Abstract
In plant cells the free radical nitric oxide (NO) interacts both with anti- as well as prooxidants. This review provides a short survey of the central roles of ascorbate and glutathione—the latter alone or in conjunction with S-nitrosoglutathione reductase—in controlling NO bioavailability. Other major topics include the regulation of antioxidant enzymes by NO and the interplay between NO and reactive oxygen species (ROS). Under stress conditions NO regulates antioxidant enzymes at the level of activity and gene expression, which can cause either enhancement or reduction of the cellular redox status. For instance chronic NO production during salt stress induced the antioxidant system thereby increasing salt tolerance in various plants. In contrast, rapid NO accumulation in response to strong stress stimuli was occasionally linked to inhibition of antioxidant enzymes and a subsequent rise in hydrogen peroxide levels. Moreover, during incompatible Arabidopsis thaliana-Pseudomonas syringae interactions ROS burst and cell death progression were shown to be terminated by S-nitrosylation-triggered inhibition of NADPH oxidases, further highlighting the multiple roles of NO during redox-signaling. In chemical reactions between NO and ROS reactive nitrogen species (RNS) arise with characteristics different from their precursors. Recently, peroxynitrite formed by the reaction of NO with superoxide has attracted much attention. We will describe putative functions of this molecule and other NO derivatives in plant cells. Non-symbiotic hemoglobins (nsHb) were proposed to act in NO degradation. Additionally, like other oxidases nsHb is also capable of catalyzing protein nitration through a nitrite- and hydrogen peroxide-dependent process. The physiological significance of the described findings under abiotic and biotic stress conditions will be discussed with a special emphasis on pathogen-induced programmed cell death (PCD).
Introduction
Exposure of plants to abiotic and biotic stress can cause a deregulation, over-flow or even disruption of electron transport chains (ETC) in mitochondria and chloroplasts. Under these conditions molecular oxygen (O2) acts as an electron acceptor giving rise to the accumulation of reactive oxygen species (ROS). Singlet oxygen (1O2), the hydroxyl radical (OH), the superoxide radical (O−2) and hydrogen peroxide (H2O2) are all strongly oxidizing compounds and therefore potentially harmful for cell integrity. Among them, H2O2 is the most stable ROS being formed in the reaction of 1O2 with O−2 and as a product of spontaneous dismutation of O−2 (Foyer and Noctor, 2009).
During evolution, land plants have developed sophisticated measures for controlling ROS levels amongst others by the antioxidant system or—as named after their discoverers—Foyer-Halliwell-Asada cycle (Figure 1) (Buchanan et al., 2002; Foyer and Noctor, 2009). Central elements of the system are the two redox couples ascorbate (AsA)/dehydroascorbate (DHA) and glutathione (GSH)/glutathione disulfide (GSSG). In the detoxification part of the antioxidant system superoxide dismutase (SOD) converts O−2 to O2 and H2O2. The latter then can be degraded by catalase (CAT), ascorbate peroxidase (APX) and several other enzymes (Figure 1). In the course of H2O2 degradation by APX AsA is oxidized to monodehydroascorbate (MDHA) and DHA. AsA and GSH can also directly be oxidized by ROS, although with slower kinetics. In the regeneration pathway MDHA reductase (MDHAR), DHA reductase (DHAR) and glutathione reductase (GR) recycle the antioxidants from their oxidized back to the reduced form. MDHAR and GR use NADPH as a reducing equivalent whereas DHAR uses GSH (Figure 1).
Figure 1
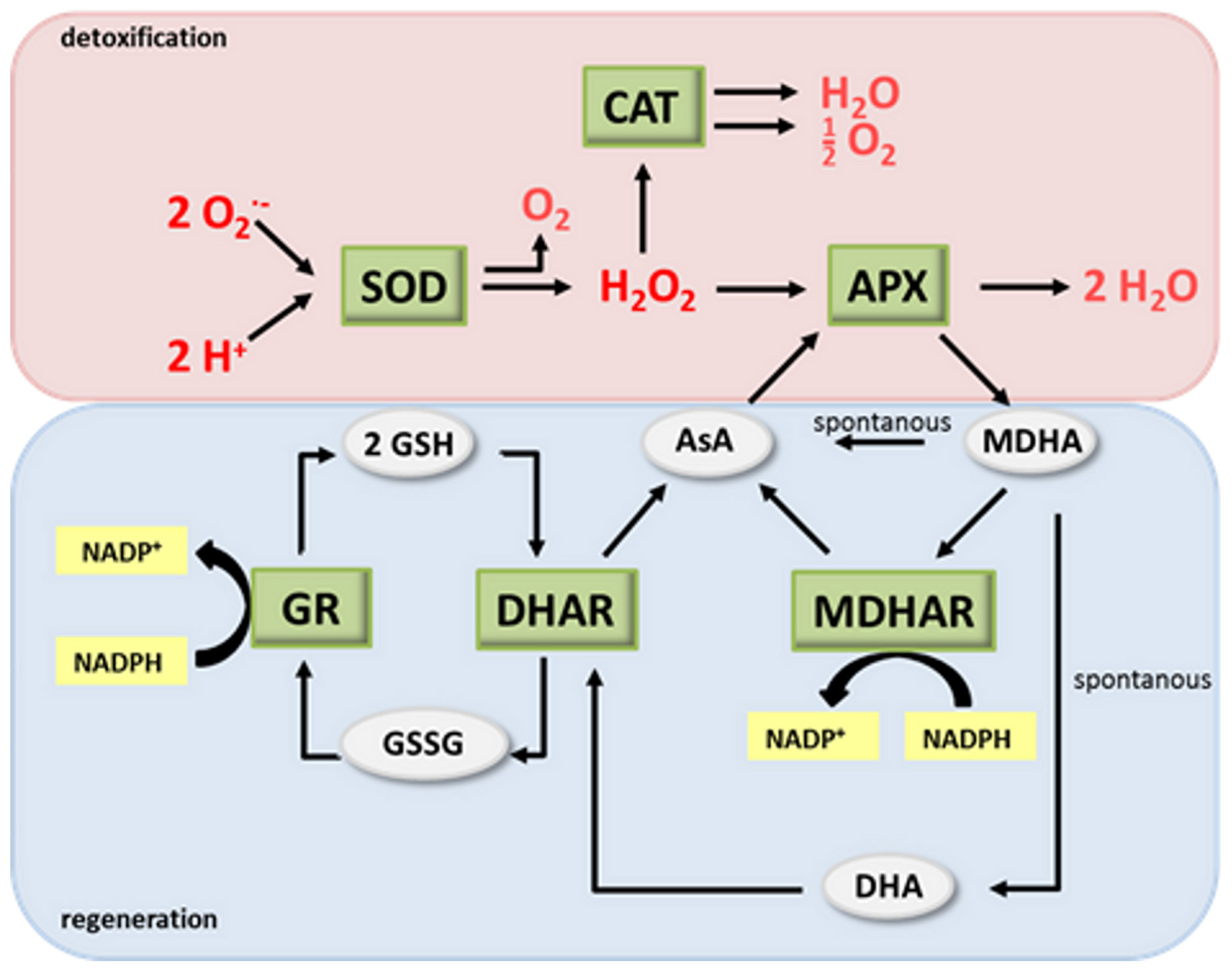
The antioxidant system. (modified after Buchanan et al., 2002). AsA, ascorbate; DHA, dehydroascorbate; SOD, superoxide dismutase; CAT, catalase; APX, ascorbate peroxidase; MDHA, monodehydroascorbate; MDHAR, MDHA reductase; DHAR, DHA reductase; GR, glutathione reductase; GSH glutathione; GSSG, glutathione disulphide.
However, apart from being toxic by-products of energy metabolism, ROS have also essential functions in primary and secondary metabolism, development, and stress responses. For instance, H2O2 acts as a signal in the regulation of stomatal closure and serves as a substrate of peroxidases during cell wall synthesis and fortification (Neill et al., 2008; O'brien et al., 2012). To date, O−2 and H2O2 are the best studied ROS, mainly because of well-established detection techniques. During signaling processes, ROS arises from the ETC but are also enzymatically produced by various peroxidases and oxidases (Foyer and Noctor, 2009; Mittler et al., 2011). Here, we will assign the term prooxidants for ROS and ROS-producing enzymes and the term antioxidants for elements of the antioxidant system. During stress signaling, the redox homeostasis of plant cells is tightly controlled. Antioxidants modulate timing and extent of ROS accumulation and additionally function as signals by their own rights. ROS levels increase either by up-regulation of prooxidant enzyme activity, (de−) regulation of electron flow or down-regulation of the antioxidant system. Redox signals are probably transduced by oxidation of proteins such as ROS-activated transcription factors and kinases (Foyer and Noctor, 2009; Mittler et al., 2011). Also other molecules including lipids and fatty acids are modified by ROS with implications for their signaling functions (Farmer and Mueller, 2013).
Similar to ROS, NO is a small redox signal with versatile chemistry. It is a relatively stable radical but rapidly reacts with other radicals including ROS (Hill et al., 2010). Products of these reactions are reactive nitrogen species (RNS) such as the nitrosonium cation (NO+), the nitroxyl anion (NO−) and higher oxides of NO including ONOO−, NO2, and N2O3. RNS have chemical properties different from their precursors and may trigger specific physiological responses. Like ROS, NO is an important messenger in many physiological processes. It is a stress signal involved in plant responses to high salt, excess light, cold, heat, ozone, UV-B and various pathogens (Leitner et al., 2009; Gaupels et al., 2011a; Mur et al., 2013). Despite the ever-growing importance of NO in plant research, only little is known about enzymatic sources and molecular receptors of NO. Best characterized is the role of NO in stomatal closure and pathogen defence (Mur et al., 2013). In both processes, NO interacts with H2O2 without exact molecular mechanisms deciphered.
The aim of this review is to summarize current knowledge on the interaction of NO with ROS and the antioxidant system in plant stress responses. We will explore how NO can chemically react with pro- and antioxidants and how NO might regulate activity and expression of pro- and antioxidant enzymes. Additionally, functions of non-symbiotic hemoglobins, SOD, GSNOR and peroxiredoxins in regulating RNS homeostasis will be discussed. The last section of this review will detail the roles of individual NO and redox messengers in signaling during stress-induced programmed cell death (PCD).
Manipulation of the NO level has an impact on the antioxidant system
The relevance of NO in stress-induced redox signaling was repeatedly investigated by treatment of plants with NO donors before or during exposure to abiotic stress conditions (Hasanuzzaman et al., 2010; Saxena and Shekhawat, 2013). Table 1 summarizes selected literature reporting the impact of NO donor treatment on H2O2 level, antioxidants and activity of antioxidant enzymes in stressed plants. The authors studied 14 different plant species, 11 stressors, and 6 different NO donors providing a comprehensive overview of the current literature on this topic. A common effect of all stress treatments was the accumulation of H2O2 often accompanied by an increase in malondialdehyde (MDA) levels pointing to ROS-dependent oxidation of lipids. In 19 of the 23 studies activities of all or at least some of the analyzed antioxidant enzymes were up-regulated. These data suggest that stress causes accumulation of ROS, which may then trigger enhancement of the antioxidant defence system.
Table 1
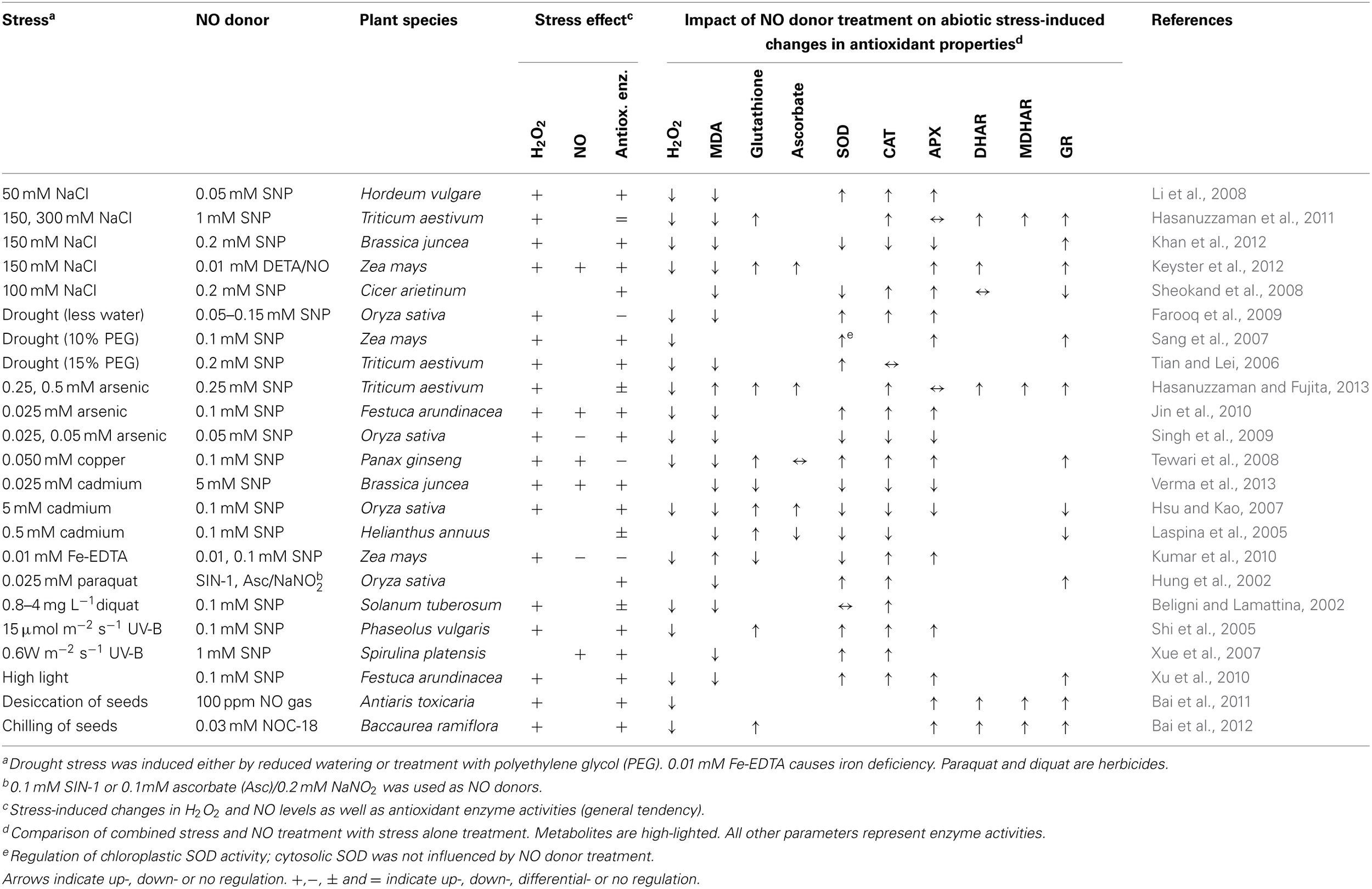
NO donors induce stress tolerance by effecting on the antioxidant properties of plant tissues.
aDrought stress was induced either by reduced watering or treatment with polyethylene glycol (PEG). 0.01 mM Fe-EDTA causes iron deficiency. Paraquat and diquat are herbicides.
b0.1 mM SIN-1 or 0.1mM ascorbate (Asc)/0.2 mM NaNO2 was used as NO donors.
cStress-induced changes in H2O2 and NO levels as well as antioxidant enzyme activities (general tendency).
dComparison of combined stress and NO treatment with stress alone treatment. Metabolites are high-lighted. All other parameters represent enzyme activities.
eRegulation of chloroplastic SOD activity; cytosolic SOD was not influenced by NO donor treatment.
