- Department of Life Science, Sogang University, Seoul, South Korea
To cope with environmental stresses, plants have developed various stress tolerance mechanisms that involve the induction of many stress responsive genes through stress-specific and common signaling pathways. Stress-specific/common transcription factors, rather than general basal factors, were considered important in this stress tolerance. The Arabidopsis STABILIZED1 (STA1) gene encodes a putative pre-mRNA splicing factor that is similar to the human U5 snRNP-associated 102-kDa protein and the yeast pre-mRNA splicing factors, PRP1p and Prp6p. As pre-mRNA splicing is a necessary process for proper gene expression in eukaryotes, STA1 is expected to be constantly functional in all conditions. Interestingly, STA1 expression is induced by temperature stresses, and STA1 recessive mutation (sta1-1) resulted in temperature stress-specific hypersensitivity. This suggests STA1’s stress specific function in addition to its presumed “housekeeping” role. In order to establish the genetic system to understand the regulation of STA1 expression in temperature stresses, we generated a bioluminescent Arabidopsis plant harboring the STA1 promoter fused to the firefly luciferase coding sequence (STA1p-LUC). Through genetic analysis, the bioluminescent Arabidopsis homozygous for one-copy STA1p-LUC was isolated and characterized. In this STA1p-LUC line, the expression patterns of STA1p-LUC were similar to those of the endogenous STA1 gene under cold and heat stresses. The STA1p-LUC line was then chemically mutagenized and screened to isolate the genetic loci of STA1 regulators under cold or heat stresses. Mutants with altered STA1p-LUC luminescence were identified and further confirmed through luminescence imaging in the next generation and analysis of endogenous STA1 expression. The categorization of STA1p-LUC deregulated mutants implicated the existence of cold or heat stress-specific as well as common genetic regulators for STA1 expression. Interestingly, some mutants showed opposite-directional deregulation of STA1 expression depending on the type of thermal stress, suggesting that the loci may represent important switch factors which determine the direction of signaling pathways for STA1 expression in response to temperature.
Introduction
To cope with environmental extremes, plants have evolved a variety of strict controls on gene regulation to induce stress tolerance genes (Ingram and Bartels, 1996; Zhu, 2002; Chinnusamy et al., 2007; von Koskull-Döring et al., 2007). For example, most temperate plants develop freezing tolerance after prior-exposure to non-freezing temperatures; this phenomenon is known as cold acclimation (Guy, 1990). During cold acclimation, many physiological and molecular changes take place including the remodeling of cell/tissue structures, the re-programming of metabolism, and most importantly, changes in gene expression (Guy, 1990; Chinnusamy et al., 2007). The regulation of gene expression changes by cold stress is controlled by multiple transcriptional and translational programs (Chinnusamy et al., 2007; Zhu et al., 2007; Knight and Knight, 2012; Zhao et al., 2015). So far, our knowledge on regulation of cold-induced gene expression includes ICE1 and CBF transcription factors which bind the promoters of its target genes and induce the cold-responsive genes, respectively (Chinnusamy et al., 2007; Knight and Knight, 2012; Zhao et al., 2015). This transcriptional program is one of the most well-known regulatory programs in cold signal transduction for cold-induced gene expression.
Similarly, heat stress signaling for tolerance acquisition is mediated by MBF1c (Multiprotein Bridging Factor 1c), HSFs (Heat Shock Factors), and some other transcription factors (Busch et al., 2005; Kotak et al., 2007; von Koskull-Döring et al., 2007; Saidi et al., 2011; Mittler et al., 2012). The heat signaling pathways through transcription factors are not linear, but rather complex and interconnected. These transcription regulators bind either directly or indirectly to promoter cis-elements of target genes to elicit heat stress response and tolerance (Kotak et al., 2007; Saidi et al., 2011; Mittler et al., 2012).
In fact, it is well known that stress signaling pathways for stress tolerance gene transcription are interconnected (Xiong et al., 2002; Shinozaki et al., 2003; Yamaguchi-Shinozaki and Shinozaki, 2006). Thus, identification of common and specific signaling factors would help in understanding the complex controls of stress tolerance gene regulation. Indeed, many shared signaling components have been isolated in cold and osmotic stresses (Yamaguchi-Shinozaki and Shinozaki, 2006). Recently, it was reported that some sets of metabolites were altered under cold and heat stresses, commonly as well as specifically (Kaplan et al., 2004). Despite this, cold- and heat-shared/specific signaling pathways and their target gene expressions have not been intensively investigated.
The Arabidopsis nuclear STABILIZED1 (STA1) gene encodes a putative pre-mRNA splicing factor that is similar to the human U5 snRNP-associated 102-kDa protein (PRPF6) and the pre-mRNA splicing factors, PRP1p and Prp6p of fission and budding yeast; respectively. The STA1 gene is expressed throughout the whole plant body and the STA1 protein is localized in the nuclei (Lee et al., 2006; Dou et al., 2013). STA1 was shown to be involved in RNA stability, pre-mRNA and primary microRNA processing (Lee et al., 2006; Ben Chaabane et al., 2013). Recently, STA1 was reported to play a role in RNA-directed DNA methylation (Dou et al., 2013). Complete knock-out of this gene has resulted in embryo lethal (Lee et al., 2006), and there is no paralog of STA1 gene in the Arabidopsis genome (i.e., one copy gene in Arabidopsis). Taken together, these results implicated the basal and constant roles of the PRP6 splicing factor homolog STA1 in crucial and basic cellular processes (i.e., RNA metabolism and epigenetic regulations). Thus, it appears that STA1 belongs to a group of so-called “housekeeping” genes that play basal roles in cellular systems (Butte et al., 2001; She et al., 2009).
Interestingly, the sta1-1 mutant, a weak mutant allele of STA1 gene, showed stress hypersensitive phenotypes mainly in temperature stresses (Lee et al., 2006). This raised an interesting question: how does the defect in this apparent “housekeeping” gene show stress-specific hypersensitivity? In this regard, induction of STA1 mainly by low or high temperature stresses may be related to stress-specific function. Therefore, understanding the mechanisms of STA1 induction by stresses is important to elucidate the specific functions of STA1. Also, studies of STA1 regulation would provide clues about the common and specific signaling pathways for cold and heat stresses.
In order to understand the gene regulation and functional specificity of STA1, we developed an Arabidopsis STA1 gene expression monitoring system by generating bioluminescent Arabidopsis plant harboring the transgene of STA1 promoter-driven luciferase (STA1p-LUC). We confirmed that one copy of the STA1p-LUC transgene was inserted into the Arabidopsis genome and that this insertion did not interfere with neighboring gene expression. We also generated a STA1p-LUC derived mutant pool through chemical mutagenesis and isolated STA1p-LUC deregulated mutants showing similar alterations in both STA1p-LUC expression and endogenous STA1 expression under temperature stress. Categorization of STA1p-LUC deregulated mutants indicated the existence of heat or cold stress-specific regulators in addition to common genetic regulators for STA1 gene expression.
Materials and Methods
Plant Growth and Stress Treatment
Arabidopsis seeds were surface-sterilized with bleach solution (commercial bleach solution with 0.01% Tween 20) for 5 min and rinsed five times with sterile water. The seeds were planted on Murashige and Skoog (MS) medium plates that were made with full strength MS salts (Caisson Laboratories, US), 0.3% gelite (Duchefa, Netherland) and 2% sucrose (pH = 5.8). For selection plates, hygromycin was added to a final concentration of 25 mg/L to the MS media. After planting, the plates were kept at 4°C for 2 days before being transferred to 22°C under constant illumination with 70% relative humidity for germination and growth. For cold and heat treatments, 11–13 days old seedlings on MS/agar plates were incubated at 0°C (cold) or 37°C (heat) for designated time.
