- 1International Crops Research Institute for the Semi-Arid Tropics, Hyderabad, India
- 2University School of Biotechnology, Guru Gobind Singh Indraprastha University, Delhi, India
- 3Agronomy Department, University of Florida, Gainesville, FL, USA
- 4Etude de Polymorphisme des Génomes Végétaux, INRA, Evry, France
- 5UMR AGAP, CIRAD, Montpellier Cedex, France
- 6UMR AGAP, Montpellier SupAgro, Montpellier, France
Chickpea is an important food legume crop for the semi-arid regions, however, its productivity is adversely affected by various biotic and abiotic stresses. Identification of candidate genes associated with abiotic stress response will help breeding efforts aiming to enhance its productivity. With this objective, 10 abiotic stress responsive candidate genes were selected on the basis of prior knowledge of this complex trait. These 10 genes were subjected to allele specific sequencing across a chickpea reference set comprising 300 genotypes including 211 genotypes of chickpea mini core collection. A total of 1.3 Mbp sequence data were generated. Multiple sequence alignment (MSA) revealed 79 SNPs and 41 indels in nine genes while the CAP2 gene was found to be conserved across all the genotypes. Among 10 candidate genes, the maximum number of SNPs (34) was observed in abscisic acid stress and ripening (ASR) gene including 22 transitions, 11 transversions and one tri-allelic SNP. Nucleotide diversity varied from 0.0004 to 0.0029 while polymorphism information content (PIC) values ranged from 0.01 (AKIN gene) to 0.43 (CAP2 promoter). Haplotype analysis revealed that alleles were represented by more than two haplotype blocks, except alleles of the CAP2 and sucrose synthase (SuSy) gene, where only one haplotype was identified. These genes can be used for association analysis and if validated, may be useful for enhancing abiotic stress, including drought tolerance, through molecular breeding.
Introduction
Chickpea (Cicer arietinum L., 2n = 16), a self-pollinated, diploid annual species which ranks second worldwide as a food legume crop, is primarily a crop of developing countries contributing to a larger part of human food and animal feed in these areas. Chickpea is a major source of nutrients to a vegetarian diet as it contain 20–30% protein, ~40% carbohydrates and is also a good source of several minerals like calcium, magnesium, potassium, phosphorus, iron, zinc, and manganese. Global chickpea production is 11.6 million t from 12.3 million ha area with an average yield of less than one t/ha (FAO, 2012), much lower than its estimated potential of 6 t/ha under optimum growing conditions. Productivity of chickpea is adversely affected by several abiotic stresses of which drought, heat and cold are the major constraints affecting seed yield (Ruelland et al., 2002). Plant stress responses are generally controlled by a network of specialized genes through intricate regulation by specific transcription factors (Chen and Zhu, 2004). Application of available approaches to improve crop productivity under adverse environmental conditions requires a better understanding of the mechanisms involved during crop's response to abiotic stress. Genomic technologies and comparative genomics approaches that have emerged during the past decade can be exploited to identify some of the genes involved in drought tolerance mechanisms. Candidate genes for stress tolerance may be used in crop improvement programs directly (transgenic approach) or indirectly (through identification of linked SNPs) (Schena et al., 1995; Kudapa et al., 2013). The “chickpea mini core” comprising of 211 diverse genotypes (Upadhyaya and Ortiz, 2001) is a subset of the core collection (Upadhyaya et al., 2001) which represents the entire collection conserved in the ICRISAT Genebank. The reference set (Upadhyaya et al., 2008) includes four C. reticulatum genotypes and three C. echinospermum genotypes, but the majority (293 genotypes) is C. arietinum (Upadhyaya et al., 2006).
Although several genes have been found to be involved in abiotic stress tolerance in other crops, few studies have been carried out in chickpea. Candidate genes can be selected on the basis of prior knowledge from mutational analysis, biochemical pathways or linkage analysis of the trait of interest (Zhu et al., 2008). The candidate genes we selected were; Snf-1 related kinase (AKIN), amino-aldehyde dehydrogenase (AMADH), abscisic acid stress and ripening (ASR) gene, a homolog of the DREB2A gene, known as the CAP2 gene, dehydrin (DHN), drought responsive element binding protein (DREB), ERECTA, Myb transcription factor (MYB), sucrose phosphate synthase (SPS), and sucrose synthase (SuSy).
The AKIN (SNF1 related protein kinase) gene belongs to the CDPK–SnRK superfamily, which serves as important regulators modulating fundamental metabolic pathways in response to nutritional and environmental stresses in plants (Halford and Hey, 2009). An AMADH gene in sorghum was found to be related to osmotic stress tolerance, dehydration and salt stress tolerance (Wood et al., 1996) and the activity of AMADH in response to stress caused by mechanical damage in pea seedlings was evaluated by Petrivalský et al. (2007). AMADH is expected to play a role in physiological processes and metabolic pathways controlling response to abiotic stresses by detoxification of toxic aminoaldehydes (Stiti et al., 2011). ASR gene is a stress-inducible gene that has been reported exclusively in plants and belongs to a small gene family characterized by the presence of an ABA/WDS domain. Members of the ASR gene family are induced by abscisic acid (ABA), various abiotic stresses including water stress and during the process of fruit ripening (Carrari et al., 2004). ASR genes in various species respond to different abiotic stress factors including drought, salt, cold and limited light (Joo et al., 2013). Over-expression of ASR in transgenic Arabidopsis was shown to increase tolerance to drought and salt and decrease sensitivity to exogenous ABA (Yang et al., 2005). Characterization of the ASR gene family in rice identified the ASR3 gene as a candidate for association studies related to drought tolerance (Philippe et al., 2010). The potential importance of the ASR1 gene in drought tolerance in common bean was reported by Cortés et al. (2012a) who found low nucleotide diversity suggestive of strong purifying selection, in wild and cultivated accessions.
Dehydrins (DHNs) are among the most commonly observed proteins induced by environmental stress associated with dehydration or low temperature (Hanin et al., 2011). The DHN proteins have been estimated to comprise up to 4% of the total seed protein, and are thought to be involved in protecting the embryo and other seed tissues from osmotic stresses associated with the low water content of the mature seed (Wise and Tunnacliffe, 2004). A positive correlation between accumulation of DHN proteins and tolerance to freezing, drought, and salinity has been shown (Close, 1996; Allagulova et al., 2003). Transgenic plants overexpressing DHN showed better growth and tolerance to drought and freezing stress compared to controls (Puhakainen et al., 2004). DREB are transcription factors that induce a set of abiotic stress-related genes and impart stress endurance to plants. DREBs belong to the ERF (ethylene responsive element binding factors) clade of the APETALA2 (AP2) family are distinctive to plants. Transcription factors DREB1A/CBF3 and DREB2A were identified as cold and drought stress–responsive genes expressed in Arabidopsis thaliana (Sakuma et al., 2006). Constitutively activated DREB2A resulted in significant drought stress tolerance in transgenic Arabidopsis plants and expression analysis revealed that DREB2A transcriptionally regulates many water stress-inducible genes (Sakuma et al., 2006). In rice, expression of OsDREB2A was induced by dehydration and high-salt stresses (Matsukura et al., 2010; Mallikarjuna et al., 2011). Based on physiological studies in several crop species, the DREB2A transcription factor is one of the most promising candidate genes for drought tolerance. Low sequence diversity of DREB2A was found in five crop species studied; chickpea, common bean, rice, sorghum, and barley (Nayak et al., 2009) as well as in studies of wild and cultivated common bean (Cortés et al., 2012b).
