- 1Southern Crop Protection and Food Research Centre, Agriculture & Agri-Food Canada, London, ON, Canada
- 2Department of Biology, University of Western Ontario, London, ON, Canada
- 3Department of Microbiology and Immunology, University of Western Ontario, London, ON, Canada
As a special phytopathogen, Agrobacterium tumefaciens infects a wide range of plant hosts and causes plant tumors also known as crown galls. The complexity of Agrobacterium–plant interaction has been studied for several decades. Agrobacterium pathogenicity is largely attributed to its evolved capabilities of precise recognition and response to plant-derived chemical signals. Agrobacterium perceives plant-derived signals to activate its virulence genes, which are responsible for transferring and integrating its Transferred DNA (T-DNA) from its Tumor-inducing (Ti) plasmid into the plant nucleus. The expression of T-DNA in plant hosts leads to the production of a large amount of indole-3-acetic acid (IAA), cytokinin (CK), and opines. IAA and CK stimulate plant growth, resulting in tumor formation. Agrobacterium utilizes opines as nutrient sources as well as signals in order to activate its quorum sensing (QS) to further promote virulence and opine metabolism. Intriguingly, Agrobacterium also recognizes plant-derived signals including γ-amino butyric acid and salicylic acid (SA) to activate quorum quenching that reduces the level of QS signals, thereby avoiding the elicitation of plant defense and preserving energy. In addition, Agrobacterium hijacks plant-derived signals including SA, IAA, and ethylene to down-regulate its virulence genes located on the Ti plasmid. Moreover, certain metabolites from corn (Zea mays) also inhibit the expression of Agrobacterium virulence genes. Here we outline the responses of Agrobacterium to major plant-derived signals that impact Agrobacterium–plant interactions.
Introduction
Agrobacterium is a genus of Gram-negative bacteria that uses horizontal gene transfer to cause tumors in many plant species with agricultural and economic importance including woody ornamental shrubs (rose), vines (grape), shade trees, fruit trees (cherry, berry, walnut), and herbaceous perennials. Agrobacterium tumefaciens is the most commonly studied species in this genus. A. tumefaciens causes typical crown-gall diseases. The disease manifests as a tumor-like growth or gall usually at the junction of the root and shoot. Infection by the species Agrobacterium vitis results in cane gall on grapevines while A. rhizogenes causes excessive formation of hairy roots or root tumors. Agrobacterium–plant interaction is an excellent paradigm for studying both plant and bacterial responses, as well as the role of chemical signaling in these processes. A. tumefaciens–plant interaction is now relatively well-understood as a result of significant findings made over the past four decades (for reviews refer to Gelvin, 2003; Brencic and Winans, 2005; McCullen and Binns, 2006; Yuan and Williams, 2012; Pitzschke, 2013). The virulence proficiency of A. tumefaciens is dependent on the presence of the Tumor-inducing (Ti) plasmid, which harbors a Transferred DNA (T-DNA) defined by two direct repeat sequences of approximately 25 base pairs, termed the left and right borders. Most studies have made use of nopaline metabolizing strains C58 and T37 (carrying plasmids pTiC58 and pTiT37, respectively) or the octopine utilizing strain A6 (carrying pTiA6). As a ubiquitous soil bacterium, Agrobacterium is capable of two lifestyles: independent free-living or acting as a pathogen in association with a plant host. When living independently, Agrobacterium virulence is essentially silent. Upon detection of plant-derived signals in the rhizosphere, Agrobacterium activates its chromosomal virulence genes (chv genes) and Ti plasmid encoded virulence genes (vir genes). Vir genes are directly involved in T-DNA cleavage from the Ti plasmid, T-DNA processing, transferring and integration into plant nuclei, conversely, Chv genes are not directly involved in the T-DNA transfer process. Instead, chv genes play important roles in signal transduction necessary for Agrobacterium pathogenicity. Since T-DNA carries genes for the synthesis of indole-3-acetic acid (IAA) and cytokinin (CK; also called oncogenic genes), their expression in plants leads to the production of a large amount of plant hormones that promote uncontrolled cell division and undifferentiated growth of plant tissues, resulting in the formation of a plant tumor and permanent plant genetic transformation.
In addition to genes responsible for IAA and CK production, T-DNA also contains genes for the synthesis of opines (unusual amino acid and sugar condensates). Opines produced by transformed plant cells can be metabolized by Agrobacterium as a source of nutrients. In addition, opines act as signals that activate Agrobacterium quorum sensing (QS). QS is a special form of cell-to-cell communication by which microorganisms synthesize, release, and perceive diffusible signals such as N-(3-oxooctanoyl)-DL-homoserine lactone (3OC8-HSL). QS enables a single cell to sense the number of surrounding cells (cell density) and coordinates their collective behavior. In Agrobacterium, QS plays important roles in interaction with plant hosts, which will be discussed in later sections.
Interestingly, T-DNA encoded oncogenic genes are neither physiologically nor biologically required for the T-DNA transfer process. Therefore, T-DNA encoded genes can be deleted and replaced with genes of interest, and such genetically modified “T-DNA” can still be transferred, integrated and expressed in the plant cell. This unique ability of inter-kingdom DNA transfer makes Agrobacterium an important tool for genetically modifying plants, allowing for incorporation of useful traits like resistance to insects and herbicides, production of recombinant vaccines, proteins, etc. In addition, T-DNA is distally located from vir genes required for T-DNA transfer process. Thus, T-DNA and vir genes can be separated onto two plasmids without affecting T-DNA transfer into plant hosts. This feature prompted the design and construction of binary vectors that greatly facilitate DNA manipulation and plant transformation, especially considering the large size of the Ti plasmid (over 200 kb).
Agrobacterium is capable of infecting/transforming a wide variety of plant species including long-lived woody plants and cultivated plants. However, plants vary greatly in their ability to be infected/transformed by Agrobacterium, even among ecotypes within species, and the underlying molecular mechanisms are poorly understood (Nam et al., 1997). To mount a successful infection in nature, it is important for Agrobacterium to precisely and specifically recognize and respond to a combination of plant-derived signals in the rhizosphere including acidity, plant released sugars and plant-derived phenolic compounds (Stachel et al., 1985; Brencic and Winans, 2005; McCullen and Binns, 2006; Gelvin, 2012; Yuan and Williams, 2012; Pitzschke, 2013). Agrobacterium virulence programing and associations with plant hosts are stringently and synergistically regulated by a combination of plant-derived chemicals.
Agrobacterium Responses to Acidic Signals Caused by Plant-Derived Chemicals in the Rhizosphere
The rhizosphere is the narrow region (within millimeter range of roots) of soil that is directly influenced by root exudates and is densely populated by soil microorganisms. Rhizosphere is rich in not only plant-derived but microbe-derived signals as well (Winans, 1992; Phillips et al., 2004; Bais et al., 2005). Plants routinely secrete organic acids such as lactic, citric, oxalic, and malic acids as well as other secondary metabolites, resulting in acidic rhizosphere conditions (Rivoal and Hanson, 1994; Xia and Roberts, 1994; Walker et al., 2003; Phillips et al., 2004; Bais et al., 2005; Wang et al., 2006; Huckelhoven, 2007; Badri and Vivanco, 2009). Upon wounding, plants release phenolic compounds as well as neutral and acidic sugars necessary to repair damaged tissue acidifying the rhizosphere (Winans, 1992). Therefore the rhizosphere, where Agrobacterium primarily infects plant hosts, is typically an acidic niche driven by various plant-released chemicals.
Upon close proximity to a suitable plant host in the rhizosphere, acidic conditions and plant-derived chemicals play important roles in initiating the Agrobacterium virulence program, which involves various Agrobacterium regulatory factors and signaling pathways (Winans, 1992). A chromosomally encoded che cluster (chemotaxis) allows A. tumefaciens to be attracted to plant-derived chemicals in the rhizosphere (Wright et al., 1998). In addition, three Agrobacterium chromosomally encoded genes chvA, chvB, and exoC are involved in synthesis of extracellular oligosaccharides, such as cyclic 1,2-b-D-glucan, that allows Agrobacterium to attach to plant hosts (Cangelosi et al., 1989). Upon perception of acidity characteristic of the rhizosphere, Agrobacterium mounts both a conserved response as well as a signaling specific response to infect plant hosts. This conserved response allows Agrobacterium to adapt to the rhizosphere niche by modulating metabolism and cellular adaptation, such as the induction of genes coding for cell envelope synthesis, stress response, transporters of sugars and peptides (Yuan et al., 2008b).
The signaling specific response to acidity is mediated by the chromosomally encoded ChvG/ChvI two-component system, as well as other genes that allow Agrobacterium to initiate its early virulence program (Yuan et al., 2008a). ChvG acts as the sensor kinase while ChvI functions as the response regulator (Winans, 1990, 1992; Chen and Winans, 1991; Charles and Nester, 1993; Mantis and Winans, 1993; Li et al., 2002). The ChvG/I system is believed to recognize acidity in the rhizosphere and activates the expression of several virulence factors including chvI, aopB encoding an outer membrane protein, katA encoding a catalase, pckA encoding phosphoenol carboxykinase, and the imp gene cluster encoding a type VI secretion system (T6SS; Yuan et al., 2008a). A more recent study confirmed that Agrobacterium T6SS is indeed induced by acidity in a ChvG/ChvI dependent manner (Wu et al., 2012). Perhaps most interestingly, it was found that upon perception of acidic signals, several vir genes were also induced including virG, virE0, and virH (Yuan et al., 2008a), consistent with the observation that the ChvG/ChvI system activates the proximal promoter (P2) of virG (Li et al., 2002). However, to be functional, VirG requires phosphorylation signaling from another plant-derived signal, e.g., plant-derived phenolic compounds, which will be discussed in the following section. It is noteworthy that in addition to ChvG/I, another chromosomally encoded virulence gene, chvE, is also involved in Agrobacterium response to acidity and plant-derived sugars in the rhizosphere, which will also be discussed later.