Arrows indicate up-, down- or no regulation. +,−, ± and = indicate up-, down-, differential- or no regulation.
Most of the published studies demonstrated accumulation of NO under stress conditions (Hasanuzzaman et al., 2010; Saxena and Shekhawat, 2013). However, results given in Table 1 as well as other data imply that NO cannot be considered to be a general stress signal. For instance, comparing the effect of 25 μM arsenic between two studies, NO production was induced in Festuca arundinaceae but decreased in Oryza sativa (Table 1) (Singh et al., 2009; Jin et al., 2010). During plant responses to cadmium stress, NO was increased or decreased acting as inducer or inhibitor of stress tolerance, depending on plant species and experimental setup (Arasimowicz-Jelonek et al., 2011a). Moreover, iron deficiency triggered NO signaling in Arabidopsis thaliana (Chen et al., 2010) but repressed basal NO synthesis in Zea mays (Table 1) (Kumar et al., 2010). In this context it is interesting that recent studies revealed NO being a modulator rather than an essential signal in the adaptation of A. thaliana to iron deficiency (Meiser et al., 2011). Together, these findings demonstrate that the link between stress perception and NO signaling is seemingly rather indirect whereas stress can directly cause ROS accumulation by disturbing the mitochondrial and plastidic ETC. Further studies are needed for investigating the biological background of the observed species-specific differences in NO regulation under stress conditions. In sum, the above findings support the notion that endogenous NO is often but not always involved in stress tolerance.
Exogenous NO always improved abiotic stress tolerance concomitant with a decrease in H2O2 and MDA levels (Table 1). This held true, even when endogenous NO was down-regulated, implying that the tested NO donors do not necessarily mimic functions of NO under natural conditions. In the displayed 23 studies, NO treatments either reversed the stress-induced decline or even further amplified up-regulation of the antioxidant system. NO donors never caused a down-regulation of antioxidant enzymes as compared to untreated control plants. For instance, salt stress stimulated SOD, CAT, and APX activities, and this effect was enhanced by SNP co-treatment, whereas copper uptake repressed the same enzymes in Panax ginseng, which was prevented by SNP (Table 1) (Li et al., 2008; Tewari et al., 2008). Again the same enzyme activities were enhanced after arsenic poisoning of O. sativa but SNP application prevented this stress effect (Table 1) (Singh et al., 2009). These findings were explained by NO acting either (I) as a direct scavenger of ROS or (II) inducer of the antioxidant system. In the first case NO would take over functions of the antioxidant system and thereby prevent its activation, like e.g. in arsenic-exposed rice as described above. In the second case NO would trigger antioxidant gene expression or activate antioxidant enzymes e.g., by posttranslational modifications. Previously, NO donors were reported to repress antioxidant enzyme activities. Particularly, SNP inhibited APX and CAT, decreased GSH/GSSG ratio and induced PCD in Arabidopsis suspension cultured cells (Murgia et al., 2004a). However, the research summarized in Table 1 was focussed on investigating mechanisms of NO-mediated stress tolerance. Therefore, NO donors were probably applied in such a way as to prevent any severe stress or damage to the plants although sometimes up to 5 mM SNP was used. We will discuss later in this review the dose dependent effects of NO on the antioxidant system and cell death initiation.
A direct chemical interaction of NO with ROS is only possible if cells or plant parts are being loaded with active NO donor solution from start of the stress treatment until sampling as was the case for Spirulina platensis cells exposed to UV-B and SNP and Brassica junceae leaf discs incubated in salt and DETA/NO donors (Table 1) (Xue et al., 2007; Khan et al., 2012). In other studies, however, measurements were done after NO donors were exhausted suggesting that NO released from the donor did not have a direct influence on ROS levels but might be rather involved in the induction of signaling events controlling the cellular redox status. Farooq et al. (2010) reported that imbibition of seeds in SNP solution rendered adult rice plants more tolerant to drought stress. Hence, NO pre-treatment could induce a primed state, which prepares plants to respond more efficiently to future stress episodes (Conrath, 2011). Alternatively, NO treatment itself could impose stress to the plants acting as the priming stimulus. Exogenous NO might also induce synthesis of endogenous NO, which then can exert signaling or scavenger functions even long after the NO donor is exhausted.
NO donors can have undesired side-effects on the plant's physiology. Therefore, NO accumulating transgenic and mutant plant lines were used for assessing the involvement of NO in development and stress signaling. Transgenic Nicotiana tabacum and A. thaliana expressing the rat neuronal nitric oxide synthase (NOS) behind a 35S promoter accumulated high levels of NO concomitant with developmental defects and altered stress resistance (Chun et al., 2012; Shi et al., 2012). 35S::nNOS lines of Arabidopsis constitutively expressed pathogenesis related (PR) genes, which correlated with enhanced pathogen resistance toward virulent Pseudomonas syringae DC3000 (Shi et al., 2012). These plants also had improved salt and drought tolerance due to reduced stomatal aperture, and were delayed in flowering. The H2O2 content was not determined, but MDA levels were found to be lowered. By comparison, nNOS-expressing tobacco showed growth retardation and constitutive inhibition of CAT, which caused an increase in H2O2 levels (Chun et al., 2012). Probably as a consequence of high NO and H2O2 levels, these plants developed spontaneous lesions, strongly elevated salicylic acid (SA) levels and PR gene expression. Reduced growth, increased oxidative stress and spontaneous lesions was not observed in nNOS expressing A. thaliana plants indicating that they either were less sensitive to NO or accumulated lower levels of NO than the corresponding tobacco transgenic lines.
Collectively, the discussed research argues for ROS being a general stress signal whereas NO signaling depends on the plant species and stress conditions investigated. It can be speculated that NO or the interaction between ROS and NO adds some degree of specificity to the stress signaling by ROS alone. Treatment of plants with NO donors caused a decrease in stress-induced ROS levels and a concomitant enhancement of abiotic stress tolerance. In this process NO might act as a scavenger of ROS or as a signal stimulating the antioxidant potential and/or a primed state of stress defence. Interpretation of the data is complicated by the fact that most of the studies are rather descriptive without exploring the underlying signaling cascades. Moreover, the biological significance of some observed weak effects of NO on ROS and the antioxidant system is ambiguous because slight changes in the cellular redox status could be just a stress marker.
Sources and cellular localization of NO and ROS production
NO and certain ROS cooperate in stress signaling, which is partly independent of their respective production sites because both molecules are supposed to be mobile intra- as well as intercellularly (Foyer and Noctor, 2009; Frohlich and Durner, 2011). Therefore, apoplastic sources can contribute to NO and ROS signal transduction within the cell (Table 2). Important ROS producing enzymes are the members of the NADPH oxidase family (NOX or Respiratory burst oxidase homolog, RBOH). These plasma membrane-associated enzymes synthesize O−2 in the apoplast through transfer of electrons from NADPH to molecular oxygen (Mittler et al., 2011). A rapid ROS burst, frequently observed during plant responses to pathogen infection, is usually mediated by the NOX isoforms D and F (Torres et al., 2002). Further oxidases and cell wall-associated peroxidases are present in the apoplast but their roles in stress responses are less well-defined. In comparison to ROS only little is known about NO formation in the extracellular space (Table 2). At the acidic pH of the apoplast exogenous NO−2 was non-enzymatically reduced to NO, which was accelerated by AsA and phenolics (Bethke et al., 2004). The pathway has been investigated in the barley aleuron layer but might occur also in other tissues. A stress-induced NO burst derived from this spontaneous reaction seems only feasible if NO−2 levels could be rapidly up-regulated, which has not been observed so far. NO−2 could also be reduced to NO by a membrane-associated nitrite:NO reductase (NiNOR) as described for tobacco (Stöhr et al., 2001). However, NiNOR cannot be considered a major player in NO signaling because it is exclusively present in roots functioning in the regulation of NO−3 uptake. Copper amine oxidase 1 (CuAO1) is another candidate enzyme involved in NO synthesis (Wimalasekera et al., 2011). The A. thaliana cuao1 mutant is impaired in polyamine- and abscisic acid-induced NO production. The molecular background underlying this interesting phenotype is still unknown.
Table 2
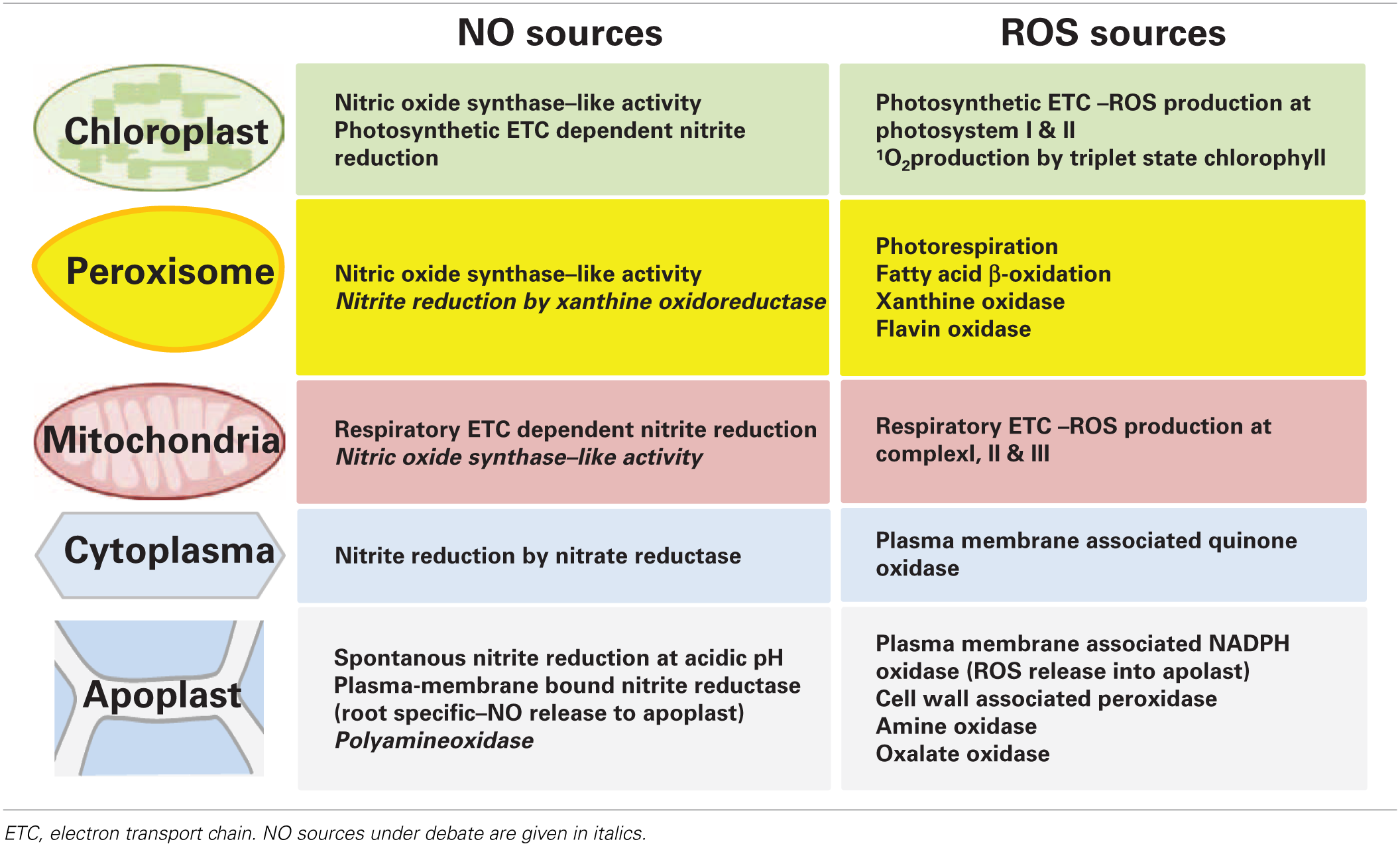
Localization of NO and ROS sources in plant cells.
ETC, electron transport chain. NO sources under debate are given in italics.
