- 1Institute of Biological, Environmental and Rural Science, Aberystwyth University, Aberystwyth, UK
- 2Department of Molecular Biology and Genetics, Aarhus University, Slagelse, Denmark
- 3Department of Plant Sciences, University of Oxford, Oxford, UK
Nitric oxide (NO) influences many aspects of plant development and responses to stress.The concentration of NO can play an important role in influencing its action (for example, in stomatal regulation; Wilson et al., 2009) so that the mechanisms through which NO content is modulated must be an important facet of NO research. Whilst NO generation mechanisms are clearly important, NO removal is of equal relevance, especially as plants will be continually exposed to NOx (NO + NO2) gases derived from soil microbial activity (Mur et al., 2013). Establishing and regulating a poise between NO generation, NO fumigation from external sources and NO scavenging, which also needs to be flexible enough to change in response to a variety of physiological cues, is an under-considered aspect of plant NO biology.
How are no Generation Mechanisms Integrated and Regulated?
Initially, many sought to find an equivalent to the mammalian Nitric Oxide Synthase (NOS) enzymes in plants. NOS is a cytochrome P450-like enzyme which oxidizes arginine to citrulline to generate NO (Gorren and Mayer, 2007). However, the existence of this enzyme in higher plants is still debatable (Frohlich and Durner, 2011) and is mainly based on pharmacological evidence and assays for NOS-like enzyme activity reviewed by Frohlich and Durner (2011) and Mur et al. (2013). In this context, it is also relevant that arginase mutants in Arabidopsis also displayed increased NO levels (Flores et al., 2008). However, NOS-activity has not been linked to a given gene. Resolution of this conundrum may derive from the observation that polyamine leads to NO production from Arabidopsis roots (Tun et al., 2006). As L-arginine is a precursor to polyamine biosynthesis, any perturbance of L-arginine metabolism would affect any polyamine-mediated NO generation mechanism and would explain the effects of NOS-inhibitors without needing NOS. Such a mechanism would be easily linked to the most well-characterized plant NO mechanism which is based on nitrate reductase (NR). NR acts by reducing nitrite to NO with NAD(P)H acting as an electron donor. NR-generated NO has been shown to regulate floral development, root formation, stomatal opening, and responses to biotic and abiotic stresses [reviewed in Mur et al. (2013)]. NR has high affinity for nitrate but switches to its lower affinity substrate nitrite to produce NO (Planchet et al., 2005). Therefore, NR requires high nitrite concentrations to produce NO; and a low pH is also required. Considering both NR and NOS-like NO generation mechanisms together it is possible to suggest some regulatory nodes. Thus, NO generation can be regulated at the level of NO3− uptake via nitrate channels, post-translational modification of NR activity (Mur et al., 2013), influencing NO2 availability, pH and the expression and/or activity of any of the amino acid and polyamine biosynthetic enzymes. These potential regulatory mechanisms need to be systematically assessed.
Interestingly, NO3− also plays a central role in anoxic/hypoxic NO generation. Under hypoxia, the resulting energy crisis leads to a decrease in pH which inhibits plasidal NiR, leading to NO3− accumulation and NO production (Ferrari and Varner, 1971). NADH-dependent NO3− reduction occurs at the mitochondrial inner membrane, via cytochrome c oxidase and/or reductase and possibly by alternative oxidase (AOX) leading to the production of NO and ATP (Stoimenova et al., 2007). NO production via this mechanism occurs below 1% oxygen with a Ki value of 0.05% (0.6 μ M) (Gupta and Igamberdiev, 2011). Again NO3− and now also NADH are limiting factors and represent possible important regulatory steps and could be the mechanism through which nitrite is transported to mitochondria which is currently not known.
Regulating the availability of NO3− also seems to be important in other less well-characterized NO generation mechanisms. NO may be generated in the peroxisome by a xanthine oxidoreductase (XOR) which can reduce NO3− to NO (Del Rio et al., 2004). NO is also generated by a plasma membrane nitrite:NO reductase (NiNOR) where NO3− is supplied by an apoplasmic, plasma membrane-bound NR.
