- Instituto de Biotecnología, Universidad Nacional Autónoma de México, Cuernavaca, México
The importance of the family of high affinity ammonium transporters is demonstrated by the presence of these proteins in all domains of life, including bacteria, archaea, fungi, plants, and humans. The majority of the proteins that have been studied from this family show high affinity and selectivity for ammonium, are impermeable to alkaline cations, saturate rapidly at low millimolar concentrations and most of them, are also permeable to methylammonium. Crystallization of homologue proteins from bacteria and archaea has demonstrated that the functional entity corresponds to a trimer, with each monomer maintaining a conductive pore. Through molecular modeling, it has been demonstrated that even though the identity of the proteins between bacteria/archaea with those from plants is below 25%, the latter seem to maintain similar tertiary and quaternary structures, an observation that has helped to address the functionality of conserved residues by means of mutational analysis. Results have shown that changes in the extracellular binding site of some plant transporters may result in their inhibition or reduction in transport activity, while in Escherichia coli, dissimilar replacements like Phe/Ala or Ser/Leu that eliminate possible π-interactions or H-bonds with ammonium, respectively, lead to more active transporters. Active mutants with changes in the pair of conserved His in the center of the transporter suggest these residues are dispensable. Additional mutations have identified other important amino acids, both in the entrance of the pore and in cytoplasmic loops. Regulation of this family of transporters can be achieved by interactions of the C-terminal with cytoplasmic loops within the same monomer, or with a neighbor in the trimer. Depending on the interacting residues, these contacts may lead to the activation or inhibition of the protein. The aim of this review is to critically evaluate the newest findings on the role of the proposed amino acids that structure the ammonium pathway, as well as highlight the importance of additional residues that have been identified through mutational analyses.
Introduction
Ammonium is the preferred nitrogen source for many plants, and its transport across cellular membranes is carried out by a family of integral membrane proteins that is conserved throughout all domains of life, the AMT/MEP/Rh family (Ammonium Transporter/Methylammonium Permease/mammalian Rhesus proteins; Wirén and Merrick, 2004; Winkler, 2006). Members of this family of transporters share several properties, among them, high affinity (Km in the micromolar range) and high selectivity for ammonium [transporting exclusively ammonium and its methylated analog, methylammonium (MeA), but no other monovalent cation] and saturation at ammonium concentrations ≤1 mM (Gazzarrini et al., 1999; Shelden et al., 2001; Ludewig et al., 2002; D’Apuzzo et al., 2004; Wood et al., 2006; Couturier et al., 2007; Boeckstaens et al., 2008; Ortiz-Ramirez et al., 2011). In spite of these common characteristics, it is not clear what is the chemical species that is transported, ammonium or ammonia, or the actual transport mechanism represented by these proteins, either a channel or a transporter (Khademi et al., 2004; Conroy et al., 2007). However, important progress in the understanding of ammonium transporters has been derived from the crystal structure of AmtB from Escherichia coli (Khademi et al., 2004; Zheng et al., 2004; Conroy et al., 2007) and Amt1 from Archaeoglobus fulgidus (Andrade et al., 2005). These studies have demonstrated that the ammonium transporters are conformed as homotrimers, with each monomer containing a central hydrophobic channel, proposed to be the pathway through which the uncharged ammonia is transported (Khademi et al., 2004; Zheng et al., 2004; Andrade et al., 2005). Taking as a reference the structure for the E. coli ammonium transporter (EcAmtB), it is proposed that the putative selective ammonium pathway is structured by a binding/recruitment site facing the periplasmic side; a so-called phenylalanine gate composed by two conserved Phe residues; a central section characterized by the presence of two highly conserved His and a cytoplasmic vestibule (Khademi et al., 2004; Zheng et al., 2004). It is suggested that in the recruitment site, S219 and W148, and well conserved in the AMT/MEP/Rh family, can bind ammonium through a hydrogen bonding by the former and through the π system of the latter (Khademi et al., 2004), although in the crystal structure of the Amt1 homologue from A. fulgidus (AfAmt1;1) this site is occupied by water (Andrade et al., 2005). The Phe gate is structured by the F107 and F215 pair that seems to block the entrance to the putative channel. Deeper into the channel of EcAmtB, the central section is partially structured by His168 and His318 which are proposed to deprotonate ammonium and to bind the substrate ammonia, facilitating thus the transport of the latter. Finally, in the cytoplasmic vestibule, the side chain of V314 occupies the pore exit, and depending on the crystal structure, it may exist in a closed (P63 structure) or open (R3 structure) state, which may suggest V314 plays a role in controlling ammonium transport, that in the case of EcAmtB, may induce the binding of the regulatory GlnK protein inhibiting ammonium transport (Zheng et al., 2004).
Sequence Analysis of AMT/MEP/RH Transporters
The genome of higher plants contain two families of ammonium transporters, AMT1 and AMT2 (Sohlenkamp et al., 2000; Simon-Rosin et al., 2003; Couturier et al., 2007). Both family members show a high affinity for ammonium (in the micromolar range), but those from the AMT2 family are unable to transport the ammonium analog, methylammonium; possess a smaller transport capacity, and ammonium transport by AtAMT2 has been shown to be electroneutral, suggesting that the transported species corresponds to the uncharged NH3 and not NH4+ (Sohlenkamp et al., 2000, 2002; Simon-Rosin et al., 2003; Neuhäuser et al., 2009). From a pair wise analysis (Figure 1) of the sequences shown in Figure 2, it can be observed that among the plant AMT1 proteins there exists a high degree of identity between them, varying between 65 and 91%, with the exception of Lycopersicon esculentum LeAMT1;3 which on average, shows a slightly lower identity with the other members, ≤60%. In comparison, identity between the AMT1 and AMT2 families is lower and accounting for only 20–25%, as has previously been reported (Sohlenkamp et al., 2000). Identity of the plant AMT1 proteins with their homologues from fungi and bacteria is also low with values close to 20–25% (Figure 1). Identity between AMT2 homologues from plants and those from fungi and bacteria falls within similar low values (≤20%). These observations are better appreciated in the phylogenetic tree in Figure 3, which clearly shows that the AMT1 family members group in a closely related cluster, while those from the AMT2, fungi, archaea and bacteria branch off and form a separate cluster. In the latter, the low identity levels among these members (Figure 3), as represented by the length of the branches, clearly indicates that plant AMT2, archaea, bacteria, and fungi proteins form distantly related groups. In spite of these differences, and as mentioned before, the majority of these proteins show high affinity and high selectivity for ammonium, which indicates that among the 20–25% conserved residues must reside those directly involved in the mechanism of transport that may correspond to those proposed to structure the ammonium pathway (Khademi et al., 2004; Zheng et al., 2004; Andrade et al., 2005).