Cellular compartments simultaneously producing NO and ROS might be focal points of stress signaling (Table 2). While chloroplasts and mitochondria are major sources of ROS from photosynthetic and respiratory ETC these organelles are also capable of NO synthesis, one proposed mechanism being the transfer of electrons from the ETCs to NO−2 by a nitrite: NO-reductase activity. Such ETC-dependent NO formation was observed in isolated choroplasts from tobacco supplied with 25–100 μM NO−2 and in mitochondria of tobacco suspension cells under anoxia (Planchet et al., 2005; Jasid et al., 2006). More work is needed for investigating if this pathway is active also in stress responses under normoxic conditions. Mammalian NOS oxidizes arginine to citrulline and NO. Although NOS-like activity is considered the most important source of NO accumulation in plant reactions to various stresses the corresponding plant NOS still awaits identification (Leitner et al., 2009; Mur et al., 2013). Recent publications reported on the detection of a NOS-like activity in chloroplasts (Jasid et al., 2006; Tewari et al., 2013). In A. thaliana and Brassica napus protoplasts NO generation was highest immediately after the isolation procedure and decreased during culture. Experiments with a NOS activity assay and specific enzyme inhibitors suggested that NO originated from a NOS-like source. Moreover, simultaneous accumulation of NO and ROS resulted in the formation of ONOO− as detected by the fluorescent dye aminophenyl fluorescein (APF) (Tewari et al., 2013). In line with this, treatment with the fungal elicitor cryptogein also triggered rapid accumulation of both NO and ROS in tobacco epidermal cells (Foissner et al., 2000). The above data imply that stress induces the accumulation of ROS and RNS in the chloroplast, which could then locally effect on photosynthesis or diffuse out of the chloroplast to other cellular compartments.
To date, there is no convincing proof of NOS-like activity in mitochondria (Table 2; Gupta et al., 2011). In contrast, peroxisomes are a source of NO both during salt stress as well as developmental processes such as lateral root growth (Corpas et al., 2009; Schlicht et al., 2013). In A. thaliana transgenic lines expressing GFP linked to peroxisomal targeting signal 1 (PTS1) fluorescence of the NO-specific dye diaminorhodamine co-localized with GFP fluorescence in the peroxisomes. Isolated peroxisomes displayed NOS-like activity, which was calcium dependent and could be inhibited by NOS inhibitors (Table 2). 100 mM NaCl stimulated NO synthesis in peroxisomes, which spread into the cytosol, where it probably contributed to ONOO− formation and protein tyrosine nitration (Corpas et al., 2009). Peroxisomes are active sites of ROS scavenging as well as formation. The main function of peroxisomes is the removal of ROS originating from photosynthetic and mitochondrial ETCs. For this purpose, peroxisomes contain large amounts of CAT but also APX and other antioxidant enzymes. However, after a stress stimulus antioxidant enzymes can be down-regulated possibly by S-nitrosylation or nitration rendering peroxisomes a ROS source rather than a sink (Sandalio et al., 2013). Peroxisomes are often closely associated with mitochondria and/or chloroplasts. Such functional units are essential for efficient ROS scavenging but it can be speculated that they also represent “reaction vessels” for enhancing ROS/RNS signal interaction.
In the past, microscopic studies with NO-specific dyes suggested higher stress-induced NO accumulation in chloroplasts and peroxisomes than in the cytoplasm (e.g., Foissner et al., 2000; Gaupels et al., 2008; Corpas et al., 2009). One possible explanation for this finding would be that the cytoplasm has a rather low capacity of NO synthesis. While NOS-like activity was not detected, nitrate reductase (NR) is the only confirmed NO source in the cytoplasm (Table 2). However, under normal growth conditions NR preferably reduces NO−3 to NO−2, which is then further reduced by nitrite reductase to NH+4. Only under special conditions such as anoxia when NO−2 reaches high levels NR reduces NO−2 to NO at considerable rates (Gupta et al., 2011; Mur et al., 2013). For this reason, it seems unlikely that NR significantly contributes to rapid stress signaling by NO. Overall, chloroplasts and peroxisomes are probably the most important sources of NO and ROS during stress responses. Available data indicate that both signal molecules are produced simultaneously giving rise to the formation of RNS such as ONOO−. ROS mainly originated from NADPH oxidases and ETCs. The NO burst was driven by a yet unidentified NOS-like activity in chloroplasts and peroxisomes. Nitrite reduction to NO either non-enzymatically or by various reductases is thought to contribute comparably less to the NO burst.
Interactions between NO and ROS
Chemical interactions between NO and ROS influence concentration, composition and signaling functions of both reaction partners. For instance, H2O2 was proposed to react with NO yielding 1O2 and NO−in vitro (Noronha-Dutra et al., 1993). If this chemical pathway occurs in vivo is still ambiguous since NO is a rather stable radical, which does not easily bind non-radical species such as H2O2. Physiologically more significant is the fusion of NO with O−2 to give ONOO− (Table 3) (Hill et al., 2010). This radical-radical reaction has a high rate constant and is favored instead of O−2 dismutation to H2O2. As a result, highly cytotoxic and long-lived ROS are replaced by ONOO−, which is short-lived in the cellular environment (Pryor et al., 2006). The exact pathway of ONOO− and ONOOH (peroxynitrous acid) decay to NO−2 and NO−3 at neutral pH is still debated (Table 3). It was suggested that ONOOH isomerises to NO−3 and H+ either directly or indirectly via the radical intermediates NO2 and OH (Goldstein and Merenyi, 2008; Koppenol et al., 2012). The peroxynitrite anion on the other hand yields the RNS NO2, NO, and N2O3 during its degradation to NO−2 (Goldstein and Merenyi, 2008). At neutral pH ONOO− and ONOOH are both present in cells and together form peroxynitrate (O2NOO−/O2NOOH), which decays to NO−2 and O2 as well as 1O2 and NO− (Khan et al., 2000; Jourd'heuil et al., 2001; Gupta et al., 2009; Miyamoto et al., 2009). Meanwhile it is widely accepted that CO2 is an important modulator of ONOO− chemistry in cells. The atmospheric gas rapidly reacts with ONOO− resulting in NO−3 and the radicals NO2 and CO−3 (carbonate anion radical Bonini et al., 1999; Pryor et al., 2006).
Table 3
| ROS | RNS |
|---|---|
| Hydrogen peroxide: H2O2 | Nitric oxide: NO |
| Superoxide: O−2 | Peroxynitrite: ONOO− |
| Singlet oxygen: 1O2 | Peroxynitrous acid: ONOOH |
| Hydroxyl radical: OH | Peroxynitrate: O2NOO− |
| Oxygen: O2 | Peroxynitric acid: O2NOOH |
| Nitrosonium cation: NO+ | |
| Nitroxyl anion:NO− | |
| Nitrogen dioxide: NO2 | |
| Dinitrogentrioxide: N2O3 | |
| Nitrosoglutathione: GSNO | |
| REACTION STOICHIOMETRY | References |
| NO−2 + 2 H+ ↔ NO + H2O | Pryor et al., 2006 |
| NO+ + H2O2 → ONOO− + 2 H+ | Beligni and Lamattina, 2002 |
| NO + O−2 → ONOO− | Miyamoto et al., 2009 |
| 2 NO + O2 → 2 NO2 | Moller et al., 2007 |
| NO2 + NO ↔ N2O3 | Moller et al., 2007 |
| N2O3 + H2O → 2 NO−2 + 2 H+ | Moller et al., 2007 |
| ONOOH → ONOO− + H+ (Ionisation) | Koppenol et al., 2012 |
| ONOOH → NO−3 + H+ (Isomerisation) | Koppenol et al., 2012 |
| ONOOH → NO2 + HO (Homolysis) | Koppenol et al., 2012 |
| ONOO−→ NO + O−2 (Homolysis) | Koppenol et al., 2012 |
| O2NOO− ↔ NO2 + O−2(Homolysis) | Gupta et al., 2009 |
| ONOOH + ONOO− → O2NOO−+ NO−2 + H+ | Gupta et al., 2009 |
| CO2+ ONOO− → CO−3 +NO2 | Pryor et al., 2006 |
Reaction stoichiometry between ROS and RNS.
High levels of NO can react with O2 giving rise to the NO2 radical (Table 3). This pathway is slow in the cytosol but might be efficient in membrane-rich cellular compartments such as chloroplasts and mitochondria owing to the lipophilic nature of NO and O2 (Liu et al., 1998; Pryor et al., 2006). Under continuous NO production NO2 will further react to N2O3 (Pryor et al., 2006; Moller et al., 2007). All reactive nitrogen oxides decompose to the stable derivatives NO−2 and NO−3 within cells. However, as described in the previous section, under acidic conditions e.g., in macrophages and in the plant apoplast N2O3, NO, and NO+ can also originate from NO−2 upon enzymatic or non-enzymatic reduction (Table 3) (Pryor et al., 2006; Combet et al., 2010; Frohlich and Durner, 2011). Hence, dependent on the prevailing cellular environment NO and ROS can interact resulting in the formation of intermediates with distinct molecular properties. For instance, NO, NO−, NO+, and N2O3 bind to nucleophilic residues of proteins causing nitrosation (covalently bound nitroso/-NO adduct) and cysteine- as well as metal S-nitrosylation (coordinate nitrosyl/..NO adduct) (Hill et al., 2010; Fukuto and Carrington, 2011). In contrast, ONOO− and the NO2 radical are involved in oxidation and nitration (covalently bound nitro/-NO2 adduct) of proteins the best studied modifications being 3-nitro-tyrosine residues (Arasimowicz-Jelonek and Floryszak-Wieczorek, 2011; Gaupels et al., 2011a; Radi, 2013). NO2 has less nitrating power than ONOO− except with protein radicals, which result from the reaction of proteins with ROS or CO−3 radicals (Bonini et al., 1999; Pryor et al., 2006). To date, the CO−3 catalyzed binding of NO2 to tyrosyl residues is thought to be the major route of protein nitration.
NO-dependent protein modifications are reversible, which is important for efficient recovery of NO receptors during stress signaling. In mammalian cells, thioredoxins (TRX) denitrosylate proteins (Tada et al., 2008; Benhar et al., 2009). Recently, the central redox switch NPR1 was suggested to be denitrosylated by TRX-h-3 and -5 during incompatible A. thaliana/P. syringae interactions, which caused its monomerisation from oligomers, transfer into the nucleus and subsequent induction of PR genes (Tada et al., 2008). However, the exact mechanism of NPR1 regulation by S-nitrosylation and TRX is still debated (Lindermayr et al., 2010). Denitration of proteins in A. thaliana is probably mediated by peptide methionine sulfoxide reductase (PMSR) under normal growth conditions since pmsr2-1 mutants displayed elevated protein nitration in the night (Bechtold et al., 2009). This enzyme reduces oxidized protein methionine residues using TRX as a co-substrate but how it can function as a denitratase is not yet resolved. Future research will uncover if additional reductases, peroxiredoxin oxidases and peroxidases such as TRX peroxidase are involved in stress signaling by NO-dependent protein modifications.
Apart from proteins many other molecules can be nitrated including lipids, fatty acids, amino acids and nucleotides (Arasimowicz-Jelonek and Floryszak-Wieczorek, 2011). Recently, 8-nitro-cGMP was uncovered as a down-stream signal of ABA, NO, and ROS in inducing stomatal closure at daytime, whereas cGMP regulated stomatal opening at night (Joudoi et al., 2013). 8-nitro-cGMP is now a prime example of how NO, ROS, and cGMP can be integrated in one signaling cascade triggering a physical response.
NO and ROS influence each other's biosynthesis and degradation
ROS are well-known inducers of NO synthesis in various plant species, plant parts and tissues. For example, treatment with 100 μM H2O2 triggered NO synthesis in roots of A. thaliana, which was used in a screen for identification of mutants defective in NO accumulation. This way, the prohibitin PHB3 was uncovered as a regulatory element of ABA- and auxin-induced NO signaling (Wang et al., 2010). Moreover, H2O2 elicited a rapid NO burst in guard cells of mung bean leaves (Phaseolus aureus) (Lum et al., 2002) as well as NOS activity along with PCD in tobacco BY-2 cells (De Pinto et al., 2006). The interplay between ROS, NO and the antioxidant system will be discussed in more detail in the last section of this review. Exposure to ozone (O3) led to high ROS levels and rapid NO production in the leaves of A. thaliana plants (Ahlfors et al., 2009). During the O3 response NO acted as a signal in the onset of the hypersensitive response (HR) and in the regulation of defence-related genes thereby interacting with jasmonic acid (JA), ethylene and SA. In the phloem of Vicia faba NO accumulation upon treatment with 10 and 100 μM H2O2 was dependent on Ca2+ and NOS-like enzyme activity (Gaupels et al., 2008). Although induction of NO biosynthesis through H2O2 and Ca2+ is widely accepted, exact signaling cascades and enzymatic sources of NO are still not well-understood. Effects of H2O2 on NO scavenging enzymes such as GSNOR and hemoglobins were not yet investigated.
NO is not just a down-stream signal of H2O2 but was also reported to influence ROS production and degradation, which hints at complex feed-back regulation between both signal molecules. NO limits ROS accumulation for instance by inhibition of the ROS producing enzyme NADPH oxidase (Yun et al., 2011). After infection of A. thaliana with avirulent pathogens the elevated SNO content inhibited the NADPH oxidase isoform AtRBOHD by S-nitrosylation at Cys 890. According to the author's hypothesis this regulatory process constrains ROS accumulation and subsequent cell death progression (Yun et al., 2011). A means of enhancing antioxidant enzyme activities is the induction of the corresponding genes by NO. Accordingly, 2D-electrophoresis and Western blot analyses revealed that pre-treatment with the NO donor SNAP further increased the Al3+-induced protein levels and activities of APX, SOD, and GR, whereas NOS inhibitor and cPTIO suppressed both the Al3+ and the SNAP effect (Yang et al., 2013). Alternatively, NO could directly modify protein functions. In Antiaris toxicaria NO fumigation improved desiccation tolerance of recalcitrant seeds, which correlated with a decrease in H2O2 levels. The authors proposed that S-nitrosylation enhanced the activities of the antioxidant enzymes GR, APX, and DHAR by preventing their oxidation/carbonylation during desiccation (Bai et al., 2011). Moreover, in salt stressed B. juncea S-nitrosylation of a Fe-SOD caused an increase in its enzyme activity (Sehrawat et al., 2013).