The ERECTA gene codes for a protein kinase receptor which mediates plants' responses to disease, predation and stress. ERECTA is involved in leaf organogenesis and reduces the density of stomata on the leaf under-surface, thereby reducing the evapotranspiration. In Arabidopsis, the ERECTA gene has been shown to control organ growth and flower development by promoting cell proliferation (Shpak et al., 2004). The contribution of ERECTA gene toward water use efficiency was confirmed using complementation assays on wilting mutant Arabidopsis plants (Masle et al., 2005). The ZmERECTA genes from maize are patented by Pioneer Hi-Bred International, Inc., which were involved in improving plant growth, transpiration efficiency and drought tolerance in crop plants (www.freepatentsonline.com/y2008/0078004.html). The Myb transcription factor family constitutes the largest and diverse class of DNA-binding transcription factors in plants (Riechmann et al., 2000). The roles of Myb genes in response to biotic and abiotic stress have been studied in a number of plant species (Romero et al., 1998; Du et al., 2012; Volpe et al., 2013). SuSy, a glycosyltransferase, and SPS are key enzymes involved in sugar metabolism. Sucrose-synthase transcript and protein levels have been shown to be modulated by dehydration and rehydration (Kleines et al., 1999) and the Arabidopsis AtSUS3 gene in particular was shown to be strongly induced by drought and mannitol, thus behaving as a marker of dehydrating tissues (Baud et al., 2004).
Genetic diversity, representing the overall genetic makeup of a species, serves as a basis for a population to adapt to changing environments (Ross-Ibarra et al., 2007). Single nucleotide polymorphisms (SNPs) have gained much popularity in assessing the diversity because of automation and abundance. Though biallelic SNPs are generally less informative than multi-allelic simple sequence repeats (SSRs), their sheer abundance makes the development of high density SNP genetic maps possible, providing the foundation for subsequent population-based genetic analysis (Rafalski, 2002). In addition, a SNP is of great importance if it affects gene function and the function of the gene in stress response is known/understood and the SNP is associated with differences in plant performance. Assessing genetic diversity for stress responsive candidate gene sequences leads to the identification of a specific allele of the particular gene in that species associated with performance in response to a corresponding abiotic stress. Such information can therefore be further used in breeding programs to develop better varieties using modern molecular breeding approaches like marker assisted recurrent selection (MARS) or gene pyramiding. Allelic diversity (richness), one of the most important and commonly used estimators of genetic diversity in populations, mainly depends on the effective population size and past evolutionary history (Petit et al., 1998). However, the number of alleles identified and their frequency distribution also depend on the genetic marker system used in these investigations. In the present study, the allelic diversity of candidate genes for abiotic stress tolerance was assessed in the chickpea reference set.
Materials and Methods
Plant Material and DNA Extraction
Young leaf tissues of each accession of the reference set from the greenhouse grown plants were harvested and immediately stored in 96-well plate and the total genomic DNA of all the genotypes was isolated using high-throughput mini- DNA extraction method (Cuc et al., 2008). The quality and quantity of extracted DNA was checked on 0.8% agarose gel. The DNA was normalized to 20 ng/μl concentration for further use.
Identification of Abiotic Stress Responsive Genes and Primer Designing
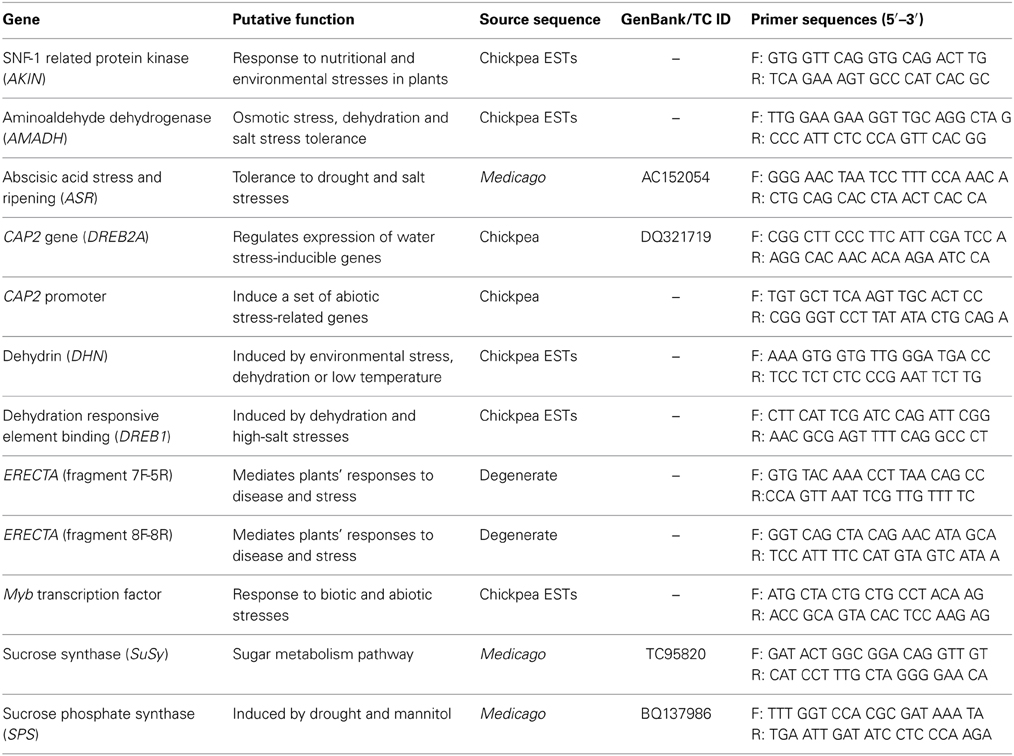
A set of 10 abiotic stress responsive genes conferring abiotic stress tolerance in model plants (Arabidopsis and Rice) and other crop species (Glycine max and Medicago spp.) were chosen based on available literature (Table 1). Different approaches were used for primer designing based on availability of gene sequence information in chickpea. In the first approach, heterologous primers were designed for ASR, SuSy, and SPS genes from corresponding Medicago sequences. The ERECTA gene in chickpea was isolated using consensus/degenerate primers designed at INRA, EPGV, France. In the second approach sequence-specific primers were designed, where in chickpea homologs of genes were isolated using chickpea ESTs developed for abiotic stress (Varshney et al., 2009) and available in NCBI EST database (DbEST- http://www.ncbi.nlm.nih.gov/dbEST/) (Roorkiwal and Sharma, 2012). The details of primers used in isolation of abiotic stress responsive candidate genes in chickpea are given in Table 1.
Polymerase Chain Reaction (PCR) and Sequencing of Amplicons
In order to amplify these candidate genes and confirm their presence, a pilot experiment was set to sequence amplicons from eight diverse genotypes of chickpea consisting of Annigeri, ICCV 2, ICC 4958, ICC 1882, ICC 283, ICC 8261, ICC 4411, and ICC 10029. PCR was set up with 20 μl reaction mixture comprising 5 ng of template DNA, 5 picomoles each of forward and reverse primers, 2 mM dNTP, 20 mM MgCl2, 1X PCR buffer (AmpliTaq Gold) and 0.25 U of Taq polymerase (Ampli Taq Gold). PCR cycles comprising of denaturation of 94°C for 5 min, followed by 40 cycles of 94°C for 30 s annealing at temperature specific for each target gene for 40 s and 72°C for 1 min 30 s and a final extension was carried out at 72°C for 20 min. The amplified product (about 2 μl) was loaded on 1.2% agarose. The remaining PCR amplicons were purified using 1 unit of Exonuclease I and 1 unit of shrimp alkaline phosphatase (SAP) per 5 μ l of PCR product. The Exo/SAP added PCR products were incubated for 45 min at 37°C followed by denaturing at 80°C for 15 min in the thermal cycler for deactivating unused exonuclease enzyme. The Exo/SAP treated amplicons were mixed with 1 μ l of BigDye Terminator V3.1 (Applied Biosystems, California, USA), 2 μ l of 5X sequencing dilution buffer and 3.2 μ M of primer (forward and reverse, separately) and the volume was made to 10 μ l by adding water. The sequencing PCR profile included an initial denaturation of 96°C for 30 s, followed by 60 cycles of 96°C for 10 s, 50°C for 5 s, and 60°C for 4 min. The PCR products were stored at 4°C until further use. Before sequencing, the PCR products were treated with 2.5 μ l of 125 mM EDTA and 25 μ l of absolute ethanol and incubated for 15 min at room temperature to precipitate the DNA. The plate containing the PCR product was centrifuged at 4000 rpm for 30 min at 4°C. The ethanol/ EDTA mix was poured off by inverting the plate, without losing the pellet. To each well, 60 μ l of 70% ethanol was added and again spun at 4000 rpm for 20 min at 4°C. The ethanol was poured off as earlier. The plate was air-dried and 10 μ l of HiDi formamide (Applied Biosystems, California, USA) was added and the products were denatured (94°C for 5 min, then immediately cooled to 4°C for 5 min) and sequenced using an ABI3700/ABI3130 automated sequencer (Applied Biosystems, California, USA).
