahrig, L., Ferraz, G., Hector, A., Holt, R. D., et al. (2011). A large-scale forest fragmentation experiment: the stability of altered forest ecosystems project. Philos. Trans. R Soc. Lond. B Biol. Sci. 366, 3292–3302. doi: 10.1098/rstb.2011.0049
Fanin, N., Fromin, N., and Bertrand, I. (2016). Functional breadth and home-field advantage generate functional differences among soil microbial decomposers. Ecology 97, 1023–1037. doi: 10.1890/15-1263.1
Fernandez, C. W., and Kennedy, P. G. (2016). Revisiting the ‘Gadgil effect': do interguild fungal interactions control carbon cycling in forest soils? New Phytol. 209, 1382–1394. doi: 10.1111/nph.13648
Gholz, H. L., Wedin, D. A., Smitherman, S. M., Harmon, M. E., and Parton, W. J. (2000). Long-term dynamics of pine and hardwood litter in contrasting environments: toward a global model of decomposition. Glob. Chang. Biol. 6, 751–765. doi: 10.1046/j.1365-2486.2000.00349.x
Gibbs, H. K., Ruesch, A. S., Achard, F., Clayton, M. K., Holmgren, P., Ramankutty, N., et al. (2010). Tropical forests were the primary sources of new agricultural land in the 1980s and 1990s. Proc. Natl. Acad. Sci. U.S.A. 107, 16732–16737. doi: 10.1073/pnas.0910275107
Gießelmann, U. C., Martins, K. G., Brändle, M., Schädler, M., Marques, R., and Brandl, R. (2011). Lack of home-field advantage in the decomposition of leaf litter in the Atlantic Rainforest of Brazil. Appl. Soil Ecol. 49, 5–10. doi: 10.1016/j.apsoil.2011.07.010
Groemping, U. (2006). Relative importance for linear regression in R: the package relaimpo. J. Stat. Softw. 17:39439. doi: 10.18637/jss.v017.i01
Gui, H., Hyde, K., Xu, J., and Mortimer, P. (2017). Arbuscular mycorrhiza enhance the rate of litter decomposition while inhibiting soil microbial community development. Sci. Rep. 7:42184. doi: 10.1038/srep42184
Hobbie, S. E., and Vitousek, P. M. (2000). Nutrient limitation of decomposition in hawaiian forests. Ecology 81, 1867–1877. doi: 10.1890/0012-9658(2000)081[1867:NLODIH]2.0.CO
Ihrmark, K., Bodeker, I. T. M., Cruz-Martinez, K., Friberg, H., Kubartova, A., Schenck, J., et al. (2012). New primers to amplify the fungal ITS2 region - evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 82, 666–677. doi: 10.1111/j.1574-6941.2012.01437.x
Kaspari, M., Garcia, M. N., Harms, K. E., Santana, M., Wright, S. J., and Yavitt, J. B. (2008). Multiple nutrients limit litterfall and decomposition in a tropical forest. Ecol. Lett. 11, 35–43. doi: 10.1111/j.1461-0248.2007.01124.x
Keiser, A. D., Keiser, D. A., Strickland, M. S., and Bradford, M. A. (2014). Disentangling the mechanisms underlying functional differences among decomposer communities. J. Ecol. 102, 603–609. doi: 10.1111/1365-2745.12220
Keiser, A. D., Strickland, M. S., Fierer, N., and Bradford, M. A. (2011). The effect of resource history on the functioning of soil microbial communities is maintained across time. Biogeosciences 8, 1477–1486. doi: 10.5194/bg-8-1477-2011
Kerfahi, D., Tripathi, B. M., Lee, J., Edwards, D. P., and Adams, J. M. (2014). The impact of selective-logging and forest clearance for oil palm on fungal communities in Borneo. PLoS ONE 9:e111525. doi: 10.1371/journal.pone.0111525
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K., and Schloss, P. D. (2013). Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the miseq illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120. doi: 10.1128/AEM.01043-13
