- Energy Biosciences Institute, University of California, Berkeley, CA, USA
Mass spectrometry is a selective and powerful technique to obtain identification and structural information on compounds present in complex mixtures. Since it requires only small sample amount it is an excellent tool for researchers interested in detecting changes in composition of complex carbohydrates of plants. This mini-review gives an overview of common mass spectrometry techniques applied to the analysis of plant cell wall carbohydrates. It presents examples in which mass spectrometry has been used to elucidate the structure of oligosaccharides derived from hemicelluloses and pectins and illustrates how information on sequence, linkages, branching, and modifications are obtained from characteristic fragmentation patterns.
Introduction
Plant cell wall carbohydrates are the most abundant biopolymers on Earth. In addition to specialized functions in plant development and physiology, the use of specific cell wall polysaccharides and proteoglyans in food chemistry and nutrition, pharmaceuticals, biomaterials, biofuels, and green chemistry is increasing. Determination of, and detection of changes in, polymer composition is of interest to an ever-widening array of researchers studying their synthesis or degradation. Even with only a limited number of monosaccharides used as building blocks, variations in sequence, linkage, branching, distribution of side chains, and functional group modifications result in a wide structural diversity. No single analytical method alone is capable of resolving carbohydrate structures, especially when present in complex mixtures. Nuclear magnetic resonance (NMR) spectroscopy is widely used in structural elucidation but requires rather pure samples and lacks the ability to reveal sequence information (Duus et al., 2000). Although some methods (e.g., nanoprobe-NMR) have significantly lowered concentration requirements for analysis, mass spectrometry (MS) provides high sensitivity and can even be applied to mixtures of carbohydrates. This allows detailed characterization of changes in cell wall polymers due to changes in environmental, developmental, and/or genetic factors. Because the technique is mass-based, isotope labeling can be used in a variety of creative ways to elucidate time-resolved changes in polymer synthesis and modification.
Mass Spectrometry for Polysaccharide Analysis
To be amenable for MS polysaccharides must first be converted into smaller oligosaccharides either by partial acid hydrolysis or enzymatic degradation. The latter has the advantage of being specific to one polymer alone and can therefore be applied to a mixture of polysaccharides or even the whole cell wall eliminating the need for purification which often removes modifications (e.g., acetylation). Enzymes are also specific for certain cleavage sites and anomeric linkages, information which can be exploited in structure elucidation. The key to the analysis of oligosaccharides is to find suitable MS fragments that are specific for one structure and allow the exclusion of alternatives. Mass spectra give ambiguous results when a product ion can originate from two or more precursor ions. Uncertainties in MS/MS spectra can often be removed by applying MSn where product ions can be further examined and assigned to a specific structure.
In recent years, fast atom bombardment (FAB) has been more and more displaced by techniques such as matrix-assisted laser desorption ionization (MALDI) and electrospray ionization (ESI). In ESI, the sample solution is highly charged and sprayed through a capillary into a strong electric field to form a fine mist of charged droplets. The solvent evaporates and produces gas-phase ions. Because samples have to be crystallized with a matrix (e.g., 2,5-dihydroxybenzoic acid or α-cyano-4-hydroxycinnamic acid) that absorbs the energy of the laser and ionizes the sample, MALDI cannot be coupled directly with separation systems (Brüll et al., 1998). MALDI is relatively salt tolerant and offers a large mass range and high sensitivity when coupled to time-of-flight (TOF) analyzers. MALDI-TOF techniques have been proven excellent and simple tools for profiling mixtures of oligosaccharides and can also easily identify sugar modifications such as methylation, acetylation, and sulfation.
The use of powerful mass analyzers with MS/MS (triple quadrupole, TOF/TOF) or even MSn (ion trap, IT) capabilities in conjunction with reverse-phase, normal phase (NP), porous graphitized carbon, size exclusion, ion exchange, or high-performance anion-exchange, liquid chromatographic (LC) or capillary electrophoretic separation methods has increased the role of mass spectrometry for oligosaccharide structural characterization (Schols et al., 1994; Deery et al., 2001; Kabel et al., 2001; Mazumder et al., 2005; Hilz et al., 2006; Coenen et al., 2008). The main function of these coupling techniques is reduction of complexity by separation of isobaric structures that are not resolved by MS.