Agrobacterium Responses to Plant-Derived Phenolic Compounds
Originally it was believed that plant wounding was necessary for Agrobacterium infection and pathogenicity. However, recent advances have found that plant wounding is in fact not essential for Agrobacterium pathogenicity since unwounded plants can also infected by Agrobacterium pathogens (Brencic and Winans, 2005). Besides acidic signals, plant-derived phenolic compounds are essential for the induction of Agrobacterium virulence (Stachel et al., 1986). Moreover, phenolics serve as chemoattractants for Agrobacterium (Parke et al., 1987; Melchers et al., 1989). Structural specificities of virulence inducing phenolics include the presence of a benzene ring with a hydroxyl group at position 4 and a methoxy group at position 3 (Dixon and Paiva, 1995). 3,5-dimethoxyacetophenone (acetosyringone) and hydroxyacetosyringone were the first identified inducers of Agrobacterium virulence (Stachel et al., 1985; Hess et al., 1991). The Agrobacterium VirA/VirG two-component system located on the Ti plasmid has been suggested to recognize acetosyringone as a host specific signal and activate vir gene expression (Winans et al., 1986; Leroux et al., 1987; Shaw et al., 1988; Winans, 1990). The membrane receptor VirA functions as a dimer with four domains; the periplasmic, cytoplasmic linker, kinase, and receiver domains. Upon phenolic signal perception, the linker domain of one VirA subunit activates the kinase domain of the opposite dimerized subunit by intermolecular phosphorylation (Chang and Winans, 1992; Turk et al., 1994; Toyoda-Yamamoto et al., 2000). However, a previous study has shown the binding of radiolabelled phenolic compounds to two small proteins other than VirA and controversy remains regarding the exact mechanism involved in phenolic detection by VirA (Lee et al., 1992). Nevertheless, the auto-phosphorylated sensor kinase VirA phosphorylates the cytosolic response regulator VirG at the conserved Asp52 (Morel et al., 1990; Lee et al., 1995). Phosphorylated VirG binds to a 12-bp vir box located upstream of transcription start sites of vir genes, thereby activating their transcription (Stachel et al., 1985; Stachel and Nester, 1986; Jin et al., 1990a,b,c; Pazour and Das, 1990; Roitsch et al., 1990). In fact, phosphorylated VirG also activates its own expression by activating virG transcription at the distal promoter (p1; Chang and Winans, 1992; Liu et al., 1992, 2005; Jia et al., 2002; Li et al., 2002; Yuan et al., 2008a; Wise et al., 2010).
Vir genes of Agrobacterium are organized in several vir operons. There are eight vir operons on the octopine-type Ti plasmid and relatively fewer vir genes on the nopaline-type Ti plasmid (Stachel and Nester, 1986; Rogowsky et al., 1987; Kalogeraki and Winans, 1998; Kalogeraki et al., 2000; Brencic and Winans, 2005). The vir operons are typically organized as virH, virA, virB, virG, virC, virD, virE, and virF transcriptional units (Brencic and Winans, 2005). Vir genes code for a set of proteins with different functions such as T-DNA excision and processing (virC and virD), coating and protecting T-DNA during transfer (virE), formation of the type IV secretion system (T4SS) responsible for the delivery of T-DNA to plant cells (virB operon), and T-DNA integration into plant nucleus (virE2 and virD4). A study by Cho and Winans (2005) revealed that each gene on the Ti plasmid was modestly induced by plant-derived phenolic signals, while the repABC operon, responsible for Ti plasmid replication/partitioning, was significantly induced by phenolic signals. This suggests that the copy number of the Ti plasmid is induced by plant-derived phenolics, which is confirmed by direct binding of phosphorylated VirG to a 12-bp vir box upstream of the repABC operon (Zhu et al., 2000; Pappas and Winans, 2003; Cho and Winans, 2005). Apparently an increase in Ti plasmid copy number enhances the dosage of vir genes responsible for T-DNA transfer. A proteomic study corroborated VirA/VirG dependent induction of vir genes by identifying 11 proteins that were significantly induced in response to acetosyringone, including proteins constituting the T4SS, the single strand binding protein VirE2 that is exported to the plant nucleus, and the trans-zeatin synthesizing protein Tzs (Lai et al., 2006). Moreover, responses to phenolic inducers may be modulated by detoxification of these compounds by VirH2. VirH2 was shown to play a role in the metabolism of several phenolic compounds including ferulic acid, another inducer of vir genes (Brencic et al., 2004).
Agrobacterium Responses to Plant Released Sugars in the Rhizosphere
Agrobacterium detects and responds to plant-derived sugars through a distinct signaling pathway involving VirA and a chromosomally encoded periplasmic protein, ChvE (Cangelosi et al., 1990). Expression of chvE is regulated by the LysR transcriptional regulator (TraR) galactose-binding protein regulator (GbpR) in the presence of sugars (Doty et al., 1993; Peng et al., 1998). ChvE mediates Agrobacterium chemotaxis in response to aldose monosaccharides such as galactose, glucose, arabinose, fucose, xylose, and sugar acids. Importantly, ChvE binds plant-derived sugars and subsequently interacts with the periplasmic domain of VirA to stimulate vir gene expression (Cangelosi et al., 1990; He et al., 2009; Hu et al., 2013). Mutations in the periplasmic domain of VirA present the same phenotype as a ChvE mutant with both mutants unable to infect specific plant hosts (Cangelosi et al., 1990; Chang and Winans, 1992; Banta et al., 1994; Peng et al., 1998; Gao and Lynn, 2005). Recent studies also suggest that the ability of ChvE to recognize and bind different plant-derived sugars is vital in determining the host range of Agrobacterium (Hu et al., 2013). Interestingly, the sugar response in Agrobacterium has been found to be linked with the acidity responses since the absence of sugars or mutations in chvE disrupted acidic signaling. In addition, the affinity of ChvE for sugar acids increases with a decrease in pH (Hu et al., 2013), which reinforces an important role for acidity in modulating Agrobacterium virulence. It has been proposed that acidic conditions, together with the presence of sugars and a functional ChvE, promotes VirA–ChvE interactions required for efficient vir gene induction (Shimoda et al., 1993; Toyoda-Yamamoto et al., 2000; Gao and Lynn, 2005; Nair et al., 2011). However, mutations in chvE that abolish sugar sensing do not abolish vir gene induction by acetosyringone, although ChvE is known to interact with the periplasmic domain of VirA. This suggests ChvE and associated sugar perception play additive roles that further promote vir gene expression in response to sugars and phenolic compounds, while phenolics are essential vir gene inducing signals (Cangelosi et al., 1990; He et al., 2009; Hu et al., 2013). Apart from its signaling role, ChvE also has a role in sugar utilization as it delivers sugars to the ABC transporter MmsAB (Hu et al., 2013).
Bacterial and Plant Molecules Involved in T-DNA Transfer and Integration
T-DNA transfer and integration into the plant nucleus is mediated by a complex set of Agrobacterium and host proteins. As discussed in previous sections, Agrobacterium recognizes three main plant-derived signals (acidity, phenolics, and sugars) and activates vir genes. VirD1, a helicase, and VirD2, a site specific endonuclease, are essential for nicking the Ti plasmid and the release of T-DNA as a single stranded DNA (referred to as T-strand; Yanofsky et al., 1986; Wang et al., 1990). VirD4, the coupling protein, and VirB1–VirB11, the mating-pore-formation components, together constitute the trans-envelope channel and pilus of Agrobacterium T4SS apparatus. In depth studies have assigned functional roles to each protein of the T4SS complex; VirD4, VirB3, VirB4, and VirB11 constitute the ATP-dependent translocation machinery, VirB6–VirB10 form the channel and VirB1, VirB2, VirB5, and VirB7 form the pilus (Cascales and Christie, 2003). The T-strand is covalently attached by VirD2, which is subsequently bound by VirD4 and VirB11 forming the T-complex. Current findings suggest that such DNA binding by associated virulence proteins (VirD4 and VirB11) stimulates ATP hydrolysis to produce a structural transition in the membrane channel protein VirB10. This allows passage of the T-complex to the cell surface where it can be directed to the T4SS pili, followed by delivery into the plant cell by the T4SS apparatus (Cascales and Christie, 2003; Cascales et al., 2013). Notably, many Gram-negative plant and animal pathogenic bacteria employ a type III secretion system (T3SS) to inject effector proteins directly into the cytosol of eukaryotic cells and thus allow the manipulation of host cellular activities to the benefit of the pathogen (Buttner and He, 2009). However, no T4SS organelles in Agrobacterium reminiscent of the basal body of flagella or needle complexes of the T3SS were evident (Christie, 2004). The exact process is still unknown regarding how the T-DNA complex is delivered by T4SS into plant cells, especially how the T-DNA complex passes through cell wall and plasma membrane, subsequently moving through the cytoplasm to the plant cell nucleus (Cascales and Christie, 2003; Gelvin, 2003).
T-DNA integration into plant nuclei is thought to occur by hijacking various host systems including defense signaling, cytoskeletal networking, molecular motors, nuclear import, proteolytic degradation, chromatin targeting, and DNA repair to ensure successful plant transformation (for review see Citovsky et al., 2007). In the plant cytoplasm, the T-strand/VirD2 complex is coated along its entire length by the VirE2 ssDNA-binding protein that is transported into the plant cell independently of the T-strand complex (Vergunst et al., 2000; Citovsky et al., 2007). Both VirD2 and VirE2 carry plant nuclear localization signals and together with host protein VIP1 (VirE2 interacting protein 1), facilitate T-complex import into the plant cell for host chromatin targeting (Citovsky et al., 1992; Tzfira et al., 2001; Li et al., 2005; Djamei et al., 2007; Lacroix et al., 2008). T-DNA is thought to attach to chromatin by interacting with nucleosomal proteins and is released from the T-complex by proteolytic removal of associated proteins (Magori and Citovsky, 2011). VirF, a bacterial F-box protein, also targets both VIP1 and VirE2 for proteasome dependent degradation. The mechanism of T-DNA integration into the plant genome is thought to occur by illegitimate recombination; however, the details of many of the molecular events within the plant cell and nucleus are still unclear.
Agrobacterium Metabolizes Plant-Derived Opines as a Source of Nutrients
Besides IAA and CK, infected plant cells produce over 20 different kinds of opines that can be classified into four families: octopine, nopaline, mannopine, and agrocinopine families (Beck von Bodman et al., 1992; Fuqua and Winans, 1996b; Piper et al., 1999). In fact, the most intensively studied Ti plasmids are the octopine and nopaline types, named after the predominant opines synthesized by transformed plant cells. Octopine is synthesized by the T-DNA-encoded enzyme octopine synthase (Ocs), which condenses pyruvate with different amino acids to produce octopine, lysopine, histopine, or octopinic acid (Dessaux et al., 1998). Nopaline is generated by nopaline synthase (Nos) in a similar condensation reaction involving αα-ketoglutaric acid and either arginine or ornithine. Opines of the mannopine and agrocinopine families are structurally more heterogeneous, which contain sugar and phosphate groups in the case of agrocinopine. Since plants cannot metabolize opines, transformed plant cells accumulate and release opines into the rhizosphere. The precise mechanism by which opines are exuded from plant cells is unknown, although the exudation of octopine and nopaline appears to depend upon the product of T-DNA gene 6a. Nevertheless, opines are present on the plant (or tumor) surface and are part of the soluble plant exudates released into the phylloplane and rhizoplane (Savka and Farrand, 1992).