Allele Sequencing and SNP Detection
For allele sequencing, of candidate genes across the 300 genotypes of the reference set, PCR and purification were carried out as described above. Sequencing was carried out at MACROGEN, Korea using BigDye terminator cycle sequencing chemistry. Raw sequences were used to obtain contigs by assembling the forward and reverse sequences of each genotype using DNA Baser V 2.9 tool and gene identities were confirmed using BLAST (blastn and blastx). The sequences of each candidate gene were aligned using CLUSTALW (http://www.ebi.ac.uk/Tools/clustalw2/index.html). Multiple sequence alignment (MSA) files and fasta files were further used for identifying equence related parameters such as number of genotypes sequenced; length of sequences; number of indels; indel frequency; number of SNPs and their types (transition or transversion); SNP frequency; nucleotide and haplotype diversity and polymorphic information content (PIC) of SNPs and haplotypes using an in-house tool developed at ICRISAT called “DIVersity ESTimator” module (DIVEST) (Jayashree et al., 2009). Further, in order to identify if any of the haplotypes could be associated with the country of origin of the genotypes under study, NETWORK programme version 4.516 was used to determine haplotype networks for each candidate gene studied.
Results
Isolation and Sequence Analysis of Abiotic Stress Responsive Candidate Genes
An AKIN homolog was amplified using the gene specific primer pair designed considering unigene sequence showing match with Arabidopsis AKIN (SNF-1 related protein kinase). The approximate amplicon size of AKIN was ~800 bp. Amplification of an AMADH homolog yielded a product of ~900 bp. The ABA stress and ripening (ASR) gene was isolated using the heterologous primers derived from Medicago sequence AC152054. A single amplicon of 700 bp was obtained for the chickpea genotypes used. A DREB2A homolog (also known as CAP2 gene) and its promoter (CAP2 promoter) were amplified using a primer pair as described by Nayak et al. (2009). The approximate amplicon size of the CAP2 gene was 1000 bp while the CAP2 promoter was ~700 bp. A dehydrin homolog of chickpea was amplified using a primer pair designed for known dehydrin gene using chickpea unigene. The approximate amplicon size of dehydrin gene was ~380 bp. A DREB1 (Dehydration response element binding) homolog in chickpea was also amplified using a primer pair designed using unigene showing match against DREB1 gene. The approximate amplicon size of the DREB1 gene was ~800 bp. About 4300 bp long ERECTA gene fragments were isolated from eight chickpea genotypes using consensus primers. An ~350 bp long MYB gene was amplified using unigene sequence having match against Glycine max Myb transcription factor. For isolating the SuSy gene in chickpea, heterologous primers were designed from Medicago sequences TC95820 (homolog to SUS2 Pea) and AJ131964 (Medicago truncatula SUS1 gene). An ~1500 bp amplicon was obtained for TC95820- derived sequences, while a 900 bp amplicon was obtained with AJ131964- derived sequences. Heterologous primers designed using Medicago sequence BQ137986 and CB893717 were used to isolate SPS in chickpea. Amplification across eight genotypes in chickpea yielded products of 400 bp in both cases (Table 2).
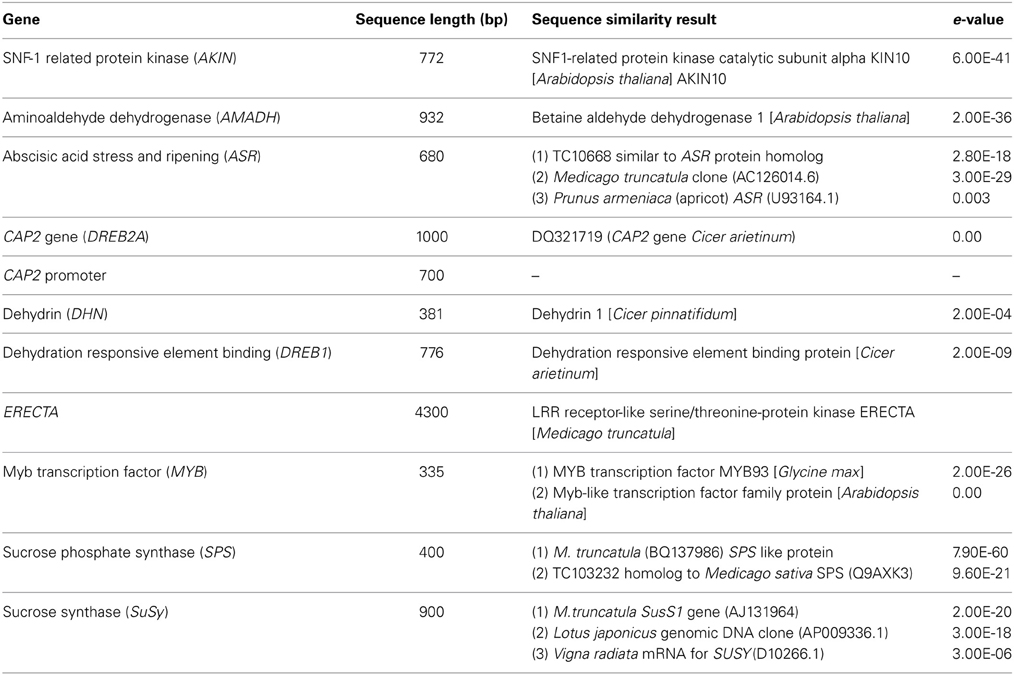
Table 2. Summary of abiotic stress responsive candidate genes showing match with previously reported accession/gene in other crop species.
Sequence Diversity Analysis of Candidate Genes
Forward and reverse sequences for all 10 abiotic stress responsive candidate genes and the CAP2 gene promoter, were used for contig construction. The number of genotypes for which good quality sequences were obtained varied from 79 (ERECTA fragment obtained from 7f-5r primer pairs) to 236 genotypes (SPS gene), out of the 300 genotypes. Diversity analysis of the candidate genes using the DIVersity ESTimator (DIVEST) tool is presented in Table 3.
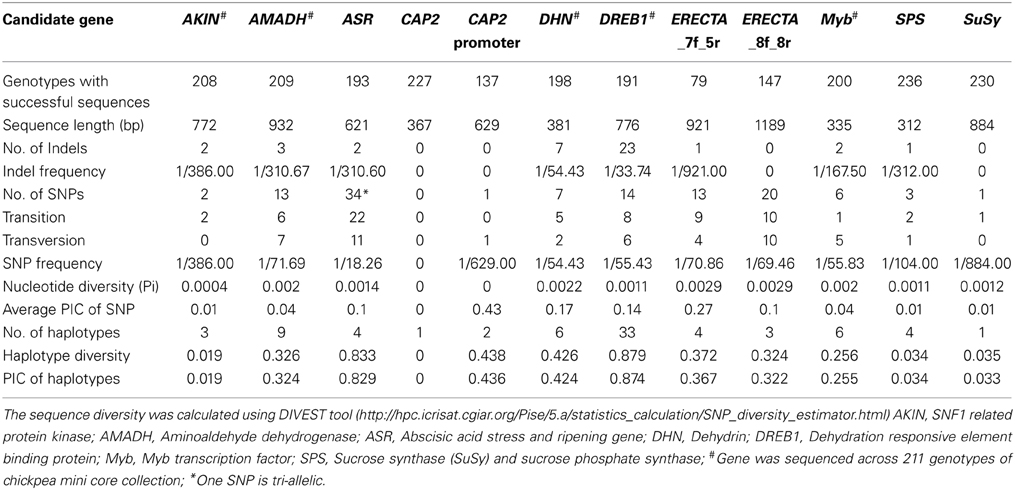
Table 3. Estimation of sequence diversity in chickpea reference set/mini core collection using 10 abiotic stress responsive genes.
