- 1State Key Laboratory of Plant Cell and Chromosome Engineering, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, China
- 2Graduate University of Chinese Academy of Sciences, Beijing, China
Plants and animals have evolved intracellular nucleotide-binding domain and leucine-rich repeat-containing immune receptors (NLRs) to perceive non-self and trigger immune responses. Plant NLRs detect strain-specific pathogen effectors and activate immune signaling leading to extensive transcriptional reprogramming and termination of pathogen infection. Here we review the recent findings in barley MLA immune receptor mediated immune responses against the barley powdery mildew fungus. We focus on nucleocytoplasmic partitioning of immune receptor, bifurcation of immune signaling, transcriptional repression and derepression connecting receptor activation to immune responses. We also discuss similar findings from other plant NLRs where appropriate.
Introduction
Plants have evolved two major classes of immune receptors to detect non-self and defend themselves against pathogen infection (Jones and Dangl, 2006). The surface resident pattern recognition receptors (PRRs) mainly recognize conserved microbe-associated molecular patterns (MAMPs) while the intracellular nucleotide-binding and leucine-rich repeat receptors (NLRs) perceive strain-specific pathogen effectors that are delivered inside host cells (Zipfel, 2009; Dodds and Rathjen, 2010). Both PRR and NLR mediated cellular defense responses share an overlapping signaling network (Tsuda et al., 2009; Tsuda and Katagiri, 2010) but differ quantitatively and kinetically in nature (Tao et al., 2003; Caldo et al., 2004), nevertheless, NLR-triggered immunity is usually associated with rapid and localized host cell-death, termed hypersensitive reaction (HR), at the attempted pathogen infection sites (Shen and Schulze-Lefert, 2007; Boller and Felix, 2009; Maekawa et al., 2011b).
Plant NLRs are typically modular-structured, consisting of a central nucleotide-binding domain, C-terminal leucine-rich repeats, and a diversified N-terminal domain of either coiled-coil (CC) or TOLL/interleukin-1 receptor (TIR) subtype. The NLR receptors act as molecular switches to regulate immune responses by switching from an inactive form to an active form upon recognition of pathogen effector(s) and induced conformational changes from ADP- to ATP-bound state (Collier and Moffett, 2009; Lukasik and Takken, 2009; Takken and Goverse, 2012). The N-terminal CC or TIR domain may act as a signaling module for triggering host cell death (Swiderski et al., 2009; Krasileva et al., 2010; Bernoux et al., 2011; Collier et al., 2011; Maekawa et al., 2011a; Bai et al., 2012).
The barley MLA locus is highly polymorphic encoding a large number of allelic CC-subtype NLRs, each conferring isolate-specific disease resistance against the barley powdery mildew fungus, Blumeria graminis f. sp. hordei (Bgh; Seeholzer et al., 2010). The N-terminal CC domains of MLA are highly conserved in sequence (Seeholzer et al., 2010; Jordan et al., 2011), containing an EDVID motif shared with many other CC-subtype NLRs (Collier and Moffett, 2009). The more diversified C-terminal LRR region of MLA was shown to confer recognition specificity (Shen et al., 2003). Here we summarize our recent progresses towards understanding MLA-triggered immune signaling, emphasizing on receptor partitioning, signaling bifurcation, interacting transcription factors (TFs) linking receptor activation to defense response regulations. We also touch upon analogies in other plant NLR-mediated immune signaling pathways.
Dynamic Nucleocytoplasmic Partitioning of MLA Immune Receptors
The barley intracellular MLA immune receptor has been shown to distribute between the nucleus and the cytoplasm (Shen et al., 2007). Using stable transgenic barley lines expressing a single copy of MLA1-HA fusion under the control of native 5′ regulatory sequences, fractionation experiments revealed that the majority of MLA1 is located in the cytoplasm and a small fraction (~5%) resides in the nucleus; and interestingly, its nuclear pool is increased upon inoculation of an incompatible Bgh isolate (Shen et al., 2007). Transient expression of a YFP-tagged natural MLA variant, MLA10, revealed that a MLA10-YFP fusion resides in both compartments in barley leaf epidermal cells (Shen et al., 2007; Bai et al., 2012). A mutation in the P-loop motif of MLA10 resulted in apparent increase of overall YFP signal intensity of MLA10-YFP in both compartments for unknown reasons (Bai et al., 2012), excluding the possibility that the P-loop motif of MLA10 is involved in nucleocytoplasmic partitioning. Similar nucleocytoplasmic distribution of the MLA10-YFP fusion was observed in the heterologous N. benthamiana system upon Agrobacterium-mediated transient expression and confocal imaging (Bai et al., 2012). Interestingly, similar nucleocytoplasmic partitioning of MLA1 was observed in Arabidopsis using a transgenic lines expressing MLA1-HA in a triple mutant background (Maekawa et al., 2012). Whether MLA immune receptors are regulated by conserved or distinct import/export machinery in these two plant species is currently unknown.