Agrobacterium Ti plasmids also contain genes for opine uptake and catabolism that are located in the non-transferrable region, e.g., occ and noc regions for octopine- and nopaline-type Ti plasmids. In addition, Ti plasmids contain chemotaxic genes for their corresponding opines (Beck von Bodman et al., 1992; Kim and Farrand, 1997). Agrobacterium LysR-type transcriptional activator OccR (octopine catabolic regulator) and NocR (nopaline catabolic regulator) recognize and bind to opines, subsequently activating the expression of opine catabolic genes (Beck von Bodman et al., 1992; Wang et al., 1992). Agrobacterium metabolism of agrocinopine is much more complicated. When agrocinopines are present, the repressor agrocinopine catabolic regulator (AccR) dissociates from the promoter, allowing for expression of the acc operon responsible for agrocinopine metabolism. In addition, the acc operon is activated in response to phosphate limitation (Kim et al., 2008). Some of opine catabolic genes are also under regulation of other factors, for example the presence of certain substrates such as succinate (Hong et al., 1993). Although the rhizosphere contains species other than Agrobacterium that are capable of utilizing opines, they comprise of a very small minority of the bacterial population (Nautiyal and Dion, 1990). Therefore, the ability to use opines as a carbon, nitrogen, and energy source provides distinct advantages to Agrobacterium in the rhizosphere niche.
Agrobacterium Quorum Sensing is Activated by Plant-Derived Opines
In addition to serving as a nutrition source for Agrobacterium, opines produced by transformed plant cells also activate Agrobacterium QS. In fact, the original study of Agrobacterium QS was relevant to Ti plasmid conjugation. In soil or cultivation at temperatures greater than 30°C, the Ti plasmid is rapidly lost from Agrobacterium populations. Once infected by Agrobacterium, plant cells produce opines. In addition to activating genes for opine metabolism, the NocR- or OccR-opine complex also activates a LuxR-type TraR located on the Ti plasmid (Li and Farrand, 2000). When the diffusible QS signal N-(3-oxooctanoyl)-DL-homoserine lactone (3OC8-HSL) reaches a threshold level with high population density, TraR binds to 3OC8-HSL. The TraR–3OC8-HSL complex activates traI, a LuxI-type 3OC8-HSL synthase, and tra/trb genes coding for a second T4SS responsible for conjugal transfer of Ti plasmids (Piper et al., 1993; Zhang et al., 1993; Fuqua and Winans, 1994; Hwang et al., 1994). Since the Ti plasmid carries genes responsible for plant infection and opine metabolism, avirulent Agrobacterium lacking the Ti plasmid becomes infectious and capable of opine metabolism by acquiring the Ti plasmid through conjugation. Additionally, the TraR–3OC8-HSL complex activates the repABC operon thereby enhancing the replication and copy number of the Ti plasmid (Fuqua and Winans, 1996a; White and Winans, 2007). TraR–3OC8-HSL complex also activates the transcription of TraM, a TraR antiactivator in both octopine- and nopaline-type strains of Agrobacterium. TraM further modulates QS and Ti plasmid conjugation in the rhizosphere (Hwang et al., 1999). Therefore, the initial infection and T-DNA transfer leads to the synthesis of opines in plant cells. Opines activate the Agrobacterium TraR/TraI QS system, which activates Ti plasmid conjugation and enhances Ti plasmid copy number (up to eightfold). This typical positive feedback regulation is advantageous for maximal infection of plant hosts and opine metabolism (Zhu and Winans, 1999, 2001; Pappas and Winans, 2003; Cho and Winans, 2005; Pinto et al., 2012).
Agrobacterium QS is Further Modulated by Other Plant-Derived Signals
Agrobacterium QS is well regulated not only for QS signal production, but also for QS signal degradation, also known as quorum quenching. γ-amino butyric acid (GABA) significantly increases in wounded plant tissues and acidic conditions. GABA also accumulates in plants infected by Agrobacterium (Chevrot et al., 2006). In addition, proline significantly accumulates in plant tumors but neither in wounded nor healthy tissues (Deeken et al., 2006). The Agrobacterium proline/GABA receptor atu2422 and ABC-transporter braE (atu2427) are required for GABA and proline uptake. GABA activates transcription of the attKLM operon located on the second plasmid of Agrobacterium. AttK is a NAD dependent dehydrogenase. AttL is a alcohol dehydrogenase and AttM, a γ-butyrolactonase, breaks down the Agrobacterium QS signal 3OC8-HSL. Proline is a competitive antagonist of GABA and is also taken up through the Atu2422-Bra ABC transporter system (Wachter et al., 2003; Haudecoeur et al., 2009). It was found that plants with relatively higher proline levels present bigger tumors and severe disease symptoms, whereas those with relatively high GABA attenuated Agrobacterium pathogenesis. This is likely a result of the pathogen’s enhanced virulence through QS that is negatively regulated by GABA (Brugiere et al., 1999). Furthermore, it was found that a short interfering RNA, AbcR1, targets the ribosome binding site of atu2422 and negatively affects its translation (Wilms et al., 2011, 2012).
Recent studies revealed that the plant defense signal salicylic acid (SA) also activates the attKLM operon thereby down-regulating Agrobacterium QS (Yuan et al., 2007, 2008a). It was suggested that down-regulation of QS during the initial stages of infection benefits Agrobacterium pathogenicity, since high levels of QS signals are known to trigger a defense response in eukaryotic hosts (Ritchie et al., 2005; Wagner et al., 2007). Therefore, Agrobacterium QS is under tight and complex modulation by plant-derived opine, SA, and GABA to ensure optimum infection of plant hosts and to avoid the elicitation of plant defense responses by high levels of QS signals, reflecting an evolutionary advantage. In addition, quorum quenching induced by SA and GABA might function to prevent unnecessary energy expenditure after T-DNA transfer. Moreover, since the AttM lactonase has a broad substrate range, the activation of Agrobacterium quorum quenching by GABA and SA likely confers Agrobacterium a competitive advantage by degrading QS signals from unrelated competitive bacteria occupying the rhizosphere niche (Mathesius et al., 2003; Carlier et al., 2004; Chevrot et al., 2006; Yuan et al., 2007). Furthermore, induction of attKLM genes allows Agrobacterium to metabolize other plant-released compounds such as gamma-butyrolactone to produce succinic acid for the central metabolism (tricarboxylic acid cycle; Carlier et al., 2004; Chevrot et al., 2006; Chai et al., 2007).
Agrobacterium Virulence Modulated by Plant Hormones and Plant-Derived Chemicals
Plant hormones play important roles in plant defense and stress resistance. IAA and ethylene (ET) levels in plant tissues are elevated at the initial stages of infection by Agrobacterium. Following T-DNA integration, SA, IAA, and ET levels are elevated (Lee et al., 2009), while jasmonic acid (JA) levels were unchanged. However, in tumors, IAA and ET signaling pathways were activated, while JA and SA signaling pathways remained inactivated. Synthesis of IAA in crown gall occurs as a two-step process from tryptophan via indoleacetamide, mediated by-products of the T-DNA encoded iaaM and iaaH genes (Thomashow et al., 1986). T-DNA also carries an ipt gene responsible for CK synthesis. The ipt product condenses isopentenyl pyrophosphate and AMP to produce isopentenyl-AMP, which is later converted to CK by host enzymes. The elevated level of IAA and CK promote plant cell growth and tumor formation. It is now becoming evident that key phytohormones significantly influence Agrobacterium pathogenicity and tumor formation through both plant signaling pathways as well as direct modulation of bacterial processes (Veselov et al., 2003; Yuan et al., 2007; Zottini et al., 2007; Anand et al., 2008). The following sections are focussed on the effects of plant hormones on Agrobacterium pathogenicity, in particular, how Agrobacterium responds to these plant hormones.
Agrobacterium Responses to Indole-3-Acetic Acid (IAA)
IAA is important for plant growth and development, where its functions are mediated by the asymmetric distribution of IAA both systemically and locally (Korbei and Luschnig, 2011). IAA produced by Agrobacterium infected cells not only contributes to tumor growth, but also affects Agrobacterium pathogenicity. It was found that IAA, at 25 μM concentrations, inhibits Agrobacterium vir gene expression while not significantly affecting Agrobacterium growth. This is thought to occur by competition between IAA and phenolic inducers of vir genes for their interaction with the VirA/VirG two-component system (Liu and Nester, 2006). Further studies have indicated that IAA likely competes with vir gene inducing signals, such as acetosyringone, for association with the VirA linker domain, which is strengthened by the related molecular structures of acetosyringone and IAA. Activation of vir genes by acetosyringone and IAA-mediated inhibition of vir genes have never been genetically separated. Moreover, the inhibition of vir genes by IAA can be rescued by higher level of acetosyringone or incorporation of a constitutive virA expressing plasmid. Furthermore, IAA inhibits Agrobacterium growth at higher concentrations (over 50 μM) yet does not kill Agrobacterium (Liu and Nester, 2006). It was proposed that after successful transformation of a plant host, the synthesis of large amounts of IAA in infected plant tissues represses vir gene expression for energy conservation. Yet it remains unclear if the local concentration of IAA in fresh tumors reaches the inhibitory range (Liu and Nester, 2006).