More commonly, however, NO was associated with inhibition rather than activation of antioxidant enzymes. In vitro, tobacco APX and CAT were reversibly inhibited by GSNO, SNAP, and NOC-9 but irreversibly inactivated by SIN-1 (Clark et al., 2000). Inhibition of APX and CAT by NO donors was confirmed in isolated pea mitochondria, leaves of Pelargonium peltatum and suspension cultured cells of A. thaliana and N. tabacum (Murgia et al., 2004a; Arasimowicz-Jelonek et al., 2011b; Marti et al., 2013). SNP and SNAP were the most effective NO donors, whereas GSNO produced variable results. The chemical properties of the donors is an important issue because SNP releases NO+ and SIN-1 simultaneously O−2 and NO whereas most other donors deliver NO. Thus, dependent on the NO donor used and the prevailing redox conditions antioxidant enzyme activity could be affected due to oxidation, S-nitrosylation, nitrosation or nitration. Unfortunately, NO- and ROS-dependent protein modifications were not investigated in the above studies.
Any of the enzymes APX, SOD, MDHAR, DHAR, GR, and CAT was proposed to be S-nitrosylated and/or tyrosine nitrated in vivo in unstressed A. thaliana, salt-stressed citrus (Citrus aurantium), GSNO-treated potato or rice injected with H2O2 for eliciting cell death (Tanou et al., 2009, 2010; Fares et al., 2011; Kato et al., 2012; Lin et al., 2012). S-nitrosylation, however, was only confirmed for APX from GSNO-treated potato leaves (Kato et al., 2012). In the same study DHAR was demonstrated to be S-nitrosylated and inhibited by NO. A possible target Cys essential for enzymatic function was revealed by point mutation of candidate Cys residues. Human manganese SOD is a mitochondrial protein that undergoes site-specific nitration at Tyr34 during inflammation. Inactivation of Mn-SOD by nitration provokes oxidative stress and ultimately dysfunction of mitochondria (Radi, 2013). It would be interesting to elucidate if plant SODs are targets of nitrating species with possible roles e.g., in PCD. Collectively, the discussed data suggest that APX, CAT, and DHAR are good candidates for NO-regulated antioxidant enzymes in plants. A systematic approach is needed for deciphering, which antioxidant enzymes are controlled by NO under stress conditions, and what are the underlying molecular mechanisms.
We mentioned before that NO bioactivity has been implicated both in increased as well as decreased antioxidant enzyme activities and ROS levels. One way of explaining the contradictory findings is based on the hypothesis that NO has a dose-dependent effect on the cellular redox status (Figure 2) (Thomas et al., 2008). At low concentrations NO might stimulate the antioxidant system and promote cell survival while high concentrations of NO cause severe cell damage and even death. In this model trace NO would preferably react with nucleophiles such as lipids, DNA and metal centered proteins but also with oxygen species forming oxidizing and nitrating species including ONOO− and NO2. Little damage and NO-induced signaling will be perceived by the cell triggering antioxidant defence and repair mechanisms. Profound NO production, on the other hand, would promote secondary reactions of NO2 and ONOO− with NO and consequently the accumulation of N2O3. This would shift conditions in the cell from weak oxidative stress toward heavy nitrosative stress, which—according to the hypothesis of Thomas et al. (2008)—inflicts severe damage ultimately leading to cell death. For some biological effects the duration of NO production is decisive because certain target molecules bind NO very slowly or need sequential NO and ROS modifications (Thomas et al., 2008). Thus, in addition to the chemical environment of the cell, which defines the RNS/ROS composition, the extent of NO production is critical in shaping stress signaling by NO.
Figure 2
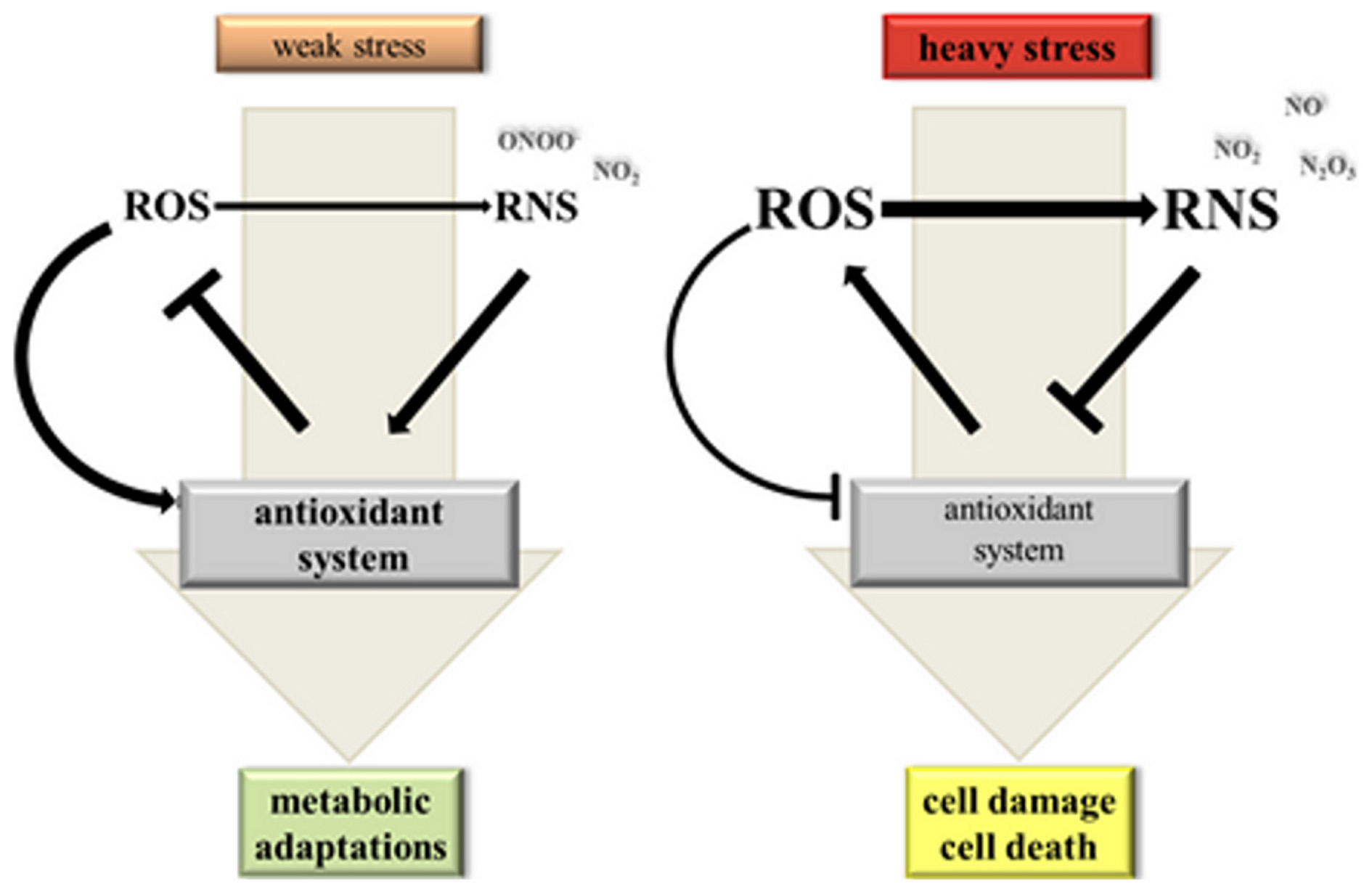
Hypothetical model on the dynamic interaction between NO, ROS and the antioxidant system under stress conditions. Weak stress triggers a moderate elevation of ROS (reactive oxygen species) and NO levels. ROS act as signals inducing NO synthesis and activation of the antioxidant system for improved metabolic adaptation. If ROS is produced at a somewhat higher rate than NO there would be mainly formation of oxidizing and nitrating RNS (reactive nitrogen species) imposing a weak oxidative stress to the cell. Heavy stress leads to a strong ROS and RNS burst. High NO levels promote formation of N2O3 from NO2 and NO and consequently nitrosative stress. Under these conditions ROS and RNS inhibit the antoxidant system causing damage and ultimately death of plant cells.
Interactions between NO and antioxidants
The versatility of signaling by RNS and ROS is further extended by their interaction with antioxidants. Reduced ascorbate does not react with NO but with nitrosating species NO+, N2O3 and with S-nitrosothiols (Scorza et al., 1997; Kytzia et al., 2006). Consequently, NO is released and AsA is converted to DHA (Combet et al., 2010). DHA spontaneously decays to the ascorbyl radical, which can combine with NO to give O-nitrosoascorbate. The latter finally undergoes hydrolysis to ascorbate and NO−2 (Kytzia et al., 2006). AsA can also scavenge ONOO− with rather slow kinetics at neutral pH but rapid kinetics at pH 5.8 yielding NO−2 and NO−3 via unknown intermediates (Kurz et al., 2003). Likewise, GSH affects ONOO− levels either by reduction to NO−2 or by radical-radical interactions of NO2 with the glutathiyl radical resulting in the formation of nitroglutathione GSNO2, which in turn can release NO (Balazy et al., 1998). Moreover, GSH effectively prevents ONOO− mediated tyrosine nitration by re-reducing tyrosyl radicals and catalysing the formation of non-nitrating O2NOO− from NO2 and O−2 (Kirsch et al., 2001). The biological significance of the above proposed pathways of ONOO− degradation remains to be investigated. However, the high concentrations of GSH and AsA in plant cells could contribute to maintaining low levels of NO derivatives under non-stress conditions.
Other known plant scavengers of ONOO− include gamma-tocopherol (vitamin E; Desel et al., 2007), carotenoids and the flavonoids ebselen, epicatechin and quercetin (Haenen et al., 1997). Some of the above compounds are not specific for ONOO− but scavenge NO and ROS, too. Recently, cytokinins were demonstrated to be involved in controlling NO levels in A. thaliana (Liu et al., 2013). Continuous root-uptake of 120 μM SNP severely inhibited growth of A. thaliana WT plants whereas the mutant line cnu-1/amp1 was resistant to the same NO treatment. Further characterization of the mutant revealed a correlation between NO resistance and elevated cytokinin levels. Accordingly, WT plants infiltrated with the cytokinin zeatin displayed improved growth on SNP-loaded agar medium. In vitro, zeatin was nitrated by peroxynitrite, which produced 8-nitro-zeatin. In vivo, SNP caused strong accumulation of 8-nitro-zeatin in cnu-1 as compared to WT. From these results, the authors concluded that cytokinins regulate NO levels by binding the NO derivative ONOO− (Liu et al., 2013).
NO interacts with glutathione in various ways. At the transcriptional level SNP and GSNO stimulated genes involved in GSH synthesis causing elevated levels of total glutathione in Medicago truncatula roots (Innocenti et al., 2007). Accordingly, NO donor treatment triggered an increase in total glutathione in 8 of 10 studies summarized in Table 1. In contrast, SNP had no strong effect on GSH concentrations in tobacco BY-2 cells (De Pinto et al., 2002). At the level of chemical interactions GSH binds NO by S-nitrosylation. GSNO is formed either after (1) ROS-induced accumulation of glutathiyl radicals, which bind NO with rate constants near the diffusion-controlled limit (Madej et al., 2008) or after (2) S-nitrosylation of GSH by nitrogen oxides such as NO+ and N2O3 (Broniowska et al., 2013). GSNO then functions as storage and transport form of NO. It is regarded as an endogenous NO donor, which releases free NO (2 GSNO → 2 NO + GSSG) or S-nitrosylates proteins by transferring the nitroso adduct (Broniowska et al., 2013; Mur et al., 2013).
Enzymatic regulation of NO homeostasis by GSNOR, hemoglobin and pro- as well as antioxidant enzymes
Levels of the S-nitrosylated tripeptide GSNO are tightly controlled by the enzyme GSNOR. This GSH-dependent formaldehyde dehydrogenase catalyzes the transformation of GSNO to GSSG and hydroxylamine (NH2NO) in the presence of GSH and NADH as the reducing species (Figure 3) (Liu et al., 2001; Sakamoto et al., 2002). In A. thaliana silencing or mutation of GSNOR1 caused accumulation of S-nitrosothiols, NO and NO−3 indicating that the corresponding enzyme is a major player in NO homeostasis (Sakamoto et al., 2002). GSNOR1 deficient plants were severely affected in growth and development (Kwon et al., 2012). They also showed increased resistance to the herbicide paraquat and altered responses toward heat stress and pathogen infection (Diaz et al., 2003; Feechan et al., 2005; Rusterucci et al., 2007; Lee et al., 2008; Chen et al., 2009; Holzmeister et al., 2011). In addition to control of NO levels, GSNOR is also indirectly involved in protein denitrosylation because GSNO and S-nitrosylated proteins are in equilibrium (Benhar et al., 2009; Malik et al., 2011). For more information on GSNOR functions refer to recent reviews (Leitner et al., 2009; Gaupels et al., 2011a; Mur et al., 2013). In mammalian/human cells CuZn-SOD and GPX (glutathione peroxidase) were proposed to use GSNO as a substrate and might act in protein denitrosylation without physiological functions being well-established yet (Benhar et al., 2009).