In recent years several plant NLR immune receptors have been shown to distribute between cytoplasm and nucleus (Deslandes et al., 2002; Burch-Smith et al., 2007; Shen et al., 2007; Wirthmueller et al., 2007; Cheng et al., 2009; Slootweg et al., 2010; Tameling et al., 2010; Hoser et al., 2013; Inoue et al., 2013; Ma et al., 2013). Some of them possess a canonical or predicted nuclear localization signal (NLS), for example the Arabidopsis RPS4/RRS1-R receptor pair and snc1, tobacco N and tomato I-2 resistance protein; while others, like MLA and potato Rx, do not harbor any discernible NLS signal. In this regard, it remains to be shown how the nucleocytoplasmic partitioning is regulated for most of these NLRs (Meier and Somers, 2011; Wirthmueller et al., 2013).
Bifurcation of MLA-Triggered Cell Death and Disease Resistance Signaling
Forced localization of MLA10 to either the cytoplasm or the nucleus, by adding either nuclear export signal (NES) or NLS to its C-terminus (CT), revealed distinct receptor activities in signaling (Shen et al., 2007; Bai et al., 2012). The nuclear pool of MLA10 is essential for powdery mildew disease resistance as transient expression of the MLA10-YFP-NES fusion, that is depleted from the nucleus, fails to restrict the growth of an avirulent Bgh isolate (Shen et al., 2007). Further, expression and enforced nuclear localization of the MLA10-NLS fusion revealed that the MLA nuclear pool alone is sufficient to confer disease resistance against Bgh in barley (Bai et al., 2012). Unexpectedly, upon transient expression in the heterologous N. benthamiana leaves, the MLA10-NES fusion was able to trigger markedly enhanced cell death signaling, whereas MLA10-NLS was unable to induce cell death (Bai et al., 2012). Although MLA10-triggered cell death in the heterologous N. benthamiana system is effector-independent, combined with functional analysis in barley these data strongly suggest a model for bifurcation of MLA signaling, in which MLA triggers cell death signaling in the cytoplasm but mediates disease resistance signaling in the nucleus, and these signaling activities of MLA can be uncoupled in a cell compartment-dependent manner (Figure 1A).
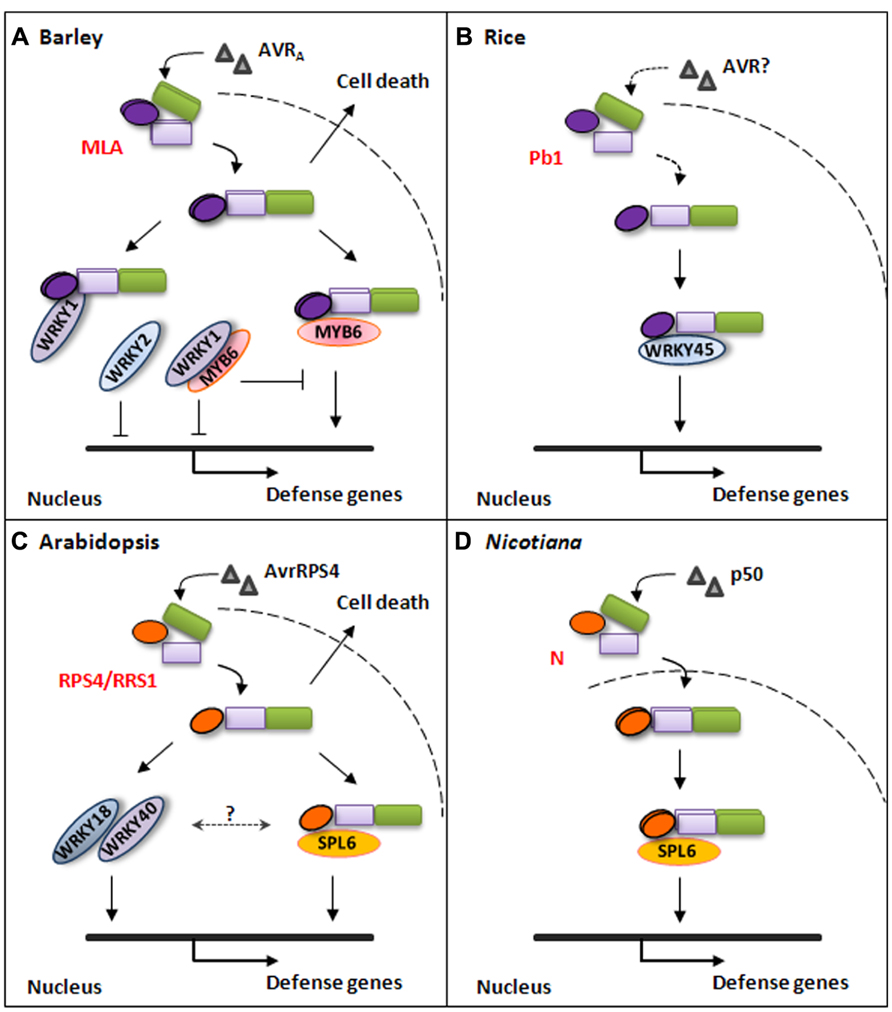
FIGURE 1. Simplified models for plant NLR-triggered immune signaling pathways. (A) Barley MLA immune receptor recognizes cognate AVRA effector from B. graminis fungal pathogen and triggers disease resistance signaling in the nucleus or cell-death signaling in the cytoplasm. The activated MLA interacts with WRKY1 through its N-terminal CC domain to release MYB6 and by itself directly interacts with MYB6 to initiate defense gene expression. Barley WRKY1 and WRKY2 are repressors of defense responses. (B) Rice atypical NLR Pb1 interacts with WRKY45 to mediate immune responses against the rice blast fungal pathogen. The Pb1-WRKY45 association can prevent the TF from being degraded by the ubiquitin/proteasome system. (C) Arabidopsis NLR pair RPS4/RRS1 mediate disease resistance signaling against PstDC3000(avrRPS4) through direct interaction with At-SPL6 or through WRKY18 and WRKY40 in the nucleus. RPS4 can also trigger cell-death signaling in the cytoplasm. (D) Nicotiana N immune receptor specifically recognizes a 50KD helicase domain (p50) from Tobacco mosaic virus (TMV) in the cytoplasm and activated N associates with SPL6 within distinct nuclear compartments to mediate immune responses against TMV.