Kumagai, T. O., and Porporato, A. (2012). Drought-induced mortality of a Bornean tropical rain forest amplified by climate change. J. Geophys. Res. Biogeosci. 117:G02032. doi: 10.1029/2011JG001835
Laurance, W. F., Sayer, J., and Cassman, K. G. (2014). Agricultural expansion and its impacts on tropical nature. Trends Ecol. Evol. 29, 107–116. doi: 10.1016/j.tree.2013.12.001
Lin, D. M., Pang, M., Fanin, N., Wang, H. J., Qian, S. H., Zhao, L., et al. (2019). Fungi participate in driving home-field advantage of litter decomposition in a subtropical forest. Plant Soil 434, 467–480. doi: 10.1007/s11104-018-3865-5
López-Mondéjar, R., Brabcová, V., Štursová, M., Davidová, A., Jansa, J., Cajthaml, T., et al. (2018). Decomposer food web in a deciduous forest shows high share of generalist microorganisms and importance of microbial biomass recycling. ISME J. 12, 1768–1778. doi: 10.1038/s41396-018-0084-2
Luke, S. H., Purnomo, D., Advento, A. D., Aryawan, A. A. K., Naim, M., et al. (2019). Effects of understory vegetation management on plant communities in oil palm plantations in Sumatra, Indonesia. Front. Forests Glob. Change 2:13. doi: 10.3389/ffgc.2019.00033
Makkonen, M., Berg, M. P., Handa, I. T., Hattenschwiler, S., Van Ruijven, J., Van Bodegom, P. M., et al. (2012). Highly consistent effects of plant litter identity and functional traits on decomposition across a latitudinal gradient. Ecol. Lett. 15, 1033–1041. doi: 10.1111/j.1461-0248.2012.01826.x
Marsh, C. W., and Greer, A. G. (1992). Forest land-use in Sabah, Malaysia: an introduction to Danum Valley. philosophical transactions of the royal society of London. Series B Biol. Sci. 335, 331–339. doi: 10.1098/rstb.1992.0025
Martiny, J. B., Martiny, A. C., Weihe, C., Lu, Y., Berlemont, R., Brodie, E. L., et al. (2017). Microbial legacies alter decomposition in response to simulated global change. ISME J. 11, 490–499. doi: 10.1038/ismej.2016.122
McGuire, K. L., D'angelo, H., Brearley, F. Q., Gedallovich, S. M., Babar, N., Yang, N., et al. (2015). Responses of soil fungi to logging and oil palm agriculture in Southeast Asian Tropical Forests. Microb. Ecol. 69, 733–747. doi: 10.1007/s00248-014-0468-4
McGuire, K. L., and Treseder, K. K. (2010). Microbial communities and their relevance for ecosystem models: decomposition as a case study. Soil Biol. Biochem. 42, 529–535. doi: 10.1016/j.soilbio.2009.11.016
McGuire, K. L., Zak, D. R., Edwards, I. P., Blackwood, C. B., and Upchurch, R. (2010). Slowed decomposition is biotically mediated in an ectomycorrhizal, tropical rain forest. Oecologia 164, 785–795. doi: 10.1007/s00442-010-1686-1
Meentemeyer, V. (1978). Macroclimate and lignin control of litter decomposition rates. Ecology 59, 465–472. doi: 10.2307/1936576
Milcu, A., and Manning, P. (2011). All size classes of soil fauna and litter quality control the acceleration of litter decay in its home environment. Oikos 120, 1366–1370. doi: 10.1111/j.1600-0706.2010.19418.x
Mueller, R. C., Rodrigues, J. L. M., Nüsslein, K., and Bohannan, B. J. M. (2016). Land use change in the Amazon rain forest favours generalist fungi. Funct. Ecol. 30, 1845–1853. doi: 10.1111/1365-2435.12651
Nannipieri, P., Ascher, J., Ceccherini, M. T., Landi, L., Pietramellara, G., and Renella, G. (2003). Microbial diversity and soil functions. Eur. J. Soil Sci. 54, 655–670. doi: 10.1046/j.1351-0754.2003.0556.x