Construction of STA1p-LUC Transgenic Plants
The 1475 bp DNA fragment (-1475 ∼ 0 upstream from the STA1 translation start codon) of the STA1 promoter was obtained by polymerase chain reaction using two primers: F9H3.5pBamH1-F; 5′-GTGGATCC ACTTATTGTAGCAATACTTGTTCTTA-3′, and F9H3.5pH3CAM-R; 5′-CCGGT AAGCTTAACCAAACTA TAAAAATCTCT-3′. The promoter fragment was inserted into the BamHI and HindIII site of the binary plant transformation vector pCAMBIA1381Z-LUC, which contains the firefly LUC coding sequence (Millar et al., 1992) instead of its original GUS coding sequence. Arabidopsis thaliana Columbia gl1 (Col-gl1) plants lacking trichomes were transformed with the resultant STA1p-LUC construct using Agrobacterium tumefaciens GV1301 strain through floral dipping (Clough and Bent, 1998). Trichome-deficient Arabidopsis was selected due to the possible interference of trichomes with the luminescence. The T1 seeds were collected from floral-dipped plants and the transformed T1 seedlings were selected on MS/agar plates containing hygromycin 25 μg/mL (Harrison et al., 2006). Hygromycin tolerant seedlings were transferred into soil pots for continuous growth under normal growth conditions.
Genetic Analysis of STA1p-LUC Transgenic Plants
Copy numbers of STA1p-LUC transgene in the selected lines were determined in the progeny derived from either self-crossing or back-crossing. For back-crossing, the selected STA1p-LUC plants were back-crossed to the wild-type Col-gl1 plants and the resultant F1 plants were allowed to self-pollinate. The F2 progeny from either self-crossing or back-crossing were scored for segregation by their luminescence in response to low temperature (0°C for 48 h) and by their hygromycine tolerance on MS/agar plates containing hygromycin 25 μg/mL (Harrison et al., 2006).
TAIL PCR and Confirmation PCR
For TAIL PCR, three T-DNA specific primers and arbitrary degenerate primers were designed as follows: LB1, 5′-TCCGA GGGCAAAGAAATAGA-3′; LB2, 5′-TTCCTATAGGGTTTCG CTCA-3′; LB3, 5′-TTCTAATTCCTAAAACCAAAATCCA-3′ and DEG1, 5′-WGCNAGTNAGWANAA G-3′; DEG2, 5′-AWGCANGNCWGANATA-3′ (W = A or T; N = A, C, G, or T). The first, second, and third round of tail PCR reactions were carried out as described previously (Liu et al., 1995). For TAIL PCR result confirmation, each PCR was conducted with a primer combination of LB1/2/3, At3g23165-F (5′-CCGGAGGGAATGGAAAATAA-3′) and K14B15-24.42K-R (5′-GGGTCAAACTTGTTTTTCTCG-3′).
Mutagenesis and Mutant Isolation
Approximately 10,000 seeds from the selected STA1p-LUC homozygous line were imbibed overnight in water at 4°C and then soaked in the 0.35% ethyl methanesulfonate (EMS) solution. The seed tube was placed on a rotary shaker set at 30 RPM for 13 h. Treated seeds were rinsed extensively (13 times, 30 min each) in autoclaved water to remove residual EMS. The resulting M1 seeds were sown and grown to set M2 seeds. Approximately 30,000 M2 seedlings from the 326 pools of 20 plants were screened. For imaging screening, the M2 seeds were surface-sterilized and planted individually on MS plates. After the cold and heat treatments, the plates were sprayed immediately with 1 mM luciferin and placed under the luminescence imaging charge-coupled device (CCD) camera (Roper Scientific, US). Luminescence images from M2 seedlings were collected and putative mutants with altered STA1p-LUC luminescence were transferred to grow in soil and the resultant M3 seeds were used for further mutant confirmation.
Measurement of STA1p-LUC Luminescence Intensity and Gene Expression
STA1p-LUC luminescence was imaged with the lumazone luminescence imaging system (Roper Scientific, US) and luminescence intensity was quantified with the WinView software provided by the camera manufacturer (Chinnusamy et al., 2002). An equal number of pixels of each seedling were selected and total intensity was obtained from a seedling. More than 20 seedlings were quantified for statistical analysis. For gene expression analysis, total RNA was isolated with RNeasy kit (Qiagen, Germany) from 11 to 13 days old seedlings on MS/agar plates with or without stress treatment. Reverse transcription (RT)-PCR was carried out with One-step RT-PCR kit (Qiagen, Germany) for each gene with the following primers; STA1 (STA1CDS-F2, 5′-CAAGAGTCTGA CCCAGTCGAA-3′; STA1CDS-R2, 5′-AGCCAGAGAACCTCA GCTTG-3′); At3g23165 (At3g23165-F, 5′-CCGGAGGGAATGG AAAATAA-3′; At3g23165-R, 5′-TGTGTTCTTGGTTGGAACT GA-3′); At3g23167 (At3g23167-F, 5′-GCAATCAAACATGCAA TCACA-3′; At3g23167-R, 5′-GCAAAAATGGCATGCAAAC-3′) and Protein phosphatase 2A (PP2A, At1g13320) (PP2A361-F, 5′-GCGTACATCAGGAAATTCGTC-3′; PP2A361-R, 5′-GCGT GTGCGTTATATGGTTG-3′).
Results
Construction and Selection of the STA1p-LUC Arabidopsis
The 1475 bp of DNA sequence upstream from the STA1 translation initiation codon was fused to the firefly luciferase coding sequence (STA1p-LUC) and used for Agrobacterium-mediated Arabidopsis transformation to generate T1 seeds for STA1p-LUC lines. This STA1 upstream area was used as a promoter sequence because successful molecular complementation of sta1-1 by this promoter-driven STA1 coding sequence demonstrated functional activity of this region (Lee et al., 2006).
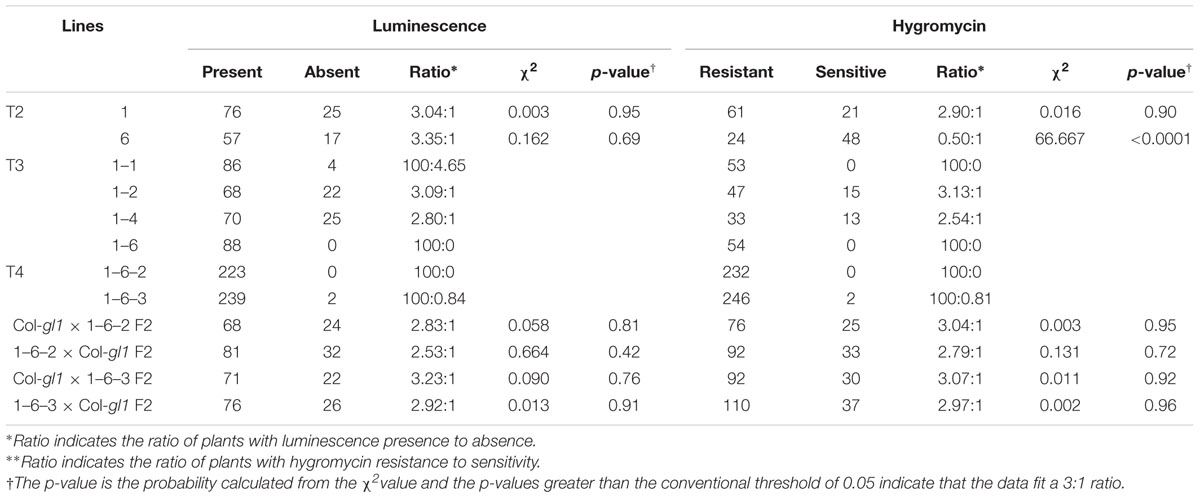
The T1 seeds were planted on MS agar media with hygromycin, and hygromycin-tolerant T1 transgenic plants were selected and transplanted to soil. These T1 plants were likely hemizygous for the STA1p-LUC transgene and the hygromycin resistance gene on the transformation vector. The T2 progeny from the hygromycin-tolerant T1 plants were further analyzed for the zygosity of functional STA1p-LUC transgene and its cosegregation with the hygromycin resistance gene. Eleven day-old T2 seedlings on plates were cold-treated at 0°C for 2 days, and two T2 lines (#1 and #6) showing a 3:1 segregation ratio of luminescence presence to absence were selected (Table 1). This 3:1 segregation suggested a single insertion of STA1p-LUC in these lines. Hygromycin resistance was also tested with these two lines (#1 and #6) and resulted in a 3:1 segregation of hygromycin resistance to non-resistance in only line #1, indicating a single insertion of the resistance gene in this line (Table 1). PCR analysis confirmed that all luminescent seedlings of line #1 contained the hygromycin resistance gene and all hygromycin-resistant seedlings of line #1 contained the luciferase transgene (data not shown) which suggested a cosegregation of STA1p-LUC and the hygromycin resistance gene. The luminescence and hygromycin resistance segregation was further examined in the T3 and T4 generations derived from line #1. In these generations, we expected to identify lines homozygous for STA1p-LUC. Lines #1–2 and #1–4 showed an approximate 3:1 segregation of hygromycin resistance to non-resistance, while nearly all seedlings of #1–1 and #1–6 were tolerant to hygromycin, indicating that the lines of #1–1 and #1–6 were homozygotes for hygromycin resistance and likely also for STA1p-LUC. Indeed, the seedlings of #1–1 and #1–6 also emitted luminescence from almost all seedlings, whereas the seedlings of #1–2 and #1–4 produced a 3:1 ratio of luminescence presence to absence. The two lines of T4 generation derived from #1–6 (#1–6–2 and #1–6–3) showed near-perfect luminescence and hygromycin resistance (Figures 1A,B). In addition, we could not detect meaningful segregation of luminescence or hygromycin resistance in the progeny seedlings of these lines in the next generation (T5; Supplementary Table S1). Among the progeny seedlings of these lines, the few seedlings that did not show luminescence and hygromycin resistance appeared to be physiological variations because PCR analysis of the seedlings revealed specific bands for both luciferase coding sequence and hygromycin resistance gene (data not shown). Taken together, genetic analysis through several generations demonstrated that the progeny lines were homozygous for STA1p-LUC and hygromycin resistance (i.e., #1–1, #1–6, #1–6–2, and #1–6–3).