Signaling bifurcation was also shown for a TIR-type immune receptor, the Arabidopsis RPS4 (Heidrich et al., 2011), which recognizes the type III effector AvrRps4 secreted by Pseudomonas syringae (Gassmann et al., 1999) and triggers EDS1-dependent transcriptional reprogramming and disease resistance (García et al., 2010; Heidrich et al., 2011). RPS4 was detected in association with EDS1 in complexes in Arabidopsis or N. benthamiana upon coexpression (Bhattacharjee et al., 2011; Heidrich et al., 2011). AVR effector-dependent activation of RPS4 in Arabidopsis nuclei restricted P. syringae growth without inducing cell death, however, it triggered weak cell death if the cognate AVR was forced to localize in the cytoplasm (Heidrich et al., 2011). It was proposed that nuclear or cytoplasmic RPS4-EDS1 pools specify distinct subcellular defense signaling branches, and that coordinated action of both defense signals is required for full defense responses (García et al., 2010; Heidrich et al., 2011, Heidrich et al., 2012; Figure 1C).
Several recent reports have shown uncoupling of host cell death from disease resistance for both TIR- and CC-subtype NLR immune receptors (Coll et al., 2010; Heidrich et al., 2011; Bai et al., 2012; Sohn et al., 2012; Gao et al., 2013), together these add unambiguous evidence to support a model that for some NLRs HR-cell death and disease resistance are distinct but interconnected subcellular functions.
MLA CC Domain as a Platform for Interacting and Signaling
MLA fragments harboring the N-terminal CC domain or other domains have been used for identifying MLA interactors in yeast two-hybrid screenings. The CC domain containing fragments identified the most MLA interactors, and interestingly, almost all of them interacted with the CC domain but not with the MLA full-length protein in further analysis in yeast and in planta upon transient coexpression (Shen et al., 2007; Chang et al., 2013; Chang and Shen, unpublished data), suggesting that the MLA CC domain alone can serve as a platform for interacting with or docking to signaling partners post MLA activation.
A crystal structure of MLA10 CC reveals that this domain can form a homodimer and this dimer configuration is shown to be critical for MLA activity (Maekawa et al., 2011a). In the heterologous N. benthamiana system, a role of the MLA10 CC domain in cell death signaling has been established by Agrobacterium-mediated transient expression (Maekawa et al., 2011a; Bai et al., 2012). MLA10 full-length protein triggered cell death requires an intact P-loop motif; and mutations in the MHD motif render MLA10 autoactive, triggering cell death in N. benthamiana and barley (Bai et al., 2012), together these findings point to a likely scenario in which MLA activation involves conformational changes driven by ATP-binding and hydrolysis cycles and releasing of the N-terminal CC domain, which adopts a homodimer conformation that could serve as a platform for signaling initiation (Maekawa et al., 2011a; Takken and Goverse, 2012). Since the MLA cytoplasmic pool alone is sufficient to trigger cell death we envisage the death signaling might first initiate from the cytoplasm and then transduced by as yet unknown signaling components.
Transcription Factors as Direct Downstream Component in MLA-Activated Signaling
Earlier studies thoroughly characterized the association between MLA and two barley WRKY TFs, WRKY1 and WRKY2 (Shen et al., 2007). WRKY1 and WRKY2 interact with the MLA CC domain but not with the full-length MLA protein in yeast, importantly, an AVRA effector-dependent association of full-length MLA10 with WRKY2 was detected in the nucleus of barley cells using fluorescence life time imaging-fluorescence resonance energy transfer (FLIM-FRET) analysis (Shen et al., 2007). Barley WRKY1 and WRKY2 were demonstrated to act as repressors of basal immunity against the Bgh fungus in barley. It was hypothesized that MLA immune receptors target these WRKY repressors to derepress PAMP-triggered immunity thus potentiating defense responses (Shen et al., 2007; Shen and Schulze-Lefert, 2007).
