- Department of Plant Pathology, Physiology, and, Weed Science, Virginia Tech, Blacksburg, VA, USA
For the past decade, our understanding of the plant purine uptake permease (PUP) transporter family was primarily oriented on purine nucleobase substrates and their tissue-specific expression patterns in Arabidopsis. However, a tobacco PUP-like homolog demonstrating nicotine uptake permease activity was recently shown to affect both nicotine metabolism and root cell growth. These new findings expand the physiological role for PUP-like transporters to include plant secondary metabolism. Molecular evolution analyses of PUP-like transporters indicate they are distinct group within an ancient super family of drug and metabolite transporters (DMTs). The PUP-like family originated during terrestrial plant evolution sometime between the bryophytes and the lycophytes. A phylogenetic analysis indicates that the PUP-like transporters were likely derived from a pre-existing nucleotide-sugar transporter family within the DMT super family. Within the lycophyte Selaginella, there are three paralogous groups of PUP-like transporters. One of the three PUP-like paralogous groups showed an extensive pattern of gene duplication and diversification within the angiosperm lineage, whereas the more ancestral PUP-like paralogous groups did not. Biochemical characterization of four closely related PUP-like paralogs together with model-based phylogenetic analyses indicate both subfunctionalization and neofunctionalization during the molecular evolution of angiosperm PUP-like transporters. These findings suggest that members of the PUP-like family of DMT transporters are likely involved in diverse primary and secondary plant metabolic pathways.
Introduction
The purpose of this mini review is to integrate recent molecular and phylogenetic findings about a tobacco nicotine uptake permease (NUP) with prior knowledge about the closely related plant-specific purine uptake permease (PUP) transporter family. The founding member of the PUP-like transporter family was Arabidopsis thaliana PUP1 (AtPUP1) that was identified in a cDNA complementation screen of a yeast adenine uptake mutant (Gillissen et al., 2000). Yeast expressing the AtPUP1 cDNA show high affinity adenine uptake activity that is abrogated by protonophore-inhibitors, indicating a proton-facilitated substrate symport mechanism. This adenine uptake activity is also efficiently competed by structurally-related purine ring-containing cytokinins and caffeine. Nicotine also reduces AtPUP1-mediated adenine uptake rates, though considerably less efficiently than the above purine compounds.
In Arabidopsis, the PUP-like genes comprise a moderate size gene family. There are 21 PUP-like genes in Arabidopsis, including one pseudogene. PUP-like transporters typically have 10 predicted transmembrane spanning domains. To date, only three Arabidopsis PUP-like genes have been the focus of detailed studies into expression profiles and substrate specificity (Gillissen et al., 2000; Burkle et al., 2003). This limited characterization of the Arabidopsis PUP-like transporter family nevertheless demonstrates evolutionary patterns of both subfunctionalization and neofunctionalization. In a parsimony phylogenetic analysis comprised of only Arabidopsis PUP-like proteins, AtPUP1 and AtPUP2 showed closest sequence identity (64%) and phylogenetic association to each other, than to any other AtPUP transporter. Both AtPUP1 and AtPUP2 transport adenine and cytokinins, albeit with different kinetics. AtPUP1 gene expression localizes to leaf hydathode tissue (specifically epithem cells) and the stigmatic surface, whereas AtPUP2 localizes to vascular tissue, specifically to the phloem. Therefore, AtPUP1 and AtPUP2 have diversified in terms of tissue specificity, but not in terms of ligand specificity. Thus, AtPUP1 and AtPUP2 display a pattern of subfunctionalization (Force et al., 1999). In contrast, AtPUP3 is more distantly related to AtPUP1 (29% protein identity) and AtPUP3 gene expression is restricted to pollen. Heterologously expressed recombinant AtPUP3 does not transport adenine during the same conditions as AtPUP1 and AtPUP2. AtPUP3 is presumed to transport another, as yet to be determined, substrate into pollen cells. Thus, in terms of both cell type-specific expression and substrate recognition, AtPUP3 exhibits patterns of neofunctionalization relative to AtPUP1 and AtPUP2. AtPUP1 is presumed to be involved in purine and/or cytokinin uptake during leaf guttation and at the stigma, whereas AtPUP2 is postulated to be involved with uptake of these substrates into the phloem. These predicted physiological roles await confirmation using plants with altered AtPUP1 or AtPUP2 expression levels. Given these patterns of both subfunctionalization and neofunctionalization, together with the relatively low degree of sequenced identity between these Arabidopsis transporters, ranging from 16% to 64% identity, it is likely that other PUP-like transporters recognize different substrates involved in either primary or secondary metabolism.