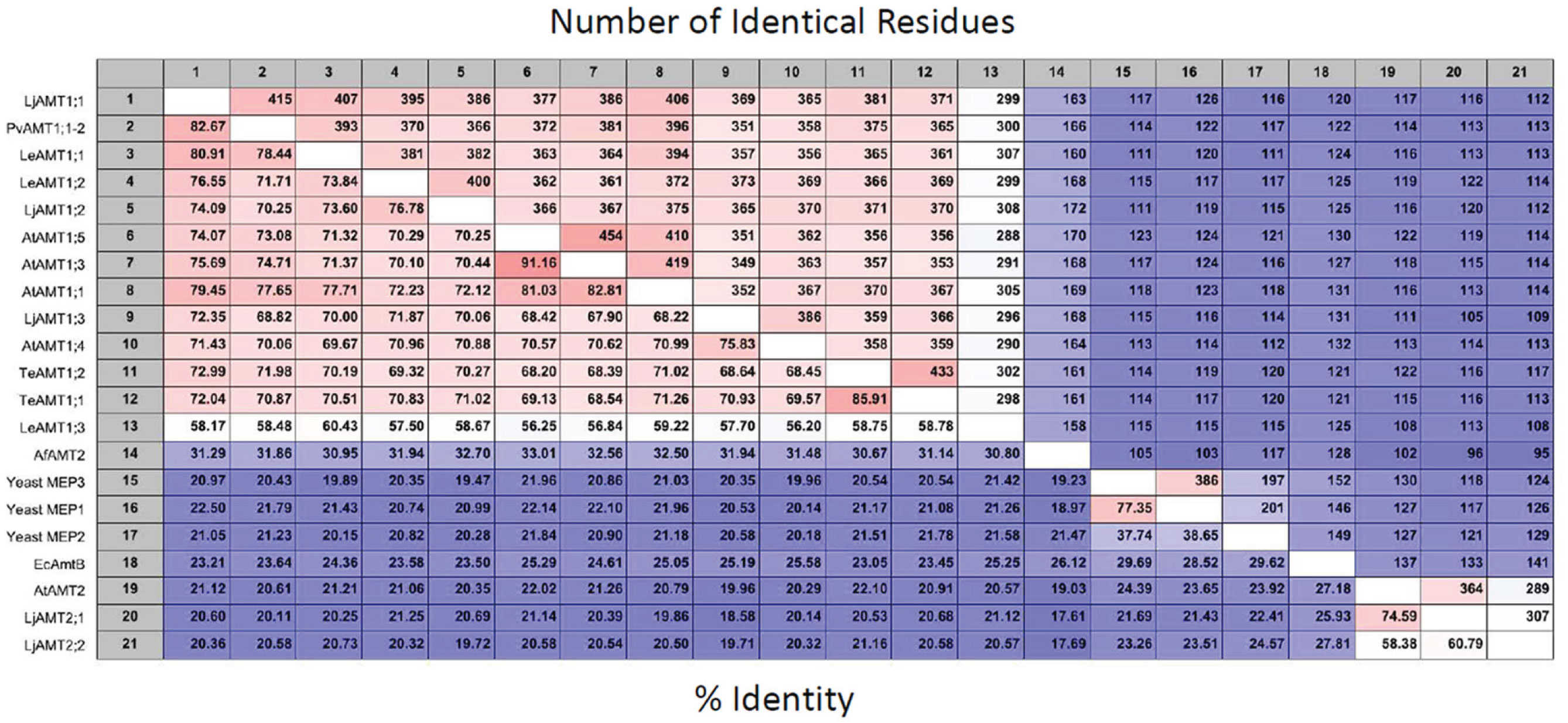
Figure 1. Pairwise analysis of AMT/MEP transporters. Employing the sequence from PvAMT1;1 and the CLC Main Workbench software (CLC bio, Katrinebjerg, Aarhus N, Denmark) 106 most conserved sequences related to AMTs were obtained from NCBI using BLAST. Twenty-one sequences that are mentioned in the text were aligned, and compared (Neighbor Joining; 100 bootstraps), to obtain the number of conserved residues, upper half and percent identity between the proteins, bottom half. The shades of red and blue color represent, respectively, the maximal and minimal number of identical residues, upper half; or per cent of identity, lower half, between the AMT/MEP transporters.
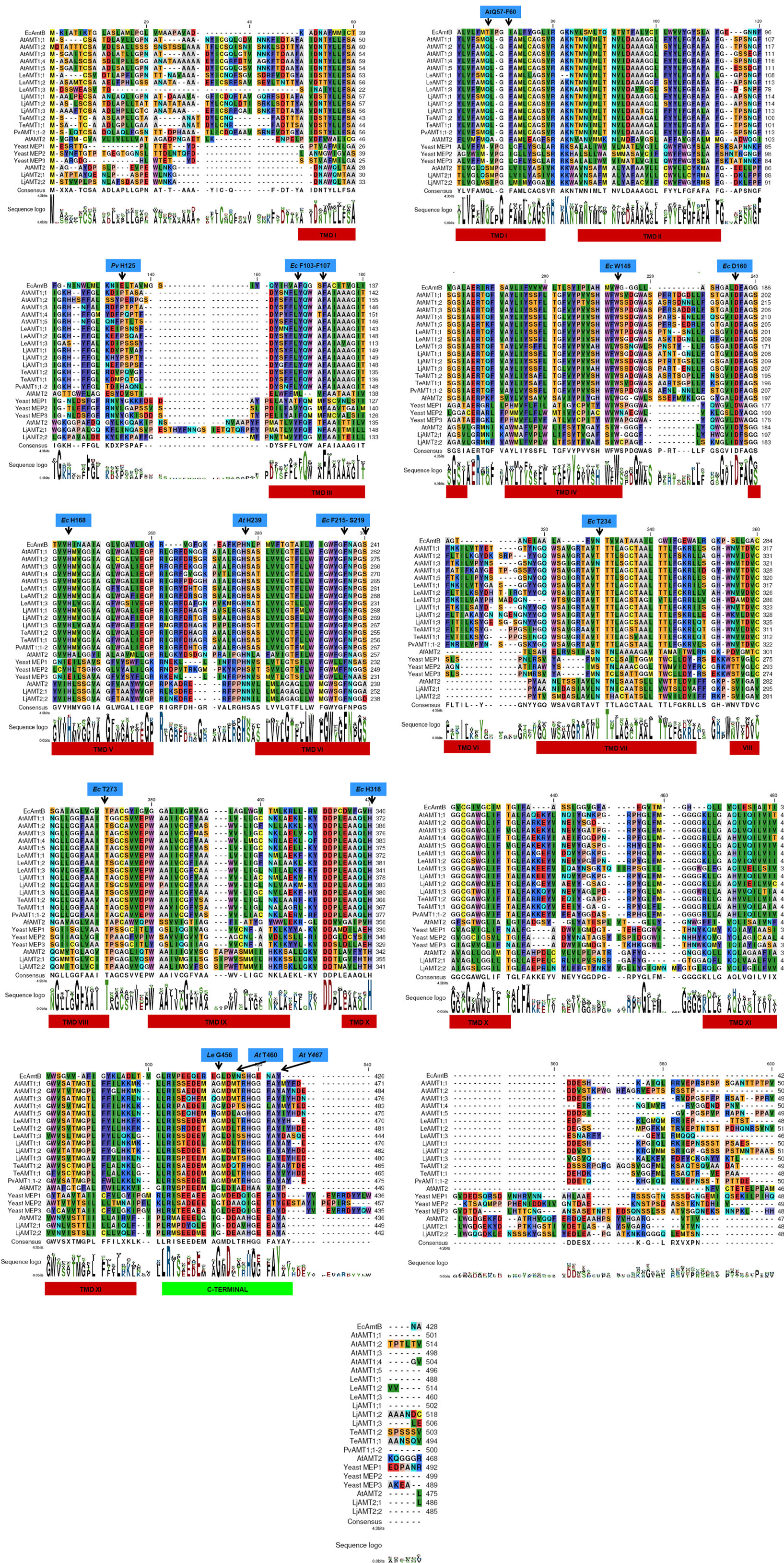
Figure 2. Alignment of 21 AMT/MEP transporters. Alignment of the 21 proteins reviewed in the text were aligned with the CLC Main Workbench software (CLC bio, Katrinebjerg, Aarhus N, Denmark). Amino acids identified as constitutive of the pore in E. coli AmtB are marked at the top of the sequences in blue squares; transmembrane domains are shown in red rectangle at the bottom of the sequences; the C-terminal is highlighted by a green rectangle.