Agrobacterium Responses to Salicylic Acid (SA)
SA is a well-known phytohormone activating plant defense responses to incompatible interactions (Zottini et al., 2007). During Agrobacterium–plant interactions, SA produced in infected plants modulates the Agrobacterium virulence program by several mechanisms (Yuan et al., 2007; Lee et al., 2009). Apart from mounting plant defense responses, SA at biologically relevant concentrations (8–10 μM) limits Agrobacterium growth, represses vir gene expression, and dampens Agrobacterium QS as discussed in the previous section (Yuan et al., 2007, 2008b). In fact, SA inhibits all the vir genes including the repABC operon, suggesting SA likely prevents the increase of Ti plasmid copy number. This is consistent with the observation that SA-overproducing plants display recalcitrance to Agrobacterium infection, whereas mutant plants impaired in SA biosynthesis and accumulation are more susceptible to tumor growth (Yuan et al., 2007; Lee et al., 2009). Similar to IAA, the inhibition of vir gene expression by SA can be rescued by either increasing levels of acetosyringone or incorporation of a constitutive virA expressing plasmid. SA likely functions as an allosteric competitive inhibitor and interferes with the interaction between the kinase domain of VirA and acetosyringone since the constitutively expressed VirA activates vir gene expression independent of acetosyringone (Yuan et al., 2007).
Agrobacterium Responses to Ethylene (ET)
ET, unlike other plant hormones, is a volatile hormone that affects many aspects of plant growth and development (Wang et al., 2013). ET also acts as a plant stress signal. ET signaling pathways are induced by various biotic and abiotic stresses including osmotic stress, salt stress, wounding, pathogen attack and flooding. These stress-induced ET signaling pathways have substantial roles in defense responses and disease resistance by accelerating senescence, abscission of infected organs and induction of specific defense proteins (Chang and Shockey, 1999). Plant tissues rich in ET, such as melons, are recalcitrant to Agrobacterium transformation, yet the cause for the transformation recalcitrance remains unclear. Thus, various strategies have been employed to reduce ET level to improve Agrobacterium transformation efficiency, including the application of an antisense ACC oxidase gene (pAP4), the final enzyme in the ET biosynthetic pathway. Recent studies have found that ET is another important factor modulating Agrobacterium virulence programing and determining crown gall morphogenesis (Nonaka et al., 2008). In particular, Agrobacterium-mediated genetic transformation was inhibited in ET-sensing melon but enhanced in ET-insensitive mutants. Further studies also revealed that Agrobacterium growth was not affected by ET, but the presence of ET at the beginning of Agrobacterium infection displays significant inhibitory activity on vir gene expression. Such inhibitory effects can be rescued by supplementation with acetosyringone, a vir gene inducer (Nonaka et al., 2008). Although the ET levels are up-regulated during Agrobacterium infection, plant genes for ET receptors and downstream signaling are not induced (Lee et al., 2009). These observations suggest that ET impacts Agrobacterium–plant interactions largely through its inhibitory effects on bacterial virulence programming.
Additional Plant-Derived Vir Gene Inhibitors in Nature
In addition to the universal phytohormones SA, ET, and IAA, monocots contain special chemicals acting as natural inhibitors of Agrobacterium virulence. Maize, along with other monocots, are notoriously resistant to Agrobacterium transformation and the cause for this has been delimited to the inhibition of Agrobacterium vir genes (Heath et al., 1997). In particular, metabolites derived from corn seedlings (Zea mays) such as 2,4-dihydroxy-7-methoxy-2H-1,4-benzoxazin-3(4H)-one (DIMBOA) and 2-hydroxy-4,7-dimethoxybenzoxazin-3-one (MDIBOA) inhibit the expression of Agrobacterium vir genes in the presence of vir inducing signals (Sahi et al., 1990; Zhang et al., 2000). In addition, Agrobacterium mutants resistant to either DIMBOA or MDIBOA were much more effective in infecting plant hosts. Moreover, Agrobacterium carrying constitutively active virA are insensitive to MDIBOA in terms of the inhibition of vir genes. These observations suggest that DIMBOA and MDIBOA, similar to SA, ET, and IAA, probably affect signal perception by the VirA sensor kinase prior to the VirA/G phosphorylation signal relay events.
Conclusion and Future Perspectives
Signaling Integration and Cascade Activation of Agrobacterium Virulence by Plant-Derived Signals
Several lines of evidence suggests a hierarchical activation of Agrobacterium virulence by a combination of plant-derived signals, as illustrated in Figure 1. Rhizospheric acidity activates the ChvG/I system, which subsequently activates virG transcription at the proximal promoter (P2) to allow basal level expression of virG. Therefore, the ChvG/I system functions upstream of VirA/VirG system (Li et al., 2002; Yuan et al., 2008b). Upon recognizing phenolic signals such as acetosyringone, VirA becomes auto-phosphorylated and subsequently phosphorylates VirG. Phosphorylated VirG activates the expression of vir genes responsible for T-DNA transfer and integration. Phosphorylated VirG also activates virG expression at the distal promoter (p1) to further promote virulence. ChvE binds to plant-derived sugars and subsequently interacts with the periplasmic domain of VirA to allow for the maximal expression of vir genes. Thus, the VirA/G system couples and integrates three rhizosphere signals: acidity, sugars (monosaccharides) and phenolic compounds. Such signaling integration and cascade activation of Agrobacterium virulence ensures precise perception of suitable plant hosts in the rhizosphere and maximal infection, reflecting an evolutionary advantage (Li et al., 2002; Yuan et al., 2008b). Although ChvG/I regulates VirA/G, chvG/chvI are expressed independent of the VirA/G system and plant-derived phenolic signals (Charles and Nester, 1993; Peng et al., 1998).
Plant-Derived Signals Function Additively and Play Redundant Roles in Modulating Agrobacterium Virulence, Ti Plasmid Copy Number and Qs
Since Ti plasmid harbors vir genes as well as genes for opine metabolism, Ti plasmid copy number directly influences pathogenicity and the efficiency of opine metabolism. It was established that the repABC operon responsible for Ti plasmid replication and partition is induced by plant-derived phenolics through the VirA/G system. In addition, opine activated-QS further promotes the expression of the repABC operon. On the other hand, plant-derived SA and GABA activate Agrobacterium quorum quenching, which has negative impacts on Ti plasmid copy number. Furthermore, SA, IAA, and ET inhibit vir genes including the repABC operon, thereby preventing the increase of Ti plasmid copy number. The signaling complexity also applies to the modulation of Agrobacterium vir genes, which are activated by tripartite signals in the rhizosphere, acidity, phenolics, and plant-derived sugars, but down-regulated by SA, IAA, and ET and other natural vir gene inhibitors such as DIMBOA and MDIBOA (not shown in the Figure 1). In fact, the modulation of Ti plasmid copy number and conjugation also influences the overall Agrobacterium pathogenicity. It is noteworthy that Agrobacterium mounts distinct but overlapping cellular responses to SA, IAA, and GABA, despite the absence of any structural relation (Yuan et al., 2008a). Therefore, it is plausible that in nature, different plant-derived signals act in concert and function additively, playing redundant roles in tailoring Agrobacterium virulence, Ti plasmid copy number and QS (Yuan et al., 2008a).
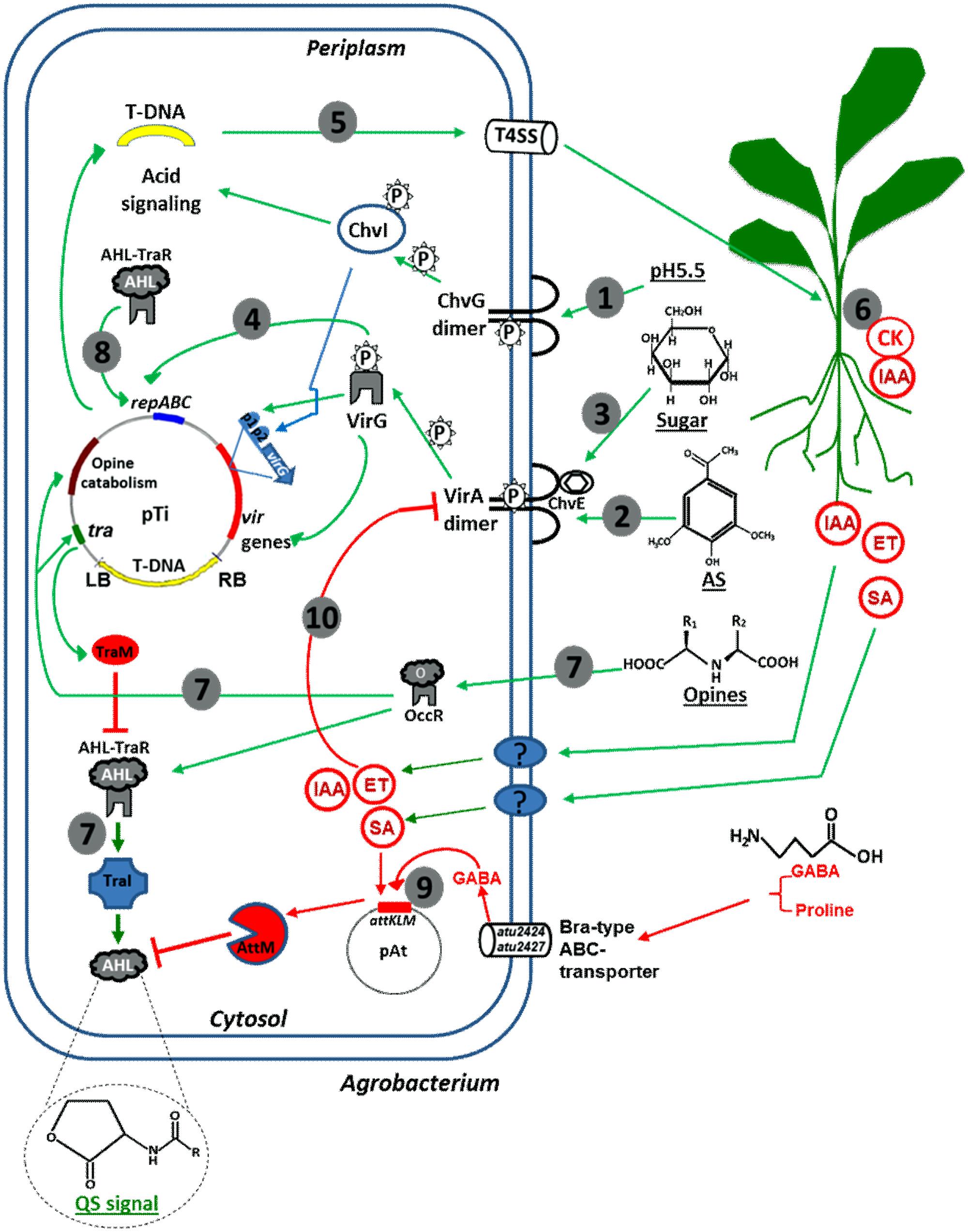
FIGURE 1. Schematic drawing of Agrobacterium responses to plant-derived signals. (1) Upon perception of acidic conditions in the rhizosphere, the ChvG/I two-component system activates the expression of several virulence genes including chvI and virG; (2) Upon perception of plant-derived phenolics, the VirA/G two-component system activates all vir genes including virG to further promote vir gene expression; (3) ChvE binds plant-released sugars and interacts with the VirA to allow maximal vir gene expression; (4) Agrobacterium Ti plasmid copy number is up-regulated in response to phenolic compounds; (5) Vir gene products process and deliver T-DNA into plant nuclei; (6) Expression of T-DNA encoded genes in plants leads to the production of IAA, CK, and opines; (7) Opine activates Agrobacterium genes for opine metabolism, as well as TraR/TraI QS system that subsequently induces Ti plasmid conjugation; (8) QS also up-regulates Ti plasmid copy number for maximal pathogenicity; (9) Agrobacterium quorum quenching (attKLM operon) is activated by plant-derived GABA and SA thereby down-regulates QS; (10) Agrobacterium hijacks plant-derived SA, IAA, and ET to down-regulate vir gene expression.