A PUP-Like Transporter NUP1 Affects Pyridine Alkaloid Metabolism and Physiology
The physiological impact of PUP-like transporters was recently expanded to include plant secondary metabolism. Transcriptional profiling of a tobacco mutant affecting total alkaloid accumulation levels and nicotine biosynthetic gene expression levels identified transcripts in roots that are coordinately regulated with several nicotine biosynthetic genes (Kidd et al., 2006). One such cDNA fragment encodes a predicted peptide with 56% identity and 67% similarity to AtPUP1. The corresponding full-length Nicotiana tabacum cDNA encodes a protein with nicotine uptake permease activity, called NUP1 (Hildreth et al., 2011). In contrast to AtPUP1, NUP1-mediated nicotine uptake activity is not efficiently competed by either purines or cytokinins, neither it is inhibited by closely-related pyridine alkaloids nor less-related tropane alkaloids. Therefore, NUP1 shows a high degree of substrate specificity for nicotine and is not an ortholog of AtPUP1. A NUP1–GFP fusion localizes primarily to the tobacco plasma membrane, suggesting NUP1 transports apoplastic nicotine into the cytoplasm. Steady state NUP1 mRNA levels accumulate to highest levels at root tips, where nicotine biosynthesis is localized (Dawson, 1942a,b; Baldwin, 1988).
The association of NUP1 with nicotine metabolism was confirmed using tobacco lines with reduced steady state NUP1 mRNA levels. Transgenic NUP1-RNAi tobacco plants have reduced foliar nicotine levels (Hildreth et al., 2011). However, the lower foliar nicotine accumulation levels are not due to altered nicotine transport from roots to shoots per se, because NUP1-RNAi plants are not compromised in their ability to proportionally transport exogenous nicotine fed to roots into leaves. Rather, less foliar nicotine is correlated with lower nicotine levels in the roots. During non-induced conditions NUP1-RNAi hairy root lines have lower nicotine levels in the hairy roots. Thus, reduced NUP1 transcript levels correlate with significant reductions in nicotine accumulation levels in leaves, roots, and hairy roots. During non-inducing culture conditions the NUP1-RNAi hairy root cultures showed significantly more nicotine accumulation in the spent culture media, compared to wild type. It was not determined whether the increased nicotine in the culture media of NUP1-RNAi lines was due to nicotine released from the open xylem vessels at the ends of the hairy roots, nicotine release from the root epidermis, or diminished nicotine re-uptake from the rhizosphere. While there were consistent patterns of decreased nicotine accumulation levels in NUP1-RNAi roots, during several conditions examined there was an overall poor correlation of reduced nicotine levels with corresponding reductions the steady state transcript levels of several essential nicotine biosynthetic genes. Thus, while NUP1 expression levels clearly affect overall nicotine metabolism, the molecular mechanism by which NUP1 affects nicotine accumulation levels is currently poorly understood.
Nicotine biosynthesis is one of a few plant specialized metabolic pathways that are integrated with ongoing root growth. On one hand, root growth is essential for net nicotine accumulation levels (Solt, 1957; Baldwin, 1988). On the other hand, tobacco roots are not immune to the toxic effects of either exogenous nicotine treatment (Baldwin and Callahan, 1993; Baldwin and Ohnmeiss, 1994; Shoji et al., 2009) or the stimulation of endogenous nicotine biosynthesis (Hildreth et al., 2011). Both conditions result in reduced wild type root growth. This indicates there are complex interactions between root growth and nicotine metabolism/accumulation that could have homeostatic properties. Another interesting phenotype of NUP1-RNAi lines is they exhibit increased seedling root elongation rates during both control conditions and several treatments that increase overall nicotine accumulation levels. This latter result highlights the current lack of knowledge in the field about the molecular mechanisms of how nicotine accumulation levels are coordinated with root growth. Therefore, it is noteworthy that NUP1 is the first molecular component demonstrated to affect both root growth and nicotine metabolism.