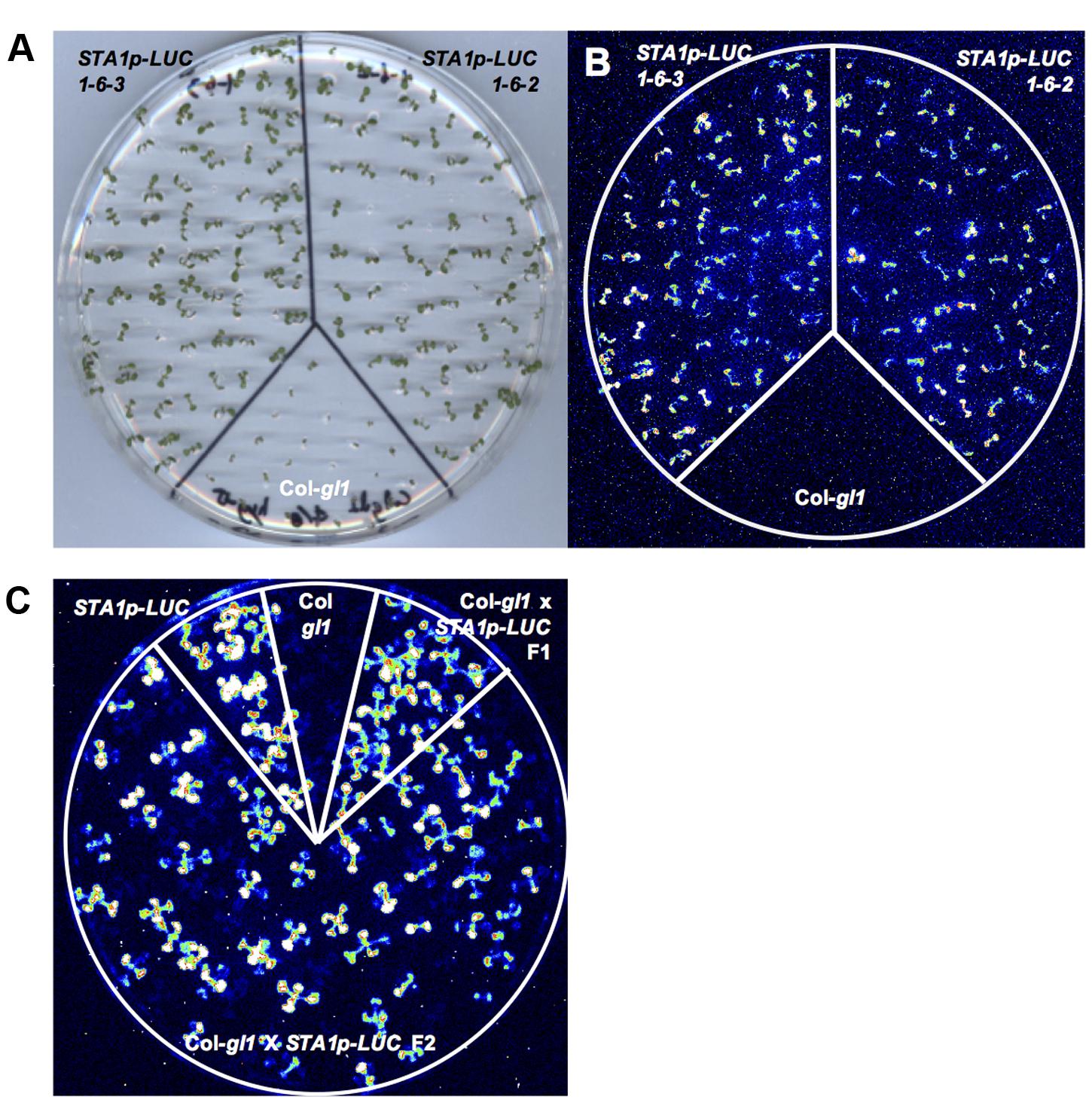
FIGURE 1. Luminescence analysis of STA1p-LUC T4 lines. (A) Plate containing 11 day-old T4 seedlings, used for luminescence image (B). (B) Luminescence image of T4 seedlings after cold stress treatment (0°C for 2 days). (C) STA1p-LUC luminescence in progeny lines derived from Col-gl1 x STA1p-LUC cross. Luminescence image of 13 days old seedlings was taken after cold stress treatment (0°C for 2 days). Col-gl1 was a negative control.
The single insertion of T-DNA was genetically reconfirmed by crossing the STA1p-LUC hymozygote lines to the original background line (Col-gl1). F2 generation from the cross between STA1p-LUC lines (#1–6–2 or #1–6–3) and Col-gl1 showed an approximate 3:1 segregation in both luminescence to no-luminescence and hygromycin-resistance to non-resistance (Table 1 and Figure 1C), which confirmed that both #1–6–2 and #1–6–3 lines contained a single homozygous insertion of STA1p-LUC transgene in each genome. For further analysis, we used #1–6–2 line as our STA1p-LUC line.
Identification of the STA1p-LUC Insertion Position
Thermal Asymmetric Interlaced (TAIL) PCR is a simple and powerful method to identify DNA sequence adjacent to known sequences (Liu et al., 1995). In order to locate the position of the STA1p-LUC transgene insertion, the flanking region adjacent to the left border of the T-DNA of STA1p-LUC was amplified by TAIL PCR. We targeted left-border neighboring DNA because left borders of T-DNA tend to remain intact more frequently than right borders during T-DNA insertion (Weigel et al., 2000). Three nested T-DNA left border specific primers (LB1, LB2, and LB3) with approximately 100 bp distance were used together with arbitrary degenerate primers (DEG1 and DEG2; Figure 2A). The resultant PCR products of each round reaction were fractionated in agarose gel, and gel-eluted DNAs from distinct bands were sequenced (Figure 2B). Blast search revealed that DNA sequences corresponded to the sequences starting from the 8,265,304th nucleotide on the chromosome 3. The locus was located between At3g23165 and At3g23167, both of which encode a member of a family of small, secreted cysteine-rich proteins similar in sequence to a pollen coat protein. Predicted T-DNA insertion was verified by PCR using two pairs of primers specific for T-DNA left border and the T-DNA-flanking Arabidopsis DNA sequences. (LB1, LB2, or LB3 and K14B15-24.42-R; At3g23165-F and K14B15-24.42K-R; Figure 2C). As shown in Figure 2D, the primer pair spanning T-DNA left border and the left-border flanking region showed distinct PCR bands, the sizes of which were gradually reduced according to the expected size reduction for the LB primers (LB1, LB2, and LB3). Also, the primers located on left and right T-DNA flanking region (At3G23165-F and K14B15-24.42K-R) produced specific PCR bands in control DNA (Col-gl1) but not in STA1p-LUC line (Figure 2D). These results confirmed that the STA1p-LUC transgene was homozygously inserted between At3g23165 and At3g21167. The insertion position of the STA1p-LUC trangene was 244 bp away from the translation start of Atg323165 and 914 bp away from the 3′ end of At3g23167 coding sequence (Figure 2C). Additionally, we tested whether the transgene altered the expression of adjacent genes (AT3G23165 and AT3G23167) using semi-quantitative RT-PCR and found that no meaningful expression alteration was caused by the STA1p-LUC transgene insertion (Figure 3). Thus, we concluded that the STA1p-LUC line was successfully generated without disrupting neighboring genes.