Fragmentation
Oligosaccharides can generate ions via glycosidic bond cleavages and cross-ring fragmentation. Soft fragmentation techniques such as post-source decay (PSD) do usually not produce abundant cross-ring fragments which are observed in high-energy collision induced dissociation (CID) spectra. By a common convention, ions containing the non-reducing end are labeled Ai (cross-ring), Bi, and Ci (glycosidic), and ions containing the reducing end are labeled Xj (cross-ring), Yj, and Zj (glycosidic) (Domon and Costello, 1988). The subscript i indicates the number of glycosidic bonds cleaved counted from the non-reducing end and j refers to the number of interglycosidic bonds counted from the reducing end (Figure 1A). The superscript (e.g., 0,2A) specifies the particular ring bonds cleaved. Ions from side chains are supplemented by a Greek letter (α, β, γ) assigned with decreasing weight of the side chain (Figure 1B). The numbering from the backbone continues into the side chain. Depending on the ionization and the oligosaccharide structure, glycosidic bond fragments of one or more series (B, C, Y, or Z) are predominantly or even exclusively formed. Additional fragments (Figure 1C) resulting from “rearrangement-elimination” or double cleavages characteristic of 1,2-linked (E) or 1,3-linked (D, F, G) residues and “sugar lactones” (H, W) are diagnostic for the branching pattern (Harvey, 1999; Chai et al., 2001; Spina et al., 2004; Maslen et al., 2007).
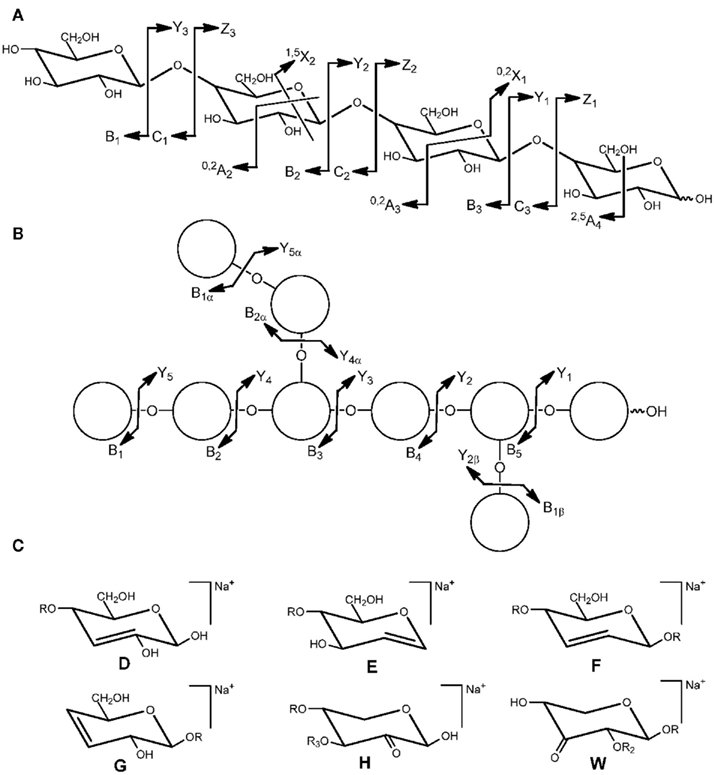
Figure 1. Nomenclature for the cleavages of linear oligosaccharides (A) and branched oligosaccharides (B) and structure of fragment ions (D–H, W) observed via “rearrangement-elimination” and double cleavage (C). R = backbone residue, R2 = H or side chain, R3 = H or side chain (adapted from Domon and Costello, 1988; Spina et al., 2004; Maslen et al., 2007).
Whereas glycosidic bond fragmentation gives information on sequence, cross-ring ions are highly diagnostic for linkage positions and are also indicative for the position of modifications (e.g., acetyl groups). Rules for 1–2, 1–3, 1–4, and 1–6 linkages have been established (Garozzo et al., 1990; Zhou et al., 1990; Hofmeister et al., 1991). For instance, the presence of ions 0,2Ai 60 Da (loss of C2H4O2) and 2,4Ai 120 Da (loss of C4H6O4) lower than the molecular peak (or a Ci ion) in combination with the absence of a loss of 90 Da (C3O3H6) is typical for a 1–4 linkage to hexoses at the reducing end of the molecule or the respective precursor (Ci). An O-2-linked acetyl group would be indicated by an 0,2Ai ion 102 (60 + 42) Da lower. 0,2Xj ions with a loss of 104 Da from Ci ions characteristic for 1–2 linked deoxyhexoses [e.g., rhamnose (Rha)] were observed in pectic rhamnogalacturonan I oligosaccharides (Ralet et al., 2009).
Various algorithms for extracting sequence information from, and assignment of fragment ions in, mass spectra of oligosaccharides are available but are mainly focusing on protein glycans (Gaucher et al., 2000; Lapadula et al., 2005; Tang et al., 2005; Ceroni et al., 2008; Böcker et al., 2011).
Derivatization
The hydroxyl groups of sugars can be naturally O-methylated in the plant or chemically per-O-methylated through a simple derivatization process (Ciucanu and Kerek, 1984; Ciucanu and Caprita, 2007). Besides a significant increase in signal strength, permethylation is applied in order to locate side chains in isomers (e.g., in arabinoxylans). Since the branch points of backbone residues are not methylated during derivatization they can be distinguished from non-branched residues by a mass difference of 14 (–CH3 + H) per branching point. For oligosaccharides containing natively methylated sugars common in pectins and xylans [e.g., 4-O-methyl glucuronic acid (4-O-Me-GlcA)], per-O-deuteromethylation is applied (Dell et al., 1989). The mass difference allows the natively methylated sugars to be distinguished from their artificially methylated counterparts.