SNPs were manually inspected for possible sequencing errors and only those SNPs with clear peaks were considered further (Figure 1A). Sequences for each gene were aligned using CLUSTALW and positions of SNPs were identified (Figure 1B). The highest number of SNPs (34) was obtained for the ASR gene, amongst which 22 were transitions, 11 were transversions and one was tri-allelic. Apart from SNPs, two indels were also detected. The CAP2 gene was found to be conserved across all 227 genotypes with no SNPs and indels. In the case of CAP2 promoter, one SNP was found (which was the same observed when eight chickpea genotypes were sequenced as a pilot experiment). For the ERECTA gene, two fragments obtained from 7f-5r and 8f-8r primer pairs were sequenced. In total, 13 SNPs (9 transitions and 4 transversions) and one indel were obtained for ERECTA 7f-5r fragments while 20 SNPs (10 transitions and 10 transversions) were observed for ERECTA 8f-8r gene fragments. One indel and 3 SNPs were observed across SPS gene sequences. The AKIN gene showed the presence of two SNPs and two indels. A total of 13 SNPs (6 transitions and 7 transversions) and 3 indels were identified in the AMADH gene, while in the DHN gene 7 SNPs (five transitions and two transversions) were identified among 198 sequences analyzed. For the MYB gene only 6 SNPs (one transition and five transversions) and 2 indels were found in 200 Myb sequences under study. No nucleotide diversity was observed for the CAP2 gene and promoter while in the case of AKIN it was 0.0004 and 0.0029 for both ERECTA fragments. The average polymorphic information content (PIC) value of SNPs ranged from 0 (CAP2 gene) to 0.43 (CAP2 promoter). Haplotype diversity ranged from 0.019 (AKIN) to 0.879 (DREB1). Average (PIC) of haplotypes values ranged from 0.019 (AKIN) to 0.874 (DREB1) (Table 3).
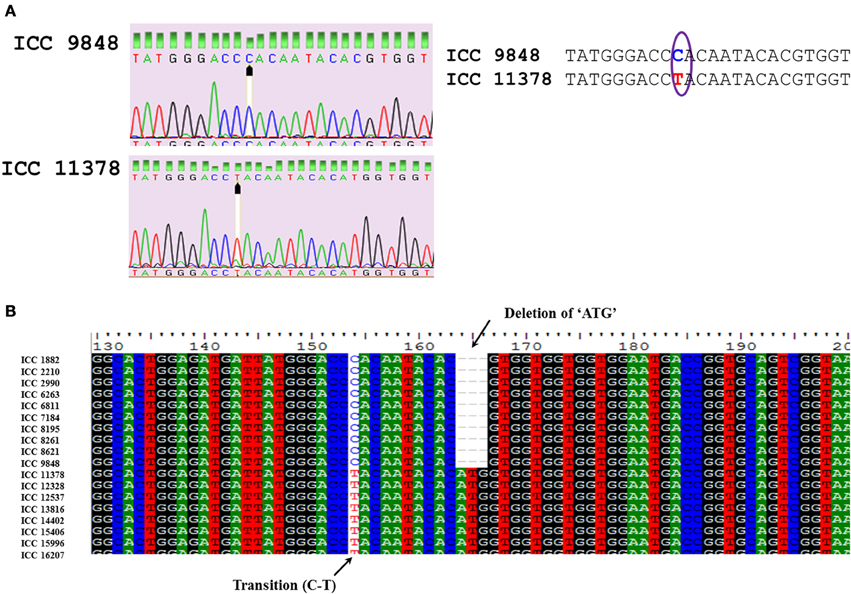
Figure 1. (A) Comparison of sequence quality to confirm the true SNP using peak quality. The presence of SNP in DHN gene using sequence chromatogram is highlighted. (B). Alignment of nucleotide sequences encoding DHN across various chickpea genotypes.
Haplotype Networks for Candidate Genes
Based on the sequence information, haplotype networks were drawn using the NETWORK program. The network figures show the number of haplotypes observed for each gene and the SNP position which separates one haplotype from the other. Network diagrams can be drawn only with the presence of more than two haplotype blocks. Haplotype frequency is depicted by circles, for example, the larger the haplotype circle, more genotypes are represented by that haplotype. The color code is given as per the country of origin of the genotypes (Figures 2A–I). CAP2 and SuSy gene represented only one haplotype with all the genotypes sequenced while the CAP2 promoter had only one SNP, forming two haplotype blocks. Hence haplotype network graphs could not be drawn for CAP2 gene, its promoter and SuSy gene. The network analysis showed a linear relationship between haplotypes for most of the genes except for transcription factors DREB1 and Myb, which showed network relationships between larger numbers of haplotypes.
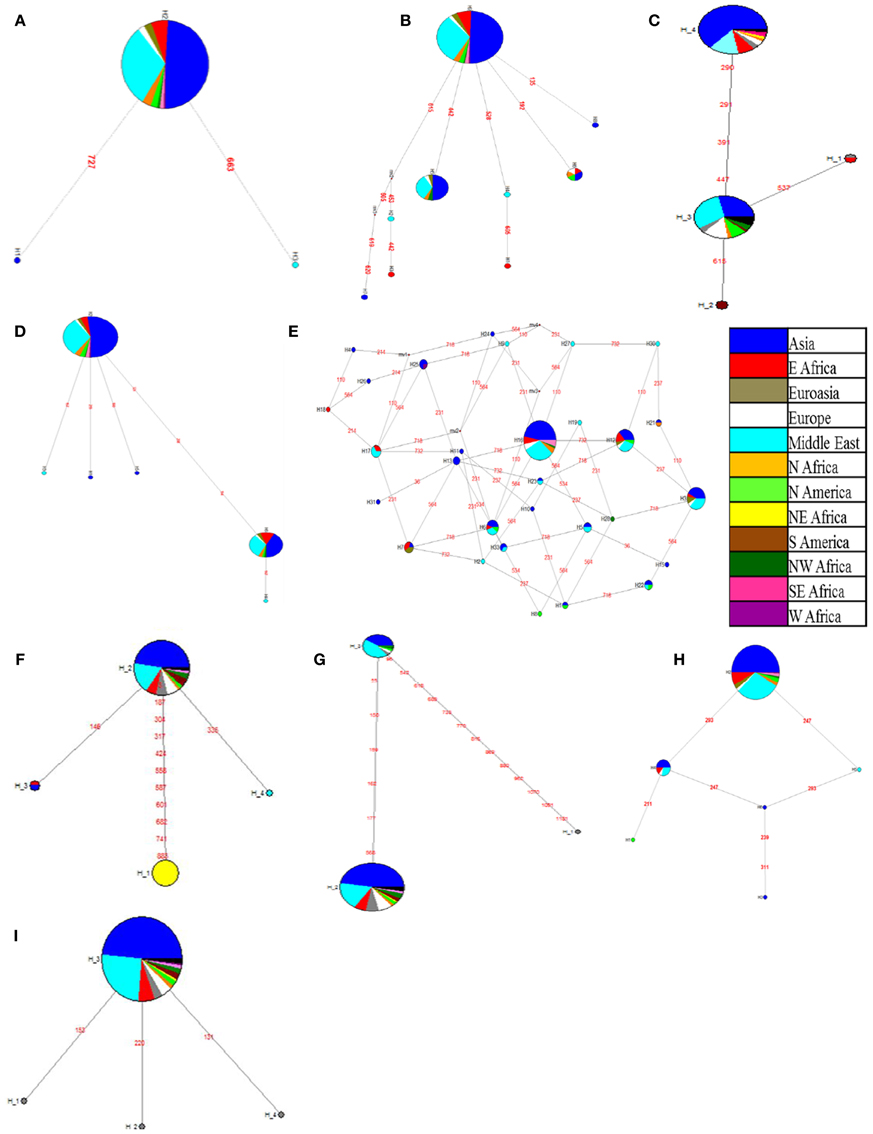
Figure 2. Haplotype network of candidate genes developed based on country of origin of genotypes of the chickpea reference set. (A) AKIN gene; (B) AMADH gene; (C) ASR gene; (D) DHN gene; (E) DREB1 gene; (F) ERECTA (7f-5r) gene; (G) ERECTA (8f-8r) gene; (H) MYB gene; (I) SPS gene; Each circle represents a haplotype and is labeled accordingly. Colors in the circles represent the countries of origin of chickpea genotypes. Circle size is in proportion to frequency (the larger the circle the more genotypes in the haplotype). Numbers in red represent the position of mutations separating the haplotypes.