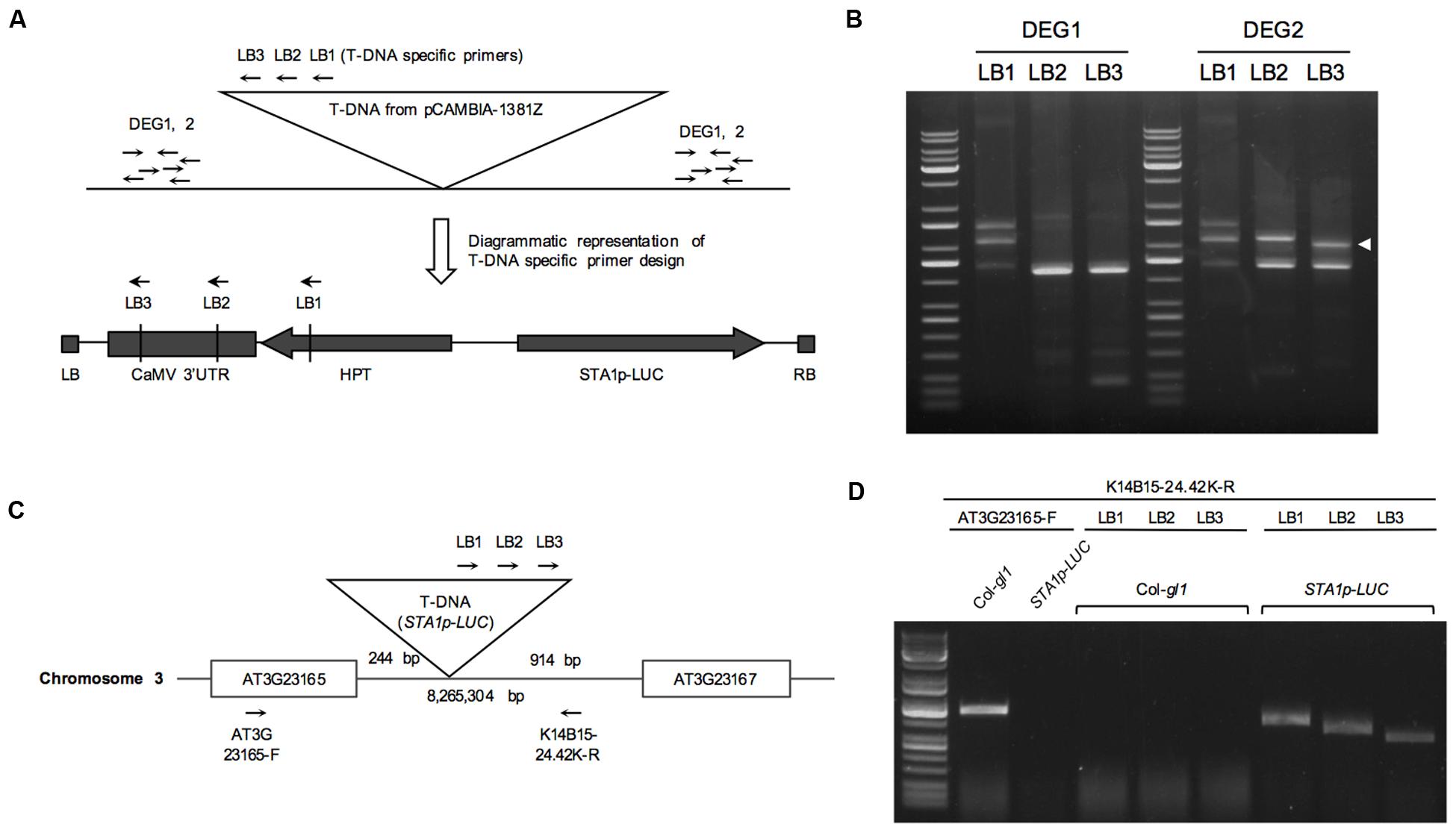
FIGURE 2. Thermal asymmetric interlaced (TAIL) PCR of STA1p-LUC. (A) Schematic diagram of TAIL PCR to show primer binding sites. (B) TAIL PCR results of STA1p-LUC. White arrowhead indicates a third round PCR product showing a TAIL PCR-typical shifted band pattern along with the first and second round PCR products in gel, which is likely a specific PCR band from the T-DNA flanking region. PCR primer pairs were labeled on the top. (C) Schematic diagram of positions of T-DNA insertion and primers designed for confirmation of TAIL PCR results. T-DNA insertion occurred at 8,265,304th bp of chromosome 3, and 244 bp and 914 bp away from AT3G23165 and AT3G23167, respectively. (D) TAIL PCR confirmation. Primer pairs used are shown on the lines and plant genomic DNA used are on top of the gel picture.
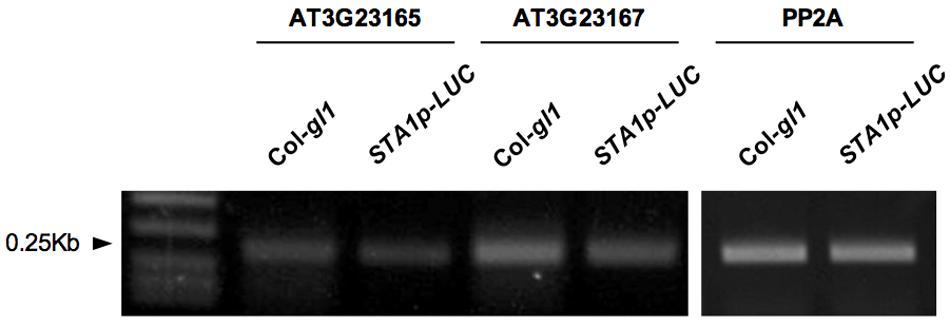
FIGURE 3. Semi-quantitative RT-PCR for STA1p-LUC neighboring genes expression. Expression of two STA1p-LUC T-DNA adjacent genes (AT3G23165 and AT3G23167) was analyzed by semi-quantitative RT-PCR. PP2A (At1g13320) was used as control.
Expression of STA1p-LUC under Temperature Stresses
Despite its “housekeeping” functions, STA1 is a temperature stress-inducible gene (Lee et al., 2006). To test whether the expression patterns of STA1p-LUC are consistent with those of the endogenous STA1 expression, we measured the STA1p-LUC luminescence intensity and the endogenous STA1 expression levels in STA1p-LUC plants under heat and cold stress conditions.
Under heat conditions, STA1p-LUC intensity increased until 24 h and then decreased gradually (Figures 4A,B). The expression patterns of endogenous STA1 gene under heat stress were similar to those of STA1p-LUC (Figure 4C). Under cold stress, the STA1p-LUC expression was gradually induced until 72 h (Figure 4E). Similarly, the endogenous STA1 expression was upregulated with a slight fluctuation under cold stress (Figure 4F). However, cold-induction of both STA1p-LUC and the endogenous STA1 was not as high as heat induction (Figure 4). Taken together, these results demonstrated that STA1p-LUC expression of the STA1p-LUC line reliably reflected endogenous STA1 expression.