Balancing the Equation: Mechanisms of no Removal
In planta NO content must represent the net of rates of production minus scavenging. These scavenging mechanisms must be highly efficient in order to maintain appropriate NO poise in crop species where the extensive use of nitrogen-fertilizers can result in external fumigation at rates that may be in excess of 20 nmol m−2 h−1 (Voldner et al., 1986; Benkovitz et al., 1996). Various means to reduce NO content have recently emerged; perhaps the most important being nonsymbiotic forms of hemoglobin (Hb). Oxygenated ferrous (Fe2+) Hb converts NO to NO3− and becomes MetHb (ferric, Fe3+) (metamoglobin) form which is then reduced to oxygenated ferrous (Fe2+) by metamoglobin reductase (MetHb) (Hill, 2012). NO oxidation by Hb plays an important role in NO accumulation during stress (Hebelstrup et al., 2012; Mur et al., 2012) thus the regulation of Hb expression is vitally important to understanding how NO poise is established (Mur et al., 2013). It is highly relevant that NO3− induces Hb (Wang et al., 2000) again showing how NO3− regulates NO content, on this occasion by influencing NO scavenging.
Other enzymes through which NO effects are modulated include S-Nitrosoglutathione Reductase (GSNOR). NO reacts with glutathione GSH and forms S-nitrosoglutathione (GSNO), which represents a significant reservoir for NO (Sakamoto et al., 2002). GSNO levels are controlled by GSNOR with converts GSNO into glutathione and sulphinamide using NADH as electron donor. Thus, GSNOR represents a means through which NO signaling may be suppressed as has been demonstrated using GSNOR mutants (Feechan et al., 2005). Additionally, under aerobic conditions mitochondria are highly efficient NO scavengers (87% of supplied NO −180 pmol) (Gupta et al., 2005). Mechanistically, this has been linked to AOX via leaking electron flow from the electron transport chain to terminal electron acceptor oxygen or nitrite in the cytochrome pathway (Cvetkovska and Vanlerberghe, 2012).
This opinion piece seeks to highlight some key questions regarding how in planta NO content is regulated (Figure 1). In developing these questions we have highlighted the role of NO3−. We suggest that understanding the regulation of NO3− uptake, assimilation and processing into a myriad of biosynthetic pathways will be central to understanding how in planta NO content is established.
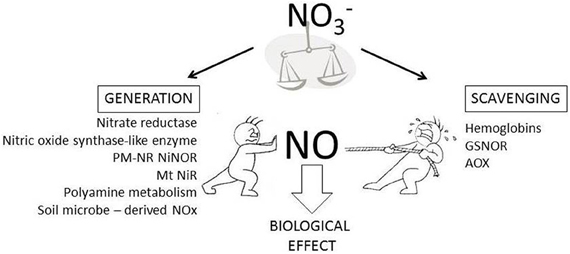
Figure 1. In planta sources of NO generation and scavenging. In planta NO content reflects the net effect of NO generation (shown as “push” in the Figure) and scavenging (shown as “pull” in the Figure) mechanisms. NO generation can involve the listed pathway (PM-NR NiNOR, plasma membrane associated nitrate reductase coupled to nitrite reductase; Mt NiR, mitochondrial nitrite reductase). The likely role of NO3− in regulating in planta NO content is highlighted.
Acknowledgments
Kapuganti J. Gupta was supported by Marie Curie Intra European Fellowship for Career Development from EU FP 7.
References
Benkovitz, C. M., Scholtz, M. T., Pacyna, J., Tarrason, L., Dignon, J., Voldner, E. C., et al. (1996). Global gridded inventories of anthropogenic emissions of sulfur and nitrogen. J. Geophys. Res. Atmos. 101, 29239–29253. doi: 10.1029/96JD00126
Cvetkovska, M., and Vanlerberghe, G. C. (2012). Alternative oxidase modulates leaf mitochondrial concentrations of superoxide and nitric oxide. New Phytol. 195, 32–39. doi: 10.1111/j.1469-8137.2012.04166.x
Del Rio, L. A., Corpas, F. J., and Barroso, J. B. (2004). Nitric oxide and nitric oxide synthase activity in plants. Phytochemistry 65, 783–792. doi: 10.1016/j.phytochem.2004.02.001
Feechan, A., Kwon, E., Yun, B. W., Wang, Y., Pallas, J. A., and Loake, G. J. (2005). A central role for S-nitrosothiols in plant disease resistance. Proc. Natl. Acad. Sci. U.S.A. 102, 8054–8059. doi: 10.1073/pnas.0501456102
Ferrari, T. E., and Varner, J. E. (1971). Intact tissue assay for nitrite reductase in barley aleurone layers. Plant Physiol. 47, 790–794. doi: 10.1104/pp.47.6.790
Flores, T., Todd, C. D., Tovar-Mendez, A., Dhanoa, P. K., Correa-Aragunde, N., Hoyos, M. E., et al. (2008). Arginase-negative mutants of Arabidopsis exhibit increased nitric oxide signaling in root development. Plant Physiol. 147, 1936–1946. doi: 10.1104/pp.108.121459
Frohlich, A., and Durner, J. (2011). The hunt for plant nitric oxide synthase (NOS): is one really needed? Plant Sci. 181, 401–404.