Figure 3
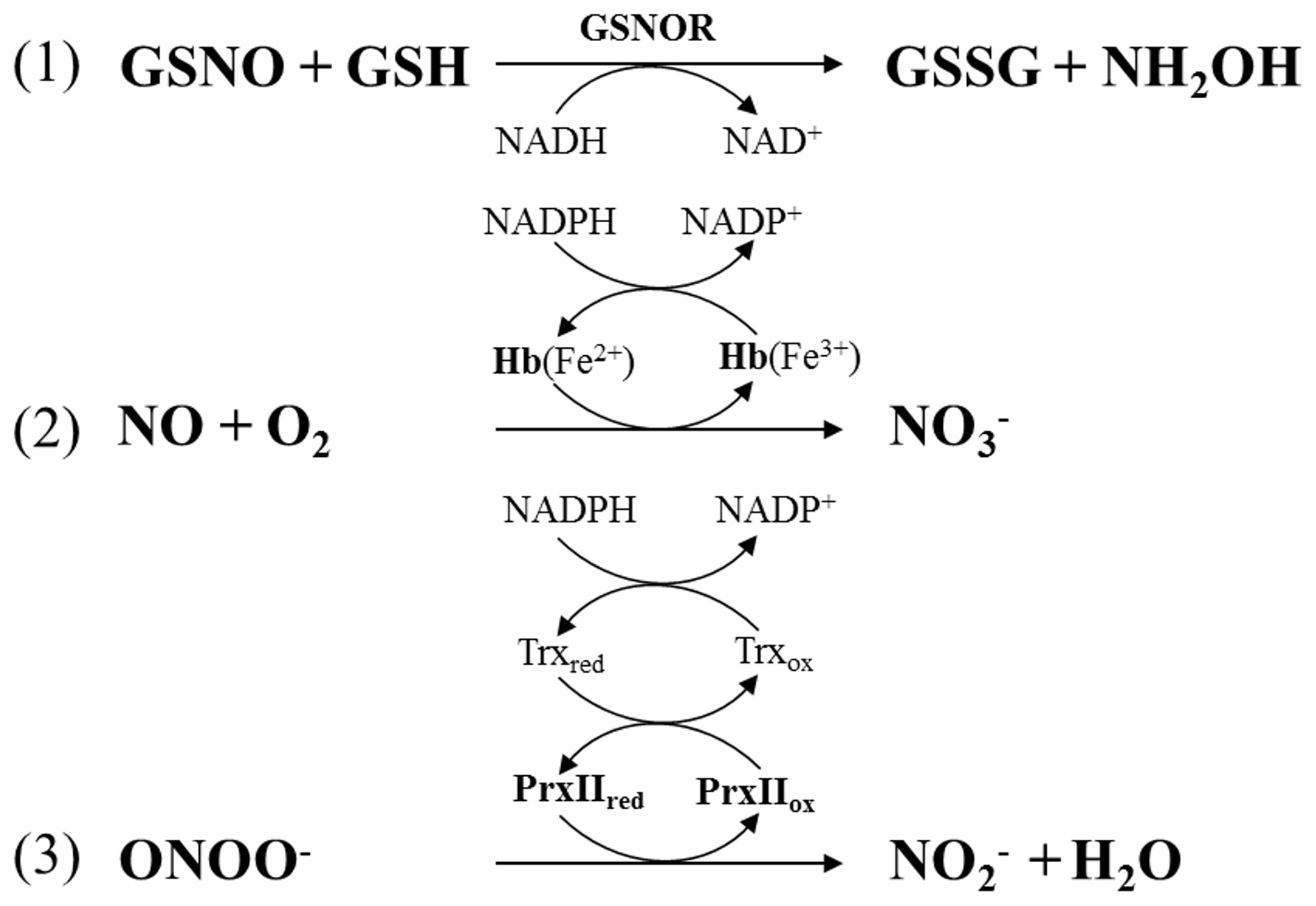
Enzymatic regulation of NO homeostasis by (1) S-nitrosogutathione reductase (GSNOR), (2) hemoglobin (Hb), and (3) peroxiredoxin IIE (PrxIIE). PrxIIE is reduced by thioredoxin (Trx).
Another upcoming topic is the modulation of NO homeostasis by plant hemoglobins. Class-1 Hb1 catalyse the turnover of NO to NO−3 thereby influencing growth, development and stress responses (Figure 3) (Hill et al., 2010; Hebelstrup et al., 2012). Particularly, the role of alfalfa and A. thaliana Hb1 in hypoxia has been studied in more detail (Dordas et al., 2003; Perazzolli et al., 2004; Hebelstrup et al., 2012). It was shown that hypoxia triggered expression of the Hb1-coding gene in roots, probably for confining the stress-induced accumulation of NO. Reduced expression of Hb1 in transgenic and mutant lines caused an increase in NO levels concomitant with decreased plant growth whereas Hb1 over-expression improved plant fitness during hypoxia. By scavenging NO the plant might suppress a costly defence response for saving energy and valuable nitrogen under limited oxygen availability (Hebelstrup et al., 2012). Recently, Hb1 was found to be involved in pathogen resistance. A. thaliana mutants defective in the Hb1-coding gene GLB1 were more resistant to the hemibiotrophic P. syringae and the necrotrophic fungus Botrytis cinerea (Mur et al., 2012). The mutant phenotype was reversed by over-expression of GLB1 under control of the 35S promoter. The enhanced resistance in the glb1 mutant correlated with accumulation of SA, JA, and ET. GLB1 was down-regulated in WT plants during infection, which probably facilitated the induction of defence responses by NO accumulation.
Notably, human hemoglobin degrades ONOO− to NO−3in vitro further extending possible functions of hemoglobins in NO signaling (Romero et al., 2003). By comparison plants have evolved efficient mechanisms for enzymatic detoxification of ONOO− by thiol-dependent peroxidases. The A. thaliana peroxiredoxin IIE (PrxII E) and glutathione peroxidase 5 (Gpx5) of poplar both reduce ONOO− to NO−2 (Figure 3) (Sakamoto et al., 2003; Romero-Puertas et al., 2008; Ferrer-Sueta and Radi, 2009). Both enzymes are then reactivated by thioredoxin in a NADPH-consuming manner. Hence, thioredoxin functions include ROS and ONOO− scavenging as well as protein denitrosylation illustrating again the essential roles of this enzyme in ROS and RNS control.
At neutral (but not acidic) pH NO−2 is a rather stable decomposition product of NO and its derivatives. However, a number of plant enzymes can convert NO−2 to RNS most prominent examples being nitrite reductase and nitrate reductase, which reduce NO−2 to NO (Stöhr et al., 2001; Morot-Gaudry-Talarmain et al., 2002; Gupta et al., 2011). During severe hypoxia deoxygenated A. thaliana Hb1 might act as nitrite reductase although with rather slow kinetics (Tiso et al., 2012). Given the high concentrations of NO−2 in hypoxic plant tissues Hb1 might still significantly contribute to NO accumulation (Sturms et al., 2011). A more wide-spread phenomenon could be the nitration-promoting activity of peroxidases. For instance, three A. thaliana hemoglobins and Hb1 of Medicago sativa were capable of mediating protein nitration via NO−2 oxidation to NO2 by a H2O2-dependent peroxidase activity (Sakamoto et al., 2004; Maassen and Hennig, 2011). Sakihama et al. (2003) demonstrated the enzymatic nitration of p-coumaric acid by action of horseradish peroxidase in the presence of NO−2 and H2O2. All the above data on Hb1 acting as nitrite reductase and enzymatic nitration by peroxidases were obtained in vitro and it is difficult to draw any meaningful conclusions for the in vivo situation.
NO and redox signaling in cell death
ROS and RNS are major players in plant stress signaling. In this section we will survey current knowledge on the roles of ROS, RNS and elements of the antioxidant system in cell death events induced by biotic and abiotic stressors. Plant PCD was described as a genetically controlled cell suicide exhibiting marked similarities but also considerable differences to apoptosis in animal/human cells (Mur et al., 2008; De Pinto et al., 2012). Plants attacked by an avirulent pathogen develop HR, which is a defence mechanism for restricting the spread of pathogens by cell wall reinforcement, production of defensive secondary metabolites and ultimately cell death (Mur et al., 2008).
Almost 20 years ago Chris Lamb and his co-workers discovered that soybean cells infected with avirulent Pseudomonas syringae pv. glycinea accumulated high levels of H2O2, which functioned as a cell death inducer during the HR (Levine et al., 1994). Suppression of the pathogen-induced H2O2 burst by the NADPH oxidase inhibitor diphenylene iodonium (DPI) prevented cell death whereas low millimolar concentrations of exogenous H2O2 triggered HR-PCD in a calcium-dependent manner (Levine et al., 1994, 1996). Later, researchers of the same group demonstrated that NO was another essential messenger in cell death execution (Delledonne et al., 1998). Application of a NO scavenger and a NOS activity inhibitor both reduced HR-PCD of soybean suspension cells infected with avirulent bacterial pathogens. Importantly, SNP triggered cell death most efficiently in conjunction with ROS but not in the presence of DPI or CAT. ROS donors in turn efficiently killed soybean cells only if applied together with SNP (Delledonne et al., 1998). Comparable results were obtained with tobacco BY-2 cells. Simultaneous application of SNP and the H2O2-generating donor system glucose/glucose oxidase but not each individual donor alone caused a drop in ascorbate and glutathione levels, inhibition of APX and consequently PCD of tobacco BY-2 cells (De Pinto et al., 2002). Therefore, it was postulated that NO and ROS cooperate in cell death signaling (Figure 2).
Recent studies have begun to unravel the underlying modes of interactions between NO, ROS and the antioxidant system during PCD. It was shown that ONOO− arose in A. thaliana plants challenged by avirulent Pseudomonas syringae (Gaupels et al., 2011b). The peak of ONOO− formation from NO and O−2 coincided with the onset of the PCD. In unstressed plants ONOO− was continuously scavenged by PrxIIE, which was inhibited by S-nitrosylation in course of the HR (Romero-Puertas et al., 2007). The fact that ONOO− levels are controlled in a sophisticated manner would imply an important role of this RNS in the induction of cell death and pathogen resistance. However, contrary to mammalian cells this RNS does not kill plant cells (Delledonne et al., 2001). It was demonstrated that SOD, GR, CAT, and APX, which are all involved in ROS depletion, can be tyrosine nitrated by ONOO− (Chaki et al., 2009; Lozano-Juste et al., 2011). If this is a significant process in vivo remains to be proven.
H2O2 rather than O−2 was proposed to be a pivotal signal in regulating PCD. This particular ROS acts as an inducer of NO synthesis in tobacco cells (De Pinto et al., 2006) and in mutant plants with disturbed redox homeostasis. For instance, rice knock-out mutants defective in a CAT-coding gene showed increased H2O2 levels, nitrate reductase-dependent accumulation of NO and spontaneous leaf cell death (Lin et al., 2012). Application of the NO scavenger PTIO mitigated the cell death phenotype. The importance of a down-regulation of ROS detoxifying enzymes during PCD was further corroborated by the finding that overexpression of thylakoidal APX led to a higher resistance against SNP induced cell death (Murgia et al., 2004b). In A. thaliana WT plants 5mM SNP triggered H2O2 accumulation and cell death, which was both reduced in the transgenic line probably because H2O2 was degraded by the elevated APX activity in these plants. The antioxidant enzymes CAT and APX control H2O2 levels under mild stress conditions. Severe cadmium stress triggered NO as well as H2O2 accumulation and senescence-like PCD of A. thaliana suspension cultured cells (De Michele et al., 2009). However, co-treatment with the NOS inhibitor L-NMMA prevented the NO-dependent inhibition of CAT and APX, which in turn reduced H2O2 levels and increased cell viability under cadmium stress.
Mechanical wounding provokes cell damage, which could serve as a point of entry into the plant e.g., for pathogenic bacteria. To avoid this, PCD is triggered in intact cells nearby the damaged cells for sealing the wound site. In wounded leaves of Pelargonium peltatum NO accumulation was restricted to the site of injury (Arasimowicz et al., 2009). Treatment with cPTIO confirmed that NO inhibited APX and CAT activity thereby temporarily enhancing the H2O2 content at the edge of the wound. Pre-treatment of leaves with NO donors before wounding prevented the H2O2 burst and reduced necrotic cell death in sweet potato (Lin et al., 2011). The exact mechanism of NO action was not determined but available data suggest that APX, GR, MDHAR and thioredoxin are S-nitrosylated during PCD, which could affect their activity (Murgia et al., 2004b; Lin et al., 2012). Inhibition of GR and MDHAR would also impact on the redox status of the glutathione and ascorbate pools. It should be considered that enzymatic activity can also be influenced by ROS-dependent modifications, which was proposed for oxidation-triggered inhibition of APX (Figure 2) (De Pinto et al., 2006). The latter enzyme was also suppressed in gene expression during PCD (De Pinto et al., 2006).
The role of NO in incompatible interactions between A. thaliana and avirulent Pseudomonas syringae was investigated using transgenic plant lines expressing a bacterial NO dioxygenase (NOD, flavohemoglobin) (Zeier et al., 2004). NOD expression attenuated the pathogen-induced NO accumulation. As a consequence the H2O2 burst was diminished and transgenic plants developed less HR-PCD and were delayed in SA-dependent PR1 expression. These results support again the hypothesis that high levels of NO amplify redox signaling during PCD by inhibiting the plant antioxidant machinery (Zeier et al., 2004). NO and H2O2 might mutually enhance each other's accumulation by positive feed-back regulation. To this end, NO and ROS producing enzymes as well as elements of the antioxidant system must be regulated in a highly coordinate fashion for initiation of PCD. The exact signaling pathways remain to be deciphered in future studies.
However, the plant must also constrain stress signaling by NO, ROS and the antioxidant system for avoiding excessive damage by runaway cell death. Therefore, it is worth mentioning that both ROS as well as NO were found to induce genes involved in cell protection such as a gene coding for glutathione S-transferase (Levine et al., 1994). Yun and colleagues (Yun et al., 2011) even demonstrated inhibition of the ROS-producing enzyme AtRBOHD by NO in A. thaliana challenged by avirulent bacteria. The authors proposed a model, in which the early burst of ROS and NO initiates HR-PCD but at later stages of the defence response the SNO levels exceed a certain threshold and subsequently the AtRBOHD is inactivated by S-nitrosylation at Cys 890, which terminates the HR. In contrast to R gene-mediated resistance against avirulent pathogens, bacterial lipopolysaccharides (LPS) elicit basal pathogen resistance without onset of HR-PCD. LPS-induced NO synthesis by an arginine-dependent enzymatic source even protected plant cells against oxidative stress and cell death by enhancing the activities of CAT, SOD, and POD. The changed cellular redox status contributed to the regulation of NPR1-dependent expression of defence genes (Sun et al., 2012). In sum, NO can either act as an inducer or suppressor of plant PCD dependent on its local cellular levels and its tightly controlled interaction with ROS and elements of the antioxidant system (Figure 2).