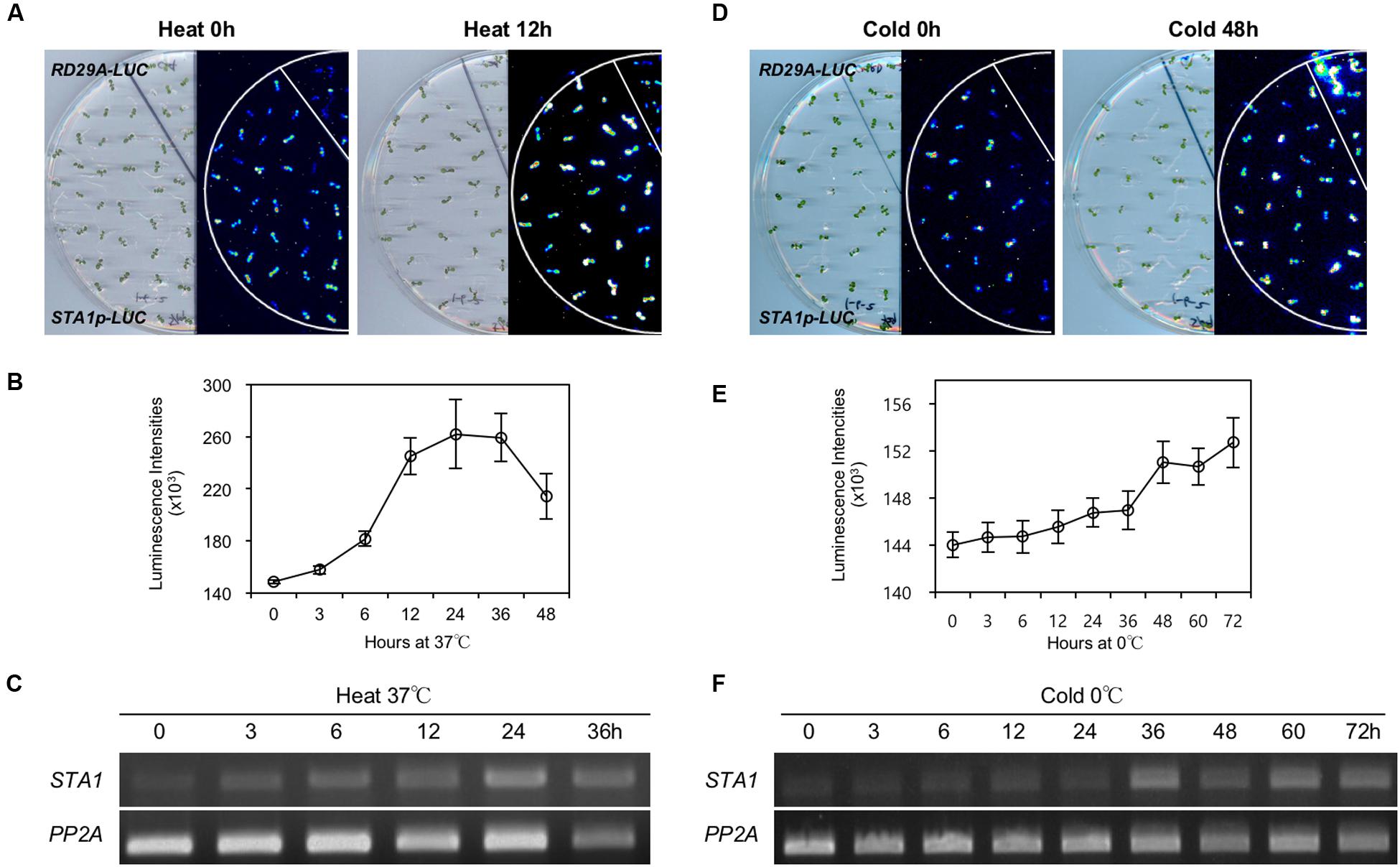
FIGURE 4. STA1p-LUC luminescence and endogenous STA1 expression in STA1p-LUC seedlings after heat or cold treatment. (A) and (D) Plate and corresponding luminescence images of STA1p-LUC seedlings before and after heat or cold treatment. RD29A-LUC was used as a positive control and did not show the heat-induction of luminescence. (B,E) Quantification of luminescence intensities of STA1p-LUC. Luminescence intensities from at least 20 seedlings were measured and averaged. Error bars represent standard deviation. (C,F) Semi-quantitative RT-PCR for the endogenous STA1 expression. PP2A (At1g13320) was used as a loading control.
Isolation of Mutants with Altered STA1p-LUC Expression under Temperature Stresses
As the STA1p-LUC line faithfully reflected endogenous STA1 expression, we decided to use the STA1p-LUC line to isolate the genetic loci for the regulators of STA1 expression. Thus, STA1p-LUC seeds were chemically mutagenized with EMS, and the resulting plants (M1 generation) were allowed to self-pollinate. The following M2 generation was screened to isolate the mutants with altered STA1p-LUC expression under cold or heat stress using an luminescence imaging system (Figures 5A–F; Chinnusamy et al., 2002). Putative M2 mutant individuals emitting altered STA1p-LUC luminescence under temperature stresses were transferred to soil and the next generation (M3) seeds from each M2 mutant were individually harvested. After the first screening with approximately 30,000 M2 seedlings from the 326 pools of 20 plants, 528 putative mutants were selected. Among these, two hundred lines survived and set seeds (M3).
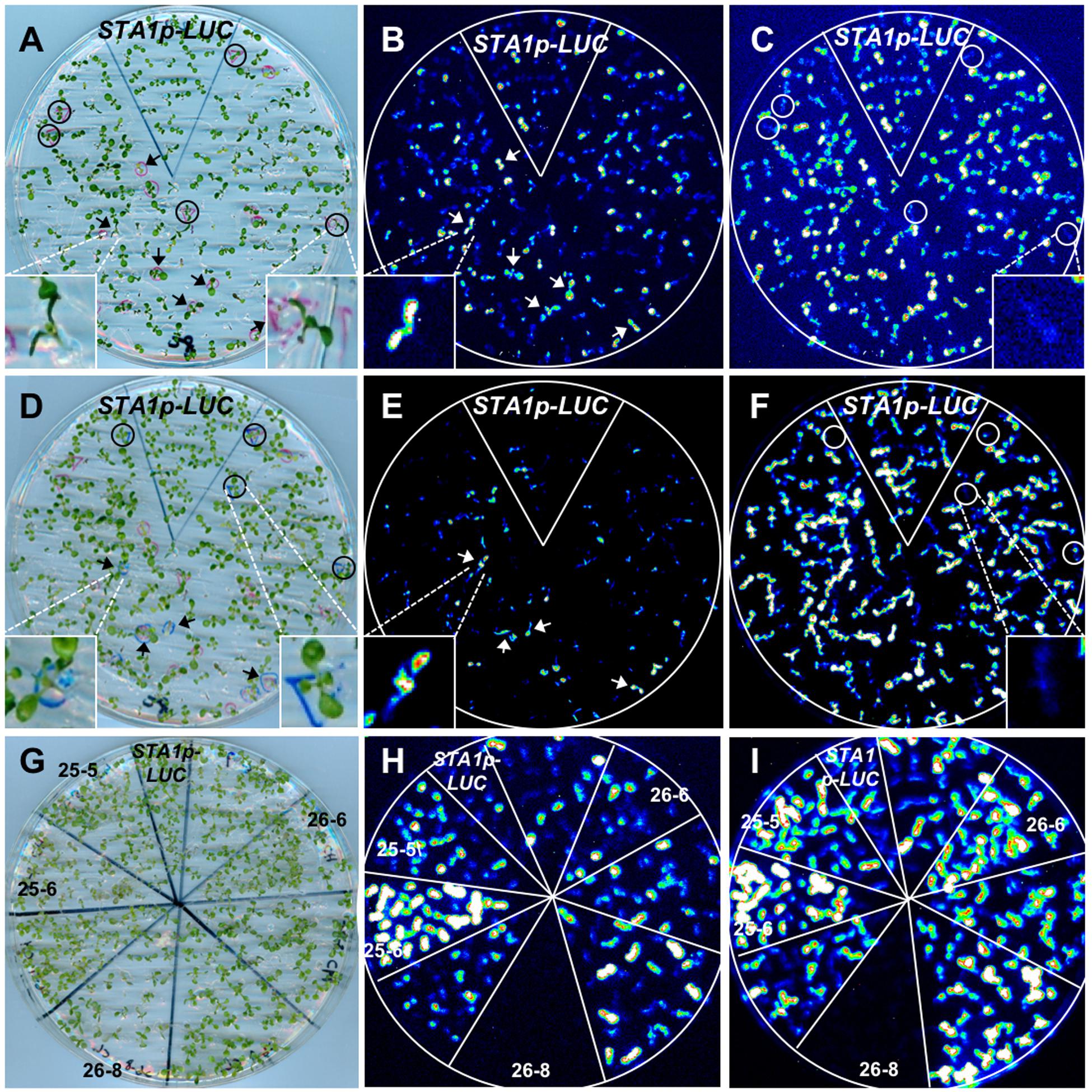
FIGURE 5. STA1p-LUC deregulated mutant screening. (A) Plate containing 11 day-old M2 seedlings for (B) and (C). (B) and (C) Luminescence image taken after cold stress treatment (0°C for 36 h). (D) Plate containing 13 day-old seedlings for (E) and (F). The same, cold-treated plate (A) was used for heat stress after 1 day adaptation at normal temperature. (E) and (F) Luminescence image taken after heat treatment (37°C for 15 h). In comparison with STA1p-LUC, putative mutants with higher luminescence were marked with arrows and those with lower luminescence were marked with circles. Insets in (A–F) are the representative images of either seedlings or luminescence. Left inset in (D) shows a seedling with one leaf slightly overlapped by a leaf of a nearby seedling. (G) Plate containing 11 day-old putative mutant seedlings (M3 generation) for confirmation (H,I). (H) and (I) Luminescence images taken after cold stress (H) and heat stress (I). The confirmed mutants without segregation were labeled with mutant numbers and the results showed two tis (25-5 and 25-6), one his (26-6), and one trs (26-8).