Recently, we reported the identification of barley MYB6 as another MLA interactor (Chang et al., 2013). MYB6 interacts with the CC domain of MLA receptors, MLA1, MLA6 and MLA10, and interestingly MYB6 appears to specifically interact with the homodimeric form of the functional CC domain (Chang et al., 2013). Since the full-length MLA protein was unable to interact with MYB6 we interpret the association of MLA CC with MYB6 as event post MLA receptor activation, somewhat analogous to the interaction between MLA and WRKY1/2. Nevertheless, contrary to the WRKY1/2 repressor, MYB6 acts as a positive regulator in basal and MLA-triggered disease resistance against the powdery mildew fungus, demonstrated by virus-induced gene silencing (VIGS) and functional gene expression analysis in barley (Chang et al., 2013).
Since WRKY1/2 and MYB6 interact with the MLA CC domain, the potential interaction between WRKY1/2 and MYB6 was tested. Significantly, WRKY1, but not WRKY2, interacts with MYB6 and interferes with MYB6 DNA binding activity (Chang et al., 2013). It is noteworthy that barley WRKY1 and WRKY2 share the same domain structure and 72% sequence similarity, and their Arabidopsis homologues, At-WRKY18, At-WRKY40 and At-WRKY60, act redundantly as negative regulators in disease resistance against the bacterial pathogen Pseudomonas syringae pv. tomato DC3000 (PstDC3000) and the Arabidopsis-infecting powdery mildew fungus Golovinomyces orontii (Xu et al., 2006; Shen et al., 2007; Pandey et al., 2010). However, surprisingly, it was recently reported that At-WRKY18 and At-WRKY40 are specifically required for mediating disease resistance against PstDC3000 expressing effector AvrRPS4, shown by the specific susceptibility phenotype of the wrky18 wrky40 mutant line infected with PstDC3000(avrRPS4) but not with other tested PstDC3000 strains (Schön et al., 2013). These findings indicate that WRKY18 and WRKY40 may function redundantly as positive regulators downstream of the RPS4/RRS1 pair, or alternatively that these WRKYs may be targeted and modified by AvrRPS4 which can be perceived by RPS4/RRS1, although the direct physical interaction between RPS4 and WRKY18 or WRKY40 was not detected in the presence or absence of the AVR effector (Schön et al., 2013; Figure 1C).
Several other TFs have recently been reported to function in NLR-mediated immune signaling (Inoue et al., 2013; Padmanabhan et al., 2013; Figure 1). The Nicotiana SPL6 TF was demonstrated to interact with the N immune receptor in subnuclear bodies once immune signaling is activated and SPL6 functions as a positive regulator in N-mediated immunity against Tobacco mosaic virus in Nicotiana plants (Figure 1D); Interestingly, like At-WRKY18 and At-WRKY40, the SPL6 paralog in Arabidopsis is also specifically required for RPS4-triggered disease resistance against PstDC3000 (avrRPS4; Padmanabhan et al., 2013; Figure 1C). The rice WRKY45 was demonstrated to interact with Pb1, an CC-NB-LRR protein conferring panicle blast resistance in rice, and this interaction prevents ubiquitin/proteasome-mediated degradation of WRKY45, which is believed to be involved in Pb1-triggered blast resistance (Hayashi et al., 2010; Inoue et al., 2013; Figure 1B).
Repressing and Derepressing: Transcriptional Regulations in MLA-Triggered Immune Signaling
The R2R3-type MYB TF family members have undergone expansion in different plant lineages and are involved in regulating diverse biological processes (Stracke et al., 2001; Dubos et al., 2010; Feller et al., 2011; Raffaele and Rivas, 2013). One of the best characterized MYB TF is Arabidopsis At-MYB30 that plays a critical role in executing hypersensitive cell death in defense response to the bacterial pathogen Xanthomonas (Vailleau et al., 2002; Raffaele et al., 2008). Significantly, the transcriptional activity of At-MYB30 and resistance function is negatively regulated not only by the host protein AtsPLA2-α through physical association in the nucleus (Froidure et al., 2010), but also by the Xanthomonas Type III effector XopD by relocalizing it to nuclear foci (Canonne et al., 2011).
Barley MYB6 is also a R2R3-type MYB TF that binds to the cognate cis-element MBS I and acts as a transcriptional activator to regulate gene expression (Chang et al., 2013). MYB6 activity in DNA-binding was evaluated in the presence of WRKY1 or MLA CC in electrophoretic mobility shift assay (EMSA) or Arabidopsis protoplast transfection assay. Interestingly, WRKY1 could suppress MYB6 DNA-binding activity, whereas the MLA10 CC domain markedly stimulated this activity, suggesting that MYB6 activity is antagonistically regulated by WRKY1 and MLA CC domain (Chang et al., 2013).
The tripartite interaction among WRKY1, MYB6 and MLA were dissected in details using yeast three-hybrid, in planta and in vitro protein interaction assays. It was demonstrated that the WRKY1-MYB6 association can be abrogated by the MLA10 CC domain in a WRKY1 CT-dependent manner, and subsequently MLA10 CC forms a complex with MYB6 in the nucleus. Importantly, MLA10 CC and an autoactive MLA10 full-length variant with a mutation in the MHD motif can antagonize WRKY1 suppression and markedly stimulates MYB6 DNA-binding activity, thus increases MYB6-dependent gene expressions in the Arabidopsis protoplast transfection system (Chang et al., 2013).