Concluding remarks
ROS and NO are increasingly recognized signaling molecules in plant physiology. While research on ROS has a long history NO came into focus only 15 years ago. In the present paper we reviewed recent literature dealing with the interaction between ROS, NO and the antioxidant system during stress defence. As one interesting outcome we found that exposure of plants to unfavorable conditions inevitably induced ROS but not necessarily NO accumulation. ROS can arise as a toxic by-product of disturbed energy metabolism and/or can be produced for signaling purposes. In contrast, NO is rather a highly specialized second messenger, which modifies ROS signaling or acts independently of ROS. Significantly, ROS and NO bursts are often triggered simultaneously—sometimes even in the same cellular compartment. Particularly chloroplasts and peroxisomes are hotspots of NO-ROS interactions. NO, ROS and antioxidants chemically react resulting in the formation of RNS such as ONOO−, NO2, N2O3, and GSNO. More indirect interactions include induction of NO synthesis by H2O2 and accumulation of ROS due to inhibition of antioxidant enzymes by NO-dependent protein modifications. Uncontrolled self-amplification of ROS/RNS signaling might provoke nitrosative stress and ultimately PCD. Therefore, plants have developed efficient measures for controlling NO levels by GSNOR, hemoglobins and other RNS scavenging enzymes. This review was also aimed at investigating the extreme versatility of possible reactions between NO, ROS and the antioxidant system. Many of the discussed findings originate from in vitro systems or animal/human models. More basic research is urgently needed for defining chemical reactions and their products actually occurring in planta.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Statements
Acknowledgments
We thank Werner Heller for helpful discussions and critical reading of the manuscript. This work was supported by the Deutsche Forschungsgemeinschaft (grant GA 1358/3-2 to Frank Gaupels).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1
Ahlfors R. Brosche M. Kollist H. Kangasjarvi J. (2009). Nitric oxide modulates ozone-induced cell death, hormone biosynthesis and gene expression in Arabidopsis thaliana. Plant J. 58, 1–12. 10.1111/j.1365-313X.2008.03756.x
2
Arasimowicz-Jelonek M. Floryszak-Wieczorek J. (2011). Understanding the fate of peroxynitrite in plant cells - from physiology to pathophysiology. Phytochemistry72, 681–688. 10.1016/j.phytochem.2011.02.025
3
Arasimowicz-Jelonek M. Floryszak-Wieczorek J. Gwozdz E. A. (2011a). The message of nitric oxide in cadmium challenged plants. Plant Sci. 181, 612–620. 10.1016/j.plantsci.2011.03.019
4
Arasimowicz-Jelonek M. Floryszak-Wieczorek J. Kosmala A. (2011b). Are nitric oxide donors a valuable tool to study the functional role of nitric oxide in plant metabolism. Plant Biol. 13, 747–756. 10.1111/j.1438-8677.2010.00430.x
5
Arasimowicz M. Floryszak-Wieczorek J. Milczarek G. Jelonek T. (2009). Nitric oxide, induced by wounding, mediates redox regulation in pelargonium leaves. Plant Biol. 11, 650–663. 10.1111/j.1438-8677.2008.00164.x
6
Bai X. Yang L. Tian M. Chen J. Shi J. Yang Y. et al . (2011). Nitric oxide enhances desiccation tolerance of recalcitrant Antiaris toxicaria seeds via protein S-nitrosylation and carbonylation. PLoS ONE6:e20714. 10.1371/journal.pone.0020714
7
Bai X.-G. Chen J.-H. Kong X.-X. Todd C. D. Yang Y.-P. Hu X.-Y. et al . (2012). Carbon monoxide enhances the chilling tolerance of recalcitrant Baccaurea ramiflora seeds via nitric-oxide-mediated glutathione homeostasis. Free Radic. Biol. Med. 53, 710–720. 10.1016/j.freeradbiomed.2012.05.042
8
Balazy M. Kaminski P. M. Mao K. Tan J. Wolin M. S. (1998). S-Nitroglutathione, a product of the reaction between peroxynitrite and glutathione that generates nitric oxide. J. Biol. Chem. 273, 32009–32015. 10.1074/jbc.273.48.32009
9
Bechtold U. Rabbani N. Mullineaux P. M. Thornalley P. J. (2009). Quantitative measurement of specific biomarkers for protein oxidation, nitration and glycation in Arabidopsis leaves. Plant J. 59, 661–671. 10.1111/j.1365-313X.2009.03898.x
10
Beligni M. V. Lamattina L. (2002). Nitric oxide interferes with plant photo-oxidative stress by detoxifying reactive oxygen species. Plant Cell Environ. 25, 737–748. 10.1046/j.1365-3040.2002.00857.x
11
Benhar M. Forrester M. T. Stamler J. S. (2009). Protein denitrosylation: enzymatic mechanisms and cellular functions. Nat. Rev. Mol. Cell Biol. 10, 721–732. 10.1038/nrm2764
12
Bethke P. C. Badger M. R. Jones R. L. (2004). Apoplastic synthesis of nitric oxide by plant tissues. Plant Cell16, 332–341. 10.1105/tpc.017822
13
Bonini M. G. Radi R. Ferrer-Sueta G. Ferreira A. M. Augusto O. (1999). Direct EPR detection of the carbonate radical anion produced from peroxynitrite and carbon dioxide. J. Biol. Chem. 274, 10802–10806. 10.1074/jbc.274.16.10802
14
Broniowska K. A. Diers A. R. Hogg N. (2013). S-nitrosoglutathione. Biochim. Biophys. Acta1830, 3173–3181. 10.1016/j.bbagen.2013.02.004
15
Buchanan B. Gruissem W. Jones R. (2002). Biochemistry and Molecular Biology of Plants. Hoboken, NJ: John Wiley and Sons.
16
Chaki M. Valderrama R. Fernandez-Ocana A. M. Carreras A. Lopez-Jaramillo J. Luque F. et al . (2009). Protein targets of tyrosine nitration in sunflower (Helianthus annuus L.) hypocotyls. J. Exp. Bot. 60, 4221–4234. 10.1093/jxb/erp263
17
Chen R. Sun S. Wang C. Li Y. Liang Y. An F. et al . (2009). The Arabidopsis PARAQUAT RESISTANT2 gene encodes an S-nitrosoglutathione reductase that is a key regulator of cell death. Cell Res. 19, 1377–1387. 10.1038/cr.2009.117
18
Chen W. W. Yang J. L. Qin C. Jin C. W. Mo J. H. Ye T. et al . (2010). Nitric oxide acts downstream of auxin to trigger root ferric-chelate reductase activity in response to iron deficiency in Arabidopsis. Plant Physiol. 154, 810–819. 10.1104/pp.110.161109
19
Chun H. J. Park H. C. Koo S. C. Lee J. H. Park C. Y. Choi M. S. et al . (2012). Constitutive expression of mammalian nitric oxide synthase in tobacco plants triggers disease resistance to pathogens. Mol. Cells34, 463–471. 10.1007/s10059-012-0213-0
20
Clark D. Durner J. Navarre D. A. Klessig D. F. (2000). Nitric oxide inhibition of tobacco catalase and ascorbate peroxidase. Mol. Plant Microbe Interact. 13, 1380–1384. 10.1094/MPMI.2000.13.12.1380
21
Combet E. El Mesmari A. Preston T. Crozier A. McColl K. E. (2010). Dietary phenolic acids and ascorbic acid: influence on acid-catalyzed nitrosative chemistry in the presence and absence of lipids. Free Radic. Biol. Med. 48, 763–771. 10.1016/j.freeradbiomed.2009.12.011
22
Conrath U. (2011). Molecular aspects of defence priming. Trends Plant Sci. 16, 524–531. 10.1016/j.tplants.2011.06.004
23
Corpas F. J. Hayashi M. Mano S. Nishimura M. Barroso J. B. (2009). Peroxisomes are required for in vivo nitric oxide accumulation in the cytosol following salinity stress of Arabidopsis plants. Plant Physiol. 151, 2083–2094. 10.1104/pp.109.146100
24
De Michele R. Vurro E. Rigo C. Costa A. Elviri L. Di Valentin M. et al . (2009). Nitric oxide is involved in cadmium-induced programmed cell death in Arabidopsis suspension cultures. Plant Physiol. 150, 217–228. 10.1104/pp.108.133397
25
De Pinto M. C. Locato V. De Gara L. (2012). Redox regulation in plant programmed cell death. Plant Cell Environ. 35, 234–244. 10.1111/j.1365-3040.2011.02387.x
26
De Pinto M. C. Paradiso A. Leonetti P. De Gara L. (2006). Hydrogen peroxide, nitric oxide and cytosolic ascorbate peroxidase at the crossroad between defence and cell death. Plant J. 48, 784–795. 10.1111/j.1365-313X.2006.02919.x
27
De Pinto M. C. Tommasi F. De Gara L. (2002). Changes in the antioxidant systems as part of the signaling pathway responsible for the programmed cell death activated by nitric oxide and reactive oxygen species in tobacco Bright-Yellow 2 cells. Plant Physiol. 130, 698–708. 10.1104/pp.005629
28
Delledonne M. Xia Y. Dixon R. A. Lamb C. (1998). Nitric oxide functions as a signal in plant disease resistance. Nature394, 585–588. 10.1038/29087
29
Delledonne M. Zeier J. Marocco A. Lamb C. (2001). Signal interactions between nitric oxide and reactive oxygen intermediates in the plant hypersensitive disease resistance response. Proc. Natl. Acad. Sci. U.S.A. 98, 13454–13459. 10.1073/pnas.231178298
30
Desel C. Hubbermann E. M. Schwarz K. Krupinska K. (2007). Nitration of gamma-tocopherol in plant tissues. Planta226, 1311–1322. 10.1007/s00425-007-0552-9
31
Diaz M. Achkor H. Titarenko E. Martinez M. C. (2003). The gene encoding glutathione-dependent formaldehyde dehydrogenase/GSNO reductase is responsive to wounding, jasmonic acid and salicylic acid. FEBS Lett. 543, 136–139. 10.1016/S0014-5793(03)00426-5
32
Dordas C. Hasinoff B. B. Igamberdiev A. U. Manac'h N. Rivoal J. Hill R. D. (2003). Expression of a stress-induced hemoglobin affects NO levels produced by alfalfa root cultures under hypoxic stress. Plant J. 35, 763–770. 10.1046/j.1365-313X.2003.01846.x
33
Fares A. Rossignol M. Peltier J. B. (2011). Proteomics investigation of endogenous S-nitrosylation in Arabidopsis. Biochem. Biophys. Res. Commun. 416, 331–336. 10.1016/j.bbrc.2011.11.036
34
Farmer E. E. Mueller M. J. (2013). ROS-mediated lipid peroxidation and RES-activated signaling. Annu. Rev. Plant Biol. 64, 429–450. 10.1146/annurev-arplant-050312-120132
35
Farooq M. Basra S. M. A. Wahid A. Ahmad N. Saleem B. A. (2009). Improving the drought tolerance in rice (Oryza sativa L.) by exogenous application of salicylic acid. J. Agron. Crop Sci. 195, 237–246. 10.1111/j.1439-037x.2009.00365.x
36
Farooq M. Kobayashi N. Ito O. Wahid A. Serraj R. (2010). Broader leaves result in better performance of indica rice under drought stress. J. Plant Physiol. 167, 1066–1075. 10.1016/j.jplph.2010.03.003
37
Feechan A. Kwon E. Yun B. W. Wang Y. Pallas J. A. Loake G. J. (2005). A central role for S-nitrosothiols in plant disease resistance. Proc. Natl. Acad. Sci. U.S.A. 102, 8054–8059. 10.1073/pnas.0501456102
38
Ferrer-Sueta G. Radi R. (2009). Chemical biology of peroxynitrite: kinetics, diffusion, and radicals. ACS Chem. Biol. 4, 161–177. 10.1021/cb800279q
39
Foissner I. Wendehenne D. Langebartels C. Durner J. (2000). In vivo imaging of an elicitor-induced nitric oxide burst in tobacco. Plant J. 23, 817–824. 10.1046/j.1365-313X.2000.00835.x