In this study, although we could find more than two haplotype blocks in some of the candidate genes like AKIN, AMADH, ASR, DHN, DREB, MYB, SPS, ERECTA (7f-5r), and ERECTA (8f-8r), there was no clear distinction between the origin of the genotypes and the haplotype information. Haplotype network analysis for the AKIN gene reported three haplotypes, including one major (H2) and two minor haplotypes (H1 and H3) (Figure 2A). The AMADH gene showed the presence of nine haplotypes across the reference set of which, one major haplotype (H9) is connected to eight other haplotypes (Figure 2B). There were three minor haplotypes (H1, H2, and H4) derived from a major haplotype (H3) as observed in ASR haplotype networks with SNPs ranging from one to four (Figure 2C). DHN gene haplotype network indicated the presence of six haplotypes, of which one major haplotype (H2) was connected to three minor haplotypes (H1, H3, and H5) with one SNP and another haplotype (H6) with three SNPs which was further connected to one minor haplotype (H4) with one SNP (Figure 2D). The DREB1 gene exhibits a complex haplotype network owing to the presence of 33 different haplotypes, which were connected to each other with 1–4 SNPs (Figure 2E). Three major haplotypes (H3, H12, and H16) covers 17, 19, and 62 individuals respectively (Figure 2E). Similarly, in ERECTA- 7f-5r gene fragment, one major haplotype (H1) defined by 10 SNPs and two minor haplotypes (H3 and H4) defined by single SNP were derived from major haplotype H2 (Figure 2F). In the case of the other ERECTA fragment (8f-8r) two haplotypes (H1 and H2) derived from H3 with 6 and 13 SNPs respectively (Figure 2G). Haplotype network of Myb gene showed the presence of six haplotypes, of which two major haplotypes (H2 and H4) are connected to four minor haplotypes with 1–2 SNPs (Figure 2H). SPS gene haplotype network showed presence of three minor haplotypes (H1, H2, and H4) derived from H3 with single nucleotide variation (Figure 2I). Accessions representing each haplotype were color coded according to their country of origin. In the present study, accessions in the major haplotypes were coming from Asia and Middle East in all the genes. The haplotype for ERECTA 7f-5r is unique to NE Africa. The network analysis showed linear relation between haplotypes in most of the genes except for DREB1 and Myb, which are transcription factors. It is also interesting to note that these are the transcription factors which regulate many downstream genes in plant system.
Discussion
The present study was initiated with the objective of the identification of favorable alleles in abiotic stress responsive genes in the chickpea reference set. These gene-based SNPs may be used to identify the suitable allele of a gene that enable the plant to survive in a stress environment. Due to lack of genome sequence information of the chickpea genome until recently (Varshney et al., 2013), identification of genes responsible for complex traits like drought tolerance was a daunting task at the time of initiation of this study. Identification of candidate genes responsible for drought tolerance was a part of an international collaborative project funded by the Generation Challenge Programme (GCP) entitled “Allelic Diversity at Orthologous Candidate genes (ADOC) in seven GCP crops”- one among them was chickpea. An extensive literature survey was carried out to identify possible candidate genes responsible for abiotic stress tolerance, which might have a consensus role in abiotic stress tolerance mechanism in model crops and other legume crops.
Most of the genes analyzed here, have not been previously studied in chickpea. Therefore, systematic efforts by using comparative genomics and bioinformatics approaches were made to determine the corresponding gene sequences in chickpea. For instance, a DREB homolog of chickpea was isolated by using sequence information available from chickpea. As Medicago truncatula is the known taxonomic ally of chickpea, the genomic information about Medicago was searched from different databases including NCBI, TIGR, and Medicago sequence repository (www.medicago.org). Putative candidate genes in chickpea namely ASR, SuSy and SPS were isolated using respective sequence information obtained from the Medicago candidate gene sequences. In addition, the remaining abiotic stress responsive genes (AKIN, AMADH, DHN, and MYB) were identified using a sequence similarity approach against the homolog genes present in model crops like Arabidopsis and Medicago. A large body of evidence demonstrated that the Snf1-related protein kinases (AKIN) serve as important regulators modulating fundamental metabolic pathways in response to nutritional and environmental stresses in yeast and mammalian cells (Hardie, 2007). To identify the AKIN homolog, chickpea ESTs were used for designing the primers for PCR amplification in eight chickpea genotypes based on sequence similarity with Arabidopsis thaliana (Table 2). Researchers have isolated the AKIN homolog in various plant species including Arabidopsis, wheat, rice, potato and tobacco and have established their role in abiotic stress response (Coello et al., 2012). The AKIN gene encodes two types of domains, catalytic kinase (highly conserved) domain and regulatory domain (highly divergent). In the present study, the AKIN gene was found to be mostly conserved except two unique alleles each reported in specific genotype, which indicates that in the present study we were able to amplify the conserved part of AKIN gene, i.e., catalytic kinase. Researchers can target the divergent regulatory domain to identify the SNPs actively involved in abiotic stress response. Similarly, a protective/curative role of the AMADH gene in response to stress events caused by mechanical injury was reported by Petrivalský et al. (2007) in pea seedlings. Since AMADHs works on degradation of reactive metabolites that show considerable toxicity, this enzyme was thought to serve as a detoxification enzyme. An AMADH homolog was amplified using primers designed from chickpea ESTs and BLASTN analysis confirmed its presence (Table 3). Over expression of the AMADH genes from Arabidopsis have been shown to affect stress responses (Missihoun et al., 2011). Based on various functional and characterization studies of the AMADH gene in rice, Arabidopsis and other crop species (Skibbe et al., 2002; Tsuji et al., 2003) makes this gene a suitable candidate for studying its similar role in chickpea. In our study, AMADH showed the second highest number of SNPs (13) across the chickpea mini core collection.
Expression of the ASR gene is regulated by water stress, salt stress and plant hormone ABA. Over-expression of the ASR gene in transgenic plants is known to induce water- and salt- stress tolerance (Kalifa et al., 2004). Although ASR gene function is not published in the case of Medicago, ASR-like sequences that were similar to some of the reported ASR sequences in other crops were used to design primers and amplified in chickpea. The sequence diversity across chickpea genotypes (193 sequences) showed 34 SNPs and two indels, highest among the candidate genes studied in the present study. The nucleotide diversity was found to be 0.0014 while haplotype diversity was 0.833. Cortés et al. (2012b) also analyzed the diversity of two ASR genes in a set of wild and cultivated beans and found two contrasting diversity patterns, most particularly for wild beans. A similar study in rice was carried out, where the polymorphism of four members of the ASR gene family was studied in a worldwide collection of 204 accessions of Oryza sativa and 14 accessions of wild relatives (O. rufipogon and O. nivara). This study provided a thorough description of the organization of the ASR family, and the nucleotide and haplotype diversity of four ASR genes in O. sativa (Philippe et al., 2010).
The chickpea CAP2 gene (a homolog of DREB2A) and its promoter, known to enhance tolerance to dehydration and salt stress, were isolated, characterized and expression studies were carried out in transgenic tobacco (Shukla et al., 2006). The sequence information was used to design nested primers in order to isolate the full-length CAP2 gene during the present study. The study also showed extreme conservation of the AP2 domain of the DREB2 genes across five species studied (Nayak et al., 2009). DREB transcription factors bind to the dehydration responsive element (DRE) of the genes at the promoter region and regulate the expression of downstream genes. The DRE containing core sequence A/GCCGAC was identified as a cis-acting promoter element, which regulates gene expression in response to drought, high salinity and cold stresses in Arabidopsis (Yamaguchi-Shinozaki and Shinozaki, 1994). The CAP2 gene and its promoter were sequenced in 300 diverse chickpea genotypes. The occurrence of a SNP within a regulatory region, accounting for the loss of function of a seed shattering gene has been already shown in rice, which indicates that single sequence variants can cause major effects on the function of gene(s) (Konishi et al., 2006). Conservation of the AP2 domain of the DREB2A gene was observed, not only within chickpea sequences, but also across other crop species; common bean, rice, sorghum and barley (Nayak et al., 2009). DREB2A diversity analysis in common bean (Cortés et al., 2012a) revealed a very high diversity level compared to DREB2B in these other species, indicative of adaptive selection and population expansion.