NUP1 may also have implications in nicotine physiology far beyond the root tip. Whole plant nicotine levels are held at allometric set points relative to total nicotine-free plant biomass (Ohnmeiss and Baldwin, 1994; Baldwin and Karb, 1995). How allometric control of whole plant nicotine levels is enforced, when the site of nicotine biosynthesis is restricted to growing root tips yet most nicotine is unequally distributed throughout the plant, is currently an enigma. Clearly, simple models of metabolic control based upon homeostatic apoplastic nicotine levels cannot account for the observed whole plant nicotine allometry. Perhaps allometric control of nicotine biosynthesis is enforced by the integration of nicotine biosynthesis with root tip growth. An appealing aspect of this model for whole plant nicotine allometric control is that growing root tips are a classic sink tissue whose growth is a function of shoot primary productivity that in turn is scaled to shoot biomass. If this is the case, NUP1 has several properties that make it a good candidate for participation in whole plant nicotine allometric set points. NUP1 presumably transports apoplastic nicotine into cells near the root tip, reduced NUP1 expression levels results in reduced root nicotine accumulation levels, and concomitant increased root elongation rates. The remainder of this article will focus on the molecular evolution of the PUP-like transporter family.
PUP-Like Transporters Originated Early During Terrestrial Plant Evolution, Possibly from a DMT UDP-Sugar Transporter Family
Two previous molecular evolution studies indicated that PUP-like homologs were restricted to plant taxa (Gillissen et al., 2000; Hildreth et al., 2011). Another study, based upon reciprocal BLASTP searches, assigned the PUP transporters to an ancient drug and metabolite transporter (DMT) super family (Jack et al., 2001). The plant PUP-like transporters comprise a distinct family within the DMT super family named the plant organocation permease (POP) family and were given the transporter classification (TC) code designation 2.A.7.14. A NUP1 BLAST query on the NCBI Conserved Domain Database (CDD) identified a conserved pfam 03151 domain (BLASTP expectation value = 7.1 e-28). On CDD the pfam 03151 domain is associated with the EamA super family, citing Jack et al. (2001) as the primary reference for the EamA super family, despite the said article does not use the EamA nomenclature. This highlights the considerable diversity in nomenclature associated with PUP-like transporters, both in the literature and promulgated in electronic databases. Henceforth, this mini review will use both the International Union of Biochemistry and Molecular Biology-approved TC coding (Jack et al., 2001; Saier Jr. et al., 2009) and the PUP terminology to discuss the molecular evolution of the PUP/2.A.7.14 family of transporters.