Through Infection of Plants, Agrobacterium Converts Plant Cell Into a Factory to Secure Nutrients and Maintain the Genetic Integrity in Nature
The evolution and survival of Agrobacterium as a bacterial species depends on an intricate balance of two populations of cells, those which actively maintain and those which passively lose the Ti-plasmid. Both forms are necessary for the species to sustain competitive lifestyles in either the absence or presence of a plant host. In the absence of a plant host, Agrobacterium harboring the Ti plasmid are at a growth disadvantage to those Agrobacterium lacking the plasmid, which is ascribed to the metabolic burden needed to maintain such a large Ti plasmid. In the presence of a host plant and opines, the advantage is shifted to Ti plasmid-retaining Agrobacterium since the Ti plasmid contains genes responsible for opine uptake and metabolism. Opines not only activates genes responsible for opine metabolism, but also activate QS-dependent functions such as induction of Ti plasmid conjugation and enhancement of Ti plasmid copy number, promoting maximal infection. In fact, the increase of Ti plasmid copy number may be advantageous for Ti plasmid conjugation. Moreover, Agrobacterium hijacks SA and GABA signaling to activate the AttKLM operon which also degrades plant-derived GABA, gamma-butyrolactone, and gammahydroxy butyrate to provide even more nutrients for the tricarboxylic acid cycle. Therefore, it is reasonable to believe that through plant transformation, Agrobacterium converts infected plant cells into a factory to secure nutrients, in particular opines, nutrients almost exclusive for Agrobacterium. In addition, QS activates Ti plasmid conjugation enabling Agrobacterium to maintain the Ti plasmid and genetic integrity in nature.
In summary, Agrobacterium pathogenicity is largely attributed to its evolved capabilities of precise recognition, response to and hijacking of plant-derived chemical signals for its own benefit. The complex inter-kingdom signaling interplay and regulatory circuits highlight elegant mechanisms of Agrobacterium–host co-evolution. Plant roots secrete and release a wide range of chemical cues into the rhizosphere (Bais et al., 2005), admittedly, only a limited number of plant-derived chemicals have been intensively studied for their roles in Agrobacterium–plant interactions. For future studies, it will be worthwhile to identify additional plant-derived chemicals that impact Agrobacterium pathogenicity and rhizospheric fitness. In addition, it will be very interesting to elucidate Agrobacterium signaling pathways and underlying regulatory mechanisms responsible for the precise perception and response to these plant-derived signals, especially at the early stage of Agrobacterium–plant interaction.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We apologize to colleagues whose original work was not cited due to space limitation. This study was funded by the Agriculture and Agri-Food Canada, Growing Forward-AgriFlex funding (RBPI number 2555) and conducted by the authors as part of their duties. We also thank Alexander William Eastman and Brian Weselowski for critical reading of the manuscript. We also thank Dr. Stanton Gelvin for critical reading and editing of the manuscript.
Abbreviations
AS, acetosyringone; chv genes, chromosome encoded virulence genes; CK, cytokinin; DIMBOA, 2,4-dihydroxy-7-methoxy-2H-1,4-benzixazin-3(4H)-one; ET, ethylene; GABA, γ-amino butyric acid; IAA, indole acetic acid; MDIBOA, 2-hydroxy-4,7-dimethoxybenzoxazin-3-one; QS, quorum sensing; SA, salicylic acid; T3SS, type III secretion system; T4SS, type IV secretion system, T6SS, type VI secretion system; T-DNA, transferred DNA; Ti-plasmid, tumor-inducing plasmid; vir genes, virulence genes.
References
Anand, A., Uppalapati, S. R., Ryu, C. M., Allen, S. N., Kang, L., Tang, Y.,et al. (2008). Salicylic acid and systemic acquired resistance play a role in attenuating crown gall disease caused by Agrobacterium tumefaciens. Plant Physiol. 146, 703–715. doi: 10.1104/pp.107.111302
Badri, D. V., and Vivanco, J. M. (2009). Regulation and function of root exudates. Plant Cell Environ. 32, 666–681. doi: 10.1111/j.1365-3040.2008.01926.x
Bais, H. P., Prithiviraj, B., Jha, A. K., Ausubel, F. M., and Vivanco, J. M. (2005). Mediation of pathogen resistance by exudation of antimicrobials from roots. Nature 434, 217–221. doi: 10.1038/nature03356
Banta, L. M., Joerger, R. D., Howitz, V. R., Campbell, A. M., and Binns, A. N. (1994). Glu-255 outside the predicted ChvE binding site in VirA is crucial for sugar enhancement of acetosyringone perception by Agrobacterium tumefaciens. J. Bacteriol. 176, 3242–3249.
Beck von Bodman, S., Hayman, G. T., and Farrand, S. K. (1992). Opine catabolism and conjugal transfer of the nopaline Ti plasmid pTiC58 are coordinately regulated by a single repressor. Proc. Natl. Acad. Sci. U.S.A. 89, 643–647. doi: 10.1073/pnas.89.2.643
Brencic, A., Eberhard, A., and Winans, S. C. (2004). Signal quenching, detoxification and mineralization of vir gene-inducing phenolics by the VirH2 protein of Agrobacterium tumefaciens. Mol. Microbiol. 51, 1103–1115. doi: 10.1046/j.1365-2958.2003.03887.x
Brencic, A., and Winans, S. C. (2005). Detection of and response to signals involved in host–microbe interactions by plant-associated bacteria. Microbiol. Mol. Biol. Rev. 69, 155–194. doi: 10.1128/MMBR.69.1.155-194.2005
Brugiere, N., Dubois, F., Limami, A. M., Lelandais, M., Roux, Y., Sangwan, R. S.,et al. (1999). Glutamine synthetase in the phloem plays a major role in controlling proline production. Plant Cell 11, 1995–2012. doi: 10.1105/tpc.11.10.1995
Buttner, D., and He, S. Y. (2009). The T3SS, type III protein secretion in plant pathogenic bacteria. Plant Physiol. 150, 1656–1664. doi: 10.1104/pp.109.139089
Cangelosi, G. A., Ankenbauer, R. G., and Nester, E. W. (1990). Sugars induce the Agrobacterium virulence genes through a periplasmic binding protein and a transmembrane signal protein. Proc. Natl. Acad. Sci. U.S.A. 87, 6708–6712. doi: 10.1073/pnas.87.17.6708
Cangelosi, G. A., Martinetti, G., Leigh, J. A., Lee, C. C., Thienes, C., and Nester, E. W. (1989). Role for Agrobacterium tumefaciens ChvA protein in export of beta-1,2-glucan. J. Bacteriol. 171, 1609–1615.
Carlier, A., Chevrot, R., Dessaux, Y., and Faure, D. (2004). The assimilation of gamma-butyrolactone in Agrobacterium tumefaciens C58 interferes with the accumulation of the N-acyl-homoserine lactone signal. Mol. Plant Microbe Interact. 17, 951–957. doi: 10.1094/MPMI.2004.17.9.951
Cascales, E., Atmakuri, K., Sarkar, M. K., and Christie, P. J. (2013). DNA substrate-induced activation of the Agrobacterium VirB/VirD4 type IV secretion system. J. Bacteriol. 195, 2691–2704. doi: 10.1128/JB.00114-13
Cascales, E., and Christie, P. J. (2003). The versatile bacterial type IV secretion systems. Nat. Rev. Microbiol. 1, 137–149. doi: 10.1038/nrmicro753
Chai, Y., Tsai, C. S., Cho, H., and Winans, S. C. (2007). Reconstitution of the biochemical activities of the AttJ repressor and the AttK, AttL, and AttM catabolic enzymes of Agrobacterium tumefaciens. J. Bacteriol. 189, 3674–3679. doi: 10.1128/JB.01274-06
Chang, C., and Shockey, J. A. (1999). The ethylene-response pathway: signal perception to gene regulation. Curr. Opin. Plant Biol. 2, 352–358. doi: 10.1016/S1369-5266(99)00004-7
Chang, C. H., and Winans, S. C. (1992). Functional roles assigned to the periplasmic, linker, and receiver domains of the Agrobacterium tumefaciens VirA protein. J. Bacteriol. 174, 7033–7039.
Charles, T. C., and Nester, E. W. (1993). A chromosomally encoded two-component sensory transduction system is required for virulence of Agrobacterium tumefaciens. J. Bacteriol. 175, 6614–6625.
Chen, C. Y., and Winans, S. C. (1991). Controlled expression of the transcriptional activator gene virG in Agrobacterium tumefaciens by using the Escherichia coli lac promoter. J. Bacteriol. 173, 1139–1144.