We propose a model in which WRKY1 repressor physically sequesters barley MYB6 from binding to the promoter of downstream target genes to prevent uncontrolled cell death and defense responses; upon perception of cognate effector activated MLA interacts with WRKY1 and releases MYB6 from suppression and stimulates its binding to cognate cis-acting elements to initiate disease resistance signaling (Chang et al., 2013; Figure 1A).
Conclusions and Perspectives
Data from barley MLA and other plant NLRs discussed here underlines the importance of nucleocytoplasmic trafficking and transcriptional regulation in plant NLR-mediated immune responses. Emerging evidence indicates that parallel mechanistics of regulation exist in mammalian NLR-mediated immunity. NLRC5 was recently presented as a transcription regulator to cooperate with TFs to induce MHC class I gene expression (Meissner et al., 2010, 2012), while CIITA was previously identified as a master transcription coactivator in regulating MHC class II gene expression (Ting and Davis, 2005); both NLRs shuttle between the cytosol and nucleus.
Specific and fundamental questions remain to be addressed to fill the gaps in MLA-activated immune signaling: what are the target genes commonly and distinctively regulated by WRKY1/2 and/or MYB6? How are MLA, WRKY1/2 and MYB6 regulated at post-translational level? What are the components/pathways involved in MLA-triggered cell death signaling in the cytoplasm? How does MLA regulate distinct immune activities in the nucleus and cytoplasm?
So far only a limited numbers of NLRs were shown to trigger defense signaling through direct association with TFs, which is likely downstream of AVR effector perception (Figure 1). Nevertheless, analogous mechanistics appears to be engaged with by both CC- and TIR-subtype of NLR receptors from either monocots or dicots to coordinate defense responses against diverse pathogens, including viral, bacterial and fungal pathogens (Figure 1). It is reasonable to envisage that NLRs are partially nuclear localized or translocated into the nucleus upon activation may orchestrate defense gene expression through transcriptional regulation. We are only at the beginning to unravel the dynamics of NLR-mediated signaling in the cytoplasm and the nucleus.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank all members in Shen’s lab for their contribution and Paul Schulze-Lefert at the Max Planck Institute, Cologne and colleagues for collaboration. This work was supported by the National Basic Research Program of China (2011CB100700), and National Natural Science Foundation of China (31030007).
References
Bai, S., Liu, J., Chang, C., Zhang, L., Maekawa, T., Wang, Q., et al. (2012). Structure-function analysis of barley NLR immune receptor MLA10 reveals its cell compartment specific activity in cell death and disease resistance. PLoS Pathog. 8:e1002752. doi: 10.1371/journal.ppat.1002752
Bernoux, M., Ve, T., Williams, S., Warren, C., Hatters, D., Valkov, E., et al. (2011). Structural and functional analysis of a plant resistance protein TIR domain reveals interfaces for self-association, signaling, and autoregulation. Cell Host Microbe 9, 200–211. doi: 10.1016/j.chom.2011.02.009
Bhattacharjee, S., Halane, M. K., Kim, S. H., and Gassmann, W. (2011). Pathogen effectors target Arabidopsis EDS1 and alter its interactions with immune regulators. Science 334, 1405–1408. doi: 10.1126/science.1211592
Boller, T., and Felix, G. (2009). A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 60, 379–406. doi: 10.1146/annurev.arplant.57.032905.105346
Burch-Smith, T. M., Schiff, M., Caplan, J. L., Tsao, J., Czymmek, K., and Dinesh-Kumar, S. P. (2007). A novel role for the TIR domain in association with pathogen-derived elicitors. PLoS Biol. 5:e68. doi: 10.1371/journal.pbio.0050068
Caldo, R. A., Nettleton, D., and Wise, R. P. (2004). Interaction-dependent gene expression in MLA-specified response to barley powdery mildew. Plant Cell 16, 2514–2528. doi: 10.1105/tpc.104.023382
Canonne, J., Marino, D., Jauneau, A., Pouzet, C., Brière, C., Roby, D., et al. (2011). The Xanthomonas type III effector XopD targets the Arabidopsis transcription factor MYB30 to suppress plant defense. Plant Cell 23, 3498–3511. doi: 10.1105/tpc.111.088815
Chang, C., Yu, D., Jiao, J., Jing, S., Schulze-Lefert, P., and Shen, Q.-H. (2013). Barley MLA immune receptors directly interfere with antagonistically acting transcription factors to initiate disease resistance signaling. Plant Cell 25, 1158–1173. doi: 10.1105/tpc.113.109942