Published PUP and NUP similarity searches and phylogenies are consistent with the PUP/2.A.7.14 family originating in terrestrial plants after the bryophytes (Gillissen et al., 2000; Hildreth et al., 2011). However, it is improbable that a progenitor PUP/2.A.7.14 transporter formed entirely de novo in plants. Rather, it is more likely that it was derived from a different DMT family pre-existing in the plantae. Jack et al. (2001) used the program GAP (Devereux et al., 1984) to assign binary affinity of PUP/2.A.7.14 family to the DMT 2.A.7.3 family of prokaryotic DMT transporters. This binary association barely met the minimum GAP score SD value cutoff of 9, and this purported association has not been independently corroborated by phylogenetic analyses. Since a BLASTP e-value cutoff of e-4 identified only plant PUP/2.A.7.14 homologs (Hildreth et al., 2011), a new PUP/2.A.7.14 phylogenetic analysis was performed that used the transitive nature of protein homology sensu stricto to identify distantly related DMT homologs. Briefly, three PUP-like homologs representing three well-supported clades in Selaginella were used as queries in the SSEARCH program on a local NCBI RefSeq database. Proteins returned by at least two searches established homology, regardless of expectation value. A total of 185 proteins meeting this criterion were aligned and subjected to four independent Bayesian runs. The resulting joint Bayesian phylogeny is displayed in Figure 1. Again, no PUP/2.A.7.14 homologs were identified in bryophyte or algal taxa, further supporting the hypothesis that the PUP/2.A.7.14 family originated during terrestrial plant evolution. Plant PUP/2.A.7.14 transporters formed a well-supported monophyletic group (posterior probability equal to 1.0). Because this phylogeny contained eukaryotic taxa other than plants, it effectively rooted the monophyletic PUP/2.A.7.14 clade and established PUP/2.A.7.14 paralogous relationships within the plantae. The most basal plant PUP/2.A.7.14 clade included both Selaginella and angiosperm homologs and may represent the earliest PUP/2.A.7.14 orthologous group. The large PUP/2.A.7.14 clade was rooted within a tritomy of other eukaryotic DMT transporters, which precludes establishing the relative branch order. Nevertheless, 78% of these eukaryotic DMT transporters were most similar to 2.A.7.12 family members that are presumed to transport nucleotide–sugar conjugates. The next more basal eukaryotic clade was likewise comprised of 2.A.7.12 family members and it also defined the bacteria – eukaryotic split with strong Bayesian support. Eighty-six percent of