Chevrot, R., Rosen, R., Haudecoeur, E., Cirou, A., Shelp, B. J., Ron, E.,et al. (2006). GABA controls the level of quorum-sensing signal in Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. U.S.A. 103, 7460–7464. doi: 10.1073/pnas.0600313103
Cho, H., and Winans, S. C. (2005). VirA and VirG activate the Ti plasmid repABC operon, elevating plasmid copy number in response to wound-released chemical signals. Proc. Natl. Acad. Sci. U.S.A. 102, 14843–14848. doi: 10.1073/pnas.0503458102
Christie, P. J. (2004). Type IV secretion: the Agrobacterium VirB/D4 and related conjugation systems. Biochim. Biophys. Acta 1694, 219–234. Review. doi: 10.1016/j.bbamcr.2004.02.013
Citovsky, V., Kozlovsky, S. V., Lacroix, B., Zaltsman, A., Dafny-Yelin, M., Vyas, S.,et al. (2007). Biological systems of the host cell involved in Agrobacterium infection. Cell Microbiol. 9, 9–20. doi: 10.1111/j.1462-5822.2006.00830.x
Citovsky, V., Zupan, J., Warnick, D., and Zambryski, P. (1992). Nuclear localization of Agrobacterium VirE2 protein in plant cells. Science 256, 1802–1805. doi: 10.1126/science.1615325
Deeken, R., Engelmann, J. C., Efetova, M., Czirjak, T., Müller, T., Kaiser, W. M.,et al. (2006). An integrated view of gene expression and solute profiles of Arabidopsis tumors: a genome-wide approach. Plant Cell 18, 3617–3634. doi: 10.1105/tpc.106.044743
Dessaux, Y., Petit, A., Farrand, S. K., and Murphy, P. J. (1998). “Opines and opine-like molecules involved in plant–Rhizobiaceae interactions,” in The Rhizobiaceae: Molecular Biology of Model Plant-associated Bacteria, eds H. P. Spaink, A. Kondorosi, and P. J. J. Hooykaas (Dordrecht: Kluwer Academic Publishers), 173–197.
Dixon, R. A., and Paiva, N. L. (1995). Stress-induced phenylpropanoid metabolism. Plant Cell 7, 1085–1097. doi: 10.1105/tpc.7.7.1085
Djamei, A., Pitzschke, A., Nakagami, H., Rajh, I., and Hirt, H. (2007). Trojan horse strategy in Agrobacterium transformation: abusing MAPK defense signaling. Science 318, 453–456. doi: 10.1126/science.1148110
Doty, S. L., Chang, M., and Nester, E. W. (1993). The chromosomal virulence gene, chvE, of Agrobacterium tumefaciens is regulated by a LysR family member. J. Bacteriol. 175, 7880–7886.
Fuqua, C., and Winans, S. C. (1996a). Conserved cis-acting promoter elements are required for density-dependent transcription of Agrobacterium tumefaciens conjugal transfer genes. J. Bacteriol. 178, 435–440.
Fuqua, C., and Winans, S. C. (1996b). Localization of OccR-activated and TraR-activated promoters that express two ABC-type permeases and the traR gene of Ti plasmid pTiR10. Mol. Microbiol. 20, 1199–1210. doi: 10.1111/j.1365-2958.1996.tb02640.x
Fuqua, W. C., and Winans, S. C. (1994). A LuxR-LuxI type regulatory system activates Agrobacterium Ti plasmid conjugal transfer in the presence of a plant tumor metabolite. J. Bacteriol. 176, 2796–2806.
Gao, R., and Lynn, D. G. (2005). Environmental pH sensing: resolving the VirA/VirG two-component system inputs for Agrobacterium pathogenesis. J. Bacteriol. 187, 2182–2189. doi: 10.1128/JB.187.6.2182-2189.2005
Gelvin, S. B. (2003). Agrobacterium-mediated plant transformation: the biology behind the “Gene-Jockeying” tool. Microbiol. Mol. Biol. Rev. 67, 16–37. doi: 10.1128/MMBR.67.1.16-37.2003
Gelvin, S. B. (2012). Traversing the cell: Agrobacterium T-DNA’s journey to the host genome. Front. Plant Sci. 3:52. doi: 10.3389/fpls.2012.00052
Haudecoeur, E., Planamente, S., Cirou, A., Tannieres, M., Shelp, B. J., Morera, S.,et al. (2009). Proline antagonizes GABA-induced quenching of quorum-sensing in Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. U.S.A. 106, 14587–14592. doi: 10.1073/pnas.0808005106
He, F., Nair, G. R., Soto, C. S., Chang, Y., Hsu, L., Ronzone, E.,et al. (2009). Molecular basis of ChvE function in sugar binding, sugar utilization, and virulence in Agrobacterium tumefaciens. J. Bacteriol. 191, 5802–5813. doi: 10.1128/JB.00451-09
Heath, J. D., Boulton, M. I., Raineri, D. M., Doty, S. L., Mushegian, A. R., Charles, T. C.,et al. (1997). Discrete regions of the sensor protein virA determine the strain-specific ability of Agrobacterium to agroinfect maize. Mol. Plant Microbe Interact. 10, 221–227. doi: 10.1094/MPMI.1997.10.2.221
Hess, K. M., Dudley, M. W., Lynn, D. G., Joerger, R. D., and Binns, A. N. (1991). Mechanism of phenolic activation of Agrobacterium virulence genes: development of a specific inhibitor of bacterial sensor/response systems. Proc. Natl. Acad. Sci. U.S.A. 88, 7854–7858. doi: 10.1073/pnas.88.17.7854
Hong, S. B., Dessaux, Y., Chilton, W. S., and Farrand, S. K. (1993). Organization and regulation of the mannopine cyclase-associated opine catabolism genes in Agrobacterium tumefaciens 15955. J. Bacteriol. 175, 401–410.
Hu, X., Zhao, J., Degrado, W. F., and Binns, A. N. (2013). Agrobacterium tumefaciens recognizes its host environment using ChvE to bind diverse plant sugars as virulence signals. Proc. Natl. Acad. Sci. U.S.A. 110, 678–683. doi: 10.1073/pnas.1215033110
Huckelhoven, R. (2007). Transport and secretion in plant-microbe interactions. Curr. Opin. Plant Biol. 10, 573–579. doi: 10.1016/j.pbi.2007.08.002
Hwang, I., Li, P. L., Zhang, L., Piper, K. R., Cook, D. M., Tate, M. E.,et al. (1994). TraI, a LuxI homologue, is responsible for production of conjugation factor, the Ti plasmid N-acylhomoserine lactone autoinducer. Proc. Natl. Acad. Sci. U.S.A. 91, 4639–4643. doi: 10.1073/pnas.91.11.4639
Hwang, I., Smyth A. J., Luo Z. Q., and Farrand S. K. (1999). Modulating quorum sensing by antiactivation: TraM interacts with TraR to inhibit activation of Ti plasmid conjugal transfer genes. Mol. Microbiol. 34, 282–294. doi: 10.1046/j.1365-2958.1999.01595.x
Jia, Y. H., Li, L. P., Hou, Q. M., and Pan, S. Q. (2002). An Agrobacterium gene involved in tumorigenesis encodes an outer membrane protein exposed on the bacterial cell surface. Gene 284, 113–124. doi: 10.1016/S0378-1119(02)00385-2
Jin, S., Roitsch, T., Ankenbauer, R. G., Gordon, M. P., and Nester, E. W. (1990a). The VirA protein of Agrobacterium tumefaciens is autophosphorylated and is essential for vir gene regulation. J. Bacteriol. 172, 525–530.
Jin, S. G., Prusti, R. K., Roitsch, T., Ankenbauer, R. G., and Nester, E. W. (1990b). Phosphorylation of the VirG protein of Agrobacterium tumefaciens by the autophosphorylated VirA protein: essential role in biological activity of VirG. J. Bacteriol. 172, 4945–4950.
Jin, S. G., Roitsch, T., Christie, P. J., and Nester, E. W. (1990c). The regulatory VirG protein specifically binds to a cis-acting regulatory sequence involved in transcriptional activation of Agrobacterium tumefaciens virulence genes. J. Bacteriol. 172, 531–537.
Kalogeraki, V. S., and Winans, S. C. (1998). Wound-released chemical signals may elicit multiple responses from an Agrobacterium tumefaciens strain containing an octopine-type Ti plasmid. J. Bacteriol. 180, 5660–5667.
Kalogeraki, V. S., Zhu, J., Stryker, J. L., and Winans, S. C. (2000). The right end of the vir region of an octopine-type Ti plasmid contains four new members of the vir regulon that are not essential for pathogenesis. J. Bacteriol. 182, 1774–1778. doi: 10.1128/JB.182.6.1774-1778.2000
Kim, H., and Farrand, S. K. (1997). Characterization of the acc operon from the nopaline-type Ti plasmid pTiC58, which encodes utilization of agrocinopines A and B and susceptibility to agrocin 84. J. Bacteriol. 179, 7559–7572.
Kim, H. S., Yi, H., Myung, J., Piper, K. R., and Farrand, S. K. (2008). Opine-based Agrobacterium competitiveness: dual expression control of the agrocinopine catabolism (acc) operon by agrocinopines and phosphate levels. J. Bacteriol. 190, 3700–3711. doi: 10.1128/JB.00067-08
Korbei, B., and Luschnig, C. (2011). Cell polarity: PIN it down! Curr. Biol. 21, R197–R199. doi: 10.1016/j.cub.2011.01.062
Lacroix, B., Loyter, A., and Citovsky, V. (2008). Association of the Agrobacterium T-DNA-protein complex with plant nucleosomes. Proc. Natl. Acad. Sci. U.S.A. 105, 15429–15434. doi: 10.1073/pnas.0805641105
Lai, E. M., Shih, H. W., Wen, S. R., Cheng, M. W., Hwang, H. H., and Chiu, S. H. (2006). Proteomic analysis of Agrobacterium tumefaciens response to the Vir gene inducer acetosyringone. Proteomics 6, 4130–4136. doi: 10.1002/pmic.200600254
Lee, C. W., Efetova, M., Engelmann, J. C., Kramell, R., Wasternack, C., Ludwig-Muller, J.,et al. (2009). Agrobacterium tumefaciens promotes tumor induction by modulating pathogen defense in Arabidopsis thaliana. Plant Cell 21, 2948–2962. doi: 10.1105/tpc.108.064576
Lee, K., Dudley, M. W., Hess, K. M., Lynn, D. G., Joerger, R. D., and Binns, A. N. (1992). Mechanism of activation of Agrobacterium virulence genes: identification of phenol-binding proteins. Proc. Natl. Acad. Sci. U.S.A. 89, 8666–8670. doi: 10.1073/pnas.89.18.8666
Lee, Y. W., Jin, S., Sim, W. S., and Nester, E. W. (1995). Genetic evidence for direct sensing of phenolic compounds by the VirA protein of Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. U.S.A. 92, 12245–12249. doi: 10.1073/pnas.92.26.12245
Leroux, B., Yanofsky, M. F., Winans, S. C., Ward, J. E., Ziegler, S. F., and Nester, E. W. (1987). Characterization of the virA locus of Agrobacterium tumefaciens: a transcriptional regulator and host range determinant. EMBO J. 6, 849–856.