In the M3 generation, confirmation imaging was conducted to validate the putative mutants (Figures 5G–I). M3 putative mutant lines were confirmed as mutants when all siblings of the lines showed the altered STA1p-LUC expression. In addition, we tested the expression levels of endogenous STA1 in these confirmed lines (M3) and identified 23 STA1p-LUC deregulated mutants having the same alterations of endogenous STA1 expression under temperature stresses (Figure 6). These mutants were then classified on the basis of luminescence intensities under either low or high temperatures (Table 2).
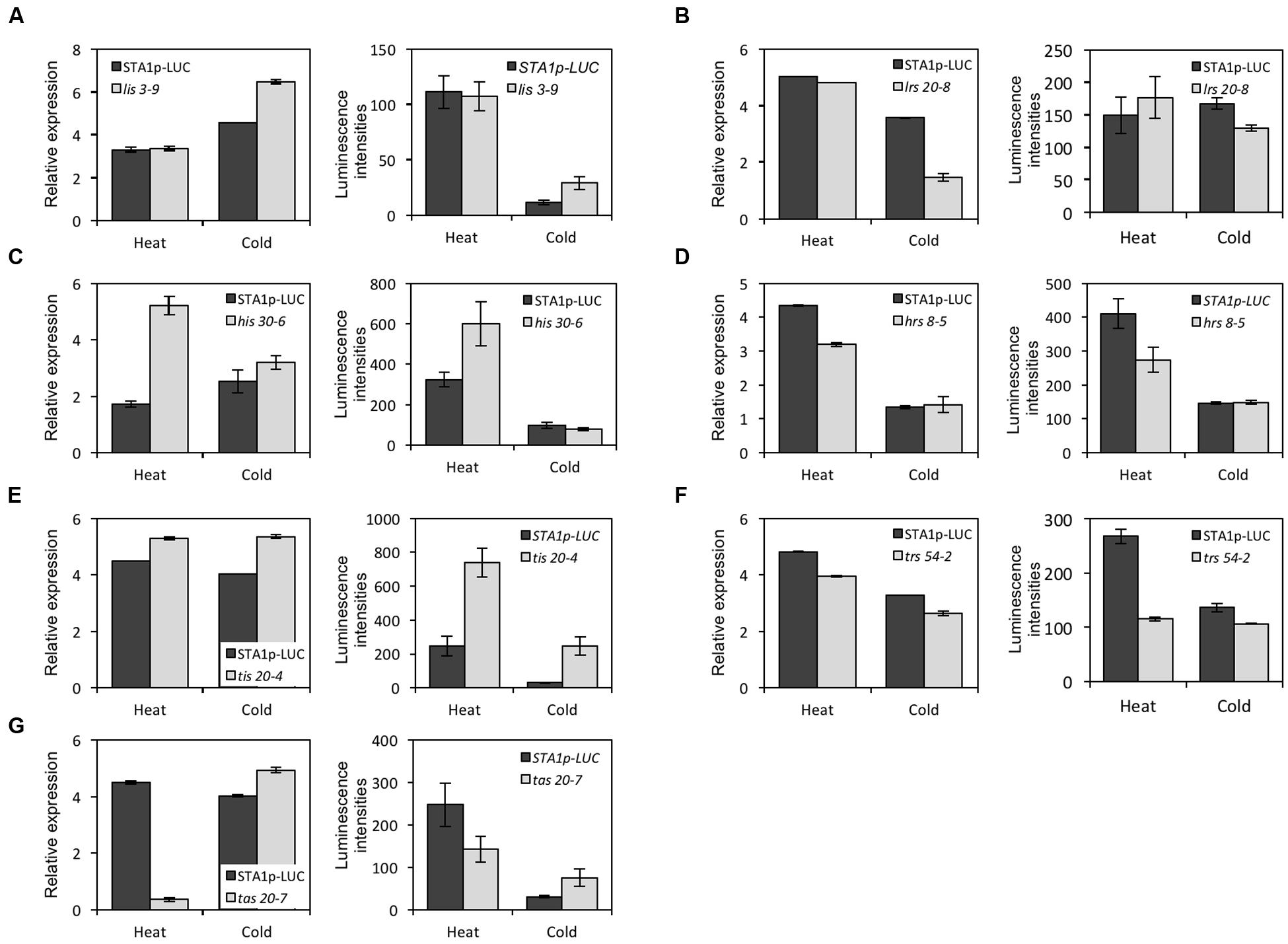
FIGURE 6. The endogenous STA1 expression and the STA1p-LUC luminescence in confirmed mutants in response to heat and cold stresses. Graphs in the left column in each letter-labeled set are the endogenous STA1 expression levels by quantitative RT-PCR (three replicates; bar = standard deviation) and graphs in the right column are STA1p-LUC luminescence intensities. 20–25 seedlings were used to measure expression levels. Error bars represent standard deviation. (A) lis (low temperature increased STA1p-LUC expression). (B) lrs (low temperature reduced STA1p-LUC expression). (C) his (high temperature increased STA1p-LUC expression). (D) hrs (high temperature reduced STA1p-LUC expression). (E) tis (temperature increased STA1p-LUC expression). (F) trs (temperature reduced STA1p-LUC expression). (G) tas (temperature altered STA1p-LUC expression).
The classified mutants were named as follows: lis (Low-temperature Increased STA1 promoter-driven luciferase expression), lrs (Low temperature Reduced STA1 promoter-driven luciferase expression), his (High temperature Increased STA1 promoter-driven luciferase expression), hrs (High temperature Reduced STA1 promoter-driven luciferase expression), tas (Temperature Altered STA1 promoter-driven luciferase expression), tis (Temperature Increased STA1 promoter-driven luciferase expression), trs (Temperature Reduced STA1 promoter-driven luciferase expression).
Some mutants showed alteration of STA1p-LUC expression by both cold and heat stresses (tis, trs, and tas), while others showed the deregulation of STA1p-LUC specifically by either cold or heat (lis, lrs, his, and hrs). These observations suggested the existence of common genetic regulators as well as heat or cold stress-specific regulators for STA1 expression. Interestingly, tas mutants were identified due to their opposite-directional deregulation of STA1p-LUC depending on the kind of thermal stress. Two mutants showed higher luminescence intensities of STA1p-LUC under cold and lower STA1p-LUC expression under heat in comparison to its background line, and these mutants also displayed similar alterations of endogenous STA1 expression (i.e., increased STA1 expression in cold and reduced STA1 expression in heat). This result suggests that these tas mutants may be defective in important switch genes directing thermal-stress signaling pathways for STA1 expression.