Gorren, A. C. F., and Mayer, B. (2007). Nitric-oxide synthase: a cytochrome P450 family foster child. Biochim. Biophys. Acta 1770, 432–445. doi: 10.1016/j.bbagen.2006.08.019
Gupta, K. J., and Igamberdiev, A. U. (2011). The anoxic plant mitochondrion as a nitrite: NO reductase. Mitochondrion 11, 537–543. doi: 10.1016/j.mito.2011.03.005
Gupta, K. J., Stoimenova, M., and Kaiser, W. M. (2005). In higher plants, only root mitochondria, but not leaf mitochondria reduce nitrite to NO, in vitro and in situ. J. Exp. Bot. 56, 2601–2609. doi: 10.1093/jxb/eri252
Hebelstrup, K. H., van Zanten, M., Mandon, J., Voesenek, L. A. C. J., Harren, F. J. M., Cristescu, S. M., et al. (2012). Haemoglobin modulates NO emission and hyponasty under hypoxia-related stress in Arabidopsis thaliana. J. Exp. Bot. 63, 5581–5591. doi: 10.1093/jxb/ers210
Hill, R. D. (2012). Non-symbiotic haemoglobins-What's happening beyond nitric oxide scavenging? AoB Plants 2012:pls004. doi: 10.1093/aobpla/pls004
Mur, L. A., Mandon, J., Persijn, S., Cristescu, S. M., Moshkov, I. E., Novikova, G. V., et al. (2013). Nitric oxide in plants: an assessment of the current state of knowledge. AoB Plants 5:pls052. doi: 10.1093/aobpla/pls052
Mur, L. A. J., Sivakumaran, A., Mandon, J., Cristescu, S. M., Harren, F. J. M., and Hebelstrup, K. H. (2012). Haemoglobin modulates salicylate and jasmonate/ethylene-mediated resistance mechanisms against pathogens. J. Exp. Bot. 63, 4375–4387. doi: 10.1093/jxb/ers116
Planchet, E., Gupta, K. J., Sonoda, M., and Kaiser, W. M. (2005). Nitric oxide emission from tobacco leaves and cell suspensions: rate limiting factors and evidence for the involvement of mitochondrial electron transport. Plant J. 41, 732–743. doi: 10.1111/j.1365-313X.2005.02335.x
Sakamoto, A., Ueda, M., and Morikawa, H. (2002). Arabidopsis glutathione-dependent formaldehyde dehydrogenase is an S-nitrosoglutathione reductase. FEBS Lett. 515, 20–24. doi: 10.1016/S0014-5793(02)02414-6
Stoimenova, M., Igamberdiev, A. U., Gupta, K. J., and Hill, R. D. (2007). Nitrite-driven anaerobic ATP synthesis in barley and rice root mitochondria. Planta 226, 465–474. doi: 10.1007/s00425-007-0496-0
Tun, N. N., Santa-Catarina, C., Begum, T., Silveira, V., Handro, W., Floh, E. I., et al. (2006). Polyamines induce rapid biosynthesis of nitric oxide (NO) in Arabidopsis thaliana seedlings. Plant Cell Physiol. 47, 346–354. doi: 10.1093/pcp/pci252
Voldner, E. C., Barrie, L. A., and Sirois, A. (1986). A literature-review of dry deposition of oxides of sulfur and nitrogen with emphasis on long-range transport modeling in North-America. Atmos. Environ. 20, 2101–2123. doi: 10.1016/0004-6981(86)90305-7
Wang, R. C., Guegler, K., LaBrie, S. T., and Crawford, N. M. (2000). Genomic analysis of a nutrient response in arabidopsis reveals diverse expression patterns and novel metabolic and potential regulatory genes induced by nitrate. Plant Cell 12, 1491–1509.
Citation: Mur LAJ, Hebelstrup KH and Gupta KJ (2013) Striking a balance: does nitrate uptake and metabolism regulate both NO generation and scavenging? Front. Plant Sci. 4:288. doi: 10.3389/fpls.2013.00288
Received: 28 June 2013; Accepted: 12 July 2013;
Published online: 30 July 2013.
Edited by:
John Hancock, University of the West of England, UKCopyright © 2013 Mur, Hebelstrup and Gupta. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence:bHVtQGFiZXIuYWMudWs=;SmFnYWRpcy5rYXB1Z2FudGlAcGxhbnRzLm94LmFjLnVr