40
Foyer C. H. Noctor G. (2009). Redox regulation in photosynthetic organisms: signaling, acclimation, and practical implications. Antioxid. Redox Signal. 11, 861–905. 10.1089/ars.2008.2177
41
Frohlich A. Durner J. (2011). The hunt for plant nitric oxide synthase (NOS): is one really needed. Plant Sci. 181, 401–404. 10.1016/j.plantsci.2011.07.014
42
Fukuto J. M. Carrington S. J. (2011). HNO signaling mechanisms. Antioxid. Redox Signal. 14, 1649–1657. 10.1089/ars.2010.3855
43
Gaupels F. Furch A. C. Will T. Mur L. A. Kogel K. H. Van Bel A. J. (2008). Nitric oxide generation in Vicia faba phloem cells reveals them to be sensitive detectors as well as possible systemic transducers of stress signals. New Phytol. 178, 634–646. 10.1111/j.1469-8137.2008.02388.x
44
Gaupels F. Kuruthukulangarakoola G. T. Durner J. (2011a). Upstream and downstream signals of nitric oxide in pathogen defence. Curr. Opin. Plant Biol. 14, 707–714. 10.1016/j.pbi.2011.07.005
45
Gaupels F. Spiazzi-Vandelle E. Yang D. Delledonne M. (2011b). Detection of peroxynitrite accumulation in Arabidopsis thaliana during the hypersensitive defense response. Nitric Oxide25, 222–228. 10.1016/j.niox.2011.01.009
46
Goldstein S. Merenyi G. (2008). The chemistry of peroxynitrite: implications for biological activity. Meth. Enzymol. 436, 49–61. 10.1016/S0076-6879(08)36004-2
47
Gupta D. Harish B. Kissner R. Koppenol W. H. (2009). Peroxynitrate is formed rapidly during decomposition of peroxynitrite at neutral pH. Dalton Trans. 29, 5730–5736. 10.1039/b905535e
48
Gupta K. J. Fernie A. R. Kaiser W. M. Van Dongen J. T. (2011). On the origins of nitric oxide. Trends Plant Sci. 16, 160–168. 10.1016/j.tplants.2010.11.007
49
Haenen G. R. Paquay J. B. Korthouwer R. E. Bast A. (1997). Peroxynitrite scavenging by flavonoids. Biochem. Biophys. Res. Commun. 236, 591–593. 10.1006/bbrc.1997.7016
50
Hasanuzzaman M. Fujita M. (2013). Exogenous sodium nitroprusside alleviates arsenic-induced oxidative stress in wheat (Triticum aestivum L.) seedlings by enhancing antioxidant defense and glyoxalase system. Ecotoxicology22, 584–596. 10.1007/s10646-013-1050-4
51
Hasanuzzaman M. Hossain M. A. Fujita M. (2010). Physiological and biochemical mechanism of nitric oxide induced abiotic stress tolerance in plants. Am. J. Plant Physiol. 5, 295–324. 10.3923/ajpp.2010.295.324
52
Hasanuzzaman M. Hossain M. A. Fujita M. (2011). Nitric oxide modulates antioxidant defense and the methylglyoxal detoxification system and reduces salinity-induced damage of wheat seedlings. Plant Biotechnol. Rep. 5, 353–365. 10.1007/s11816-011-0189-9
53
Hebelstrup K. H. Van Zanten M. Mandon J. Voesenek L. A. Harren F. J. Cristescu S. M. et al . (2012). Haemoglobin modulates NO emission and hyponasty under hypoxia-related stress in Arabidopsis thaliana. J. Exp. Bot. 63, 5581–5591. 10.1093/jxb/ers210
54
Hill B. G. Dranka B. P. Bailey S. M. Lancaster J. R. Jr. Darley-Usmar V. M. (2010). What part of NO don't you understand. Some answers to the cardinal questions in nitric oxide biology. J. Biol. Chem. 285, 19699–19704. 10.1074/jbc.R110.101618
55
Holzmeister C. Frohlich A. Sarioglu H. Bauer N. Durner J. Lindermayr C. (2011). Proteomic analysis of defense response of wildtype Arabidopsis thaliana and plants with impaired NO- homeostasis. Proteomics11, 1664–1683. 10.1002/pmic.201000652
56
Hsu Y. T. Kao C. H. (2007). Toxicity in leaves of rice exposed to cadmium is due to hydrogen peroxide accumulation. Plant Soil298, 231–241. 10.1007/s11104-007-9357-7
57
Hung K. T. Chang C. J. Kao C. H. (2002). Paraquat toxicity is reduced by nitric oxide in rice leaves. J. Plant Physiol. 159, 159–166. 10.1078/0176-1617-00692
58
Innocenti G. Pucciariello C. LeGleuher M. Hopkins J. De Stefano M. Delledonne M. et al . (2007). Glutathione synthesis is regulated by nitric oxide in Medicago truncatula roots. Planta225, 1597–1602. 10.1007/s00425-006-0461-3
59
Jasid S. Simontacchi M. Bartoli C. G. Puntarulo S. (2006). Chloroplasts as a nitric oxide cellular source. Effect of reactive nitrogen species on chloroplastic lipids and proteins. Plant Physiol. 142, 1246–1255. 10.1104/pp.106.086918
60
Jin J.-W. Xu Y.-F. Huang Y.-F. (2010). Protective effect of nitric oxide against arsenic-induced oxidative damage in tall fescue leaves. Afr. J. Biotechnol. 9, 1619–1627. ISSN: 1684-5315
61
Joudoi T. Shichiri Y. Kamizono N. Akaike T. Sawa T. Yoshitake J. et al . (2013). Nitrated cyclic GMP modulates guard cell signaling in Arabidopsis. Plant Cell25, 558–571. 10.1105/tpc.112.105049
62
Jourd'heuil D. Jourd'heuil F. L. Kutchukian P. S. Musah R. A. Wink D. A. Grisham M. B. (2001). Reaction of superoxide and nitric oxide with peroxynitrite. Implications for peroxynitrite-mediated oxidation reactions in vivo. J. Biol. Chem. 276, 28799–28805. 10.1074/jbc.M102341200
63
Kato H. Takemoto D. Kawakita K. (2012). Proteomic analysis of S-nitrosylated proteins in potato plant. Physiol. Plantarum. 148, 371–386. 10.1111/j.1399-3054.2012.01684.x
64
Keyster M. Klein A. Ludidi N. (2012). Caspase-like enzymatic activity and the ascorbate-glutathione cycle participate in salt stress tolerance of maize conferred by exogenously applied nitric oxide. Plant Signal. Behav. 7, 349–360. 10.4161/psb.18967
65
Khan A. U. Kovacic D. Kolbanovskiy A. Desai M. Frenkel K. Geacintov N. E. (2000). The decomposition of peroxynitrite to nitroxyl anion (NO-) and singlet oxygen in aqueous solution. Proc. Natl. Acad. Sci. U.S.A. 97, 2984–2989. 10.1073/pnas.97.7.2984
66
Khan M. N. Siddiqui M. H. Mohammad F. Naeem M. (2012). Interactive role of nitric oxide and calcium chloride in enhancing tolerance to salt stress. Nitric Oxide27, 210–218. 10.1016/j.niox.2012.07.005
67
Kirsch M. Lehnig M. Korth H. G. Sustmann R. De Groot H. (2001). Inhibition of peroxynitrite-induced nitration of tyrosine by glutathione in the presence of carbon dioxide through both radical repair and peroxynitrate formation. Chemistry7, 3313–3320. 10.1002/1521-3765(20010803)7:15<3313::AID-CHEM3313>3.0.CO;2-7
68
Koppenol W. H. Bounds P. L. Nauser T. Kissner R. Ruegger H. (2012). Peroxynitrous acid: controversy and consensus surrounding an enigmatic oxidant. Dalton Trans. 41, 13779–13787. 10.1039/c2dt31526b
69
Kumar P. Tewari R. K. Sharma P. N. (2010). Sodium nitroprusside-mediated alleviation of iron deficiency and modulation of antioxidant responses in maize plants. AoB plants2010:plq002. 10.1093/aobpla/plq002
70
Kurz C. R. Kissner R. Nauser T. Perrin D. Koppenol W. H. (2003). Rapid scavenging of peroxynitrous acid by monohydroascorbate. Free Radic. Biol. Med. 35, 1529–1537. 10.1016/j.freeradbiomed.2003.08.012
71
Kwon E. Feechan A. Yun B. W. Hwang B. H. Pallas J. A. Kang J. G. et al . (2012). AtGSNOR1 function is required for multiple developmental programs in Arabidopsis. Planta236, 887–900. 10.1007/s00425-012-1697-8
72
Kytzia A. Korth H. G. Sustmann R. De Groot H. Kirsch M. (2006). On the mechanism of the ascorbic acid-induced release of nitric oxide from N-nitrosated tryptophan derivatives: scavenging of NO by ascorbyl radicals. Chemistry12, 8786–8797. 10.1002/chem.200600405
73
Laspina N. Groppa M. Tomaro M. Benavides M. (2005). Nitric oxide protects sunflower leaves against Cd-induced oxidative stress. Plant Sci. 169, 323–330. 10.1016/j.plantsci.2005.02.007
74
Lee U. Wie C. Fernandez B. O. Feelisch M. Vierling E. (2008). Modulation of nitrosative stress by S-nitrosoglutathione reductase is critical for thermotolerance and plant growth in Arabidopsis. Plant Cell20, 786–802. 10.1105/tpc.107.052647
75
Leitner M. Vandelle E. Gaupels F. Bellin D. Delledonne M. (2009). NO signals in the haze: nitric oxide signalling in plant defence. Curr. Opin. Plant Biol. 12, 451–458. 10.1016/j.pbi.2009.05.012
76
Levine A. Pennell R. I. Alvarez M. E. Palmer R. Lamb C. (1996). Calcium-mediated apoptosis in a plant hypersensitive disease resistance response. Curr. Biol. 6, 427–437. 10.1016/S0960-9822(02)00510-9
77
Levine A. Tenhaken R. Dixon R. Lamb C. (1994). H2O2 from the oxidative burst orchestrates the plant hypersensitive disease resistance response. Cell79, 583–595. 10.1016/0092-8674(94)90544-4
78
Li Q. Y. Niu H. B. Yin J. Wang M. B. Shao H. B. Deng D. Z. et al . (2008). Protective role of exogenous nitric oxide against oxidative-stress induced by salt stress in barley (Hordeum vulgare). Colloids Surf. B Biointerfaces65, 220–225. 10.1016/j.colsurfb.2008.04.007
79
Lin A. Wang Y. Tang J. Xue P. Li C. Liu L. et al . (2012). Nitric oxide and protein S-nitrosylation are integral to hydrogen peroxide-induced leaf cell death in rice. Plant Physiol. 158, 451–464. 10.1104/pp.111.184531
80
Lin C. C. Jih P. J. Lin H. H. Lin J. S. Chang L. L. Shen Y. H. et al . (2011). Nitric oxide activates superoxide dismutase and ascorbate peroxidase to repress the cell death induced by wounding. Plant Mol. Biol. 77, 235–249. 10.1007/s11103-011-9805-x
81
Lindermayr C. Sell S. Muller B. Leister D. Durner J. (2010). Redox regulation of the NPR1-TGA1 system of Arabidopsis thaliana by nitric oxide. Plant Cell22, 2894–2907. 10.1105/tpc.109.066464
82
Liu L. Hausladen A. Zeng M. Que L. Heitman J. Stamler J. S. (2001). A metabolic enzyme for S-nitrosothiol conserved from bacteria to humans. Nature410, 490–494. 10.1038/35068596
83
Liu W. Z. Kong D. D. Gu X. X. Gao H. B. Wang J. Z. Xia M. et al . (2013). Cytokinins can act as suppressors of nitric oxide in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 110, 1548–1553. 10.1073/pnas.1213235110
84
Liu X. Miller M. J. Joshi M. S. Thomas D. D. Lancaster J. R. Jr. (1998). Accelerated reaction of nitric oxide with O2 within the hydrophobic interior of biological membranes. Proc. Natl. Acad. Sci. U.S.A. 95, 2175–2179. 10.1073/pnas.95.5.2175
85
Lozano-Juste J. Colom-Moreno R. Leon J. (2011). In vivo protein tyrosine nitration in Arabidopsis thaliana. J. Exp. Bot. 62, 3501–3517. 10.1093/jxb/err042
86
Lum H. K. Butt Y. K. Lo S. C. (2002). Hydrogen peroxide induces a rapid production of nitric oxide in mung bean (Phaseolus aureus). Nitric Oxide6, 205–213. 10.1006/niox.2001.0395
87
Maassen A. Hennig J. (2011). Effect of Medicago sativa Mhb1gene expression on defense response of Arabidopsis thaliana plants. Acta Biochim. Pol. 58, 427–432.