Nguyen, N. H., Song, Z., Bates, S. T., Branco, S., Tedersoo, L., Menke, J., et al. (2016). FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 20, 241–248. doi: 10.1016/j.funeco.2015.06.006
Phosri, C., Rodriguez, A., Sanders, I. R., and Jeffries, P. (2010). The role of mycorrhizas in more sustainable oil palm cultivation. Agric. Ecosyst. Environ. 135, 187–193. doi: 10.1016/j.agee.2009.09.006
Potapov, P., Hansen, M. C., Laestadius, L., Turubanova, S., Yaroshenko, A., Thies, C., et al. (2017). The last frontiers of wilderness: Tracking loss of intact forest landscapes from 2000 to 2013. Sci. Adv. 3:e1600821. doi: 10.1126/sciadv.1600821
Reed, H. E., and Martiny, J. B. H. (2007). Testing the functional significance of microbial composition in natural communities. FEMS Microbiol. Ecol. 62, 161–170. doi: 10.1111/j.1574-6941.2007.00386.x
Rinnan, R., and Bååth, E. (2009). Differential utilization of carbon substrates by bacteria and fungi in Tundra Soil. Appl. Environ. Microbiol. 75, 3611–3620. doi: 10.1128/AEM.02865-08
Riutta, T., Malhi, Y., Kho, L. K., Marthews, T. R., Huasco, W. H., Khoo, M., et al. (2018). Logging disturbance shifts net primary productivity and its allocation in Bornean tropical forests. Glob. Chang. Biol. 24:2915–2916. doi: 10.1111/gcb.14068
Rodrigues, J. L. M., Pellizari, V. H., Mueller, R., Baek, K., Jesus, E. D. C., Paula, F. S., et al. (2013). Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc. Natl. Acad. Sci. U.S.A. 110, 988–993. doi: 10.1073/pnas.1220608110
Ruegg, J., Quezada, J. C., Santonja, M., Ghazoul, J., Kuzyakov, Y., Buttler, A., et al. (2019). Drivers of soil carbon stabilization in oil palm plantations. Land Degrad. Dev. 30, 1904–1915. doi: 10.1002/ldr.3380
Shi, L. L., Dossa, G. G. O., Paudel, E., Zang, H. D., Xu, J. C., and Harrison, R. D. (2019). Changes in fungal communities across a forest disturbance gradient. Appl. Environ. Microbiol. 85:e00080–19. doi: 10.1128/AEM.00080-19
Sodhi, N. S., Koh, L. P., Brook, B. W., and Ng, P. K. L. (2004). Southeast Asian biodiversity: an impending disaster. Trends Ecol. Evol. 19, 654–660. doi: 10.1016/j.tree.2004.09.006
St. John, M. G., Orwin, K. H., and Dickie, I. A. (2011). No ‘home' versus ‘away' effects of decomposition found in a grassland–forest reciprocal litter transplant study. Soil Biol. Biochem. 43, 1482–1489. doi: 10.1016/j.soilbio.2011.03.022
Strickland, M. S., Lauber, C., Fierer, N., and Bradford, M. A. (2009a). Testing the functional significance of microbial community composition. Ecology 90, 441–451. doi: 10.1890/08-0296.1
Strickland, M. S., Osburn, E., Lauber, C., Fierer, N., and Bradford, M. A. (2009b). Litter quality is in the eye of the beholder: initial decomposition rates as a function of inoculum characteristics. Funct. Ecol. 23, 627–636. doi: 10.1111/j.1365-2435.2008.01515.x
Tripathi, B. M., Edwards, D. P., Mendes, L. W., Kim, M., Dong, K., Kim, H., et al. (2016). The impact of tropical forest logging and oil palm agriculture on the soil microbiome. Mol. Ecol. 25, 2244–2257. doi: 10.1111/mec.13620
Tripathi, B. M., Kim, M., Singh, D., Lee-Cruz, L., Lai-Hoe, A., Ainuddin, A. N., et al. (2012). Tropical soil bacterial communities in Malaysia: pH dominates in the equatorial tropics too. Microb. Ecol. 64, 474–484. doi: 10.1007/s00248-012-0028-8