Li, J., Krichevsky, A., Vaidya, M., Tzfira, T., and Citovsky, V. (2005). Uncoupling of the functions of the Arabidopsis VIP1 protein in transient and stable plant genetic transformation by Agrobacterium. Proc. Natl. Acad. Sci. U.S.A. 102, 5733–5738. doi: 10.1073/pnas.0404118102
Li, L., Jia, Y., Hou, Q., Charles, T. C., Nester, E. W., and Pan, S. Q. (2002). A global pH sensor: Agrobacterium sensor protein ChvG regulates acid-inducible genes on its two chromosomes and Ti plasmid. Proc. Natl. Acad. Sci. U.S.A. 99, 12369–12374. doi: 10.1073/pnas.192439499
Li, P. L., and Farrand, S. K. (2000). The replicator of the nopaline-type Ti plasmid pTiC58 is a member of the repABC family and is influenced by the TraR-dependent quorum-sensing regulatory system. J. Bacteriol. 182, 179–188. doi: 10.1128/JB.182.1.179-188.2000
Liu, C. N., Li, X. Q., and Gelvin, S. B. (1992). Multiple copies of virG enhance the transient transformation of celery, carrot and rice tissues by Agrobacterium tumefaciens. Plant Mol. Biol. 20, 1071–1087. doi: 10.1007/BF00028894
Liu, P., and Nester, E. W. (2006). Indoleacetic acid, a product of transferred DNA, inhibits vir gene expression and growth of Agrobacterium tumefaciens C58. Proc. Natl. Acad. Sci. U.S.A. 103, 4658–4662. doi: 10.1073/pnas.0600366103
Liu, P., Wood, D., and Nester, E. W. (2005). Phosphoenolpyruvate carboxykinase is an acid-induced, chromosomally encoded virulence factor in Agrobacterium tumefaciens. J. Bacteriol. 187, 6039–6045. doi: 10.1128/JB.187.17.6039-6045.2005
Magori, S., and Citovsky, V. (2011). Agrobacterium counteracts host-induced degradation of its effector F-box protein. Sci. Signal. 4:ra69. doi: 10.1126/scisignal.2002124
Mantis, N. J., and Winans, S. C. (1993). The chromosomal response regulatory gene chvI of Agrobacterium tumefaciens complements an Escherichia coli phoB mutation and is required for virulence. J. Bacteriol. 175, 6626–6636.
Mathesius, U., Mulders, S., Gao, M., Teplitski, M., Caetano-Anolles, G., Rolfe, B. G.,et al. (2003). Extensive and specific responses of a eukaryote to bacterial quorum-sensing signals. Proc. Natl. Acad. Sci. U.S.A. 100, 1444–1449. doi: 10.1073/pnas.262672599
McCullen, C. A., and Binns, A. N. (2006). Agrobacterium tumefaciens and plant cell interactions and activities required for interkingdom macromolecular transfer. Annu. Rev. Cell Dev. Biol. 22, 101–127. doi: 10.1146/annurev.cellbio.22.011105.102022
Melchers, L. S., Regensburg-Tuink, A. J., Schilperoort, R. A., and Hooykaas, P. J. (1989). Specificity of signal molecules in the activation of Agrobacterium virulence gene expression. Mol. Microbiol. 3, 969–977. doi: 10.1111/j.1365-2958.1989.tb00246.x
Morel, P., Powell, B. S., and Kado, C. I. (1990). Demonstration of 3 functional domains responsible for a kinase activity in VirA, a transmembrane sensory protein encoded by the Ti plasmid of Agrobacterium tumefaciens. C. R. Acad. Sci. III 310, 21–26.
Nair, G. R., Lai, X., Wise, A. A., Rhee, B. W., Jacobs, M., and Binns, A. N. (2011). The integrity of the periplasmic domain of the VirA sensor kinase is critical for optimal coordination of the virulence signal response in Agrobacterium tumefaciens. J. Bacteriol. 193, 1436–1448. doi: 10.1128/JB.01227-10
Nam, J., Matthysse, A. G., and Gelvin, S. B. (1997). Differences in susceptibility of Arabidopsis ecotypes to crown gall disease may result from a deficiency in T-DNA integration. Plant Cell 9, 317–333. doi: 10.1105/tpc.9.3.317
Nautiyal, C. S., and Dion, P. (1990). Characterization of the opine-utilizing microflora associated with samples of soil and plants. Appl. Environ. Microbiol. 56, 2576–2579.
Nonaka, S., Yuhashi, K., Takada, K., Sugaware, M., Minamisawa, K., and Ezura, H. (2008). Ethylene production in plants during transformation suppresses vir gene expression in Agrobacterium tumefaciens. New Phytol. 178, 647–656. doi: 10.1111/j.1469-8137.2008.02400.x
Pappas, K. M., and Winans, S. C. (2003). A LuxR-type regulator from Agrobacterium tumefaciens elevates Ti plasmid copy number by activating transcription of plasmid replication genes. Mol. Microbiol. 48, 1059–1073. doi: 10.1046/j.1365-2958.2003.03488.x
Parke, D., Ornston, L. N., and Nester, E. W. (1987). Chemotaxis to plant phenolic inducers of virulence genes is constitutively expressed in the absence of the Ti plasmid in Agrobacterium tumefaciens. J. Bacteriol. 169, 5336–5338.
Pazour, G. J., and Das, A. (1990). Characterization of the VirG binding site of Agrobacterium tumefaciens. Nucleic Acids Res. 18, 6909–6913. doi: 10.1093/nar/18.23.6909
Peng, W. T., Lee, Y. W., and Nester, E. W. (1998). The phenolic recognition profiles of the Agrobacterium tumefaciens VirA protein are broadened by a high level of the sugar binding protein ChvE. J. Bacteriol. 180, 5632–5638.
Phillips, D. A., Fox, T. C., King, M. D., Bhuvaneswari, T. V., and Teuber, L. R. (2004). Microbial products trigger amino acid exudation from plant roots. Plant Physiol. 136, 2887–2894. doi: 10.1104/pp.104.044222
Pinto, U. M., Pappas, K. M., and Winans, S. C. (2012). The ABCs of plasmid replication and segregation. Nat. Rev. Microbiol. 10, 755–765. doi: 10.1038/nrmicro2882
Piper, K. R., Beck Von Bodman, S., and Farrand, S. K. (1993). Conjugation factor of Agrobacterium tumefaciens regulates Ti plasmid transfer by autoinduction. Nature 362, 448–450. doi: 10.1038/362448a0
Piper, K. R., Beck Von Bodman, S., Hwang, I., and Farrand, S. K. (1999). Hierarchical gene regulatory systems arising from fortuitous gene associations: controlling quorum sensing by the opine regulon in Agrobacterium. Mol. Microbiol. 32, 1077–1089. doi: 10.1046/j.1365-2958.1999.01422.x
Pitzschke, A. (2013). Infection and plant defense-transformation success hangs by a thread. Front. Plant Sci. 4:519. doi: 10.3389/fpls.2013.00519
Ritchie, A. J., Jansson, A., Stallberg, J., Nilsson, P., Lysaght, P., and Cooley, M. A. (2005). The Pseudomonas aeruginosa quorum-sensing molecule N-3-(oxododecanoyl)-L-homoserine lactone inhibits T-cell differentiation and cytokine production by a mechanism involving an early step in T-cell activation. Infect. Immun. 73, 1648–1655. doi: 10.1128/IAI.73.3.1648-1655.2005
Rivoal, J., and Hanson, A. D. (1994). Metabolic control of anaerobic glycolysis (overexpression of lactate dehydrogenase in transgenic tomato roots supports the Davies–Roberts hypothesis and points to a critical role for lactate secretion. Plant Physiol. 106, 1179–1185.
Rogowsky, P. M., Close, T. J., Chimera, J. A., Shaw, J. J., and Kado, C. I. (1987). Regulation of the vir genes of Agrobacterium tumefaciens plasmid pTiC58. J. Bacteriol. 169, 5101–5112.
Roitsch, T., Wang, H., Jin, S. G., and Nester, E. W. (1990). Mutational analysis of the VirG protein, a transcriptional activator of Agrobacterium tumefaciens virulence genes. J. Bacteriol. 172, 6054–6060.
Sahi, S. V., Chilton, M. D., and Chilton, W. S. (1990). Corn metabolites affect growth and virulence of Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. U.S.A. 87, 3879–3883. doi: 10.1073/pnas.87.10.3879
Savka, M. A., and Farrand S. K. (1992). Mannityl opine accumulation and exudation by transgenic tobacco. Plant Physiol. 98, 784–789. doi: 10.1104/pp.98.2.784
Shaw, C. H., Ashby, A. M., Brown, A., Royal, C., and Loake, G. J. (1988). virA and virG are the Ti-plasmid functions required for chemotaxis of Agrobacterium tumefaciens towards acetosyringone. Mol. Microbiol. 2, 413–417. doi: 10.1111/j.1365-2958.1988.tb00046.x
Shimoda, N., Toyoda-Yamamoto, A., Aoki, S., and Machida, Y. (1993). Genetic evidence for an interaction between the VirA sensor protein and the ChvE sugar-binding protein of Agrobacterium. J. Biol. Chem. 268, 26552–26558.
Stachel, S. E., Messens, M., Van Montagu, A., and Zambryski, P. (1985). Identification of the signal molecules produced by wounded plant cells that activate T-DNA transfer in Agrobacterium tumefaciens. Nature 318, 624–629. doi: 10.1038/318624a0
Stachel, S. E., and Nester, E. W. (1986). The genetic and transcriptional organization of the vir region of the A6 Ti plasmid of Agrobacterium tumefaciens. EMBO J. 5, 1445–1454.