Discussion
The Arabidopsis STA1 gene is present as a single copy in the genome and encodes a pre-mRNA splicing factor with high homology to the budding yeast pre-mRNA splicing factors, Prp6p and the human U5 snRNP-associated 102-kDa protein, PRPF6. Arabidopsis is known to have a total of 14 U5 snRNP specific proteins including STA1 (Wang and Brendel, 2004). Among them, our database search revealed that only STA1 and one Brr2 homolog (At2g42270) were highly induced by thermal stresses. Although the sta1-1 mutants have shown temperature stress-hypersensitivity, the mutant phenotypes of the temperature-induced Brr2 homolog (At2g42270) have not been reported so far. Thus, STA1 provides a good opportunity to study how this seemingly “housekeeping” gene is specifically required under unfavorable temperatures, and how thermal stress signaling pathways are interconnected. One possible explanation for thermal stress-specific phenotypes of sta1-1 is that sta1-1 is a temperature sensitive allele, but sta1-1 has shown developmental defects under normal conditions (Lee et al., 2006). Thus, sta1-1 mutation does not seem to be a temperature sensitive allele which should otherwise display normal phenotypes under normal conditions. Another explanation is that STA1 itself is specifically required under unfavorable temperatures; thus, thermal stress-induction of STA1 might be correlated with its function in cold or heat stress.
As a first step toward understanding STA1 induction and specificity under cold and heat, we generated the STA1p-LUC bioluminescent Arabidopsis plant. The faithfulness of our STA1p-LUC line system was verified by comparing the STA1p-LUC expression patterns with endogenous STA1 expressions. In addition, the homozygous single-copy STA1p-LUC insertion was confirmed not to interfere with the insertion-neighboring gene expression (Figure 3). In some cases, foreign gene insertion with 35S promoters resulted in altered expression of the genes adjacent to the insertion (Yoo et al., 2005; Zheng et al., 2007; Singer et al., 2010, 2011).
Several kinds of bioluminescent Arabidopsis plants have been developed to study gene expression regulation (Ishitani et al., 1997; Chinnusamy et al., 2002; Yao et al., 2014). In our study, the mutagenesis of STA1p-LUC lines and mutant screening identified many luminescence-altered mutants under cold and heat stresses. While some mutants altered STA1p-LUC expression under only cold or heat stress conditions, others affected STA1p-LUC expression under both. These results suggested the existence of independent signaling pathways for each stress, and also the presence of diverse cross-talks and shared signaling pathways between cold and heat stress responses. Heat shock transcription factors (HSF) and heat shock proteins (HSP) are induced by multiple stresses and are thought to be an interacting point between heat and non-heat stress responses (Swindell et al., 2007). Swindell et al. (2007) suggested that HSF and HSP induction by multiple stresses might be mediated by secondary oxidative stress. Interestingly, STA1 was strongly induced primarily by temperature stresses but remained almost unaffected by other stresses, implicating that STA1 induction by thermal stresses might not be mediated by oxidative stress. Taken together, our STA1p-LUC mutants might only represent the genetic factors involved in more direct thermal-stress signaling pathways for STA1 regulation rather than indirect oxidative pathways.
Multiple cross-talks and specific signaling pathways among the stress signaling pathways are not uncommon. For example, Arabidopsis full-length cDNA microarray analyses revealed that more than half of drought-induced genes were also induced by high salt stress, implicating the presence of cross-talks between salt and drought stress signaling for gene induction (Seki et al., 2002). However, to our knowledge, direct comparisons of whole genome expression profiles between heat and cold stresses have not been carried out. This might be because of the assumption that plants may not undergo such dramatic temperature changes (e.g., -5 to 35°C). Nevertheless, public genomics data suggested that there are some common gene sets that are regulated by both cold and heat. Indeed, the comparative metabolomics revealed that many metabolites were commonly altered by heat and cold; some were changed only by each specific stress (Kaplan et al., 2004). These findings suggest that plants have mechanisms which use common and specific signaling networks for such dramatic temperature changes. Changes in membrane fluidity, internal Ca2+ levels and protein unfolding levels are among the common cellular responses that happen early during both heat and cold stresses (Sung et al., 2003; Ruelland and Zachowski, 2010). Therefore, some of our STA1p-LUC mutants deregulated by both cold and heat might have defects in these interconnection points. Interestingly, we isolated two tas mutants which showed STA1 up-regulation by cold, but down-regulation by heat. These genetic loci may represent the decision-making branching points in the signaling pathways. Cloning of genes responsible for STA1p-LUC deregulation in our mutants will help in identifying common and specific thermal-stress signaling pathways for STA1 regulation and shed light on the regulation of temperature stress-induction of this “housekeeping” gene.
Conclusion
In this study, we have generated a bioluminescent Arabidopsis plant harboring a single copy of a STA1 promoter-driven firefly luciferase (STA1p-LUC), which faithfully reflected the gene expression patterns of endogenous STA1 under cold and heat stresses. Mutagenesis was performed using STA1p-LUC as a background line, and mutants showing deregulation of the transgenic STA1p-LUC and endogenous STA1 gene under cold and heat stresses were successfully isolated. The isolated mutants suggested the existence of genetic loci for stress-specific and shared signaling components of STA1 regulation. These genetic loci may also include important genetic switches to determine the direction of cold and heat signaling pathways.
Author Contributions
B-hL conceived and designed the research; S-iY, CC, and J-HH performed the experiments; S-iY, J-HH and B-hL discussed the results and wrote the paper.
Funding
The research was supported by Next-Generation BioGreen21 Program (PJ011006), Rural Development Administration, Republic of Korea.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Authors thank the lab members for advice and help, and Dr. Dae-Jin Yun (Gyeongsang National University, Korea) for pCAMBIA1381Z-LUC.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2016.00618
References
Ben Chaabane, S., Liu, R., Chinnusamy, V., Kwon, Y., Park, J.-H., Kim, S. Y., et al. (2013). STA1, an Arabidopsis pre-mRNA processing factor 6 homolog, is a new player involved in miRNA biogenesis. Nucleic Acids Res. 41, 1984–1997. doi: 10.1093/nar/gks1309
Busch, W., Wunderlich, M., and Schöffl, F. (2005). Identification of Novel Heat Shock Factor-Dependent Genes and Biochemical Pathways in Arabidopsis thaliana - Busch - 2004 - The Plant Journal. Hoboken, NJ: Wiley Online Library.