the prokaryotic DMT transporters belonged to the 2.A.7.3 family. This is the same family identified in the binary association analysis reported in the initial characterization of the DMT super family (Jack et al., 2001). Given the abundance of 2.A.7.12 family members in both the basal and sister groups to the monophyletic plant PUP/2.A.7.14 transporters, it is likely that the PUP/2.A.7.14 family arose from a pre-existing 2.A.7.12 family of nucleotide-sugar transporters sometime after the bryophytes, and was well established by the lycophytes.
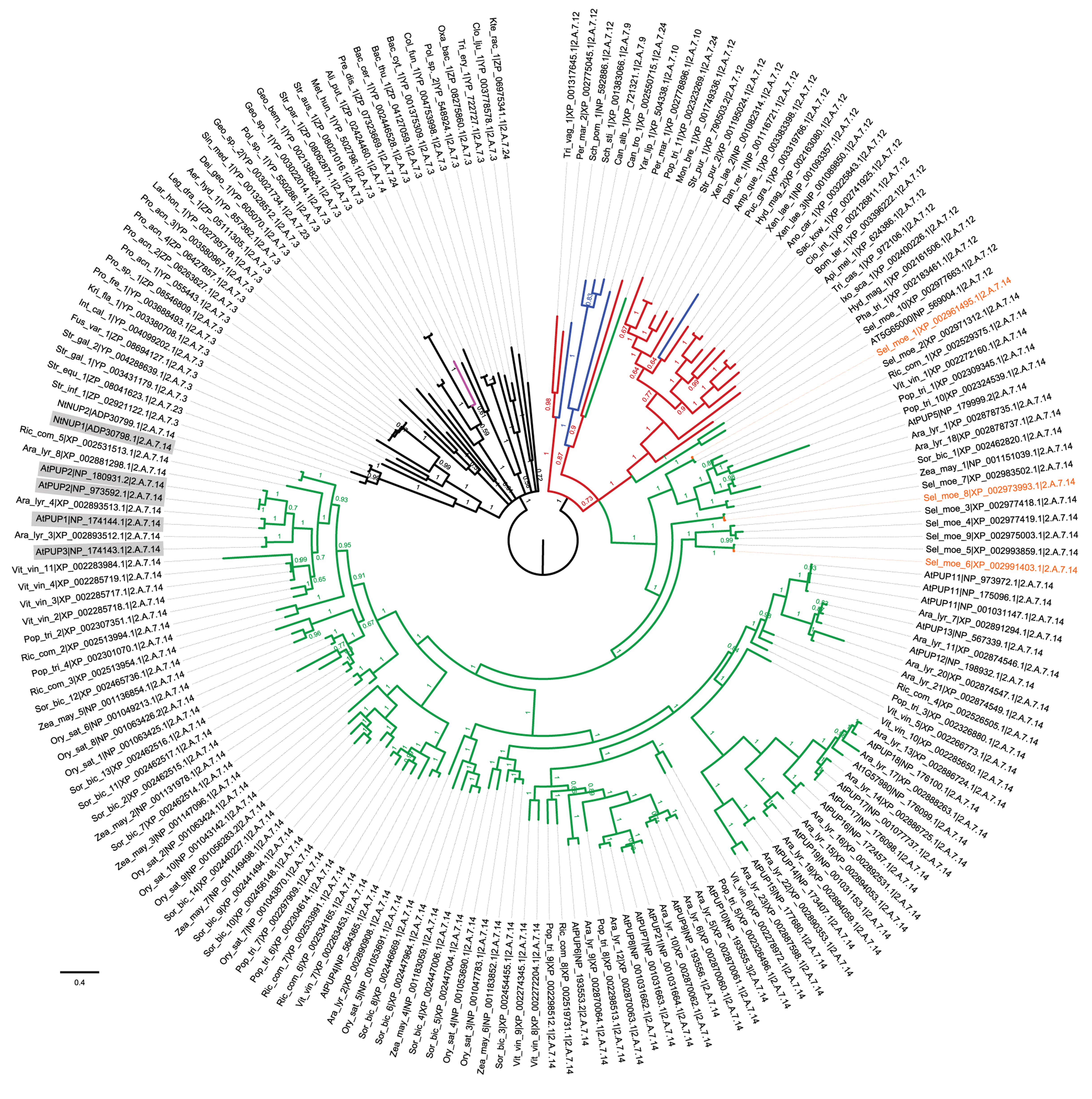
FIGURE 1. Plant PUP-like transporters are closely related to the DMT 2.A.7.12 family of nucleotide-sugar transporters. Bayesian phylogram of DMT homologs related to the PUP/2.A.7.14 family. Taxonomic coding was as follows: black, bacteria; pink, archaea; red, diverse eukaryota; green, plantae; and blue, fungi. Orange indicates three Selaginella homolog queries used in SSEARCH. Arabidopsis and tobacco PUP-like transporters were labeled by their published protein names, whereas other PUP-like homologs were identified by a three letter abbreviated Genus_species_serial number|GenBank accession|TC code. TC codes were assigned to each transporter based upon the TC code of the optimal BLASTP homolog identified at http://www.tcdb.org. Biochemically characterized PUP-like transporters were highlighted in gray. The tree was rooted on the node separating prokaryotes from eukaryotes. Methods: Homology was defined by three Selaginella query sequences (orange) using SSEARCH (Pearson, 1996) with the BLOSUM 50 matrix on a local copy of the NCBI RefSeq database. Proteins identified in any two of the three queries defined homology sensu stricto. These were combined into a non-redundant dataset that was aligned using MAFFT (Katoh et al., 2005) and analyzed using MrBayes ver. 3.1.2 (Ronquist and Huelsenbeck, 2003) implementing a LG + G + F model (determined by ProtTest; Darriba et al., 2011) with four independent runs each with eight heated chains (temp 0.1) for 10 million generations, and a 2.5 million generation burn-in. Convergence diagnostics were evaluated using Tracer (Rambaut and Drummond, 2003). Skyline plots of all parameters showed stable distributions of posterior parameter estimates. The effective sample sizes (ESS) of the LnL values ranged from 837 to 948, and the ESS values for the TL parameter ranged from 450 to 626. Therefore, the four runs did not show evidence of non-convergence, so a joint analysis was performed. The resulting consensus tree was displayed and annotated using Figtree (Rambaut, 2009).