Stachel, S. E., Nester, E. W., and Zambryski, P. C. (1986). A plant cell factor induces Agrobacterium tumefaciens vir gene expression. Proc. Natl. Acad. Sci. U.S.A. 83, 379–383. doi: 10.1073/pnas.83.2.379
Thomashow, M. F., Hugly S., Buchholz W. G., and Thomashow L. S. (1986). Molecular basis for the auxin-independent phenotype of crown gall tumor tissues. Science 231, 616–618. doi: 10.1126/science.3511528
Toyoda-Yamamoto, A., Shimoda, N., and Machida, Y. (2000). Genetic analysis of the signal-sensing region of the histidine protein kinase VirA of Agrobacterium tumefaciens. Mol. Gen. Genet. 263, 939–947. doi: 10.1007/PL00008694
Turk, S. C., Van Lange, R. P., Regensburg-Tuink, T. J., and Hooykaas, P. J. (1994). Localization of the VirA domain involved in acetosyringone-mediated vir gene induction in Agrobacterium tumefaciens. Plant Mol. Biol. 25, 899–907. doi: 10.1007/BF00028884
Tzfira, T., Vaidya, M., and Citovsky, V. (2001). VIP1, an Arabidopsis protein that interacts with Agrobacterium VirE2, is involved in VirE2 nuclear import and Agrobacterium infectivity. EMBO J. 20, 3596–3607. doi: 10.1093/emboj/20.13.3596
Vergunst, A. C., Schrammeijer, B., Den Dulk-Ras, A., De Vlaam, C. M., Regensburg-Tuink, T. J., and Hooykaas, P. J. (2000). VirB/D4-dependent protein translocation from Agrobacterium into plant cells. Science 290, 979–982. doi: 10.1126/science.290.5493.979
Veselov, D., Langhans, M., Hartung, W., Aloni, R., Feussner, I., Götz, C.,et al. (2003). Development of Agrobacterium tumefaciens C58-induced plant tumors, and impact on host shoots are controlled by a cascade of jasmonic acid, auxin, cytokinin, ethylene, and abscisic acid. Planta 216, 512–522.
Wachter, R., Langhans, M., Aloni, R., Gotz, S., Weilmunster, A., Koops, A.,et al. (2003). Vascularization, high-volume solution flow, and localized roles for enzymes of sucrose metabolism during tumorigenesis by Agrobacterium tumefaciens. Plant Physiol. 133, 1024–1037. doi: 10.1104/pp.103.028142
Wagner, C., Zimmermann, S., Brenner-Weiss, G., Hug, F., Prior, B., Obst, U.,et al. (2007). The quorum-sensing molecule N-3-oxododecanoyl homoserine lactone (3OC12-HSL) enhances the host defence by activating human polymorphonuclear neutrophils (PMN). Anal. Bioanal. Chem. 387, 481–487. doi: 10.1007/s00216-006-0698-695
Walker, T. S., Bais, H. P., Grotewold, E., and Vivanco, J. M. (2003). Root exudation and rhizosphere biology. Plant Physiol. 132, 44–51. doi: 10.1104/pp.102.019661
Wang, F. F., Cui, X. K., Sun, Y., and Dong, C. H. (2013). Ethylene signaling and regulation in plant growth and stress responses. Plant Cell Rep. 32, 1099–1109. doi: 10.1007/s00299-013-1421-6
Wang, K., Herrera-Estrella, A., and Van Montagu, M. (1990). Overexpression of virD1 and virD2 genes in Agrobacterium tumefaciens enhances T-complex formation and plant transformation. J. Bacteriol. 172, 4432–4440.
Wang, L., Helmann, J. D., and Winans, S. C. (1992). The A. tumefaciens transcriptional activator OccR causes a bend at a target promoter, which is partially relaxed by a plant tumor metabolite. Cell 69, 659–667. doi: 10.1016/0092-8674(92)90229-6
Wang, P., Bi, S., Ma, L., and Han, W. (2006). Aluminum tolerance of two wheat cultivars (Brevor and Atlas66) in relation to their rhizosphere pH and organic acids exuded from roots. J. Agric. Food Chem. 54, 10033–10039. doi: 10.1021/jf0611769
White, C. E., and Winans, S. C. (2007). Cell–cell communication in the plant pathogen Agrobacterium tumefaciens. Philos. Trans. R. Soc. Lond. B Biol. Sci. 362, 1135–1148. doi: 10.1098/rstb.2007.2040
Wilms, I., Moller, P., Stock, A. M., Gurski, R., Lai, E. M., and Narberhaus, F. (2012). Hfq influences multiple transport systems and virulence in the plant pathogen Agrobacterium tumefaciens. J. Bacteriol. 194, 5209–5217. doi: 10.1128/JB.00510-12
Wilms, I., Voss, B., Hess, W. R., Leichert, L. I., and Narberhaus, F. (2011). Small RNA-mediated control of the Agrobacterium tumefaciens GABA binding protein. Mol. Microbiol. 80, 492–506. doi: 10.1111/j.1365-2958.2011.07589.x
Winans, S. C. (1990). Transcriptional induction of an Agrobacterium regulatory gene at tandem promoters by plant-released phenolic compounds, phosphate starvation, and acidic growth media. J. Bacteriol. 172, 2433–2438.
Winans, S. C. (1992). Two-way chemical signaling in Agrobacterium–plant interactions. Microbiol. Rev. 56, 12–31.
Winans, S. C., Ebert, P. R., Stachel, S. E., Gordon, M. P., and Nester, E. W. (1986). A gene essential for Agrobacterium virulence is homologous to a family of positive regulatory loci. Proc. Natl. Acad. Sci. U.S.A. 83, 8278–8282. doi: 10.1073/pnas.83.21.8278
Wise, A. A., Fang, F., Lin, Y. H., He, F., Lynn, D. G., and Binns, A. N. (2010). The receiver domain of hybrid histidine kinase VirA: an enhancing factor for vir gene expression in Agrobacterium tumefaciens. J. Bacteriol. 192, 1534–1542. doi: 10.1128/JB.01007-09
Wright, E. L., Deakin, W. J., and Shaw, C. H. (1998). A chemotaxis cluster from Agrobacterium tumefaciens. Gene 220, 83–89. doi: 10.1016/S0378-1119(98)00438-7
Wu, C. F., Lin, J. S., Shaw, G. C., and Lai, E. M. (2012). Acid-induced type VI secretion system is regulated by ExoR-ChvG/ChvI signaling cascade in Agrobacterium tumefaciens. PLoS Pathog. 8:e1002938. doi: 10.1371/journal.ppat.1002938
Xia, J. H., and Roberts, J. (1994). Improved cytoplasmic pH regulation, increased lactate efflux, and reduced cytoplasmic lactate levels are biochemical traits expressed in root tips of whole maize seedlings acclimated to a low-oxygen environment. Plant Physiol. 105, 651–657.
Yanofsky, M. F., Porter, S. G., Young, C., Albright, L. M., Gordon, M. P., and Nester, E. W. (1986). The virD operon of Agrobacterium tumefaciens encodes a site-specific endonuclease. Cell 47, 471–477. doi: 10.1016/0092-8674(86)90604-5
Yuan, Z. C., Edlind, M. P., Liu, P., Saenkham, P., Banta, L. M., Wise, A. A.,et al. (2007). The plant signal salicylic acid shuts down expression of the vir regulon and activates quormone-quenching genes in Agrobacterium. Proc. Natl. Acad. Sci. U.S.A. 104, 11790–11795. doi: 10.1073/pnas.0704866104
Yuan, Z. C., Haudecoeur, E., Faure, D., Kerr, K. F., and Nester, E. W. (2008a). Comparative transcriptome analysis of Agrobacterium tumefaciens in response to plant signal salicylic acid, indole-3-acetic acid and gamma-amino butyric acid reveals signalling cross-talk and Agrobacterium–plant co-evolution. Cell Microbiol. 10, 2339–2354. doi: 10.1111/j.1462-5822.2008.01215.x
Yuan, Z. C., Liu, P., Saenkham, P., Kerr, K., and Nester, E. W. (2008b). Transcriptome profiling and functional analysis of Agrobacterium tumefaciens reveals a general conserved response to acidic conditions (pH 5.5) and a complex acid-mediated signaling involved in Agrobacterium–plant interactions. J. Bacteriol. 190, 494–507. doi: 10.1128/JB.01387-1387
Yuan, Z. C., and Williams, M. (2012). A really useful pathogen, Agrobacterium tumefaciens. Plant Cell 24:tpc.112.tt1012. doi: 10.1105/tpc.112.tt1012
Zhang, J., Boone, L., Kocz, R., Zhang, C., Binns, A. N., and Lynn, D. G. (2000). At the maize/Agrobacterium interface: natural factors limiting host transformation. Chem. Biol. 7, 611–621. doi: 10.1016/S1074-5521(00)00007-7
Zhang, L., Murphy, P. J., Kerr, A., and Tate, M. E. (1993). Agrobacterium conjugation and gene regulation by N-acyl-L-homoserine lactones. Nature 362, 446–448. doi: 10.1038/362446a0
Zhu, J., Oger, P. M., Schrammeijer, B., Hooykaas, P. J., Farrand, S. K., and Winans, S. C. (2000). The bases of crown gall tumorigenesis. J. Bacteriol. 182, 3885–3895. doi: 10.1128/JB.182.14.3885-3895.2000
Zhu, J., and Winans, S. C. (1999). Autoinducer binding by the quorum-sensing regulator TraR increases affinity for target promoters in vitro and decreases TraR turnover rates in whole cells. Proc. Natl. Acad. Sci. U.S.A. 96, 4832–4837. doi: 10.1073/pnas.96.9.4832
Zhu, J., and Winans, S. C. (2001). The quorum-sensing transcriptional regulator TraR requires its cognate signaling ligand for protein folding, protease resistance, and dimerization. Proc. Natl. Acad. Sci. U.S.A. 98, 1507–1512. doi: 10.1073/pnas.98.4.1507
Keywords: Agrobacterium tumefaciens, virulence, signaling mechanism, gene regulation, quorum sensing
Citation: Subramoni S, Nathoo N, Klimov E and Yuan Z-C (2014) Agrobacterium tumefaciens responses to plant-derived signaling molecules. Front. Plant Sci. 5:322. doi: 10.3389/fpls.2014.00322
Received: 24 February 2014; Accepted: 18 June 2014;
Published online: 08 July 2014.
Edited by:
Stanton B. Gelvin, Purdue University, USAReviewed by:
Tina Britta Jordan, Eberhard Karls University Tübingen, GermanyRosalia Deeken, University of Wuerzburg, Germany
Copyright © 2014 Subramoni, Nathoo, Klimov and Yuan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ze-Chun Yuan, Department of Microbiology and Immunology, University of Western Ontario, London, ON N6A 5C1, Canada e-mail:enl1YW4yN0B1d28uY2E=;eXVhbnpAYWdyLmdjLmNh
 Sujatha Subramoni1
Sujatha Subramoni1 Naeem Nathoo
Naeem Nathoo Ze-Chun Yuan
Ze-Chun Yuan