The DHNs are one of the several proteins that have been specifically associated with qualitative and quantitative changes in cold hardiness (Close, 1996). Arabidopsis plants engineered for DHN over-expression, showed improved survival when exposed to low temperature (Puhakainen et al., 2004). Similarly, transgenic tobaccos with increased level of expression of a citrus dehydrin protein have shown tolerance to low temperature (Hara et al., 2003) making DHN a suitable candidate gene for study in chickpea. Researchers have distinguished five different DHN genes in silico, which could be grouped into two types-K2 and SKn. Three of the dehydrin genes reported several sequence variants which differ by multiple or single amino acid substitutions (Velasco-Conde et al., 2012). The role of ERECTA genes in drought tolerance pertains to their involvement in stomatal density and evapotranspiration (Shpak et al., 2004; Masle et al., 2005). Two fragments of ERECTA genes were isolated in the present study. In chickpea, a total of 33 SNPs (13 from fragment obtained from ERECTA-7f-5r and 20 from fragment obtained from ERECTA-8f-8r) making 7 haplotypes (4 in ERECTA-7f-5r and 3 in ERECTA-8f-8r) were observed. Nucleotide diversity was found to be 0.0029 which was high compared to all other candidate genes under study. The sequence diversity studies across the reference set of chickpea, provides the insights regarding existing haplotypes, which could be involved in drought tolerance mechanism. The role of plant Myb-proteins has been well characterized by using different genetic approaches. In most of the cases, the Myb domain binds to a specific DNA sequence (C/TAACG/TG) to facilitate transcriptional activation (Biedenkapp et al., 1988). A rice R2R3-type MYB transcription factor gene, JAmyb, whose overexpression causes tolerance to high salinity has been identified (Yokotani et al., 2013).
The SuSy and SPS genes encode for the enzymes involved in sugar metabolism and are known to be up-regulated in dehydration stress. The SuSy gene in chickpea is also associated with increased seed size (Kumar and Turner, 2009). A partial SuSy gene was isolated here, and sequencing discovered only 1 SNP across the chickpea reference set. The SuSy gene is a candidate gene for drought tolerance in many plant species (Gonzalez et al., 1995; Baud et al., 2004), and the SPS gene was found to be involved with drought tolerance in maize (Abdel-latif, 2007) and wheat (Fresneau et al., 2007). An SPS homolog was identified in chickpea in the present study. Diversity analysis of this gene on the reference set of chickpea showed the presence of three SNPs and one indel represented as four haplotypes across 235 chickpea genotypes. This observation indicates the conservation of this gene across chickpea genotypes. Studies on sequence diversity on the SPS gene are limited to date. Sequence diversity of an SPS gene was studied for two cultivars of sugarcane and 10 SNPs were identified in a 400 bp sequenced region. These SNPs were screened on a mapping population derived from the two cultivars. The SNP frequency did not vary in the two bulked DNA samples, suggesting that SNPs from this SPS gene family are not associated with variation in sucrose content. Estimation of genetic diversity serves many purposes concerning breeder's interest, like identification of distinct genetic groups for retention in germplasm, identification of genes responsible for phenotypic variation accrued during domestication (Ross-Ibarra et al., 2007) and inference of crop evolution. Allelic diversity studied through NETWORK indicated the distribution of different alleles across the globe based on the origin of the accessions. For some genes (ex: ERECTA 7F-5r), haplotypes identified were coming from particular geographic area (ex: H1 from NE Africa). Such haplotypes indicate a historical constraint as a result of selection, domestication or adaptation. In rice, a haplotype study of three genes revealed the difference in domestication pattern of cultivated and wild rice cultivars (Londo et al., 2006; Kovach et al., 2007). In the present study, linear haplotype networks were found in all genes except for transcription factors DREB1 and Myb. Diversity of transcription factors at a sequence and functional level may affect downstream genes and their expression. Knowledge about genetic diversity and relationships within the diverse germplasm is also useful for breeders as it facilitates their decisions on the selection of the parents for hybridization when widening the genetic basis of breeding programs. Molecular variation in the germplasm can help in the selection of superior genotypes for the generation of new varieties for several agronomic traits. A total of 114 SNPs and 41 indels have been identified in these abiotic stress responsive genes across the chickpea reference set. These SNPs and indels were used for diversity estimation using DIVersity ESTimator (DIVEST). Among the 114 SNPs detected, 66 SNPs regions were transitions, whereas the other 49 were transversions, and one SNP was reported tri-allelic. The nucleotide diversity across the chickpea mini core collection ranged from 0.0004 to 0.0022 with overall mean diversity of 0.0015. The possibilities of association mapping can be explored further by linking sequence diversity with the phenotype diversity in order to identify favorable alleles or haplotypes conferring drought tolerance in chickpea.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study was funded by grants from CGIAR Generation Challenge Programme (GCP), Mexico and Department of Biotechnology (DBT), Government of India. Authors are thankful to Dr. Julie Hoffer for her comments/suggestions to improve the MS. This work has been undertaken as part of the CGIAR Research Program on Grain Legumes. ICRISAT is a member of CGIAR Consortium. Thanks are also due to several colleagues at ICRISAT, GGSIPU and partners in collaborating centers.
References
Abdel-latif, A. (2007). Response of maize leaf sucrose phosphate synthase to salinity. Res. J. Agri. Biol. Sci. 3, 930–933.