Hydrolysis, reduction, and O-acetylation of the per-O-methylated oligosaccharides yields partially methylated alditol acetates that are commonly analyzed by gas chromatography(GC)/MS in order to reveal neutral monosaccharide identity and linkage positions, though sequence information is not obtained (Björndal et al., 1967). For acid-containing polysaccharides, reduction of free uronic groups is achieved by activation with a water-soluble carbodiimide (Kim and Carpita, 1992) or reduction of the permethylated oligosaccharides with lithium triethylborodeuteride (Lerouge et al., 1993). A GC/MS method for simultaneous determination of neutral, acidic, deoxy-, methyl- and even acid-labile 2-keto-3-deoxy-sugars and aceric acid (AceA) via their trimethylsilyl methyl glycosides has also been developed (Doco et al., 2001).
The assignment of ions to a particular series is enabled by oligosaccharide labeling. For example, unsaturated linear oligogalacturonides in ESI negative mode mainly exhibit ions corresponding to the C- and Z-series which are isobaric and therefore undistinguishable. By labeling the reducing end with 18O, all ions comprising the reducing end (X, Y, and Z) will show a two mass unit increase (Körner et al., 1999). Xyloglucan oligosaccharides are often reduced to the corresponding alcohols (Vierhuis et al., 2001).
Introduction of a permanent charge (e.g., quartenary ammonium groups) or an ionizable function (amino or carboxy group) not only increases sensitivity and often attaches a chromophore group to the molecule but also fixes the charge state at one side of the molecule (Harvey, 2000; Gouw et al., 2002; Coenen et al., 2008). This greatly enhances the intensities of ions containing the charged label and supports structure confirmation.
Relative Quantification
Quantification of oligosaccharides by MS is difficult because ionization efficiencies can differ significantly. Other compounds present in a mixture may compete or interfere with ionization. In order to compensate for these effects, internal standards with properties almost identical to the analytes but differing in their mass are used, in practice fulfilled by isotopologues. Absolute quantification requires pure standards but is constrained by the vast variety of possible oligosaccharides structures. For relative quantification of the levels of individual oligosaccharides, each sample is derivatized by an isotopic tag with a unique mass. An example is the labeling with deuterium (D4 and D6) labeled 2-aminopyridine via reductive amination (Yuan et al., 2005). Aliquots of the samples are pooled and analyzed by MS directly or after separation. For chromatography coupling, 13C-isomers are preferred over deuterated ones since the latter can show slight differences in retention time (Zhang et al., 2001). Labeling with commercially available 12C6 and 13C6 isotopes of aniline was recently introduced (Ridlova et al., 2008).
Selected Examples
The following are a few examples of how various MS techniques have been applied to elucidate structural features in various cell wall polysaccharides.
Xyloglucan
Xyloglucan is a highly branched hemicellulose which is an important component of the primary cell wall of higher plants. The branch pattern and composition is variable and has specific physiological functions in the plant including modulating expansion in growing cells (Fry, 1989). Xyloglucan-specific endoglucanases with known specific activity release typical building blocks from xyloglucan. This enabled the development of high-throughput MALDI-TOF screening methods (Oligosaccharide Mass Profiling, OLIMP) for oligosaccharide fingerprinting of plant cell walls (Lerouxel et al., 2002). The method reveals detailed side chain distribution patterns including acetylation. The procedure can be directly applied to specific leaf tissues (Obel et al., 2009) and has been further expanded to characterize a more comprehensive set of cell wall polysaccharides (Westphal et al., 2010). Although structural isomers such as XXLG and XLXG (for nomenclature see Fry et al., 1993) are not resolved by MS they can be distinguished by MALDI-PSD or -TOF/TOF analysis using their distinct ion intensities (Yamagaki et al., 1998). ESI-MS2 was found to be superior to MALDI-PSD-TOF for the analysis of XXFG from black currants and revealed that acetyl groups were located on the galactose residue (Hilz et al., 2006). The glycosyl sequence of unusual oligosaccharides from olive fruit with arabinose substitution (XXSG and XLSG) has been confirmed by MALDI-PSD-TOF MS (Vierhuis et al., 2001).