There is asymmetry in the pattern of PUP/2.A.7.14 transporter evolution within the plantae. Of the three paralogous PUP/2.A.7.14 groups in Selaginella: one group is restricted only to Selaginella, one group has just one or two homologs in diverse taxa, whereas the third paralogous group comprises a sister group to an extensive pattern of gene duplication during angiosperm evolution. The widespread diversification of the PUP/2.A.7.14 gene family in the angiosperms likely promoted increased complexity in both plant development and plant chemical diversity. It follows that the phylogenetically more derived homologs may be involved with correspondingly more derived plant traits/characters. Consistent with this idea, the phylogenetically more derived AtPUP1, AtPUP2, APUP3 transporters show a pattern of subfunctionalization with respect to their restricted cell-type expression patterns in Arabidopsis, whereas AtPUP3 also shows neofunctionalization in substrate specificity. Similarly, tobacco NUP1 shows neofunctionalization with respect to both tissue localization and substrate specificity. These evolutionary patterns suggests that PUP/2.A.7.14 transporters were important for the evolution of highly specialized plant tissue/cell types. Assuming the patterns of subfunctionalization and neofunctionalization observed with these four characterized PUP/2.A.7.14 members are typical for this transporter family, then the other phylogenetically highly-derived PUP/2.A.7.14 family members shown in Figure 1 are expected to show considerable diversity in both tissue-specific expression and/or ligand specificity/selectivity.
The discovery that NUP1 affects pyridine alkaloid metabolism has implications for other plant alkaloid biosynthetic pathways. Both tropane and monoterpenoid indole alkaloid (MIA) biosynthetic pathways are cell non-autonomous processes, in which specific biosynthetic enzymes are expressed in non-adjacent specialized cell types (reviewed in Ziegler and Facchini, 2008). This means that alkaloid intermediates must be transported both out of and into dispersed cells. Despite the recognized importance of these inferred transport processes during tropane and MIA biosynthesis, specific transporters mediating these processes are a conspicuous gap in current knowledge. While it is currently uncertain if nicotine biosynthesis is a cell autonomous process, NUP1 nevertheless provides an important precedent that a member of the PUP/2.A.7.14 transporter family can impact the metabolism of at least one plant alkaloid. This may have important implications for the biosynthesis other plant alkaloids for which intercellular transport of alkaloid intermediates is believed to be an essential process.
Outlook for PUP-Like Transporter Research
The discovery that NUP1 is involved in secondary metabolism significantly broadens the scope of the under-investigated PUP/2.A.7.14 family of DMT transporters. The molecular evolution of the PUP/2.A.7.14 family in angiosperms combined with the narrow substrate specificity/selectivity profiles of the few biochemically characterized homologs support the hypothesis that other PUP-like transporters are likely involved in other highly specialized metabolic pathways. Characterization of additional PUP/2.A.7.14 homologs will be required to determine if this hypothesis is well-supported or not. Toward that end, there are many PUP/2.A.7.14 transporters that await investigation.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Alex Weisberg provided scripting to reformat tree taxa labels and Sarah Kashanian assigned TC codes to tree taxa labels. Dr. Angus Murphy provided helpful comments during the early drafting of the manuscript. I am also grateful for the helpful suggestions of the reviewers. This work was supported by a USDA award administered through the Virginia Agriculture and Experiment Station Biodesign and Bioprocessing Research Center to John G. Jelesko.
References
Baldwin, I. T. (1988). Damage-induced alkaloids in tobacco: pot-bound plants are not inducible. J. Chem. Ecol. 14, 1113–1120.
Baldwin, I. T., and Callahan, P. (1993). Autotoxicity and chemical defense: nicotine accumulation and carbon gain in solanaceous plants. Oecologia 94, 534–541.
Baldwin, I. T., and Karb, M. J. (1995). Plasticity in allocation of nicotine to reproductive parts in Nicotiana attenuata. J. Chem. Ecol. 21, 897–909.
Baldwin, I. T., and Ohnmeiss, T. E. (1994). Swords into plowshares? Nicotiana sylvestris does not use nicotine as a nitrogen source under nitrogen-limited growth. Oecologia 98, 385–392.
Burkle, L., Cedzich, A., Dopke, C., Stransky, H., Okumoto, S., Gillissen, B., Kuhn, C., and Frommer, W. B. (2003). Transport of cytokinins mediated by purine transporters of the PUP family expressed in phloem, hydathodes, and pollen of Arabidopsis. Plant J. 34, 13–26.