Allagulova, C. R., Gimalov, F. R., Shakirova, F. M., and Vakhitov, V. A. (2003). The plant dehydrins: structure and putative functions. Biochemistry (Mosc.) 68, 1157–1165. doi: 10.1023/A:1026077825584
Baud, S., Vaultier, M. N., and Rochat, C. (2004). Structure and expression profile of the sucrose synthase multigene family in Arabidopsis. J. Exp. Bot. 55, 397–409. doi: 10.1093/jxb/erh047
Biedenkapp, H., Borgmeyer, U., Sippel, A. E., and Klempnauer, K. H. (1988). Viral myb oncogene encodes a sequence-specific DNA-binding activity. Nature 335, 835–837. doi: 10.1038/335835a0
Carrari, F., Fernie, A. R., and Iusem, N. D. (2004). Heard it through the grapevine? ABA and sugar cross-talk: the ASR story. Trends Plant Sci. 9, 57–59. doi: 10.1016/j.tplants.2003.12.004
Chen, W. J., and Zhu, T. (2004). Networks of transcription factors with roles in environmental stress response. Trends Plant Sci. 9, 591–596. doi: 10.1016/j.tplants.2004.10.007
Close, T. J. (1996). Dehydrins: emergence of a biochemical role of a family of plant dehydration proteins. Physiol. Plant 97, 795–803. doi: 10.1111/j.1399-3054.1996.tb00546.x
Coello, P., Hirano, E., Hey, S. J., Muttucumaru, N., Martinez-Barajas, E., Parry, M. A., et al. (2012). Evidence that abscisic acid promotes degradation of SNF1-related protein kinase (SnRK) 1 in wheat and activation of a putative calcium-dependent SnRK2. J. Exp. Bot. 63, 913–924. doi: 10.1093/jxb/err320
Cortés, A. J., Chavarro, M. C., Madriñán, S., This, D., and Blair, M. W. (2012a). Molecular ecology and selection in the drought-related Asr gene polymorphisms in wild and cultivated common bean (Phaseolus vulgaris L.). BMC Genet. 13:58. doi: 10.1186/1471-2156-13-58
Cortés, A. J., This, D., Chavarro, C., Madriñán, S., and Blair, M. W. (2012b). Nucleotide diversity patterns at the drought-related DREB2 encoding genes in wild and cultivated common bean (Phaseolus vulgaris L.). Theor. Appl. Genet. 125, 1069–1085. doi: 10.1007/s00122-012-1896-5
Cuc, L. M., Mace, E. S., Crouch, J. H., Quang, V. D., Long, T. D., and Varshney, R. K. (2008). Isolation and characterization of novel microsatellite markers and their application for diversity assessment in cultivated groundnut (Arachis hypogaea). BMC Plant Biol. 8:55. doi: 10.1186/1471-2229-8-55
Du, H., Yang, S. S., Liang, Z., Feng, B. R., Liu, L., Huang, Y. B., et al. (2012). Genome-wide analysis of the MYB transcription factor superfamily in soybean. BMC Plant Biol. 12:106. doi: 10.1186/1471-2229-12-106
FAO. (2012). Available online at: http://faostat.fao.org/site/567/default.aspx#ancor
Fresneau, C., Ghashghaie, J., and Cornic, G. (2007). Drought effect on nitrate reductase and sucrose-phosphate synthase activities in wheat (Triticum durum L.): role of leaf internal CO2. J. Exp. Bot. 58, 2983–2992. doi: 10.1093/jxb/erm150
Gonzalez, E. M., Gordon, A. J., James, C. L., and Arrese-Igor, C. (1995). The role of sucrose synthase in the response of soybean nodules to drought. J. Exp. Bot. 46, 1515–1523. doi: 10.1093/jxb/46.10.1515
Halford, N. G., and Hey, S. J. (2009). Snf1-related protein kinases (SnRKs) act within an intricate network that links metabolic and stress signaling in plants. Biochem. J. 419, 247–259. doi: 10.1042/BJ20082408
Hanin, M., Brini, F., Ebel, C., Toda, Y., Takeda, S., and Masmoudi, K. (2011). Plant dehydrins and stress tolerance: versatile proteins for complex mechanisms. Plant Signal. Behav. 6, 1503–1509. doi: 10.4161/psb.6.10.17088
Hara, M., Terashima, S., Fukaya, T., and Kuboi, T. (2003). Enhancement of cold tolerance and inhibition of lipid peroxidation by citrus dehydrin in transgenic tobacco. Planta 217, 290–298. doi: 10.1007/s00425-003-0986-7
Hardie, D. G. (2007). AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat. Rev. Mol. Cell. Biol. 8, 774–785. doi: 10.1038/nrm2249
Jayashree, B., Bhanuprakash, A., Jami, A., Reddy, S. P., Nayak, S., and Varshney, R. K. (2009). Perl module and PISE wrappers for the integrated analysis of sequence data and SNP features. BMC Res. Notes 2:92. doi: 10.1186/1756-0500-2-92
Joo, J., Lee, Y. H., Kim, Y. K., Nahm, B. H., and Song, S. I. (2013). Abiotic stress responsive rice ASR1 and ASR3 exhibit different tissue-dependent sugar and hormone-sensitivities. Mol. Cells 35, 421–435. doi: 10.1007/s10059-013-0036-7
Kalifa, Y., Perlson, E., Gilad, A., Konrad, Z., Scolnik, P. A., and Bar-Zvi, D. (2004). Over-expression of the water and salt stress-regulated Asr1 gene confers an increased salt tolerance. Plant Cell Environ. 27, 1459–1468. doi: 10.1111/j.1365-3040.2004.01251.x
Kleines, M., Elster, R.-C., Rodrigo, M.-J., Blervacq, A.-S., Salamini, F., and Bartels, D. (1999). Isolation and expression analysis of two stress-responsive sucrose-synthase genes from the resurrection plant Craterostigma plantagineum Hochst. Planta 209, 13–24. doi: 10.1007/s004250050602
Konishi, S., Izawa, T., Lin, S. Y., Ebana, K., Fukuta, Y., Sasaki, T., et al. (2006). An SNP caused loss of seed shattering during rice domestication. Science 312, 1392–1396. doi: 10.1126/science.1126410
Kovach, M. J., Sweeney, M. T., and McCouch, S. R. (2007). New insights into the history of rice domestication. Trends Genet. 23, 578–587. doi: 10.1016/j.tig.2007.08.012
Kudapa, H., Ramalingam, A., Nayakoti, S., Chen, X., Zhuang, W., Liang, X., et al. (2013). Functional genomics to study stress responses in crop legumes: progress and prospects. Funct. Plant Biol. 40, 1221–1233. doi: 10.1071/FP13191
Kumar, A., and Turner, N. C. (2009). Growth and sucrose synthase activity of developing chickpea (Cicer arietinum L.) seeds under field conditions. Austr. J. Crop Sci. 3, 20–27.
Londo, J. P., Chiang, Y. C., Hung, K. H., Chiang, T. Y., and Schaal BA. (2006). Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa. Proc. Natl. Acad. Sci. U.S.A. 103, 9578–9583. doi: 10.1073/pnas.0603152103
Mallikarjuna, G., Mallikarjuna, K., Reddy, M. K., and Kaul, T. (2011). Expression of OsDREB2A transcription factor confers enhanced dehydration and salt stress tolerance in rice (Oryza sativa L.). Biotechnol. Lett. 33, 1689–1697. doi: 10.1007/s10529-011-0620-x
Masle, J., Gilmore, S. R., and Farquhar, G. D. (2005). The ERECTA gene regulates plant transpiration efficiency in Arabidopsis. Nature 436, 866–870. doi: 10.1038/nature03835
Matsukura, S., Mizoi, J., Yoshida, T., Todaka, D., Ito, Y., Maruyama, K., et al. (2010). Comprehensive analysis of rice DREB2-type genes that encode transcription factors involved in the expression of abiotic stress-responsive genes. Mol. Genet. Genomics 283, 185–196. doi: 10.1007/s00438-009-0506-y
Missihoun, T. D., Schmitz, J., Klug, R., Kirch, H. H., and Bartels, D. (2011). Betaine aldehyde dehydrogenase genes from Arabidopsis with different sub-cellular localization affect stress responses. Planta 233, 369–382. doi: 10.1007/s00425-010-1297-4
Nayak, S. N., Jayashree, B., Upadhyaya, H. D., Hash, C. T., Kavi Kishor, P. B., Chattopadhyay, D., et al. (2009). Isolation and sequence analysis of DREB2A homologues in three cereals and two legume species. Plant Sci. 117, 460–467. doi: 10.1016/j.plantsci.2009.07.009
Petit, R. J., El Mousadik, A., and Pons, O. (1998). Identifying populations for conservation on the basis of genetic markers. Conserv. Biol. 12, 844–855. doi: 10.1046/j.1523-1739.1998.96489.x
Petrivalský, M., Brauner, F., Luhová, L., Gagneul, D., and Sebela, M. (2007). Aminoaldehyde dehydrogenase activity during wound healing of mechanically injured pea seedlings. J. Plant Physiol. 164, 1410–1418. doi: 10.1016/j.jplph.2007.01.018