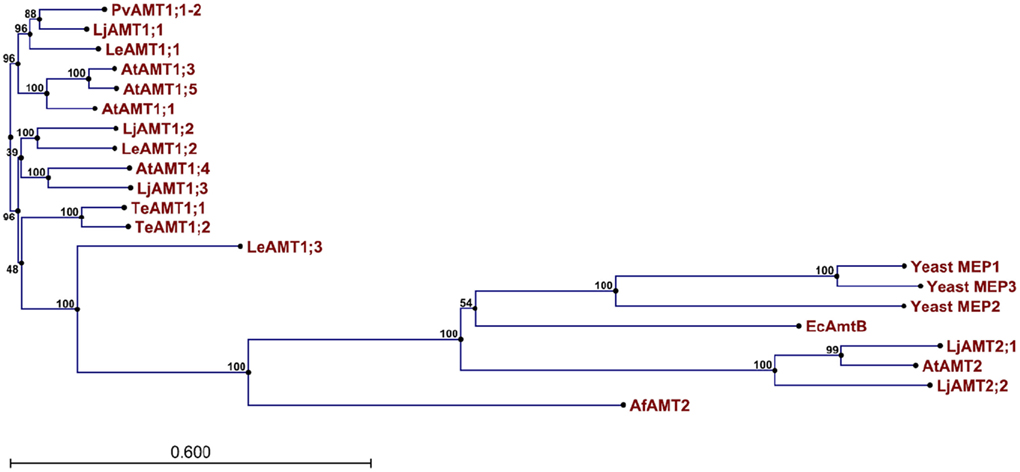
Figure 3. Phylogenetic tree of AMT/MEP transporters. The phylogenetic tree was calculated with the CLC Main Workbench software (CLC bio, Katrinebjerg, Aarhus N, Denmark) employing the 21 sequences described in the text.
The Ammonium Binding/Recruiting Site
As postulated by several groups, among the conserved amino acids are W148 and S219, in EcAmtB, proposed to structure the ammonium binding site (Khademi et al., 2004; Zheng et al., 2004; Andrade et al., 2005). According to the alignment shown in Figure 2, most of these residues are well conserved, with the exception of S219 which is replaced by Ala or Asp in plant AMT2 proteins. The more detailed analyses on the putative pore have been made on the E. coli homologue EcAmtB. Most results on the role of the residues localized in the ammonium recruiting/binding site have demonstrated that substitution of F107, W148, or S219 with either Ala or Leu, do not modify the expression level of EcAmtB, although some changes were observed in the transport properties (Fong et al., 2007; Javelle et al., 2008; Hall and Kustu, 2011). Mutations in F107A, W148A-L, or S219A increased transport activity between 2- and 10-fold while the rest of the mutants showed similar levels to those of the wild type (wt; Javelle et al., 2008; Hall and Kustu, 2011). Inhibition of MeA transport by thallium in all these mutants was similar to the wt, indicating that the microenvironment is not severely affected for MeA binding to occur. Inhibition by thallium however, was reduced (15–20%) in the triple mutant F107A/W148A/S219A. Crystal analyses of some of these variants showed that the mutations did not modify the general structure of the transporter, with the changes being limited to the mutated residue or with minor changes on neighboring amino acids (Javelle et al., 2008). These results indicate that the proposed role for the aromatic amino acids in NH4+/MeA binding through cation-π interactions or to S219 by H-bonding (Khademi et al., 2004) is dispensable, and that these substrates can apparently, move freely through the periplasmic/extracellular region. An additional residue, D160 has been shown to play an important role by mutational analysis, where substitution by Ala renders the protein inactive, and thus, proposed to be the ammonium binding site (Javelle et al., 2004). However, in the crystal structure of EcAmtB (Khademi et al., 2004), a density peak associated with the presence of high ammonium or MeA concentrations was not observed close to D160, that besides being inaccessible to the extracellular solution, suggest that the role of this residue may be structural (Khademi et al., 2004). Mutations in the equivalent residue in yeast (Saccharomyces cerevisiae) MEP transporters (ScMEP2 D186N; Marini et al., 2006) and Arabidopsis thaliana (AtAMT1;1 D198N; Loqué et al., 2007) also inhibited ammonium transport, thus confirming the importance of this Asp located in the periplasmic/extracellular side of the proteins.
In plants, attempts have been made to ascertain the importance of W178 and Y133 in LeAMT1;1 from tomato (W148 and F103 in EcAmtB). The mutation W178L transformed the transporter into an inactive mechanism, unable to generate currents with ammonium or to transport the analog MeA (Mayer et al., 2006). On the contrary, the change Y133I induced an increase in ammonium affinity (10-fold), together with a significant decrease in transport capacity (10-fold smaller currents; Mayer et al., 2006). The mutations W178L and Y133I were made to mimic the binding site found in the human Rh homologue glycoproteins, transporters that carry out low affinity and electroneutral ammonium transport (Mayer et al., 2006). The properties shown by the W178L and Y133I mutants indicate that substitutions of these two amino acids does not transform the tomato transporter into an Rh-like protein and that the transport properties of the latter must depend on residues located elsewhere.