Cheng, Y. T., Germain, H., Wiermer, M., Bi, D., Xu, F., García, A. V., et al. (2009). Nuclear pore complex component MOS7/Nup88 is required for innate immunity and nuclear accumulation of defense regulators in Arabidopsis. Plant Cell 21, 2503–2516. doi: 10.1105/tpc.108.064519
Coll, N. S., Vercammen, D., Smidler, A., Clover, C., Van Breusegem, F., Dangl, J. L., et al. (2010). Arabidopsis type I metacaspases control cell death. Science 330, 1393–1397. doi: 10.1126/science.1194980
Collier, S. M., Hamel, L.-P., and Moffett, P. (2011). Cell death mediated by the N-terminal domains of a unique and highly conserved class of NB-LRR protein. Mol. Plant Microbe Interact. 24, 918–931. doi: 10.1094/MPMI-03-11-0050
Collier, S. M., and Moffett, P. (2009). NB-LRRs work a “bait and switch” on pathogens. Trends Plant Sci. 14, 521–529. doi: 10.1016/j.tplants.2009.08.001
Deslandes, L., Olivier, J., Theulières, F., Hirsch, J., Feng, D. X., Bittner-Eddy, P., et al. (2002). Resistance to Ralstonia solanacearum in Arabidopsis thaliana is conferred by the recessive RRS1-R gene, a member of a novel family of resistance genes. Proc. Natl. Acad. Sci. U.S.A. 99, 2404–2409. doi: 10.1073/pnas.032485099
Dodds, P. N., and Rathjen, J. P. (2010). Plant immunity: towards an integrated view of plant-pathogen interactions. Nat. Rev. Genet. 11, 539–548. doi: 10.1038/nrg2812
Dubos, C., Stracke, R., Grotewold, E., Weisshaar, B., Martin, C., and Lepiniec, L. (2010). MYB transcription factors in Arabidopsis. Trends Plant Sci. 15, 573–581. doi: 10.1016/j.tplants.2010.06.005
Feller, A., Machemer, K., Braun, E. L., and Grotewold, E. (2011). Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. 66, 94–116. doi: 10.1111/j.1365-313X.2010.04459.x
Froidure, S., Canonne, J., Daniel, X., Jauneau, A., Brière, C., Roby, D., et al. (2010). AtsPLA2-α nuclear relocalization by the Arabidopsis transcription factor AtMYB30 leads to repression of the plant defense response. Proc. Natl. Acad. Sci. U.S.A. 107, 15281–15286. doi: 10.1073/pnas.1009056107
Gao, X., Chen, X., Lin, W., Chen, S., Lu, D., Niu, Y., et al. (2013). Bifurcation of Arabidopsis NLR Immune signaling via Ca2+-dependent protein kinases. PLoS Pathog. 9:e1003127. doi: 10.1371/journal.ppat.1003127
García, A. V., Blanvillain-Baufumé, S., Huibers, R. P., Wiermer, M., Li, G., Gobbato, E., et al. (2010). Balanced nuclear and cytoplasmic activities of EDS1 are required for a complete plant innate immune response. PLoS Pathog. 6:e1000970. doi: 10.1371/journal.ppat.1000970
Gassmann, W., Hinsch, M. E., and Staskawicz, B. J. (1999). The Arabidopsis RPS4 bacterial-resistance gene is a member of the TIR-NBS-LRR family of disease-resistance genes. Plant J. 20, 265–277. doi: 10.1046/j.1365-313X.1999.t01-1-00600.x
Hayashi, N., Inoue, H., Kato, T., Funao, T., Shirota, M., Shimizu, T., et al. (2010). Durable panicle blast-resistance gene Pb1 encodes an atypical CC-NBS-LRR protein and was generated by acquiring a promoter through local genome duplication. Plant J. 64, 498–510. doi: 10.1111/j.1365-313X.2010.04348.x
Heidrich, K., Blanvillain-Baufumé, S., and Parker, J. E. (2012). Molecular and spatial constraints on NB-LRR receptor signaling. Curr. Opin. Plant Biol. 15, 385–391. doi: 10.1016/j.pbi.2012.03.015
Heidrich, K., Wirthmueller, L., Tasset, C., Pouzet, C., Deslandes, L., and Parker, J. E. (2011). Arabidopsis EDS1 connects pathogen effector recognition to cell compartment-specific immune responses. Science 334, 1401–1404. doi: 10.1126/science.1211641
Hoser, R., Żurczak, M., Lichocka, M., Zuzga, S., Dadlez, M., Samuel, M. A., et al. (2013). Nucleocytoplasmic partitioning of tobacco N receptor is modulated by SGT1. New Phytol. 200, 158–171. doi: 10.1111/nph.12347
Inoue, H., Hayashi, N., Matsushita, A., Xinqiong, L., Nakayama, A., Sugano, S., et al. (2013). Blast resistance of CC-NB-LRR protein Pb1 is mediated by WRKY45 through protein–protein interaction. Proc. Natl. Acad. Sci. U.S.A. 110, 9577–9582. doi: 10.1073/pnas.1222155110
Jones, J. D. G., and Dangl, J. L. (2006). The plant immune system. Nature 444, 323–329. doi: 10.1038/nature05286
Jordan, T., Seeholzer, S., Schwizer, S., Töller, A., Somssich, I. E., and Keller, B. (2011). The wheat MLA homologue TmMla1 exhibits an evolutionarily conserved function against powdery mildew in both wheat and barley. Plant J. 65, 610–621. doi: 10.1111/j.1365-313X.2010.04445.x