Xylan
Xylans are hemicelluloses typically found in the secondary cell walls such as those in structural and water conducting tissues. The backbone of α-(1,4)-linked xylosyl (Xyl) residues is decorated with glucuronic acid (GlcA), 4-O-Me-GlcA, and/or arabinose (Ara), depending on the plant species and cell tissue. The biological function of these side chains is unknown but they have a big influence on the polymer properties and also influence the enzymatic degradation (Ebringerová and Heinze, 2000). MALDI-TOF analysis of glucuronoxylan from Arabidopsis after endo-xylanase digestion revealed that the irx8 mutant is devoid of [M + Na]+ ions corresponding to structures with unmethylated GlcA side chains (Persson et al., 2007). For the gux1 gux2 double mutant side chains were even completely absent (Mortimer et al., 2010). The distribution patterns of 4-O-Me-GlcA residues of glucuronoxylans oligosaccharides up to m/z 6000 from softwood and hardwood were determined by MALDI-TOF-MS analysis after partial acid hydrolysis (Jacobs et al., 2001).
MALDI-TOF/TOF fragmentation produced a series of Yn and 1,5X as well as D, E, F, and G ions that confirmed branching and linkage positions of a short complex oligosaccharide sequence β-Xyl-(1,4)-β-Xyl-(1,3)-α-Rha-(1,2)-α-GalA-(1,4)-Xyl. It is typically found in Arabidopsis xylan but is absent in irx7, irx8 and parvus-3 mutants (Brown et al., 2007).
A study of various underivatized linear xylo-oligosaccharides and nine isomeric mono- and disubstituted arabino-xylo-oligosaccharides [up to degree of polymerization (DP) 9] with known arabinose substitution pattern from enzymatic wheat arabinoxylan was performed using negative ESI-QTOF-MS (Quéméner et al., 2006). Two ions m/z 299 (Yiα/C2) and m/z 263 (Ziα/C2) and their ratio were found to be highly indicative of mono- (ratio ∼3–5) or disubstitution (ratio ∼0.1–0.2) at the penultimate xylose. In addition, cross-ring cleavage ions 0,2A2 were only present in linear oligosaccharides but not in the mono-substituted isomers. The absence or presence of a cross-ring fragment 0,2A3 also enabled the discrimination between structures with mono-substitution at the penultimate xylose and mono-substitution on either of its two interior neighbors.
Characteristic D, F, G, and W ions for O-3-linked arabinose and E and H ions for O-2-linked arabinose have been observed for NP-LC isolated 2-aminobenzoic acid-labeled wheat arabinoxylan (DP 4–6) oligosaccharides in MALDI-TOF/TOF MS (Maslen et al., 2007). Together with characteristic cross-ring ions 1,5X, 0,2X, and 3,5A the sequence information and the position of arabinose substitution was obtained.
In another study, endo-xylanase released arabinoxylan oligosaccharides from Switchgrass were analyzed after size-exclusion separation by linkage analysis and ESI-MSn. The interpretation of B and Y ions of spectra up to MS4 of the per-O-methylated compounds confirmed structures with either 5, 6, or 8 pentosyl residues and a backbone of β-(1-4)-Xyl with either single α-Ara or α-Ara-(1-2)-α-Ara side chains at O-3 at the second xylose from the non-reducing end (Mazumder and York, 2010).
An impressive example demonstrating the strength of ESI-CID-MSn using ion trap up to MS4 is the characterization of complex mixtures of enzymatically released wheat arabinoxylan oligosaccharides without further separation steps (Matamoros Fernandéz et al., 2003). The ion m/z 1029.7 (DP 6, [M + Na]+) of the per-O-methylated sample was generated by at least four identified branched isomers with either one (at the 2nd, 3rd, and 4th xylose from the reducing end, respectively) or two arabinose units (at the 3rd xylose).
Pectins
Pectins are the most complex polysaccharides in plant cell walls. With charged residues, complicated and poorly understood branching, and highly variable modifications, structural analysis is particularly difficult. Determining the type and pattern of substitution and modification is critical to understanding the many functional aspects of pectins ranging from structural adhesive components in the wall to whole plant signaling, self-incompatibility, and defense against pathogens (Ridley et al., 2001). General fragmentation patterns of saturated and unsaturated oligogalacturonides (oligoGalA) by MALDI-PSD-TOF-MS have been determined. In positive mode, saturated and unsaturated oligoGalA gave ions of the Bi-, Ci- and Yi-series but the Ci-ion series disappeared in saturated methyl-esterified oligoGalA. Unsaturated methyl-esterified oligoGalA additionally showed 1,5Xi cross-ring cleavage ions (Van Alebeek et al., 2000). Negative-mode ESI-IT-MS/MS produces mainly C and Z-series ions which enables the location of the methyl-esterification and allows sequencing of partially methyl-esterified oligoGalA up to DP 10 (Körner et al., 1999). A loss of 32 Da from the precursor ion can be attributed to methyl-esterification at the GalA at the reducing end (Quéméner et al., 2003b).