Changes in the putative ammonium binding site occur naturally in the members of the AMT2 plant family, where the residue equivalent to S219 in EcAmtB and proposed to form an H-bond with ammonium, is substituted either, by Ala in AtAMT2;1 (A239) and (Lotus japonicus) LjAMT2;1 (A252) or Asp in LjAMT2;2 (D238; see Figure 2). LjAMT2;1 and AtAMT2;1 seem to transport uncharged ammonia, with a low transport capacity (low Vmax) and are impermeable to MeA (Simon-Rosin et al., 2003; Neuhäuser et al., 2009). Interestingly, LjAMT2;2 a mycorrhizal specific ammonium transporter is also electroneutral, but it does transport MeA with a relatively high affinity (Km = 0.83 mM). The natural substitution of S219 by Ala in AtAMT2;1 and LjAMT2;1 together with the results from the EcAmtBS219A mutant challenge the proposed role of S219 in ammonium binding through an H-bond and suggests that other amino acids must be involved in ammonium binding.
The Phenylalanine Gate
The Phe gate, structured in EcAmtB by the F107 and F215 pair, it seems to block the entrance to the putative channel (Khademi et al., 2004; Andrade et al., 2005). Both residues are well conserved in all the analyzed proteins (Figure 2). Interestingly, while the single mutant F215A, was well expressed in the membrane, did not carry out MeA uptake (Javelle et al., 2008) nor grew on low ammonium concentrations (Hall and Kustu, 2011), the F107A mutant behaved very much like the wt transporter (Javelle et al., 2008; Hall and Kustu, 2011). Further mutations in F215 (H, Q, S, or W) did not transport MeA (Javelle et al., 2008), just like the mutant F215L (Javelle et al., 2008; Hall and Kustu, 2011), which however, grew slightly slower than the wt in micromolar ammonium (Hall and Kustu, 2011). It is important to point out the contrasting results with the F215L mutant which led to the conclusion by one group that F215 plays a crucial role in the functioning of the transporter based on its inability to transport MeA (Javelle et al., 2008), while another group proposed that F215 and the associated π-interactions that they may establish with ammonium are not required for the functioning of EcAmtB, as indicated by the similar growth rates shown by the F215L mutant and the wt (Hall and Kustu, 2011). Interestingly, the double (W148L/F215L) and triple (F107A/W148L/F215L) mutants did not differ much from the wt when grown in low ammonium, although both did not achieve MeA transport (Hall and Kustu, 2011).
Intuitively, mutation on the Phe gate to F107A/F215A has potential structural effects, generating an open pore, which would suggest that ammonium and other ions with similar radius could freely permeate through the mutant transporter. However, neither MeA, K+, H+ nor water were transported by the F107A/F215A double mutant or by EcAmtB. It remains to be demonstrated if this mutant is capable to grow at low ammonium concentrations before reaching a definitive conclusion on the effects of this double substitution (Javelle et al., 2008).
The Role of His168 and His318 in Ammonium Deprotonation and pH-Dependence
The pair of His postulated to deprotonate NH4+ and facilitate the movement of uncharged NH3 (H168 and H318) is also well conserved, excluding the yeast ScMEP1 and ScMEP3 proteins where the residue equivalent to His168 corresponds to E181 and E180, respectively (Figure 3). Analysis of 14 mutants in EcAmtB where the His residues were replaced by aliphatic, aromatic, or polar residues, demonstrated that most of them were expressed in the membrane and maintained their trimeric structure. Six of the polar mutants were also expressed in the membrane but the apparent formation of monomers and dimers indicated that they were rather unstable (Javelle et al., 2006). From those mutants that were crystallized, it was observed that the structural changes were restricted to the mutated His, with small effects on adjacent residues (Javelle et al., 2006). From all the analyzed mutants, only H168E and H168D showed reduced MeA uptake that accounted for 15–25% of the wt activity (Javelle et al., 2006; Hall and Kustu, 2011), although growth in low ammonium was normal for the H168E mutant (Hall and Kustu, 2011), Functionality of the H168E mutant should not be surprising since this substitution is naturally present in the ScMEP1 and ScMEP3 homologues from yeast (Figure 2), both being able to transport ammonium and MeA (Marini et al., 2000). Recently, it was reported that the same substitution (H211E) in PvAMT1;1 from bean (Phaseolus vulgaris), caused an increase in ammonium current and a large decrease in MeA transport (Ortiz-Ramirez et al., 2011). PvAMT1;1, like most of the AMT1 family members shows high affinity (Km = 15 μM and Vmax = 140 nA), and high selectivity for ammonium, and saturates at concentrations below 1 mM; however, it is pH-dependent, being more active in acidic extracellular environments. Associated to the higher activity at acidic external pH, ammonium transport (currents) in Xenopus oocytes was accompanied by cytoplasm acidification. Changes in both, extracellular ammonium and proton concentration induced changes in the reversal potential of the ammonium currents with slopes of 33 and 26 mV/decade, respectively, values that were used to calculate an stoichiometry of 1 NH4+/1 H+, indicating that PvAMT1;1 operates as an NH4+/H+ symporter (Ortiz-Ramirez et al., 2011). The mutation H211E transformed the protein into a pH-independent mechanism, with a decreased affinity and increased transport capacity for ammonium (Km = 1.3 mM and Vmax = 300 nA), accounting for 40- and 2-fold increases in these two parameters, respectively. Additionally, the reversal potential of the ammonium currents changed according to the Nernst equation, indicating that ammonium was the only ion transporter by the mutant PvAMT1;1H211E, and confirmed by the absence of cytoplasm acidification (Ortiz-Ramirez et al., 2011). These results indicate that H211 positioned in the middle of the channel can be substituted by Glu, as it is naturally for other transporters. The ScMEP1 and ScMEP3 homologues from yeast present a similar arrangement to the H211E mutant, in which the first His is substituted by Glu, and show similar transport properties. ScMep1 and ScMep3 are pH-independent, have a low MeA affinity and apparently, larger transport capacity as evidenced by the toxic effects of MeA (Boeckstaens et al., 2008). In contrast, ScMep2 maintains the first conserved His, like the wt bean transporter, and also shares the high affinity for ammonium and pH-dependence (Boeckstaens et al., 2008). These properties become similar to those of its paralogs or the H211E bean mutant if the first His is mutated to Glu (Boeckstaens et al., 2008). Analogous manipulation in the EcAmtB protein lead to similar results to those observed for ScMep2 (Boeckstaens et al., 2008). All these observations confirm that the first conserved histidine can be replaced by Glu without inhibiting ammonium transport, and suggest that this residue in association with as yet, unidentified amino acids, are responsible for the pH sensitivity of some members of this family.