88
Madej E. Folkes L. K. Wardman P. Czapski G. Goldstein S. (2008). Thiyl radicals react with nitric oxide to form S-nitrosothiols with rate constants near the diffusion-controlled limit. Free Radic. Biol. Med. 44, 2013–2018. 10.1016/j.freeradbiomed.2008.02.015
89
Malik S. I. Hussain A. Yun B. W. Spoel S. H. Loake G. J. (2011). GSNOR-mediated de-nitrosylation in the plant defence response. Plant Sci. 181, 540–544. 10.1016/j.plantsci.2011.04.004
90
Marti M. C. Florez-Sarasa I. Camejo D. Pallol B. Ortiz A. Ribas-Carbo M. et al . (2013). Response of mitochondrial antioxidant system and respiratory pathways to reactive nitrogen species in pea leaves. Physiol. Plant. 147, 194–206. 10.1111/j.1399-3054.2012.01654.x
91
Meiser J. Lingam S. Bauer P. (2011). Posttranslational regulation of the iron deficiency basic helix-loop-helix transcription factor FIT is affected by iron and nitric oxide. Plant Physiol. 157, 2154–2166. 10.1104/pp.111.183285
92
Mittler R. Vanderauwera S. Suzuki N. Miller G. Tognetti V. B. Vandepoele K. et al . (2011). ROS signaling: the new wave. Trends Plant Sci. 16, 300–309. 10.1016/j.tplants.2011.03.007
93
Miyamoto S. Ronsein G. E. Correa T. C. Martinez G. R. Medeiros M. H. Di Mascio P. (2009). Direct evidence of singlet molecular oxygen generation from peroxynitrate, a decomposition product of peroxynitrite. Dalton Trans. 29, 5720–5729. 10.1039/b905560f
94
Moller M. N. Li Q. Lancaster J. R. Jr. Denicola A. (2007). Acceleration of nitric oxide autoxidation and nitrosation by membranes. IUBMB Life59, 243–248. 10.1080/15216540701311147
95
Morot-Gaudry-Talarmain Y. Rockel P. Moureaux T. Quillere I. Leydecker M. T. Kaiser W. M. et al . (2002). Nitrite accumulation and nitric oxide emission in relation to cellular signaling in nitrite reductase antisense tobacco. Planta215, 708–715. 10.1007/s00425-002-0816-3
96
Mur L. A. Kenton P. Lloyd A. J. Ougham H. Prats E. (2008). The hypersensitive response; the centenary is upon us but how much do we know. J. Exp. Bot. 59, 501–520. 10.1093/jxb/erm239
97
Mur L. A. Mandon J. Persijn S. Cristescu S. M. Moshkov I. E. Novikova G. V. et al . (2013). Nitric oxide in plants: an assessment of the current state of knowledge. AoB Plants5:pls052. 10.1093/aobpla/pls052
98
Mur L. A. Sivakumaran A. Mandon J. Cristescu S. M. Harren F. J. Hebelstrup K. H. (2012). Haemoglobin modulates salicylate and jasmonate/ethylene-mediated resistance mechanisms against pathogens. J. Exp. Bot. 63, 4375–4387. 10.1093/jxb/ers116
99
Murgia I. De Pinto M. C. Delledonne M. Soave C. De Gara L. (2004a). Comparative effects of various nitric oxide donors on ferritin regulation, programmed cell death, and cell redox state in plant cells. J. Plant Physiol. 161, 777–783. 10.1016/j.jplph.2003.12.004
100
Murgia I. Tarantino D. Vannini C. Bracale M. Carravieri S. Soave C. (2004b). Arabidopsis thaliana plants overexpressing thylakoidal ascorbate peroxidase show increased resistance to Paraquat-induced photooxidative stress and to nitric oxide-induced cell death. Plant J. 38, 940–953. 10.1111/j.1365-313x.2004.02092.x
101
Neill S. Barros R. Bright J. Desikan R. Hancock J. Harrison J. et al . (2008). Nitric oxide, stomatal closure, and abiotic stress. J. Exp. Bot. 59, 165–176. 10.1093/jxb/erm293
102
Noronha-Dutra A. A. Epperlein M. M. Woolf N. (1993). Reaction of nitric oxide with hydrogen peroxide to produce potentially cytotoxic singlet oxygen as a model for nitric oxide-mediated killing. FEBS Lett. 321, 59–62. 10.1016/0014-5793(93)80621-Z
103
O'brien J. A. Daudi A. Butt V. S. Bolwell G. P. (2012). Reactive oxygen species and their role in plant defence and cell wall metabolism. Planta236, 765–779. 10.1007/s00425-012-1696-9
104
Perazzolli M. Dominici P. Romero-Puertas M. C. Zago E. Zeier J. Sonoda M. et al . (2004). Arabidopsis nonsymbiotic hemoglobin AHb1 modulates nitric oxide bioactivity. Plant Cell16, 2785–2794. 10.1105/tpc.104.025379
105
Planchet E. Jagadis Gupta K. Sonoda M. Kaiser W. M. (2005). Nitric oxide emission from tobacco leaves and cell suspensions: rate limiting factors and evidence for the involvement of mitochondrial electron transport. Plant J. 41, 732–743. 10.1111/j.1365-313X.2005.02335.x
106
Pryor W. A. Houk K. N. Foote C. S. Fukuto J. M. Ignarro L. J. Squadrito G. L. et al . (2006). Free radical biology and medicine: it's a gas, man!American journal of physiology. Regul. Integr. Comp. Physiol. 291, R491–R511. 10.1152/ajpregu.00614.2005
107
Radi R. (2013). Protein tyrosine nitration: biochemical mechanisms and structural basis of functional effects. Acc. Chem. Res. 46, 550–559. 10.1021/ar300234c
108
Romero-Puertas M. C. Campostrini N. Matte A. Righetti P. G. Perazzolli M. Zolla L. et al . (2008). Proteomic analysis of S-nitrosylated proteins in Arabidopsis thaliana undergoing hypersensitive response. Proteomics8, 1459–1469. 10.1002/pmic.200700536
109
Romero-Puertas M. C. Laxa M. Matte A. Zaninotto F. Finkemeier I. Jones A. M. et al . (2007). S-nitrosylation of peroxiredoxin II E promotes peroxynitrite-mediated tyrosine nitration. Plant Cell19, 4120–4130. 10.1105/tpc.107.055061
110
Romero N. Radi R. Linares E. Augusto O. Detweiler C. D. Mason R. P. et al . (2003). Reaction of human hemoglobin with peroxynitrite. Isomerization to nitrate and secondary formation of protein radicals. J. Biol. Chem. 278, 44049–44057. 10.1074/jbc.M305895200
111
Rusterucci C. Espunya M. C. Diaz M. Chabannes M. Martinez M. C. (2007). S-nitrosoglutathione reductase affords protection against pathogens in Arabidopsis, both locally and systemically. Plant Physiol. 143, 1282–1292. 10.1104/pp.106.091686
112
Sakamoto A. Sakurao S. H. Fukunaga K. Matsubara T. Ueda-Hashimoto M. Tsukamoto S. et al . (2004). Three distinct Arabidopsis hemoglobins exhibit peroxidase-like activity and differentially mediate nitrite-dependent protein nitration. FEBS Lett. 572, 27–32. 10.1016/j.febslet.2004.07.005
113
Sakamoto A. Tsukamoto S. Yamamoto H. Ueda-Hashimoto M. Takahashi M. Suzuki H. et al . (2003). Functional complementation in yeast reveals a protective role of chloroplast 2-Cys peroxiredoxin against reactive nitrogen species. Plant J. 33, 841–851. 10.1046/j.1365-313X.2003.01669.x
114
Sakamoto A. Ueda M. Morikawa H. (2002). Arabidopsis glutathione-dependent formaldehyde dehydrogenase is an S-nitrosoglutathione reductase. FEBS Lett. 515, 20–24. 10.1016/S0014-5793(02)02414-6
115
Sakihama Y. Tamaki R. Shimoji H. Ichiba T. Fukushi Y. Tahara S. et al . (2003). Enzymatic nitration of phytophenolics: evidence for peroxynitrite-independent nitration of plant secondary metabolites. FEBS Lett. 553, 377–380. 10.1016/S0014-5793(03)01059-7
116
Sandalio L. M. Rodriguez-Serrano M. Romero-Puertas M. C. Del Rio L. A. (2013). Role of peroxisomes as a source of reactive oxygen species (ROS) signaling molecules. Subcell. Biochem. 69, 231–255. 10.1007/978-94-007-6889-5_13
117
Sang J. R. Jiang M. Y. Lin F. Li J. Xu S. C. (2007). Role of nitric oxide in abscisic acid-induced subcellular antioxidant defense of maize leaves. Zhi Wu Sheng Li Yu Fen Zi Sheng Wu Xue Xue Bao33, 553–566.
118
Saxena I. Shekhawat G. S. (2013). Nitric oxide (NO) in alleviation of heavy metal induced phytotoxicity and its role in protein nitration. Nitric Oxide32, 13–20. 10.1016/j.niox.2013.03.004
119
Schlicht M. Ludwig-Muller J. Burbach C. Volkmann D. Baluska F. (2013). Indole-3-butyric acid induces lateral root formation via peroxisome-derived indole-3-acetic acid and nitric oxide. New Phytol. 200, 473–482. 10.1111/nph.12377
120
Scorza G. Pietraforte D. Minetti M. (1997). Role of ascorbate and protein thiols in the release of nitric oxide from S-nitroso-albumin and S-nitroso-glutathione in human plasma. Free Radic. Biol. Med. 22, 633–642. 10.1016/S0891-5849(96)00378-4
121
Sehrawat A. Abat J. K. Deswal R. (2013). RuBisCO depletion improved proteome coverage of cold responsive S-nitrosylated targets in Brassica juncea. Front. Plant Sci. 4:342. 10.3389/fpls.2013.00342
122
Sheokand S. Kumari A. Sawhney V. (2008). Effect of nitric oxide and putrescine on antioxidative responses under NaCl stress in chickpea plants. Physiol. Mol. Biol. Plants14, 355–362. 10.1007/s12298-008-0034-y
123
Shi H. T. Li R. J. Cai W. Liu W. Wang C. L. Lu Y. T. (2012). Increasing nitric oxide content in Arabidopsis thaliana by expressing rat neuronal nitric oxide synthase resulted in enhanced stress tolerance. Plant Cell Physiol. 53, 344–357. 10.1093/pcp/pcr181
124
Shi S. Wang G. Wang Y. Zhang L. Zhang L. (2005). Protective effect of nitric oxide against oxidative stress under ultraviolet-B radiation. Nitric Oxide13, 1–9. 10.1016/j.niox.2005.04.006
125
Singh H. P. Kaur S. Batish D. R. Sharma V. P. Sharma N. Kohli R. K. (2009). Nitric oxide alleviates arsenic toxicity by reducing oxidative damage in the roots of Oryza sativa (rice). Nitric Oxide20, 289–297. 10.1016/j.niox.2009.02.004
126
Stöhr C. Strube F. Marx G. Ullrich W. R. Rockel P. (2001). A plasma membrane-bound enzyme of tobacco roots catalyses the formation of nitric oxide from nitrite. Planta212, 835–841. 10.1007/s004250000447
127
Sturms R. Dispirito A. A. Hargrove M. S. (2011). Plant and cyanobacterial hemoglobins reduce nitrite to nitric oxide under anoxic conditions. Biochemistry50, 3873–3878. 10.1021/bi2004312
128
Sun X. Li Y. Cai H. Bai X. Ji W. Ding X. et al . (2012). The Arabidopsis AtbZIP1 transcription factor is a positive regulator of plant tolerance to salt, osmotic and drought stresses. J. Plant Res. 125, 429–438. 10.1007/s10265-011-0448-4
129
Tada Y. Spoel S. H. Pajerowska-Mukhtar K. Mou Z. Song J. Wang C. et al . (2008). Plant immunity requires conformational charges of NPR1 via S-nitrosylation and thioredoxins. Science321, 952–956. 10.1126/science.1156970
130
Tanou G. Job C. Belghazi M. Molassiotis A. Diamantidis G. Job D. (2010). Proteomic signatures uncover hydrogen peroxide and nitric oxide cross-talk signaling network in citrus plants. J. Proteome Res. 9, 5994–6006. 10.1021/pr100782h
131
Tanou G. Job C. Rajjou L. Arc E. Belghazi M. Diamantidis G. et al . (2009). Proteomics reveals the overlapping roles of hydrogen peroxide and nitric oxide in the acclimation of citrus plants to salinity. Plant J. 60, 795–804. 10.1111/j.1365-313X.2009.04000.x
132
Tewari R. K. Hahn E. J. Paek K. Y. (2008). Function of nitric oxide and superoxide anion in the adventitious root development and antioxidant defence in Panax ginseng. Plant Cell Rep. 27, 563–573. 10.1007/s00299-007-0448-y
133
Tewari R. K. Prommer J. Watanabe M. (2013). Endogenous nitric oxide generation in protoplast chloroplasts. Plant Cell Rep. 32, 31–44. 10.1007/s00299-012-1338-5
134
Thomas D. D. Ridnour L. A. Isenberg J. S. Flores-Santana W. Switzer C. H. Donzelli S. et al . (2008). The chemical biology of nitric oxide: implications in cellular signaling. Free Radic. Biol. Med. 45, 18–31. 10.1016/j.freeradbiomed.2008.03.020
135
Tian X. Lei Y. (2006). Nitric oxide treatment alleviates drought stress in wheat seedlings. Biol. Plant. 50, 775–778. 10.1007/s10535-006-0129-7
136
Tiso M. Tejero J. Kenney C. Frizzell S. Gladwin M. T. (2012). Nitrite reductase activity of nonsymbiotic hemoglobins from Arabidopsis thaliana. Biochemistry51, 5285–5292. 10.1021/bi300570v
137
Torres M. A. Dangl J. L. Jones J. D. (2002). Arabidopsis gp91phox homologues AtrbohD and AtrbohF are required for accumulation of reactive oxygen intermediates in the plant defense response. Proc. Natl. Acad. Sci. U.S.A. 99, 517–522. 10.1073/pnas.012452499
138
Verma K. Mehta S. Shekhawat G. (2013). Nitric oxide (NO) counteracts cadmium induced cytotoxic processes mediated by reactive oxygen species (ROS) in Brassica juncea: cross-talk between ROS, NO and antioxidant responses. Biometals26, 255–269. 10.1007/s10534-013-9608-4
139
Wang Y. Ries A. Wu K. Yang A. Crawford N. M. (2010). The arabidopsis prohibitin gene PHB3 functions in nitric oxide-mediated responses and in hydrogen peroxide-induced nitric oxide accumulation. Plant Cell22, 249–259. 10.1105/tpc.109.072066
140
Wimalasekera R. Villar C. Begum T. Scherer G. F. (2011). COPPER AMINE OXIDASE1 (CuAO1) of Arabidopsis thaliana contributes to abscisic acid- and polyamine-induced nitric oxide biosynthesis and abscisic acid signal transduction. Molecular Plant4, 663–678. 10.1093/mp/ssr023
141
Xu Y. Sun X. Jin J. Zhou H. (2010). Protective effect of nitric oxide on light-induced oxidative damage in leaves of tall fescue. J. Plant Physiol. 167, 512–518. 10.1016/j.jplph.2009.10.010
142
Xue L. Li S. Sheng H. Feng H. Xu S. An L. (2007). Nitric oxide alleviates oxidative damage induced by enhanced ultraviolet-B radiation in cyanobacterium. Curr. Microbiol. 55, 294–301. 10.1007/s00284-006-0621-5
143
Yang L. Tian D. Todd C. D. Luo Y. Hu X. (2013). Comparative proteome analyses reveal that nitric oxide is an important signal molecule in the response of rice to aluminum toxicity. J. Proteome Res. 12, 1316–1330. 10.1021/pr300971n
144
Yun B. W. Feechan A. Yin M. Saidi N. B. Le Bihan T. Yu M. et al . (2011). S-nitrosylation of NADPH oxidase regulates cell death in plant immunity. Nature478, 264–268. 10.1038/nature10427
145
Zeier J. Delledonne M. Mishina T. Severi E. Sonoda M. Lamb C. (2004). Genetic elucidation of nitric oxide signaling in incompatible plant-pathogen interactions. Plant Physiol. 136, 2875–2886. 10.1104/pp.104.042499
Summary
Keywords
nitric oxide, reactive oxygen species, signaling, peroxynitrite, glutathione, ascorbate, antioxidant system, programmed cell death
Citation
Groß F, Durner J and Gaupels F (2013) Nitric oxide, antioxidants and prooxidants in plant defence responses. Front. Plant Sci. 4:419. doi: 10.3389/fpls.2013.00419
Received
28 June 2013
Accepted
01 October 2013
Published
29 October 2013
Volume
4 - 2013
Edited by
John Hancock, University of the West of England, Bristol, UK
Reviewed by
Christine H. Foyer, University of Leeds, UK; Radhika Desikan, Imperial College London, UK
Copyright
© 2013 Groß, Durner and Gaupels.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Frank Gaupels, German Research Center for Environmental Health, Institute of Biochemical Plant Pathology, Helmholtz Zentrum München, Ingolstädter Landstr. 1, Oberschleißheim, D-85764 Munich, Germany e-mail: frank.gaupels@helmholtz-muenchen.de
This article was submitted to Plant Physiology, a section of the journal Frontiers in Plant Science.
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.