Pectins are often also acetylated. The acetyl groups can be assigned by negative ESI-IT-MSn through sequential trapping and fragmentation of C-ions (Quéméner et al., 2003a) and precise location of the acetyl group on O-2 or O-3 can be specified by cross-ring fragments 0,2Ai. The use of anion-exchange chromatography fractionation allows the structural determination of oligoGalA up to DP 5 with multiple methyl and/or acetyl groups at O-2 and/or O-3 (Ralet et al., 2005) and enabled the construction of a hypothetical sugar beet homogalacturonan with a proposed blockwise distribution of acetyl groups (Ralet et al., 2008).
Side-chain determination is particularly complicated in pectins. MALDI-TOF-MS and HPAEC has been used to provide evidence of the complex xylogalacturonan in Arabidopsis thaliana (Zandleven et al., 2007). The ESI-IT-MS fragmentation pattern resulting from MS3 of GalAXyl2 and MS2 of GalA2Xyl3 from apple and potato indicated a dimeric xylose side chain (Zandleven et al., 2006).
Mild acid hydrolysis releases the side chains A–D of the complex pectin rhamnogalacturonan II (RG-II) from citrus pectin at their acid-labile branching points (Séveno et al., 2009). Y-type fragmentation patterns in positive ESI-MS/MS provide sequence information and enabled the structural identification while diagnostic ions derived by a loss of 296 and 340 Da resulting from cleavage of the AceA-rhamnose bond and ring-cleavage of AceA were found upon B-chain fragmentation. This acidic fingerprinting methology was applied to the analysis of RG-II from Arabidopsis mur1 mutant that is deficient in L-fucose (Fuc) synthesis. The [M + Na]+ ion as well as the fragmentation pattern confirmed that the 2-O-Me-Xyl-L-Fuc segment was replaced by L-galactose (Gal) in the A-chain and 2-O-Me-L-Gal instead of 2-O-Me-L-Fuc was incorporated into the B chain.
Conclusion and Outlook
Mass spectrometry has been proven to be a sensitive, fast, and powerful tool for the screening of cell wall changes as well as for the structural analysis of carbohydrates in complex mixtures providing sequence and branching information including the location of modifications. MS analysis benefits from readily available specific enzymes that selectively release oligosaccharides. Although efforts in this direction have been made (Bauer et al., 2006), discovery of new enzymes with specific activity on specialized linkages in cell wall compounds which cannot be efficiently hydrolyzed or for which enzymes have not been identified (e.g., RG-II) would greatly improve polysaccharide analysis by MS. The complexity of mixtures with many isobaric structures can further be overcome by coupling mass spectrometers to ion mobility (IM) spectrometers that are capable of separating gas-phase ions by their size/charge ratio as well as their interaction with a buffer gas (Kanu et al., 2008). One can envision interesting 3-dimensional separation techniques by using various combinations of chromatography, IM, and MS. Through an expanding repertoire of sensitive techniques, MS will play an ever increasing role in the field of plant cell wall analysis.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The author acknowledges funding from the Energy Biosciences Institute and critical discussions and assistance from Valerie Mitchell and Dr Heather Youngs.
References
Bauer, S., Vasu, P., Persson, S., Mort, A. J., and Somerville, C. R. (2006). Development of a suite of polysaccharide-degrading enzymes for analyzing plant cell walls. Proc. Natl. Acad. Sci. U.S.A. 103, 11417–11422.
Björndal, H., Lindberg, B., and Svensson, S. (1967). Mass spectrometry of partially methylated alditol acetates. Carbohydr. Res. 5, 433–440.
Böcker, S., Kehr, B., and Rasche, F. (2011). Determination of glycan structure from tandem mass spectra. Trans. Comput. Biol. Bioinformatics 8, 976–986.
Brown, D. M., Goubet, F., Wong, V. W., Goodacre, R., Stephens, E., Dupree, P., and Turner, S. R. (2007). Comparison of five xylan synthesis mutants reveals new insight into the mechanisms of xylan synthesis. Plant J. 52, 1154–1168.
Brüll, L., Huisman, M., Schols, H., Voragen, F., Critchley, G., Thomas-Oates, J., and Haverkamp, J. (1998). Rapid molecular mass and structural determination of plant cell wall-derived oligosaccharides using off-line high-performance anion-exchange chromatography/mass spectrometry. J. Mass. Spectrom. 33, 713–720.
Ceroni, A., Maass, K., Geyer, H., Geyer, R., Dell, A., and Haslam, S. M. (2008). GlycoWorkbench: A tool for the computer-assisted annotation of mass spectra of glycans. J. Proteome Res. 7, 1650–1659.
Chai, W., Piskarev, V., and Lawson, A. M. (2001). Negative electrospray mass spectrometry of neutral underivatized oligosaccharides. Anal. Chem. 73, 651–657.
Ciucanu, I., and Caprita, R. (2007). Per-O-methylation of neutral carbohydrates directly from aqueous samples for gas chromatography and mass spectrometry analysis. Anal. Chim. Acta 585, 81–85.