Analysis of the four double mutants (H168E/H318E, H168D/H318D, H168D/H318E, and H168E/H318D) demonstrated that all were severely affected, unable to grow in low ammonium or to transport MeA, as a result of low levels of expression. Employing these mutants to isolate suppressor mutants led to the identification of I110N as an important modification (Hall and Kustu, 2011). The mutants H168D/H318D, H168D/H318E were both rescued by I110N, showing a slightly longer duplication rates in low ammonium medium than the wt, but significantly better that the parental lines. It was proposed that Asn, being present in close proximity to the two acidic amino acids, at the other side of the pore, may help to decrease the strong electrostatic effect of Asp and Glu by means of H-bond interactions (Hall and Kustu, 2011). It is important to point out that neither of the rescued mutants was able to transport MeA, demonstrating, that the two assays for the evaluation of AMT activity, growth in low ammonium and MeA transport, do not give similar information. The studies on the pair of His show that when one of them is mutated to an acidic amino acid, the transporter is functional, but not when both are, implying that presence of two carboxy groups in the center, destabilize the protein, as suggested by the low levels of expression of the H168D/H318D, H168D/H318E mutants. The fact that ammonium and MeA can be transported by the suppressor I110N in the presence of hydrophilic residues in the pore suggests that presence of Asn may help to the stabilization of the protein and that charged  seems to be the transported species, implying that an hydrophobic pore is not a requisite for ammonium transport to occur.
seems to be the transported species, implying that an hydrophobic pore is not a requisite for ammonium transport to occur.
The Importance of the C-Terminal in the Regulation of Ammonium Transporters
Although the similarity of plant ammonium transporters with those from E. coli or A. fulgidus is somewhat low (20–25%), it is remarkable that molecular modeling has shown that plant proteins seem to maintain similar tertiary and quaternary structures, particularly the trans membrane domains (TMD), the cytoplasmic loops and the carboxy terminal (C-terminal), with the extracellular loops showing some divergence, mainly as a result of some of them being longer (Mayer et al., 2006; Loqué et al., 2007; Neuhäuser et al., 2007, 2009; Guether et al., 2009; Ortiz-Ramirez et al., 2011).
Confirmation that plant ammonium homologues can be modeled very closely to the crystal structure of bacteria and archaea, has open up the possibility to perform directed mutational analysis for the identification of additional amino acids that may be involved in the functioning of these transporters, as well as to verify the importance of the conserved residues present in the putative ammonium pathway.
One of the first studies where the knowledge derived from the quaternary structure obtained for the ammonium transporter from A. fulgidus was applied, is that reported by Loqué et al. (2007) where modeling of A. thaliana AtAMT1;1 demonstrated that the C-terminal established interactions with cytoplasmic loops of the same monomer, as well as with loops from neighbor monomers in the homotrimer. Based on the observed interactions of the C-terminal with cytoplasmic loops between TMDI–II, TMDIII–IV, and TMDV–VI of its own monomer, as well as with cytoplasmic loops connecting TMDI–II, TMDV–VI, and TMDVII–VIII of a neighboring monomer reported for A. fulgidus (Andrade et al., 2005), Loqué et al. (2007) analyzed the predicted interaction between Y467 in AtAMT1;1 (Y448 in AfAMT1) located at the C-terminal and H239 (H234 in AfAMT1) positioned in the loop between TMDV–VI by mutating both residues. These authors observed that both mutants, Y467F and H239C, independently, were inactive and trans-inhibitory. More importantly, through a targeted suppressor screen, the interaction between these two residues was confirmed, since Y467F was only rescued by H239C, and vice versa. Thus, demonstrating that the interaction between the two regions of the protein (C-terminal and the cytoplasmic loop linking TMDV–VI), either through H or hydrophobic bonding is important in the activation of the transporter. Conservation of these two amino acids in most of the AMT proteins from plants (Figure 2), indicates that this regulatory mechanism is well conserved, and presence of the same pair of residues in E. coli and A. fulgidus suggests that it may be more widely common. Similar analyses on Thr460 from AtAMT1;1, that seems to establish H-bonding within the C-terminal, showed that the partially inactive mutant T460A was rescued by mutations in residues located at cytoplasmic loops between TMDIII–IV (I146M and Q151E) of the same monomer and TMDV–VI (S242) and TMDVII–VIII (V313L) from a neighbor monomer, confirming the interactions observed in the crystal structure of A. fulgidus, and reinforcing the role of these associations in the regulation of the transporter (Loqué et al., 2007).
The importance of T460 in AtAMT1;1 has been shown to be a result of its post-translational modification by phosphorylation (Lanquar et al., 2009). Initial studies had demonstrated that proteins carrying the mutation T460D were inactive, that when co-expressed with wt functioning transporters AtAMT1;1 (Loqué et al., 2007) or AtAMT1;2 (Neuhäuser et al., 2007), caused the trans-inhibition of the latter. Phosphorylation of T460 has been demonstrated to be dependent on ammonium concentration and time of exposure, as a mechanism proposed to prevent the continuous accumulation of this cation, that may become toxic if accumulated in the cytoplasm (Lanquar et al., 2009). Importantly, phosphorylation of T460 was caused rapidly and at low ammonium concentration (50 μM), and only induced by extracellular ammonium but not by any other cation, suggesting the existence of an ammonium receptor that directly or indirectly modifies the activity of the transporter. Alternatively, the same transporter could be functioning as a transceptor that relies the signal to an associated membrane kinase (Lanquar et al., 2009). Conservation of this regulatory mechanism may be common among the plant AMT1 homologues, where T460 is well conserved, with the exception of AtAMT1;5 (A463). An Ala is also present in the equivalent position of plant AMT2 family members which indicates that in these homologues the regulatory role associated with T460 is not present. It remains to be shown if other mechanisms or residues in the C-terminal of the AMT2 transporters or AtAMT1;5 participate in a similar regulatory mechanism.
An additional residue located at the C-terminal of plant transporters that plays an important role in maintaining the functionality of these proteins is G456 in LeAMT1;1 (Ludewig et al., 2003), first identified in the yeast ScMEP1 homologue (G413; Marini et al., 2000). Mutation of G456 to Asp transforms LeAMT1;1 into an inactive transporter that also trans-inactivates its own wt or its paralog LeAMT1;2, evidence that first suggested that the AMT transporters could oligomerize. In contrast to T460, G456 is highly conserved among all the members of the MEP/AMT family (Figure 2), highlighting the importance of this residue in establishing trans-interactions between monomers that results in the activation of the transporters.
In view of all these results, it is important to highlight the role of the C-terminal in regulating the activity of plant AMT1 ammonium transporters, that as has been described above, it seems to be through direct interactions within the same or an adjacent monomer (Loqué et al., 2007; Neuhäuser et al., 2007). In bacteria, in contrast, the role of the C-terminal is dependent on its association with the non-uridylylated form of the cytoplasmic protein GlnK, interaction that results in the inhibition of EcAmtB and ammonium transport, a mechanism that serves to regulate the uptake of this potentially toxic nutrient (Coutts et al., 2002). Interestingly, deletion of the C-terminal reduces the activity of EcAmtB while completely inhibiting the functioning of AtAMT1;1 (Marini et al., 2000; Loqué et al., 2007).