Darriba, D., Taboada, G. L., Doallo, R., and Posada, D. (2011). ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165.
Dawson, R. F. (1942a). Accumulation of nicotine in reciprocal grafts of tomato and tobacco. Am. J. Bot. 29, 66–71.
Devereux, J., Haeberli, P., and Smithies, O. (1984). A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 12, 387–395.
Force, A., Lynch, M., Pickett, F. B., Amores, A., Yan, Y. L., and Postlethwait, J. (1999). Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545.
Gillissen, B., Burkle, L., Andre, B., Kuhn, C., Rentsch, D., Brandl, B., and Frommer, W. B. (2000). A new family of high-affinity transporters for adenine, cytosine, and purine derivatives in Arabidopsis. Plant Cell 12, 291–300.
Hildreth, S. B., Gehman, E. A., Yang, H., Lu, R.-H., Ritesh, K. C., Harich, K. C., Yu, S., Lin, J., Sandoe, J. L., Okumoto, S., Murphy, A. S., and Jelesko, J. G. (2011). Tobacco nicotine uptake permease (NUP1) affects alkaloid metabolism. Proc. Natl. Acad. Sci. U.S.A. 108, 18179–18184.
Jack, D. L., Yang, N. M., and H. Saier, M. (2001). The drug/metabolite transporter superfamily. Eur. J. Biochem. 268, 3620–3639.
Katoh, K., Kuma, K., Toh, H., and Miyata, T. (2005). MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518.
Kidd, S. K., Melillo, A. A., Lu, R. H., Reed, D. G., Kuno, N., Uchida, K., Furuya, M., and Jelesko, J. G. (2006). The A and B loci in tobacco regulate a network of stress response genes, few of which are associated with nicotine biosynthesis. Plant Mol. Biol. 60, 699–716.
Ohnmeiss, T. E., and Baldwin, I. T. (1994). The allometry of nitrogen allocation to growth and an inducible defense under nitrogen-limited growth. Ecology 75, 995–1002.
Rambaut, A. (2009). FigTree, ver. 1.2.2. Available at: http://tree.bio.ed.ac.uk/software/figtree
Rambaut, A., and Drummond, A. J. (2003). Tracer, ver. 1.5. Available at: http://tree.bio.ed.ac.uk/software/tracer
Ronquist, F., and Huelsenbeck, J. P. (2003). MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574.
Saier, M. H. Jr., Yen, M. R., Noto, K., Tamang, D. G., and Elkan, C. (2009). The Transporter Classification Database: recent advances. Nucleic Acids Res. 37, D274–D278.
Shoji, T., Inai, K., Yazaki, Y., Sato, Y., Takase, H., Shitan, N., Yazaki, K., Goto, Y., Toyooka, K., Matsuoka, K., and Hashimoto, T. (2009). Multidrug and toxic compound extrusion-type transporters implicated in vacuolar sequestration of nicotine in tobacco roots. Plant Physiol. 149, 708–718.
Solt, M. L. (1957). Nicotine production and growth of excised tobacco root cultures. Plant Physiol. 32, 480–484.
Keywords: adenine, alkaloid, evolution, nicotine, transport
Citation: Jelesko JG (2012) An expanding role for purine uptake permease-like transporters in plant secondary metabolism. Front. Plant Sci. 3:78. doi:10.3389/fpls.2012.00078
Received: 23 December 2011; Accepted: 09 April 2012;
Published online: 10 May 2012.
Edited by:
Heven Sze, University of Maryland, USAReviewed by:
Roland Krause, Max Planck Institute of Molecular Cell Biology and Genetics, GermanyMarcelo Desimone, Instituto Multidisciplinario de Biología Vegetal, Argentina
Copyright: © 2012 Jelesko. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: John G. Jelesko, Department of Plant Pathology, Physiology, and Weed Science, Virginia Tech, 548 Latham Hall, AgQuad Drive, Blacksburg, VA 24061-0390, USA. e-mail:amVsZXNrb0B2dC5lZHU=