Ciucanu, I., and Kerek, F. (1984). A simple and rapid method for the permethylation of carbohydrates. Carbohydr. Res. 131, 209–217.
Coenen, G.-J., Kabel, M. A., Schols, H. A., and Voragen, A. G. J. (2008). CE-MSn of complex pectin-derived oligomers. Electrophoresis 29, 2101–2111.
Deery, M. J., Stimson, E., and Chappell, C. G. (2001). Size exclusion chromatography/mass spectrometry applied to the analysis of polysaccharides. Rapid Commun. Mass Spectrom. 15, 2273–2283.
Dell, A., Khoo, K.-H., Panico, M., McDowell, R. A., Etienne, A., Reason, A. J., and Morris, H. R. (1989). “FAB-MS and ESI-MS of glycoproteins,” in Glycobiology. A Practical Approach, Vol. 125, eds M. Fukuda, and A. Kobata (Oxford: Oxford University Press), 187–222.
Doco, T., O’Neill, M. A., and Pellerin, P. (2001). Determination of the neutral and acidic glycosyl-residue compositions of plant polysaccharides by GC-EI-MS analysis of the trimethylsilyl methyl glycoside derivatives. Carbohydr. Polym. 46, 249–259.
Domon, B., and Costello, C. E. (1988). A systematic nomenclature for carbohydrate fragmentation in FAB-MS/MS spectra of glycoconjugates. Glycoconjugate J. 5, 397–409.
Duus, J. Ø., Gotfredsen, C. H., and Bock, K. (2000). Carbohydrate structural determination NMR spectroscopy: Modern methods and limitations. Chem. Rev. 100, 4589–4614.
Ebringerová, A., and Heinze, T. (2000). Xylan and xylan derivatives – biopolymers with valuable properties, 1. Naturally occurring xylans structures, isolation procedures and properties. Macromol. Rapid Commun. 21, 542–556.
Fry, S. C., York, W. S., Albersheim, P., Darvill, A., Hayashi, T., Joseleau, J.-P., Kato, Y., Pérez Lorences, E., Maclachlan, A. J., Grant Reid, J. S., Seitz, H. U., Selvendran, R. R., Voragen, A. G. J., and White, A. R. (1993). An umambiuos nomenclature for xyloglucan-derived oligosaccharides. Physiol Plant 89, 1–3.
Garozzo, D., Giuffrida, M., Impallomeni, G., Ballistreri, A., and Montaudo, G. (1990). Determination of linkage position and identification of the reducing end in linear oligosaccharides by negative ion fast atom bombardment mass spectrometry. Anal. Chem. 62, 279–286.
Gaucher, S. P., Morrow, J., and Leary, J. A. (2000). STAT: A saccharide topology analysis tool used in combination with tandem mass spectrometry. Anal. Chem. 72, 2331–2336.
Gouw, J. W., Burgers, P. C., Trikoupis, M. A., and Terlouw, J. K. (2002). Derivatization of small oligosaccharides prior to analysis by matrix-assisted laser desorption/ionization using glycidyltrimethylammonium chloride and Girard’s reagent T. Rapid Commun. Mass Spectrom. 16, 905–912.
Harvey, D. J. (1999). Matrix-assisted laser desoption/ionization mass spectrometry of carbohydrates. Mass Spectrom. Rev. 18, 349–450.
Harvey, D. J. (2000). Electrospray mass spectrometry and fragmentation of N-linked carbohydrates derivatized at the reducing terminus. J. Am. Soc. Mass Spectrom. 11, 900–915.
Hilz, H., de Jong, L. E., Kabel, M. A., Schols, H. A., and Voragen, A. G. J. (2006). A comparison of liquid chromatography, capillary electrophoresis, and mass spectrometry methods to determine xyloglucan structures in black currants. J. Chromatogr. A 1133, 275–286.
Hofmeister, G., Zhou, Z., and Leary, J. A. (1991). Linkage position determination in lithium-cationized disaccharides: Tandem mass spectrometry and semiempirical calculations. J. Am. Chem. Soc. 113, 5964–5970.
Jacobs, A., Larsson, P. T., and Dahlman, O. (2001). Distribution of uronic acids in xylans from various species of soft- and hardwood as determined by MALDI mass spectrometry. Biomacromolecules 9, 979–990.
Kabel, M. A., Schols, H. A., and Voragen, A. G. J. (2001). Mass determination of oligosaccharides by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry following HPLC, assisted by on-line desalting and automated sample handling. Carbohydr. Polym. 44, 161–165.
Kanu, A. B., Dwivedi, P., Tam, M., Matz, L., and Hill, H. H. Jr. (2008). Ion mobility-mass spectrometry. J. Mass. Spectrom. 43, 1–22.