Identification of Additional Amino Acids Important in the Functioning of Plant AMT’s
Among the plant AMT1 family members, an additional common property is the pH-independence of their activity (Ludewig et al., 2002, 2003; Wood et al., 2006), with the exception, so far, of PvAMT1;1 and Triticum aestivum TaAMT1;1 (Søgaard et al., 2009; Ortiz-Ramirez et al., 2011). In the wheat homologue, transport was stimulated by acidic external pH, with a half-maximal response (EC50) of 50 μM ammonium that was constant at all levels of pH. Moreover, TeAMT1;1 does not seem to transport H+, as indicated by the unchanged reversal potential in response to external pH changes (Søgaard et al., 2009). A mutation in the cytoplasmic side of the pore (L56F), caused the transporter to become pH-independent, to lose affinity for ammonium, and also associated with an increase in transport activity (Søgaard et al., 2009). From these results, Søgaard et al. (2009) postulated that the increased ammonium transport was a result of a larger diameter of the pore in the cytoplasmic side together with a facilitated H+ transport. However, if this were the case, it would be expected that changes in both ammonium and H+ concentrations would cause changes in the reversal potential of the currents, as has been reported for PvAMT1;1 (Ortiz-Ramirez et al., 2011). Unfortunately, this information was not presented in the report by Søgaard et al. (2009). What is intriguing is that a point mutation on the cytoplasmic side of the protein senses pH changes on the extracellular side, when it is expected that those residues directly involved in binding and/or sensing of regulatory factors, should be located near to the exposure side.
Studies on the role of pore structuring residues is still limited in plant ammonium transporters, however, participation of other amino acids in the mechanism of action of these proteins has been obtained by isolating suppressors of non-functional mutants. Recovering of the mutant T460A and the truncated AtAMT1;1 (G456stop) protein was achieved individually by the mutations F60S, I136F, A137D, postulated to transform the transporter into a deregulated constantly active mechanism (Loqué et al., 2007). Two further suppressor mutants, Q57H and A138P, recovered the non-functional T460A and T460D proteins, showing a twofold larger ammonium uptake, and as a consequence, sensitivity of the yeast transformants to relative low concentrations of ammonium (Loqué et al., 2007, 2009). Interestingly, the Q57H and A138P mutants also conferred resistance to the toxic effects of MeA. Confirmation of these properties was obtained for AtAMT1;1Q57H in Xenopus oocytes, were large and non-saturable ammonium currents were recorded, while recording an important decrease in the magnitude of the currents (10-fold lower) generated with MeA (Loqué et al., 2009). The kinetic properties of Q57H were substantially different to those of AtAMT1;1, showing a 65-fold larger Km for ammonium and a 10-fold larger Vmax (Loqué et al., 2009). All these modified residues are located in the middle of TMDI (Q57 and F60) or TMDIII (I136, A137, and A138), forming part of the putative pore, an observation that was used to suggest that modifications to the structure of the pore not only release the regulation of the proteins by the C-terminal, but also change the mechanism of action, transforming the transporter into an ion channel (Loqué et al., 2007). A proposal that was partially confirmed by the electrical properties of the Q57H mutant, although failure in recording single channel activity from this protein (Loqué et al., 2009), leaves still open the confirmation of such proposal.
Identification of a unique His in PvAMT1;1, which is replaced by Pro in all other plant homologues (Figure 2), prompted the authors to ascertain if this residue was responsible for the pH-dependence showed by the bean transporter. The mutation H125A transformed the transporter into a less active mechanism, but still pH-dependent, with higher activity under an acidic environment. In contrast, the mutant H125R, generated a more active protein with a Vmax twice as large as that observed for the wt, but maintaining its stimulation by low pH (Ortiz-Ramirez et al., 2011). Interestingly, H125 is located in the extracellular loop joining TMDII and TMDIII indicating that substantial changes in the properties of the transporter can be caused by modification of residues not directly lining or structuring the putative conductive pore.
Continuing with the analysis of other conserved amino acids, it is important to recall an observation made by Zheng et al. (2004) who have pointed out that T234 and T273 are also well conserved as shown in Figure 2, and suggested that these residues may participate in structuring the side chains of the conserved residues surrounding the periplasmic side of the pore (ammonium binding site). Studying the crystal structure 1U7C and 1XQE of EcAmtB (Khademi et al., 2004; Zheng et al., 2004), it was observed that T273 may form an H-bond with A162 located in the periplasmic side as well as with H168 (Figure 4), which indicates that T273 may have an important role in the functioning of the transporter. Unfortunately, so far, there is no evidence on the participation of T273 or A162 in the functioning of any of the AMT/MEP/Rh proteins which leaves open the analysis of the importance of these two residues. According to the pseudo twofold symmetry of the ammonium transporters (Khademi et al., 2004; Zheng et al., 2004; Andrade et al., 2005), the amino acid equivalent to D160, D310, is also very well conserved (Figure 2), and preceded by another well conserved D309, both located in the cytoplasmic side of the protein where it has been postulated to participate in structuring the backbone of trans membrane domain X (TMDX) through H-bonding (Zheng et al., 2004). So far, evidence on the importance of D310 (or D309) in EcAmtB or its equivalent in any of the homologues is still lacking.
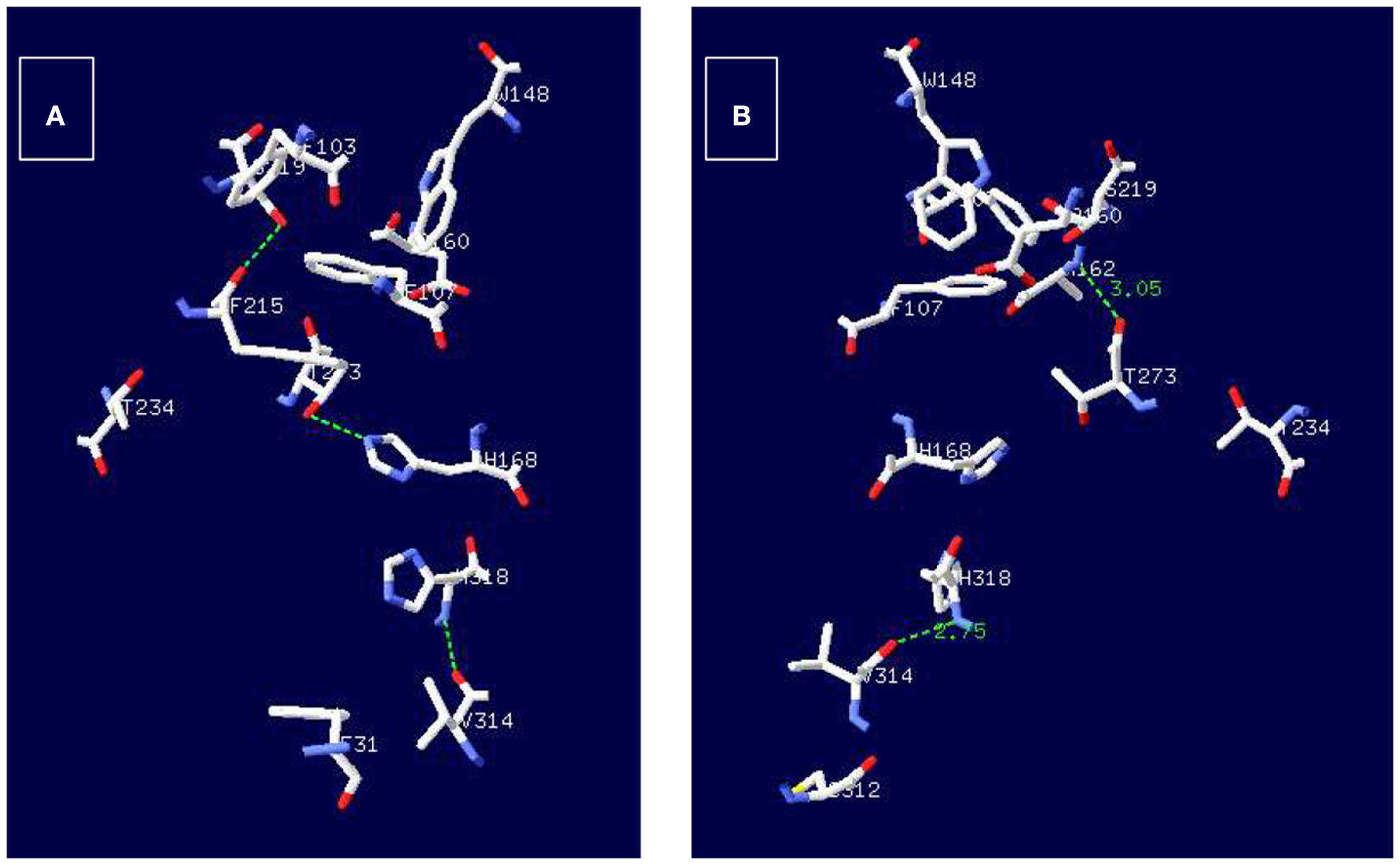
Figure 4. Potential H-bond interactions of T273 with amino acids structuring the conductive ammonium pore in EcAmtB. Employing the structures 1XQE (A) and 1U7C (B) of EcAmtB and the program DeepView/Swiss-PdbViewer (http://www.expasy.org.spdbv) H-bonds were calculated and are shown as broken green lines between T273 and H128 (A) or T273 and A162.