Butte, A. J., Dzau, V. J., and Glueck, S. B. (2001). Further defining housekeeping, or “maintenance,” genes Focus on ‘A compendium of gene expression in normal human tissues’. Physiol. Genomics 7, 95–96. doi: 10.1152/physiolgenomics.00040.2001
Chinnusamy, V., Stevenson, B., Lee, B., and Zhu, J. K. (2002). Screening for gene regulation mutants by bioluminescence imaging. Sci. Signal. 2002:l10. doi: 10.1126/stke.2002.140.pl10
Chinnusamy, V., Zhu, J., and Zhu, J.-K. (2007). Cold stress regulation of gene expression in plants. Trends Plant Sci. 12, 444–451. doi: 10.1016/j.tplants.2007.07.002
Clough, S. J., and Bent, A. F. (1998). Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743. doi: 10.1046/j.1365-313x.1998.00343.x
Dou, K., Huang, C. F., Ma, Z. Y., Zhang, C. J., Zhou, J. X., Huang, H. W., et al. (2013). The PRP6-like splicing factor STA1 is involved in RNA-directed DNA methylation by facilitating the production of Pol V-dependent scaffold RNAs. Nucleic Acids Res. 41, 8489–8502. doi: 10.1093/nar/gkt639
Guy, C. L. (1990). Cold acclimation and freezing stress tolerance: role of protein metabolism. Annu. Rev. Plant Physiol. Plant Mol. Biol. 41, 187–223. doi: 10.1146/annurev.pp.41.060190.001155
Harrison, S. J., Mott, E. K., Parsley, K., Aspinall, S., Gray, J. C., and Cottage, A. (2006). A rapid and robust method of identifying transformed Arabidopsis thaliana seedlings following floral dip transformation. Plant Methods 2:19. doi: 10.1186/1746-4811-2-19
Ingram, J., and Bartels, D. (1996). The molecular basis of dehydration tolerance in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 377–403. doi: 10.1146/annurev.arplant.47.1.377
Ishitani, M., Xiong, L., Stevenson, B., and Zhu, J. K. (1997). Genetic analysis of osmotic and cold stress signal transduction in Arabidopsis: interactions and convergence of abscisic acid-dependent and abscisic acid-independent pathways. Plant Cell 9, 1935–1949. doi: 10.1105/tpc.9.11.1935
Kaplan, F., Kopka, J., Haskell, D. W., Zhao, W., Schiller, K. C., Gatzke, N., et al. (2004). Exploring the temperature-stress metabolome of Arabidopsis. Plant Physiol. 136, 4159–4168. doi: 10.1104/pp.104.052142
Knight, M. R., and Knight, H. (2012). Low-temperature perception leading to gene expression and cold tolerance in higher plants. New Phytol. 195, 737–751. doi: 10.1111/j.1469-8137.2012.04239.x
Kotak, S., Larkindale, J., Lee, U., von, P., Vierling, E., and Scharf, K.-D. (2007). Complexity of the heat stress response in plants. Curr. Opin. Plant Biol. 10, 310–316. doi: 10.1016/j.pbi.2007.04.011
Lee, B.-H., Kapoor, A., Zhu, J., and Zhu, J.-K. (2006). STABILIZED1, a stress-upregulated nuclear protein, is required for pre-mRNA splicing, mRNA turnover, and stress tolerance in Arabidopsis. Plant Cell 18, 1736–1749. doi: 10.1105/tpc.106.042184
Liu, Y. G., Mitsukawa, N., Oosumi, T., and Whittier, R. F. (1995). Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J. 8, 457–463. doi: 10.1046/j.1365-313X.1995.08030457.x
Millar, A. J., Short, S. R., Hiratsuka, K., Chua, N.-H., and Kay, S. A. (1992). Firefly luciferase as a reporter of regulated gene expression in higher plants. Plant Mol. Biol. Rep. 10, 324–337. doi: 10.1007/BF02668909
Mittler, R., Finka, A., and Goloubinoff, P. (2012). How do plants feel the heat? Trends Biochem. Sci. 37, 118–125. doi: 10.1016/j.tibs.2011.11.007
Ruelland, E., and Zachowski, A. (2010). How plants sense temperature. Environ. Exp. Bot. 69, 225–232. doi: 10.1016/j.envexpbot.2010.05.011
Saidi, Y., Finka, A., and Goloubinoff, P. (2011). Heat perception and signalling in plants: a tortuous path to thermotolerance. New Phytol. 190, 556–565. doi: 10.1111/j.1469-8137.2010.03571.x
Seki, M., Narusaka, M., Ishida, J., Nanjo, T., Fujita, M., Oono, Y., et al. (2002). Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J. 31, 279–292. doi: 10.1046/j.1365-313X.2002.01359.x
She, X., Rohl, C. A., Castle, J. C., Kulkarni, A. V., Johnson, J. M., and Chen, R. (2009). Definition, conservation and epigenetics of housekeeping and tissue-enriched genes. BMC Genomics 10:269. doi: 10.1186/1471-2164-10-269
Shinozaki, K., Yamaguchi-Shinozaki, K., and Seki, M. (2003). Regulatory network of gene expression in the drought and cold stress responses. Curr. Opin. Plant Biol. 6, 410–417. doi: 10.1016/S1369-5266(03)00092-X
Singer, S. D., Cox, K. D., and Liu, Z. (2010). Both the constitutive Cauliflower Mosaic Virus 35S and tissue-specific AGAMOUS enhancers activate transcription autonomously in Arabidopsis thaliana. Plant Mol. Biol. 74, 293–305. doi: 10.1007/s11103-010-9673-9
Singer, S. D., Liu, Z., and Cox, K. D. (2011). Minimizing the unpredictability of transgene expression in plants: the role of genetic insulators. Plant Cell Rep. 31, 13–25. doi: 10.1007/s00299-011-1167-y
Sung, D. Y., Kaplan, F., Lee, K.-J., and Guy, C. L. (2003). Acquired tolerance to temperature extremes. Trends Plant Sci. 8, 179–187. doi: 10.1016/S1360-1385(03)00047-5
Swindell, W. R., Huebner, M., and Weber, A. P. (2007). Transcriptional profiling of Arabidopsis heat shock proteins and transcription factors reveals extensive overlap between heat and non-heat stress response pathways. BMC Genomics 8:125. doi: 10.1186/1471-2164-8-125
von Koskull-Döring, P., Scharf, K.-D., and Nover, L. (2007). The diversity of plant heat stress transcription factors. Trends Plant Sci. 12, 452–457. doi: 10.1016/j.tplants.2007.08.014
Wang, B.-B., and Brendel, V. (2004). The ASRG database: identification and survey of. Genome Biol. 5:R102. doi: 10.1186/gb-2004-5-9-r69
Weigel, D., Ahn, J. H., Blázquez, M. A., Borevitz, J. O., Christensen, S. K., Fankhauser, C., et al. (2000). Activation tagging in Arabidopsis. Plant Physiol. 122, 1003–1013. doi: 10.1104/pp.122.4.1003
Xiong, L., Schumaker, K. S., and Zhu, J.-K. (2002). Cell signaling during cold, drought, and salt stress. Plant Cell 14(Suppl.), S165–S183. doi: 10.1105/tpc.000596
Yamaguchi-Shinozaki, K., and Shinozaki, K. (2006). Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol. 57, 781–803. doi: 10.1146/annurev.arplant.57.032905.105444
Yao, Y., Dong, C.-H., Yi, Y., Li, X., Zhang, X., and Liu, J. (2014). Regulatory function of AMP1 in ABA biosynthesis and drought resistance in arabidopsis. J. Plant Biol. 57, 117–126. doi: 10.1007/s12374-013-0475-x
Yoo, S. Y., Bomblies, K., Yoo, S. K., Yang, J. W., Choi, M. S., Lee, J. S., et al. (2005). The 35S promoter used in a selectable marker gene of a plant transformation vector affects the expression of the transgene. Planta 221, 523–530. doi: 10.1007/s00425-004-1466-4
Zhao, C., Lang, Z., and Zhu, J.-K. (2015). Cold responsive gene transcription becomes more complex. Trends Plant Sci. 20, 466–468. doi: 10.1016/j.tplants.2015.06.001
Zheng, X., Deng, W., Luo, K., Duan, H., Chen, Y., McAvoy, R., et al. (2007). The cauliflower mosaic virus (CaMV) 35S promoter sequence alters the level and patterns of activity of adjacent tissue- and organ-specific gene promoters. Plant Cell Rep. 26, 1195–1203. doi: 10.1007/s00299-007-0307-x
Zhu, J., Dong, C.-H., and Zhu, J.-K. (2007). Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr. Opin. Plant Biol. 10, 290–295. doi: 10.1016/j.pbi.2007.04.010
Keywords: cold stress, heat stress, thermal stress, STA1, STABILIZED1, luminescence screening, housekeeping gene
Citation: Yu S-i, Han J-H, Chhoeun C and Lee B-h (2016) Genetic Screening for Arabidopsis Mutants Defective in STA1 Regulation under Thermal Stress Implicates the Existence of Regulators of Its Specific Expression, and the Genetic Interactions in the Stress Signaling Pathways. Front. Plant Sci. 7:618. doi: 10.3389/fpls.2016.00618
Received: 20 March 2016; Accepted: 22 April 2016;
Published: 10 May 2016.
Edited by:
Mohammad Anwar Hossain, Bangladesh Agricultural University, BangladeshReviewed by:
Jianhua Zhu, University of Maryland, College Park, USAZofia Szweykowska-Kulinska, Adam Mickiewicz University, Poland
Copyright © 2016 Yu, Han, Chhoeun and Lee. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Byeong-ha Lee, Ynllb25naGFAc29nYW5nLmFjLmty
 Si-in Yu
Si-in Yu Jin-Hee Han
Jin-Hee Han Chanvotey Chhoeun
Chanvotey Chhoeun Byeong-ha Lee
Byeong-ha Lee