Kim, J. B., and Carpita, N. C. (1992). Changes in esterification of the uronic acid groups of cell wall polysaccharides during elongation of maize coleoptiles. Plant Physiol. 98, 646–653.
Körner, R., Limberg, G., Christensen, T. M. I. E., Mikkelsen, J. D., and Roepstorff, P. (1999). Sequencing of partially methyl-esterified oligogalacturonates by tandem mass spectrometry and its use to determine pectinase specificity. Anal. Chem. 71, 1421–1427.
Lapadula, A. J., Hatcher, P. J., Hanneman, A. J., Ashline, D. J., Zhang, H., and Reinhold, V. N. (2005). Congruent strategies for carbohydrate sequencing. 3. OSCAR: an algorithm for assigning oligosaccharide topology from MSn data. Anal. Chem. 77, 6271–6279.
Lerouge, P., O’Neill, M. A., Darvill, A. G., and Albersheim, P. (1993). Structural characterization of endo-glycanase-generated oligoglycosyl side chains of rhamnogalacturonan I. Carbohydr. Res. 243, 359–371.
Lerouxel, O., Choo, T. S., Séveno, M., Usadel, B., Faye, L., Lerouge, P., and Pauly, M. (2002). Rapid structural phenotyping of plant cell wall mutants by enzymatic oligosaccharide fingerprinting. Plant Physiol. 130, 1754–1763.
Maslen, S. L., Goubet, F., Adam, A., Dupree, P., and Stephens, E. (2007). Structure elucidation of arabinoxylan isomers by normal phase HPLC-MALDI-TOF/TOF-MS/MS. Carbohydr. Res. 342, 724–735.
Matamoros Fernandéz, L. E., Obel, N., Scheller, H. V., and Roepstorff, P. (2003). Characterization of plant oligosaccharides by matrix-assisted laser desorption/ionization and electrospray mass spectrometry. J. Mass. Spectrom. 38, 427–437.
Mazumder, K., and York, W. S. (2010). Structural analysis of arabinoxylans isolated from ball-milled switchgrass biomass. Carbohydr. Res. 345, 2183–2193.
Mazumder, S., Lerouge, P., Loutelier-Bourhis, C., Driouich, A., and Ray, B. (2005). Structural characterization of hemicellulosic polysaccharides from Benincasa hispida using specific enzyme hydrolysis, ion exchange chromatography and MALDI-TOF mass spectrometry. Carbohydr. Polym. 59, 231–238.
Mortimer, J. C., Miles, G. P., Brown, D. M., Zhang, Z., Segura, M. P., Weimar, T., Yu, X., Seffen, K. A., Stephens, E., Turner, S. R., and Dupree, P. (2010). Absence of branches from xylan in Arabidopsis gux mutants reveals potential for simplification of lignocellulosic biomass. Proc. Natl. Acad. Sci. U.S.A. 107, 17409–17414.
Obel, N., Erben, V., Schwarz, T., Kühnel, S., Fodor, A., and Pauly, M. (2009). Microanalysis of plant cell wall polysaccharides. Mol. Plant 2, 922–932.
Persson, S., Caffall, K. H., Freshour, G., Hilley, M. T., Bauer, S., Poindexter, P., Hahn, M. G., Mohnen, D., and Somerville, C. (2007). The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19, 237–255.
Quéméner, B., Cabrera Pino, J. C., Ralet, M.-C., Bonnin, E., and Thibault, J.-F. (2003a). Assignment of acetyl groups to O-2 and/or O-3 of pectic oligogalacturonides using negative electrospray ionization ion trap mass spectrometry. J. Mass. Spectrom. 38, 641–648.
Quéméner, B., Désiré, C., and Lahaye, M. (2003b). Structural characterisation by both positive- and negative-ion electrospray mass spectrometry of partially methyl-esterified oligogalacturonides purified by semi-preparative high-performance anion-exchange chromatography. Eur. J. Mass. Spectrom 9, 45–60.
Quéméner, B., Ordaz-Ortiz, J. J., and Saulnier, L. (2006). Structural characterization of underivatized arabino-xylo-oligosaccharides by negative-ion electrospray mass spectrometry. Carbohydr. Res. 341, 1834–1847.
Ralet, M.-C., Cabrera, J. C., Bonnin, E., Quéméner, B., Hellín, P., and Thibault, J.-F. (2005). Mapping sugar beet pectin acetylation pattern. Phytochemistry 66, 1832–1843.
Ralet, M.-C., Crépeau, M.-J., and Bonnin, E. (2008). Evidence for a blockwise distribution of acetyl groups onto homogalacturonans from commercial sugar beet (Beta vulgaris) pectin. Phytochemistry 69, 1903–1909.
Ralet, M.-C., Lerouge, P., and Quéméner, B. (2009). Mass spectro-metry for pectin structure analysis. Carbohydr. Res. 344, 1798–1807.