Krasileva, K. V., Dahlbeck, D., and Staskawicz, B. J. (2010). Activation of an Arabidopsis resistance protein is specified by the in planta association of its leucine-rich repeat domain with the cognate oomycete effector. Plant Cell 22, 2444–2458. doi: 10.1105/tpc.110.075358
Lukasik, E., and Takken, F. L. W. (2009). STANDing strong, resistance proteins instigators of plant defence. Curr. Opin. Plant Biol. 12, 427–436. doi: 10.1016/j.pbi.2009.03.001
Ma, L., Cornelissen, B. J. C., and Takken, F. L. W. (2013). A nuclear localization for Avr2 from Fusarium oxysporum is required to activate the tomato resistance protein I-2. Front. Plant Sci. 4:94. doi: 10.3389/fpls.2013.00094
Maekawa, T., Cheng, W., Spiridon, L. N., Töller, A., Lukasik, E., Saijo, Y., et al. (2011a). Coiled-coil domain-dependent homodimerization of intracellular barley immune receptors defines a minimal functional module for triggering cell death. Cell Host Microbe 9, 187–199. doi: 10.1016/j.chom.2011.02.008
Maekawa, T., Kufer, T. A., and Schulze-Lefert, P. (2011b). NLR functions in plant and animal immune systems: so far and yet so close. Nat. Immunol. 12, 817–826. doi: 10.1038/ni.2083
Maekawa, T., Kracher, B., Vernaldi, S., Ver Loren van Themaat, E., and Schulze-Lefert, P. (2012). Conservation of NLR-triggered immunity across plant lineages. Proc. Natl. Acad. Sci. U.S.A. 109, 20119–20123. doi: 10.1073/pnas.1218059109
Meier, I., and Somers, D. E. (2011). Regulation of nucleocytoplasmic trafficking in plants. Curr. Opin. Plant Biol. 14, 538–546. doi: 10.1016/j.pbi.2011.06.005
Meissner, T. B., Li, A., Biswas, A., Lee, K.-H., Liu, Y.-J., Bayir, E., et al. (2010). NLR family member NLRC5 is a transcriptional regulator of MHC class I genes. Proc. Natl. Acad. Sci. U.S.A. 107, 13794–13799. doi: 10.1073/pnas.1008684107
Meissner, T. B., Liu, Y.-J., Lee, K.-H., Li, A., Biswas, A., van Eggermond, M. C. J. A., et al. (2012). NLRC5 cooperates with the RFX transcription factor complex to induce MHC Class I gene expression. J. Immunol. 188, 4951–4958. doi: 10.4049/jimmunol.1103160
Padmanabhan, M. S., Ma, S., Burch-Smith, T. M., Czymmek, K., Huijser, P., and Dinesh-Kumar, S. P. (2013). Novel positive regulatory role for the SPL6 transcription factor in the N TIR-NB-LRR receptor-mediated plant innate immunity. PLoS Pathog. 9:e1003235. doi: 10.1371/journal.ppat.1003235
Pandey, S. P., Roccaro, M., Schön, M., Logemann, E., and Somssich, I. E. (2010). Transcriptional reprogramming regulated by WRKY18 and WRKY40 facilitates powdery mildew infection of Arabidopsis. Plant J. 64, 912–923. doi: 10.1111/j.1365-313X.2010.04387.x
Raffaele, S., and Rivas, S. (2013). Regulate and be regulated: integration of defence and other signals by the AtMYB30 transcription factor. Front. Plant Sci. 4:98. doi: 10.3389/fpls.2013.00098
Raffaele, S., Vailleau, F., Léger, A., Joubès, J., Miersch, O., Huard, C., et al. (2008). A MYB transcription factor regulates very-long-chain fatty acid biosynthesis for activation of the hypersensitive cell death response in Arabidopsis. Plant Cell 20, 752–767. doi: 10.1105/tpc.107.054858
Schön, M., Töller, A., Diezel, C., Roth, C., Westphal, L., Wiermer, M., et al. (2013). Analyses of wrky18 wrky40 plants reveal critical roles of SA/EDS1 signaling and indole-glucosinolate biosynthesis for Golovinomyces orontii resistance and a loss-of resistance towards Pseudomonas syringae pv. tomato AvrRPS4. Mol. Plant Microbe Interact. 26, 758–767. doi: 10.1094/MPMI-11-12-0265-R
Seeholzer, S., Tsuchimatsu, T., Jordan, T., Bieri, S., Pajonk, S., Yang, W., et al. (2010). Diversity at the Mla powdery mildew resistance locus from cultivated barley reveals sites of positive selection. Mol. Plant Microbe Interact. 23, 497–509. doi: 10.1094/MPMI-23-4-0497
Shen, Q. H., Saijo, Y., Mauch, S., Biskup, C., Bieri, S., Keller, B., et al. (2007). Nuclear activity of MLA immune receptors links isolate-specific and basal disease-resistance responses. Science 315, 1098–1103. doi: 10.1126/science.1136372
Shen, Q. H., and Schulze-Lefert, P. (2007). Rumble in the nuclear jungle: compartmentalization, trafficking, and nuclear action of plant immune receptors. EMBO J. 26, 4293–4301. doi: 10.1038/sj.emboj.7601854
Shen, Q. H., Zhou, F. S., Bieri, S., Haizel, T., Shirasu, K., and Schulze-Lefert, P. (2003). Recognition specificity and RAR1/SGT1 dependence in barley MLA disease resistance genes to the powdery mildew fungus. Plant Cell 15, 732–744. doi: 10.1105/tpc.009258
