One of the last mysteries in plant physiology is the efflux of sugars from mesophyll cells into the apoplasmic space. Depending on the strategy of long-distance assimilate transport, text books divide plants into symplasmic and apoplasmic phloem loaders. In both cases large concentration gradients are built up between mesophyll cells and the sieve element–companion cell complex (SE–CCC) of the phloem. In case of apoplasmic loaders this gradient is achieved by active transport of sucrose from the apoplasm into the SE–CCC. This uptake is mediated by well characterized plasma membrane sucrose–H+ symporters (Kühn and Grof, 2010). As a consequence of this model, sucrose synthesized within mesophyll cells must be exported into the apoplasmic space, but the efflux mechanism is as yet unknown (Lemoine, 2000; Williams et al., 2000). In addition, sugar efflux must play a crucial role in fuelling pollen and embryo development as well as nectar production and secretion of carbohydrates to the rhizosphere. Furthermore, phytopathogenic bacteria usually colonize the apoplasmic space and are dependent on nutrient uptake from the host. Hence, it is very likely that efflux carriers from the host are required for bacterial growth. Despite these important functions, attempts to isolate and characterize sugar efflux carriers failed so far, probably because of unsuitable screening systems.
Using a new and elegant screening system, Chen et al. (2010) were able to isolate and characterize the first sugar efflux carrier from plants and later on could demonstrate that these efflux carriers belong to a novel transporter family with homologs in Caenorhabditis elegans and humans. The screening system was based on a mammalian cell line (HEK293T) co-expressing a genetically encoded glucose sensor (Frommer et al., 2009) and a library of polytopic A. thaliana membrane proteins of unknown function as predicted by the membrane protein database Aramemnon (Schwacke et al., 2003). This system allowed the sensitive detection of glucose uptake and let to the identification of AtSWEET1. In Arabidopsis thaliana, AtSWEET1 is a member of a gene family comprising 17 members. Mutant phenotypes of some members of the SWEET family have already been described in A. thaliana and rice. Mutation of AtSWEET8 (ruptured pollen grain 1, RPG1) for example, prevents microsporocyte development by inhibiting glucose efflux from tapetum cells (Guan et al., 2008). Interestingly, several members of the AtSWEET gene family are inducible by pathogens. This let to the hypothesis that some pathogens might hijack host efflux systems to support their own growth in the apoplastic space of infected host plants. Support for this hypothesis comes from rice. Mutations in OsSWEET11 (Xa13) results in resistance against late blight (Chu et al., 2006). Moreover, in wild-type rice plants, Xanthomonas oryzae, the causative agent of late blight, activates OsSWEET11 expression by binding of TAL effectors to promoter elements of the OsSWEET11 gene (Römer et al., 2010). Mutations in these binding sites lead to the failure of gene induction and hence prevent colonization of the host. Interestingly, mutations in the same gene also result in reduced fertility of pollen. Thus pathogens seem to be able to reprogram host cells to support their nutrient supply by interfering with the host transport system.
Taken together, assigning a function to the SWEET gene family opens up exiting perspectives for both plant physiology and plant–pathogen interaction. It could clearly be shown that SWEETS are in involved in glucose efflux and play a crucial role during pollen development and plant–pathogen interaction. Since the SWEETS transport glucose, this still leaves the question which proteins are responsible for sucrose efflux around the phloem or from the seed coat. However the base has been laid – one could use the FRET sucrose sensors (Lager et al., 2006) to use the same principal approach as was used for identifying the SWEETS, namely to screen a library of unknowns for novel sucrose transporters that might be responsible for sugar efflux around the phloem. But it seems to be clear that on the basis of the important role of members of the SWEET family during plant–pathogen interaction, novel strategies in plant protection become feasible. Therefore, the article by Chen et al. (2010) represents an exiting milestone in furthering our understanding of transport processes in plants and their relevance in plant–microbe interactions (for illustration see Figure 1).
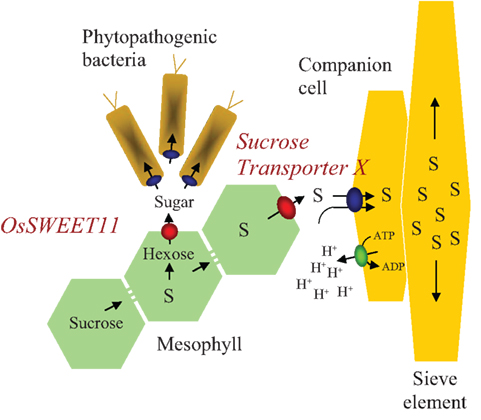
Figure 1. Schematic representation of sugar transport processes during plant–bacterium interaction and phloem loading. S, sucrose; H+, proton.
References
Chen, L.-Q., Hou, B.-H., Lalonde, S., Takanaga, H., Hartung, M. L., Qu, X.-Q., Guo, W.-J., Kim, J.-G., Underwood, W., Chaudhuri, B., Chermak, D., Antony, G., White, F. F., Sommerville, S. C., Mudgett, M. B., and Frommer, W. B. (2010). Sugar transporters for intercellular exchange and nutrition of pathogens. Nature 468, 527–534.
Chu, Z., Yuan, M., and Yao, J. (2006). Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev. 20, 1250–1255.
Frommer, W. B., Davidson, M. W., and Campbell, R. E. (2009). Genetically encoded biosensors based on engineered fluorescent proteins. Chem. Soc. Rev. 38, 2833–2841.
Guan, Y. F., Huang, X. Y., Zhu, J., Gao, J. F., Zhang, H. X., and Yang, Z. N. (2008). Ruptured pollen grain, a member of the MtN3/saliva gene family, is crucial for exine pattern formation and cell integrity of microspores in Arabidopsis. Plant Physiol. 147, 852–863.
Kühn, C., and Grof, C. P. (2010). Sucrose transporters of higher plants. Curr. Opin. Plant Biol. 13, 288–298.
Lager, I., Looger, L. L., Hilpert, M., Lalonde, S., and Frommer, W. B. (2006). Conversion of a putative Agrobacterium sugar-binding protein into a FRET sensor with high selectivity for sucrose. J. Biol. Chem. 281, 30875–30883.
Lemoine, R. (2000). Sucrose transporters in plants: update on function and structure. Biochim. Biophys. Acta 1465, 246–262.
Römer, P., Recht, S., Strauss, T., Elsaesser, J., Schornack, S., Boch, J., Wang, S., and Lahaye, T. (2010). Promoter elements of rice susceptibility genes are bound and activated by specific TAL effectors from the bacterial blight pathogen, Xanthomonas oryzae pv. oryzae. New Phytol. 187, 1048–1054.
Schwacke, R., Schneider, A., van der Graff, E., Fischer, K., Catoni, E., Desimone, M., Frommer, W. B., Flügge, U.-I., and Kunze, R. (2003). ARAMEMNON, a novel database for Arabidopsis integral membrane proteins. Plant Physiol. 131, 16–26.
Citation: Sonnewald U (2011) SWEETS – the missing sugar efflux carriers. Front. Plant Sci. 2:7. doi: 10.3389/fpls.2011.00007
Received: 18 March 2011;
Accepted: 21 March 2011;
Published online: 31 March 2011.
Copyright: © 2011 Sonnewald. This is an open-access article subject to a non-exclusive license between the authors and Frontiers Media SA, which permits use, distribution and reproduction in other forums, provided the original authors and source are credited and other Frontiers conditions are complied with.
*Correspondence:dXNvbm5lQGJpb2xvZ2llLnVuaS1lcmxhbmdlbi5kZQ==