Van Der Heijden, M. G. A., Bardgett, R. D., and Van Straalen, N. M. (2008). The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol. Lett. 11, 296–310. doi: 10.1111/j.1461-0248.2007.01139.x
Veen, G. F., Freschet, G. T., Ordonez, A., and Wardle, D. A. (2015). Litter quality and environmental controls of home-field advantage effects on litter decomposition. Oikos 124, 187–195. doi: 10.1111/oik.01374
Veen, G. F., Keiser, A. D., Van Der Putten, W. H., and Wardle, D. A. (2018). Variation in home-field advantage and ability in leaf litter decomposition across successional gradients. Funct. Ecol. 32, 1563–1574. doi: 10.1111/1365-2435.13107
Wall, D. H., Bradford, M. A., St John, M. G., Trofymow, J. A., Behan-Pelletier, V., Bignell, D. E., et al. (2008). Global decomposition experiment shows soil animal impacts on decomposition are climate-dependent. Glob. Chang. Biol. 14, 2661–2677. doi: 10.1111/j.1365-2486.2008.01672.x
Walsh, R. P., and Newbery, D. M. (1999). The ecoclimatology of Danum, Sabah, in the context of the world's rainforest regions, with particular reference to dry periods and their impact. Philos Trans. R Soc. Lond. B Biol. Sci. 354, 1869–1883. doi: 10.1098/rstb.1999.0528
Wang, Q., Zhong, M., and He, T. (2013). Home-field advantage of litter decomposition and nitrogen release in forest ecosystems. Biol. Fertil. Soils 49, 427–434. doi: 10.1007/s00374-012-0741-y
Wardle, D. A., Barker, G. M., Bonner, K. I., and Nicholson, K. S. (1998). Can comparative approaches based on plant ecophysiological traits predict the nature of biotic interactions and individual plant species effects in ecosystems? J. Ecol. 86, 405–420. doi: 10.1046/j.1365-2745.1998.00268.x
Whitaker, J., Ostle, N., Mcnamara, N. P., Nottingham, A. T., Stott, A. W., Bardgett, R. D., et al. (2014). Microbial carbon mineralization in tropical lowland and montane forest soils of Peru. Front. Microbiol. 5:720. doi: 10.3389/fmicb.2014.00720
White, D. C., Davis, W. M., Nickels, J. S., King, J. D., and Bobbie, R. J. (1979). Determination of the sedimentary microbial biomass by extractible lipid phosphate. Oecologia 40, 51–62. doi: 10.1007/BF00388810
Willers, C., Jansen Van Rensburg, P. J., and Claassens, S. (2015). Phospholipid fatty acid profiling of microbial communities–a review of interpretations and recent applications. J. Appl. Microbiol. 119, 1207–1218. doi: 10.1111/jam.12902
Keywords: tropical forests, logging, oil palm, leaf litter decomposition, soil microbial communities, functional dissimilarity
Citation: Elias DMO, Robinson S, Both S, Goodall T, Majalap-Lee N, Ostle NJ and McNamara NP (2020) Soil Microbial Community and Litter Quality Controls on Decomposition Across a Tropical Forest Disturbance Gradient. Front. For. Glob. Change 3:81. doi: 10.3389/ffgc.2020.00081
Received: 30 August 2019; Accepted: 09 June 2020;
Published: 16 July 2020.
Edited by:
Edward T. A. Mitchard, University of Edinburgh, United KingdomReviewed by:
Tomas Ferreira Domingues, University of São Paulo, BrazilFrances Claire Manning, University of Aberdeen, United Kingdom
Copyright © 2020 Elias, Robinson, Both, Goodall, Majalap-Lee, Ostle and McNamara. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Dafydd M. O. Elias, ZGFmaWFzQGNlaC5hYy51aw==
 Dafydd M. O. Elias
Dafydd M. O. Elias Samuel Robinson
Samuel Robinson Sabine Both
Sabine Both Tim Goodall5
Tim Goodall5 Nick J. Ostle
Nick J. Ostle Niall P. McNamara
Niall P. McNamara