Slootweg, E., Roosien, J., Spiridon, L. N., Petrescu, A.-J., Tameling, W., Joosten, M., et al. (2010). Nucleocytoplasmic distribution is required for activation of resistance by the potato NB-LRR receptor Rx1 and is balanced by its functional domains. Plant Cell 22, 4195–4215. doi: 10.1105/tpc.110.077537
Sohn, K. H., Hughes, R. K., Piquerez, S. J., Jones, J. D. G., and Banfield, M. J. (2012). Distinct regions of the Pseudomonas syringae coiled-coil effector AvrRps4 are required for activation of immunity. Proc. Natl. Acad. Sci. U.S.A. 109, 16371–16376. doi: 10.1073/pnas.1212332109
Stracke, R., Werber, M., and Weisshaar, B. (2001). The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 4, 447–456. doi: 10.1016/S1369-5266(00)00199-0
Swiderski, M. R., Birker, D., and Jones, J. D. G. (2009). The TIR domain of TIR-NB-LRR resistance proteins is a signaling domain involved in cell death induction. Mol. Plant Microbe Interact. 22, 157–165. doi: 10.1094/MPMI-22-2-0157
Takken, F. L. W., and Goverse, A. (2012). How to build a pathogen detector: structural basis of NB-LRR function. Curr. Opin. Plant Biol. 15, 375–384. doi: 10.1016/j.pbi.2012.05.001
Tameling, W. I. L., Nooijen, C., Ludwig, N., Boter, M., Slootweg, E., Goverse, A., et al. (2010). RanGAP2 mediates nucleocytoplasmic partitioning of the NB-LRR immune receptor Rx in the Solanaceae, thereby dictating Rx function. Plant Cell 22, 4176–4194. doi: 10.1105/tpc.110.077461
Tao, Y., Xie, Z., Chen, W., Glazebrook, J., Chang, H.-S., Han, B., et al. (2003). Quantitative nature of Arabidopsis responses during compatible and incompatible interactions with the bacterial pathogen Pseudomonas syringae. Plant Cell 15, 317–330. doi: 10.1105/tpc.007591
Ting, J. P.-Y., and Davis, B. K. (2005). CATERPILLER: a novel gene family important in immunity, cell death, and diseases. Annu. Rev. Immunol. 23, 387–414. doi: 10.1146/annurev.immunol.23.021704.115616
Tsuda, K., and Katagiri, F. (2010). Comparing signaling mechanisms engaged in pattern-triggered and effector-triggered immunity. Curr. Opin. Plant Biol. 13, 459–465. doi: 10.1016/j.pbi.2010.04.006
Tsuda, K., Sato, M., Stoddard, T., Glazebrook, J., and Katagiri, F. (2009). Network properties of robust immunity in plants. PLoS Genet. 5:e1000772. doi: 10.1371/journal.pgen.1000772
Vailleau, F., Daniel, X., Tronchet, M., Montillet, J. L., Triantaphylidès, C., and Roby, D. (2002). A R2R3-MYB gene, AtMYB30, acts as a positive regulator of the hypersensitive cell death program in plants in response to pathogen attack. Proc. Natl. Acad. Sci. U.S.A. 99, 10179–10184. doi: 10.1073/pnas.152047199
Wirthmueller, L., Roth, C., Banfield, M. J., and Wiermer, M. (2013). Hop-on hop-off: importin-a-guided tours to the nucleus in innate immune signaling. Front. Plant Sci. 4:149. doi: 10.3389/fpls.2013.00149
Wirthmueller, L., Zhang, Y., Jones, J. D. G., and Parker, J. E. (2007). Nuclear accumulation of the Arabidopsis immune receptor RPS4 is necessary for triggering EDS1-dependent defense. Curr. Biol. 17, 2023–2029. doi: 10.1016/j.cub.2007.10.042
Xu, X., Chen, C., Fan, B., and Chen, Z. (2006). Physical and functional interactions between pathogen-induced Arabidopsis WRKY18, WRKY40, and WRKY60 transcription factors. Plant Cell 18, 1310–1326. doi: 10.1105/tpc.105.037523
Keywords: plant NLRs, MLA, barley, cell death, immune signaling, transcription factors, transcription regulation
Citation: Chang C, Zhang L and Shen Q-H (2013) Partitioning, repressing and derepressing: dynamic regulations in MLA immune receptor triggered defense signaling. Front. Plant Sci. 4:396. doi: 10.3389/fpls.2013.00396
Received: 10 June 2013; Paper pending published: 01 July 2013;
Accepted: 16 September 2013; Published online: 08 October 2013.
Edited by:
Susana Rivas, Centre National de la Recherche Scientifique, Laboratoire des Interactions Plantes-Microorganismes, FranceReviewed by:
Aska Goverse, Wageningen University, NetherlandsLennart Wirthmueller, John Innes Centre, UK
Copyright © 2013 Chang, Zhang and Shen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Qian-Hua Shen, State Key Laboratory of Plant Cell and Chromosome Engineering, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing 100101, China e-mail:cWhzaGVuQGdlbmV0aWNzLmFjLmNu
 Cheng Chang
Cheng Chang Ling Zhang
Ling Zhang Qian-Hua Shen
Qian-Hua Shen