Philippe, R., Courtois, B., McNally, K. L., Mournet, P., El-Malki, R., Paslier, M. C. L., et al. (2010). Structure, allelic diversity and selection of Asr genes, candidate for drought tolerance, in Oryza sativa L. and wild relatives. Theor. Appl. Genet. 121, 769–787. doi: 10.1007/s00122-010-1348-z
Puhakainen, T., Hess, M. W., Mäkelä, P., Svensson, J., Heino, P., and Palva, E. T. (2004). Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Mol. Biol. 54, 743–753. doi: 10.1023/B:PLAN.0000040903.66496.a4
Rafalski, A. (2002). Applications of single nucleotide polymorphisms in crop genetics. Curr. Opin. Plant Biol. 5, 94–100. doi: 10.1016/S1369-5266(02)00240-6
Riechmann, J. L., Heard, J., Martin, G., Reuber, L., Jiang, C., Keddie, J., et al. (2000). Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290, 2105–2110. doi: 10.1126/science.290.5499.2105
Romero, I., Fuertes, A., Benito, M. J., Malpica, J. M., Leyva, A., and Paz-Ares, J. (1998). More than 80R2R3-MYB regulatory genes in the genome of Arabidopsis thaliana. Plant J. 14, 273–284. doi: 10.1046/j.1365-313X.1998.00113.x
Roorkiwal, M., and Sharma, P. C. (2012). Sequence similarity based identification of abiotic stress responsive genes in chickpea. Bioinformation 8, 92–97. doi: 10.6026/97320630008092
Ross-Ibarra, J., Morrell, P. L., and Gaut, B. S. (2007). Plant domestication, a unique opportunity to identify the genetic basis of adaptation. Proc. Natl. Acad. Sci. USA 15:104. doi: 10.1073/pnas.0700643104
Ruelland, E., Cantrel, C., Gawer, M., Kader, J. C., and Zachowski, A. (2002). Activation of phospholipases C and D is an early response to a cold exposure in Arabidopsis suspension cells. Plant Physiol. 130, 999–1007. doi: 10.1104/pp.006080
Sakuma, Y., Maruyama, K., Osakabe, Y., Qin, F., Seki, M., Shinozaki, K., et al. (2006). Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18, 1292–1309. doi: 10.1105/tpc.105.035881
Schena, M., Shalon, D., Davis, R. W., and Brown, P. O. (1995). Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470. doi: 10.1126/science.270.5235.467
Shpak, E. D., Berthiaume, C. T., Hill, E. J., and Torii, K. U. (2004). Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131, 1491–1501. doi: 10.1242/dev.01028
Shukla, R. K., Raha, S., Tripathi, V., and Chattopadhyay, D. (2006). Expression of CAP2, an AP2- family transcription factor from chickpea enhances growth and tolerance to dehydration and salt tress in transgenic tobacco. Plant Physiol. 142, 113–123. doi: 10.1104/pp.106.081752
Skibbe, D. S., Liu, F., Wen, T. J., Yandeau, M. D., Cui, X., Cao, J., et al. (2002). Characterization of the aldehyde dehydrogenase gene families of Zea mays and Arabidopsis. Plant Mol. Biol. 48, 751–764. doi: 10.1023/A:1014870429630
Stiti, N., Missihoun, T. D., Kotchoni, S. O., Kirch, H. H., and Bartels, D. (2011). Aldehyde dehydrogenases in Arabidopsis thaliana: biochemical requirements, metabolic pathways, and functional analysis. Front. Plant Sci. 2:65. doi: 10.3389/fpls.2011.00065
Tsuji, G., Fujii, S., Fujihara, N., Hirose, C., Tsuge, S., Shiraishi, T., et al. (2003). Agrobacterium tumefaciens-mediated transformation for random insertional mutagenesis in Colletotrichum lagenarium. J. Gen. Plant Pathol. 69, 230–239. doi: 10.1007/s10327-003-0040-4
Upadhyaya, H. D., Bramel, P. J., and Singh, S. (2001). Development of a chickpea core subset using geographic distribution and quantitative traits. Crop Sci. 41, 206–210. doi: 10.2135/cropsci2001.411206x
Upadhyaya, H. D., Dwivedi, S. L., Baum, M., Varshney, R. K., Udupa, S. M., Gowda, C. L. L., et al. (2008). Genetic structure, diversity, and allelic richness in composite collection and reference set in chickpea (Cicer arietinum L.). BMC Plant Biol. 8:106. doi: 10.1186/1471-2229-8-106
Upadhyaya, H. D., Furman, B. J., Dwivedi, S. L., Udupa, S. M., Gowda, C. L. L., Baum, M., et al. (2006). Development of a composite collection for mining germplasm possessing allelic variation for beneficial traits in chickpea. Plant Genet. Resour. 4, 13–19. doi: 10.1079/PGR2005101
Upadhyaya, H. D., and Ortiz, R. (2001). A mini core subset for capturing diversity and promoting utilization of chickpea genetic resources in crop improvement. Theor. Appl. Genet. 102, 1292–1298. doi: 10.1007/s00122-001-0556-y
Varshney, R. K., Hiremath, P. J., Lekha, P., Kashiwagi, J., Balaji, J., Deokar, A. A., et al. (2009). A comprehensive resource of drought- and salinity- responsive ESTs for gene discovery and marker development in chickpea (Cicer arietinum L.). BMC Genomics 10:523. doi: 10.1186/1471-2164-10-523
Varshney, R. K., Song, C., Saxena, R. K., Azam, S., Yu, S., Sharpe, A. G., et al. (2013). Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat. Biotechnol. 31, 240–246. doi: 10.1038/nbt.2491
Velasco-Conde, T., Yakovlev, I., Majada, J. P., Aranda, I., and Johnsen, Ø. (2012). Dehydrins in maritime pine (Pinus pinaster) and their expression related to drought stress response. Tree Genet. Gen. 8, 957–973. doi: 10.1007/s11295-012-0476-9
Volpe, V., Dell'Aglio, E., Giovannetti, M., Ruberti, C., Costa, A., Genre, A., et al. (2013). An AM-induced, MYB-family gene of Lotus japonicus (LjMAMI) affects root growth in an AM-independent manner. Plant J. 73, 442–455. doi: 10.1111/tpj.12045
Wise, M. J., and Tunnacliffe, A. (2004). POPP the question: what do LEA proteins do? Trends Plant Sci. 9, 13–17. doi: 10.1016/j.tplants.2003.10.012
Wood, A. J., Saneoka, H., Rhodes, D., Joly, R. J., and Goldsbrough, P. B. (1996). Betaine aldehyde dehydrogenase in sorghum. Plant Physiol. 110, 1301–1308. doi: 10.1104/pp.110.4.1301
Yamaguchi-Shinozaki, K., and Shinozaki, K. (1994). A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low temperature, or high salt stress. Plant Cell 6, 251–264. doi: 10.1105/tpc.6.2.251
Yang, C. Y., Chen, Y. C., Jauh, G. Y., and Wang, C. S. (2005). A Lily ASR protein involves abscisic acid signaling and confers drought and salt resistance in Arabidopsis. Plant Physiol. 139, 836–846. doi: 10.1104/pp.105.065458
Yokotani, N., Ichikawa, T., Kondou, Y., Iwabuchi, M., Matsui, M., Hirochika, H., et al. (2013). Role of the rice transcription factor JAmyb in abiotic stress response. J. Plant Res. 126, 131–139. doi: 10.1007/s10265-012-0501-y
Keywords: chickpea, abiotic stress, single nucleotide polymorphism, genetic diversity, candidate genes
Citation: Roorkiwal M, Nayak SN, Thudi M, Upadhyaya HD, Brunel D, Mournet P, This D, Sharma PC and Varshney RK (2014) Allele diversity for abiotic stress responsive candidate genes in chickpea reference set using gene based SNP markers. Front. Plant Sci. 5:248. doi: 10.3389/fpls.2014.00248
Received: 10 January 2014; Accepted: 15 May 2014;
Published online: 05 June 2014.
Edited by:
Mukesh Jain, National Institute of Plant Genome Research, IndiaReviewed by:
David M. Rhoads, California State University, USAShailesh Tripathi, Indian Agricultural Research Institute, India
Copyright © 2014 Roorkiwal, Nayak, Thudi, Upadhyaya, Brunel, Mournet, This, Sharma and Varshney. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Prakash C. Sharma, University School of Biotechnology, Guru Gobind Singh Indraprastha University, AFR 109, A-Block, Dwarka Sec 16C, Delhi 110078, India e-mail:cHJvZi5wY3NoYXJtYUBnbWFpbC5jb20=;
Rajeev K. Varshney, Center of Excellence in Genomics, International Crops Research Institute for the Semi-Arid Tropics, Building No 300, Patancheru, Hyderabad 502324, India e-mail:ci5rLnZhcnNobmV5QGNnaWFyLm9yZw==
†These authors have contributed equally to this work.
 Manish Roorkiwal
Manish Roorkiwal Spurthi N. Nayak
Spurthi N. Nayak Mahendar Thudi
Mahendar Thudi Hari D. Upadhyaya
Hari D. Upadhyaya Dominique Brunel
Dominique Brunel Pierre Mournet5
Pierre Mournet5 Dominique This
Dominique This Prakash C. Sharma
Prakash C. Sharma Rajeev K. Varshney
Rajeev K. Varshney