Observations on the Use of MeA as an Analog of Ammonium
Based on the results that have recently been reported on the modified properties of mutant transporters, where differences have been observed in their capacity to grow on low ammonium but unable to take up MeA, it is important to draw attention to the latter response. Radio-labeled MeA has been widely used for the evaluation of ammonium transport activity in a variety of organisms due to the availability of 14C-labeled MeA and the short life of radioactive 13N. The use on this ammonium analog is also attractive because it allows the parallel measurements of 14C-MeA uptake and the currents associated to this transport, when electrogenic, allowing the estimation of the stoichiometry of the transporter, as it has been used by Ludewig et al. to demonstrated that NH4+ is the transported substrate by plant AMT1’s (Mayer et al., 2006). However, the results reported for mutants like EcAmtBH138E or H138D (Hall and Kustu, 2011), AtAMT1;1Q57H (Loqué et al., 2009) and PvAMT1;1H211E (Ortiz-Ramirez et al., 2011) that show an increased selectivity toward ammonium in comparison to MeA, underscore potential misinterpretations if the results are obtained exclusively from MeA uptake experiments. Further work on the analysis of AMT/MEP transporters or mutants should include evidence obtained from different methodologies to clearly demonstrate that the proteins do or do not carry out ammonium transport.
Conclusion
Our knowledge on the functioning of AMT/MEP/Rh proteins has advanced considerably in the last few years, and resolution of the crystal structure from the bacteria and archaea homologues, has also been of great help in guiding the research focused on identifying crucial amino acids in the functioning of these transporters. It seems that the residues initially postulated as directly involved in the mechanism of ammonium transport may not play a central role, as it has been reviewed here. Modifications to the putative binding site by residues with properties substantially different to the conserved aromatic amino acids that structure this extracellular domain, result in changes that do not completely abolished the activity of the transporter or that generated a more active mechanism (Loqué et al., 2009; Hall and Kustu, 2011). Likewise, it now appears that the deprotonation role assigned to the twin His may not be necessary, and that a combination of more hydrophilic residues in the pore is also functional, as demonstrated by the triple mutants H168D/H318D/I110N and H168D/H318E/I110N (Hall and Kustu, 2011) which also suggest that the charged NH4+ is the transported molecule. However, it is also possible that the properties of an individual ammonium transporter are not due to the presence or absence of a particular amino acid, but most likely, are a result of a concerted group of residues that establish singular conditions proper to each member of this family.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The support from CONACyT-México (grants 42664 and 79191) and Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México (Grant IN218308) is acknowledged.
References
Andrade, S. L. A., Dickmanns, A., Ficner, R., and Einsle, O. (2005). Crystal structure of the archaeal ammonium transporter Amt-1 from Archaeoglobus fulgidus. Proc. Natl. Acad. Sci. U.S.A. 102, 14994–14999.
Boeckstaens, M., Bruno, A., and Marini, A. M. (2008). Distinct transport mechanisms in yeast ammonium transport/sensor proteins of the Mep/Amt/Rh family and impact on filamentation. J. Biol. Chem. 283, 21362–21370.
Conroy, M. J., Durand, A., Lupo, D., Li, X.-D., Bullough, P. A., Winkler, F. K., and Merrick, M. (2007). The crystal structure of the Escherichia coli AmtB–GlnK complex reveals how GlnK regulates the ammonia channel. Proc. Natl. Acad. Sci. U.S.A. 104, 1213.
Coutts, G., Thomas, G., Blakey, D., and Merrick, M. (2002). Membrane sequestration of the signal transduction protein GlnK by the ammonium transporter AmtB. EMBO J. 21, 536–545.
Couturier, J., Montanini, B., Martin, F., Brun, A., Blaudez, D., and Chalot, M. (2007). The expanded family of ammonium transporters in the perennial poplar plant. New Phytol. 174, 137–150.
D’Apuzzo, E., Rogato, A., Simon-Rosin, U., El Alaoui, H., Barbulova, A., Betti, M., Dimou, M., Katinakis, P., Marquez, A., Marini, A. M., Udvardi, M. K., and Chiurazzi, M. (2004). Characterization of three functional high-affinity ammonium transporters in Lotus japonicus with differential transcriptional regulation and spatial expression. Plant Physiol. 134, 1763–1774.
Fong, R. N., Kim, K.-S., Yoshihara, C., Inwood, W. B., and Kustu, S. (2007). The W148L substitution in the Escherichia coli ammonium channel AmtB increases flux and indicates that the substrate is an ion. Proc. Natl. Acad. Sci. U.S.A. 104, 18706–18711.
Gazzarrini, S., Lejay, L., Gojon, A., Ninnemann, O., Frommer, W. B., and von Wirén, N. (1999). Three functional transporters for constitutive, diurnally regulated, and starvation-induced uptake of ammonium into Arabidopsis roots. Plant Cell 11, 937–948.