Ridley, B. L., O’Neill, M. A., and Mohnen, D. (2001). Pectins: structure, biosynthesis, and oligogalacturonide-related signaling. Phytochemistry 57, 929–967.
Ridlova, G., Mortimer, J. C., Maslen, S. L., Dupree, P., and Stephens, E. (2008). Oligosaccharides relative quantification using isotope tagging and normal-phase liquid chromatography/mass spectrometry. Rapid Commun. Mass Spectrom. 22, 2723–2730.
Schols, H. A., Mutter, M., Voragen, A. G. J., Niessen, W. M. A., van der Hoeven, R. A. M., van der Greef, J., and Bruggink, C. (1994). The use of combined high-performance anion-exchange chromatography-thermospray mass spectrometry in the structural analysis of pectic oligosaccharides. Carbohydr. Res. 261, 335–342.
Séveno, M., Voxeur, A., Rihouey, C., Wu, A.-M., Ishii, T., Chevalier, C., Ralet, M.-C., Driouich, A., Marchant, A., and Lerouge, P. (2009). Structural characterization of the pectic polysaccharide rhamnogalacturonan II using an acidic fingerprinting methodology. Planta 230, 947–957.
Spina, E., Sturiale, L., Romeo, D., Impallomeni, G., Garozzo, D., Waidelich, D., and Glueckmann, M. (2004). New fragmentation mechanisms in matrix-assisted laser desorption/ionization time-of-flight tandem mass spectrometry of carbohydrates. Rapid Commun. Mass Spectrom. 18, 392–398.
Tang, H., Mechref, Y., and Novotny, M. V. (2005). Automated interpretation of MS/MS spectra of oligosaccharides. Bioinformatics 21(Suppl. 1), i431–i439.
Van Alebeek, G.-J. W.M., Zabotina, O., Beldman, G., Schols, H. A., and Voragen, A. G. J. (2000). Structural analysis of (methyl-esterified) oligogalacturonides using post-source decay matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J. Mass. Spectrom. 35, 831–840.
Vierhuis, E., York, W. S., Kolli, V. S. K., Vincken, J.-P., Schols, H. A., Van Alebeek, G.-J. W. M., and Voragen, A. G. J. (2001). Structural analysis of two arabinose containing oligosaccharides derived from olive fruit xyloglucan: XXSG and XLSG. Carbohydr. Res. 332, 285–297.
Westphal, Y., Schols, H. A., Voragen, A. G. J., and Gruppen, H. (2010). MALDI-TOF MS and CE-LIF fingerprinting of plant cell wall polysaccharide digests as a screening tool for Arabidopsis cell wall mutants. J. Agric. Food Chem. 58, 4644–4652.
Yamagaki, T., Mitsuishi, Y., and Nakanishi, H. (1998). Determination of structural isomers of xyloglucan octasaccharide using post-source decay fragment analysis in MALDI-TOF mass spectrometry. Tetrahedron Lett. 39, 4051–4054.
Yuan, J., Hashii, N., Kawasaki, N., Itoh, S., Kawanishi, T., and Hayakawa, T. (2005). Isotope tag method for quantitative analysis of carbohydrates by liquid chromatography-mass spectrometry. J. Chromatogr. A 1067, 145–152.
Zandleven, J., Beldman, G., Bosveld, M., Schols, H. A., and Voragen, A. G. J. (2006). Enzymatic degradation studies of xylogalacturonans from apple and potato, using xylogalacturonan hydrolase. Carbohydr. Polym. 65, 495–503.
Zandleven, J., Oxenbøll Sørensen, S., Harholt, J., Beldman, G., Schols, H. A., Scheller, H. V., and Voragen, A. G. J. (2007). Xylogalacturonan exists in cell walls from various tissues of Arabidopsis thaliana. Phytochemistry 68, 1219–1226.
Zhang, R., Sioma, C. S., Wang, S., and Regnier, F. E. (2001). Fractionation of isotopically labeled peptides in quantitative proteomics. Anal. Chem. 73, 5142–5149.
Keywords: mass spectrometry, cell wall, carbohydrates, oligosaccharides
Citation: Bauer S (2012) Mass spectrometry for characterizing plant cell wall polysaccharides. Front. Plant Sci. 3:45. doi: 10.3389/fpls.2012.00045
Received: 30 January 2012; Paper pending published: 16 February 2012;
Accepted: 23 February 2012; Published online: 12 March 2012.
Edited by:
Jose Manuel Estevez, University of Buenos Aires and Conicet, ArgentinaCopyright: © 2012 Bauer. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Stefan Bauer, Energy Biosciences Institute, University of California, 204 Cavin Lab, MC 5230, Berkeley, CA 94720, USA. e-mail:c3RlZmFuLmJhdWVyQGJlcmtlbGV5LmVkdQ==