Guether, M., Neuhäuser, B., Balestrini, R., Dynowski, M., Ludewig, U., and Bonfante, P. (2009). A mycorrhizal-specific ammonium transporter from Lotus japonicus acquires nitrogen released by arbuscular mycorrhizal fungi. Plant Physiol. 150, 73–83.
Hall, J. A., and Kustu, S. (2011). The pivotal twin histidines and aromatic triad of the Escherichia coli ammonium channel AmtB can be replaced. Proc. Natl. Acad. Sci. U.S.A. 108, 13270–13274.
Javelle, A., Lupo, D., Ripoche, P., Fulford, T., Merrick, M., and Winkler, F. K. (2008). Substrate binding, deprotonation, and selectivity at the periplasmic entrance of the Escherichia coli ammonia channel AmtB. Proc. Natl. Acad. Sci. U.S.A. 105, 5040–5045.
Javelle, A., Severi, E., Thornton, J., and Merrick, M. (2004). Ammonium sensing in Escherichia coli. Role of the ammonium transporter AmtB and AmtB-GlnK complex formation. J. Biol. Chem. 279, 8530–8538.
Javelle, A., Lupo, D., Zheng, L., Li, X. D., Winkler, F. K., and Merrick, M. (2006). An unusual twin-his arrangement in the pore of ammonia channels is essential for substrate conductance. J. Biol. Chem. 281, 39492–39498.
Khademi, S., O’Connell, J. III, Remis, J., Robles-Colmenares, Y., Miercke, L. J. W., and Stroud, R. M. (2004). Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 Å. Science 305, 1587–1594.
Lanquar, V., Loqué, D., Hörmann, F., Yuan, L., Bohner, A., Engelsberger, W. R., Lalonde, S., Schulze, W. X., von Wirén, N., and Frommer, W. B. (2009). Feedback inhibition of ammonium uptake by a phospho-dependent allosteric mechanism in Arabidopsis. Plant Cell 21, 3610–3622.
Loqué, D., Lalonde, S., Looger, L. L., von Wirén, N., and Frommer, W. B. (2007). A cytosolic trans-activation domain essential for ammonium uptake. Nature 446, 195–198.
Loqué, D., Mora, S. I., Andrade, S. L. A., Pantoja, O., and Frommer, W. B. (2009). Pore mutations in ammonium transporter AMT1 with increased electrogenic ammonium transport activity. J. Biol. Chem. 284, 24988–24995.
Ludewig, U., von Wirén, N., and Frommer, W. B. (2002). Uniport of NH4+ by the root hair plasma membrane ammonium transporter LeAMT1; 1. J. Biol. Chem. 277, 13548–13555.
Ludewig, U., Wilken, S., Wu, B., Jost, W., Obrdlik, P., El Bakkoury, M., Marini, A. M., André, B., Hamacher, T., Boles, E., Von Wirén, N., and Frommer, W. B. (2003). Homo-and hetero-oligomerization of ammonium transporter-1 uniporters. J. Biol. Chem. 278, 45603–45610.
Marini, A. M., Boeckstaens, M., Benjelloun, F., Chérif-Zahar, B., and André, B. (2006). Structural involvement in substrate recognition of an essential aspartate residue conserved in Mep/Amt and Rh-type ammonium transporters. Curr. Gen. 49, 364–374.
Marini, A. M., Springael, J. Y., Frommer, W. B., and André, B. (2000). Cross-talk between ammonium transporters in yeast and interference by the soybean SAT1 protein. Mol. Microbiol. 35, 378–385.
Mayer, M., Dynowski, M., and Ludewig, U. (2006). Ammonium ion transport by the AMT/Rh homologue LeAMT1; 1. Biochemical J. 396, 431–437.
Neuhäuser, B., Dynowski, M., and Ludewig, U. (2009). Channel-like NH3 flux by ammonium transporter AtAMT2. FEBS Lett. 583, 2833–2838.
Neuhäuser, B., Dynowski, M., Mayer, M., and Ludewig, U. (2007). Regulation of NH4+ transport by essential cross talk between AMT monomers through the carboxyl tails. Plant Physiol. 143, 1651–1659.
Ortiz-Ramirez, C., Mora, S. I., Trejo, J., and Pantoja, O. (2011). PvAMT1;1, a highly selective ammonium transporter that functions as H+/NH4+ symporter. J. Biol. Chem. 286, 31113–31122.
Shelden, M. C., Dong, B., de Bruxelles, G. L., Trevaskis, B., Whelan, J., Ryan, P. R., Howitt, S. M., and Udvardi, M. K. (2001). Arabidopsis ammonium transporters, AtAMT1; 1 and AtAMT1; 2, have different biochemical properties and functional roles. Plant Soil 231, 151–160.
Simon-Rosin, U., Wood, C., and Udvardi, M. K. (2003). Molecular and cellular characterisation of LjAMT2;1, an ammonium transporter from the model legume. Lotus japonicus. Plant Mol. Biol. 51, 99–108.
Søgaard, R., Alsterfjord, M., Macaulay, N., and Zeuthen, T. (2009). Ammonium ion transport by the AMT/Rh homolog TaAMT1;1 is stimulated by acidic pH. Pflügers Arch. Eur. J. Physiol.458, 733–743.
Sohlenkamp, C., Shelden, M., Howitt, S., and Udvardi, M. (2000). Characterization of Arabidopsis AtAMT2, a novel ammonium transporter in plants. FEBS Lett. 467, 273–278.
Sohlenkamp, C., Wood, C. C., Roeb, G. W., and Udvardi, M. K. (2002). Characterization of Arabidopsis AtAMT2, a high-affinity ammonium transporter of the plasma membrane. Plant Physiol. 130, 1788–1796.
Winkler, F. K. (2006). Amt/MEP/Rh proteins conduct ammonia. Pflügers Arch. Eur. J. Physiol. 451, 701–707.
Wirén, N. V., and Merrick, M. (2004). Molecular mechanisms controlling transmembrane transport. Topics in Current Genetics 1–26.
Wood, C. C., Porée, F., Dreyer, I., Koehler, G. J., and Udvardi, M. K. (2006). Mechanisms of ammonium transport, accumulation, and retention in ooyctes and yeast cells expressing Arabidopsis AtAMT1; 1. FEBS Lett. 580, 3931–3936.
Keywords: ammonium, transport, AMT, high affinity
Citation: Pantoja O (2012) High affinity ammonium transporters: molecular mechanism of action. Front. Plant Sci. 3:34. doi: 10.3389/fpls.2012.00034
Received: 11 October 2011; Accepted: 06 February 2012;
Published online: 28 March 2012.
Edited by:
Angus S. Murphy, Purdue University, USAReviewed by:
Yi-Fang Tsay, Academia Sinica, TaiwanLi-Qing Chen, Carnegie Institution for Science, USA
Copyright: © 2012 Pantoja. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Omar Pantoja, Instituto de Biotecnología, Universidad Nacional Autónoma de México, Apdo. Postal 510-3, Cuernavaca, Morelos 62250, México. e-mail:b21hckBpYnQudW5hbS5teA==
